- Agency for Science, Technology and Research, Genome Institute of Singapore, Singapore.
In the nervous system, several key steps in cellular complexity and development are regulated by non-coding RNAs (ncRNAs) and the repressor element-1 silencing transcription factor/neuron-restrictive silencing factor (REST/NRSF). REST recruits gene regulatory complexes to regulatory sequences, among them the repressor element-1/neuron-restrictive silencer element, and mediates developmental stage-specific gene expression or repression, chromatin (re-)organization or silencing for protein-coding genes as well as for several ncRNAs like microRNAs, short interfering RNAs or long ncRNAs. NcRNAs are far from being just transcriptional noise and are involved in chromatin accessibility, transcription and post-transcriptional processing, trafficking, or RNA editing. REST and its cofactor CoREST are both highly regulated through various ncRNAs. The importance of the correct regulation within the ncRNA network, the ncRNAome, is demonstrated when it comes to a deregulation of REST and/or ncRNAs associated with molecular pathophysiology underlying diverse disorders including neurodegenerative diseases or brain tumors.
Introduction
Both experimental and computational evidence suggest that many of the newly discovered non-coding transcripts and processing products, such as the non-coding (ncRNA) small interfering (siRNA) or microRNAs (miRNA), are functional. Likely, we underestimate the complexity of the regulatory circuitry by assuming a hierarchical structure of well-separated layers in transcriptional regulation. At least a large fraction of the non-coding transcripts themselves are functional and are far from being just transcriptional noise – begging the question “What’s regulated in gene regulation?”. Many of these transcripts show a differential expression in differentiation and disease; there is also sequence conservation and many of the highly conserved elements give rise to ncRNAs of heretofore-unknown function. It is of high importance to understand the fundamental principles in neuroscience, viz. how cellular complexity, diversity, and development are regulated. Such understanding in both healthy and diseased conditions will pave the way to target various diseases and to develop novel drugs and therapeutics. Further, genomic and epigenomic studies provide insights into the complex mechanisms that underlie the aforementioned processes, including the important roles of ncRNAs and the repressor element-1 silencing transcription factor/ neuron-restrictive silencing factor (REST/NRSF). Looking at the complexity of the genome, the whole genome may be transcribed with each nucleotide potentially serving as a multifunctional unit within multiple interconnected genomic layers or elements (Birney et al., 2007). Individual cell types are characterized by their unique repertoire of protein- and non-coding transcripts and a cell’s identity and function is determined by a specific series of temporal and spatial extracellular cues, combinatorial transcription factor codes, epigenetic regulatory networks, and complex chromatin dynamics. In this context, REST and its cofactor CoREST play important roles as they bind to genomic regulatory sequences, including the repressor element-1/neuron-restrictive silencer element (RE1/NRSE). REST and CoREST modulate a huge variety of protein-coding (Johnson et al., 2007; Otto et al., 2007) and non-coding genes (Wu and Xie, 2006; Johnson et al., 2009), regulate gene expression or repression as well as long-term gene silencing (Zheng et al., 2009). Within the network regulated by REST and CoREST, neural genes are connecting points and this network is highly integrated with the ncRNAome that orchestrates aspects as diverse as lineage restriction, stem cell maintenance, differentiation, fate determination, or homeostasis (Mehler, 2008). Given the involvement in these key regulatory pathways, it is not surprising that the perturbation of REST and CoREST activity or the related ncRNAs are implicated in the molecular pathophysiology that underlies various diseases. Among these diseases are tumors like medullo-, glio-, or neuroblastoma (Coulson, 2005; Blom et al., 2006; Westbrook et al., 2008), in addition to ischemia (Formisano et al., 2007), epilepsy (Garriga-Canut et al., 2006; Bassuk et al., 2008), or neurodegenerative diseases (Zuccato et al., 2007; Benn et al., 2008; Packer et al., 2008).
