Department of Neurobiology, University of Southern California, Los Angeles, CA, USA
We describe in this paper the structure and main features of a domain specific ontology for neuroscience, the BAMS Neuroanatomical Ontology. The ontology includes a complete set of concepts that describe the parts of the rat nervous system, a growing set of concepts that describe neuron populations identified in different brain regions, and relationships between concepts. The ontology is linked with a complex representation of structural and physiological variables used to classify neurons, which is encoded in BAMS. BAMS Neuroanatomical Ontology is accessible on the web and includes an interface that allows browsing terms, viewing criteria for classification, and accessing associated information.
The problem of neuron classification (i.e., a taxonomy of neuron populations and the variables used to classify them) is fundamental for understanding the structure and function of different brain regions, and for modeling purposes (Bota and Swanson, 2007
). Moreover, the large quantity of inherently complex data, and the need for data and information sharing between research groups, drives creation of accessible neuroinformatic systems to define different neuron types and store their morphological and physiological properties.
Neuroscience is characterized by an explosion of terminologies both at the regional and individual neuron levels of organization associated with the vertebrate central nervous system (CNS). The advent of new experimental techniques often leads to new brain parcellation schemes, and the discovery of new cell populations, or the renaming of known populations. Thus, neuroinformatics systems that include terms and their definitions, as well as relationships between terms (i.e., ontologies; Gruber, 1993
), are needed.
As in any research field, specialized ontologies are needed to describe a specified domain – in this case a neuroanatomical account of the nervous system (NS). While not denying the need to align and create more general ontologies, neuroscience ontologies must be constructed first as stand-alone applications because they are designed primarily for use by neuroscientists. Because structural features of the NS are the domain of neuroanatomy, it can be seen as a natural starting point or framework for the construction of neuroscience ontologies.
An ideal neuroscience ontology might be viewed as including first a complete set of structural parts (gray and white matter, and ventricles) and neuron types making up the structural parts, unequivocally defined. Second, it would include axonal projections between regions and neuron types, and an inventory of molecules expressed in them – in the form of relations and properties of the ontology classes. Third, the ideal neuroscience ontology would include morphological, connectional, and electrophysiological properties of neurons modeled as relations and attributes of classes. This ontology would have to allow different types of semantic relationships and be written in a language allowing automatic inferences. By allowing different types of semantic relations, the ideal ontology would also address the problem of multiple names for different NS parts and neuron types. And finally, the ideal neuroscience ontology must be species specific. This is necessary both for capturing what is specific about the NS in each species, and for explicit comparisons between species (homology evaluation). Nervous system parts and neuron types in different species must be considered as separate objects because they can and do differ in their properties and attributes.
Such an ontology cannot be created by a single research group, but is instead the long-term goal of a community project. We have begun construction of a structural neuroscience ontology by providing concepts that name different parts of the rat NS and its component neuron populations. The concepts of the ontology are ordered by “part of” and “is a” relationships, and the ontology includes semantic relations between neuron population concepts and other terms used in the literature.
In this paper we describe the design and main features of the Brain Architecture Knowledge Management System (BAMS) Neuroanatomical Ontology, which is an evolving neuroscience ontology. It is species specific and includes concepts defining the parts that, in the work described here, specifically make up the rat NS. It also includes an evolving and expanding set of concepts and terms that define the neuron types and classes identified in different rat NS regions. And finally, it deals with the problem of multiple nomenclatures for the same neuron populations (Bota and Swanson, 2007
), and is associated with a complex classification schema for neurons.
Ontology Schema
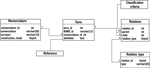
Like other component modules of BAMS, the Neuroanatomical Ontology is implemented in MySQL, and its backend relational schema is shown in Figure 1
.
Figure 1. The OR schema of BAMS Neuroanatomical Ontology.
