- Department of Materials Science and Biotechnology, Applied Chemistry, Graduate School of Science and Engineering, Ehime University, Matsuyama, Japan
To date, more than 90 modified nucleosides have been found in tRNA and the biosynthetic pathways of the majority of tRNA modifications include a methylation step(s). Recent studies of the biosynthetic pathways have demonstrated that the availability of methyl group donors for the methylation in tRNA is important for correct and efficient protein synthesis. In this review, I focus on the methylated nucleosides and tRNA methyltransferases. The primary functions of tRNA methylations are linked to the different steps of protein synthesis, such as the stabilization of tRNA structure, reinforcement of the codon-anticodon interaction, regulation of wobble base pairing, and prevention of frameshift errors. However, beyond these basic functions, recent studies have demonstrated that tRNA methylations are also involved in the RNA quality control system and regulation of tRNA localization in the cell. In a thermophilic eubacterium, tRNA modifications and the modification enzymes form a network that responses to temperature changes. Furthermore, several modifications are involved in genetic diseases, infections, and the immune response. Moreover, structural, biochemical, and bioinformatics studies of tRNA methyltransferases have been clarifying the details of tRNA methyltransferases and have enabled these enzymes to be classified. In the final section, the evolution of modification enzymes is discussed.
Introduction
The first tRNA sequence was determined in 1965 and numerous modifications were identified at various positions within the sequence (Holley et al., 1965). At almost the same time, several tRNA methyltransferase activities were detected in Escherichia coli cell extract (Hurwitz et al., 1964), which suggested that diverse enzymes are involved in tRNA modification. To date, more than 90 modified nucleosides have been identified in tRNA (Machnicka et al., 2013). Thus, the majority of modified nucleosides that have been discovered in different RNA species are found in tRNA. In the twenty-first century, the major modification pathways of tRNA have been elucidated on the basis of genome sequence data. These studies have demonstrated that the pathways of tRNA modification show diversity among living organisms. In this review, I focus on the methylated nucleosides in tRNA, together with tRNA methyltransferases, and introduce their basic roles as well as their more complex functions.
The Primary Role of tRNA Modifications is the Regulation of Protein Synthesis
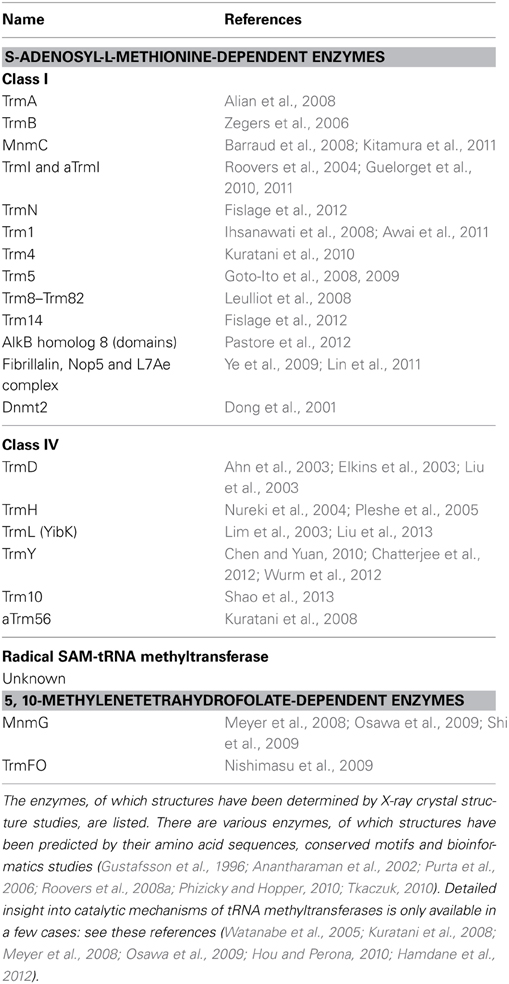
Transfer RNA is an adaptor molecule that enables the genetic code of nucleic acids to be converted to amino acids in protein. Consequently, the primary functions of individual tRNA modifications are linked to the different steps of protein synthesis. In fact, if a tRNA remains unmodified, it becomes charged with a non-cognate amino acid, the corresponding codon in the mRNA is mistranslated, and a mutation is introduced. Table 1 summarizes the typical methylated nucleosides and their positions within the tRNA, their distributions in the three domains of life, the corresponding tRNA methyltransferases, their contributions to tRNA structure, their functions in addition to structural roles, and related publications. (Phillips and Kjellin-Straby, 1967; Taya and Nishimura, 1973; Yaniv and Folk, 1975; Delk et al., 1976; Watanabe et al., 1976, 2005, 2006; Pierre et al., 1978, 2003; Pope et al., 1978; Raba et al., 1979; Greenberg and Dudock, 1980; Ny and Bjork, 1980; Osorio-Almeida et al., 1980; Byström and Björk, 1982; Hopper et al., 1982; Walker, 1983; Gupta, 1984; Johnson et al., 1985; Ellis et al., 1986; Reinhart et al., 1986; van Tol et al., 1987; Ny et al., 1988; Björk et al., 1989, 2001; Jakab et al., 1990; Keith et al., 1990; Perret et al., 1990; Edmonds et al., 1991; Gu and Santi, 1991; Gustafsson and Björk, 1993; Hagervall et al., 1993; Edqvist et al., 1994; Kowalak et al., 1994; Martin and Hopper, 1994; Grosjean et al., 1995, 1996, 2008; Durand et al., 1997; Jiang et al., 1997; Li et al., 1997; Persson et al., 1997, 1998; Anderson et al., 1998, 2000; Constantinesco et al., 1998, 1999a,b; Helm et al., 1998; Hori et al., 1998, 2002, 2003; Matsuyama et al., 1998; Qian et al., 1998; Tomita et al., 1998; Cavaillé et al., 1999; Farabaugh and Björk, 1999; Liu et al., 1999, 2003, 2013; Motorin and Grosjean, 1999; Niederberger et al., 1999; Liu and Straby, 2000; Nordlund et al., 2000; Clouet-d'Orval et al., 2001, 2005; Dong et al., 2001; Urbonavicius et al., 2001, 2002, 2003, 2005; Yasukawa et al., 2001; Alexandrov et al., 2002, 2005, 2006; Johansson and Byström, 2002; King and Redman, 2002; Pintard et al., 2002; Suzuki et al., 2002, 2007, 2011a; Ahn et al., 2003; Bortolin et al., 2003; De Bie et al., 2003; Droogmans et al., 2003; Elkins et al., 2003; Jackman et al., 2003; Kalhor and Clarke, 2003; Kaneko et al., 2003; Kierzek and Kierzek, 2003; Takai and Yokoyama, 2003; Armengaud et al., 2004; Brulé et al., 2004; Bujnicki et al., 2004; Christian et al., 2004, 2013; Freude et al., 2004; Kadaba et al., 2004; Nasvall et al., 2004; Nureki et al., 2004; O'Dwyer et al., 2004; Okamoto et al., 2004; Roovers et al., 2004, 2008a,b, 2012; Singh et al., 2004; Cartlidge et al., 2005; Chen et al., 2005, 2011; Durant et al., 2005; Huang et al., 2005; Kalhor et al., 2005; Kirino et al., 2005; Leipuviene and Björk, 2005; Lu et al., 2005; Pleshe et al., 2005; Purushothaman et al., 2005; Renalier et al., 2005; Sakurai et al., 2005a,b; Umeda et al., 2005; Waas et al., 2005, 2007; Brzezicha et al., 2006; Goll et al., 2006; McCrate et al., 2006; Noma et al., 2006, 2010, 2011; Ote et al., 2006; Purta et al., 2006; Shigi et al., 2006; Takano et al., 2006; Takeda et al., 2006; Yim et al., 2006; Zegers et al., 2006; Auxilien et al., 2007, 2011, 2012; Begley et al., 2007; Christian and Hou, 2007; Choudhury et al., 2007a,b; Lee et al., 2007; Matsumoto et al., 2007; Ozanick et al., 2007; Walbott et al., 2007; Wilkinson et al., 2007; Alian et al., 2008; Barraud et al., 2008; Chernyakov et al., 2008; Goto-Ito et al., 2008, 2009; Ihsanawati et al., 2008; Klassen et al., 2008; Kurata et al., 2008; Kuratani et al., 2008, 2010; Leulliot et al., 2008; Meyer et al., 2008, 2009; Tomikawa et al., 2008, 2010, 2013; Toyooka et al., 2008; Awai et al., 2009, 2011; Lai et al., 2009; Moukadiri et al., 2009, 2014; Nishimasu et al., 2009; Osawa et al., 2009; Shi et al., 2009; Shimada et al., 2009; Umitsu et al., 2009; Ye et al., 2009; Arragain et al., 2010; Atta et al., 2010, 2012; Benítez-Páez et al., 2010; Böhme et al., 2010; Chen and Yuan, 2010; de Crécy-Lagard et al., 2010; Fu et al., 2010; Guelorget et al., 2010, 2011; Kempenaers et al., 2010; Mazauric et al., 2010; Ochi et al., 2010, 2013; Songe-Møller et al., 2010; Tkaczuk, 2010; D'Silva et al., 2011; Hamdane et al., 2011a,b, 2012, 2013; Joardar et al., 2011; Kitamura et al., 2011, 2012; Leihne et al., 2011; Liger et al., 2011; Lin et al., 2011; Menezes et al., 2011; Pearson and Carell, 2011; Qiu et al., 2011; van den Born et al., 2011; Wei et al., 2011; Armengod et al., 2012; Chan et al., 2012; Chatterjee et al., 2012; Chujo and Suzuki, 2012; Dewe et al., 2012; Fislage et al., 2012; Gehrig et al., 2012; Guy et al., 2012; Jöckel et al., 2012; Novoa et al., 2012; Pastore et al., 2012; Patil et al., 2012a,b; Perche-Letuvée et al., 2012; Sakaguchi et al., 2012; Towns and Begley, 2012; Vilardo et al., 2012; Wurm et al., 2012; Yamagami et al., 2012; Edelheit et al., 2013; Fujimori, 2013; Igoillo-Esteve et al., 2013; Kim and Almo, 2013; Ohira et al., 2013; Paris et al., 2013; Preston et al., 2013; Shao et al., 2013; Swinehart et al., 2013). In Table 1, several important tRNA modifications such as pseudouridine (ψ), lysidine, agmatidine, queosine (Q), and 2-thiouridine (s2U) are not listed because their biosynthetic pathways do not include any methylation steps. Nevertheless, Table 1 outlines the roles of key tRNA modifications, and demonstrates that methylated nucleosides and tRNA methyltransferases are very important for such functions. The structures of typical methylated nucleosides are shown in Figure 1. It is impossible to depict all methylated nucleosides in Figure 1 due to limitations of space. Please visit the database (http://modomics.genesilico.pl/modifications/) to obtain additional structural information (Machnicka et al., 2013). The structure of tRNA and positions of the methylated nucleotides are shown in Figure 2. As for tRNA stabilization by methylated nucleosides, see this review (Motorin and Helm, 2010). Even today, the contributions to tRNA structure and/or function in protein synthesis of many methylated nucleosides remain unknown (Table 1). However, various tRNA methyltransferases and their corresponding disruptant strains have been analyzed, and their functions are gradually being elucidated. Among the phenotypes of the gene disruptant strains, many phenomena have been reported that are difficult to understand directly in terms of enzyme function or effects on protein synthesis. For example, E. coli miaA mutant strains, which contain A37 instead of ms2i6A37 in the tRNA, show a moderate mutator phenotype that results in an increased rate of GC -> AT transversion (Zhao et al., 2001). Furthermore, inosine 34 modification in fission yeast is essential for cell cycle progression (Tsutsumi et al., 2007). These phenomena might be caused by changes in the amount of certain protein(s), such as transcription factors, in the disruptant strains. In fact, recently, it has been reported that Trm9-specific tRNA modifications enhance codon-specific elongation of translation and promote increased levels of DNA damage response proteins (Begley et al., 2007). Furthermore, several eukaryotic tRNA methyltransferases (for example, human ALKBH8 Shimada et al., 2009; Fu et al., 2010 and yeast Trm2 Choudhury et al., 2007a,b) are involved directly in DNA repair and carcinogenesis because they exist as fusion proteins with other enzyme(s). However, it remains possible that some of the phenotypes observed in the disruptant strains are linked to unknown biological phenomena.
