- Department of Molecular Virology, Immunology and Medical Genetics, The Ohio State University Comprehensive Cancer Center, Columbus, OH, USA
Prostate cancer (PCA) still represents a leading cause of death. An increasing number of studies have documented that microRNAs (miRNAs), a subgroup of non-coding RNAs with gene regulatory functions, are differentially expressed in PCA respect to the normal tissue counterpart, suggesting their involvement in prostate carcinogenesis and dissemination. Interestingly, it has been shown that miRNAs undergo the same regulatory mechanisms than any other protein coding gene, including epigenetic regulation. In turn, miRNAs can also affect the expression of oncogenes and tumor suppressor genes by targeting effectors of the epigenetic machinery, therefore indirectly affecting the epigenetic controls on these genes. Among the genes that undergo this complex regulation, there is the androgen receptor (AR), a key therapeutic target for PCA. This review will focus on the role of epigenetically regulated and epigenetically regulating miRNAs in PCA and on the fine regulation of AR expression, as mediated by this miRNA–epigenetics interaction.
Introduction
Epigenetics is the study of changes in gene expression caused by mechanisms other than changes in the DNA sequence. Accumulation of genetic and epigenetic alterations is one of the most important factors in the tumorigenesis and progression of prostate cancer (PCA). DNA hypermethylation, loss of imprinting and altered histone modification patterns have been frequently reported in PCA. The transcriptional control of gene expression is regulated by a complex machinery composed of DNA methyltransferases (DNMTs), histone acetyltransferases (HATs), histone deacetylases (HDACs), histone methyltransferases (HMTs), histone demethylases (HDMTs), and chromatin remodeling enzymes.
Hypermethylation of a 5′-cytosine to guanosine in CpG rich areas located in the promoter region of several genes occurs frequently in PCA, but rarely in normal prostate tissue, resulting in silencing of methylated genes. Many of these epigenetically regulated genes are tumor suppressor genes (TSGs), whose silencing promotes a carcinogenetic effect. In addition to this mechanism, also upregulation of effectors of the epigenetic machinery, such as the HMT enhancer of zeste homolog 2 (EZH2) has been associated with high proliferation rate and aggressive tumor subgroups in PCA. These data highlight the importance of epigenetic factors in PCA genesis and progression.
MicroRNAs (miRNAs) are small non-coding RNAs (ncRNAs) with gene regulatory functions (Ambros, 2001). The advent of high throughput techniques has revealed that aberrations of the miRNome (defined as the full spectrum of miRNA expression in a given genome) occur in almost all human tumors, with respect to the normal tissue counterpart, including PCA (Volinia et al., 2006; Dillhoff et al., 2008; Croce, 2009; Fabbri and Croce, 2011). In addition to the “traditional” mechanism of action of miRNAs (according to which these ncRNAs mainly silence the target gene by binding to the 3′-untranslated region, 3′-UTR of its mRNA; Bartel, 2004), miRNAs can also increase the translation of their target gene, by binding to the 5′-UTR of the targeted mRNA (Vasudevan et al., 2007), and it has also been observed that they can affect gene expression by modifying their epigenetic control (Fabbri et al., 2007a). In particular, our group was able to show that in cancer, a specific family of miRNAs (namely, the miR-29 family) directly targets DNMT3A and DNMT3B (Fabbri et al., 2007a), and indirectly targets DNMT1 (Garzon et al., 2009), leading to a global reduction in cancer cell methylation and re-expression of TSGs (such as WWOX and FHIT), normally epigenetically silenced in tumors, and to an anti-tumoral effect. Moreover, several groups have previously shown that miRNAs undergo the same epigenetic regulatory mechanisms that affect the expression of any other protein coding gene (PCG), revealing a complex level of interaction between epigenetics and miRNAs. This review will focus on the implications of this miRNA–epigenetics network for PCA, and will provide the state-of-the-art knowledge on how the intertwined relationship between miRNA and epigenetics regulates the expression of the androgen receptor (AR), one of the main actors in the development and progression of PCA, therefore providing the rationale for new possible therapeutic avenues for the treatment of this still very debilitating type of cancer.
