- 1Key Laboratory of Forest Protection, Research Institute of Forest Ecology, Environment and Protection, Chinese Academy of Forestry, State Forestry Administration, Beijing, China
- 2Key Laboratory of Insect Developmental and Evolutionary Biology, Institute of Plant Physiology and Ecology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai, China
- 3Department of Entomology, University of Kentucky, Lexington, KY, USA
The pine caterpillar moth, Dendrolimus punctatus, is a devastating forest pest. Genetic manipulation of this insect pest is limited due to the lack of genomic and functional genomic toolsets. Recently, CRISPR/Cas9 technology has been demonstrated to be a promising approach to modify the genome. To investigate gene functions during the embryogenesis, we introduced CRISPR/Cas9 system in D. punctatus to precisely and effectively manipulate gene expressions inmutant embryos. Compared to controls, knocking out of DpWnt-1, a gene well known for its role in the early body planning, led to high embryonic mortality. Among these mutants, 32.9% of the embryos and larvae showed an abnormal development. DpWnt-1 mutants predominantly exhibited abnormal posterior segments. In addition, multiple phenotypes were observed, including the loss of limbs and the head deformation, suggesting that DpWnt-1 signaling pathway is necessary for anterior segmentation and appendage development. Overall, our results demonstrate that CRISPR/Cas9 system is feasible and efficient in inducing mutations at a specific locus in D. punctatus. This study not only lays the foundation for characterizing gene functions in a non-model species, but also facilitates the future development of pest control alternatives for a major defoliator.
Introduction
The pine caterpillar moth Dendrolimus punctatus (Lepidoptera: Lasiocampidae) is one of the most destructive forest pests in China and Southeast Asia, where it attacks a variety of pine species and causes extensive forest damages (Billings, 1991; Zeng et al., 2010). Through the years, D. punctatus management has relied primarily on synthetic insecticides. The advent of Genomic Era facilitates the development of environmentally friendly and sustainable control alternatives. The sterile insect technique (SIT) is an environmentally friendly insect control technology that relies on the release of large numbers of sterile males to mate with wild females to suppress pest population (Benedict and Robinson, 2003). The application of this method, however, is limited by the production of undesired females which need to be separated and eliminated. A modified SIT technique, the release of insects carrying a conditional dominant lethal gene (RIDL) can overcome this issue by inducing repressible female-specific lethality (Heinrich and Scott, 2000; Horn and Wimmer, 2003; Fu et al., 2007; Windbichler et al., 2008; Tan A. et al., 2013). This concept has been proofed in the mosquito control, both in laboratory and confined field tests (Thomas et al., 2000; Alphey and Andreasen, 2002; Alphey et al., 2002).
RNAi and genome editing are the primary tools to elucidate gene functions (Mao et al., 2013; Ma et al., 2014; Xu et al., 2014, 2015; Hammond et al., 2016). However, RNAi efficiency is highly variable in lepidopterans which underlying mechanisms are still unknown. More importantly, heritable RNAi effects have yet to be documented in lepidopterans (Bettencourt et al., 2002; Terenius et al., 2011; Swevers and Smagghe, 2012). In contrast, genome editing can achieve target gene mutagenesis by inducing irreversible DNA breaks (Corrigan-Curay et al., 2015). Genome editing tools, including customized zinc-finger nucleases (ZFN), transcription activator-like effector nucleases (TALEN) or clustered regularly interspaced short palindromic repeats-associated nuclease 9 (CRISPR-Cas9), can effectively modify the genomic DNA of organisms. By inducing DNA double-stranded breaks (DSBs), these tools stimulate subsequent homologous recombination (HR) and/or non-homologous ends-joining (NHEJ), which facilitate genome manipulation at a target locus (Harrison et al., 2014). Although ZFN and TALEN have been used for gene targeting, the complexity of module construction and the costs associated with these tools limit their applications. Recently, a bacteria-derived CRISPR/Cas9 system, consisting of CRISPR RNAs and Cas proteins, circumvents some of these issues. With the aid of two short RNA molecules, namely CRISPR RNA (crRNA) and trans-encoded CRISPR RNA (tracrRNA), the Cas9 endonuclease can cleave a specific sequence that is targeted by the RNAs. These two RNA molecules can be fused artificially to form a chimeric RNA molecule called single guide RNA (sgRNA). CRISPR/Cas9 system has been used to produce heritable mutations in non-model organisms, including RNAi-recalcitrant Lepidoptera, such as Bombyx mori, Danaus plexippus, Spodoptera litura, Plutella xylostella, Spodoptera littoralis, and Helicoverpa armigera (Wang et al., 2013, 2016; Daimon et al., 2014; Huang et al., 2016; Koutroumpa et al., 2016; Markert et al., 2016; Zhu et al., 2016).
To facilitate the construction of RIDL, we focus on the search of targeting genes, including lethal genes. In Drosophila, wingless also called Wnt Family Member 1 (Wnt-1), is associated with wing development (Sharma and Chopra, 1976). Wnt/β-catenin signalingis highly conserved in insects, can control cell fate and proliferation, and determine body plan in vertebrate embryos (Hikasa and Sokol, 2013). While Wnt/β-catenin signaling is required for segmentation during the early embryogenesis (Bolognesi et al., 2008; Petersen and Reddien, 2009; Fu et al., 2012), it also involves in the renewal of epithelial tissue (Sahai-Hernandez et al., 2012), antero-posterior brain patterning (Kobayashi et al., 2007), long-term memory (Tan Y. et al., 2013), neural plate and planarian regeneration (Niehrs, 2010) and head formation (Posnien et al., 2010). In Tribolium, Wnt signaling plays important roles in leg development during the embryonic stage, also involves in leg and wing regeneration, and in metamorphosis (Ober and Jockusch, 2006; Shah et al., 2011). In Lepidoptera, including Manduca sexta and B. mori, Wnt-1 contributes to the posterior growth and segmentation processes (Kraft and Jäckle, 1994; Zhang et al., 2015). In other species of vertebrate and invertebrate, Wnt-signaling genes are involved in the head morphogenesis and appendage development (Heisenberg et al., 2001; Müller et al., 2007; Lewis et al., 2008; Eroshkin et al., 2016).
The segmentation process involves multiple genes' interactions. In Drosophila, Wnt suppressed hedgehog (hh) and engrailed (en) expression in intercalary stripe and antennal stripe, but initiated en expression in ocular segment (Gallitano-Mendel and Finkelstein, 1997). A cephalic gap genes Orthodenticle (otd) represses wg expression in the antennal segment and all segments posterior to it, but activates wg expression in ocular segment (Gallitano-Mendel and Finkelstein, 1998). In Tribolium, complementary cross-regulation of Wnt and Hh pathways play an opposite interaction in the head and trunk development (Oberhofer et al., 2014). Knockout of Axin, a negative regulator of the Wnt pathway, led to missing head and thorax (Fu et al., 2012). A similar phenotype was obtained from Masterblind/Axin1 mutation, which showed smaller head and eyes in zebrafish (Heisenberg et al., 2001). In Xenopus laevis, Noggin4 regulates head development by inhibiting Wnt8 signaling (Eroshkin et al., 2016). In mouse, DKK (Dickkopf -related protein 1) as one of Wnt antagonists, is expressed anteriorly to repress Wnt signaling in the head (Lewis et al., 2008). In Hydractinia, activation of Wnt signaling by blocking GSK-3β(Glycogen Synthase Kinase 3β) affected regeneration, the patterning of growing polyps and the asexual formation of new polyps in the colony (Müller et al., 2007).