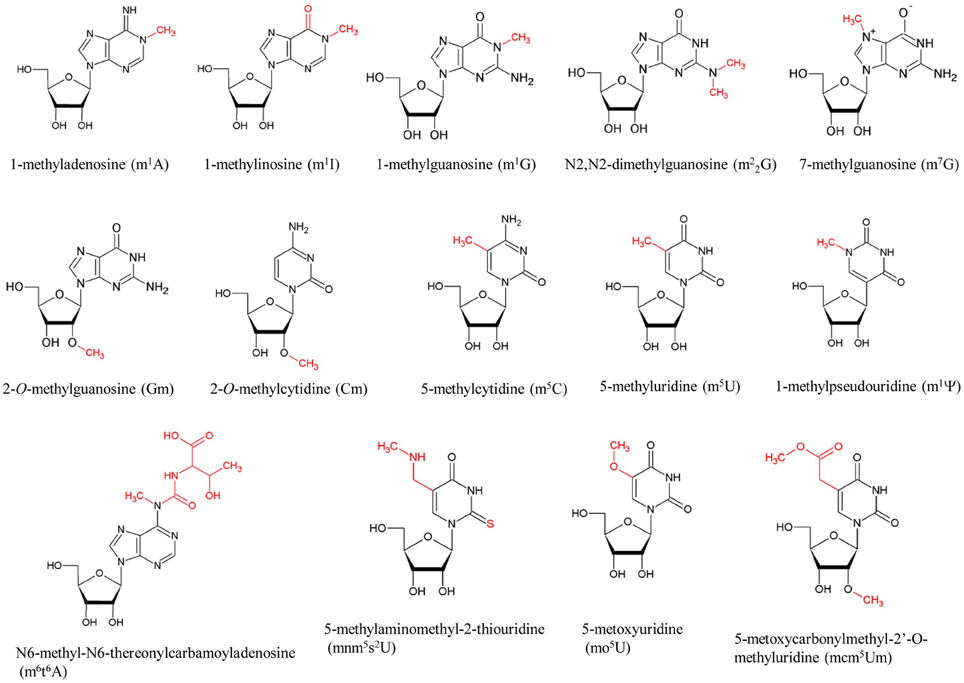
Figure 1. Typical methylated nucleosides in tRNA. The modification sites are colored in red. The abbreviations of the modified nucleosides are shown in parentheses.
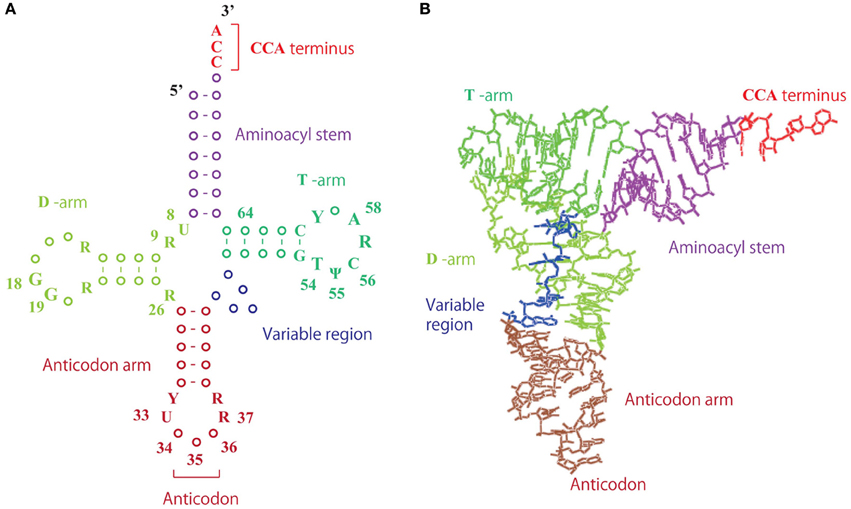
Figure 2. Structure of tRNA. (A) Clover-leaf structure of tRNA. The numbers show the positions of the nucleotides. (B) L-shaped tRNA structure. Transfer RNA forms an L-shaped structure, in which the D- and T-arms interact by tertiary base pairs.
Multiple Regulation of tRNA Modification Pathways and Importance of the Availability of Methyl Donors
In living cells, more than 50% of the high energy compounds such as ATP, that are produced by respiration are consumed by protein synthesis. Furthermore, the most important metabolic pathway of amino acids is protein synthesis. The metabolic pathways of energy and amino acids are closely linked. Studies on the pathways of tRNA modification have revealed that the RNA modification systems are located downstream of the pathways of energy and amino acid metabolism and that they are regulated at multiple steps (Herbig et al., 2002; Iwata-Reuyl, 2003; Ikeuchi et al., 2008, 2010; Shigi et al., 2008; Suzuki and Miyauchi, 2010; Phillips et al., 2012; Laxman et al., 2013; Miyauchi et al., 2013; Perrochia et al., 2013 Figure 3 and Table 1). Thus, depletion of a certain compound (for example, one of the amino acids) or disruption of a metabolic pathway can result in incomplete modification of tRNA and thus an increased frequency of translational errors.
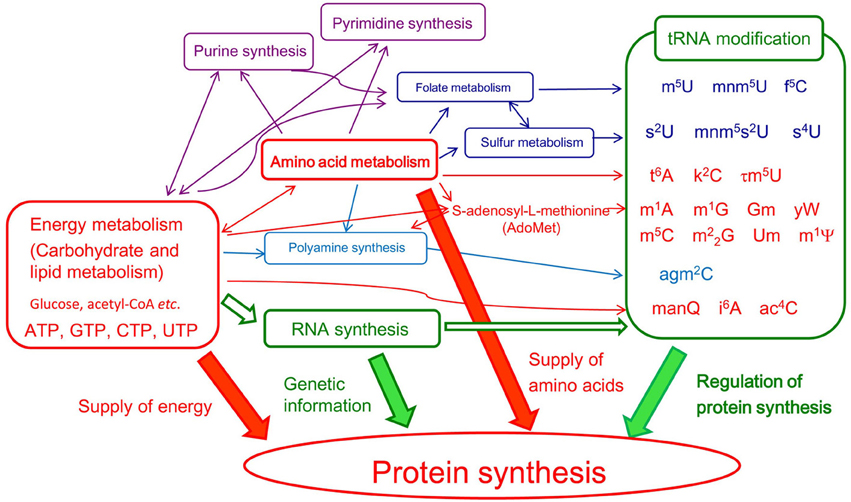
Figure 3. Transfer RNA modifications are regulated by various metabolic pathways. In this figure, conversions of chemical compounds and conceptual effects are shown by thin and thick arrows, respectively. To emphasis the relationships among energy metabolism, amino acid metabolism, tRNA modification, and protein synthesis, DNA replication and so on are omitted.
The structures of identified modified nucleosides suggest that the majority of tRNA modifications require a methylation step(s) (Table 1 and Figure 1). The methyl-transfer reaction by majority of tRNA methyltransferases consumes S-adenosyl-L-methionine (AdoMet) as the methyl-group donor. Thus, the depletion of AdoMet leads to multiple incomplete modifications in tRNA. The precursors of AdoMet are ATP and methionine. These facts seem to provide an answer for the question, “Why do living organisms use the methionine codon as the initiation codon for protein synthesis?” Under conditions where methionine is limited and the tRNA contains multiple incomplete modifications, to avoid increase of frequency of translational error, the methionine codon is selected the initiation codon of protein synthesis. Analogously, the fact that eubacterial methionyl-initiator tRNAMet is formylated and formylation is the transfer of one carbon atom suggests that the supply of sources of single carbon atoms is very important for efficient and accurate protein synthesis in bacteria.
Structures of tRNA Methyltransferases
Transfer RNA methyltransferases can be divided into two types on the basis of their methyl donor: one class uses AdoMet whereas the other utilizes 5, 10-methylenetetrahydrofolate (Table 2). As mentioned above, the majority of tRNA methyltransferases are AdoMet-dependent. For information on the catalytic mechanisms of tRNA methyltransferases, (see Watanabe et al., 2005; Kuratani et al., 2008; Meyer et al., 2008; Osawa et al., 2009; Hou and Perona, 2010; Hamdane et al., 2012). Recently, a radical SAM enzyme was identified as a ribosomal RNA methyltransferase (Atta et al., 2010); radical SAM enzymes utilize a 4Fe-4S cluster to generate a reactive radical from AdoMet. No radical SAM enzymes that act as tRNA methyltransferases have been identified as yet. However, three types of radical SAM enzymes are involved in tRNA modifications (2-methylthiotransferases that generate ms2t6A derivatives, 2-methylthiotransferases that generate ms2i6A derivatives, and enzymes involved in the biosynthesis of yW37 derivatives) (Suzuki et al., 2007; Atta et al., 2010, 2012; de Crécy-Lagard et al., 2010; Fujimori, 2013 and Table 1). Radical SAM tRNA methyltransferase(s) might be identified in the near future, because there are many methylated nucleosides, for which the responsible enzyme(s) have not yet been identified (Table 1).