miRNAs Regulate the Epigenetic Machinery in Prostate Cancer
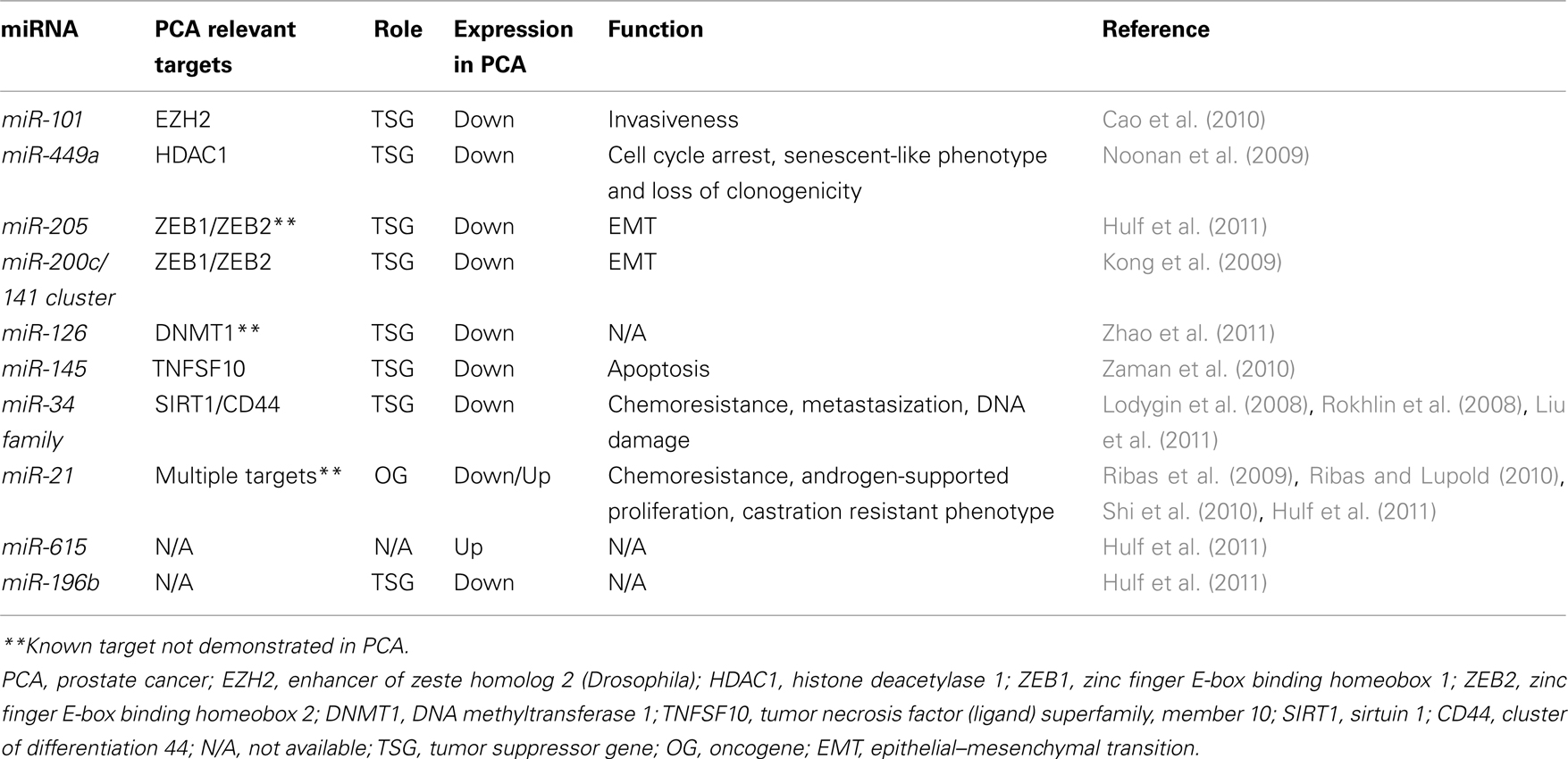
As previously observed in lung cancer (Fabbri et al., 2007a) and hematological malignancies (Garzon et al., 2009), also in PCA miRNAs have been shown to target key effectors of the epigenetic machinery (which might act either as OGs or as TSGs themselves), or they could affect the expression of epigenetically regulated OGs or TSGs. These two scenarios have been disclosed for miR-101, regulating EZH2, and miR-449a, regulating HDAC1 (cfr. Table 1).
Enhancer of zeste homolog 2 is part of multiprotein complex called polycomb repressive complex 2 (PRC2). These proteins regulate the tri-methylation of histone H3K27 of several target gene promoters leading to their epigenetic silencing. EZH2 is considered an OG since, when overexpressed, it induces cell proliferation, increases tumor colony formation and increases tumor invasion both in vitro and in vivo (Varambally et al., 2008). EZH2 is overexpressed in hormone refractory, metastatic PCA (Varambally et al., 2008). MiR-101 expression is frequently reduced in PCA (Lu et al., 2005; Volinia et al., 2006), and it has been demonstrated that miR-101 negatively regulates EZH2 expression. Restoration of miR-101 expression in PCA cells attenuates tumor invasiveness. Intriguingly, both AR and HIF-1-alpha and -beta seem to be able to induce miR-101 expression, suggesting that miR-101 levels depend on specific physiological conditions, and are affected by changes in the surrounding microenvironment (Cao et al., 2010).
Also HDAC1 has been demonstrated to be overexpressed in PCA (Patra et al., 2001; Halkidou et al., 2004; Weichert et al., 2008), and an inverse correlation has been observed between HDAC1 and miR-449a expression in PCA (Noonan et al., 2009). In the PCA cell line PC3, miR-449a directly targets HDAC1, and miR-449a restoration leads to cell cycle arrest, acquisition of a senescent-like phenotype, and loss of clonogenicity (Noonan et al., 2009). Part of this observed phenotype could be also mediated by over-expression of p27 (a known target of HDCA1), upon HDCA1 silencing by miR-449a (Noonan et al., 2009).
Despite some of the miRNAs regulating proteins of the epigenetic machinery have been identified, there are still several of these effectors for which such a regulation can be hypothesized and the involved miRNAs are being investigated. Therefore, it is likely that in the near future an increasing number of these miRNAs will be identified and will contribute to a better understanding of the actual function of the targeted epigenetic effectors themselves.
miRNAs are Epigenetically Regulated in Prostate Cancer
MicroRNAs undergo the same regulatory mechanisms of any other PCG, therefore the genes encoding for miRNAs can also be epigenetically regulated (Fabbri, 2008). In several types of cancer (including PCA), aberrations of this epigenetic regulation of miRNAs are able to explain, at least in part, some of the specific miRNome abnormalities which characterize that specific malignant phenotype.
