- 1 Department of Surgery, Scott & White Memorial Clinic and Hospital, Temple, TX, USA
- 2 Department of Veterinary Pathobiology, Texas A&M University, College Station, TX, USA
- 3 Department of Plant Pathology and Microbiology, Texas A&M University, College Station, TX, USA
- 4 Department of Molecular and Cellular Medicine, Texas A&M Health Science Center College of Medicine, College Station, TX, USA
- 5 Center for Microencapsulation and Drug Delivery, Texas A&M Health Science College of Medicine, College Station, TX, USA
- 6 Department of Microbial and Molecular Pathogenesis, Texas A&M Health Science Center College of Medicine, College Station, TX, USA
A comprehensive system for national security and defense against terror threats must include biosecurity elements. The latter must encompass guidelines and regulations for monitoring and controlling the use of potentially infectious biothreat agents by investigators, laboratories, or research institutions. Molecular techniques to aid and enhance this monitoring effort could greatly strengthen biosecurity systems. Here, we suggest that natural or induced genomic polymorphisms in research strains of infectious agents may provide “inherent watermarks (IWs)” by which agents can be monitored and tracked. In addition, we describe methods by which the integrity of an IW system can be maintained in the context of a decentralized research infrastructure.
We recently proposed an infectious agent control and tracking system that employs genetically engineered, synthetic, strain-specific DNA sequences – “synthetic watermarks (SWs)” – that allow organisms associated with a particular research entity to be distinguished from those of others in the research community (Jupiter et al., 2010). In the event of release, the offending pathogen can be interrogated for the presence of a registered SW. If such a watermark is present, then information about the possible source becomes immediately available. The SW system requires the designation of a trusted authorizing entity to ensure that SWs, and the strains that carry them, are managed appropriately. The authorizing entity is charged with distributing organisms containing unique SW sequences to individual research entities, cataloging existing SWs, and acting as an intermediary in the sharing of watermarked agents.
The SW system is attractive because it provides a framework that can potentially reduce the possibility of mistaken source assignment in the event of malicious or accidental release of an infectious agent. In addition, a proactive effort by the research community to develop watermarking strategies provides an opportunity to restore the public’s confidence in research entities working with highly infectious agents, which had become tarnished due to the still unresolved anthrax release case. However, the reliance of the SW system on the use of genetically tractable organisms raised several concerns. For example, the approach is costly because each organism requires individualized watermarking, which in turn, requires specialized expertise and organism-specific reagents for the genomic integration of the synthetic DNA sequences. In addition, many infectious agents are not genetically tractable, and hence, cannot be monitored under an SW system.
To address these issues, we propose here a novel approach that exploits IWs to monitor and track infectious agents (Figure 1). An IW is a virtual string of strain-specific polymorphic sequences (e.g., single nucleotide polymorphisms, insertions, deletions, inversions) that are embedded within the genome of an infectious agent. Like an SW, an IW provides a mechanism by which a unique genomic sequence (and corresponding strain) can be linked to a specific research entity. However, unlike its SW counterpart, an IW is a virtual signature that is computationally assembled following the analysis of the whole genome sequence of a particular strain of an agent. A watermarking system based on IWs therefore combines insights about natural or induced genetic variation in research strains with next generation sequencing technology (NGS) to circumvent the genetic engineering of research strains that is required in an SW system (Metzker, 2010).