AdoMet-dependent methyltransferases are classified by their catalytic domain (Schubert et al., 2003). Two different classes (classes I and IV) have been identified among the tRNA methyltransferases (Table 2). Class I enzymes contain the Rossmann fold in the catalytic domain (Figure 4A), whereas class IV enzymes have the topological-knot structure (Figure 4B). Class IV enzymes were predicted initially by bioinformatics studies to be members of the SpoU-TrmD (SPOUT) superfamily (Anantharaman et al., 2002). Subsequently, crystallographic studies (Table 2) revealed that these enzymes have a topological knot structure. YibK was predicted initially to be an RNA methyltransferase of unknown function (Gustafsson et al., 1996). Determination of the crystal structure revealed the presence of the topological-knot structure in the catalytic domain of YibK (Lim et al., 2003). Later, YibK was shown to function as tRNA (Cm34/cmnm5Um34) methyltransferase and was renamed TrmL (Benítez-Páez et al., 2010; Liu et al., 2013). At almost the same time, three groups independently reported the crystal structures of TrmD proteins and revealed that TrmD proteins also contain the topological-knot structure (Ahn et al., 2003; Elkins et al., 2003; Liu et al., 2003). In 1997, SpoU was found to have tRNA (Gm18) 2′-O-methyltransferase activity and was renamed as TrmH (Persson et al., 1997). We solved the crystal structure of TrmH in 2004 and confirmed that it is a class IV enzyme with the topological-knot structure (Nureki et al., 2004 and Figure 4C). These studies established the structural foundation of SPOUT enzymes (Anantharaman et al., 2002; Tkaczuk et al., 2007), which can be identified on the basis of the topological-knot structure. To date, several tRNA methyltransferases have been identified as members of the SPOUT superfamily on the basis of crystal structures (Kuratani et al., 2008; Chen and Yuan, 2010; Chatterjee et al., 2012; Wurm et al., 2012; Shao et al., 2013) or structures predicted from amino acid sequences and conserved motifs (Renalier et al., 2005; Purta et al., 2006; Tkaczuk et al., 2007; Kempenaers et al., 2010, and Figure 4B). Furthermore, the SPOUT superfamily is expanding beyond the SpoU and TrmD families: novel enzymes such as an archaeal Trm10 homolog (Kempenaers et al., 2010) and TrmY (Chen and Yuan, 2010; Chatterjee et al., 2012; Wurm et al., 2012) have been identified. These enzymes cannot be simply classified into the SpoU or TrmD families. Therefore, it might be necessary to reclassify the enzymes of the SPOUT superfamily on the basis of their structure, the methylated nucleosides produced, and their reaction mechanisms.
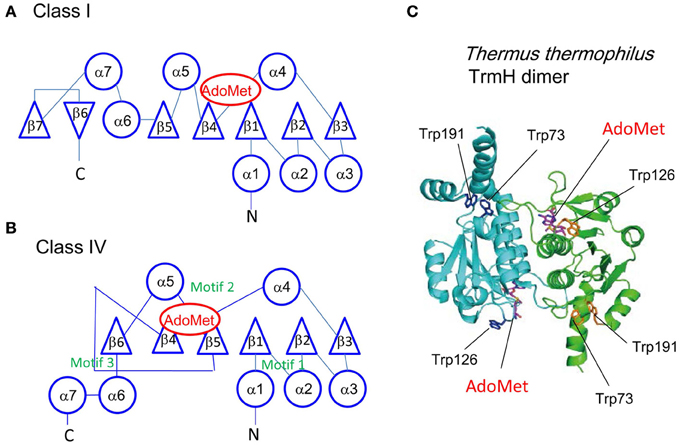
Figure 4. Structures of Rossman fold (Class I) and topological knot fold (Class IV). The topologies of class I (A) and IV (B) folds are compared. Circles and triangles show α-helices and β-strands, respectively. The AdoMet binding sites and three conserved motifs in the class IV are shown in red and green, respectively. The known class IV enzymes work as a dimer. (C) The dimer structure of T. thermophilus TrmH. T. thermophilus TrmH is a typical class IV enzyme. Fluorescence derived from three tryptophan residues (Trp73, Trp126, and Trp191) was monitored in the stopped-flow pre-steady state kinetic analysis as described in the main text. This figure is based on these publications Clouet-d'Orval et al. (2005), Ochi et al. (2013) with slight modifications.
The number of identified class I methyltransferases has also increased. Crystal structures of class I enzymes have been reported, as shown in Table 2; however, for many of the enzymes, structures have been predicted from their amino acid sequences and conserved motifs. The difficulty with crystallographic studies is that the eukaryotic and archaeal enzymes often require other subunit(s) to regulate (or stabilize) their activities (Anderson et al., 1998; Alexandrov et al., 2002, 2005; Purushothaman et al., 2005; Mazauric et al., 2010; Liger et al., 2011; Noma et al., 2011, and Table 1). Only a few structural studies of the multisubunit complexes have been performed, namely Trm8–Trm82 (Leulliot et al., 2008), and the Fibrillalin, Nop5 and L7Ae complex (Ye et al., 2009; Lin et al., 2011). In addition, structures for the tRNA bound-form of Trm5 (Goto-Ito et al., 2009) and T-arm-like RNA bound-form of TrmA (Alian et al., 2008) have been reported. Furthermore, several eukaryotic tRNA methyltransferases are fused with other functional domains and are involved in other processes such as DNA repair (Choudhury et al., 2007a,b; Shimada et al., 2009; Fu et al., 2010; Songe-Møller et al., 2010; D'Silva et al., 2011; Leihne et al., 2011; Noma et al., 2011; van den Born et al., 2011; Pastore et al., 2012). Although the crystal structures of the RNA recognition motif and AlkB domains of ALKB8H, which also contains a methyltransferase domain, have been reported (Pastore et al., 2012), there is no entire crystal structure of a eukaryotic multidomain tRNA methyltransferase. To understand the reaction mechanisms, substrate specificity, subunit (domain) interactions, and regulation of activity of these enzymes, structural studies are necessary.
Among the enzyme complexes that are involved in tRNA methylation, the mnmEG and mnmC complexes, which are required for the mnm5U34 modification (Taya and Nishimura, 1973; Bujnicki et al., 2004; Yim et al., 2006; Meyer et al., 2008, 2009; Roovers et al., 2008b; Moukadiri et al., 2009, 2014; Osawa et al., 2009; Shi et al., 2009; Böhme et al., 2010; Kitamura et al., 2011, 2012; Pearson and Carell, 2011; Armengod et al., 2012; Kim and Almo, 2013), are only found in eubacteria, which shows the complexity of the Xm5U34 biosynthetic pathway. In eukaryotes, the biosynthetic pathways of Xm5U34 have not been completely clarified: Trm9 and the so-called “Elongator” complex are known to be involved (Huang et al., 2005; Chen et al., 2011; Leihne et al., 2011). Furthermore, although we determined recently that tRNALeu from Thermoplasma acidophilum, a thermo-acidophilic archaeon, has 5-carbamoylmethyluridine at position 34 (ncm5U34) (Tomikawa et al., 2013), the biosynthetic pathway in archaea is unknown.
As studies on eukaryotic enzymes have progressed, the number of complex enzymes identified has increased. For example, mammalian enzymes often have additional domains, regulatory subunits and/or paralogs. For information on the identification and prediction of human tRNA methyltransferases, see this review (Towns and Begley, 2012).
Transfer RNA Recognition by tRNA Methyltransferases
Transfer RNA methyltransferases strictly modify a specific nucleoside at a specific position in a tRNA. Within the field of nucleic acid-related enzymes, a common question is “How does the enzyme recognize a specific substrate and act at a specific position?” Consequently, the substrate specificities of tRNA methyltransferases have been studied by measuring activities in crude cell extracts, microinjecting labeled tRNA, biochemical studies with purified enzymes, crystallographic studies, and analyses of tRNA from disruptant strains.
In general, tRNA methyltransferases recognize the local structure around the target site in the tRNA, including tertiary structural elements such as stem-loop structure(s). TrmA from E. coli recognizes U54 in the ribose-phosphate backbone of the T-arm (Gu and Santi, 1991; Alian et al., 2008). Aquifex aeolicus TrmB requires the five nucleotides AGG*UC sandwiched between two stem-loop structures (the asterisk corresponds to the methylation site, G46) (Okamoto et al., 2004). TrmFO recognizes the G53-C61 base pair and U54U55C56 sequence in the T-arm (Yamagami et al., 2012). TrmD recognizes the purine36G37 sequence in the anticodon-arm-like microhelix (Brulé et al., 2004; Takeda et al., 2006). In some cases, tertiary interactions are required. For example, crystallographic studies of the complex between Trm5 and tRNA revealed that the enzyme requires interaction between the D- and T-loop of the tRNA (Goto-Ito et al., 2009), which is consistent with the results of biochemical studies with the purified enzyme (Christian et al., 2004; Christian and Hou, 2007).
The target site for methylation is often embedded in the L-shaped tRNA structure. Consequently, in many (or almost all) cases, recognition of tRNA by tRNA methyltransferases seems to involve multiple steps (initial binding and induced fit processes). Although it is very difficult to prepare intermediate complexes, we recently analyzed the initial binding and changes in structure of TrmH by stopped-flow presteady-state kinetic analysis (Ochi et al., 2010, 2013). TrmH binds to tRNA within 10 ms in the initial binding process, in which substrate and non-substrate (methylated) tRNAs are not distinguished. Methylated tRNA is excluded from the complex subsequently due to steric hindrance between the methyl groups in the tRNA and AdoMet before the induced-fit process occurs. The advantage of this mechanism is that methylated tRNA does not severely inhibit the methyl-transfer reaction as a competitive inhibitor. Subsequently, in the induced-fit process, which takes more than 50 ms, G18 is recognized and ribose introduced into the catalytic pocket. During the induced-fit process, movement of Trp126 in motif 2 is observed (Ochi et al., 2013 and Figure 4C).
Several tRNA methyltransferases act on multiple sites in tRNA. For example, archaeal TrmI acts on both A57 and A58 (Roovers et al., 2004; Guelorget et al., 2010). Similarly, Aquifex aeolicus Trm1 acts on both G26 and G27 (Awai et al., 2009). On the basis of biochemical studies, we determined that this eubacterial Trm1 recognizes the methylation sites (G26 and G27) from the T-arm (Awai et al., 2009, 2011) whereas archaeal Trm1 recognizes G26 from the D-stem and variable region (Constantinesco et al., 1999b). These Trm1 proteins share high sequence homology (Awai et al., 2009); however, comparison of the crystal structures revealed that the distribution of positive charges on the enzyme surface differs between archaeal (Ihsanawati et al., 2008) and eubacterial (Awai et al., 2011) Trm1. Thus, these studies show how difficult it is to predict target sites on the basis of amino acid sequences. Furthermore, in some cases, other subunits regulate the site specificity. For example, the methylation site recognized by Trm7 is determined by its partner subunit (Guy et al., 2012) and the site specificity of archaeal Trm4 changes in the presence of archease (Auxilien et al., 2007). Moreover, the m5C modifications in eukaryotic tRNA are regulated by the presence of an intron in the precursor tRNA (Motorin and Grosjean, 1999; Brzezicha et al., 2006; Auxilien et al., 2012). In addition, some 2′-O-methylated nucleosides in archaeal tRNA are introduced by the aFib, Nop5p and L7Ae complex with the BoxC/D guide RNA system (Clouet-d'Orval et al., 2001, 2005; Bortolin et al., 2003; Singh et al., 2004; Renalier et al., 2005; Ye et al., 2009; Joardar et al., 2011; Lin et al., 2011). In some cases, an intron in the precursor tRNA acts as the guide RNA (Clouet-d'Orval et al., 2001, 2005; Bortolin et al., 2003; Singh et al., 2004). This system is useful in minimizing the size of the genome. In the future, it is possible that considerable numbers of 2′-O-methylated modifications in archaeal tRNA might be identified as products of this system.
