Major Role of Surrounding Environment in Shaping Biofilm Community Composition on Marine Plastic Debris
- 1Universitá Politecnica delle Marche, Dipartimento di Scienze della Vita e dell’Ambiente, Ancona, Italy
- 2Istituto per le Risorse Biologiche e le Biotecnologie Marine, Consiglio Nazionale delle Ricerche (IRBIM-CNR), Ancona, Italy
- 3Department of Biology, Università degli Studi di Napoli “Federico II”, Naples, Italy
- 4Earth-Life Science Institute, Tokyo Institute of Technology, Tokyo, Japan
- 5Department of Marine and Coastal Science, Rutgers University, New Brunswick, NJ, United States
Plastic debris in aquatic environments is colonized by microbes, yet factors influencing biofilm development and composition on plastics remain poorly understood. Here, we explored the microbial assemblages associated with different types of plastic debris collected from two coastal sites in the Mediterranean Sea. All plastic samples were heavily colonized by prokaryotes, with abundances up to 1.9 × 107 cells/cm2. Microbial assemblages on plastics significantly differed between the two geographic areas but not between polymer types, suggesting a major role of the environment as source for the plastisphere composition. Nevertheless, plastic communities differed from those in the surrounding seawater and sediments, indicating a further selection of microbial taxa on the plastic substrates. The presence of potential pathogens on the plastic surface reflected the levels of microbial pollution in the surrounding environment, regardless of the polymer type, and confirmed the role of plastics as carriers for pathogenic microorganisms across the coastal ocean, deserving further investigations.
Introduction
Synthetic thermoplastic polymers, commonly known as plastics, are widespread in our society and constitute one of the most used materials in our everyday life (Barnes et al., 2009). The increasing global production and use of plastics has led to an accumulation of enormous amounts of plastic litter in the world’s oceans, from the coastline to the open sea and from the surface down to the seafloor (Barnes et al., 2009; Schlining et al., 2013), even reaching the most remote open ocean ecosystems (Cózar et al., 2017). Increasing concerns have focused on plastics as a cause of injuries and other health problems for marine organisms (Brouwer et al., 2017) and on their role as a vector of persistent organic pollutants (POPs; Andrady, 2011), invasive species (Barnes and Fraser, 2003; Gregory, 2009), as well as pathogenic species (Keswani et al., 2016).
Whatever their size, in the aquatic environment, plastics represent a physical substrate for microbial colonization and biofilm formation. Several studies have reported that plastics select for specific microbial communities, which differ from the organic particle-attached and free-living communities (Zettler et al., 2013; Oberbeckmann et al., 2015, 2016; Bryant et al., 2016; Dussud et al., 2018; Pinto et al., 2019), and host the so-called “core” plastisphere community (Zettler et al., 2013). However, whether microbial assemblages on marine plastics are selected in relation to the type of polymers, the geographical location, and/or seasonality remains controversial (Carson et al., 2013; Oberbeckmann et al., 2014; Amaral-Zettler et al., 2015; De Tender et al., 2015). Additionally, recent works also suggest that (micro)plastics represent novel means of transport for pathogenic microorganisms, across the marine environment, consequently acting as a possible medium for the spread of these microbes and the consequent diffusion of infectious diseases (Zettler et al., 2013; Keswani et al., 2016; Kirstein et al., 2016). Moreover, due to their known ability to adapt quickly to new environmental conditions (Wiedenbeck and Cohan, 2011), it has been hypothesized that some marine microorganisms are able to use the plastic debris as a new substrate not only for the formation of biofilm but also to further use plastic polymers as a carbon source (Balasubramanian et al., 2010; Bryant et al., 2016).
The Mediterranean basin is one of the largest plastic debris accumulation areas in the world (Ruiz-Orejón et al., 2016; Suaria et al., 2018), with values similar to those reported from the inner accumulation zones of all main oceanic gyres (Cózar et al., 2015). Within the Mediterranean Sea, the Adriatic Sea has shown a significantly lower abundance of plastic fragments in respect to the Western Mediterranean basin, likely reflecting the distinctive hydrological features of the Adriatic basin (Suaria et al., 2018). On the Tyrrhenian Sea, contrasting results have been obtained in the accumulation of plastic debris, due to seasonal and sea currents variations (Suaria and Aliani, 2014; Deudero and Alomar, 2015; Mansui et al., 2015).
In this study, we selected two sites located on the opposite coasts of Italy, Naples (Tyrrhenian Sea) and Ancona (Adriatic Sea), on the basis of their geographical distance and different levels of human impact, and we described the morphology, abundance and diversity of the prokaryotic communities living on plastic debris. Plastic samples were analyzed in order to verify whether plastics selected for a specific microbial community, and to understand to what extent the surrounding environment acts as the source for bacterial colonization. Finally, with the aim of testing the role of debris as vector of pathogens, we looked for the presence, diversity and distribution of fecal microbes and potential human pathogens within our dataset.
Materials and Methods
Study Area
Sampling activities were carried out at two sites located on the Western (in the city of Naples, 40°50′ N 14°13′ E) and Eastern (in the city of Ancona, 43°36′ N 13°27′ E) coast of Italy. Both study areas were selected as representative of anthropogenically impacted coastal sites, although at different levels. In fact, the site at Naples is an urban beach very close to the city center, characterized by typically high frequencies of tourists and the presence of potential pollution sources (sewage output pipes) in proximity of the sampled area. The site in Ancona was a recreational beach located farther from the city, in an area characterized by beach tourism during the summer season. From each site, we collected plastic debris, seawater and sediment samples. Sampling activities were carried out on January 2018 in Naples (“NAP-”, “NAPWAT-”, “NAPSED-” for plastic items, water and sediment samples, respectively) and in Ancona (“AN-”, “ANWAT-”, “ANSED-” for plastic items, seawater and sediment samples, respectively). Plastic items with different sizes (ranging from few cm2 to about 10 cm2), colors and textures (Figure 1), were randomly collected on the shore in the intertidal area (thus in direct contact with both sediment and seawater, and subjected to sun exposure) using sterilized tweezers. One liter of surface seawater and about 50 g of surface sand from the shoreline were also collected close to the plastic fragments’ collection area by using sterile procedures, and put within sterile containers until return to laboratory. After collection, plastic items were stored at 4°C for transport to the nearby laboratory. A total of seven plastic items were collected in Ancona, and 12 in Naples. Once in the laboratory, before processing, plastic items were washed for a few seconds with sterile seawater in order to remove the loosely-attached sand and microbes from the surface. Each plastic item was then cut into different scraps of 2 × 2 cm ca. surface area; each of these fragments was used for a different analysis, as described below. All procedures in the laboratory were performed under sterile conditions (i.e., under laminar flow hood and using sterile tools) in order to avoid further contamination of the samples during processing.
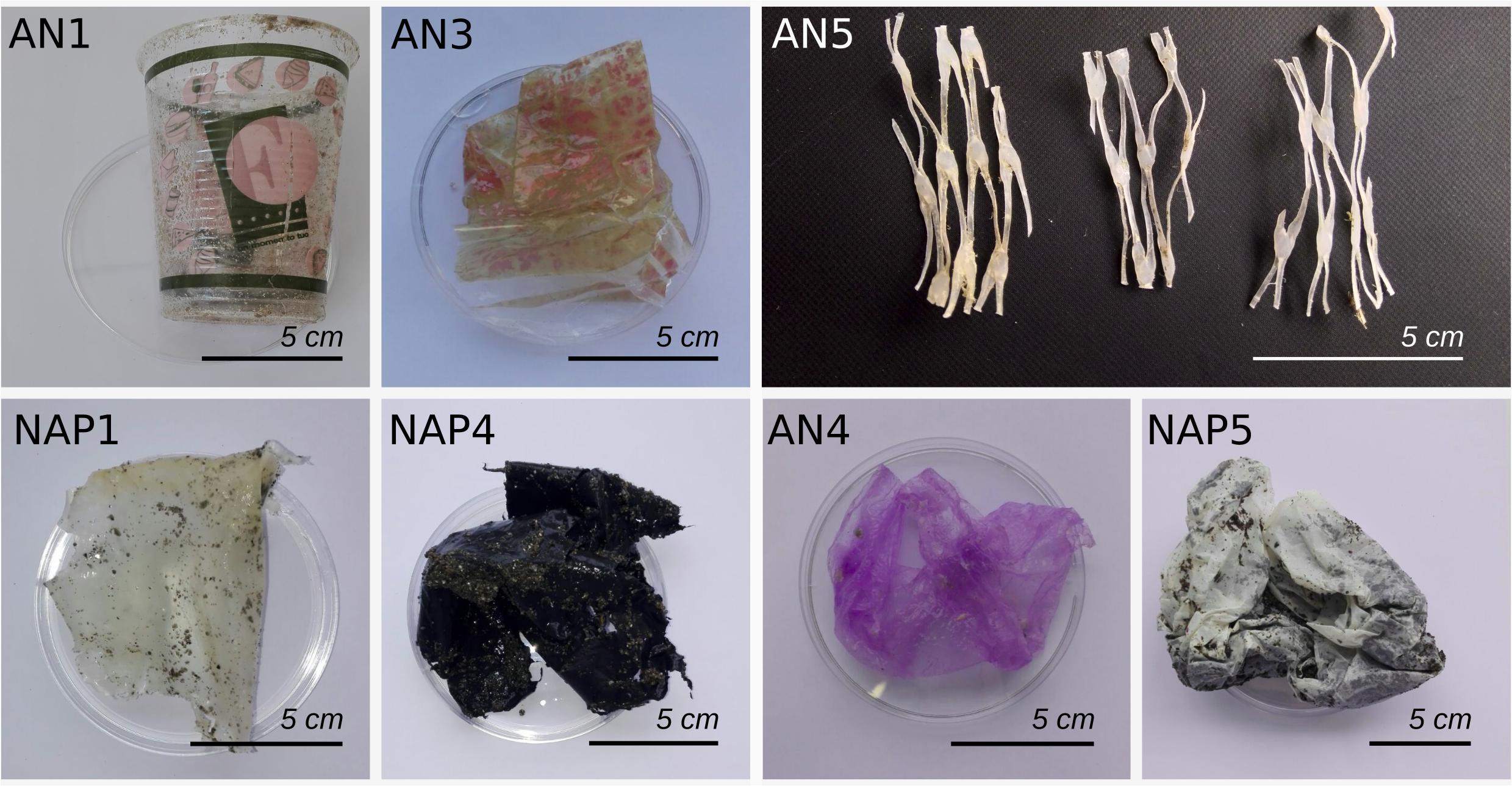

Figure 1. Images of a selected subset of plastic items collected in this study from the Naples (NAP) and Ancona (AN) sites. As shown, different types of plastic items were collected, according to color, texture and size. Scale is reported.