Recently, an integrative approach has been successfully used to show that the expression of several miRNAs is modified by epigenetic modulation in PCA (Hulf et al., 2011). By combining primary transcription analysis, genome-wide DNA methylation and H3K9Ac marks with miRNA expression, Hulf et al. (2011) demonstrated that at least three miRNAs are down-regulated (namely, miR-205, -21, and -196b) and one upregulated (miR-615) by epigenetic mechanisms in PCA. Interestingly, one of the down-regulated miRNAs, miR-205, is involved in the epithelial–mesenchymal transition (EMT) process, and together with another family of miRNAs (namely, the miR-200 family) it has been extensively studied in several types of cancers, including PCA (Kong et al., 2009). The miR-200 family includes miR-200a, miR-200b miR-200c, miR-141, and miR-429. MiRNAs of the miR-200 family are organized in two clusters, one (encoding for miR-200b, miR-200a, and miR-429) on chromosome 1, while the other (encoding for miR-200c and miR-141) is located on chromosome 12. MiR-200 family and miR-205 directly target the E-cadherin transcriptional repressors ZEB1 and ZEB2, which are implicated in EMT and affect the formation of metastases, by controlling the adhesion and invasion processes (Gregory et al., 2008). Recently, Kong et al. (2009) reported that the miR-200c/miR-141 cluster is transcribed as a polycistronic RNA, and is silenced by promoter hypermethylation in several cancer types including PCA. Indeed, treatment of PC3 cells with the de-methylating agent 5-aza-2′-deoxycytidine (5-AZA) induces re-expression of both miRNAs of the cluster, proving that aberrant epigenetic mechanisms participate in the inappropriate repression of the miR-200c/miR-141 cluster, and this could affect the metastatic ability of PCA cells.
Interestingly, the epigenetic regulation of miRNAs appears to be cancer-type specific. Indeed, while the expression of miR-196b is reduced in PCA, miR-196b is upregulated in lymphoblastic leukemia and gastric cancer cells (Schotte et al., 2010; Tsai et al., 2010) because of a reduced degree of methylation at the same CpG island which are hypermethylated in PCA, and are responsible for miR-196b silencing in PCA. This finding corroborates the dual nature or miRNAs, able to act both as OGs and TSGs (Fabbri et al., 2007b), but also indicates that different strengths of epigenetic control can be exerted by cancer cells on the same gene, and the extent of this control depends on other yet not clearly identified intracellular factors, which ultimately determine whether an epigenetically regulated miRNA is silenced or not, in a cell-type specific fashion. Interestingly both miR-615 and miR-196b are located in CpG islands within the HOX gene clusters, but while the HOXC cluster host gene has been previously demonstrated to be part of a long range hypermethylated domain (Coolen et al., 2010), miR-615, located within this cluster is actually hypomethylated in PCA. This finding confirms that miRNA epigenetic regulation is strictly “tailored” and separate for miRNAs and their host genes.
Also miR-126, frequently down-regulated in PCA and in bladder cancer, undergoes epigenetic regulation, since the treatment of several cancer cell lines with 5-AZA or PBA (4-phenylbutyric acid), which is an HDAC inhibitor strongly increases miR-126 expression (Saito et al., 2009). Interestingly, a recent study reported upregulation of miR-126 in CD4+ T cells of patients with systemic lupus erythematosus (SLE), and this favors the development of this disease (Zhao et al., 2011). Moreover, this study identified a feed-forward loop in which miR-126 directly targets DNMT1, leading to a subsequent hypomethylation of miR-126 promoter, and increased levels of miR-126 (Zhao et al., 2011).
Based on these considerations it would be interesting to investigate whether a “reversed loop” occurs in PCA, where low miR-126, associated to higher expression of DNMT1, contributes in keeping the miRNA itself under epigenetic silencing.
Another miRNA frequently down-regulated in PCA with respect to the normal tissue counterpart is miR-145. Combinatory treatment with 5-AZA and the HDAC inhibitor TCA (trichostatin-A) increases miR-145 expression, suggesting that also in this case the silencing is due to epigenetic modulation. Exogenous over-expression of miR-145 in PC3 cells induces the expression of the pro-apoptotic gene TNFSF10 leading the cells to apoptosis (Zaman et al., 2010).
In PCA also miR-193b is silenced by hypermethylation of a CpG island located about 1 kb upstream of the miRNA locus (Rauhala et al., 2010). Re-expression of miR-193b in 22Rv1 PCA cells induced a significant tumor growth inhibition, as a result of a reduced number of cells in the S-phase of the cell cycle. In addition, the anchorage independent growth of this same cell line was partially inhibited upon transient expression of miR-193b (Rauhala et al., 2010).