Regulation of the Degradation and Localization of tRNA by Methylated Nucleosides
As shown in Table 1, modifications of the anticodon loop (positions 32–38) are involved directly in protein synthesis whereas other modifications affect the structure of the tRNA. Consequently, for a long time, it was thought that modifications outside the anticodon loop acted to stabilize tRNA structure and regulate the half-life of tRNAs. Indeed, we observed in the thermophilic eubacterium Thermus thermophiles that hypomodification at multiple sites in tRNA owing to disruption of one of the modification enzymes promotes the degradation of tRNAPhe and tRNALys at high temperatures (Tomikawa et al., 2010).
In the case of eukaryotes, tRNA methylations work coordinately as stabilizing factors and markers of maturation, and the degree of modification changes in response to various stresses. Hypomodified tRNAs are degraded aggressively. For example, in the Saccharomyces cerevisiae trm4 (synthesizes m5C at multiple sites) and trm8 (produces m7G46) double knock-out strain, the half-life of tRNAVal is shortened and the strain shows a growth defect (Alexandrov et al., 2006). Therefore, tRNA modifications stabilize tRNA structure coordinately and systems to degrade hypomodified tRNAs exist in eukaryotic cells (Alexandrov et al., 2006; Chernyakov et al., 2008; Phizicky and Hopper, 2010; D'Silva et al., 2011; Dewe et al., 2012). Furthermore, in S. cerevisiae, the m1A58 modification by the Trm6–Trm61 complex regulates both the degradation of initiator tRNAMet and its transport from the nucleus to the cytoplasm (Anderson et al., 1998, 2000; Kadaba et al., 2004). The m1A58 modification functions a marker of maturation and absence of modification leads to degradation of initiator tRNAMet during transport. Thus, m1A58 is part of the RNA quality control system. Moreover, in the case of S. cerevisiae, splicing is performed in the cytoplasm (Takano et al., 2005) and precursor tRNAs are matured during repeated-transports between the nucleus and cytoplasm (Ohira and Suzuki, 2011). Therefore, some tRNA modifications might act as the markers of maturation at halfway checkpoints. In Leishmania tarentolae, a proportion of tRNAGlu and tRNAGln is transported from the cytoplasm to the mitochondria (Kaneko et al., 2003). In the cytoplasmic tRNA, U34 is modified to mcm5s2U34, whereas in the mitochondrial tRNA it is modified to mcm5Um34. These results suggest that the s2U34 modification may suppress transport from the cytoplasm to mitochondria. Given that both the s2U and Um modifications shift the equilibrium of ribose puckering to the C3′-endo form (Kawai et al., 1992), these modifications have a nearly equivalent stabilizing effect on the codon-anticodon interaction. The 5-methylcarboxymethyl (mcm) group restricts wobble base pairing (Takai and Yokoyama, 2003). Taken together, these findings suggest that a substantial number of methylated nucleosides contribute to RNA quality control systems and/or the regulation of tRNA localization, even though they were considered previously to have simply a structural role.
Adaptation of Protein Synthesis to Environmental Change through a Network between Modified Nucleosides and tRNA Modification Enzymes
tRNA Modifications in T. thermophilus
Thermus thermophilus provides an example of a living organism that utilizes changes in the structural rigidity (flexibility) of tRNA through multiple nucleoside modifications to adapt protein synthesis to environmental changes. Thermus thermophilus is an extreme thermophilic eubacterium found in hot springs and can grow at a wide range of temperatures (50~83°C). Under natural conditions, the temperature of hot springs can be changed dramatically by several factors, for instance the overflow of hot spring water, snow falling, and the influx of river water. Thermus thermophilus can synthesize proteins in response to these temperature changes. Three distinct modifications (Gm18, m5s2U54, and m1A58) are found in T. thermophilus tRNA and the combination of these modifications increases the melting temperature of tRNA by near 10°C as compared with that of the unmodified transcript (Watanabe et al., 1976; Horie et al., 1985; Shigi et al., 2006; Tomikawa et al., 2010). Although these modifications are very important as structural factors in tRNA, they do not have an effect on translational fidelity below 65°C and the level of modification is very low in tRNA from cells cultured at 50°C (Figure 5A). This change in the extent of modification reflects the adaptation of protein synthesis to temperature change (Yokoyama et al., 1987). Transfer RNAPhe from cells cultured at 80°C efficiently synthesizes poly(U) at high temperatures (above 65°C). In contrast, tRNAPhe from cells cultured at 50°C, in which the levels of the three modifications are low, works efficiently at low temperatures (Figure 5B). Thus, the levels of three modified nucleosides, Gm18, m5s2U54, and m1A58, in tRNA control the elongation of translation via the flexibility of the tRNA. These findings were reported in 1987 (Yokoyama et al., 1987). However, at the beginning of the twenty-first century, the mechanisms of regulation of these modifications remained unknown.
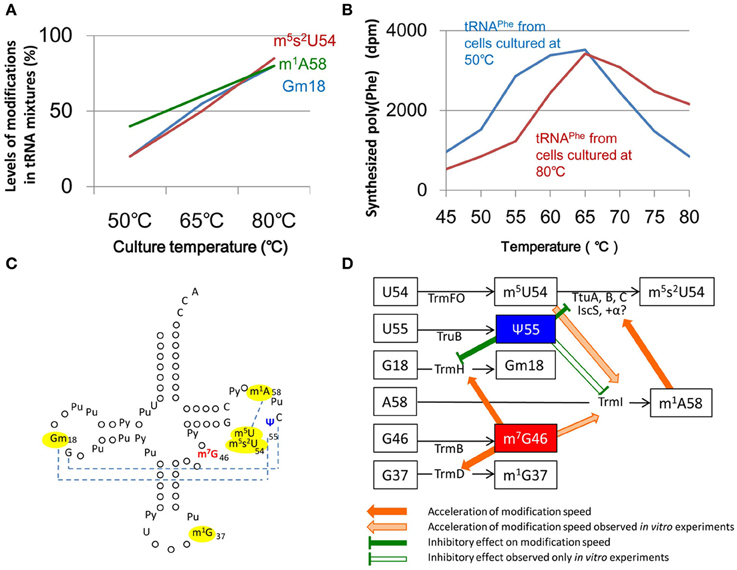
Figure 5. Network between modified nucleotides and tRNA modification enzymes observed in T. thermophilus. (A) The proportion of Gm18, m5s2U54, and m1A58 in tRNA (contents in tRNA fraction) increases with increasing culture temperature. (B) Transfer RNAPhe from cells cultured at 80°C can efficiently synthesize Poly(U) at high temperatures. In contrast, at low temperatures, tRNAPhe from cells cultured at 50°C can work more efficiently than tRNA from cells cultured at 80°C. (C) Modifications of tRNA in T. thermophilus are depicted on the clover-leaf structure. Dotted lines show the tertiary base pairs. The levels of the m7G46 and ψ55 modifications are nearly 100% at a wide range of temperatures. The levels of modifications marked by yellow are regulated by the m7G46 and ψ55 modifications. (D) At temperatures greater than 65°C, the presence of m7G46 increases the rates of modification of Gm18 by TrmH, m1A58 by TrmI and m1G37 by TrmD. The acceleration of m1A58 formation by TrmI in the presence of m7G46 and m5U54 has been confirmed only by in vitro experiments. The m1A58 modification accelerates the thio-transfer reaction by the sulfur atom exchange complex that is required for the formation of m5s2U54. Therefore, at high temperatures, m7G46, m5U54, and m1A58 coordinately promote the formation of m5s2U54 and increases tRNA stability. In contrast, at low temperatures below 65°C, the ψ55 modification increases rigidity within the local structure of the tRNA as described in the main text. This network provides a mechanism by which extreme thermophilic eubacteria adapt to temperature changes. The network regulates the order of modifications in tRNA. This figure summarizes the experimental data in these publications Yokoyama et al. (1987), Shigi et al. (2006), Tomikawa et al. (2010), Ishida et al. (2011), Yamagami et al. (2012).
Switching of Network between Modified Nucleosides and tRNA Modification Enzymes
Initially, we assumed that transcriptional and/or translational regulation of the tRNA modification enzymes was involved in the regulation of the three modifications. However, unexpectedly, we have observed that the phenomenon can be simply explained by the RNA recognition mechanisms of the tRNA modification enzymes (Shigi et al., 2002; Tomikawa et al., 2010; Ishida et al., 2011; Yamagami et al., 2012). Several common modifications (for example, m7G46 and ψ55) are found in T. thermophilus tRNA in addition to Gm18, m5s2U54, and m1A58. When the genes for the modification enzymes for m7G46 and ψ55 (trmB and truB, respectively) were disrupted individually, the levels of Gm18, m5s2U54 and m1A58 in tRNA were changed dramatically (Tomikawa et al., 2010; Ishida et al., 2011). Thus, modified nucleosides and tRNA modification enzymes form a network, and this network regulates the extent of modifications on the basis of temperature (Figures 5C,D).
At high temperatures (above 65°C), m7G46 functions as a marker of precursor tRNA and increases the reaction rates of other modification enzymes. In contrast, at low temperatures, ψ55 confers local structural rigidity and slows down the rate of formation of other modifications around ψ55 (that is, Gm18, m5s2U54, and m1A58). This inhibitory effect weakens as the temperature increases and is not observed above 65°C. Thus, the m7G46 and ψ55 modifications work as an accelerator and a brake in the network, respectively. The advantage of this mechanism is that the network does not include any transcriptional or translational regulatory steps: protein synthesis is not necessary. Thus, the response of the network to environmental changes is very rapid. This is a typical strategy in eubacteria, where genome size is limited.
Similar networks between modified nucleosides and tRNA modification enzymes have also been reported in mesophiles. For example, ms2i6A37 modification in E. coli tRNA is required for 2′-O-methylation by TrmL (Benítez-Páez et al., 2010), and the Cm32 and Gm34 modifications in S. cerevisiae tRNAPhe are required for the formation of yW37 from m1G37 (Guy et al., 2012). However, the network in T. thermophilus is distinct because the modifications are almost all in the three-dimensional core of the tRNA and the network responds to environmental changes.
Genetic Disease and tRNA Methylation
Modifications of tRNA regulate protein synthesis. Consequently, if a disruption of tRNA modification is not lethal, it can directly cause a genetic disease. In fact, there are several reports concerning the relationship between genetic disease and tRNA modification (Yasukawa et al., 2001; Suzuki et al., 2002, 2011b; Freude et al., 2004; Kirino et al., 2005; Umeda et al., 2005; Wei et al., 2011; Towns and Begley, 2012; Igoillo-Esteve et al., 2013). In particular, the number of reports of a link between diabetes and tRNA modification are increasing, which suggests that an increase in the frequency of translation errors has an effect on energy metabolism. The severe disruption of energy metabolism often damages muscle and neuronal cells, which consume large amounts of energy. This perspective enables mitochondrial diseases that are caused by a problem with mitochondrial tRNA modification to be understood (Yasukawa et al., 2001; Suzuki et al., 2002, 2011b; Kirino et al., 2005; Umeda et al., 2005). Furthermore, several tRNA methyltransferases are fused to DNA repair enzymes, which means that these enzymes are related directly to DNA repair and carcinogenesis (Choudhury et al., 2007a,b; Shimada et al., 2009). Moreover, abnormal tRNA modifications have been also reported in cancers (Kuchino and Borek, 1978; Kuchino et al., 1981; Shindo-Okada et al., 1981). These might be caused by the rearrangement of chromosomes in cancer cells.