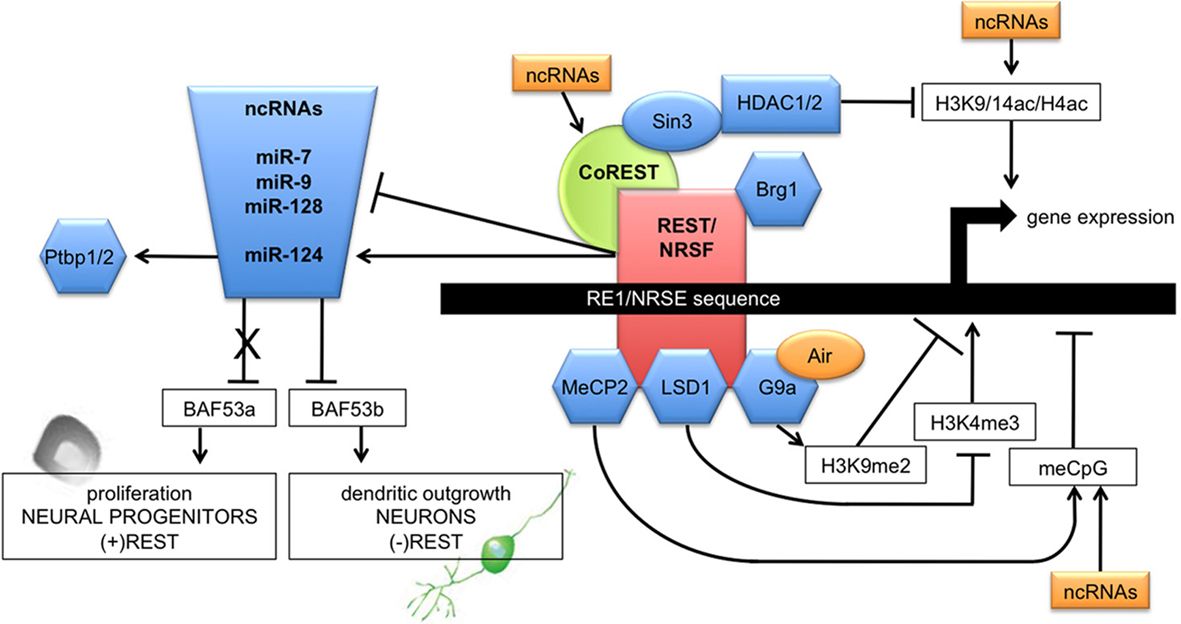
Figure 1. Schematic diagram illustrating the major components of ncRNA–REST/NRSF interactions in neural networks. A yet to identified mechanism initiates the binding of BRG1 to REST/NRSF, resulting in the exposure of the RE1/NRSE motif and the subsequent occupation of it by REST/NRSF. The exact sequential order is unknown. With the assistance of Sin3 and CoREST, the RE1-bound REST/NRSF complexes recruit histone deacetylases (HDAC1/2) and CpG methylDNA binding protein (MeCP2), to promote histone deacetylations, histone methylases (G9a), and histone demethylases (LSD1). REST/NRSF can recruit additional histone methylases and demethylases to target lysine residues of histones; arrows link cofactors to their chromatin remodeling and modifying activities. Inhibition of these histone modifying components results in an attenuation of REST/NRSF recruitment, a decrease in H3K9me2, an increase in H3K9/K14 and H4 acetylation, an increase in H3K4me3 and a decrease in binding of MeCP2 to methylated DNA. REST/NRSF-mediated regulation of neuronal gene expression relies on the dissociation of the REST/NRSF repressor complex from the RE1/NRSE site. In addition to the REST/NRSF complex at the RE1/NRSE site, CoREST and MeCP2 complexes bind to genes on adjacent methylated CpGs in their promoters throughout neuronal differentiation. In neural progenitors, REST/NRSF is expressed and inhibits miR-124a expression, allowing the persistence of non-neuronal transcripts. Upon differentiation of the neuronal progenitors into neurons, REST/NRSF dissociation from the miR-124a gene loci induces derepression of the gene, resulting in the selective degradation of non-neuronal gene transcripts. Thus, REST/NRSF links transcriptional and post-transcriptional events to fine-tune the balance of phenotype between neuronal and non-neuronal cells, controlled by miR-124 and further non-coding transcripts (ncRNA) that act in a network with miR-124. See text for references.
REST and CoREST in Neural Networks
The zinc finger protein REST binds to genomic sequences that include canonical RE1 sequences. REST acts as a modulator for the assembly of diverse transcription factors. Initially thought to repress only neuronal genes in non-neuronal cells (Schoenherr and Anderson, 1995; Ballas et al., 2001), REST is now seen as a key regulator in multifaceted pathways in various cell types and as a modulator of the cellular epigenome. In addition to CoREST, REST also recruits mSin3 (Grimes et al., 2000), a scaffold for histone deacetylases (Hdac-1, -2, -4, and -5). When associated with REST, CoREST binds to Hdac-1 and -2, histone H3K4 lysine demethylase, LSD1, histone H3K9 methyltransferases, G9a and Suv39h1, methyl-CpG binding protein 2 (Mecp2), and to a component of the SWI/SNF chromatin remodeling complex, Brg1 (Andres et al., 1999). REST also associates with a wide number of epigenetic and regulatory cofactors that include DNA methyltransferases (DNMTs), chromatin remodeling enzymes, the RNA polymerase II transcriptional Mediator subunits, Med-19 and -26 (Ding et al., 2009), the NADH-binding factor, CtBP (Garriga-Canut et al., 2006; Guardavaccaro et al., 2008), the transcription factor Sp3 (Kim et al., 2006), or the small C-terminal domain phosphatase, Scp1 (Visvanathan et al., 2007) (Figure 1). Therefore, REST acts as a platform for genomic integrity, stability, and regulation and specifies context-dependent gene repression or activation. The complexity further increases when looking at the various parameters that regulate the affinity of the REST complex through several parameters that include its affinity for binding to different RE1 and non-RE1 sites, alternative spliced variants of REST (Shimojo et al., 1999; Lee et al., 2000a), or the modulation via ncRNAs (Kuwabara et al., 2004; Visvanathan et al., 2007; Packer et al., 2008).
The locus for REST contains several regulatory elements that are targeted by transcriptional networks including Nanog and Oct4 (Singh et al., 2008) or by factors like Wnt and retinoic acid (Nishihara et al., 2003). The shuffling of REST from the nucleus to the cytoplasm is controlled by huntingtin (Htt) and is controlled by a complex of Htt, dynactin p150, and REST (Johnson et al., 2008). Besides the effects of transcriptional regulation, a role of REST in post-transcriptional processing has been shown for the μ-opioid receptor (μOR) gene (Kim et al., 2008).