In this study, we explored CRISPR/Cas9-based genome editing in a major forest pest in China, the pine caterpillar moth, D. punctatus. Our molecular target, Wnt-1, is believed to be involved in the body plan in D. punctatus. To test this functional genomics tool, we first cloned the DpWnt-1, and then generated loss-of-function mutations through microinjection at the embryonic stage. The resultant phenotypic impacts of Wnt-1 knockout included lethality, abnormal segmentation and defective appendages. This proof-of-concept study using the CRISPR/Cas9-based genome editing tool demonstrates the feasibility of the genetic manipulation in a forest insect pest, which bears promising future advances in functional genomic research in forest entomology.
Materials and Methods
Gene Identification, Motif, and Phylogenetic Analyses
To search for the Wnt-1 homolog, nucleotide sequence of BmWnt-1 (NM_001043850.1) was used as a query to BLAST against a D. punctatus transcriptome (HHL, unpublished data). RACE was used to obtain the full length cDNA of DpWnt-1. The predicted open reading frame (ORF) was subjected to motif search, pattern analysis, and phylogenetic analysis. The MEME online server was used for motif analysis, and parameters were as follows: a minimum width was 6; a maximum width was 12; and a maximum number of motif was 8 (http://meme-suite.org/tools/meme). To understand the phylogenetic relationship of DpWnt-1 with homologs from other animals, a neighbor-joining tree was constructed using MEGA5, http://www.mega-software.net/ (Tamura et al., 2011). The Wnt-1 ORFs included in the analysis are as follows: B. mori (NM_001043850), H. armigera (KJ206240), Amyelois transitella (XM_013345048), P. xylostella (XM_011569928), M. sexta (Z30280), P. xuthus (XM_013325799), Danio rerio (XP_005162280), Fopius arisanus (XM_011300877), Bombus terrestris (XM_003393116), Nasonia vitripennis (XM_001603338), Bactrocera dorsalis (XM_011204079), Drosophila willistoni (XM_002066877), Drosophila melanogaster (NM_078778), Tribolium castaneum (EFA04660), Periplaneta americana (KC311252), Gryllus bimaculatus (BAB19660), Homo sapiens (NP_005421) and Mus musculus (NP_067254).
cDNA Cloning and Sequence Analysis
Total RNA was isolated with Trizol Reagent (Invitrogen, USA) from D. punctatus pupae. Recombinant DNase I-treated (Takara, Japan) RNA was used for cDNA synthesis with the Scientific Revert Aid First Strand cDNA synthesis kit (Thermo, USA). Diluted reverse transcription products were used as templates to amplify DNA fragments. The primer sets used to obtain the DpWnt-1 ORF are listed in Table 1. Template DNA was denatured at 94°C for 2 min, followed by 35 cycles of 94°C for 15 s, 55°C for 30 s and 68°C for 1.5 min. PCR products were cloned into the pCR-Blunt vector for sequencing by ABI 3730 XL sequencer (Applied. Biosystems, USA).
Quantitative Real-Time PCR (qRT-PCR) Analysis
qRT-PCR was performed to analyse the expression profile of DpWnt-1 and 8 Hox genes during the embryonic stage. cDNA samples were prepared from embryos of different developmental stages (day 1–day 8 of wild type) and the first instar larvae of DpWnt-1 mutants. Mastercycler EP realplex (Eppendorf) was used for the qRT-PCR. The primer sets used in qRT-PCR analysis are listed in Table 1. The cycling conditions were as follows: an initial incubation at 95°C for 10 s, 40 cycles of 95°C for 15 s, and 60°C for 30 s according to SYBR Green fluorescent relative quantitative approaches (TaKaRa, Japan). The relative mRNA level of the target genes was calculated using the 2−ΔΔCt method, in which the target gene expression was normalized to an internal reference, RP32. Three independent replications for each sample were performed.
In vitro Transcription of Cas9 and sgRNA
The Cas9 gene template used in this work was provided by View Solid Biotech (Beijing, China). Cas9 mRNA was synthesized in vitro with the mMESSAGE mMACHINE® T7 kit (Ambion, USA) according to the manufacturer's instructions.
For the in vitro transcription of sgRNA driven by the T7 promoter, target sequences start with GG. With the PAM sequences in consideration, the designed sgRNA sites follow the GGN19GG rule (Wang et al., 2013). We identified two 23 bp sgRNA targeting sites at exon III of DpWnt-1 (Figure 3A). The control sgRNAs were used for targeting the EGFP gene. Two complementary oligonucleotides were annealed and cloned into pJET1.2 (Fermentas, USA). The templates for in vitro transcription were amplified from pJET1.2, and primer sets used in this study are listed in Table 1. sgRNAs were transcribed in vitro with the MAXIscript® T7 kit (Ambion, USA), following the manufacturer's recommended protocol.
Colony Maintenance and Embryonic Microinjection
Dendrolimus punctatus pupae were originally obtained from Xing'an County of Guilin city, Guangxi province, P.R. China. D. punctatus colonies were provisioned with Masson's pine, and maintained at 27 ± 1°C under a L/D cycle of 16/8 h. Fertilized eggs were collected within 2 h after oviposition, and subjected to microinjection.
The combination of Cas9 mRNA (300 ng/μl) and sgRNAs (sgRNA-a and sgRNA-b, 300 ng/μl, respectively), and Cas9 mRNA/sgRNAs (sgRNA-a and sgRNA-b) (500 ng/μl each) were co-injected into preblastoderm embryos. An exogenous gene EGFP and nuclear free water without any sgRNAs or Cas9 mRNA were used as control. These control should have none effect on the embryonic development. Injection was carried out following Tamura et al. (1990) with modification, and injection site was shown in Figure S2. As the egg is oval in shape, we lined up the egg with the micropyle on top and injected compounds to the gonad region. The microinjection was concluded within 6 h. Afterwards, the injected eggs were incubated at 25 ± 1°C in a humidified chamber for 8–10 days until hatch. All hatched larvae were collected and transferred to Masson's pine.
Phenotype Documentation and Mutation Screening
The injected embryos were dissected and checked to calculate the mutation rate and hatching rate on the seventh day of the embryonic stage, and the resultant phenotypes were documented under a multi-function zoom microscope (AZ100, Nikon). The images were recorded with a computer-controlled microscope system. The pictures of DpWnt-1 mutants, including both larvae and pupae, were taken by SLR cameras.
To calculate the efficiency of Cas9/sgRNA-mediated gene alteration in the injected generation, individuals were collected on the eighth day after injection. The DNA fragments surrounding the sgRNA targets were obtained by GBdirect PCR directly from embryos (GBI, China). The primer sets are shown in Table 1. Mutations were confirmed by sequencing.
Immunoblotting Analysis
Proteins from 7 day old embryos were used for the immunoblotting analysis. The primary antibodies, B. mori Anti-Wnt-1 and Anti-β-actin, respectively, were used at 1:1000 dilution. The secondary antibody, anti-rabbit IgG, was diluted at 1:5000. Proteins were extracted and diluted with PBS and quantified using bicinchoninic acid (BCA) protein assay kit (Thermo). A 12.5% SDS-PAGE gel was used to separate the same amount of proteins from both the wild types and mutants. The proteins were then transferred to a polyvinylidene fluoride membrane. Signal visualization was obtained using the ECL Plus Western Blotting detection kit (GE Health-care).
Results
Expression Profile of DpWnt-1 during Embryogenesis
EST sequence of DpWnt-1 (GenBank accession #:KU640201) was initially obtained from D. punctatus transcriptome. The full length cDNAs of DpWnt-1 contained 1182 nucleotides, which encodes 394 amino acids. The nucleotide sequence of DpWnt-1 was rich in cysteine residues-a character of Wnt protein family (Figure S1). Wnt-1 homologs from 18 species shared eight conserved motifs, which located between the N- and C-terminus (Figure 1). Phylogenetic relationship showed that DpWNT-1 clustered with other lepidopterans WNT-1 protein sequences (Figure S2). The expression of DpWnt-1 peaked at the very beginning, declined during the development, and reached the minimum level at the end of embryogenesis (Figure 2), suggesting that DpWnt-1 may play a vital role in D. punctatus during the early embryogenesis.