μFT-IR Spectroscopy
Each plastic item was analyzed by using a μFT-IR microscope (Spotlight i200, PerkinElmer) coupled to a spectrometer (Spectrum Two, PerkinElmer) in order to provide knowledge on their chemical composition. All the measures were made using the μATR mode; following back-ground scans, 32 scans, with a resolution of 4 cm–1, were performed for each item. The software Spectrum 10 was used for the output spectra and the identification of polymers was performed by comparison with libraries of standard spectra. Polymers matching with reference spectra for more than 70% were validated (Avio et al., 2015).
Scanning Electron Microscope
A subset of plastic fragments was selected and examined for biofilm visualization by Scanning Electron Microscope (SEM). Preparation of plastic fragment followed the protocols described by Zettler et al. (2013) and Pinto et al. (2019) with slight modifications. Briefly, plastic fragments were fixed with 4% formaldehyde for about 24 h. The samples were then washed three times using sterile seawater for 15 min to allow the removal of all the formaldehyde, and dehydrated in a graded ethanol series of 30, 50, 70, and 90% and absolute ethanol for 30 min each. Critical point drying was performed on a Leica EMCPD300. Plastic fragments were then platinum-coated using a Polaron SC7640 Sputter Coater (Thermo VG Scientific). Images were obtained with a JSM-6700F scanning electron microscope (Jeol) at an accelerating voltage of 10.00–20.00 kV.
Prokaryotic Abundance
Total prokaryotic abundance (TPA) was measured on plastic and sediment samples by epifluorescence microscopy, using acridine orange as stain as described by Luna et al. (2002). Briefly, three fragments (i.e., replicates) were used for TPA, each one placed in a sterile tube and fixed with 10 mL of pre-filtered, 2% formalin solution, previously buffered with Na2B4O7 × 10H2O until a complete coverage of samples by the solution. Fixed samples were preserved overnight at 4°C. Each sample was then sonicated three times for 1 min to detach cells from plastic fragments and sediments. Subsequently, sediment subsamples were diluted 500 times in filtered seawater, while 100 μL of the suspension of the plastic samples was diluted in 3 mL of filtered seawater. Each sample was then stained with acridine orange (final concentration, 0.025%), and the solution was filtered on a black Nucleopore polycarbonate 0.2 μm pore-size filter by vacuum pump. Each filter was analyzed using epifluorescence microscopy. For each slide, 10 randomly selected microscope fields were observed, and the microbial cells were counted. For each replicate, cell abundances were calculated as the average number of prokaryotic cells per field. The same method was used for sediment (three replicates of 1 g were diluted 100 times in filtered water).
Biodiversity and Community Composition by High-Throughput Sequencing of Microbial 16S rDNA
Microbial DNA was extracted from each plastic fragment, from 1 L of seawater, filtered on 0.22 μm membrane filter by vacuum pump, and from 1 g of sediment sample, using the PowerSoil DNA Isolation Kit (Mo Bio Laboratories, Inc., CA, United States), according to the manufacturer’s instructions with two additional vortexing steps for 2 min, followed by incubation at 70°C for 5 min and adding one more washing step with Solution C5 as an additional removal step for contaminants, as described in Quero et al. (2017), to increase the DNA yield and quality. DNA concentration was determined with Qubit fluorometer (Thermo Fisher Scientific), and the DNA was stored at −80°C until the sequencing.
For HTS library preparation, the Illumina Nextera protocol was used to obtain amplicon libraries of the V3–V4 regions of the 16S rRNA gene, which were sequenced on the Illumina MiSeq platform by LGC Genomics GmbH (Berlin) using V3 chemistry and 2 × 300 bp sequencing. The 341F (5′-CCTACGGGNGGCWGCAG-3′) and 785R (5′-GACTACHVGGGTATCTAATCC-3′) universal bacterial primer pair (Eiler et al., 2012) was used. Raw sequences were pre-cleaned by clipping synthetic primers and sequencing adapters.
All sequences were imported in R and analyzed with the DADA2 package (Callahan et al., 2016). Following the package instructions, sequences’ quality was inspected by checking the quality plots and by subsequently trimming the last 30 and 50 bp for forward and reverse reads, respectively, and by allowing a max estimated error (“maxEE” option) higher than 2 and 5 per 100 bp for forward and reverse reads, respectively. After this step, samples were pooled to estimate sequencing error rates until the convergence of the parametric error model was achieved. The amplicon sequence-variant (oligotype) inference was performed on the dereplicated sequences after pooling all samples together to reduce possible biases due to low sampling depths. Chimeric sequences were then identified, paired-ends reads merged, and prokaryotic taxonomy was assigned using a native implementation of the naive Bayesian classifier method against the silva database1 (v128). The Amplicon Sequence Variants (ASV) table was then rarefied to an even number of sequences per sample to ensure an equal sampling depth for all samples (n = 3,682). ASVs were defined as clusters sharing 100% sequence identity. ASV table is available as Supplementary Data Sheet 1. The sequences have been submitted to the SRA – Sequence Read Archive (BioProject accession number PRJNA558771, SAMN12497894 to SAMN12497918).
Analyses of Microbes of Fecal Origin and Potential Pathogens Associated to Plastics
Within each sample, we specifically looked for ASVs identified as belonging to bacterial taxa of potential fecal origin. In particular, we searched for “traditional” fecal indicators (i.e., the Enterobacteriaceae family including the genera Escherichia, and Enterococcus, used worldwide to assess fecal pollution in aquatic environments) and for “alternative” fecal indicators (Newton et al., 2013; Fisher et al., 2015; Luna et al., 2016; Feng et al., 2018), belonging to five “feces-associated” bacterial families (Bacteroidaceae, Porphyromonadaceae, Clostridiaceae, Lachnospiraceae, and Ruminococcaceae) and three “sewage-associated” bacterial genera (Acinetobacter, Arcobacter, and Trichococcus), used as potential signatures of fecal (human and non-human) and sewage contamination, respectively. We also searched for other widely recognized potential human pathogens, as reported in Supplementary Table S1.
Data Analyses
The ASV table was imported in RStudio (RStudio Team, 2016) and analyzed using the R package vegan (Oksanen et al., 2017). The sample NAP2 (474 reads) was an order of magnitude smaller in library size, and was excluded from statistical analyses. ASV richness (i.e., the number of ASVs) and Shannon diversity indexes were calculated for the analysis of alpha-diversity (estimateR and diversity commands, respectively). Non-metric multidimensional scaling (nMDS) and cluster analysis were performed using a Bray–Curtis dissimilarity matrix and average linkage approach, respectively, after Hellinger’s transformation of data. To test the presence of statistical differences between and among samples for TPA, major phyla and classes and pathogenic signature, we used Kruskal–Wallis test in R, using dplyr and FSA libraries. The presence of statistical differences between microbial communities according to site and/or polymer was calculated by using PERMANOVA through the adonis function (vegan package) in R. Venn diagrams were plotted in order to show the number of shared and unique ASVs between sampling sites and between plastic polymers, sediment and water. For this analysis, any ASV found in at least one polyethylene (PE) and one polypropylene (PP) sample was considered as “shared”. Analyses of the core microbiome were performed on QIIME v1.8.0 (Caporaso et al., 2010) using the compute_core_microbiome.py script on the normalized ASV table. Finally, shared and unique ASVs were also identified using a network-based analysis (Giovannelli et al., 2016) based on the make_otu_network.py script in QIIME v 1.9.0 (Caporaso et al., 2010) to display and analyse how ASVs were partitioned between samples. Network results were visualized and analyzed with Gephi (Bastian et al., 2009).
Results
μFT-IR Spectroscopy
Plastic samples were identified as PE, PP, and polystyrene (PS) through μFT-IR analysis. In the Ancona site, out of the seven total plastic samples collected, three were identified as PE and four as PP. In the Naples site, out of 12 total samples, 7 were identified as PE, 4 as PP and only 1 sample as PS (Supplementary Figure S1A).
SEM-Based Visualization of Microbial Biofilm
SEM revealed that samples were prevalently colonized by prokaryotic-sized assemblages in both PE and PP (Supplementary Figure S2A,B,D–F). In some cases, eukaryotic microorganisms, such as pennate diatoms (often fragmented) were observed (Supplementary Figure S2C). Rod-shaped cells were more prevalent than coccoid-shaped microorganisms and often occurred in patches, and including dividing cells, a finding that suggested an active microbial growth. Microbial “hot-spots” on plastics mainly occurred when cracks and pits were observed on the plastic surface.