The Interplay of REST and ncRNAs
Since REST contains an RE1 sequence, it may be that REST regulates its own expression in a feedback loop. In this regard, it is interesting that small double-stranded ncRNAs encoding the RE1 sequence (dsNRSEs) were found to interact with the REST complex to promote the expression of RE1-associated genes (Kuwabara et al., 2005). This fact demonstrates the close link of REST function and ncRNA expression and is particularly relevant for the determination of cell fates in the nervous system. Initially, REST was found to modulate genes for neuronal differentiation, homeostasis and plasticity, including cytokines, neurotransmitter receptors, ion channels, cell adhesion molecules, growth factors, and factors for axonal guidance or vesicle proteins at synapses (Chen et al., 1998). A similar role has been demonstrated for miRNAs and ncRNA networks in neuronal differentiation and plasticity (Ooi and Wood, 2008).
Genome-wide mapping, expression and binding profiles for REST have been examined in various studies. REST targets both the canonical and non-canonical RE1 sequence that contains insertions in the canonical sequence of variable lengths (Ooi and Wood, 2007; Otto et al., 2007). NcRNAs are believed to play an important role in the regulation of REST binding to such sequences. Among them are miRNAs that are cell type-specific and are only expressed in the brain, viz. miR-9, miR-124, or miR-128 (Ooi and Wood, 2008) (Figure 1). MiRNAs directly target the 3′-untranslated region (UTR) of target mRNAs leading to a sequestration of mRNAs in P-bodies for storage or subsequent degradation, respectively. P-bodies regulate RNA metabolism, translational repression and play a critical role in the transport of RNAs to dendrites regulating synaptic plasticity (Zeitelhofer et al., 2008). Many functions of the complex ncRNAome are yet to be uncovered, however, a large number of RE1 sites are present within close proximity of nervous system enriched miRNA genes – and the expression of those genes is regulated by REST (Wu and Xie, 2006).
The important role of critical components of the RNAi mediated gene silencing pathways has been demonstrated for factors that are associated with RE1, such as Argonaute (Ago1, Ago3, Ago4, Xpo5) or Dicer (Dicer1) proteins (Nishihara et al., 2003; Rossbach, 2010). REST itself may be targeted by multiple ncRNAs, such as the miRNAs mir-9 or miR-124 (Wu and Xie, 2006; Packer et al., 2008) and, consequently, there may be several layers of regulation connected in neurodevelopment, e.g., double-negative feedback loops for REST and ncRNAs. There are further classes of non-coding regulators of REST, like long ncRNAs; 23% of the REST binding sites lie within a 10-kb region of long ncRNA genes (Shimojo et al., 1999). For instance, when looking at the DiGeorge syndrome, the locus of the DiGeorge syndrome-associated ncRNA DGCR5 is a breakpoint region, which is characterized by malformations during neurodevelopment and causes neuropsychiatric diseases. REST binds to a binding site of DGCR5, leading to transcriptional repression (Johnson et al., 2009). Mammalian genomes may encode huge amounts of such long ncRNAs, many of them are poly-A tailed, highly regulated during development, alternately spliced, and enriched in the nervous system. In the brain and in embryonic stem cells (ESCs), long ncRNAs exhibit cell type-specific, spatial, and temporal regulated expression profiles (Dinger et al., 2008a) Besides transcription, post-transcriptional processing, RNA editing, and trafficking (Mehler, 2008), ncRNAs regulate chromosomal architecture, dynamics and plasticity by recruiting chromatin activator or repressor complexes to their targets (Mattick et al., 2009). An example is the ncRNA Air that accumulates at promoter regions where it silences gene expression by recruiting the H3K9 methyltransferase G9a (Nagano et al., 2008) (Figure 1).
Network Regulations Mediated by ncRNAs
Non-coding transcripts not only include miRNA, siRNAs, and long ncRNAs, but also small nucleolar RNAs (snoRNA) and antisense RNAs (asRNAs) – but it is quite likely that further classes of ncRNAs will emerge. Given the complex regulatory networks, REST and CoREST may function as dynamic mediators of regulation, thus regulating transcription, chromatin architecture and responding to epigenetic signals (Gan et al., 2007). REST has been implicated in cell- and tissue-specific epigenetics, and is, like CoREST, crucial in neural stem cells (NSCs) and non-neuronal cells, but also in ESCs, neuronal and glial progenitor cells, or in terminally differentiated neurons and glia. During specific lineage determination, silencing of genes involved in lineage specification and pluripotency takes place. Thus, REST is highly integrated with the network of transcription in maintenance of pluripotency, viz. Oct4, Nanog, or Sox2 (Singh et al., 2008; Jorgensen et al., 2009a). Studies suggest that REST is involved in ESC pluripotency by targeting miRNAs like miR-21 (Singh et al., 2008); other studies, however, find conflicting results and show no significant impact of REST on the maintenance of pluripotency (Buckley et al., 2009; Jorgensen et al., 2009a). Besides its potential role in pluripotency, REST is implicated in the silencing of RE1 associated genes in ESCs, among them genes important for neuronal terminal differentiation or function. Examples include, but are not limited to, Syt4, Syp, Stmn3, or Celsr3. However, Mash1 or Math1 are not on this list, suggesting that REST does not repress the potential for neural lineage commitment but silences genes important for neuronal subtype specification in ESCs (Jorgensen et al., 2009b). The role of REST in neurogenesis has been demonstrated through the interaction of REST and ncRNAs encoding the RE1 sequence (dsNRSEs); here, REST acts as an activator and induces neuronal differentiation (Kuwabara et al., 2005). REST-deficient NSCs show defective behavior in regard to migration and survival (Sun et al., 2008).