The construction principle of BAMS Neuroanatomical Ontology is that different NS parts can be parceled based on different criteria, and the smallest divisions consist of neuron types. The design of the ontology follows the principle of disjoint decomposition of classes (Gómez-Pérez et al., 2003
). Each of the concepts included in the BAMS Neuroanatomical Ontology can be associated with a textual definition, as well as with original literature references. The most abstract classes of BAMS Neuroanatomical Ontology follow the Foundational Model of Anatomy (FMA; Rosse and Mejino, 2003
; see Figure 2
). The design of the ontology also follows the criteria for inclusion in OBO foundry (Smith et al., 2005
): concepts are associated with textual definitions and uniquely identified by alphanumeric identifiers, the ontology is also in an OWL format, we reused as much as possible other existent ontologies, the “is-a” relation follows the principle of single inheritance, and the “part-of” relation is either of the mereotopological type, which can be further dissociated in the basic topological relations between two objects, or the aggregate type.
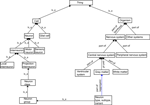
Figure 2. The general structure of BAMS Neuroanatomical Ontology.
The root of the ontology is the abstract concept called “Thing”. The concepts “Organism” and “Cell” are instances of “Thing”, and in turn are the most abstract classes for the major parts of the ontology. “Organism” is similar to the concept of “Body” in FMA. The meaning of a “class” in our ontology is the same as in any ontology: classes are formed by the set of concepts describing neuron populations with different degrees of abstraction (left hand side of Figure 2
) and NS parts related by the “part of” relationship (the right hand side of Figure 2
).
The concepts that describe different parts of the rat NS (the right hand side of Figure 2
) were adapted from Swanson (2003) and from the rat brain Atlas (Swanson, 1998
), which includes a complete, internally consistent and fully documented nomenclature of the rat NS. Thus, the class “Gray matter” of BAMS Neuroanatomical Ontology includes those brain and spinal cord parts defined in the rat brain Atlas (Swanson, 1998
). The first six levels of CNS gray matter, the first three levels of peripheral nervous system and ventricular system, and the first two levels of white matter and nerve fibers were adapted from rat brain Atlas (Swanson, 2003
; Table A, page 166), which are general enough to apply to any mammalian species. Each class that can be applied to the mammalian NS is associated with a definition. The subparts of these general regions are specific to the rat (at least at the present time), and their definitions (descriptions) were taken from the rat brain Atlas (Swanson, 1998
).
A concept (e.g., a class) is uniquely identified in BAMS Neuroanatomical Ontology by its name and nomenclature because the concept can be defined differently by different authors, or the same name can refer to different classes. Specifically, Nomenclatures refer to concepts or internally consistent sets of concepts or terms defined by an author or group of authors, and are associated with a set of references in the literature. Nomenclature is therefore an attribute of concepts and terms included in the ontology. Neuron concepts included in BAMS Neuroanatomical Ontology are those terms that best describe, based on published data and collator knowledge, the set of distinguishable neuron populations in a particular region or set of regions. Methods for constructing nomenclatures are described in detail in Bota et al. (2005) and Bota and Swanson (2007)
.
The ontology part that includes neuron classes identified in different brain regions of the rat CNS was also constructed from original literature. Because this part of the ontology was constructed using “is-a” relationships, the children of a class are considered to be its instances, and therefore inherit all its properties. The “is-a” inheritance follows the principle of single inheritance, recommended by Smith et al. (2005)
.
Below we describe the construction method that was used for the Neurons part of the ontology. The children of the class “cell” are “neuron” and “glial cell” (Figure 1
). Children of the class “neuron” are “sensory”, “motor”, “sensory-motor”, and “internuncial (interneuron)”, defined by fundamental connectional input-output relationships (Swanson, 2003
). These classes are further parceled into instances, depending on sensory modality, type of motor output, and length of interneuron axon projection – local, or projection between regions (Swanson, 2003
). All these classes are general enough to be applied to any vertebrate species. The subsequent classes, denoted generally by “neuron class” and “neuron group”, are specific to particular neural systems (Figure 1
). The lowest-level classes of BAMS Neuroanatomical Ontology are made of neuron types (or subtypes and variants), which are also parts of the smallest “Gray matter” parts, according to the ontologies construction principle. The present version of BAMS Neuroanatomical Ontology allows registration of neuron populations and not of individual neurons. Future extensions of the ontology will include the possibility of registering attributes of individual neurons, which will have distinct ID’s (see below). It is worth noting that vertebrate neurons are usually described in terms of populations, whereas invertebrate neurons are often described as individual instances.