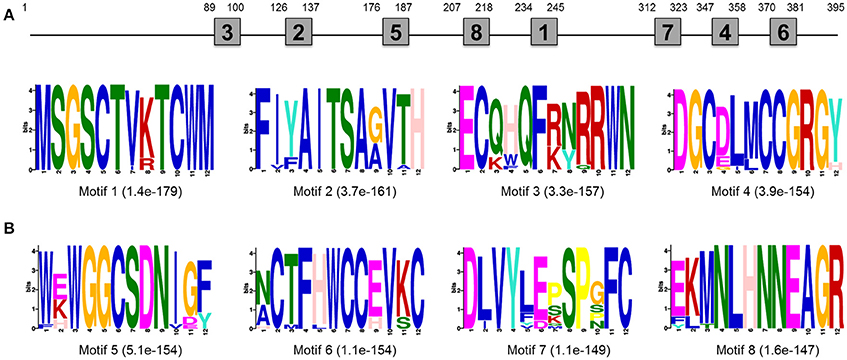
Figure 1. Motif analysis of Wnt-1 primary structure. (A) Approximate location of each motif in the protein sequence. (B) The most conserved motifs. The number in the boxes corresponds to the numbered motifs. The number in parentheses represents the e-values.
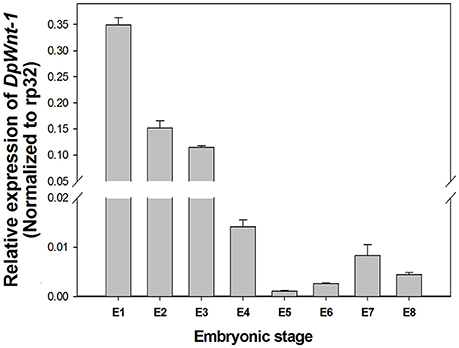
Figure 2. Temporal expression of DpWnt-1during embryonic stages. The relative mRNA levels of DpWnt-1 in embryos from day 1 to 8 (E1-8). RP32 was used as a reference gene to normalize target gene expression. The data are presented as mean values ± S.E.M (n = 3).
CRISPR/Cas9 Induced DpWnt-1 Mutations
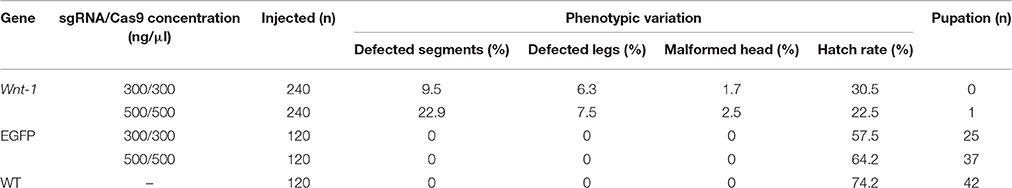
To functionally characterize DpWnt-1, CRISPR/Cas9 mutagenesis system was introduced into D. punctatus. A total of 240 D. punctatus eggs were co-injected for each concentration of Cas9 mRNA and DpWnt-1 sgRNAs, whereas 120 eggs were injected for the corresponding concentrations for the control EGFP sgRNAs (Table 2). Compared to co-injections of Cas9 protein and DpWnt-1 guide RNAs with those targeting a control gene (EGFP), D. punctatus embryos with an inactive copy of Wnt-1showed a reduced hatching rate (22.5 and 30.5% at a concentration of 500 and 300 ng/μl, respectively), and a range of phenotypic effects (e.g., various body plan defects, absence of tissue differentiation). Among the 120 control eggs injected with EGFP sgRNAs/Cas9 mRNA, 57.5 and 64.2% individuals hatched at a concentration of 500 and 300 ng/μl, respectively. In comparison, 65.8% (79/120) wild type eggs hatched.
CRISPR/Cas9 system induced mutations in the pine moth with high efficiency. Eighty percentage (8 of 10) of the dissected embryos had mutations at the target sites, and the overall mutagenesis frequency was 32.9% in the injected generation at a higher dose (500 ng/μl). Similarly, at a lower dosage (300 ng/μl), 70% (7 of 10) of the dissected embryos had mutations at the target sites and the overall mutagenesis frequency was ~17.5% (Table 2). The genotypes of the wild types and DpWnt-1 mutants were confirmed by both sequencing and Western blotting analysis (Figures 3B,C). All examined DpWnt-1 mutants, including embryos and larvae, had alterations at the target sites that led to at least five type of deletions (Figure 3D). The deletion occurred at target sites individually, simultaneously, or was absent from both sites.
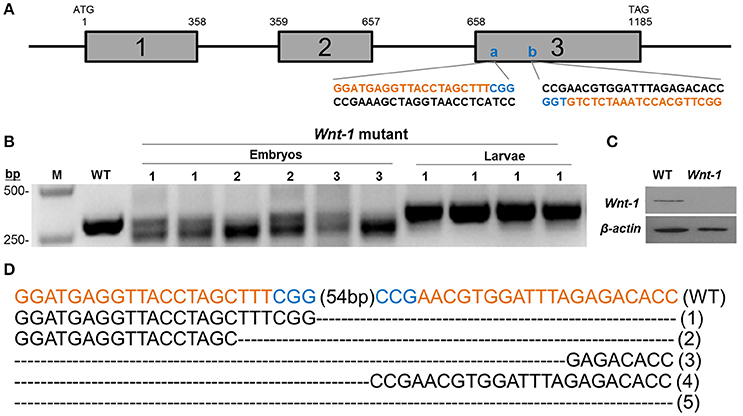
Figure 3. Cas9/sgRNA-induced DpWnt-1 mutations. (A) Schematic representation of Wnt-1 sgRNA targeting sites. The boxes indicate the three deduced exons of DpWnt-1, and the black line represents the untranslated regions and introns. The sgRNA targeting sites, (A) (74–96 bp) and (B) (151–173 bp), are located on exon 3. Wnt-1-F and Wnt-1-R were annealed to the upstream and downstream regions of the targeted site. (B–D) CRISPR/Cas9-induced mutagenesis of DpWnt-1. (B) Representative electrophoretogram of PCR products. Mutants with defective segments (1), defective legs (2), and malformed head (3) were sequenced. (C) DpWnt-1 protein was undetectable in mutants by Western blotting analysis. (D) Various deletion genotypes. The fragment flanking the two targeted sites were deleted. The indel mutation genotype is noted on the right.
Functional Characterization of DpWnt-1
Knocking out DpWnt-1 has great impact on eggs development. Most of eggs showed abdominal segments distortion and only some of them could hatch and develop into pupae, of which none reached the adult stage. When injected with 500 ng/μl of Cas9 mRNA and DpWnt-1 sgRNA, 22.9% of the embryos showed abnormal anterior-posterior (A-P) axis and abdominal segmentation phenotypes, 7.5% showed defective legs, and 2.5% showed head malformations. In contrast, when the injection concentration is 300 ng/μl, 9.5% of embryos showed abnormal A-P axis and abdominal segmentation phenotypes, 6.3% showed defective legs, and 1.7% showed head malformations. As a control, 240 eggs were co-injected with EGFP-sgRNA/Cas9 mRNA. A total of 146 eggs (60.8%) hatched, and no morphological changes were observed (Table 2).
Patterning of the Posterior Segment from Embryo to Pupa
DpWnt-1 knockout led to visible abnormal abdominal formation phenotypes and abnormal patterning of the A-P axis (Figures 4–6). Some of the embryos showed the anteriorization of segments A2/7 (Figure 4). In some mutants, the loss of DpWnt-1 led to the transformation of segments A2–A6 into more anterior abdominal segments (Figure 5). Some embryos showed a loss of epithelia on the dorsal side of the A3/5 segments, which was close to the intersegmental membrane and the dorsal mid line (Figures 4I,J). In other mutants, the boundaries between the abdominal segments and the anteroposterior body axis were discreet, as all of the abdominal segments (A2–A7) were fused together (Figure 5), indicating that DpWnt-1 plays a role in posterior segmentation and A-P axis patterning. During the development, DpWnt-1 mutants retained the posterior segment fusion and the truncated cuticle phenotypes and were unable to form posterior segments in a specific region (Figure 4).