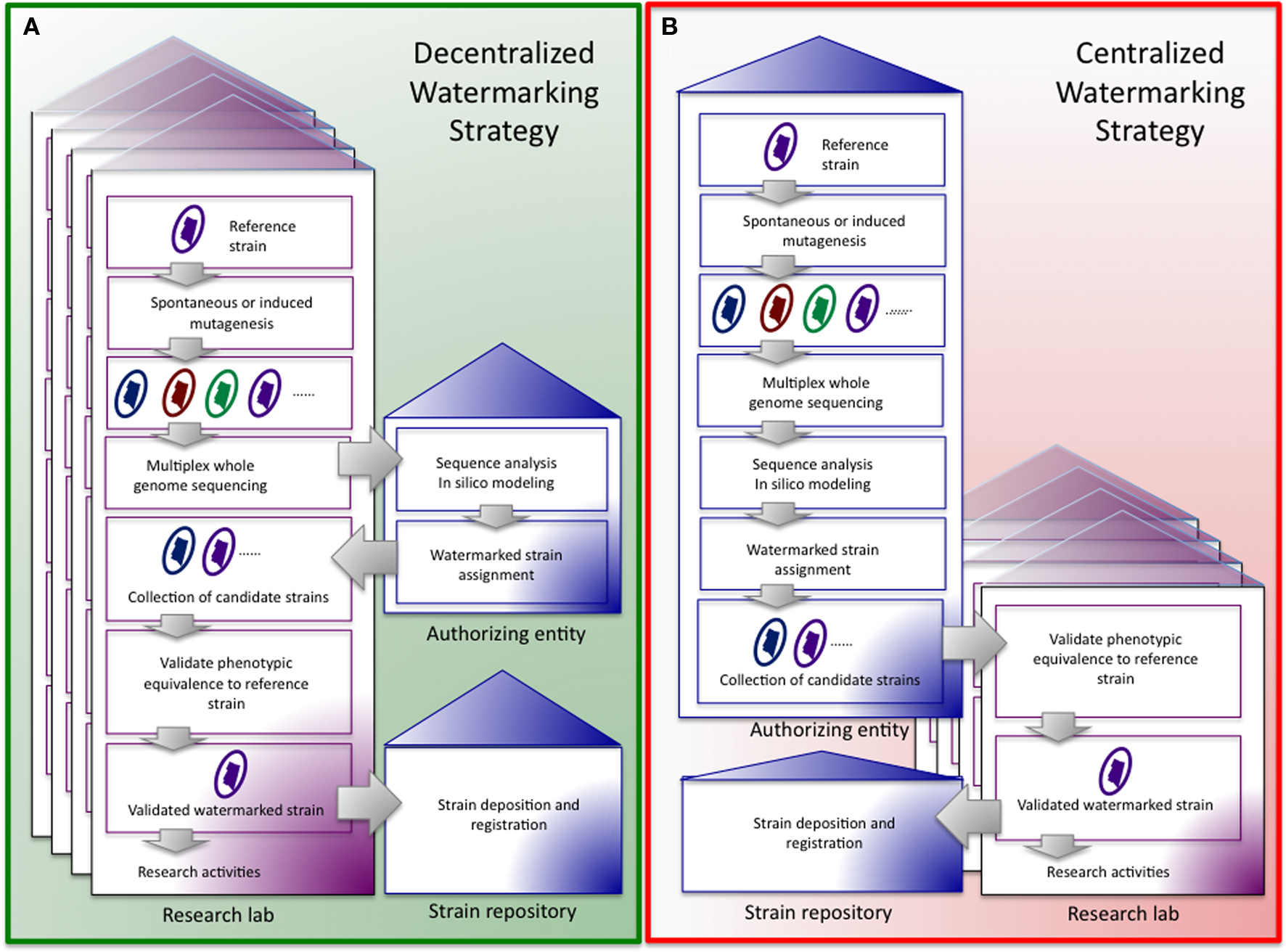
Centralized vs. decentralized management of an IW watermarking system. (A) In a decentralized system the central authorizing entity takes responsibility for sequence analysis and strain assignment, after labs have sequenced genomes and induced mutations. Research entities collect candidate strains, verify phenotypic neutrality, deposit optimal strains with a strain repository, and then perform experiments with authorized, phenotypically neutral strains carrying IWs. (B) In the centralized system the centralized authorizing entity determines the whole genome sequence of candidate strains, induces mutations, assigns and distributes strains to research entities, and collects authorized, phenotypically neutral strains carrying IWs from the same for deposition in a strain repository. Research entities verify phenotypic neutrality.
Manifold strategies for implementing an IW strategy can be envisioned. However, these strategies will likely share several key components. First, spontaneous or induced mutant variants of a particular reference strain of an infectious agent will be generated and sequenced by an approved entity. Strains that not only possess genome sequences that are sufficiently different from one another to support an IW strategy but also are phenotypically equivalent (e.g., as tested by mixed infection with wild-type strains) to the parent strains, will be identified, cataloged, and distributed to participating research laboratories.