Repressor element-1 silencing transcription factor is downregulated during NSC differentiation (Westbrook et al., 2008) and regulates the nervous system specific miRNA miR-124 that silences several non-neuronal genes; in non-neuronal cells, REST silences miR-124 (Wu and Xie, 2006). During the maturation of neurons, however, REST is suppressed and thus miR-124 levels rise. Such observations implicate that REST plays a critical role in neurodevelopment and in regional neuronal subtype and glial lineage specification, with CoREST mediating NSC maintenance and maturational functions (Qureshi and Mehler, 2009).
Increasing Complexity in Neural Networks
In healthy and diseased contexts, REST levels vary significantly, and both overexpression and downregulation of REST have been observed in various disease phenotypes. Thus, REST can act as a tumor suppressor and oncogene, respectively, depending on the cellular environment (Westbrook et al., 2005; Weissman, 2008). With respect to the ncRNAome, several altered levels of ncRNA expression have been observed in various cancer phenotypes, among them neuroblastoma or glioblastoma multiforme in the nervous system (Nicoloso and Calin, 2008; Skog et al., 2008), and such observations demonstrate how an aberrant expression of REST and CoREST is involved in pathogenesis of several disorders like cancer, neurodevelopmental, or neurodegenerative diseases.
When looking at the epigenome, it is becoming more and more evident that REST and its associated factors are critical modulators during development, lineage specification or gene regulation in general and that all these processes depend on epigenetic signals. Here, another feedback regulation is in place since REST not only mediates responses to epigenetic signal, but also targets receptor expression or components of the signaling cascade – with ncRNAs being important for the execution of these regulatory processes. During neurodevelopment, the correct modifications of RNAs are crucial, like RNA trafficking, editing, or alternative splicing. Such processes are regulated in a complex interplay of REST, ncRNAs, and cofactors like Ptbp1 that is directly targeted by miR-124. While neuronal differentiation is progressing, miR-124 downregulates Ptbp1, leading to an increase in Ptbp2. Ptbp2 is the nervous system homolog of Ptbp1, mediates neuron specific alternative RNA splicing (Makeyev et al., 2007) and is targeted by REST and CoREST during neural lineage specification.
This complexity further increases when considering the fact that REST has several isoforms with synergistic or antagonistic effects for the regulatory network REST is integrated in Coulson et al., 2000, Lee et al., 2000b, Shimojo et al., 1999. Protein-coding RNA and ncRNA modifications, trafficking, functional modifications, cellular signaling, and epigenetic signal processing seem to be modulated by REST, CoREST, and ncRNAs (Dinger et al., 2008b). However, further studies are required to elucidate the full picture of REST medicated ncRNA networks.
Concluding Remarks – The Therapeutic Implications
The emerging concepts of ncRNAs as novel therapeutics and diagnostics in neurological diseases bear both great potentials and challenges. Several mutations, deletions, amplifications, and alterations have been found in neurological and neurodegenerative diseases and were shown to alter the expression levels of ncRNA transcripts, like lncRNAs or miRNAs. A large fraction of highly conserved regions encodes for sets of ncRNAs whose expression is altered during disease progression. Besides the molecular and biochemical studies, bioinformatics, and systems biology approaches become more and more important as tools to identify associations and correlations between ncRNAs and gene regulatory networks since ncRNA profiles allow the identification of various signatures that are associated with diagnosis, prognosis, and the response to treatment of neurological, neurodegenerative diseases, or brain tumors. Detailed ncRNAome profiles are a first step toward discoveries leading to novel therapeutics since they were shown to affect brain tumor development or progression and are implicated in various diseases. Specifically addressing changes in the ncRNAome in disease contexts with tools that focus on si-, lncRNAs, and miRNAs will lead to important findings with respect to biomarker discovery and the identification of novel therapeutic targets. The targeting of the ncRNAome is becoming an important therapeutic and diagnostic strategy, a concept known as theranostics, and is therefore highly relevant for drug development and personalized medicine. Examples include the restoration of the expression of a deficient or downregulated ncRNA or, alternatively, the inhibition of an overexpressed ncRNA to reverse a disease phenotype. The silencing of oncogenic and the re-activation of tumor suppressor miRNAs, for instance, are possible therapeutic paradigms that are currently being tested in clinical trials. If successful, such compounds represent new tools for the treatment of cancer and other diseases. With their unique sequences and secondary structural elements, ncRNAs comprise potential binding sites for small molecular drugs acting either as agonists or antagonists, respectively. Further areas of intervention with small molecules, aptamers, peptides, or oligonucleotides are (i) the interaction of ncRNAs with epigenetic effector molecules or (ii) with specific RNA degradation pathways. Both therapeutic options can be used to target aberrant gene expression; however, they require further validation. Thus, the understanding of the basic processes of gene regulation may lead toward innovative therapeutic strategies for personalized treatments of various human diseases.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Andres, M. E., Burger, C., Peral-Rubio, M. J., Battaglioli, E., Anderson, M. E., Grimes, J., Dallman, J., Ballas, N., and Mandel, G. (1999). CoREST: a functional corepressor required for regulation of neural-specific gene expression. Proc. Natl. Acad. Sci. U.S.A. 96, 9873–9878.