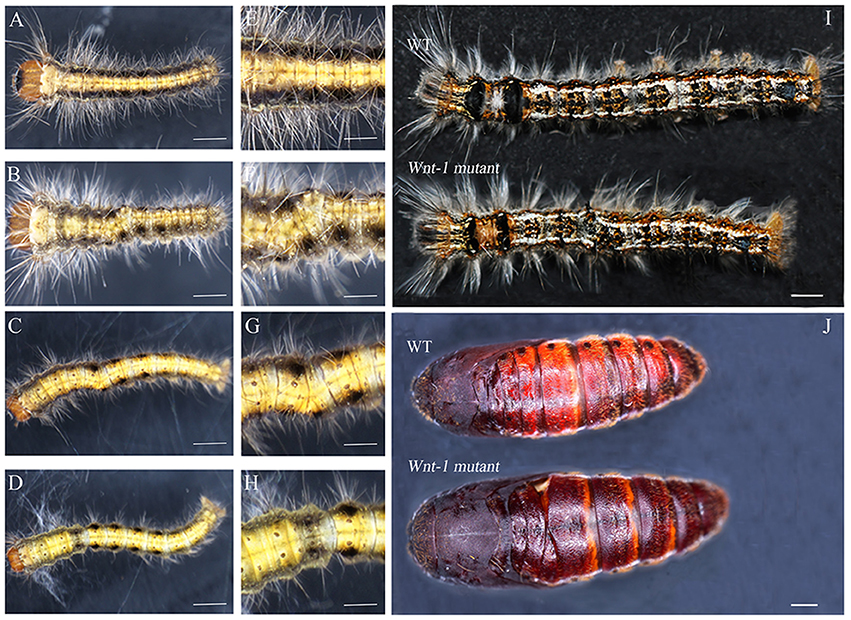

Figure 4. Cas9/sgRNA-induced posterior segment defects in D. punctatus larvae and pupae. (A,E) EGFP-specific sgRNAs/Cas9 mRNA control. (B–D,F–H) Mildly affected larvae resulting from DpWnt-1 sgRNAs/Cas9 mRNA co-injection. Transformation of the abdominal segment from posterior to anterior. (I) Fifth instar larvae, wild type (up) and DpWnt-1 mutant (down), displaying the transformation of A6/7 into A6. (J) Wild type and DpWnt-1 mutant pupae. (B,F) The mutant larvae type I showed a transformation of A3/5 into A3 and a disturbance of the anterior-posterior axis. (C,G) The mutant larvae type II showed a transformation of A2/4 into A3 and a disturbance of the anterior-posterior axis. (D,H) The mutant larvae type III has extra pigmentation at A2. (E–H) Close-up images of the wild type and mutant individuals. The scale bars represent 0.5 mm (A–D), 0.25 mm (E–H), 50.0 mm (I), and 2.0 mm (J).
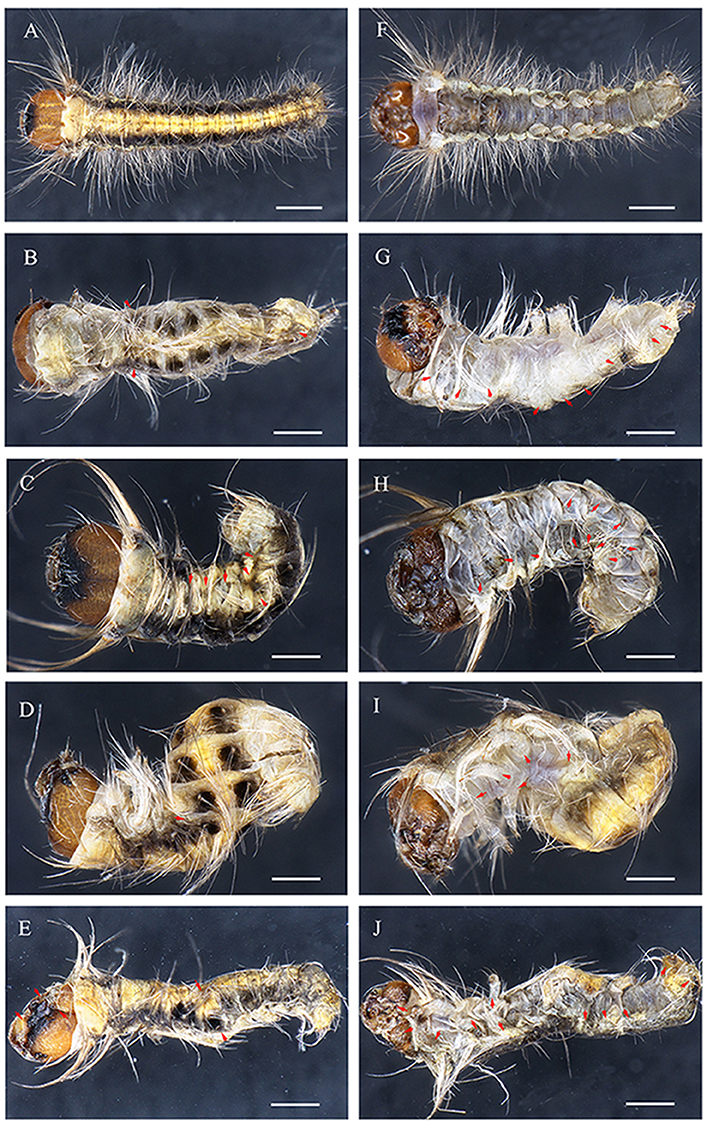
Figure 5. Embryonic phenotypes in D. punctatus. (A,F) EGFP sgRNAs/Cas9 mRNA injected control embryo. (B–E,G–J) Severely affected embryo resulting from DpWnt-1 sgRNAs/Cas9 mRNA injection. (B,G) Thoracic leg and prolegs missing on one side. (C,H) Compact body with thoracic legs and prolegs missing on both sides. (D,I) Twisted body without thoracic legs or patterning along anterior and posterior axis, with all prolegs missing. (E,J) Deformed body with malformed head, missing thoracic legs and prolegs on one side. All images were taken at the same magnification. Dorsal is on left and ventral is on right. The scale bars represent 1 mm.
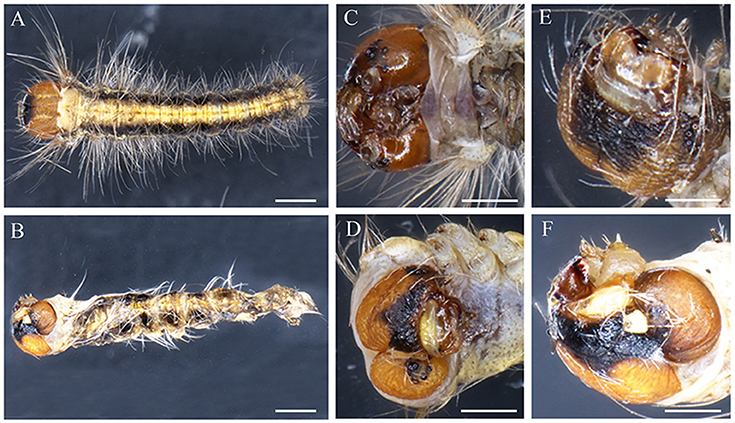

Figure 6. Head phenotypes of DpWnt-1 mutants. (A,C,E) Wild type embryo. (B,D,F) Severely affected embryo with malformed head, missing thoracic legs and prolegs on both sides. The scale bars represent 1 mm.