Another miRNA tightly controlled by promoter methylation in PCA is miR-34a. Silencing of miR-34a expression is presumably mediated by CpG methylation of a region located 100–500 base-pairs upstream of the miR-34a transcription start which includes a binding site for p53, which transactivates miR-34a transcription (Raver-Shapira et al., 2007). The different miR-34a expression levels among DU145, PC3, and LNCaP cell lines can only partially be explained by the promoter methylation status of the miRNA, therefore it can be hypothesized that the endogenous levels of p53 (different among the different cell lines) contribute to the overall endogenous expression of this miRNA. Exogenous re-expression of miR-34a in PC3 cells induces cellular senescence, and permanent cell cycle arrest (Lodygin et al., 2008). Finally, miR-34a has been involved in the control of cancer stem cells population balance in PCA. Liu et al. (2011) observed that exogenous over-expression of miR-34a in CD44 positive PCA cells (that are considered PCA stem cells) inhibits clonogenic expansion, tumor regeneration, and metastasis. In contrast, miR-34a silencing in CD44 negative PCA cells, achieved by antagomirs (Krutzfeldt et al., 2005), promoted tumor development and metastasis, confirming that miR-34a has an important TSG function in PCA.
Despite miR-21 is generally considered a miRNA which acts as an OG, its role in PCA is still unclear, given the fact that conflicting results have been published. Hulf et al. (2011) demonstrated a significant epigenetic repression of miR-21 in PCA, suggesting a TSG role for miR-21 in PCA. However, Ribas et al. (2009) found that over-expression of miR-21 was sufficient to drive androgen-independent growth in two androgen-dependent PCA cell lines. Additionally, they observed that miR-21 was enhancing androgen-supported proliferation of LNCaP cells. In mouse models, elevated levels of miR-21 improved tumor establishment, increased tumor growth, and induced a castration-resistant phenotype (Ribas and Lupold, 2010). Overall, these experiments support an OG role for miR-21, or at least suggest that miR-21 might be implicated in the switch from hormone-sensitive to hormone-resistant PCA. Recently, however Folini et al. (2010) showed that miR- 21 knockdown elicited no change in proliferative or invasive properties in PCA cells. In the same paper the authors were unable to confirm a significant difference in the expression of miR-21 in PCA versus normal tissue on a cohort of 36 patients. In summary, a clear role for miR-21 in prostate carcinogenesis has not been defined yet, and further studies are warranted to clarify this issue.
Interestingly, both miR-34 and miR-21 are also involved in chemoresistance of PCA. MiR-34 downregulation is associated with the HDAC SIRT1 over-expression, this protein has been described to be associated with chemoresistance to camptothecin and cisplatin in PC3 and DU145 cells (Fujita et al., 2008). Artificial introduction of miR-34a in PC3 cells induces apoptosis and attenuates chemoresistance to camptothecin through the targeting of SIRT1 and probably others cell cycle regulators, previously described to be targets of miR-34a such as CDK4, CDK6, Cyclin D1, Cyclin E2, E2F3, and BCL2 (Hermeking, 2007; Fujita et al., 2008).
The involvement of miR-21 in PCA chemoresistance has been demonstrated by Shi et al. (2010). They showed that miR-21 is upregulated in docetaxel resistant PC3 (PC3R) cells compared to wild type PC3 cells. This process is mediated by the ability of miR-21 to target the oncosuppressor gene PDCD4 that is known to be involved in the sensitivity to other chemotherapic agents like cisplatin and paclitaxel (Shi et al., 2010).
Several studies have reported that both miR-21 and miR-34a can act as PCA biomarkers. But while miR-34a has been shown to be silenced by promoter hypermethylation in PCA, and consequently its expression in the blood of PCA patients is reduced with respect to non-cancerous patients (Kojima et al., 2010), for miR-21 there is the conflicting evidence of low expression in primary prostate tumors (due to promoter hypermethylation) but high expression in the serum of patients with hormone-refractory PCA (Yaman Agaoglu et al., 2011; Zhang et al., 2011). Again, these findings highlights the need to better understand what is the role of miR-21 in prostate carcinogenesis and in particular how its epigenetic regulation affects the shift toward hormone-refractory PCA.
Epigenetics of the Androgen Receptor: Role of miRNAs
Androgen receptor is the main actor in the development and progression of PCA and controlling its activity represents one of the most important goals in the clinical management of PCA. Despite its relevance for PCA, the mechanisms that regulate the expression of this protein remain poorly understood. While in some cases AR is not functional because of genetic mutations, in up to 40% of PCA patients (and in the DU145 cell line as well) it has been found to be silenced by promoter hypermethylation (Jarrard et al., 1998; Kinoshita et al., 2000; Nakayama et al., 2000; Sasaki et al., 2002; Yamanaka et al., 2003; Reibenwein et al., 2007), revealing a remarkable role of epigenetics in AR expression regulation. Other described mechanisms of AR aberrant expression and function include genetic amplification, generation of ligand-independent receptor variants, downregulation of AR repressors, persistence of intratumoral androgens, or upregulation of signals that activate AR also in absence of androgens (for a complete review see Shen and Abate-Shen, 2010).
In PCA, AR can function in two opposite directions: on one side, AR signaling is crucial for prostate and PCA cell survival (Shen and Abate-Shen, 2010), however its activation can also limit cell proliferation and mediate apoptotic induction under specific circumstances (Shen and Abate-Shen, 2010). For example, under genotoxic stress AR is fundamental for p53 activation and for the subsequent induction of apoptosis (Heinlein and Chang, 2004; Rokhlin et al., 2008). Since it has been estimated that miRNAs can regulate the expression of about 30% of the human genes (Lewis et al., 2005), there has been interest in investigating whether miRNAs can regulate the expression of AR. Despite a large number of miRNAs has been shown to be able to regulate AR expression (Sikand et al., 2010; Ostling et al., 2011), there is a substantial gap between the basic research findings and the expression of these miRNAs in clinical samples, raising the question of the relevance of these results for translational research purposes.