Each concept included in the ontology has as an attribute a unique alphanumeric identifier that is constructed like Gene Ontology identifiers (Ashburner and Lewis, 2002
). The general form of the identifier is BAMSC:nnnnnnn where nnnnnnn is a unique integer.
BAMS Neuroanatomical Ontology allows several types of relationships between concepts. It first includes parent–child “is a” and “part of” relationships. The “is a” relationship is established between concepts when the child is an instance of the parent. The “part of” relationship in BAMS Neuroanatomical Ontology has two different meanings. The first meaning, that of primitive relation of parthood between two or more spatial entities (Smith et al., 2005
), is associated with the relations between NS regions. Thus, any NS region is considered as a spatial entity that can be further divided in at least two subdivisions. The second meaning of the “part of” relationship is that of aggregate and applies to the relations between cell types and the lowest level of neural regions (see Figure 2
). Any neural region can be made of a collection of cell bodies of different neuron types. This relation allows the existence of a single neuron type in a neural region.
In addition, BAMS Neuroanatomical Ontology includes semantic relationships between neuron terms and concepts, viewed as populations. The allowed semantic relationships are “identical”, “includes” with its complementary relationship “is included”, “overlap”, and “different”. The relationship “identical” is interchangeable with the “synonym” relationship used in other ontologies. We chose the term “identical” because it is more appropriate (by convention) when comparing neuron populations. However, the web interface of the ontology will display synonyms and not identical terms (Figure 4
). We allow semantic relations other than “identical” or “synonym” because of difficulties in properly defining neuron types that form the mammalian CNS. Neuron classification can be made according to different criteria (Bota and Swanson, 2007
; see below) and neuron types and classes have been defined using different techniques, each of them with relative advantages and limitations. Definitions of the same cell population using different criteria usually do not necessarily imply identity of concepts. The constant development and refinement of techniques used in neuron identification also may lead to the discovery of new types, to redefining existing types, and to the emergence of new classification schemes.
Relationships that are established in BAMS Neuroanatomical Ontology are collated from original research literature, and are associated with annotations and references.
Criteria for Classification
Any parent–child relationship can be associated in BAMS Neuroanatomical Ontology with criteria for classification that are collated from associated references. These are the major criteria used by various authors to classify neurons: morphology, specialized parts, input and output regions or neurons types, regions where the neurons were identified, expressed molecules, and physiology. The database classification criteria schema (Figure 3
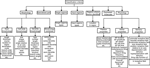
) was constructed to comply with measurements performed by a broad range of research groups, and with the most recent efforts to create a unified terminology of neuron types (Markram, 2006
), and it was implemented in BAMS. All variables included in the classification criteria are in text format, and are associated with annotations and references.
Figure 3. Schema of classification criteria associated with “is-a” relations between neurons in BAMS Neuroanatomical Ontology.
The “Morphology” criterion encodes for those structural variables used for neuron classification. It includes four subcriteria specific to each of the three major parts of neurons (cell body, axon, and dendrites). The fourth subcriterion, “Morphology combined”, includes as variables ratios between variables included in the first three subcriteria. Overall, the “Morphology” criterion includes 47 variables as of now.
The “Specialized part” criterion is specific to sensory or sensory-motor neurons and encodes for those molecules, or neuron parts, that make possible transformation of an external signal into neural signals. For example, opsin, which is present in photoreceptors, is considered a “specialized part” of these neurons.