Prokaryotic Abundance on Plastic Debris and Surrounding Environment
Epifluorescence microscopy counts revealed that all the samples were highly colonized by prokaryotes (Figure 2). TPA on plastics ranged from 0.58 to 10.35 × 108 cells/g (when considering the weight of the plastic item), and from 0.19 to 18.8 × 106 cells/cm2 (when considering the plastic item surface), with an average abundance of 6.22 × 108 ± 9.33 cells/g and 3.26 × 106 ± 5.64 cells/cm2 (weight and surface, respectively). Abundances in the sediment samples, which showed the average value of 1.30 × 107 ± 1.06 cell/g, were one order of magnitude lower than those observed in plastic samples. We found no significant difference between plastic samples considering the values of the two different sites (Kruskal–Wallis, p = 0.28). Conversely, we found a statistical difference among plastic polymers (PE, PP, and PS) (Kruskal–Wallis, p = 0.016), with higher TPA observed in NAP6 (1.88 × 107 ± 3.83 × 106 cells/cm2) and lower TPA in NAP2 (1.95 × 105 ± 8.68 × 104 cells/cm2), although this difference was mainly driven by the peak in TPA observed in NAP6 (PS).
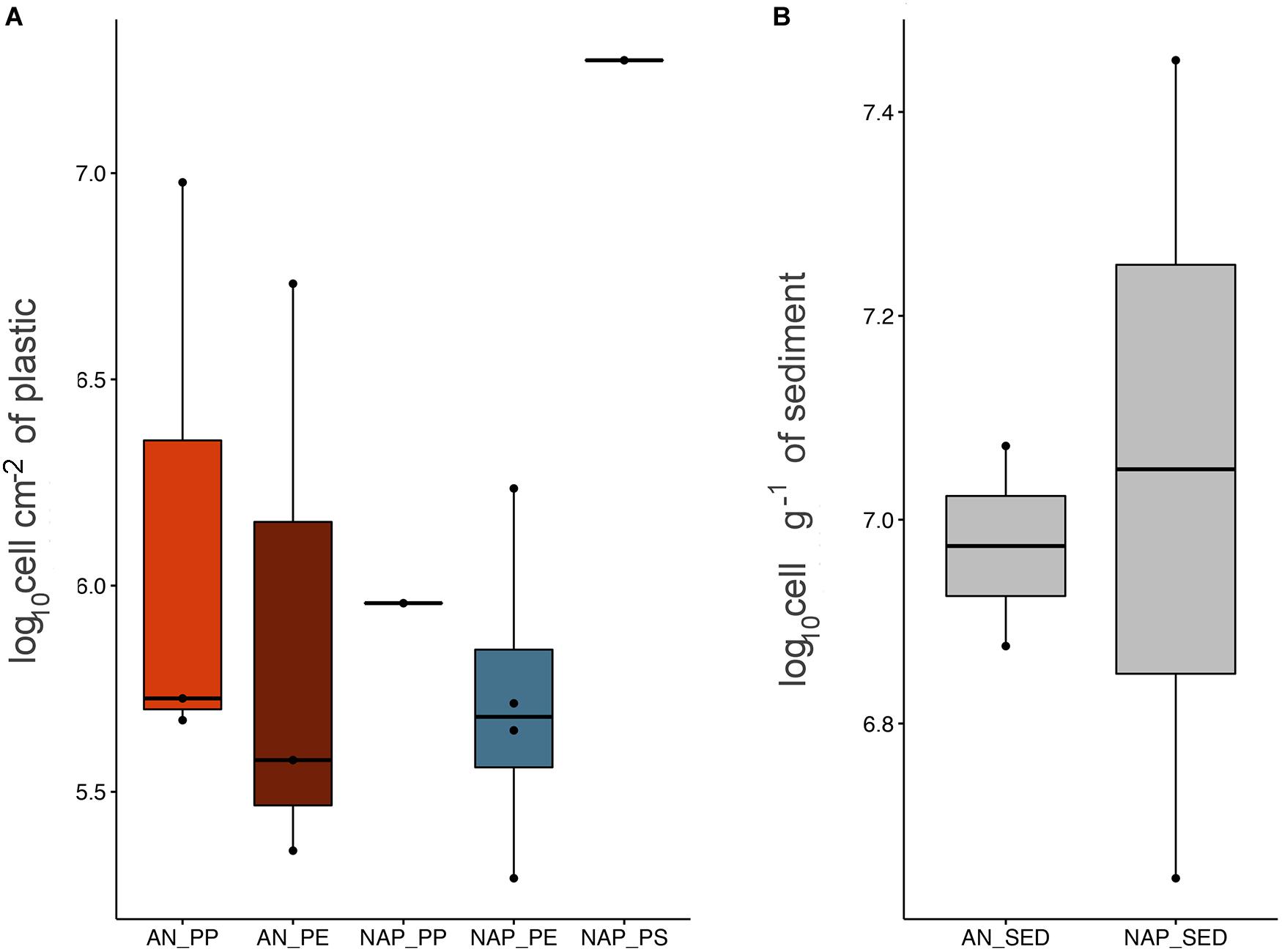
Figure 2. Boxplot showing TPA on plastic items, expressed as cells by unit of surface (A), and on sediment samples, expressed as cell per gram (B). Boxplots were calculated considering an average value derived from triplicate determinations carried out for each sample.
Prokaryotic Diversity and Community Composition
Across the whole set of samples (plastic, seawater, and sediment), we overall observed differences in the number of reads per sample (Supplementary Figure S3), ranging from 474 (NAP2) to 278,705 reads (AN7) for an average number of 62,002. Diversity analyses indicated the presence of highly diversified microbial communities present on both plastic and environmental samples.
The cumulative number of ASVs throughout the entire dataset was 8,617 ASVs, which decreased to 6,283 ASVs after subsampling and removal of NAP2 sample. Values of alpha-diversity indices of communities associated to the plastic samples (PE and PP) from Ancona (avg. ASV richness 200 ± 140, and average Shannon index 3.64 ± 0.91) were significantly lower than those from Naples (avg. 487 ± 177 for ASV richness, and 4.95 ± 0.66 for Shannon index) (Kruskal–Wallis, p < 0.05). No significant difference in prokaryotic alpha-diversity associated to the different polymers was observed. Seawater samples showed average values of alpha-diversity similar to plastics, respectively: 189 ± 63 (ASV richness) and 3.5 ± 0.05 (Shannon index) for Ancona; 468 ± 128 (ASV richness) and 4.8 ± 0.42 (Shannon index) for Naples. Consistent with the SEM images (which showed presence of microalgae, Supplementary Figure S2), a total of 177 ASVs, present in most of the plastic samples except for NAP9 and NAP11, were identified as “chloroplast,” with a relative abundance up to 13% (NAP3) (data not shown). For the subsequent analyses of prokaryotic community composition and beta-diversity, sequences identified as Eukaryote and chloroplast were removed from the ASV table.
Across the whole set of samples, the most abundant prokaryotic phylum was represented by Proteobacteria (avg., 64.83 ± 15.61%), mainly constituted by Gammaproteobacteria (37.81 ± 26.74%) and Alphaproteobacteria (22.70 ± 17.74%) (Figure 3). Other dominant phyla included Bacteroidetes (avg., 21.21 ± 13.22%), Actinobacteria (avg., 6.20 ± 7.55%), Cyanobacteria (avg., 5.5 ± 6.57%), Planctomycetes (avg., 2.00 ± 2.41%), and Firmicutes (avg., 1.50 ± 3.41%).
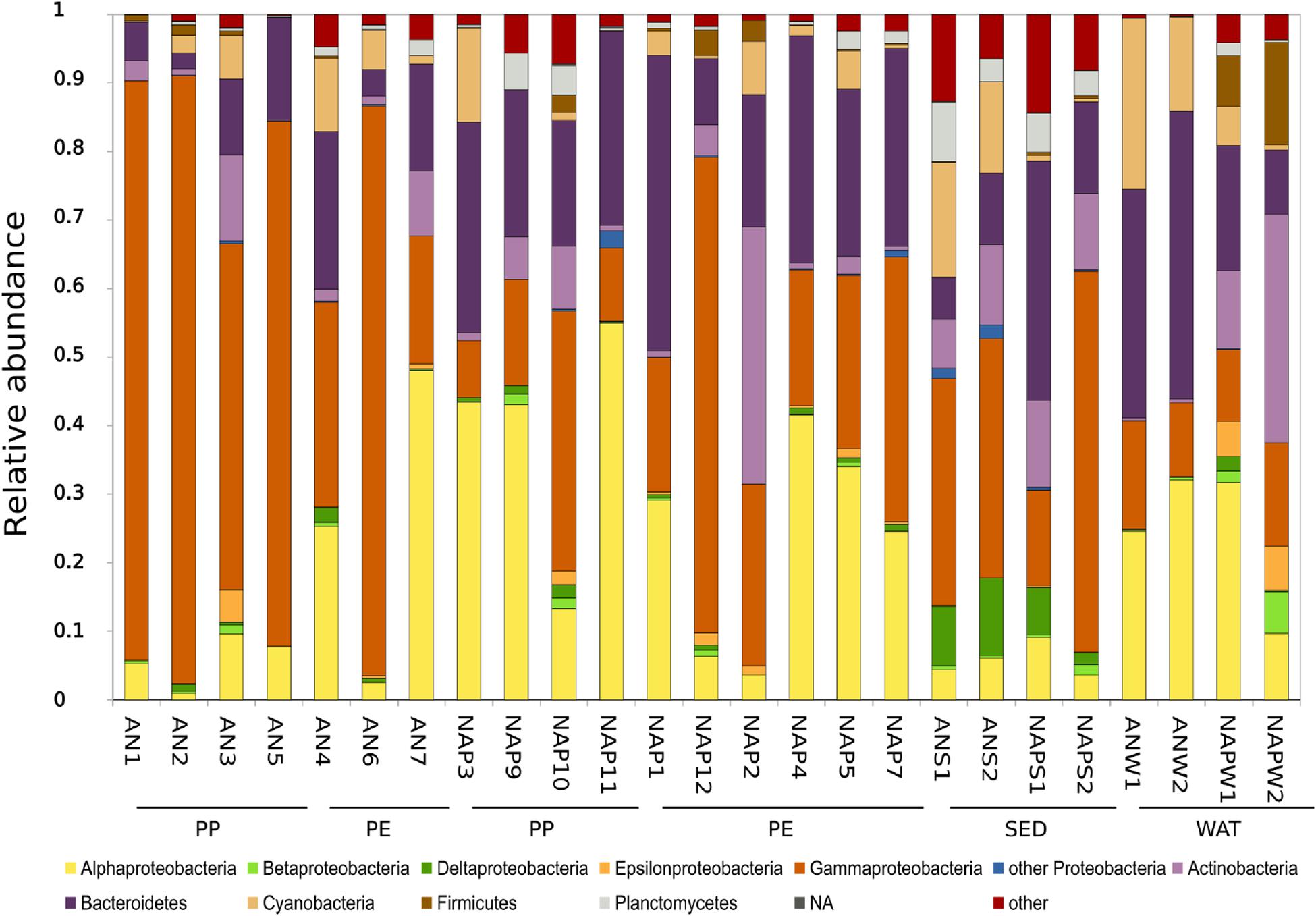
Figure 3. Prokaryotic community composition (as%) at the Phylum level (Class level for Proteobacteria only). Groups with an average relative abundance across all samples <1% were aggregated into the group reported as “Others”. “Unassigned” includes all those reads that did not match any known taxonomy. PP, polypropylene; PE, polyethylene; SED, sediment; WAT, seawater; NA, not assigned.