Conversely, it can be expected that reduced expression of AR-targeting miRNAs is responsible for AR increased expression. Recently, Ostling et al. (2011) demonstrated that AR-3′-UTR is much longer than contemplated by the most frequently used miRNA target prediction programs. In particular, the authors demonstrated that at least 71 miRNAs affect AR expression in PCA cell lines, and they confirmed that 13 of them directly bind to and regulate the 6-kb extended 3′-UTR of AR. Among them, miR-34a and miR-34c are particularly interesting, since, as we previously described, they are epigenetically regulated in PCA, and could represent the missing link between epigenetics–miRNAs and AR regulation in PCA (Figure 1). In 2008, Rokhlin et al. (2008) demonstrated that miR-34a and miR-34c are involved in the apoptotic induction after p53 activation in PCA cells, and that pro-apoptotic functions of DNA damage-induced p53 and miR-34s are dependent upon AR activation. Therefore, a central role of the epigenetically regulated miR-34 family members in the physiology of AR in PCA can be claimed (Figure 1).
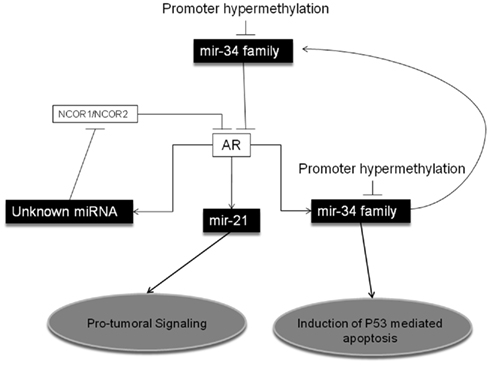
Figure 1. MicroRNAs and Epigenetic Regulation of the Androgen Receptor in Prostate Cancer. MiR-34 family has been shown to directly target the Androgen Receptor (AR) in Prostate Cancer (PCA). Also, miR-34 family can be transactivated by AR, leading to a feedback regulatory loop, and to p53-mediated pro-apoptotic effects. Promoter hypermethylation has been described as a main epigenetic regulatory mechanism for miR-34 family expression in PCA. Also miR-21 is transactivated by AR, but its over-expression promotes an oncogenic signaling. Finally, activation of AR has been shown to induce yet not-better identified miRNAs, which target NCOR1 (nuclear receptor corepressor 1) and NCOR2 (nuclear receptor corepressor 2), co-repressors of AR.
In addition to miR-34s, other miRNAs have been reported to be controlled by AR. Narayanan et al. (2010) demonstrated that by blocking miRNAs maturation most of the functions of AR are also blocked. Specifically, they observed that the silencing of DICER (a key enzyme in miRNA biogenesis) prevents DHT (dihydrotestosterone)-induced PSA production. They also demonstrated that by silencing DICER, DHT activation of AR results in upregulation of NCOR1 and NCOR2, two well characterized AR co-repressors, a mechanism bona fide miRNA-mediated, despite the actual miRNAs involved have not been identified in this study (Figure 1). Also miR-21 has been demonstrated to be directly transactivated by AR, through an AR binding site on the miRNA promoter (Ribas et al., 2009). These findings corroborate the need for a deeper investigation on the role of miR-21 in the acquisition of a hormone-resistant PCA phenotype. Finally, it has been demonstrated that a group of five miRNAs (namely miR-141, -494, 29a, -29b, and -29c) which directly target effectors of the epigenetic machinery, are also induced by DHT stimulation in VCaP and AR-expressing LNCaP cells (Waltering et al., 2010). This finding casts a new light on the interaction between AR function in PCA and miRNA controlling key enzymes ruling gene epigenetic regulation.
Conclusion
The genetics of PCA is complex and only partially understood. In recent years, the discovery that a specific subgroup of ncRNAs, called miRNAs, is implicated in the pathogenesis and dissemination of this cancer has shown that the molecular biology of PCA is more complex than initially thought. Moreover, the finding that epigenetic mechanisms are responsible for some of the aberrations of the miRNome which are hallmarks of PCA and/or its resistance to drugs has introduced a new layer of complexity to the picture. Despite this apparent intricacy, an accurate dissection of all the molecular mechanisms responsible for the pathogenesis and clinical behavior of this disease is the absolutely necessary background to understand which are the best molecular targets for the treatment of PCA. Undoubtedly, for some aspects, we are still at the dawn of our comprehension. Unconfirmed data and, in some cases, even conflicting results witness the need for a deeper insight into the intertwined mechanisms that drive cells to develop a malignant phenotype. Therefore (and almost paradoxically), the discovery of an increased complex interaction between well known OGs and TSGs, epigenetics and new characters like the miRNAs, ends up being one of the fastest ways to identify new promising therapeutic targets, especially (yet not exclusively) for those patients who still have an uncurable PCA.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Dr. Fabbri is supported by a 2009 Kimmel Scholar Award.
References
Cao, P., Deng, Z., Wan, M., Huang, W., Cramer, S. D., Xu, J., Lei, M., and Sui, G. (2010). MicroRNA-101 negatively regulates Ezh2 and its expression is modulated by androgen receptor and HIF-1alpha/HIF-1beta. Mol. Cancer 9, 108.