Criteria “Input regions” and “Output regions” encode for those NS regions that are either the input sources, or the targets of the classified neurons. They include the “Type of region” subcriterion, which allows three values: “neural” and “non-neural” region, and “part of major fiber tract.” The first two values are used to discriminate between interneurons and motor neurons. The third value is applied whenever the axons of a type or class of neurons (e.g., retinal ganglion cells) are part of, or make up, a major fiber tract, without the explicit specification of target regions. The dissociation between local and projection interneurons is made according to the brain regions where the neuron types were identified and the target regions. Variables of these criteria include the names of input sources and target regions, and the laterality of connection (ipsilateral, contralateral, and bilateral).
The “Expressed molecules” criterion encodes for those molecules expressed by the classified neurons, and used to define different neuron types and classes.
The “Regions where neuron identified” criterion encodes for those brain regions where different neuron types were identified, and that are used to define them. This criterion includes an additional subcriterion, “Location in region”, which encodes for the specific localization of neurons within the region. This criterion is used in highly laminated structures, such as the retina or the cerebral cortex, where different neuron types occupy specific layers.
The “Physiology” criterion encodes for those physiological properties used to define and identify neurons. We subdivided this criterion into three subcriteria, depending on the type of response elicited from stimulation and the corresponding measured variables: “Passive properties”, “Active properties”, and “Sensory properties”. The “Passive properties” subcriterion includes those variables associated with the cable properties of the neuronal membrane, and the “Active properties” includes action potential variables. The “Sensory properties” subcriterion shares some of its variables with “Active properties”, but it is considered as a separate class of physiological properties because the latter are elicited by sensory stimuli. Examples of variables included in each subcriterion are shown in Figure 3
. Overall, the “Physiology” criterion so far includes 45 variables divided between the three subcriteria.
Collators can associate with each “is a” relationship any number of classification criteria or subcriteria, and for each of them any number of classification variables.
Web Interface
BAMS Neuroanatomical Ontology is fully accessible on the web and it includes a complex interface that allows users to browse concepts, view criteria for classification, and also access additional information. The general design of the web interface follows that used for the other BAMS modules (Bota et al., 2005
; Bota and Swanson, 2006
).
BAMS Neuroanatomical Ontology web interface displays the concepts in an expandable tree fashion that has links associated with each concept, as shown in Figure 4
(URL: http://retzius.usc.edu/bkms/bams-ontology.html). The information that is displayed in the right frame of the interface includes the concept, its definition and the defining nomenclature, the associated BAMS identifier, concepts related by “is a” relationships, its synonyms, and a link to details included in BAMS. If the concept is a neuron, then the associated link will lead to a page that displays those brain regions where that neuron class or type was identified, related terms (semantic relations) and additional information such as associated annotations and references, as shown in Figure 5
. Otherwise, the retrieved page includes the description of the brain region as recorded in BAMS, links to its input and output patterns, chemoarchitecture profiles, and neuron types (Bota et al., 2005
; Bota and Swanson, 2006
). Clicking on the links associated with neuron types identified in the retrieved brain region will also return their parents and the associated classification criteria, if recorded in the ontology (Figure 6
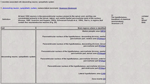
). Users can access classification criteria by clicking on the “criteria” links, which are associated with each displayed neuron class. These criteria are displayed as lists that include those variables used to classify neuron types and classes, together with the associated annotations and references (Figure 7
).
Figure 4. The web interface of BAMS Neuroanatomical Ontology allows users to browse concepts and retrieve their definitions, defining nomenclatures, parents and children, and synonyms.
Figure 5. BAMS Neuroanatomical Ontology web interface allows users to view those brain regions where the neuron type or class of interest was identified. Links are associated with each brain region abbreviation, and they return descriptions of the regions, as recorded in BAMS (Bota et al., 2005
; Bota and Swanson, 2006
).