Infection, Immunity, and tRNA Methylations—tRNA Therapy
Among the tRNA modification enzymes, tRNA guanine transglycosidase (Tgt), which is required for the production of Q34, and tRNA ψ55 synthase (TruB), which generates ψ55, are essential factors for infection by Shigella flexneri (Durand et al., 1994) and Pseudomonas aeruginosa (Saga et al., 1997), respectively. Similarly, we also found that tRNA (m7G46) methyltransferase is essential for infection by Colletotrichum lagenarium, an infectious fungus (Takano et al., 2006). Furthermore, tRNAs that contain mcm5U modifications are the target of Kluyveromyces lactis gamma-toxin (Lu et al., 2005) and Pichia acaciae killer toxin (Klassen et al., 2008). Moreover, given that retroviruses utilize host tRNA as the primer for reverse transcription, tRNA methylation and methyltransferases are involved in both reverse transcription and the packaging of virus particles. For example, human immunodeficiency virus (HIV; AIDS virus) utilizes the m1A58 modification in tRNALys3 as the terminator of reverse transcription (see reviews, Marquet, 1998; Saadatmand and Kleiman, 2012; Sleiman et al., 2012). Consequently, the regulation of tRNA modification and modification enzymes might be a powerful tool to control infectious organisms.
When an exogenous single-stranded RNA such as Haemophilus influenzae tRNA is present in humans, Toll-like receptor 7 (TLR7) forms a dimer structure and then activates the immune response systems (Figure 6). However, endogenous or E. coli tRNA does not stimulate TLR7. The mechanism of differentiation was clarified recently by two groups, who found that the Gm18 modification in E. coli tRNA suppresses immunostimulation via TLR7 (Gehrig et al., 2012; Jöckel et al., 2012). Thus, enterobacteria exploit the Gm18 modification in tRNA to avoid the host immune system. Furthermore, given that Gm18-modified tRNA acts as an antagonist of TLR7 (Jöckel et al., 2012), Gm18-modified tRNA might be an effective anti-inflammatory drug.
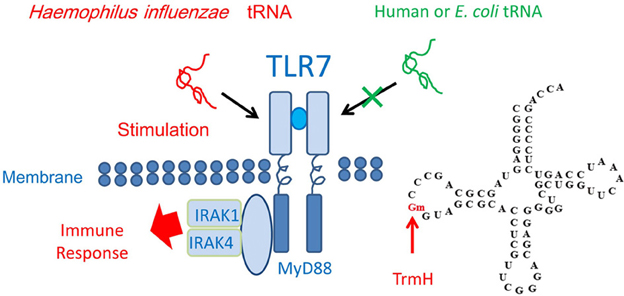
Figure 6. Immune response and tRNA methylation. Transfer RNA from H. influenzae, a respiratory infectious bacterium, induces the dimer formation by Toll like receptor-7 (TLR7) and then the immune response is stimulated via binding of the proteins, MyD88, IRAK1, and IRAK4. However, human and E. coli tRNAs do not stimulate TLR7 because these tRNA contains the Gm18 modification. The E. coli trmH gene disruptant strain does not show any obvious phenotype under laboratory culture conditions (Persson et al., 1997). The Gm18 modification and TrmH are required for survival of E. coli in the animal gut.
Evolution of Modifications in tRNA
Finally, it is worthwhile discussing the evolution of modifications in tRNA. During the early period of chemical evolution (see reviews Cermakian and Cedergren, 1998; Joyce and Orgel, 2006), inosine could be used as a basic component of RNA, because it can be synthesized from adenosine non-enzymatically. Inosine seems to have been excluded after the appearance of genes because it changes the genetic information during the replication process. Simple methylated nucleosides such as m1G became essential when the reading frame of protein synthesis was separated into three-nucleotide units (Björk et al., 1989, 2001). Thus, several methylated nucleosides seem to have appeared during the chemical evolution period (Cermakian and Cedergren, 1998). After the appearance of the reading frame, the importance of the availability of methyl groups increased and it seems that the methionine codon was selected as the translation initiation codon.
It appears that complicated enzymes were not formed during the period of chemical evolution (Joyce and Orgel, 2006). The early enzymes might have been oligopeptides and might have included metals as the catalytic center, as is the case for deaminases (Carter, 1998; Schaub and Keller, 2002). It is possible that the codons were not fixed strictly as is observed in the universal code (Jukes, 1973; Cedergren et al., 1986; Osawa et al., 1992). However, it is likely that the most basic catalytic core of tRNA methyltransferases was established when cell-like organisms began to exchange their components and genes because the basic structure of tRNA methyltransferases is shared by all living organisms found today (Figure 4 and Table 2). The structures of methyltransferases (Schubert et al., 2003) suggest that RNA methyltransferases, which were required for protein synthesis, evolved to yield DNA and protein methyltransferases many times during the evolution of life. The mechanisms to generate the complicated modified nucleotides that regulate the wobble base pair seem to have arisen after the origination of living organisms because they show considerable diversity and involve multistep reactions (Table 1).
The temperature of primordial Earth was higher than that of the Earth at present. Consequently, several nucleoside modifications in tRNA and rRNA would be necessary to stabilize the structure of the RNA (Motorin and Helm, 2010). However, it is likely that the network between modified nucleosides and tRNA modification enzymes that is observed in extreme thermophiles (Figure 5D and section Adaptation of Protein Synthesis to Environmental Change Through a Network Between Modified Nucleosides and tRNA Modification Enzymes) was established after the cooling of the Earth because it responds to low temperatures (Ishida et al., 2011). Obviously, the functions of modified nucleotides with respect to the RNA quality control system and regulation of cellular localization were acquired after the appearance of eukaryotes (see section Regulation of the Degradation and Localization of tRNA by Methylated Nucleosides).
Transfer RNA modifications are still evolving. The most powerful driving force is the existence of infectious organisms (see section Infection, Immunity, and tRNA Methylations—tRNA Therapy). Hosts need to distinguish endogenous RNA from exogenous RNA to prevent infection and infectious organisms need to avoid the host defense system to survive. Consequently, tRNA modifications and modification enzymes are still subject to evolution even today.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Ahn, H. J., Kim, H. W., Yoon, H. J., Lee, B., Suh, S. S., and Yang, J. K. (2003). Crystal structure of tRNA(m1G37)methyltransferase: insights into tRNA recognition. EMBO J. 22, 2593–2603. doi: 10.1093/emboj/cdg269
Alexandrov, A., Chernyakov, I., Gu, W., Hiley, S. L., Hughes, T. R., Grayhack, E. J., et al. (2006). Rapid tRNA decay can result from lack of nonessential modifications. Mol. Cell 21, 87–96. doi: 10.1016/j.molcel.2005.10.036
Alexandrov, A., Grayhack, E. J., and Phizicky, E. M. (2005). tRNA m7G methyltransferase Trm8p/Trm82p: evidence linking activity to a growth phenotype and implicating Trm82p in maintaining levels of active Trm8p. RNA 11, 821–830. doi: 10.1261/rna.2030705
Alexandrov, A., Martzen, M. R., and Phizicky, E. M. (2002). Two proteins that form a complex are required for 7-methylguanosine modification of yeast tRNA. RNA 8, 1253–1266. doi: 10.1017/S1355838202024019
Alian, A., Lee, T. T., Griner, S. L., Stroud, R. M., and Finer-Moore, J. (2008). Structure of a TrmA-RNA complex: a consensus RNA fold contributes to substrate selectivity and catalysis in m5U methyltransferases. Proc. Natl. Acad. Sci. U.S.A. 105, 6876–6881. doi: 10.1073/pnas.0802247105
Anantharaman, V., Koonin, E. V., and Aravind, L. (2002). SPOUT: a class of methyltransferases that includes spoU and trmD RNA methylase superfamilies, and novel superfamilies of predicted prokaryotic RNA methylases. J. Mol. Microbiol. Biotechnol. 4, 71–75.