Ballas, N., Battaglioli, E., Atouf, F., Andres, M. E., Chenoweth, J., Anderson, M. E., Burger, C., Moniwa, M., Davie, J. R., Bowers, W. J., Federoff, H. J., Rose, D. W., Rosenfeld, M. G., Brehm, P., and Mandel, G. (2001). Regulation of neuronal traits by a novel transcriptional complex. Neuron 31, 353–365.
Bassuk, A. G., Wallace, R. H., Buhr, A., Buller, A. R., Afawi, Z., Shimojo, M., Miyata, S., Chen, S., Gonzalez-Alegre, P., Griesbach, H. L., Wu, S., Nashelsky, M., Vladar, E. K., Antic, D., Ferguson, P. J., Cirak, S., Voit, T., Scott, M. P., Axelrod, J. D., Gurnett, C., Daoud, A. S., Kivity, S., Neufeld, M. Y., Mazarib, A., Straussberg, R., Walid, S., Korczyn, A. D., Slusarski, D. C., Berkovic, S. F., and El-Shanti, H. I. (2008). A homozygous mutation in human PRICKLE1 causes an autosomal-recessive progressive myoclonus epilepsy-ataxia syndrome. Am. J. Hum. Genet. 83, 572–581.
Benn, C. L., Sun, T., Sadri-Vakili, G., McFarland, K. N., DiRocco, D. P., Yohrling, G. J., Clark, T. W., Bouzou, B., and Cha, J. H. (2008). Huntingtin modulates transcription, occupies gene promoters in vivo, and binds directly to DNA in a polyglutamine dependent manner. J. Neurosci. 28, 10720–10733.
Birney, E., Stamatoyannopoulos, J. A., Dutta, A., Guigo, R., Gingeras, T. R., Margulies, E. H., Weng, Z., Snyder, M., Dermitzakis, E. T., Thurman, R. E., Kuehn, M. S., Taylor, C. M., Neph, S., Koch, C. M., Asthana, S., Malhotra, A., Adzhubei, I., Greenbaum, J. A., Andrews, R. M., Flicek, P., Boyle, P. J., Cao, H., Carter, N. P., Clelland, G. K., Davis, S., Day, N., Dhami, P., Dillon, S. C., Dorschner, M. O., Fiegler, H., Giresi, P. G., Goldy, J., Hawrylycz, M., Haydock, A., Humbert, R., James, K. D., Johnson, B. E., Johnson, E. M., Frum, T. T., Rosenzweig, E. R., Karnani, N., Lee, K., Lefebvre, G. C., Navas, P. A., Neri, F., Parker, S. C., Sabo, P. J., Sandstrom, R., Shafer, A., Vetrie, D., Weaver, M., Wilcox, S., Yu, M., Collins, F. S., Dekker, J., Lieb, J. D., Tullius, T. D., Crawford, G. E., Sunyaev, S., Noble, W. S., Dunham, I., Denoeud, F., Reymond, A., Kapranov, P., Rozowsky, J., Zheng, D., Castelo, R., Frankish, A., Harrow, J., Ghosh, S., Sandelin, A., Hofacker, I. L., Baertsch, R., Keefe, D., Dike, S., Cheng, J., Hirsch, H. A., Sekinger, E. A., Lagarde, J., Abril, J. F., Shahab, A., Flamm, C., Fried, C., Hackermuller, J., Hertel, J., Lindemeyer, M., Missal, K., Tanzer, A., Washietl, S., Korbel, J., Emanuelsson, O., Pedersen, J. S., Holroyd, N., Taylor, R., Swarbreck, D., Matthews, N., Dickson, M. C., Thomas, D. J., Weirauch, M. T., Gilbert, J., Drenkow, J., Bell, I., Zhao, X., Srinivasan, K. G., Sung, W. K., Ooi, H. S., Chiu, K. P., Foissac, S., Alioto, T., Brent, M., Pachter, L., Tress, M. L., Valencia, A., Choo, S. W., Choo, C. Y., Ucla, C., Manzano, C., Wyss, C., Cheung, E., Clark, T. G., Brown, J. B., Ganesh, M., Patel, S., Tammana, H., Chrast, J., Henrichsen, C. N., Kai, C., Kawai, J., Nagalakshmi, U., Wu, J., Lian, Z., Lian, J., Newburger, P., Zhang, X., Bickel, P., Mattick, J. S., Carninci, P., Hayashizaki, Y., Weissman, S., Hubbard, T., Myers, R. M., Rogers, J., Stadler, P. F., Lowe, T. M., Wei, C. L., Ruan, Y., Struhl, K., Gerstein, M., Antonarakis, S. E., Fu, Y., Green, E. D., Karaoz, U., Siepel, A., Taylor, J., Liefer, L. A., Wetterstrand, K. A., Good, P. J., Feingold, E. A., Guyer, M. S., Cooper, G. M., Asimenos, G., Dewey, C. N., Hou, M., Nikolaev, S., Montoya-Burgos, J. I., Loytynoja, A., Whelan, S., Pardi, F., Massingham, T., Huang, H., Zhang, N. R., Holmes, I., Mullikin, J. C., Ureta-Vidal, A., Paten, B., Seringhaus, M., Church, D., Rosenbloom, K., Kent, W. J., Stone, E. A., Batzoglou, S., Goldman, N., Hardison, R. C., Haussler, D., Miller, W., Sidow, A., Trinklein, N. D., Zhang, Z. D., Barrera, L., Stuart, R., King, D. C., Ameur, A., Enroth, S., Bieda, M. C., Kim, J., Bhinge, A. A., Jiang, N., Liu, J., Yao, F., Vega, V. B., Lee, C. W., Ng, P., Yang, A., Moqtaderi, Z., Zhu, Z., Xu, X., Squazzo, S., Oberley, M. J., Inman, D., Singer, M. A., Richmond, T. A., Munn, K. J., Rada-Iglesias, A., Wallerman, O., Komorowski, J., Fowler, J. C., Couttet, P., Bruce, A. W., Dovey, O. M., Ellis, P. D., Langford, C. F., Nix, D. A., Euskirchen, G., Hartman, S., Urban, A. E., Kraus, P., Van Calcar, S., Heintzman, N., Kim, T. H., Wang, K., Qu, C., Hon, G., Luna, R., Glass, C. K., Rosenfeld, M. G., Aldred, S. F., Cooper, S. J., Halees, A., Lin, J. M., Shulha, H. P., Xu, M., Haidar, J. N., Yu, Y., Iyer, V. R., Green, R. D., Wadelius, C., Farnham, P. J., Ren, B., Harte, R. A., Hinrichs, A. S., Trumbower, H., Clawson, H., Hillman-Jackson, J., Zweig, A. S., Smith, K., Thakkapallayil, A., Barber, G., Kuhn, R. M., Karolchik, D., Armengol, L., Bird, C. P., de Bakker, P. I., Kern, A. D., Lopez-Bigas, N., Martin, J. D., Stranger, B. E., Woodroffe, A., Davydov, E., Dimas, A., Eyras, E., Hallgrimsdottir, I. B., Huppert, J., Zody, M. C., Abecasis, G. R., Estivill, X., Bouffard, G. G., Guan, X., Hansen, N. F., Idol, J. R., Maduro, V. V., Maskeri, B., McDowell, J. C., Park, M., Thomas, P. J., Young, A. C., Blakesley, R. W., Muzny, D. M., Sodergren, E., Wheeler, D. A., Worley, K. C., Jiang, H., Weinstock, G. M., Gibbs, R. A., Graves, T., Fulton, R., Mardis, E. R., Wilson, R. K., Clamp, M., Cuff, J., Gnerre, S., Jaffe, D. B., Chang, J. L., Lindblad-Toh, K., Lander, E. S., Koriabine, M., Nefedov, M., Osoegawa, K., Yoshinaga, Y., Zhu, B., and de Jong, P. J. (2007). Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816.
Blom, T., Tynninen, O., Puputti, M., Halonen, M., Paetau, A., Haapasalo, H., Tanner, M., and Nupponen, N. N. (2006). Molecular genetic analysis of the REST/NRSF gene in nervous system tumors. Acta Neuropathol. 112, 483–490.
Buckley, N. J., Johnson, R., Sun, Y. M., and Stanton, L. W. (2009). Is REST a regulator of pluripotency? Nature 457, E5–E6.
Chen, Z. F., Paquette, A. J., and Anderson, D. J. (1998). NRSF/REST is required in vivo for repression of multiple neuronal target genes during embryogenesis. Nat. Genet. 20, 136–142.
Coulson, J. M. (2005). Transcriptional regulation: cancer, neurons and the REST. Curr. Biol. 15, R665–R668.
Coulson, J. M., Edgson, J. L., Woll, P. J., and Quinn, J. P. (2000). A splice variant of the neuron restrictive silencer factor repressor is expressed in small cell lung cancer: a potential role in derepression of neuroendocrine genes and a useful clinical marker. Cancer Res. 60, 1840–1844.
Ding, N., Tomomori-Sato, C., Sato, S., Conaway, R. C., Conaway, J. W., and Boyer, T. G. (2009). MED19 and MED26 are synergistic functional targets of the RE1 silencing transcription factor in epigenetic silencing of neuronal gene expression. J. Biol. Chem. 284, 2648–2656.
Dinger, M. E., Amaral, P. P., Mercer, T. R., Pang, K. C., Bruce, S. J., Gardiner, B. B., Askarian-Amiri, M. E., Ru, K., Solda, G., Simons, C., Sunkin, S. M., Crowe, M. L., Grimmond, S. M., Perkins, A. C., and Mattick, J. S. (2008a). Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res. 18, 1433–1445.
Dinger, M. E., Mercer, T. R., and Mattick, J. S. (2008b). RNAs as extracellular signaling molecules. J. Mol. Endocrinol. 40, 151–159.
Formisano, L., Noh, K. M., Miyawaki, T., Mashiko, T., Bennett, M. V., and Zukin, R. S. (2007). Ischemic insults promote epigenetic reprogramming of mu opioid receptor expression in hippocampal neurons. Proc. Natl. Acad. Sci. U.S.A. 104, 4170–4175.