Coolen, M. W., Stirzaker, C., Song, J. Z., Statham, A. L., Kassir, Z., Moreno, C. S., Young, A. N., Varma, V., Speed, T. P., Cowley, M., Lacaze, P., Kaplan, W., Robinson, M. D., and Clark, S. J. (2010). Consolidation of the cancer genome into domains of repressive chromatin by long-range epigenetic silencing (LRES) reduces transcriptional plasticity. Nat. Cell Biol. 12, 235–246.
Croce, C. M. (2009). Causes and consequences of microRNA dysregulation in cancer. Nat. Rev. Genet. 10, 704–714.
Dillhoff, M., Wojcik, S. E., and Bloomston, M. (2008). MicroRNAs in Solid Tumors. J. Surg. Res. 154, 349–354.
Fabbri, M., and Croce, C. M. (2011). Role of microRNAs in lymphoid biology and disease. Curr. Opin. Hematol. 18, 266–272.
Fabbri, M., Garzon, R., Cimmino, A., Liu, Z., Zanesi, N., Callegari, E., Liu, S., Alder, H., Costinean, S., Fernandez-Cymering, C., Volinia, S., Guler, G., Morrison, C. D., Chan, K. K., Marcucci, G., Calin, G. A., Huebner, K., and Croce, C. M. (2007a). MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl. Acad. Sci. U.S.A. 104, 15805–15810.
Fabbri, M., Ivan, M., Cimmino, A., Negrini, M., and Calin, G. A. (2007b). Regulatory mechanisms of microRNAs involvement in cancer. Expert Opin. Biol. Ther. 7, 1009–1019.
Folini, M., Gandellini, P., Longoni, N., Profumo, V., Callari, M., Pennati, M., Colecchia, M., Supino, R., Veneroni, S., Salvioni, R., Valdagni, R., Daidone, M. G., and Zaffaroni, N. (2010). MiR-21: an oncomir on strike in prostate cancer. Mol. Cancer 9, 12.
Fujita, Y., Kojima, K., Hamada, N., Ohhashi, R., Akao, Y., Nozawa, Y., Deguchi, T., and Ito, M. (2008). Effects of miR-34a on cell growth and chemoresistance in prostate cancer PC3 cells. Biochem. Biophys. Res. Commun. 377, 114–119.
Garzon, R., Liu, S., Fabbri, M., Liu, Z., Heaphy, C. E., Callegari, E., Schwind, S., Pang, J., Yu, J., Muthusamy, N., Havelange, V., Volinia, S., Blum, W., Rush, L. J., Perrotti, D., Andreeff, M., Bloomfield, C. D., Byrd, J. C., Chan, K., Wu, L. C., Croce, C. M., and Marcucci, G. (2009). MicroRNA-29b induces global DNA hypomethylation and tumor suppressor gene re-expression in acute myeloid leukemia by targeting directly DNMT3A and 3B and indirectly DNMT1. Blood 113, 6411–6418.
Gregory, P. A., Bert, A. G., Paterson, E. L., Barry, S. C., Tsykin, A., Farshid, G., Vadas, M. A., Khew-Goodall, Y., and Goodall, G. J. (2008). The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat. Cell Biol. 10, 593–601.
Halkidou, K., Gaughan, L., Cook, S., Leung, H. Y., Neal, D. E., and Robson, C. N. (2004). Upregulation and nuclear recruitment of HDAC1 in hormone refractory prostate cancer. Prostate 59, 177–189.
Heinlein, C. A., and Chang, C. (2004). Androgen receptor in prostate cancer. Endocr. Rev. 25, 276–308.
Hulf, T., Sibbritt, T., Wiklund, E. D., Bert, S., Strbenac, D., Statham, A. L., Robinson, M. D., and Clark, S. J. (2011). Discovery pipeline for epigenetically deregulated miRNAs in cancer: integration of primary miRNA transcription. BMC Genomics 12, 54.
Jarrard, D. F., Kinoshita, H., Shi, Y., Sandefur, C., Hoff, D., Meisner, L. F., Chang, C., Herman, J. G., and Isaacs, W. B. (1998). Nassif N Methylation of the androgen receptor promoter CpG island is associated with loss of androgen receptor expression in prostate cancer cells. Cancer Res. 58, 5310–5314.
Kinoshita, H., Shi, Y., Sandefur, C., Meisner, L. F., Chang, C., Choon, A., Reznikoff, C. R., Bova, G. S., Friedl, A., and Jarrard, D. F. (2000). Methylation of the androgen receptor minimal promoter silences transcription in human prostate cancer. Cancer Res. 60, 3623–3630.
Kojima, K., Fujita, Y., Nozawa, Y., Deguchi, T., and Ito, M. (2010). MiR-34a attenuates paclitaxel-resistance of hormone-refractory prostate cancer PC3 cells through direct and indirect mechanisms. Prostate 70, 1501–1512.
Kong, D., Li, Y., Wang, Z., Banerjee, S., Ahmad, A., Kim, H. R., and Sarkar, F. H. (2009). MiR-200 regulates PDGF-D-mediated epithelial-mesenchymal transition, adhesion, and invasion of prostate cancer cells. Stem Cells 27, 1712–1721.
Krutzfeldt, J., Rajewsky, N., Braich, R., Rajeev, K. G., Tuschl, T., Manoharan, M., and Stoffel, M. (2005). Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438, 685–689.