At the Class level, apart from the previously described proteobacterial classes, the most abundant classes were represented by Sphingobacteria (2.04%) and Flavobacteria (18.01%) for Bacteroidetes; Acidimicrobia (2.88%) and Actinobacteria (ph. Actinobacteria) (3.10%), Bacilli (0.35%) and Clostridia (0.98%) within the Firmicutes phylum. When considering only the plastic samples, we found a significantly higher abundance of Gammaproteobacteria in Ancona (Kruskal–Wallis, p < 0.05; avg., 63.93 ± 28.49%) than in Naples (avg., 27.89 ± 19.13%); conversely, plastics from Naples showed significantly higher abundances of Alphaproteobacteria (Kruskal–Wallis p < 0.05; avg., 33.64 ± 16.73%) and Bacteroidetes (p < 0.05; avg., 26.89 ± 9.76%). The most abundant ASV throughout the entire dataset was identified as belonging to the genus Pseudoalteromonas (phylum Proteobacteria, avg. 4.3 ± 10.8%).
When considering plastic samples only, we found that the same Pseudoalteromonas ASV was the most abundant in Ancona plastic samples (0–37%, with an average of 15 ± 17%). In Naples, the most abundant ASV on plastics was identified as belonging to Dokdonia genus (avg. 4.76 ± 7.1%). Among the other most abundant ASVs on plastic items, we also found other Pseudoalteromonas (2.9%), as well as other gammaproteobacterial genera such as Cocleimonas (2.2%) and Alteromonas (2.01%). ASVs identified as Dokdonia, Alteromonas, and Sulfitobacter, among the most abundant, were found only in plastics samples.
Venn diagrams (Supplementary Figure S4) revealed that only a small fraction (6.6%) of ASVs was shared (i.e., by at least one sample in each category) between Ancona and Naples (n = 401), and mainly included ASVs belonging to the Pseudoalteromonas genus and Rhodobacteraceae family. PE and PP samples shared 630 ASVs, mainly belonging to the Pseudoalteromonas genus. Conversely, a high number of ASVs was exclusive for each of the considered groupings. Only 33 ASVs (0.5%) were shared between the four substrates, and only 89 ASVs between sediment and seawater, suggesting that these communities were consistently distinct. Results from core microbiome analyses of the ASVs shared by all plastic samples at both sites showed that none of the ASVs were present in all of our plastic items, and that one ASV identified as Sulfitobacter spp. was present in 85% of the samples. Only four ASVs, namely ASV5025 (Propionibacterium spp.), ASV6676 (Psychrobacter spp.), ASV7362 (Sulfitobacter spp.), and ASV7398 (Loktanella spp.), were found to be shared by at least 75% of PE samples. We then looked for the presence of core ASVs in each of the polymer types, and we found that only one single ASV was shared by all PE samples (ASV7362, Sulfitobacter spp.), and five in at least 85% of the PE samples, namely ASV5025 (Propionibacterium spp.), ASV6676 (Psychrobacter spp.), ASV7362 (Sulfitobacter spp.), ASV3668 (Muriicola spp.) and ASV7398 (Loktanella spp.). For PP samples, none of the ASVs were shared by all PP items, while we found that shared ASVs (n = 2) were only detectable when considering at least 75% of samples, namely ASV7362 (Sulfitobacter spp.) and ASV7398 (Loktanella spp.). ASVs with similarities to the Loktanella genus appear to be present in the majority of the plastic samples, despite being represented by different ASVs.
We further analyzed the differences in community composition among samples taking into account the whole prokaryotic communities. The non-metric Multi-Dimensional Scaling (nMDS) ordination showed that prokaryotic communities grouped separately according to the sampling site along dimension 1 (Figure 4) and, within site, were segregated according to the sample type along dimension 2 (i.e., plastic communities separated from environmental samples). Overall, we found that microbial communities on plastics were significantly (adonis, p ≤ 0.001) different from those in the surrounding seawater and sediments, and that the differences in microbial community composition were mainly and significantly (adonis, p ≤ 0.001) related to the site but not to the type of plastic polymers. The subsequent cluster analysis (Supplementary Figure S5) confirmed that community composition clustered according to the sampling site but not to the polymer type, and that communities from plastic samples separated from those in other environments. This analysis also highlighted that sample AN7 behaved as a peculiar sample, since it grouped together with the NAP samples.
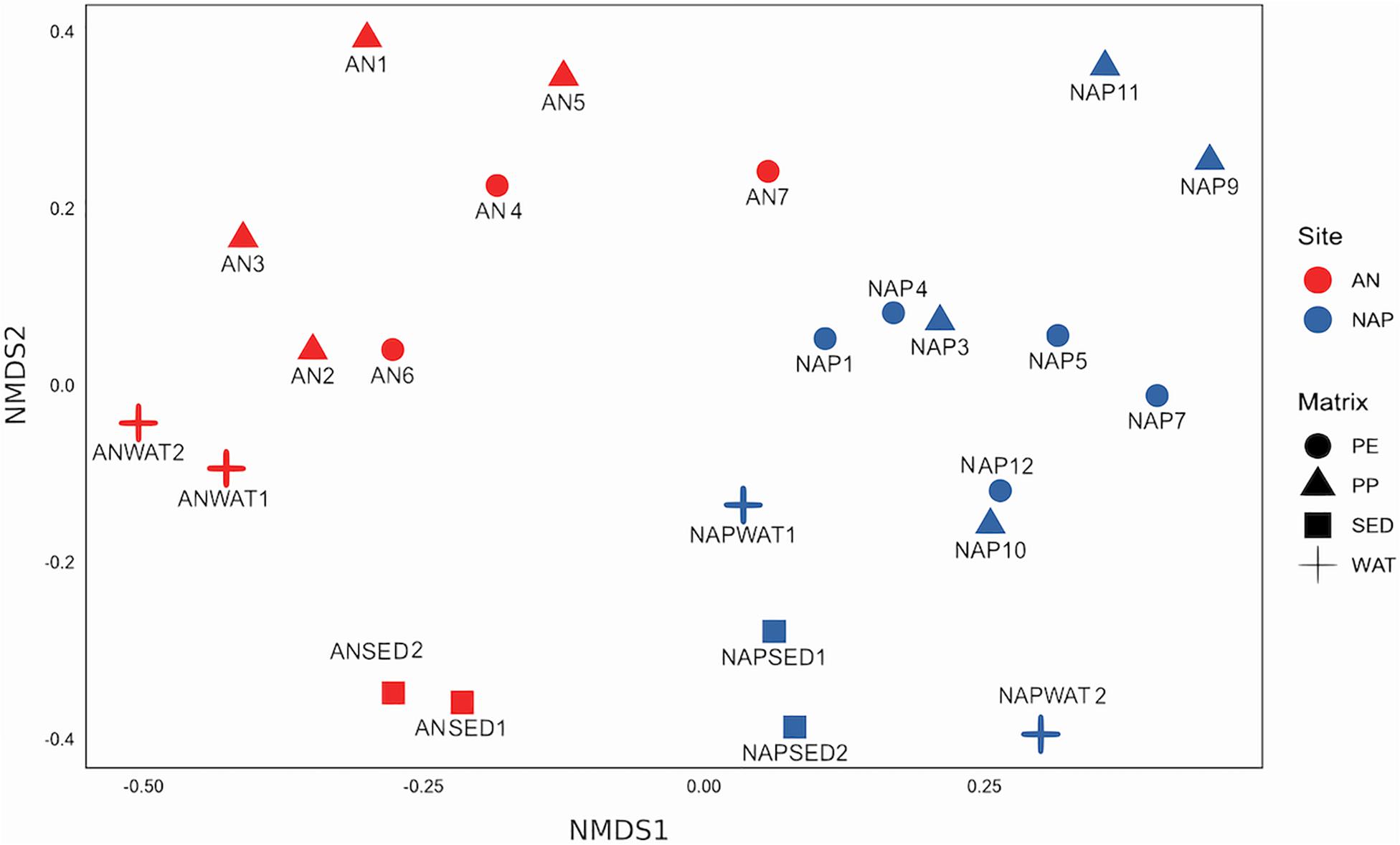
Figure 4. Non-metric MDS (nMDS) ordination of community composition (based on 16S rRNA gene analysis and values of Bray–Curtis dissimilarity) in the different sites and samples investigated. Shape represents different substrate, and color represents the sites. Stress value = 0.1860.