Figure 6. Information that can be retrieved in BAMS and is related to neuron concepts include related terms, their semantic relations, annotations, references and collators, as well as the set of parents with links to the associated classification criteria. The neuron type “C others” defined in the nomenclature “Huxlin and Goodchild” (Huxlin and Goodchild, 1997
) has two related terms in BAMS Neuroanatomical Ontology: it includes the retinal ganglion cell type III, nomenclature Perry (Perry, 1979
) and has as synonym the retinal ganglion cell C3, nomenclature Sun (Sun et al., 2004
).
Figure 7. Users can access classification criteria associated with each “is a” relationship included in BAMS Neuroanatomical Ontology. Retrieved criteria are organized as ordered lists of variables used to define and classify the neuron concepts. Associated information includes textual annotations, references, and collator names. The list of variables shown in this Figure is used to classify rat retinal ganglion cells A2 inner (Huxlin and Goodchild, 1997
) as projection interneurons, passing through the intermediary classes.1:11 PM 6/6/2008
BAMS Neuroanatomical Ontology (concepts, ID’s, definitions, nomenclatures, and relations) is available in XML format (http://retzius.usc.edu/bkms/neuron-ontology.xml), which is similar to XML versions of Cell Cycle Ontology and GO Ontology (http://www.cellcycleontology.org/html/index.html).
The brain regions and neuron concepts, as well as parent–child relationships, are also available to users and in OWL format, which was constructed by Stephen Larson (UCSD), and it can be used for automated reasoning. Finally, a link to the OWL file of the Swanson (1998)
nomenclature also is provided in our website. (http://esw.w3.org/topic/ HCLS/HCLSIG_DemoHomePage_HCLSIG_Demo).
Data and Data Entry
At present, BAMS Neuroanatomical Ontology is populated with 1,120 concepts associated with the rat NS, including more than 1,000 brain parts concepts and 128 neuron concepts.
The ontology also includes more than 160 additional terms that are semantically related to the neuron concepts, defined in 57 neuron nomenclatures, and more than 1,400 relations. The brain parts concepts are a complete description of the rat NS and the neuron concepts describe neuron types and classes identified in 46 rat brain regions. At present, BAMS Neuroanatomical Ontology includes the complete inventories of terms used to define neuron populations of the rat retina and the cerebellar cortex.
New concepts, relationships, and related terms can be inserted in BAMS Neuroanatomical Ontology by BAMS collators. The quality and completeness of web accessible information is checked by BAMS curators. At present, BAMS has eight collators and two curators. Neuroscientists can become collators by contacting the curators. Thus, experimental data can be entered directly by authors who publish them (thus becoming curators), or sent by email to current BAMS collators, or curators. The minimal information that can be inserted in BAMS Neuroanatomical Ontology includes the concept, its definition, the associated annotation and reference, and the nomenclature. If a new neuron nomenclature has to be defined, then additional information should include the name and acronym of the nomenclature, and the associated references.