Gan, Q., Yoshida, T., McDonald, O. G., and Owens, G. K. (2007). Concise review: epigenetic mechanisms contribute to pluripotency and cell lineage determination of embryonic stem cells. Stem Cells 25, 2–9.
Garriga-Canut, M., Schoenike, B., Qazi, R., Bergendahl, K., Daley, T. J., Pfender, R. M., Morrison, J. F., Ockuly, J., Stafstrom, C., Sutula, T., and Roopra, A. (2006). 2-Deoxy-d-glucose reduces epilepsy progression by NRSF-CtBP-dependent metabolic regulation of chromatin structure. Nat. Neurosci. 9, 1382–1387.
Grimes, J. A., Nielsen, S. J., Battaglioli, E., Miska, E. A., Speh, J. C., Berry, D. L., Atouf, F., Holdener, B. C., Mandel, G., and Kouzarides, T. (2000). The co-repressor mSin3A is a functional component of the REST-CoREST repressor complex. J. Biol. Chem. 275, 9461–9467.
Guardavaccaro, D., Frescas, D., Dorrello, N. V., Peschiaroli, A., Multani, A. S., Cardozo, T., Lasorella, A., Iavarone, A., Chang, S., Hernando, E., and Pagano, M. (2008). Control of chromosome stability by the beta-TrCP-REST-Mad2 axis. Nature 452, 365–369.
Johnson, D. S., Mortazavi, A., Myers, R. M., and Wold, B. (2007). Genome-wide mapping of in vivo protein-DNA interactions. Science 316, 1497–1502.
Johnson, R., Teh, C. H., Jia, H., Vanisri, R. R., Pandey, T., Lu, Z. H., Buckley, N. J., Stanton, L. W., and Lipovich, L. (2009). Regulation of neural macroRNAs by the transcriptional repressor REST. RNA 15, 85–96.
Johnson, R., Zuccato, C., Belyaev, N. D., Guest, D. J., Cattaneo, E., and Buckley, N. J. (2008). A microRNA-based gene dysregulation pathway in Huntington’s disease. Neurobiol. Dis. 29, 438–445.
Jorgensen, H. F., Chen, Z. F., Merkenschlager, M., and Fisher, A. G. (2009a). Is REST required for ESC pluripotency? Nature 457, E4–E5 [discussion E7].
Jorgensen, H. F., Terry, A., Beretta, C., Pereira, C. F., Leleu, M., Chen, Z. F., Kelly, C., Merkenschlager, M., and Fisher, A. G. (2009b). REST selectively represses a subset of RE1-containing neuronal genes in mouse embryonic stem cells. Development 136, 715–721.
Kim, C. S., Choi, H. S., Hwang, C. K., Song, K. Y., Lee, B. K., Law, P. Y., Wei, L. N., and Loh, H. H. (2006). Evidence of the neuron-restrictive silencer factor (NRSF) interaction with Sp3 and its synergic repression to the mu opioid receptor (MOR) gene. Nucleic Acids Res. 34, 6392–6403.
Kim, C. S., Hwang, C. K., Song, K. Y., Choi, H. S., Kim do, K., Law, P. Y., Wei, L. N., and Loh, H. H. (2008). Novel function of neuron-restrictive silencer factor (NRSF) for posttranscriptional regulation. Biochim. Biophys. Acta 1783, 1835–1846.
Kuwabara, T., Hsieh, J., Nakashima, K., Taira, K., and Gage, F. H. (2004). A small modulatory dsRNA specifies the fate of adult neural stem cells. Cell 116, 779–793.
Kuwabara, T., Hsieh, J., Nakashima, K., Warashina, M., Taira, K., and Gage, F. H. (2005). The NRSEsmRNAspecifies the fate of adult hippocampal neural stem cells. Nucleic Acids Symp. Ser. (Oxf.) 87–88.
Lee, J. H., Chai, Y. G., and Hersh, L. B. (2000a). Expression patterns of mouse repressor element-1 silencing transcription factor 4 (REST4) and its possible function in neuroblastoma. J. Mol. Neurosci. 15, 205–214.
Lee, J. H., Shimojo, M., Chai, Y. G., and Hersh, L. B. (2000b). Studies on the interaction of REST4 with the cholinergic repressor element-1/neuron restrictive silencer element. Brain Res. Mol. Brain Res. 80, 88–98.
Makeyev, E. V., Zhang, J., Carrasco, M. A., and Maniatis, T. (2007). The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell 27, 435–448.
Mattick, J. S., Amaral, P. P., Dinger, M. E., Mercer, T. R., and Mehler, M. F. (2009). RNA regulation of epigenetic processes. Bioessays 31, 51–59.
Mehler, M. F. (2008). Epigenetic principles and mechanisms underlying nervous system functions in health and disease. Prog. Neurobiol. 86, 305–341.
Nagano, T., Mitchell, J. A., Sanz, L. A., Pauler, F. M., Ferguson-Smith, A. C., Feil, R., and Fraser, P. (2008). The air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322, 1717–1720.
Nicoloso, M. S., and Calin, G. A. (2008). MicroRNA involvement in brain tumors: from bench to bedside. Brain Pathol. 18, 122–129.
Nishihara, S., Tsuda, L., and Ogura, T. (2003). The canonical Wnt pathway directly regulates NRSF/REST expression in chick spinal cord. Biochem. Biophys. Res. Commun. 311, 55–63.