Finally, in order to visually emphasize similarities and differences between samples in large and highly complex datasets, we used a network-based analysis to display and analyse how ASVs were partitioned between samples. Network analysis (Figure 5) strongly corroborated the main results shown previously by the other approaches, and by the description of the shared/unique fractions within communities. In fact, we found here that a partitioning of samples according to the site was evident, and that both Naples and Ancona plastic samples harbored a high number of exclusive ASVs, a pattern also visible, by means of network analysis, at the single sample level.
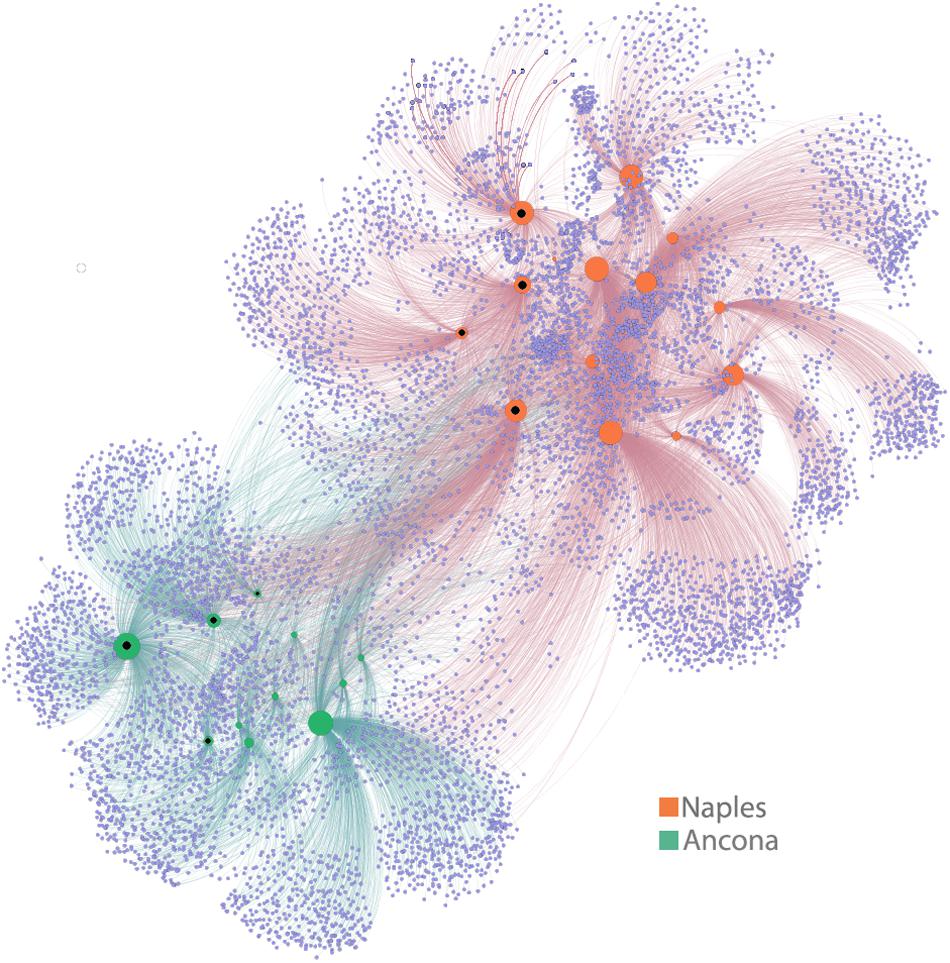

Figure 5. Network analysis. The visual output of this analysis is a clustering of samples according to their shared ASVs – i.e., samples that share more ASVs cluster closer together. The degree to which samples cluster is based on the number of ASVs shared between samples (when ASVs are found in more than one sample) and this is weighted according to the number of sequences within an ASV. In the network diagram, there are two kinds of “nodes” represented, ASV-nodes (light violet nodes) and sample-nodes (green = AN, orange = NAP samples). If an ASV is found within a sample, the two nodes (sample node and ASV node) are connected with a line (an “edge”). Inner black dots in sample nodes indicate environmental (seawater or sediment) samples. The size of the sample nodes is proportional to their degree of connection, i.e., highly connected sample nodes are bigger than those with fewer connections.
Microbial Indicators of Fecal Pollution and Potential Pathogens in Plastic and Environmental Samples
The analysis of fecal bacteria, as well as other potential pathogens revealed that, in both sites, several sequences identified as belonging to fecal and other potential pathogenic bacteria could be found (Figure 6).
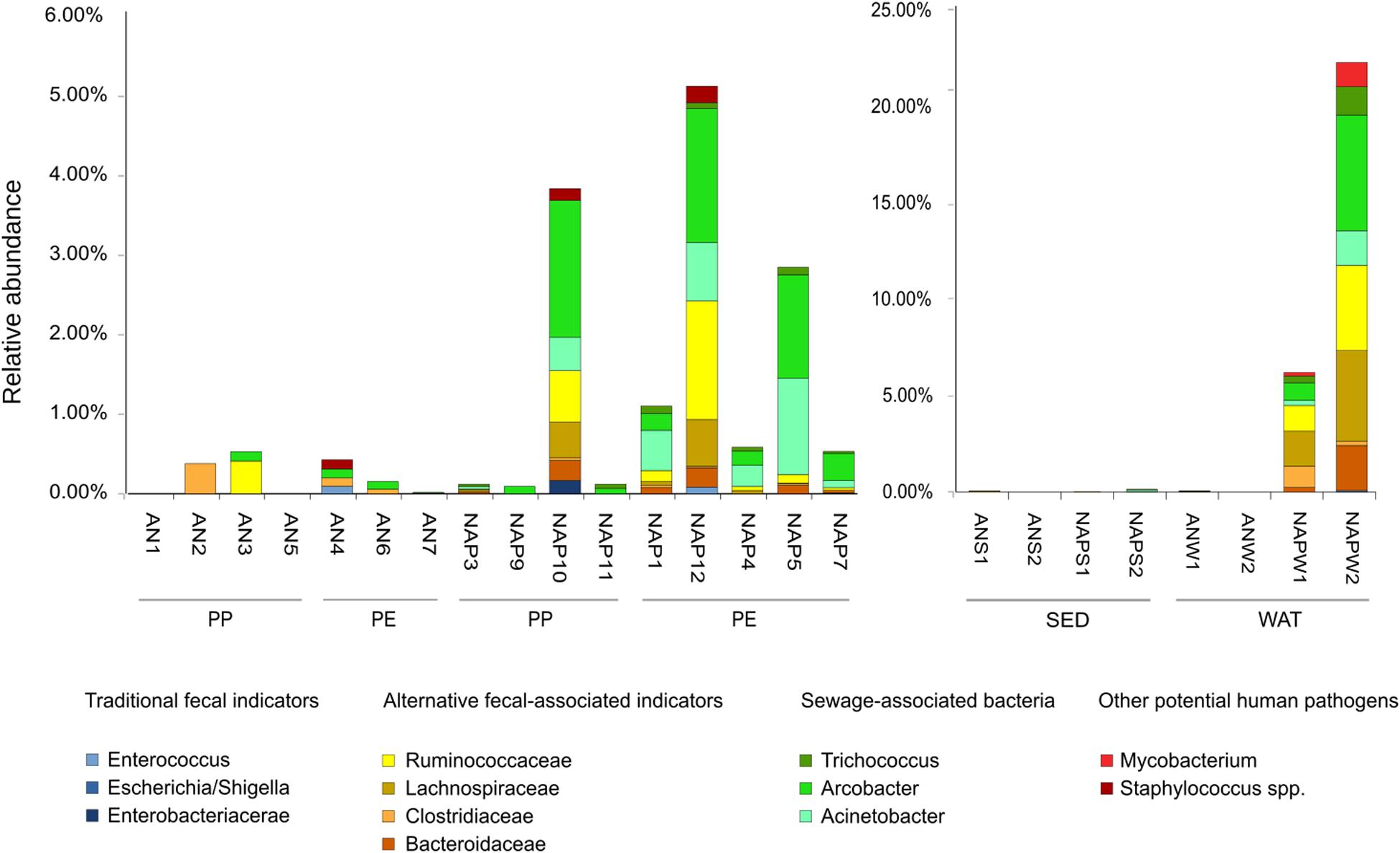
Figure 6. Bar plots of relative abundance (based on the number of reads of 16S rDNA sequences) of sequences belonging to the traditional, and alternative (feces-associated and sewage-associated) fecal indicator bacteria, in the plastic (left panel) and seawater and sediment samples (right panel). Y-axis shows the relative abundance (please, note the difference in scale between the two panels). Taxa listed in the “Materials and Methods” section as indicator bacteria or other potential pathogens not detected in our dataset are not listed in the figure legend.
The microbial fecal signature differed significantly between sites, with higher percentage observed in Naples rather than in Ancona (Kruskal–Wallis, p < 0.01). Even when considering only plastic samples, a significant difference between the two sites was found (Kruskal–Wallis, p < 0.05). Within each site, no significant difference was observed between PE and PP (Kruskal–Wallis, n.s.); however, it must be pointed out that, in Naples, microbial pollutants were often more abundant in PE samples than PP (Figure 6). Similarly to the pattern observed for plastic samples, the fecal signature in seawater was significantly higher in Naples than Ancona, with high relative abundance of microbial pollutants observed in Naples seawater (accounting for up to 14.63 and 49.81% in NAPWAT1 e NAPWAT2, respectively). In Naples, most of the fecal pollution, both on plastic and seawater samples, was attributable to sewage-associated indicators, such as Arcobacter and Acinetobacter, and by feces-associated indicators (Lachnospiraceae and Ruminococcaceae). Conversely, very few sequences belonging to the traditional indicators of fecal pollution were found, mainly constituted by the Enterococcus genus, and small fractions of other potentially pathogenic microbes, such as Staphylococcus spp. and Mycobacterium spp., were found. In plastic samples from this site, the relative abundance of microbial pollutants attached to plastic ranged from a minimum of 0.09% (NAP9) up to 5.07% (NAP12). In Ancona, the level of fecal contamination was considerably lower than in Naples (Figure 6). Within this site, we mainly detected Clostridiaceae and Ruminococcaceae, and the cumulative contribution of all fecal indicators sequences accounted from undetectable (0.0%, i.e., AN1, AN5) up to 0.48% (AN3) of the whole bacterial assemblage on plastic samples. Conversely to what we found in Naples, seawater and sediment samples in Ancona were characterized by almost complete absence of microbial fecal pollutants.
Discussion
In agreement with patterns previously described at the global and Mediterranean levels (Zettler et al., 2013; Oberbeckmann et al., 2014; Bryant et al., 2016; Suaria et al., 2016; Plastics Europe, 2018; Vianello et al., 2018), the majority of the plastic items collected in this study were made of PE and PP polymers. Among the plastic types specifically retrieved in the Adriatic Sea, we found a high number of fragments of mussel nets, a type of debris that is increasingly reported in this basin (Strafella et al., 2015; Pasquini et al., 2016).
SEM analyses indicated that microbial life on plastic samples was mainly prokaryotic and that, consistently with the molecular analyses, only a few eukaryotic organisms (i.e., diatoms) were present. This result differs from previous studies in other oceanic regions, in which diatoms and other eukaryotic organisms were found to be more abundant on plastic debris (Carson et al., 2013; Zettler et al., 2013). The density of prokaryotic cells on our plastic samples is in line with the data reported in a recent study carried out on plastic debris in the Mediterranean Sea (Dussud et al., 2018) and supports the general idea that, for microbes, plastics represents a suitable surface to attach to. However, no significant statistical relationships were found between the type of polymer and prokaryotic abundance, suggesting that the type of plastic polymer is not a strong determinant that regulates microbial abundance. It must be highlighted that the only fragment of polystyrene (PS) in our dataset was the one hosting the highest number of prokaryotes compared to the other plastic items, a finding previously reported by Carson et al. (2013). Despite being not supported by statistical analyses, this result corroborates the hypothesis that PS in the aquatic environment may be easily colonized by higher numbers of microbes, a consideration that deserves further investigations, due to its potential ecological consequences.
Analyses of the diversity and community composition highlighted a major role of the surrounding environment (e.g., seawater, sediment as a likely driving force that shapes the types of microbes found associated with plastics, providing further support to the indications by Carson et al. (2013) in the Eastern North Pacific Gyre, De Tender et al. (2015) in the North Sea, and other authors worldwide (Zettler et al., 2013; Oberbeckmann et al., 2014; Pinto et al., 2019). This evidence was accompanied by the lack of a significant difference in the plastisphere community composition between polymers, acknowledging previous observations by Bryant et al. (2016) and Pinto et al. (2019). Network analysis also confirmed that the sampling site was more important than the type of plastic polymer in shaping the plastisphere, as indicated by the high heterogeneity among and within samples. As pointed out by De Tender et al. (2015), the large variation of prokaryotic community composition on plastics, along with differences from the sediment and seawater counterparts, indicates that plastics may represent a distinct environmental niche. In fact, the environment likely acts as a source of primary (early) colonizers, which depends on the site of origin; however, different stages or dynamics of biofilm formation concur later in shaping the microbial community on plastic surfaces (Lobelle and Cunliffe, 2011; Oberbeckmann et al., 2015; Pinto et al., 2019). The dominance of both typical primary (Gamma- and Alphaproteobacteria) and secondary colonizers (e.g., Bacteroidetes; Dang and Lovell, 2000; Lee et al., 2008; Oberbeckmann et al., 2015), as also reported in biofilm from other marine environments (O’Brien et al., 2015; Patwardhan et al., 2018), led us to hypothesize that biofilms on our samples were not at an early stage of formation. Among the other highly represented taxa in our dataset, we found Actinobacteria and Cyanobacteria. Members of the phylum Actinobacteria have been reported as an abundant component of plastic debris communities (Salta et al., 2013; Pinto et al., 2019), as well as prokaryotic phototrophs, such as Cyanobacteria, which were highly represented on PP and PE items (Zettler et al., 2013).
Regardless of the polymer type, most plastics in our study shared some of the most abundant ASVs, such as those identified as belonging to Pseudoalteromonas and Alteromonas genera, known as major components of the core plastisphere of marine PP and PE (Zettler et al., 2013) and PET debris (Oberbeckmann et al., 2016), respectively. Pseudoalteromonas has also been described as a hydrocarbon-degrading bacterium (Chronopoulou et al., 2015) and, for this reason, our results suggest a possible role of plastic-inhabiting microbes in the degradation of plastic polymers. Dokdonia (Yoon et al., 2005), a genus including strictly aerobic, phototrophic and biofilm-forming marine bacteria, was particularly abundant in Naples samples. Interestingly, this genus was almost completely undetectable in sediments and water samples from both sites, suggesting that the plastic surface offered more advantageous conditions for the survival and growth of this microbe compared to the aquatic environment.
Notably, none of the ASVs was shared by all plastic samples, neither by all PP samples; a single ASV (Sulfitobacter spp.) was common to 100% of PE items. This result further supports the lack of any significant similarity between plastic communities attached to different polymer types, recently reported in a meta-analyses on microbial communities associated with PE microplastics suggesting that, although some similarities among PE samples are detectable, the core PE bacterial community still remains to be established (Oberbeckmann and Labrenz, 2020). Nevertheless, two ASVs, Sulfitobacter and Loktanella, both belonging to the Roseobacter clade, were identified as core ASVs (>75%) in PE and PP samples separately. In PE samples, an ASV identified as Muriicola (ph. Bacteroidetes) was also found. All these bacterial genera have been previously described as well adapted to a surface-associated lifestyle, likely suitable for inhabiting plastic debris (Lee et al., 2008; Jain and Krishnan, 2017). Interestingly, Loktanella has been recently described as one of the most abundant taxa (>5%) on marine PE debris by Pinto et al. (2019). Among the other core taxa, Psychrobacter has been reported in biofilms associated with different marine plastics, such as PP (Zettler et al., 2013) and poly ε-caprolactone (Sekiguchi et al., 2011). Taken together, these results suggest that the above-mentioned genera constitute a likely predictable core group of taxa occupying the niche created by marine plastic debris, even in geographically distinct areas. To the best of our knowledge, this is the first time that Propionibacterium is reported in association with marine plastics. However, since this genus includes biofilm-forming bacteria in non-marine habitats (Holmberg et al., 2009), we speculate that, once in the environment, Propionibacterium could be capable of using plastics as an attachment surface.
The surface of plastics might offer a protective niche and an ideal environment also for pathogenic microbial colonizers (Keswani et al., 2016), acting as a potential dissemination vehicle for the spreading of infectious diseases across the marine environment (Keswani et al., 2016; Quero and Luna, 2017; Silva et al., 2019). Specifically for fecal bacteria, plastics may favor their survival in the aquatic environment, offering protection from predation and sunlight, which strongly drive fecal bacterial decay in the aquatic environments (Wanjugi and Harwood, 2013). Previous studies reported the presence of potential pathogenic bacteria on (micro)plastics in the aquatic environments. Zettler et al. (2013) detected high abundances of Vibrio on PP particles sampled in the North Atlantic, whereas McCormick et al. (2014) demonstrated that microplastics in rivers were a distinct microbial habitat, and represented a novel vector for the downstream transport of unique bacterial assemblages, including pathogens. Also, the authors discussed a potential link between the input of treated wastewater and the occurrence of Campylobacteraceae, being this group of microorganisms associated with human feces and gastrointestinal disorders.
In this study, both plastic samples from Ancona and Naples showed evidences of microbial pollution, although this presence was proportional to the contrasting levels of pollution that characterized the two sites (more pollution in Naples). This finding corroborates the hypothesis of plastics as a potential dissemination vehicle for pathogens. The presence of a sewage discharge close to the sampling area in Naples was indeed reflected by the potential pathogenic signature in seawater, which showed higher abundances of sewage-associated and feces-associated indicators than Ancona samples. The same pathogenic signature was also observed in the plastic items from the same site, indicating that, in highly polluted sites, fecal microbes can easily move from seawater to plastics. It must be also pointed out that, when considering the potential pathogenic signatures alone, we could not find a correlation between polymer type and microbial pollutants, confirming a minor role of the synthetic polymer also in selecting the potential pathogenic community. It is well known that pathogenicity cannot be predictable, and that it may vary sensibly even from one strain to another (Ehrlich et al., 2008); our analysis does not provide any proof of pathogenicity of plastic-associated microbes, but instead highlights the presence of potentially pathogenic bacteria on these substrates, suggesting the need for more detailed investigations. As plastic debris can move quickly and across large spatial distances in the ocean, these findings have important implications from the health-risk perspective, and highlights important, but seldomly discussed consequences for marine plastic pollution.