Complexities of the neuron classification problem, combined with the existence of alternate nomenclatures for brain parts and neurons, make ontologies necessary tools for the explicit specification of concepts (Gruber, 1993
) and for mapping or indexing between terms. Brain parts ontologies and terminologies have been designed as stand alone applications (Bowden and Dubach, 2003
), or as parts of larger ontologies (Rosse and Mejino, 2003
) or general dictionaries (Mouse adult gross anatomy ontology; http://obofoundry.org/cgi-bin/detail.cgi?id=adult_mouse_anatomy). Cell ontologies also have been developed as components of larger bioinformatics [Gene Ontology Database (Ashburner and Lewis, 2002
; Bard et al., 2005
), DopaNet (Le Novère and Donizelli, 2004
)] and anatomy ontologies (Rosse and Mejino, 2003
), and other cell ontologies are available as well online (Cell Type: http://www.sanbi.ac.za/evoc/ontologies_html/latest/celltype.html; TissueDB: http://tissuedb.ontology.ims.u-tokyo.ac.jp:8082/tissuedb/). Ontologies that have been designed to represent parts of neurons include the Subcellular Anatomy (SAO; http://ccdb.ucsd.edu/sao.html; Larson et al., 2007
) and the Synapse Ontology (Zhang et al., 2007
). However, none of them are designed specifically for the nervous system. BAMS Neuroanatomical Ontology is thus designed specifically for representing domain knowledge in neuroscience. It is a lightweight ontology and it was designed using database technology. The ontology includes a complete and internally consistent set of concepts that describe brain parts and a growing set of concepts that describe different neuron types and classes of the rat NS. Allowed relations include parent–child “is a” and “part of” relationships, and semantic relationships that allow mapping between terms defined in different nomenclatures and thus alignment of alternate classification schemas (Bota and Swanson, 2007
). BAMS Neuroanatomical Ontology is the first to address problems of multiple cell nomenclatures in the vertebrate nervous system, allowing insertion of terms uniquely identified by species and nomenclature from the primary research literature.
BAMS Neuroanatomical Ontology is one of the first semantic frameworks that support the complex representation of criteria used to classify neuron types and classes. Other related ontologies include the NeuronDB of Senselab (http://senselab.med.yale.edu/neurondb/default.asp), NeuroMorpho (http://neuromorpho.org/neuroMorpho/index.jsp), and SAO.
A problem specific to the ontology life cycle is alignment across ontologies. The NS parts included in BAMS Neuroanatomical Ontology can easily be aligned with other ontologies, such as FMA, because they include a subset of regions that are common across vertebrates and across mammals. This is also true for the most abstract neuron classes, which can be found in any vertebrate species. Therefore, alignment of BAMS Neuroanatomical Ontology with ontologies such as the Cell Type Ontology should be easily made. BAMS Neuroanatomical Ontology can also be aligned with nervous system parts of SAO, which were designed to describe neurons and their structural and molecular properties. In turn, SAO can easily integrate neuron classes used by BAMS Neuroanatomical Ontology because it addresses the species: the rat. At present, there is an active effort to integrate BAMS and SAO and to rewrite them in RDF/OWL for automatic inferences about neuronal connections from inserted axonal projections. BAMS Neuroanatomical Ontology can also augment general anatomical dictionaries used for gene mapping, such as the Mouse adult gross anatomy ontology, because the first six levels of gray matter are general for any mammalian species.
However, the alignment of NS parts and of lower level neuron classes that are specific to the rat is more problematic because existing ontologies are not species specific, or are specific to humans (FMA). A possible solution is to attach to each concept an attribute of the form used by the Gene Ontology to indicate species. However, the alignment of concepts that are species specific should be made only after evaluation of similarities (homologies) of parts, or of neurons. Alignment of ontologies also implies adding new types of relations between concepts that will represent structural and functional similarities of parts and neurons across species.
At least six extensions and refinements of BAMS Neuroanatomical Ontology are obvious and necessary. First, the inventory of terms used for rat NS neurons needs completion. Second, the ontology needs generalizing to include concepts specific to other species. This will be accomplished initially by extending the ontology with mouse parts and neuron concepts, and then with macaque and human concepts. Third, classification criteria need to be converted to attributes of concepts and relations between them. Fourth, the ontology needs extending to include experimental data and to store information about individual neurons. Fifth, functionality must be added for tracking changes and updates. Sixth, BAMS Neuroanatomical Ontology needs to be stored in ontology repositories such as OBO (http://obofoundry.org), NCBO (http://www.bioontology.org), and the Ontology Lookup Service (http://www.ebi.ac.uk/ontology-lookup). It also needs to be made available through web services, as the other modules of BAMS (Dinov et al., 2006
).
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
We thank to Stephen Larson (UCSD) for the OWL file construction. This work was supported by NIH Grants MH61223, NS16668, and NS050792.