Standard operating procedures (SOPs) for using and sharing of IW strains will need to be devised and implemented. First, SOPs that provide for the monitoring of the integrity of the IWs in distributed laboratory strains are required. Although information about mutation rates in a few model organisms has been collected (Mukai, 1964; Kibota and Lynch, 1996; Keightley and Caballero, 1997; Wloch et al., 2001; Loewe et al., 2003), the amount of genetic variation for many biothreat agents has not been empirically determined. Moreover, how individual laboratory practices impact the selection of mutations is unknown. In addition, the expected stability of mutations will need to be experimentally determined in each model system. However, not all mutations need to be included in the virtual watermark. For example, mutations associated with genetic hotspots may be omitted because of the possibility of reversion and the consequent introduction of noise into the watermark signature. That said, a robust watermarking system would be tolerant of mutations that introduce a background rate of noise in the system. Second, for the integrity of the IW system to be maintained, protocols for sharing strains among community members would need to be implemented. In the event of sharing strains among labs, where uniqueness of watermarks is not maintained, assignment of strain to lab of origin cannot be done. Given that sharing of strains is critical to scientific research, we are presented with the question: can an IW system be successfully employed in such a setting? One way to address this need would be to task the authorizing entity with ensuring the appropriate transfer of reagents. In this case, the authorizing entity passages strains or induces mutations to generate new IWs in the genomes of the transferred material. The presence of the new IW can be verified by whole genome sequencing, confirmed by in silico modeling, and registered with the authorizing entity. The watermarked strain can then be delivered to the recipient laboratory for phenotypic testing and use. Finally, an IW system can be supported by either a centralized or decentralized management system. In a centralized system, a controlling authorizing entity is responsible for inducing mutations, computationally verifying IWs, and distributing strains to participating investigators, labs, or institutions. In a decentralized system, the burden of sequencing and mutagenesis falls to the individual research entities, and the authority entity’s role is limited to computational verification and assignment of IWs. In both settings, labs participate in the phenotypic verification of watermarked strains. The centralized approach would have the advantage of supporting standardized protocols for all strains. As costs of NGS declines, the cost of implementing a centralized IW system may also be lower than a corresponding system that relies on engineering SWs in genetically tractable organisms.
Acknowledgments
Thomas Ficht receives support from NIAID/NIH (R01-AI48496-10, U54 AI057156) and DOD/USAMRMC (W81XWH 07 1 0304). Allison Rice-Ficht receives support from USAMRMC W81XWH-07-1-0304, NIH 1 R41 AI068252-01A2 and GCE OPP1007142. James Samuel receives support from NIH (AI057768, AI078213, AI057156). Paul de Figueiredo receives support from NIH (5R21AI072446-02) and NSF (NSF0818758). This publication was also made possible by Grant number 1U54AI057156-0100 from NIAID/NIH. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the RCE Programs Office, NIAID, or NIH.
References
Jupiter, D. C., Ficht, T. A., Samuel, J., Qin, Q. M., and de Figueiredo, P. (2010). DNA watermarking of infectious agents: progress and prospects. PLoS Pathog. 6, e1000950. doi: 10.1371/journal.ppat.1000950.
Keightley, P. D., and Caballero, A. (1997). Genomic mutation rates for lifetime reproductive output and lifespan in Caenorhabditis elegans. Proc. Natl. Acad. Sci. U.S.A. 94, 3823–3827.
Kibota, T. T., and Lynch, M. (1996). Estimate of the genomic mutation rate deleterious to overall fitness in E. coli. Nature 381, 694–696.
Loewe, L., Textor, V., and Scherer, S. (2003). High deleterious genomic mutation rate in stationary phase of Escherichia coli. Science 302, 1558–1560.
Mukai, T. (1964). The genetic structure of natural populations of Drosophila melanogaster. I. Spontaneous mutation rate of polygenes controlling viability. Genetics 50, 1–19.
Citation: Jupiter D, Ficht T, Qin Q-M, Rice-Ficht A, Samuel J and de Figueiredo P (2010) Genomic polymorphisms as inherent watermarks for tracking infectious agents. Front. Microbio. 1:109. doi: 10.3389/fmicb.2010.00109
This article was submitted to Frontiers in Cellular and Infection Microbiology, a specialty of Frontiers in Microbiology.
Received: 25 August 2010;
Accepted: 28 August 2010;
Published online: 30 September 2010
Copyright: © 2010 Jupiter, Ficht, Qin, Rice-Ficht, Samuel and de Figueiredo. This is an open-access article subject to an exclusive license agreement between the authors and the Frontiers Research Foundation, which permits unrestricted use, distribution, and reproduction in any medium, provided the original authors and source are credited.
*Correspondence: pjdefigueiredo@tamu.edu