Lewis, B. P., Burge, C. B., and Bartel, D. P. (2005). Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20.
Liu, C., Kelnar, K., Liu, B., Chen, X., Calhoun-Davis, T., Li, H., Patrawala, L., Yan, H., Jeter, C., Honorio, S., Wiggins, J. F., Bader, A. G., Fagin, R., Brown, D., and Tang, D. G. (2011). The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat. Med. 17, 211–215.
Lodygin, D., Tarasov, V., Epanchintsev, A., Berking, C., Knyazeva, T., Körner, H., Knyazev, P., Diebold, J., and Hermeking, H. (2008). Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle 7, 2591–2600.
Lu, J., Getz, G., Miska, E. A., Alvarez-Saavedra, E., Lamb, J., Peck, D., Sweet-Cordero, A., Ebert, B. L., Mak, R. H., Ferrando, A. A., Downing, J. R., Jacks, T., Horvitz, H. R., and Golub, T. R. (2005). MicroRNA expression profiles classify human cancers. Nature 435, 834–838.
Nakayama, T., Watanabe, M., Suzuki, H., Toyota, M., Sekita, N., Hirokawa, Y., Mizokami, A., Ito, H., Yatani, R., and Shiraishi, T. (2000). Epigenetic regulation of androgen receptor gene expression in human prostate cancers. Lab. Invest. 80, 1789–1796.
Narayanan, R., Jiang, J., Gusev, Y., Jones, A., Kearbey, J. D., Miller, D. D., Schmittgen, T. D., and Dalton, J. T. (2010). MicroRNAs are mediators of androgen action in prostate and muscle. PLoS ONE 5, e13637.
Noonan, E. J., Place, R. F., Pookot, D., Basak, S., Whitson, J. M., Hirata, H., Giardina, C., and Dahiya, R. (2009). MiR-449a targets HDAC-1 and induces growth arrest in prostate cancer. Oncogene 28, 1714–1724.
Ostling, P., Leivonen, S. K., Aakula, A., Kohonen, P., Mäkelä, R., Hagman, Z., Edsjö, A., Kangaspeska, S., Edgren, H., Nicorici, D., Bjartell, A., Ceder, Y., Perälä, M., and Kallioniemi, O. (2011). Systematic analysis of microRNAs targeting the androgen receptor in prostate cancer cells. Cancer Res. 71, 1956–1967.
Patra, S. K., Patra, A., and Dahiya, R. (2001). Histone deacetylase and DNA methyltransferase in human prostate cancer. Biochem. Biophys. Res. Commun. 287, 705–713.
Rauhala, H. E., Jalava, S. E., Isotalo, J., Bracken, H., Lehmusvaara, S., Tammela, T. L., Oja, H., and Visakorpi, T. (2010). MiR-193b is an epigenetically regulated putative tumor suppressor in prostate cancer. Int. J. Cancer 127, 1363–1372.
Raver-Shapira, N., Marciano, E., Meiri, E., Spector, Y., Rosenfeld, N., Moskovits, N., Bentwich, Z., and Oren, M. (2007). Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol. Cell 26, 731–743.
Reibenwein, J., Pils, D., Horak, P., Tomicek, B., Goldner, G., Worel, N., Elandt, K., and Krainer, M. (2007). Promoter hypermethylation of GSTP1, AR, and 14-3-3sigma in serum of prostate cancer patients and its clinical relevance. Prostate 67, 427–432.
Ribas, J., and Lupold, S. E. (2010). The transcriptional regulation of miR-21, its multiple transcripts, and their implication in prostate cancer. Cell Cycle 9, 923–929.
Ribas, J., Ni, X., Haffner, M., Wentzel, E. A., Salmasi, A. H., Chowdhury, W. H., Kudrolli, T. A., Yegnasubramanian, S., Luo, J., Rodriguez, R., Mendell, J. T., and Lupold, S. E. (2009). MiR-21: an androgen receptor-regulated microRNA that promotes hormone-dependent and hormone-independent prostate cancer growth. Cancer Res. 69, 7165–7169.
Rokhlin, O. W., Scheinker, V. S., Taghiyev, A. F., Bumcrot, D., Glover, R. A., and Cohen, M. B. (2008). MicroRNA-34 mediates AR-dependent p53-induced apoptosis in prostate cancer. Cancer Biol. Ther. 7, 1288–1296.
Saito, Y., Friedman, J. M., Chihara, Y., Egger, G., Chuang, J. C., and Liang, G. (2009). Epigenetic therapy upregulates the tumor suppressor microRNA-126 and its host gene EGFL7 in human cancer cells. Biochem. Biophys. Res. Commun. 379, 726–731.
Sasaki, M., Tanaka, Y., Perinchery, G., Dharia, A., Kotcherguina, I., Fujimoto, S., and Dahiya, R. (2002). Methylation and inactivation of estrogen, progesterone, and androgen receptors in prostate cancer. J. Natl. Cancer Inst. 94, 384–390.
Schotte, D., Lange-Turenhout, E. A., Stumpel, D. J., Stam, R. W., Buijs-Gladdines, J. G., Meijerink, J. P., Pieters, R., and Den Boer, M. L. (2010). Expression of miR-196b is not exclusively MLL-driven but is especially linked to activation of HOXA genes in pediatric acute lymphoblastic leukemia. Haematologica 95, 1675–1682.