Anterior Body Development
DpWnt-1 signaling plays a crucial role in the development of the anterior segments in D. punctatus. DpWnt-1 mutant larvae had missing appendages and displayed asymmetric anterior segment phenotypes (Figures 5, 6). In the wild type, the ecdysial line is localized in the middle of the head, and the lateral ocelli and antennae are located on both sides of the head (Figure 5A). In comparison with wild type larvae, partial lateral ocelli, antennae and intercalary were missing on the head of DpWnt-1 mutants, while other mutants showed defective mouthparts with mandibular, maxillary and labial missing (Figures 5E,J, 6B–F).
Leg Patterning
DpWnt-1 is involved in the leg development, specifically on thoracic segments (T1–T3) and abdominal segments (A3–A6). The wild type embryo had three pairs of thoracic legs from the first to third thoracic segments and four pairs of prolegs from the third to sixth abdominal segments. In the type I mutant, some of the T1–T3 and A3–A6 segments were missing, and thoracic legs and prolegs were on one side of the segments (Figures 5B,G). In the type II mutant, some of the T1–T3 and A3–A6 segments were missing, and thoracic legs and prolegs were on both sides of the segments (Figures 5C,D,H,I). In the type III mutant, the legs on the T1–T3 thoracic segments did not follow the principle of symmetry and showed an asymmetrical distribution along the A-P axis. Moreover, the A3–A6 prolegs were missing on both sides of the segments (Figures 5E,J).
Pleiotropic Impact of DpWnt-1 Knockout
The distinct phenotypes exhibited in DpWnt-1 mutants suggested that DpWnt-1 may participate in segmentation. Hox genes are known to be involved in segmentation. qRT-PCR analysis in 8-day old DpWnt-1 mutant and wild type embryos results showed that Sex combs reduced (Scr), Deformed (Dfd), and Abdominal-b (Abd-b) were significantly upregulated while Ultrabithorax (Ubx) was downregulated in DpWnt-1 mutants. The DpWnt-1 mutants also showed slightly reduced expression levels of Labial (Lab), Abdominal-a (Abd-a), and Antennapedia (Antp), whereas Proboscipedia (Pb) was undetectable (Figure 7).
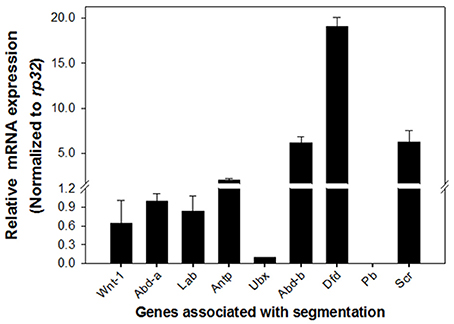
Figure 7. Expression profiling in DpWnt-1 mutants. Compared to the controls, the mRNA expression of Sex combs reduced (Scr), Deformed (Dfd), and Abdominal-b (Abd-b) increased more than 4-fold in the DpWnt-1 mutants. Others, including Labial (Lab), Proboscipedia (Pb), Antennapedia (Antp), Ultrabithorax (Ubx), and Abdominal-a (Abd-a), changed <2-fold. Rp32 was used as reference gene for RT-PCR normalization. The data are presented as mean values ± S.E.M (n = 3).
Discussion
Characteristics of Wnt-1 Homolog
Understanding the function of Wnt-1 is critical for exploring its potential role in pest management. In this study, we cloned and characterized DpWnt-1 homolog and identified one Wnt-1 gene in D. punctatus, DpWnt-1. The motif and phylogenetic analyses confirmed that DpWnt-1 is most closely related to BmWnt-1 (Dhawan and Gopinathan, 2003).
In Drosophila, with long germ embryos, Wnt-1 expression was first detected in the whole segments of the blastoderm during cellularization (Baker, 1987; Vorwald-Denholtz and De Robertis, 2011). In Tribolium, with short-germ embryos, Wnt-1 was initially detected in the blastoderm stage, expressed sequentially from anterior to posterior with the germ band elongation and at the ventral portion of each segment during the late embryonic stage (Nagy and Carroll, 1994). In short/intermediate germ embryos, Wnt-1 was detected in a broad median of the germ disk and then retracted posteriorly within segmentation process (Nakao, 2010). The expression pattern of DpWnt-1 during the embryonic stage showed the same trend with that of Bombyx (Zhang et al., 2015). BmWnt-1 was present in a maternal gradient and might play a role during the blastoderm formation (Nakao, 2010; Zhang et al., 2015). We hypothesized that D. punctatus may have a short/intermediate germ band, in which segmentation proceeds consecutively from anterior to posterior and show visible anterior and posterior segments after gastrulation.
CRISPR/Cas 9 System in D. punctatus
In this study, embryonic injection of a mixture of sgRNAs/Cas9 mRNA successfully induced mutations in DpWnt-1, demonstrating that CRISPR/Cas9-mediated genome editing can specifically and efficiently induce gene alterations in D. punctatus. Besides D. punctatus, CRISPR/Cas9 system has also been exploited in seven other Lepidoptera species, including B. mori, S. litura, S. littoralis, P. xylostella, P. xuthus, H. armigera, and D. plexippus, to manipulate genes associated with development (embryogenesis), pigmentation, metamorphosis, resistance mechanism, and adult mating (Wang et al., 2013, 2016; Li et al., 2015; Bi et al., 2016; Huang et al., 2016; Koutroumpa et al., 2016; Markert et al., 2016; Zhu et al., 2016). Moreover, the frequency of mutation is dose dependent. Knocking out DpWnt-1 led to a high embryonic mortality (~70%), and none of the DpWnt-1 mutants could developed from larva to adult, suggesting that DpWnt-1 is a potential candidate for conditional lethal gene.
Although CRISPR/Cas9 system is clearly applicable in Dendrolimus, additional experiments are needed to fully established this genome editing technology in this major forest pest. In situ hybridization study of DpWnt-1 not only will validate genome editing results at the translational level, but also provide the spatial expression pattern, and the potential Hox targets. Also, without genome information, we could not pinpoint the off-target effects, which is a routine problem for this technology. With other genomic resources (Yang et al., 2016), the potential off-target effects can be predicted.
Involvement of DpWnt-1 in Segmentation and Appendage Development
DpWnt-1 in Posterior Segmentation
Wnt-1 has been documented to play an important role in A-P axis patterning and segment development during embryogenesis. In DpWnt-1 mutants, abnormal posterior segments from Abdomen 2 (A2) to Abdomen 7 (A7) were observed along with affected A-P axis patterning. An examination of Hox genes in Wnt-1 mutants suggested that DpWnt-1 may have a connection with Hox genes in regulating insect segmentation. Our results for the function of DpWnt-1 are consistent with those of Bombyx, in which DpWnt-1 plays a role in body segmentation. However, Wnt-1 appears to have a different effect on the expression of other genes, as all Hox genes were significantly down-regulated in Bombyx (Zhang et al., 2015). Consistent with Drosophila, Wingless signaling ensures the formation of the posterior segment boundaries (Larsen et al., 2003). However, depletion of Wnt-1 in G. bimaculatus, Oncopeltus fasiatus, and Tribolium, does not reduce the number of segments, but depletion of other Wnt signaling genes like GbArm leads to abdominal segments defects in embryos, removal of OfPan results in truncates segmentation, depleting of TcWnt-8 brings about embryos lacking abdominal segments and additional removal of TcWnt-1 enhances this phenotype (Miyawaki et al., 2004; Angelini and Kaufman, 2005; Shah et al., 2011). All of these results indicate that DpWnt-1 plays a role in segmentation in D. punctatus.