Ooi, L., and Wood, I. C. (2007). Chromatin crosstalk in development and disease: lessons from REST. Nat. Rev. Genet. 8, 544–554.
Ooi, L., and Wood, I. C. (2008). Regulation of gene expression in the nervous system. Biochem. J. 414, 327–341.
Otto, S. J., McCorkle, S. R., Hover, J., Conaco, C., Han, J. J., Impey, S., Yochum, G. S., Dunn, J. J., Goodman, R. H., and Mandel, G. (2007). A new binding motif for the transcriptional repressor REST uncovers large gene networks devoted to neuronal functions. J. Neurosci. 27, 6729–6739.
Packer, A. N., Xing, Y., Harper, S. Q., Jones, L., and Davidson, B. L. (2008). The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington’s disease. J. Neurosci. 28, 14341–14346.
Qureshi, I. A., and Mehler, M. F. (2009). Regulation of non-coding RNA networks in the nervous system – what’s the REST of the story? Neurosci. Lett. 466, 73–80.
Schoenherr, C. J., and Anderson, D. J. (1995). The neuron-restrictive silencer factor (NRSF): a coordinate repressor of multiple neuron-specific genes. Science 267, 1360–1363.
Shimojo, M., Paquette, A. J., Anderson, D. J., and Hersh, L. B. (1999). Protein kinase A regulates cholinergic gene expression in PC12 cells: REST4 silences the silencing activity of neuron-restrictive silencer factor/REST. Mol. Cell. Biol. 19, 6788–6795.
Singh, S. K., Kagalwala, M. N., Parker-Thornburg, J., Adams, H., and Majumder, S. (2008). REST maintains self-renewal and pluripotency of embryonic stem cells. Nature 453, 223–227.
Skog, J., Wurdinger, T., van Rijn, S., Meijer, D. H., Gainche, L., Sena-Esteves, M., Curry, W. T. Jr., Carter, B. S., Krichevsky, A. M., and Breakefield, X. O. (2008). Glioblastoma microvesicles transport RNA and proteins that promote tumor growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476.
Sun, Y. M., Cooper, M., Finch, S., Lin, H. H., Chen, Z. F., Williams, B. P., and Buckley, N. J. (2008). Rest-mediated regulation of extracellular matrix is crucial for neural development. PLoS ONE 3, e3656.
Visvanathan, J., Lee, S., Lee, B., Lee, J. W., and Lee, S. K. (2007). The microRNA miR-124 antagonizes the anti-neural REST/SCP1 pathway during embryonic CNS development. Genes Dev. 21, 744–749.
Westbrook, T. F., Hu, G., Ang, X. L., Mulligan, P., Pavlova, N. N., Liang, A., Leng, Y., Maehr, R., Shi, Y., Harper, J. W., and Elledge, S. J. (2008). SCFbeta-TRCP controls oncogenic transformation and neural differentiation through REST degradation. Nature 452, 370–374.
Westbrook, T. F., Martin, E. S., Schlabach, M. R., Leng, Y., Liang, A. C., Feng, B., Zhao, J. J., Roberts, T. M., Mandel, G., Hannon, G. J., Depinho, R. A., Chin, L., and Elledge, S. J. (2005). A genetic screen for candidate tumor suppressors identifies REST. Cell 121, 837–848.
Wu, J., and Xie, X. (2006). Comparative sequence analysis reveals an intricate network among REST, CREB and miRNA in mediating neuronal gene expression. Genome Biol. 7, R85.
Zeitelhofer, M., Macchi, P., and Dahm, R. (2008). Perplexing bodies: the putative roles of P-bodies in neurons. RNA Biol 5, 244–248.
Zheng, D., Zhao, K., and Mehler, M. F. (2009). Profiling RE1/REST-mediated histone modifications in the human genome. Genome Biol. 10, R9.
Zuccato, C., Belyaev, N., Conforti, P., Ooi, L., Tartari, M., Papadimou, E., MacDonald, M., Fossale, E., Zeitlin, S., Buckley, N., and Cattaneo, E. (2007). Widespread disruption of repressor element-1 silencing transcription factor/neuron-restrictive silencer factor occupancy at its target genes in Huntington’s disease. J. Neurosci. 27, 6972–6983.
Keywords: non-coding RNA, REST, gene regulation, pathophysiology, nervous system
Citation: Rossbach M (2011) Non-coding RNAs in neural networks, REST-assured. Front. Gene. 2:8. doi:10.3389/fgene.2011.00008
Received: 07 December 2010;
Paper pending published: 01 January 2011;
Accepted: 15 February 2011;
Published online: 28 February 2011.
Edited by:
Muller Fabbri, The Ohio State University, USAReviewed by:
George Perry, The University of Texas at San Antonio, USAGeorge Calin, MD Anderson Cancer Center, USA
Alan G. Ramsay, Queen Mary University of London, UK
Copyright: © 2011 Rossbach. This is an open-access article subject to an exclusive license agreement between the authors and Frontiers Media SA, which permits unrestricted use, distribution, and reproduction in any medium, provided the original authors and source are credited.
*Correspondence: Michael Rossbach, Agency for Science, Technology and Research, Genome Institute of Singapore, Singapore. e-mail:cm9zc2JhY2htQGdpcy5hLXN0YXIuZWR1LnNn