Conclusion
In this study we explored the microbial life associated with different types of marine plastic debris in two geographically separated coastal sites and observed that communities associated with the plastic items significantly differed according to the site of collection, but not to the plastic polymer. Also, plastic communities differed from those living in the surrounding seawater and sediment, suggesting that the plastisphere is unique with respect to the surrounding aquatic environment. We also report that plastic debris can be carriers of fecal bacteria and other potential human pathogens originating from anthropogenic activities, and that the potential pathogenic signature on plastics reflects the level of microbial pollution in the surrounding environment, regardless of the polymer type. We conclude that microbial community composition on plastics is both driven by environmental features and biofilm formation processes, and that a small proportion of taxa are common to the plastisphere living on different polymers. Some of the most abundant taxa found of plastic debris were previously described as hydrocarbon degraders, leading to hypothesize a possible role of these plastic-inhabiting microbes in the substrate degradation. Being studies on the driving factors of microbial diversity on marine plastics still controversial, further investigations will be needed to better understand the magnitude, the factors and the interactions that drive microbial life associated with plastics, and to identify the mechanisms behind microbial colonization.
Data Availability Statement
The sequences analyzed in this study have been submitted to the SRA – Sequence Read Archive (BioProject accession number PRJNA558771, SAMN12497894 to SAMN12497918).
Author Contributions
GL and GQ designed the study. MB, GQ, EM, and GL collected the samples. MB, GQ, EM, and CA performed laboratory analyses. MB, GQ, DG, and RD performed bioinformatic and statistical analyses. MB, GQ, DG, EM, CV, and GL interpreted the data. MB, GQ, and GL wrote the manuscript. All authors provided the constructive comments, revised and edited the manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was funded by Davines (Italy; https://www.davines.com) within the Research program “Ricerca il Futuro” granted to GMQ. The authors also acknowledge support by the project Circles (funded by EU Horizon 2020 Research and Innovation Programme under grant agreement No. 818290) and the AMOBIO Unit of Stazione Zoologica “Anton Dohrn” of Naples for technical assistance and consultancy in the preparation and observation of SEM samples.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2020.00262/full#supplementary-material
Footnotes
References
Amaral-Zettler, L. A., Zettler, E. R., Slikas, B., Boyd, G. D., Melvin, D. W., Morrall, C. E., et al. (2015). The biogeography of the Plastisphere: implications for policy. Front. Ecolo. Environ. 1, 541–546. doi: 10.1890/150017
Andrady, A. L. (2011). Microplastics in the marine environment. Mar. Pollut. Bull. 62, 1596–1605. doi: 10.1016/j.marpolbul.2011.05.030
Avio, C. G., Gorbi, S., and Regoli, F. (2015). Experimental development of a new protocol for extraction and characterization of microplastics in fish tissues: first observations in commercial species from Adriatic Sea. Mar. Environ. Res. 111, 18–26. doi: 10.1016/j.marenvres.2015.06.014
Balasubramanian, V., Natarajan, K., Hemambika, B., Ramesh, N., Sumathi, C. S., Kottaimuthu, R., et al. (2010). High-density polyethylene (HDPE)-degrading potential bacteria from marine ecosystem of Gulf of Mannar, India: high-density polyethylene (HDPE)-degrading potential bacteria from marine ecosystem. Lett. Appli. Microbiol. 51, 205–211. doi: 10.1111/j.1472-765X.2010.02883.x
Barnes, D., and Fraser, K. (2003). Rafting by five phyla on man-made flotsam in the Southern Ocean. Mar. Ecol. Prog. Ser. 262, 289–291. doi: 10.3354/meps262289
Barnes, D. K. A., Galgani, F., Thompson, R. C., and Barlaz, M. (2009). Accumulation and fragmentation of plastic debris in global environments. Philos. Trans. R. Soc. B Biol. Sci. 364, 1985–1998. doi: 10.1098/rstb.2008.0205
Bastian, M., Heymann, S., and Jacomy, M. (2009). “Gephi: an open source software for exploring and manipulating networks,” in Proceedings of the Third international AAAI Conference on Weblogs and Social Media, (Menlo Park: AAAI).
Brouwer, R., Hadzhiyska, D., Ioakeimidis, C., and Ouderdorp, H. (2017). The social costs of marine litter along European coasts. Ocean Coast. Manag. 138, 38–49. doi: 10.1016/j.ocecoaman.2017.01.011
Bryant, J. A., Clemente, T. M., Viviani, D. A., Fong, A. A., Thomas, K. A., Kemp, P., et al. (2016). Diversity and activity of communities inhabiting plastic debris in the North Pacific Gyre. MSystems 1:e00024-16.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J., and Holmes, S. P. (2016). DADA2: high resolution sample inference from amplicon data. Nat. Methods 13, 581–583. doi: 10.1038/nmeth.3869
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336.
Carson, H. S., Nerheim, M. S., Carroll, K. A., and Eriksen, M. (2013). The plastic-associated microorganisms of the North Pacific Gyre. Mar. Pollut. Bull. 75, 126–132. doi: 10.1016/j.marpolbul.2013.07.054
Chronopoulou, P. M., Sanni, G. O., Silas-Olu, D. I., van der Meer, J. R., Timmis, K. N., Brussaard, C. P., et al. (2015). Generalist hydrocarbon-degrading bacterial communities in the oil-polluted water column of the North Sea: generalist marine hydrocarbon-degrading bacteria. Microb. Biotechnol. 8, 434–447. doi: 10.1111/1751-7915.12176
Cózar, A., Martí, E., Duarte, C. M., García-de-Lomas, J., Van Sebille, E., Ballatore, T. J., et al. (2017). The Arctic Ocean as a dead end for floating plastics in the North Atlantic branch of the thermohaline circulation. Sci. Adv. 3:e1600582. doi: 10.1126/sciadv.1600582
Cózar, A., Sanz-Martín, M., Martí, E., González-Gordillo, J. I, Ubeda, B., Gálvez, J. Á, et al. (2015). Plastic accumulation in the Mediterranean Sea. PLoS One 10:e0121762. doi: 10.1371/journal.pone.0121762
Dang, H., and Lovell, C. R. (2000). Bacterial primary colonization and early succession on surfaces in marine waters as determined by Amplified rRNA Gene Restriction Analysis and sequence analysis of 16S rRNA genes. Appl. Environ. Microbiol. 66, 467–475. doi: 10.1128/aem.66.2.467-475.2000
De Tender, C. A., Devriese, L. I., Haegeman, A., Maes, S., Ruttink, T., and Dawyndt, P. (2015). Bacterial community profiling of plastic litter in the belgian part of the North Sea. Environ. Sci. Technol. 49, 9629–9638. doi: 10.1021/acs.est.5b01093
Deudero, S., and Alomar, C. (2015). Mediterranean marine biodiversity under threat: reviewing influence of marine litter on species. Mar. Pollut. Bull. 98, 58–68. doi: 10.1016/j.marpolbul.2015.07.012
Dussud, C., Meistertzheim, A. L., Conan, P., Pujo-Pay, M., George, M., Fabre, P., et al. (2018). Evidence of niche partitioning among bacteria living on plastics, organic particles and surrounding seawaters. Environ. Pollut. 236, 807–816. doi: 10.1016/j.envpol.2017.12.027
Ehrlich, G. D., Hiller, N. L., and Hu, F. Z. (2008). What makes pathogens pathogenic. Genome Biol. 9:225. doi: 10.1186/gb-2008-9-6-225
Eiler, A., Heinrich, F., and Bertilsson, S. (2012). Coherent dynamics and association networks among lake bacterioplankton taxa. ISME J. 6, 330–342. doi: 10.1038/ismej.2011.113
Feng, S., Bootsma, M., and McLellan, S. L. (2018). Human-associated Lachnospiraceae genetic markers improve detection of fecal pollution sources in urban waters. Appl. Environ. Microbiol. 84:e00309-18. doi: 10.1128/AEM.00309-18
Fisher, J. C., Eren, A. M., Green, H. C., Shanks, O. C., Morrison, H. G., Vineis, J. H., et al. (2015). Comparison of sewage and animal fecal microbiomes by using oligotyping reveals potential human fecal indicators in multiple taxonomic groups. Appl. Environ. Microbiol. 81, 7023–7033. doi: 10.1128/AEM.01524-15
Giovannelli, D., d’Errico, G., Fiorentino, F., Fattorini, D., Regoli, F., Angeletti, L., et al. (2016). Diversity and distribution of prokaryotes within a shallow-water pockmark field. Front. Microbiol. 7:941. doi: 10.3389/fmicb.2016.00941
Gregory, M. R. (2009). Environmental implications of plastic debris in marine settings–entanglement, ingestion, smothering, hangers-on, hitch-hiking and alien invasions. Philos. Trans. R. Soc. B Biol. Sci. 364, 2013–2025. doi: 10.1098/rstb.2008.0265
Holmberg, A., Lood, R., Mörgelin, M., Söderquist, B., Holst, E., Collin, M., et al. (2009). Biofilm formation by Propionibacterium acnes is a characteristic of invasive isolates. Clin. Microbiol. Infect. 15, 787–795. doi: 10.1111/j.1469-0691.2009.02747.x
Jain, A., and Krishnan, K. P. (2017). Differences in free-living and particle-associated bacterial communities and their spatial variation in Kongsfjorden, Arctic. J. Basic Microbiol. 57, 827–838. doi: 10.1002/jobm.201700216
Keswani, A., Oliver, D. M., Gutierrez, T., and Quilliam, R. S. (2016). Microbial hitchhikers on marine plastic debris: human exposure risks at bathing waters and beach environments. Mar. Environ. Res. 118, 10–19. doi: 10.1016/j.marenvres.2016.04.006
Kirstein, I. V., Kirmizi, S., Wichels, A., Garin-Fernandez, A., Erler, R., Martin, L., et al. (2016). Dangerous hitchhikers? Evidence for potentially pathogenic Vibrio spp. on microplastic particles. Mar. Environ. Res. 120, 1–8. doi: 10.1016/j.marenvres.2016.07.004
Lee, J.-W., Nam, J.-H., Kim, Y.-H., Lee, K.-H., and Lee, D.-H. (2008). Bacterial communities in the initial stage of marine biofilm formation on artificial surfaces. J. Microbiol. 46, 174–182. doi: 10.1007/s12275-008-0032-3
Lobelle, D., and Cunliffe, M. (2011). Early microbial biofilm formation on marine plastic debris. Mar. Pollut. Bull. 62, 197–200. doi: 10.1016/j.marpolbul.2010.10.013
Luna, G. M., Manini, E., and Danovaro, R. (2002). Large fraction of dead and inactive bacteria in coastal marine sediments: comparison of protocols for determination and ecological significance. Appl. Environ. Microbiol. 68, 3509–3513. doi: 10.1128/aem.68.7.3509-3513.2002
Luna, G. M., Quero, G. M., and Perini, L. (2016). Next generation sequencing reveals distinct fecal pollution signatures in aquatic sediments across gradients of anthropogenic influence. Adv. Oceanogr. Limnol. 7, 115–124.