DpWnt-1 in Anterior Segmentation
The genetic regulation of the anterior development in insects is poorly understood. According to Rogers and Kaufman (1996), head was divided into three cephalic segments (ocular, antennal, and intercalary) and three gnathal segments (mandibular, maxillary, and labial). In animals, Wnt-1 is involved in the head development, including eyes, mesencephalon and metencephalon (Bally-Cuif et al., 1995; Friedrich, 2003; Lekven et al., 2003; Rossi et al., 2007). In D. melanogaster, temporal regulation of Wnt signaling is critical for the differentiation of antennal and maxillary organs (Lebreton et al., 2008). In Tribolium, Wnt/β-catenin signaling is required for the anterior development, which is needed for head patterning after cellularization (Bolognesi et al., 2008; Fu et al., 2012; Benton et al., 2013). Consistent with previous observations, both anterior and posterior sequential segmentation were affected in DpWnt-1 mutants. Besides, partial cephalic segments and gnathal segments of the mutants were missing or defected. These results support the hypothesis that Wnt signaling is an integral part of an ancestral metazoan mechanism that specify the architecture of posterior and anterior segments.
DpWnt-1 in Appendage Development
The morphological plasticity of appendages represents a crucial aspect of animal body plan. Knocking out DpWnt-1 produced defects in appendage development. No discernible defects in the appendages were found in mildly affected individuals (Figure 5). In severely affected individuals, however, lateral ocelli, antennae, the thoracic legs and prolegs were missing (Figure 6). Among these mutants, some thoracic legs or prolegs were distributed asymmetrically along the normal AP axis (Figure 6), suggesting that the specification of appendages in Dendrolimus requires DpWnt-1. Some of the defects, such as the loss of prolegs could be the indirect consequences of segmentation defects. Consistent with other holometabolous taxa, including Coleoptera, Lepidoptera, Hymenoptera and Diptera, Wnt-1 signaling is involved in post-embryonic appendage development (Bejsovec and Peifer, 1992; Siegfried et al., 1994; Sato et al., 2008; Shah et al., 2011; Zhang et al., 2015). This is different from taxa that undergo incomplete metamorphosis, of which appendage development requires Wnt-1 to interact with other genes, such as in G. bimaculatus (Miyawaki et al., 2004). Although Gbwg knockouts by RNAi showed no significant impacts on segmentation, GbWnt/GbArm signaling was involved in the posterior sequential segmentation during embryogenesis. In P. americana, Wnt signaling engaged in cross talk with caudal and Notch signaling in the regulation of growth and segmentation (Chesebro et al., 2013). In O. fasiatus, Wnt signaling played a role in body segmentation but not in appendage development (Angelini and Kaufman, 2005). Based on these results, we propose that the function of Wnt signaling is conserved among insects even though Wnt-1 gene has diverse functions in different species.
In summary, our study demonstrates that genome editing using CRISPR/Cas9 system is feasible in Dendrolimus. This provides a brand new tool for conducting functional genomic research in a major forest pest. Furthermore, the results from the functional characterization of DpWnt-1 demonstrated that this gene could potentially be utilized as a specific lethal gene in RIDL. CRISPR/Cas9 system could also be used to create transgenic lines to screen for dominant suppressors driven by specific promoters to provide candidate genes for the control of Dendrolimus.
Author Contributions
HL designed and conceived the study. XZ, HL analyzed the data. HL, XZ, YH, ZZ, and QL wrote the manuscript. All authors approved the final version of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This project was supported by strategic Priority Research Program of the Chinese Academy of Sciences (NO. XDB11010600) and a special fund for Forest Scientific Research in the Public Welfare (201504302). We would like to thank Lang You for his assistance with the micro-injection, and Rongmei Chen for colony maintenance. Special thanks go to Jun Xu, Zhongjie Zhang, and Baosheng Zeng for their comments on an earlier draft.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fphys.2016.00666/full#supplementary-material
References
Alphey, L., and Andreasen, M. (2002). Dominant lethality and insect population control. Mol. Biochem. Parasitol. 12, 173–178. doi: 10.1016/S0166-6851(02)00040-3
Alphey, L., Beard, C. B., Billingsley, P., Coetzee, M., Crisanti, A., Curtis, C., et al. (2002). Malaria control with genetically manipulated insect vectors. Science 298, 119–121. doi: 10.1126/science.1078278
Angelini, D. R., and Kaufman, T. C. (2005). Functional analyses in the milkweed bug Oncopeltus fasciatus (Hemiptera) support a role for Wnt signaling in body segmentation but not appendage development. Dev. Biol. 283, 409–423. doi: 10.1016/j.ydbio.2005.04.034
Baker, E. N. (1987). Molecular cloning of sequences from wingless, a segment polarity gene in Drosophila: the spatial distribution of a transcript in embryos. EMBO. J. 6, 1765–1773.
Bally-Cuif, L., Cholley, B., and Wassef, M. (1995). Involvement of Wnt-1 in the formation of the mes/metencephalic boundary. Mech. Dev. 53, 23–34. doi: 10.1016/0925-4773(95)00421-1
Bejsovec, A., and Peifer, M. (1992). Knowing your neighbors: cell interactions determine intrasegmental patterning in Drosophila. Trends Genet. 8, 243–249. doi: 10.1016/0168-9525(92)90394-J
Benedict, M. Q., and Robinson, A. S. (2003). The first releases of transgenic mosquitoes: an argument for the sterile insect technique. Trends Parasitol. 19, 349–355. doi: 10.1016/S1471-4922(03)00144-2
Benton, M. A., Akam, M., and Pavlopoulos, A. (2013). Cell and tissue dynamics during Tribolium embryogenesis revealed by versatile fluorescence labeling approaches. Development 140, 3210–3220. doi: 10.1242/dev.096271
Bettencourt, R., Terenius, O., and Faye, I. (2002). Hemolin gene silencing by dsRNA injected into Cecropia pupae is lethal to next generation embryos. Insect Mol. Biol. 11, 267–271. doi: 10.1046/j.1365-2583.2002.00334.x
Bi, H. L., Xu, J., Tan, A. J., and Huang, Y. P. (2016). CRISPR/Cas9-mediated targeted gene mutagenesis in Spodoptera litura. Insect Sci. 23, 69–77. doi: 10.1111/1744-7917.12341
Billings, R. F. (1991). The pine caterpillar Dendrolimus punctatus in Viet Nam Recommendations for integrated pest management. For. Ecol. Manage. 39, 97–106. doi: 10.1016/0378-1127(91)90167-T
Bolognesi, R., Farzana, L., Fischer, T. D., and Brown, S. J. (2008). Multiple Wnt genes are required for segmentation in the short-germ embryo of Tribolium castaneum. Curr. Biol. 18, 1624–1629. doi: 10.1016/j.cub.2008.09.057
Chesebro, J. E., Pueyo, J. I., and Couso, J. P. (2013). Interplay between a Wnt-dependent organiser and the Notch segmentation clock regulates posterior development in Periplaneta americana. Biol. Open 2, 227–237. doi: 10.1242/bio.20123699
Corrigan-Curay, J., O'Reilly, M., Kohn, D. B., Cannon, P. M., Bao, G., Bushman, F. D., et al. (2015). Genome editing technologies: defining a path to clinic. Mol. Ther. 23, 796–806. doi: 10.1038/mt.2015.54
Daimon, T., Kiuchi, T., and Takasu, Y. (2014). Recent progress in genome engineering techniques in the silkworm, Bombyx mori. Dev. Growth Differ. 56, 14–25. doi: 10.1111/dgd.12096