Anderson, J., Phan, L., Cuesta, R., Carison, B. A., Pak, M., Asano, K., et al. (1998). The essential Gcd10p-Gcd14p nuclear complex is required for 1-methyladenosine modification and maturation of initiator methionyl-tRNA. Genes Dev. 12, 3650–3652. doi: 10.1101/gad.12.23.3650
Anderson, J., Phan, L., and Hinnebusch, A. G. (2000). The Gcd10p/Gcd14p complex is the essential two-subunit tRNA(1-methyladenosine) methyltransferase of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. U.S.A. 97, 5173–5178. doi: 10.1073/pnas.090102597
Armengaud, J., Urbonavicius, J., Fernandez, B., Chaussinand, G., Bujnicki, J. M., and Grosjean, H. (2004). N2-methylation of guanosine at position 10 in tRNA is catalyzed by a THUMP domain-containing, S-adenosylmethionine-dependent methyltransferase, conserved in Archaea and Eukaryota. J. Biol. Chem. 279, 37142–37152. doi: 10.1074/jbc.M403845200
Armengod, M. E., Moukadiri, I., Prado, S., Ruiz-Partida, R., Benítez-Páez, A., Villarroya, M., et al. (2012). Enzymology of tRNA modification in the bacterial MnmEG pathway. Biochimie 94, 1510–1520. doi: 10.1016/j.biochi.2012.02.019
Arragain, S., Handelman, S. K., Forouhar, F., Wei, F. Y., Tomizawa, K., Hunt, J. F., et al. (2010). Identification of eukaryotic and prokaryotic methylthiotransferase for biosynthesis of 2-methylthio-N6-threonylcarbamoyladenosine in tRNA. J. Biol. Chem. 285, 28425–28433. doi: 10.1074/jbc.M110.106831
Atta, M., Arragain, S., Fontecave, M., Mulliez, E., Hunt, J. F., Luff, J. D., et al. (2012). The methylthiolation reaction mediated by the Radical-SAM enzymes. Biochim. Biophys. Acta 1824, 1223–1230. doi: 10.1016/j.bbapap.2011.11.007
Atta, M., Mulliez, E., Arragain, S., Forouhar, F., Hunt, J. F., and Fontecave, M. (2010). S-Adenosylmethionine-dependent radical-based modification of biological macromolecules. Curr. Opin. Struct. Biol. 20, 684–692. doi: 10.1016/j.sbi.2010.09.009
Auxilien, S., El Khadali, F., Rasmussen, A., Douthwaite, S., and Grosjean, H. (2007). Archease from Pyrococcus abyssi improves substrate specificity and solubility of a tRNA m5C methyltransferase. J. Biol. Chem. 282, 18711–18721. doi: 10.1074/jbc.M607459200
Auxilien, S., Guérineau, V., Szweykowska-Kuliñska, Z., and Golinelli-Pimpaneau, B. (2012). The human tRNA m (5) C methyltransferase misu is multisite-specific. RNA Biol. 9, 1331–1338. doi: 10.4161/rna.22180
Auxilien, S., Rasmussen, A., Rose, S., Brochier-Armanet, C., Husson, C., Fourmy, D., et al. (2011). Specificity shifts in the rRNA and tRNA nucleotide targets of archaeal and bacterial m5U methyltransferases. RNA 17, 45–53. doi: 10.1261/rna.2323411
Awai, T., Kimura, S., Tomikawa, C., Ochi, A., Ihsanawati, Bessho, Y., et al. (2009). Aquifex aeolicus tRNA (N2,N2-guanine)-dimethyltransferase (Trm1) catalyzes transfer of methyl groups not only to guanine 26 but also to guanine 27 in tRNA. J. Biol. Chem. 284, 20467–20478. doi: 10.1074/jbc.M109.020024
Awai, T., Ochi, A., Ihsanawati, Sengoku, T., Hirata, A., Bessho, Y., et al. (2011). Substrate tRNA recognition mechanism of a multisite-specific tRNA methyltransferase, Aquifex aeolicus Trm1, based on the X-ray crystal structure. J. Biol. Chem. 286, 35236–35246. doi: 10.1074/jbc.M111.253641
Barraud, P., Golinelli-Pimpaneau, B., Atmanene, C., Sanglier, S., Van Dorsselaer, A., Droogmans, L., et al. (2008). Crystal structure of Thermus thermophilus tRNA m1A58 methyltransferase and biophysical characterization of its interaction with tRNA. J. Mol. Biol. 377, 535–550. doi: 10.1016/j.jmb.2008.01.041
Begley, U., Dyavaiah, M., Patil, A., Rooney, J. P., DiRenzo, D., Young, C. M., et al. (2007). Trm9-catalyzed tRNA modifications link translation to the DNA damage response. Mol. Cell 28, 860–870. doi: 10.1016/j.molcel.2007.09.021
Benítez-Páez, A., Villarroya, M., Douthwaite, S., Gabaldón, T., and Armengod, M. E. (2010). YibK is the 2'-O-methyltransferase TrmL that modifies the wobble nucleotide in Escherichia coli tRNA(Leu) isoacceptors. RNA 16, 2131–2143. doi: 10.1261/rna.2245910
Björk, G. R., Jacobsson, K., Nilsson, K., Johansson, M. J., Byström, A. S., and Persson, O. P. (2001). A primordial tRNA modification required for the evolution of life? EMBO J. 20, 231–239. doi: 10.1093/emboj/20.1.231
Björk, G. R., Wikstrom, P. M., and Byström, A. S. (1989). Prevention of translational frameshifting by the modified nucleoside 1-methylguanosine. Science 244, 986–989. doi: 10.1126/science.2471265
Böhme, S., Meyer, S., Krüger, A., Steinhoff, H. J., Wittinghofer, A., and Klare, J. P. (2010). Stabilization of G domain conformations in the tRNA-modifying MnmE-GidA complex observed with double electron electron resonance spectroscopy. J. Biol. Chem. 285, 16991–17000. doi: 10.1074/jbc.M109.096131
Bortolin, M. L., Bachellerie, J. P., and Clouet-d'Orval, B. (2003). In vitro RNP assembly and methylation guide activity of an unusual box C/D RNA, cis-acting archaeal pre-tRNA(Trp). Nucleic Acids Res. 31, 6524–6535. doi: 10.1093/nar/gkg860
Brulé, H., Elliott, M., Redlak, M., Zehner, Z. E., and Holmes, W. M. (2004). Isolation and characterization of the human tRNA-(N1G37) methyltransferase (TRM5) and comparison to the Escherichia coli TrmD protein. Biochemistry 43, 9243–9255. doi: 10.1021/bi049671q
Brzezicha, B., Schmidt, M., Makalowska, I., Jarmolowski, A., Pienkowska, J., and Szweykowska-Kulinska, Z. (2006). Identification of human tRNA:m5C methyltransferase catalysing intron-dependent m5C formation in the first position of the anticodon of the pre-tRNA Leu (CAA). Nucleic Acids Res. 34, 6034–6043. doi: 10.1093/nar/gkl765
Bujnicki, J. M., Oudjama, Y., Roovers, M., Owczarek, S., Caillet, J., and Droogmans, L. (2004). Identification of a bifunctional enzyme MnmC involved in the biosynthesis of a hypermodified uridine in the wobble position of tRNA. RNA 18, 1236–1242. doi: 10.1261/rna.7470904
Byström, A. S., and Björk, G. R. (1982). Chromosomal location and cloning of the gene (trmD) responsible for the synthesis of tRNA (m1G) methyltransferase in Escherichia coli K-12. Mol. Gen. Genet. 188, 440–446. doi: 10.1007/BF00330046
Carter, C. W. Jr. (1998). “Chapter 20 nucleoside deaminases for cytidine and adenosine: comparison with deaminases acting on RNA,” in Modification and Editing of RNA, edS H. Grosjean and R. Benne(Washington, DC: ASM press), 363–375.
Cartlidge, R. A., Knebel, A., Peggie, M., Alexandrov, A., Phizicky, E. M., and Cohen, P. (2005). The tRNA methylase METTL1 is phosphorylated and inactivated by PKB and RSK in vitro and in cells. EMBO J. 24, 1696–1705. doi: 10.1038/sj.emboj.7600648
Cavaillé, J., Chetouani, F., and Bachellerie, J.-P. (1999). The yeast Saccharomyces cerevisiae YDL112w ORF encodes the putative 2'-O-ribose methyltransferase catalyzing the formation of Gm18 in tRNAs. RNA 5, 66–81. doi: 10.1017/S1355838299981475
Cedergren, R., Grosjean, H., and Larue, B. (1986). Primordial reading of genetic information. Biosystems 19, 259–266. doi: 10.1016/0303-2647(86)90002-X
Cermakian, N., and Cedergren, R. (1998). “Chapter 29 Modified nucleosides always were: an evolutionary model,” in Modification and Editing of RNA, edS H. Grosjean and R. Benne (Washington, DC: ASM press), 535–541.
Chan, C. T., Pang, Y. L., Deng, W., Babu, I. R., Dyavaiah, M., Begley, T. J., et al. (2012). Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins. Nat Commun. 3, 937. doi: 10.1038/ncomms1938
Chatterjee, K., Blaby, I. K., Thiaville, P. C., Majumder, M., Grosjean, H., Yuan, Y. A., et al. (2012). The archaeal COG1901/DUF358 SPOUT-methyltransferase members, together with pseudouridine synthase Pus10, catalyze the formation of 1-methylpseudouridine at position 54 of tRNA. RNA 18, 421–433. doi: 10.1261/rna.030841.111
Chen, C., Huang, B., Anderson, J. T., and Byström, A. S. (2011). Unexpected accumulation of ncm(5)U and ncm(5)S(2) (U) in a trm9 mutant suggests an additional step in the synthesis of mcm(5)U and mcm(5)S(2)U. PLoS ONE 6:e20783. doi: 10.1371/journal.pone.0020783
Chen, H. Y., and Yuan, Y. A. (2010). Crystal structure of Mj1640/DUF358 protein reveals a putative SPOUT-class RNA methyltransferase. J. Mol. Cell. Biol. 2, 366–374. doi: 10.1093/jmcb/mjq034
Chen, P., Crain, P. F., Näsvall, S. J., Pomerantz, S. C., and Björk, G. R. (2005). A “gain of function” mutation in a protein mediates production of novel modified nucleosides. EMBO J. 24, 1842–1851. doi: 10.1038/sj.emboj.7600666
Chernyakov, I., Whipple, J. M., Kotelawala, L., Grayhack, E. J., and Phizicky, E. M. (2008). Degradation of several hypomodified mature tRNA species in Saccharomyces cerevisiae is mediated by Met22 and the 5′-3′ exonucleases Rat1 and Xrn1. Genes Dev. 22, 1369–1380. doi: 10.1101/gad.1654308
Choudhury, S. A., Asefa, B., Kauler, P., and Chow, T. Y. (2007b). Synergistic effect of TRM2/RNC1 and EXO1 in DNA double-strand break repair in Saccharomyces cerevisiae. Mol. Cell. Biochem. 304, 127–134. doi: 10.1007/s11010-007-9493-7
Choudhury, S. A., Asefa, B., Webb, A., Ramotar, D., and Chow, T. Y. (2007a). Functional and genetic analysis of the Saccharomyces cerevisiae RNC1/TRM2: evidences for its involvement in DNA double-strand break repair. Mol. Cell. Biochem. 300, 215–226. doi: 10.1007/s11010-006-9386-1
Christian, T., Evilia, C., Williams, S., and Hou, Y. M. (2004). Distinct origins of tRNA(m1G37) methyltransferase. J. Mol. Biol. 339, 707–719. doi: 10.1016/j.jmb.2004.04.025
Christian, T., Gamper, H., and Hou, Y. M. (2013). Conservation of structure and mechanism by Trm5 enzymes. RNA 19, 1192–1199. doi: 10.1261/rna.039503.113
Christian, T., and Hou, Y. M. (2007). Distinct determinants of tRNA recognition by the TrmD and Trm5 methyl transferases. J. Mol. Biol. 373, 623–632. doi: 10.1016/j.jmb.2007.08.010
Chujo, T., and Suzuki, T. (2012). Trmt61B is a methyltransferase responsible for 1-methyladenosine at position 58 of human mitochondrial tRNAs. RNA 18, 2269–2276. doi: 10.1261/rna.035600.112
Clouet-d'Orval, B., Bortolin, M. L., Gaspin, C., and Bachellerie, J. P. (2001). Box C/D RNA guides for the ribose methylation of archaeal tRNAs. the tRNATrp intron guides the formation of two ribose-methylated nucleosides in the mature tRNATrp. Nucleic Acids Res. 29, 4518–4529. doi: 10.1093/nar/29.22.4518
Clouet-d'Orval, B., Gaspin, C., and Mougin, A. (2005). Two different mechanisms for tRNA ribose methylation in Archaea: a short survey. Biochimie 87, 889–895. doi: 10.1016/j.biochi.2005.02.004
Constantinesco, F., Benachenhou, N., Motorin, Y., and Grosjean, H. (1998). The tRNA(guanine-26,N2-N2) methyltransferase (Trm1) from the hyperthermophilic archaeon Pyrococcus furiosus: cloning, sequencing of the gene and its expression in Escherichia coli. Nucleic Acids Res. 26, 3753–3761. doi: 10.1093/nar/26.16.3753
Constantinesco, F., Motorin, Y., and Grosjean, H. (1999a). Transfer RNA modification enzymes from Pyrococcus furiosus: detection of the enzymatic activities in vitro. Nucleic Acids Res. 27, 1308–1315. doi: 10.1093/nar/27.5.1308
Constantinesco, F., Motorin, Y., and Grosjean, H. (1999b). Characterisation and enzymatic properties of tRNA(guanine 26, N (2), N (2))-dimethyltransferase (Trm1p) from Pyrococcus furiosus. J. Mol. Biol. 291, 375–392.
De Bie, L. G., Roovers, M., Oudjama, Y., Wattiez, R., Tricot, C., Stalon, V., et al. (2003). The yggH gene of Escherichia coli encodes a tRNA (m7G46) methyltransferase. J. Bacteriol. 185, 3238–3243. doi: 10.1128/JB.185.10.3238-3243.2003
de Crécy-Lagard, V., Brochier-Armanet, C., Urbonavicius, J., Fernandez, B., Phillips, G., Lyons, B., et al. (2010). Biosynthesis of wyosine derivatives in tRNA: an ancient and highly diverse pathway in Archaea. Mol. Biol. Evol. 27, 2062–2077. doi: 10.1093/molbev/msq096
Delk, A. S., Romeo, J. M., Nagle, D. P. Jr., and Rabinowitz, J. C. (1976). Biosynthesis of ribothymidine in the transfer RNA of Streptococcus faecalis and Bacillus subtilis. a methylation of RNA involving 5,10-methylenetetrahydrofolate. J. Biol. Chem. 251, 7649–7656.