Mansui, J., Molcard, A., and Ourmières, Y. (2015). Modelling the transport and accumulation of floating marine debris in the Mediterranean basin. Mar. Pollut. Bull. 91, 249–257. doi: 10.1016/j.marpolbul.2014.11.037
McCormick, A., Hoellein, T. J., Mason, S. A., Schluep, J., and Kelly, J. J. (2014). Microplastic is an abundant and distinct microbial habitat in an urban river. Environ. Sci. Technol. 48, 11863–11871. doi: 10.1021/es503610r
Newton, R. J., Bootsma, M. J., Morrison, H. G., Sogin, M. L., and McLellan, S. L. (2013). A microbial signature approach to identify fecal pollution in the waters off an urbanized coast of Lake Michigan. Microb. Ecol. 65, 1011–1023. doi: 10.1007/s00248-013-0200-9
Oberbeckmann, S., and Labrenz, M. (2020). Marine microbial assemblages on microplastics: diversity, adaptation, and role in degradation. Annu. Rev. Mar. Sci. 12:1. doi: 10.1146/annurev-marine-010419-010633
Oberbeckmann, S., Löder, M. G. J., and Labrenz, M. (2015). Marine microplastic-associated biofilms – a review. Environ. Chem. 12:551. doi: 10.1071/en15069
Oberbeckmann, S., Loeder, M. G. J., Gerdts, G., and Osborn, A. M. (2014). Spatial and seasonal variation in diversity and structure of microbial biofilms on marine plastics in Northern European waters. FEMS Microbiol. Ecol. 90, 478–492. doi: 10.1111/1574-6941.12409
Oberbeckmann, S., Osborn, A. M., and Duhaime, M. B. (2016). microbes on a bottle: substrate, season and geography influence community composition of microbes colonizing marine plastic debris. PLoS One 11:e0159289. doi: 10.1371/journal.pone.0159289
O’Brien, C., Giovannelli, D., Govenar, B., Luther, G. W., Lutz, R. A., Shank, T. M., et al. (2015). Microbial biofilms associated with fluid chemistry and megafaunal colonization at post-eruptive deep-sea hydrothermal vents. Deep Sea Res. II 121, 31–40. doi: 10.1016/j.dsr2.2015.07.020
Oksanen, J., Blanchet, F. G., Friendly, M., Kindt, R., Legendre, P., and McGlinn, D. (2017). Vegan: Community Ecology Package. R Package Version 2.44.
Pasquini, G., Ronchi, F., Strafella, P., Scarcella, G., and Fortibuoni, T. (2016). Seabed litter composition, distribution and sources in the Northern and Central Adriatic Sea (Mediterranean). Waste Manag. 58, 41–51.
Patwardhan, S., Foustoukos, D. I., Giovannelli, D., Yücel, M., and Vetriani, C. (2018). Ecological succession of sulfur-oxidizing Epsilon- and Gammaproteobacteria during colonization of a shallow-water gas vent. Front. Microbiol. 9:2970. doi: 10.3389/fmicb.2018.02970
Pinto, M., Langer, T. M., Huffer, T., Hofmann, T., and Herndl, G. J. (2019). The composition of bacterial communities associated with plastic biofilms differs between different polymers and stages of biofilm succession. PLoS One 14:e0217165. doi: 10.1371/journal.pone.0217165
Plastics Europe (2018). Plastics – the Facts 2017. An analysis of European Plastics Production, Demand and Waste Data. New Jersey: EPRO.
Quero, G. M., and Luna, G. M. (2017). Surfing and dining on the “plastisphere”: microbial life on plastic marine debris. Adv. Oceanogr. Limnol. 8, 199–207.
Quero, G. M., Perini, L., Pesole, G., Manzari, C., Lionetti, C., Bastianini, M., et al. (2017). Seasonal rather than spatial variability drives planktonic and benthic bacterial diversity in a microtidal lagoon and the adjacent open sea. Mol. Ecol. 26, 5961–5973. doi: 10.1111/mec.14363
Ruiz-Orejón, L. F., Sardá, R., and Ramis-Pujol, J. (2016). Floating plastic debris in the Central and Western Mediterranean Sea. Mar. Environ. Res. 120, 136–144. doi: 10.1016/j.marenvres.2016.08.001
Salta, M., Wharton, J. A., Blache, Y., Stokes, K. R., and Briand, J.-F. (2013). Marine biofilms on artificial surfaces: structure and dynamics. Environ. Microbiol. 15, 2879–2893. doi: 10.1111/1462-2920.12186
Schlining, K., von Thun, S., Kuhnz, L., Schlining, B., Lundsten, L., Jacobsen Stout, N., et al. (2013). Debris in the deep: using a 22-year video annotation database to survey marine litter in Monterey Canyon, central California, USA. Deep Sea Res. Part I Oceanogr. Res. Pap. 79, 96–105.
Sekiguchi, T., Sato, T., Enoki, M., Kanehiro, H., Uematsu, K., et al. (2011). Isolation and characterization of biodegradable plastic degrading bacteria from deep-sea environments. JAMSTEC Rep. Res. Dev. 11, 33–41.
Silva, M. M., Maldonado, G. C., Castro, R. O., de Sá Felizardo, J., Cardoso, R. P., dos Anjos, R. M., et al. (2019). Dispersal of potentially pathogenic bacteria by plastic debris in Guanabara Bay, RJ, Brazil. Mar. Pollut. Bull. 141, 561–568. doi: 10.1016/j.marpolbul.2019.02.064
Strafella, P., Fabi, G., Spagnolo, A., Grati, F., Polidori, P., Punzo, E., et al. (2015). Spatial pattern and weight of seabed marine litter in the northern and central Adriatic Sea. Mar. Pollut. Bull. 91, 120–127. doi: 10.1016/j.marpolbul.2014.12.018
Suaria, G., and Aliani, S. (2014). Floating debris in the Mediterranean Sea. Mar. Pollut. Bull. 86, 494–504. doi: 10.1016/j.marpolbul.2014.06.025
Suaria, G., Avio, C. G., Mineo, A., Lattin, G. L., Magaldi, M. G., Belmonte, G., et al. (2016). The Mediterranean plastic soup: synthetic polymers in Mediterranean surface waters. Sci. Rep. 6:37551.
Suaria, G., Avio, C. G., Regoli, F., and Aliani, S. (2018). “Sub-basin scale heterogeneity in the polymeric composition of floating microplastics in the Mediterranean Sea,” in Proceedings of the International Conference on Microplastic Pollution in the Mediterranean Sea, (Cham: Springer International Publishing), 1–7.
Vianello, A., Da Ros, L., Boldrin, A., Marceta, T., and Moschino, V. (2018). First evaluation of floating microplastics in the Northwestern Adriatic Sea. Environ. Sci. Pollut. Res. 25, 28546–28561. doi: 10.1007/s11356-018-2812-6
Wanjugi, P., and Harwood, V. J. (2013). The influence of predation and competition on the survival of commensal and pathogenic fecal bacteria in aquatic habitats. Environ. Microbiol. 15, 517–526. doi: 10.1111/j.1462-2920.2012.02877.x
Wiedenbeck, J., and Cohan, F. M. (2011). Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol. Rev. 35, 957–976. doi: 10.1111/j.1574-6976.2011.00292.x
Yoon, J.-H., Kang, S.-J., Lee, C.-H., and Oh, T.-K. (2005). Dokdonia donghaensis gen. nov., sp. nov., isolated from sea water. Int. J. Syst. Evol. Microbiol. 55, 2323–2328.
Keywords: plastisphere, plastic debris, microbial communities, pathogens, Mediterranean Sea
Citation: Basili M, Quero GM, Giovannelli D, Manini E, Vignaroli C, Avio CG, De Marco R and Luna GM (2020) Major Role of Surrounding Environment in Shaping Biofilm Community Composition on Marine Plastic Debris. Front. Mar. Sci. 7:262. doi: 10.3389/fmars.2020.00262
Received: 09 January 2020; Accepted: 01 April 2020;
Published: 07 May 2020.
Edited by:
Tony Gutierrez, Heriot-Watt University, United KingdomReviewed by:
Stephen Summers, Nanyang Technological University, SingaporeGunnar Gerdts, Alfred Wegener Institute Helmholtz Centre for Polar and Marine Research (AWI), Germany
Copyright © 2020 Basili, Quero, Giovannelli, Manini, Vignaroli, Avio, De Marco and Luna. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Gian Marco Luna, gianmarco.luna@cnr.it
†These authors have contributed equally to this work
 Marco Basili
Marco Basili Grazia Marina Quero
Grazia Marina Quero Donato Giovannelli
Donato Giovannelli Elena Manini2
Elena Manini2  Carlo Giacomo Avio
Carlo Giacomo Avio Rocco De Marco
Rocco De Marco Gian Marco Luna
Gian Marco Luna