Dhawan, S., and Gopinathan, K. P. (2003). Spatio-temporal expression of wnt-1 during embryonic- wing- and silkgland development in Bombyx mori. Gene Expr. Patterns 3, 559–570. doi: 10.1016/S1567-133X(03)00122-4
Eroshkin, F. M., Nesterenko, A. M., Borodulin, A. V., Martynova, N. Y., Ermakova, G. V., Gyoeva, F. K., et al. (2016). Noggin4 is a long-range inhibitor of Wnt8 signaling that regulates head development in Xenopus laevis. Sci. Rep. 6:23049. doi: 10.1038/srep23049
Friedrich, M. (2003). Evolution of insect eye development: first insights from fruit fly, grasshopper and flour beetle. Integr. Comp. Biol. 43, 508–521. doi: 10.1093/icb/43.4.508
Fu, G., Condon, K. C., Epton, M. J., Gong, P., Jin, L., Condon, G. C., et al. (2007). Female-specific insect lethality engineered using alternative splicing. Nat. Biotechnol. 25, 353–357. doi: 10.1038/nbt1283
Fu, J., Posnien, N., Bolognesi, R., Fischer, T. D., Rayl, P., Oberhofer, G., et al. (2012). Asymmetrically expressed axin required for anterior development in Tribolium. Proc. Natl. Acad. Sci. U.S.A. 109, 7782–7786. doi: 10.1073/pnas.1116641109
Gallitano-Mendel, A., and Finkelstein, R. (1997). Novel segment polarity gene interactions during embryonic head development in Drosophila. Dev. Biol. 192, 599–613. doi: 10.1006/dbio.1997.8753
Gallitano-Mendel, A., and Finkelstein, R. (1998). Ectopic orthodenticle expression alters segment polarity gene expression but not head segment identity in the Drosophila Embryo. Dev. Biol. 199, 125–137. doi: 10.1006/dbio.1998.8917
Hammond, A., Galizi, R., Kyrou, K., Simoni, A., Siniscalchi, C., Katsanos, D., et al. (2016). A CRISPR-Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nat. Biotechnol. 34, 78–83. doi: 10.1038/nbt.3439
Harrison, M. M., Jenkins, B. V., O'Connor-Giles, K. M., and Wildonger, J. (2014). A CRISPR view of development. Genes Dev. 28, 1859–1872. doi: 10.1101/gad.248252.114
Heinrich, J. C., and Scott, M. J. (2000). A repressible female-specific lethal genetic system for making transgenic insect strains suitable for a sterile-release program. Proc. Natl. Acad. Sci. U.S.A. 97, 8229–8232. doi: 10.1073/pnas.140142697
Heisenberg, C. P., Houart, C., Take-Uchi, M., Rauch, G. J., Young, N., Coutinho, P., et al. (2001). A mutation in the Gsk3-binding domain of zebrafish Masterblind/Axin1 leads to a fate transformation of telencephalon and eyes to diencephalon. Genes Dev. 15, 1427–1434. doi: 10.1101/gad.194301
Hikasa, H., and Sokol, S. Y. (2013). Wnt signaling in vertebrate axis specification. Cold Spring Harb. Perspect. Biol. 5:a007955. doi: 10.1101/cshperspect.a007955
Horn, C., and Wimmer, E. A. (2003). A transgene-based, embryo-specific lethality system for insect pest management. Nat. Biotechnol. 21, 64–70. doi: 10.1038/nbt769
Huang, Y., Chen, Y., Zeng, B., Wang, Y., James, A. A., Gurr, G. M., et al. (2016). CRISPR/Cas9 mediated knockout of the abdominal-A homeotic gene in the global pest, diamondback moth (Plutella xylostella). Insect Biochem. Mol. Biol. 75, 98–106. doi: 10.1016/j.ibmb.2016.06.004
Kobayashi, C., Saito, Y., Ogawa, K., and Agata, K. (2007). Wnt signaling is required for antero-posterior patterning of the planarian brain. Dev. Biol. 306, 714–724. doi: 10.1016/j.ydbio.2007.04.010
Koutroumpa, F. A., Monsempes, C., François, M. C., de Cian, A., Royer, C., Concordet, J. P., et al. (2016). Heritable genome editing with CRISPR/Cas9 induces anosmia in a crop pest moth. Sci. Rep. 6:29620. doi: 10.1038/srep29620
Kraft, R., and Jäckle, H. (1994). Drosophila mode of metamerization in the embryogenesis of the lepidopteran insect Manduca sexta. Proc. Natl. Acad. Sci. U.S.A. 91, 6634–6638. doi: 10.1073/pnas.91.14.6634
Larsen, C. W., Hirst, E., Alexandre, C., and Vincent, J. P. (2003). Segment boundary formation in Drosophila embryos. Development 130, 5625–5635. doi: 10.1242/dev.00867
Lebreton, G., Faucher, C., Cribbs, D. L., and Benassayag, C. (2008). Timing of Wingless signaling distinguishes maxillary and antennal identities in Drosophila melanogaster. Development 135, 2301–2309. doi: 10.1242/dev.017053
Lekven, A. C., Buckles, G. R., Kostakis, N., and Moon, R. T. (2003). Wnt1 and wnt10b function redundantly at the zebrafish midbrain–hindbrain boundary. Dev. Biol. 254, 172–187. doi: 10.1016/S0012-1606(02)00044-1
Lewis, S. L., Khoo, P. L., De Young, R. A., Steiner, K., Wilcock, C., Mukhopadhyay, M., et al. (2008). Dkk1 and Wnt3 interact to control head morphogenesis in the mouse. Development 135, 1791–1801. doi: 10.1242/dev.018853
Li, X., Fan, D., Zhang, W., Liu, G., Zhang, L., Zhao, L., et al. (2015). Outbred genome sequencing and CRISPR/Cas9 gene editing in butterflies. Nat. Commun. 6, 8212. doi: 10.1038/ncomms9212
Ma, S., Shi, R., Wang, X., Liu, Y., Chang, J., Gao, J., et al. (2014). Genome editing of BmFib-H gene provides an empty Bombyx mori silk gland for a highly efficient bioreactor. Sci. Rep. 4:6867. doi: 10.1038/srep06867
Mao, Y. B., Xue, X. Y., Tao, X. Y., Yang, C. Q., Wang, L. J., and Chen, X. Y. (2013). Cysteine protease enhances plant-mediated bollworm RNA interference. Plant Mol. Biol. 83, 119–129. doi: 10.1007/s11103-013-0030-7
Markert, M. J., Zhang, Y., Enuameh, M. S., Reppert, S. M., Wolfe, S. A., and Merlin, C. (2016). Genomic access to monarch migration using TALEN and CRISPR/Cas9-Mediated targeted mutagenesis. G3 (Bethesda) 6, 905–915. doi: 10.1534/g3.116.027029
Miyawaki, K., Mito, T., Sarashina, I., Zhang, H., Shinmyo, Y., Ohuchi, H., et al. (2004). Involvement of Wingless/Armadillo signaling in the posterior sequential segmentation in the cricket, Gryllus bimaculatus (Orthoptera), as revealed by RNAi analysis. Mech. Dev. 121, 119–130. doi: 10.1016/j.mod.2004.01.002
Müller, W., Frank, U., Teo, R., Mokady, O., Guette, C., and Plickert, G. (2007). Wnt signaling in hydroid development: ectopic heads and giant buds induced by GSK-3beta inhibitors. Int. J. Dev. Biol. 51, 211–220. doi: 10.1387/ijdb.062247wm
Nagy, L. M., and Carroll, S. (1994). Conservation of wingless patterning functions in the short-germ embryos of Tribolium castaneum. Nature 367, 460–463. doi: 10.1038/367460a0
Nakao, H. (2010). Characterization of Bombyx embryo segmentation process: expression profiles of engrailed, even-skipped, caudal, and wnt1/wingless homologues. J. Exp. Zool. B Mol. Dev. Evol. 314, 224–231. doi: 10.1002/jez.b.21328
Niehrs, C. (2010). On growth and form: a Cartesian coordinate system of Wnt and BMP signaling specifies bilaterian body axes. Development 137, 845–857. doi: 10.1242/dev.039651
Ober, K. A., and Jockusch, E. L. (2006). The roles of wingless and decapentaplegic in axis and appendage development in the red flour beetle, Tribolium castaneum. Dev. Biol. 294, 391–405. doi: 10.1016/j.ydbio.2006.02.053
Oberhofer, G., Grossmann, D., Siemanowski, J. L., Beissbarth, T., and Bucher, G. (2014). Wnt/beta-catenin signaling integrates patterning and metabolism of the insect growth zone. Development 141, 4740–4750. doi: 10.1242/dev.112797
Petersen, C. P., and Reddien, P. W. (2009). Wnt signaling and the polarity of the primary body axis. Cell 139, 1056–1068. doi: 10.1016/j.cell.2009.11.035
Posnien, N., Schinko, J. B., Kittelmann, S., and Bucher, G. (2010). Genetics, development and composition of the insect head-a beetle's view. Arthropod Struct. Dev. 39, 399–410. doi: 10.1016/j.asd.2010.08.002
Rogers, B. T., and Kaufman, T. C. (1996). Structure of the insect head as revealed by the EN protein pattern in developing embryos. Development 122, 3419–3432.