Shen, M. M., and Abate-Shen, C. (2010). Molecular genetics of prostate cancer: new prospects for old challenges. Genes Dev. 24, 1967–2000.
Shi, G. H., Ye, D. W., Yao, X. D., Zhang, S. L., Dai, B., Zhang, H. L., Shen, Y. J., Zhu, Y., Zhu, Y. P., Xiao, W. J., and Ma, C. G. (2010). Involvement of microRNA-21 in mediating chemo-resistance to docetaxel in androgen-independent prostate cancer PC3 cells. Acta Pharmacol. Sin. 31, 867–873.
Sikand, K., Slaibi, J. E., Singh, R., Slane, S. D., and Shukla, G. C. (2010). MiR 488* inhibits androgen receptor expression in prostate carcinoma cells. Int. J. Cancer. 129, 810–819.
Tsai, K. W., Hu, L. Y., Wu, C. W., Li, S. C., Lai, C. H., Kao, H. W., Fang, W. L., and Lin, W. C. (2010). Epigenetic regulation of miR-196b expression in gastric cancer. Genes Chromosomes Cancer 49, 969–980.
Varambally, S., Cao, Q., Mani, R. S., Shankar, S., Wang, X., Ateeq, B., Laxman, B., Cao, X., Jing, X., Ramnarayanan, K., Brenner, J. C., Yu, J., Kim, J. H., Han, B., Tan, P., Kumar-Sinha, C., Lonigro, R. J., Palanisamy, N., Maher, C. A., and Chinnaiyan, A. M. (2008). Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science 322, 1695–1699.
Vasudevan, S., Tong, Y., and Steitz, J. A. (2007). Switching from repression to activation: microRNAs can up-regulate translation. Science 318, 1931–1934.
Volinia, S., Calin, G. A., Liu, C. G., Ambs, S., Cimmino, A., Petrocca, F., Visone, R., Iorio, M., Roldo, C., Ferracin, M., Prueitt, R. L., Yanaihara, N., Lanza, G., Scarpa, A., Vecchione, A., Negrini, M., Harris, C. C., and Croce, C. M. (2006). A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. U.S.A. 103, 2257–2261.
Waltering, K. K., Porkka, K. P., Jalava, S. E., Urbanucci, A., Kohonen, P. J., Latonen, L. M., Kallioniemi, O. P., Jenster, G., and Visakorpi, T. (2010). Androgen regulation of micro-RNAs in prostate cancer. Prostate 71, 604–614.
Weichert, W., Röske, A., Gekeler, V., Beckers, T., Stephan, C., Jung, K., Fritzsche, F. R., Niesporek, S., Denkert, C., Dietel, M., and Kristiansen, G. (2008). Histone deacetylases 1, 2 and 3 are highly expressed in prostate cancer and HDAC2 expression is associated with shorter PSA relapse time after radical prostatectomy. Br. J. Cancer 98, 604–610.
Yaman Agaoglu, F., Kovancilar, M., Dizdar, Y., Darendeliler, E., Holdenrieder, S., Dalay, N., and Gezer, U. (2011). Investigation of miR-21, miR-141, and miR-221 in blood circulation of patients with prostate cancer. Tumour Biol. 32, 583–588.
Yamanaka, M., Watanabe, M., Yamada, Y., Takagi, A., Murata, T., Takahashi, H., Suzuki, H., Ito, H., Tsukino, H., Katoh, T., Sugimura, Y., and Shiraishi, T. (2003). Altered methylation of multiple genes in carcinogenesis of the prostate. Int. J. Cancer 106, 382–387.
Zaman, M. S., Chen, Y., Deng, G., Shahryari, V., Suh, S. O., Saini, S., Majid, S., Liu, J., Khatri, G., Tanaka, Y., and Dahiya, R. (2010). The functional significance of microRNA-145 in prostate cancer. Br. J. Cancer 103, 256–264.
Zhang, H. L., Yang, L. F., Zhu, Y., Yao, X. D., Zhang, S. L., Dai, B., Zhu, Y. P., Shen, Y. J., Shi, G. H., and Ye, D. W. (2011). Serum miRNA-21: elevated levels in patients with metastatic hormone-refractory prostate cancer and potential predictive factor for the efficacy of docetaxel-based chemotherapy. Prostate 71, 326–331.
Keywords: microRNA, prostate cancer, epigenetics, androgen receptor, methylation, histone acetylation
Citation: Paone A, Galli R and Fabbri M (2011) MicroRNAs as new characters in the plot between epigenetics and prostate cancer. Front. Gene. 2:62. doi: 10.3389/fgene.2011.00062
Received: 06 July 2011; Paper pending published: 29 July 2011;
Accepted: 18 August 2011; Published online: 06 September 2011.
Edited by:
Don L. Gibbons, University of Texas, USAReviewed by:
Alan G. Ramsay, Queen Mary University of London, UKDong Xu, University of Missouri, USA
Copyright: © 2011 Paone, Galli and Fabbri. This is an open-access article subject to a non-exclusive license between the authors and Frontiers Media SA, which permits use, distribution and reproduction in other forums, provided the original authors and source are credited and other Frontiers conditions are complied with.
*Correspondence: Muller Fabbri, 1092 Biomedical Research Tower, 460 West 12th Avenue, Columbus, OH 43210, USA. e-mail:bXVsbGVyLmZhYmJyaUBvc3VtYy5lZHU=

 Roberta Galli
Roberta Galli