Dewe, J. M., Whipple, J. M., Chernyakov, I., Jaramillo, L. N., and Phizicky, E. M. (2012). The yeast rapid tRNA decay pathway competes with elongation factor 1A for substrate tRNAs and acts on tRNAs lacking one or more of several modifications. RNA 18, 1886–1896. doi: 10.1261/rna.033654.112
Dong, A., Yoder, J. A., Zhang, X., Zhou, L., Bestor, T. H., and Cheng, X. (2001). Structure of human DNMT2, an enigmatic DNA methyltransferase homolog that displays denaturant-resistant binding to DNA. Nucleic Acids Res. 29, 439–448. doi: 10.1093/nar/29.2.439
Droogmans, L., Roovers, M., Bujnicki, J. M., Tricot, C., Hartsch, T., Stalon, V., et al. (2003). Cloning and characterization of tRNA (m1A58) methyltransferase (TrmI) from Thermus thermophilus HB27, a protein required for cell growth at extreme temperatures. Nucleic Acids Res. 31, 2148–2156. doi: 10.1093/nar/gkg314
D'Silva, S., Haider, S. J., and Phizicky, E. M. (2011). A domain of the actin binding protein Abp140 is the yeast methyltransferase responsible for 3-methylcytidine modification in the tRNA anti-codon loop. RNA 17, 1100–1110. doi: 10.1261/rna.2652611
Durand, J. M., Björk, G. R., Kuwae, A., Yoshikawa, M., and Sasakawa, C. (1997). The modified nucleoside 2-methylthio-N6-isopentenyladenosine in tRNA of Shigella flexneri is required for expression of virulence genes. J. Bacteriol. 179, 5777–5782.
Durand, J. M., Okada, N., Tobe, T., Watarai, M., Fukuda, I., Suzuki, T., et al. (1994). vacC, a virulence-associated chromosomal locus of Shigella flexneri, is homologous to tgt, a gene encoding tRNA-guanine transglycosylase (Tgt) of Escherichia coli K-12. J. Bacteriol. 176, 4627–4634.
Durant, P. C., Bajji, A. C., Sundaram, M., Kumar, R. K., and Davis, D. R. (2005). Structural effects of hypermodified nucleosides in the Escherichia coli and human tRNALys anticodon loop: the effect of nucleosides s2U, mcm5U, mcm5s2U, mnm5s2U, t6A, and ms2t6A. Biochemistry 44, 8078–8089. doi: 10.1021/bi050343f
Edelheit, S., Schwartz, S., Mumbach, M. R., Wurtzel, O., and Sorek, R. (2013). Transcriptome-wide mapping of 5-methylcytidine RNA modifications in bacteria, archaea, and yeast reveals m5C within archaeal mRNAs. PLoS Genet. 9:e1003602. doi: 10.1371/journal.pgen.1003602
Edmonds, C. G., Crain, P. F., Gupta, R., Hashizume, T., Hocart, C. H., Kowalak, J. A., et al. (1991). Posttranscriptional modification of tRNA in thermophilic archaea (Archaebacteria). J. Bacteriol. 173, 3138–3148.
Edqvist, J., Blomqvist, K., and Stråby, K. B. (1994). Structural elements in yeast tRNAs required for homologous modification of guanosine-26 into dimethylguanosine-26 by the yeast Trm1 tRNA-modifying enzyme. Biochemistry 33, 9546–9551. doi: 10.1021/bi00198a021
Elkins, P. A., Watts, J. M., Zalacain, M., van Thiel, A., Vitazka, P. R., Redlak, M., et al. (2003). Insights into catalysis by a knotted TrmD tRNA methyltransferase. J. Mol. Biol. 333, 931–949. doi: 10.1016/j.jmb.2003.09.011
Ellis, S. R., Morales, M. J., Li, J. M., Hopper, A. K., and Martin, N. C. (1986). Isolation and characterization of the TRM1 locus, a gene essential for the N2,N2-dimethylguanosine modification of both mitochondrial and cytoplasmic tRNA in Saccharomyces cerevisiae. J. Biol. Chem. 261, 9703–9709.
Farabaugh, P. J., and Björk, G. R. (1999). How translational accuracy influences reading frame maintenance. EMBO J. 18, 1427–1434. doi: 10.1093/emboj/18.6.1427
Fislage, M., Roovers, M., Tuszynska, I., Bujnicki, J. M., Droogmans, L., and Versées, W. (2012). Crystal structures of the tRNA:m2G6 methyltransferase Trm14/TrmN from two domains of life. Nucleic Acids Res. 40, 5149–5161. doi: 10.1093/nar/gks163
Freude, K., Hoffmann, K., Jensen, L. R., Delatycki, M. B., des Portes, V., Moser, B., et al. (2004). Mutations in the FTSJ1 gene coding for a novel S-adenosylmethionine-binding protein cause nonsyndromic X-linked mental retardation. Am. J. Hum. Genet. 75, 305–309. doi: 10.1086/422507
Fu, D., Brophy, J. A., Chan, C. T., Atmore, K. A., Begley, U., Paules, R. S., et al. (2010). Human AlkB homolog ABH8 is a tRNA methyltransferase required for wobble uridine modification and DNA damage survival. Mol. Cell Biol. 30, 2449–2459. doi: 10.1128/MCB.01604-09
Fujimori, D. G. (2013). Radical SAM-mediated methylation reactions. Curr. Opin. Chem. Biol. 17, 597–604. doi: 10.1016/j.cbpa.2013.05.032
Gehrig, S., Eberle, M.-E., Botschen, F., Rimbach, K., Eberle, F., Eigenbrod, T., et al. (2012). Identification of modifications in microbial, native tRNA that suppress immunostimulatory activity. J. Exp. Med. 209, 225–233. doi: 10.1084/jem.20111044
Goll, M. G., Kirpekar, F., Maggert, K. A., Yoder, J. A., Hsieh, C. L., Zhang, X., et al. (2006). Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398. doi: 10.1126/science.1120976
Goto-Ito, S., Ito, T., Ishii, R., Muto, Y., Bessho, Y., and Yokoyama, S. (2008). Crystal structure of archaeal tRNA(m(1)G37)methyltransferase aTrm5. Proteins 72, 1274–1289. doi: 10.1002/prot.22019
Goto-Ito, S., Ito, T., Kuratani, M., Bessho, Y., and Yokoyama, S. (2009). Tertiary structure checkpoint at anticodon loop modification in tRNA functional maturation. Nat. Struct. Mol. Biol. 16, 1109–1115. doi: 10.1038/nsmb.1653
Greenberg, R., and Dudock, B. (1980). Isolation and characterization of m5U-methyltransferase from Escherichia coli. J. Biol. Chem. 255, 8296–8302.
Grosjean, H., Auxilien, S., Constantinesco, F., Simon, C., Corda, Y., Becker, H. F., et al. (1996). Enzymatic conversion of adenosine to inosine and to N1-methylinosine in transfer RNAs: a review Biochimie 78, 488–501. doi: 10.1016/0300-9084(96)84755-9
Grosjean, H., Constantinesco, F., Foiret, D., and Benachenhou, N. (1995). A novel enzymatic pathway leading to 1-methylinosine modification in Haloferax volcanii tRNA. Nucleic Acids Res. 23, 4312–4319. doi: 10.1093/nar/23.21.4312
Grosjean, H., Gaspin, C., Marck, C., Decatur, W. A., and de Crecy-Lagard, V. (2008). RNomics and Modomics in the halophilic archaea Haloferax volcanii: identification of RNA modification genes. BMC Genomics 9:470. doi: 10.1186/1471-2164-9-470
Gu, X., and Santi, D. V. (1991). The T-arm of tRNA is a substrate for tRNA (m5U54)-methyltransferase. Biochemistry 30, 2999–3002. doi: 10.1021/bi00226a003
Guelorget, A., Barraud, P., Tisné, C., and Golinelli-Pimpaneau, B. (2011). Structural comparison of tRNA m(1)A58 methyltransferases revealed different molecular strategies to maintain their oligomeric architecture under extreme conditions. BMC Struct. Biol. 11:48. doi: 10.1186/1472-6807-11-48
Guelorget, A., Roovers, M., Guérineau, V., Barbey, C., Li, X., and Golinelli-Pimpaneau, B. (2010). Insights into the hyperthermostability and unusual region-specificity of archaeal Pyrococcus abyssi tRNA m1A57/58 methyltransferase. Nucleic Acids Res. 38, 6206–6218. doi: 10.1093/nar/gkq381
Gupta, R. (1984). Halobacterium volcanii tRNAs. Identification of 41 tRNAs covering all amino acids, and the sequences of 33 class I tRNAs. J. Biol. Chem. 259, 9461–9471.
Gustafsson, C., and Björk, G. R. (1993). The tRNA-(m5U54)-methyltransferase of Escherichia coli is present in two forms in vivo, one of which is present as bound to tRNA and to a 3'-end fragment of 16 s rRNA. J. Biol. Chem. 268, 1326–1331.