Rossi, E., Siwiec, F., and Yan, C. Y. (2007). Pattern of Wnt ligand expression during chick eye development. Braz. J. Med. Biol. Res. 40, 1333–1338. doi: 10.1590/S0100-879X2006005000155
Sahai-Hernandez, P., Castanieto, A., and Nystul, T. G. (2012). Drosophila models of epithelial stem cells and their niches. Wiley Interdiscip. Rev. Dev. Biol. 1, 447–457. doi: 10.1002/wdev.36
Sato, K., Matsunaga, T. M., Futahashi, R., Kojima, T., Mita, K., Banno, Y., et al. (2008). Positional cloning of a Bombyx wingless locus flugellos (fl) reveals a crucial role for fringe that is specific for wing morphogenesis. Genetics 179, 875–885. doi: 10.1534/genetics.107.082784
Shah, M. V., Namigai, E. K., and Suzuki, Y. (2011). The role of canonical Wnt signaling in leg regeneration and metamorphosis in the red flour beetle Tribolium castaneum. Mech. Dev. 128, 342–358. doi: 10.1016/j.mod.2011.07.001
Sharma, R. P., and Chopra, V. L. (1976). Effect of the Wingless (wg1) mutation on wing and haltere development in Drosophila melanogaster. Dev. Biol. 48, 461–465. doi: 10.1016/0012-1606(76)90108-1
Siegfried, E., Wilder, E. L., and Perrimon, N. (1994). Components of wingless signaling in Drosophila. Nature 367, 76–80. doi: 10.1038/367076a0
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., and Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739. doi: 10.1093/molbev/msr121
Tamura, T., Kanda, T., Takiya, S., Okano, K., and Maekawa, H. (1990). Transient expression of chimeric CAT genes injected into early embryos of the domesticated silkworm Bombyx mori. Jpn. J. Genet. 65, 401–410. doi: 10.1266/jjg.65.401
Tan, A., Fu, G., Jin, L., Guo, Q., Li, Z., Niu, B., et al. (2013). Transgene-based, female-specific lethality system for genetic sexing of the silkworm, Bombyx mori. Proc. Natl. Acad. Sci. U.S.A. 110, 6766–6770. doi: 10.1073/pnas.1221700110
Tan, Y., Yu, D., Busto, G. U., Wilson, C., and Davis, R. L. (2013). Wnt signaling is required for long-term memory formation. Cell Rep. 4, 1082–1089. doi: 10.1016/j.celrep.2013.08.007
Terenius, O., Papanicolaou, A., Garbutt, J. S., Eleftherianos, I., Huvenne, H., Kanginakudru, S., et al. (2011). RNA interference in Lepidoptera: an overview of successful and unsuccessful studies and implications for experimental design. J. Insect. Physiol. 57, 231–245. doi: 10.1016/j.jinsphys.2010.11.006
Thomas, D. D., Donnelly, C. A., Wood, R. J., and Alphey, L. S. (2000). Insect population control using a dominant, repressible, lethal genetic system. Science 287, 2474–2476. doi: 10.1126/science.287.5462.2474
Vorwald-Denholtz, P. P., and De Robertis, E. M. (2011). Temporal pattern of the posterior expression of Wingless in Drosophila blastoderm. Gene Expr. Patterns 11, 456–463. doi: 10.1016/j.gep.2011.07.004
Wang, J., Zhang, H., Wang, H., Zhao, S., Zuo, Y., Yang, Y., et al. (2016). Functional validation of cadherin as a receptor of Bt toxin Cry1Ac in Helicoverpa armigera utilizing the CRISPR/Cas9 system. Insect Biochem. Mol. Biol. 76, 11–17. doi: 10.1016/j.ibmb.2016.06.008
Wang, Y., Li, Z., Xu, J., Zeng, B., Ling, L., You, L., et al. (2013). The CRISPR/Cas system mediates efficient genome engineering in Bombyx mori. Cell Res. 23, 1414–1416. doi: 10.1038/cr.2013.146
Windbichler, N., Papathanos, P. A., and Crisanti, A. (2008). Targeting the X chromosome during spermatogenesis induces Y chromosome transmission ratio distortion and early dominant embryo lethality in Anopheles gambiae. PLoS Genet. 4:e1000291. doi: 10.1371/journal.pgen.1000291
Xu, J., Wang, Y., Li, Z., Ling, L., Zeng, B., James, A. A., et al. (2014). Transcription activator-like effector nuclease (TALEN)-mediated female-specific sterility in the silkworm, Bombyx mori. Insect Mol. Biol. 23, 800–807. doi: 10.1111/imb.12125
Xu, L. H., Zeng, B. S., Noland Jeffery, E., Huang, Y. P., and Zhou, X. G. (2015). The coming of RNA-based pest controls. J. Plant Prot. 42, 673–690. doi: 10.13802/j.cnki.zwbhxb.2015.05.001
Yang, C. H., Yang, P. C., Li, J., Yang, F., and Zhang, A. B. (2016). Transcriptome characterization of Dendrolimus punctatus and expression profiles at different developmental stages. PLoS ONE 11:e0161667. doi: 10.1371/journal.pone.0161667
Zeng, J., Ge, F., Su, J., and He, Z. (2010). Researches on the occurrences of major forest insect pests of pine caterpillar Dendrolimus spp. in China. Chinese Bull. Entomol. 47, 451–459.
Zhang, Z., Aslam, A. F., Liu, X., Li, M., Huang, Y., and Tan, A. (2015). Functional analysis of Bombyx Wnt1 during embryogenesis using the CRISPR/Cas9 system. J. Insect Physiol. 79, 73–79. doi: 10.1016/j.jinsphys.2015.06.004
Keywords: Dendrolimus punctatus, CRISPR/Cas9, Wnt-1, segmentation, embryogenesis, genome editing
Citation: Liu H, Liu Q, Zhou X, Huang Y and Zhang Z (2017) Genome Editing of Wnt-1, a Gene Associated with Segmentation, via CRISPR/Cas9 in the Pine Caterpillar Moth, Dendrolimus punctatus. Front. Physiol. 7:666. doi: 10.3389/fphys.2016.00666
Received: 30 July 2016; Accepted: 19 December 2016;
Published: 06 January 2017.
Edited by:
Graziano Fiorito, Stazione Zoologica Anton Dohrn, ItalyReviewed by:
Marjorie A. Lienard, Harvard University, USAAbderrahman Khila, Institut de Génomique Fonctionnelle, France
Copyright © 2017 Liu, Liu, Zhou, Huang and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xuguo Zhou, eHVndW96aG91QHVreS5lZHU=
Yongping Huang, eXBodWFuZ0BzaWJzLmFjLmNu
Zhen Zhang, emhhbmd6aGVuQGNhZi5hYy5jbg==
 Huihui Liu1
Huihui Liu1 Qun Liu
Qun Liu Xuguo Zhou
Xuguo Zhou Zhen Zhang
Zhen Zhang