Gustafsson, C., Reid, R., Greene, P. J., and Santi, D. V. (1996). Identification of new RNA modifying enzymes by iterative genome search using known modifying enzymes as probes. Nucleic Acids Res. 24, 3756–3762. doi: 10.1093/nar/24.19.3756
Guy, M. P., Podyma, B. M., Preston, M. A., Shaheen, H. H., Krivos, K. L., Limbach, P. A., et al. (2012). Yeast Trm7 interacts with distinct proteins for critical modifications of the tRNAPhe anticodon loop. RNA 18, 1921–1933. doi: 10.1261/rna.035287.112
Hagervall, T. G., Tuohy, T. M. F., Atkins, J. F., and Björk, G. R. (1993). Deficiency of 1-methylguanosine in tRNA from Salmonella typhimurium induces frameshifting by quadruplet translocation. J. Mol. Biol. 232, 756–765. doi: 10.1006/jmbi.1993.1429
Hamdane, D., Argentini, M., Cornu, D., Golinelli-Pimpaneau, B., and Fontecave, M. (2012). FAD/folate-dependent tRNA methyltransferase: flavin as a new methyl-transfer agent. J. Am. Chem. Soc. 134, 19739–19745. doi: 10.1021/ja308145p
Hamdane, D., Argentini, M., Cornu, D., Myllykallio, H., Skouloubris, S., Hui-Bon-Hoa, G., et al. (2011b). Insights into folate/FAD-dependent tRNA methyltransferase mechanism: role of two highly conserved cysteines in catalysis. J. Biol. Chem. 286, 36268–36280. doi: 10.1074/jbc.M111.256966
Hamdane, D., Bruch, E., Un, S., Field, M., and Fontecave M. (2013). Activation of a unique flavin-dependent tRNA-Methylating agent. Biochemistry 52, 8949–8956. doi: 10.1021/bi4013879
Hamdane, D., Guerineau, V., Un, S., and Golinelli-Pimpaneau, B. (2011a). A catalytic intermediate and several flavin redox states stabilized by folate-dependent tRNA methyltransferase from Bacillus subtilis. Biochemistry 50, 5208–5219. doi: 10.1021/bi1019463
Helm, M., Brule, H., Deqoul, F., Cepanec, C., Leroux, J. P., Giege, R., et al. (1998). The presence of modified nucleotides is required for cloverleaf folding of a human mitochondrial tRNA. Nucleic Acids Res. 26, 1636–1643. doi: 10.1093/nar/26.7.1636
Herbig, K., Chiang, E. P., Lee, L. R., Hills, J., Shane, B., and Stove, P. J. (2002). Cytoplasmic serine hydroxymethyltransferase mediates competition between folate-dependent deoxyribonucleotide and S-adenosylmethionine biosyntheses. J. Biol. Chem. 277, 38381–38389. doi: 10.1074/jbc.M205000200
Holley, R. W., Apgar, J., Everett, G. A., Madison, J. T., Marquisee, M., Merrill, S. H., et al. (1965). Structure of a ribonucleic acid. Science 147, 1462–1465. doi: 10.1126/science.147.3664.1462
Hopper, A. K., Furukawa, A. H., Pham, H. D., and Martin, N. C. (1982). Defects in modification of cytoplasmic and mitochondrial transfer RNAs are caused by single nuclear mutations. Cell 28, 543–550. doi: 10.1016/0092-8674(82)90209-4
Hori, H., Kubota, S., Watanabe, K., Kim, J. M., Ogasawara, T., Sawasaki, T., et al. (2003). Aquifex aeolicus tRNA (Gm18) methyltransferase has unique substrate specificity: tRNA recognition mechanism of the enzyme. J. Biol. Chem. 278, 25081–25090. doi: 10.1074/jbc.M212577200
Hori, H., Suzuki, T., Sugawara, K., Inoue, Y., Shibata, T., Kuramitsu, S., et al. (2002). Identification and characterization of tRNA (Gm18) methyltransferase from Thermus thermophilus HB8: domain structure and conserved amino acid sequence motifs. Genes Cells 7, 259–272. doi: 10.1046/j.1365-2443.2002.00520.x
Hori, H., Yamazaki, N., Matsumoto, T., Watanabe, Y., Ueda, T., Nishikawa, K., et al. (1998). Substrate recognition of tRNA (Guanosine-2'-)-methyltransferase from Thermus thermophilus HB27. J. Biol. Chem. 273, 25721–25727. doi: 10.1074/jbc.273.40.25721
Horie, N., Hara-Yokoyama, M., Yokoyama, S., Watanabe, K., Kuchino, Y., Nishimura, S., et al. (1985). Two tRNAIle1 species from an extreme thermophile, Thermus thermophilus HB8: effect of 2-thiolation of ribothymidine on the thermostability of tRNA. Biochemistry 24, 5711–5715. doi: 10.1021/bi00342a004
Hou, Y. M., and Perona, J. J. (2010). Stereochemical mechanisms of tRNA methyltransferases. FEBS Lett. 584, 278–286. doi: 10.1016/j.febslet.2009.11.075
Huang, B., Johansson, M. J., and Byström, A. S. (2005). An early step in wobble uridine tRNA modification requires the elongator complex. RNA 11, 424–436. doi: 10.1261/rna.7247705
Hurwitz, J., Gold, M., and Anders, M. (1964). The enzymatic methylation of ribonucleic acid and deoxyribonucleic acid. IV. The properties of the soluble ribonucleic acid-methylating enzymes. J. Biol. Chem. 239, 3474–3482.
Igoillo-Esteve, M., Genin, A., Lambert, N., Desir, J., Pirson, I., Abdulkarim, B., et al. (2013). tRNA methyltransferase homolog gene TRMT10A mutation in young onset diabetes and primary microcephaly in humans PLoS Genet. 9:e1003888. doi: 10.1371/journal.pgen.1003888
Ihsanawati, Nishimoto, M., Higashijima, K., Shirouzu, M., Grosjean, H., Bessho, Y., et al. (2008). Crystal structure of tRNA N2,N2-guanosine dimethyltransferase Trm1 from Pyrococcus horikoshii. J. Mol. Biol. 383, 871–884. doi: 10.1016/j.jmb.2008.08.068
Ikeuchi, Y., Kimura, S., Numata, T., Nakamura, D., Yokogawa, T., Ogata, T., et al. (2010). Agmatine-conjugated cytidine in a tRNA anticodon is essential for AUA decoding in archaea. Nat. Chem. Biol. 6, 277–282. doi: 10.1038/nchembio.323
Ikeuchi, Y., Kitahara, K., and Suzuki, T. (2008). The RNA acetyltransferase driven by ATP hydrolysis synthesizes N4-acetylcytidine of tRNA anticodon. EMBO J. 27, 2194–2203. doi: 10.1038/emboj.2008.154
Ishida, K., Kunibayashi, T., Tomikawa, C., Ochi, A., Kanai, T., Hirata, A., et al. (2011). Pseudouridine at position 55 in tRNA controls the contents of other modified nucleotides for low-temperature adaptation in the extreme-thermophilic eubacterium Thermus thermophilus. Nucleic Acids Res. 39, 2304–2318. doi: 10.1093/nar/gkq1180
Iwata-Reuyl, D. (2003). Biosynthesis of the 7-deazaguanosine hypermodified nucleosides of transfer RNA. Bioorg. Chem. 31, 24–43. doi: 10.1016/S0045-2068(02)00513-8
Jackman, J. E., Montange, R. K., Malik, H. S., and Phizicky, E. M. (2003). Identification of the yeast gene encoding the tRNA m1G methyltransferase responsible for modification at position 9. RNA 9, 574–585. doi: 10.1261/rna.5070303
Jakab, G., Kis, M., Pálfi, Z., and Solymosy, F. (1990). Nucleotide sequence of chloroplast tRNA(Leu)/UA m7G/from Chlamydomonas reinhardtii. Nucleic Acids Res. 18, 7444. doi: 10.1093/nar/18.24.7444
Jiang, H. Q., Motorin, Y., Jin, Y. X., and Grosjean, H. (1997). Pleiotropic effects of intron removal on base modification pattern of yeast tRNAPhe: an in vitro study. Nucleic Acids Res. 25, 2694–2701. doi: 10.1093/nar/25.14.2694
Joardar, A., Malliahgari, S. R., Skariah, G., and Gupta, R. (2011). 2'-O-methylation of the wobble residue of elongator pre-tRNA(Met) in Haloferax volcanii is guided by a box C/D RNA containing unique features. RNA Biol. 8, 782–791. doi: 10.4161/rna.8.5.16015
Jöckel, S., Nees, G., Sommer, R., Zhao, Y., Cherkasov, D., Hori, H., et al. (2012). The 2'-O-methylation status of a single guanosine controls transfer RNA-mediated Toll-like receptor 7 activation or inhibition. J. Exp. Med. 209, 235–241. doi: 10.1084/jem.20111075
Johansson, M. J., and Byström, A. S. (2002). Dual function of the tRNA(m(5)U54)methyltransferase in tRNA maturation. RNA 8, 324–335. doi: 10.1017/S1355838202027851
Johnson, G. D., Pirtle, I. L., and Pirtle, R. M. (1985). The nucleotide sequence of tyrosine tRNAQ* psi A from bovine liver. Arch. Biochem. Biophys. 236, 448–453. doi: 10.1016/0003-9861(85)90647-2
Joyce, G. F., and Orgel, L. E. (2006). “Chapter 2 Progress toward understanding the orgin of the RNA world,” in The RNA World, 3rd Edn., eds R. F. Gesteland, T. R. Cech, and J. F. Atkins (New York, NY: Cold Spring Harbor Laboratory Press), 23–56.
Jukes, T. H. (1973). Possibilities for the evolution of the genetic code from a preceding form. Nature 246, 22–26. doi: 10.1038/246022a0
Kadaba, S., Krueger, A., Trice, T., Krecic, A. M., Hinnebusch, A. G., and Anderson, J. (2004). Nuclear surveillance and degradation of hypomodified initiator tRNAMet in S. cerevisiae. Genes Dev. 18, 1227–1240. doi: 10.1101/gad.1183804
Kalhor, H. R., and Clarke, S. (2003). Novel methyltransferase for modified uridine residues at the wobble position of tRNA. Mol. Cell Biol. 23, 9283–9292. doi: 10.1128/MCB.23.24.9283-9292.2003
Kalhor, H. R., Penjwini, M., and Clarke, S. (2005). A novel methyltransferase required for the formation of the hypermodified nucleoside wybutosine in eucaryotic tRNA. Biochem. Biophys. Res. Commun. 334, 433–440. doi: 10.1016/j.bbrc.2005.06.111
Kaneko, T., Suzuki, T., Kapushoc, S. T., Rubio, M. A., Ghazvini, J., Watanabe, K., et al. (2003). Wobble modification differences and subcellular localization of tRNAs in Leishmania tarentolae: implication for tRNA sorting mechanism. EMBO J. 22, 657–667. doi: 10.1093/emboj/cdg066
Kawai, G., Yamamoto, Y., Kamimura, T., Masegi, T., Sekine, M., Hata, T., et al. (1992). Conformational rigidity of specific pyrimidine residues in tRNA arises from posttranscriptional modifications that enhance steric interaction between the base and the 2'-hydroxyl group. Biochemistry 31, 1040–1046. doi: 10.1021/bi00119a012
Keith, G., Desgrès, J., Pochart, P., Heyman, T., Kuo, K. C., and Gehrke, C. W. (1990). Eukaryotic tRNAs(Pro): primary structure of the anticodon loop; presence of 5-carbamoylmethyluridine or inosine as the first nucleoside of the anticodon. Biochim. Biophys. Acta 1049, 255–260. doi: 10.1016/0167-4781(90)90095-J
Kempenaers, M., Roovers, M., Oudjama, Y., Tkaczuk, K. L., Bujnicki, J. M., and Droogmans, L. (2010). New archaeal methyltransferases forming 1-methyladenosine or 1-methyladenosine and 1-methylguanosine at position 9 of tRNA. Nucleic Acids Res. 38, 6533–6543. doi: 10.1093/nar/gkq451
Kierzek, E., and Kierzek, R. (2003). The thermodynamic stability of RNA duplexes and hairpins containing N6-alkyladenosines and 2-methylthio-N6-alkyladenosines. Nucleic Acids Res. 31, 4472–4480. doi: 10.1093/nar/gkg633
Kim, J., and Almo, S. C. (2013). Structural basis for hypermodification of the wobble uridine in tRNA by bifunctional enzyme MnmC. BMC Struct. Biol. 13:5. doi: 10.1186/1472-6807-13-5
King, M. Y., and Redman, K. L. (2002). RNA methyltransferases utilize two cysteine residues in the formation of 5-methylcytosine. Biochemistry 41, 11218–11225. doi: 10.1021/bi026055q
Kirino, Y., Goto, Y., Campos, Y., Arenas, J., and Suzuki, T. (2005). Specific correlation between the wobble modification deficiency in mutant tRNAs and the clinical features of a human mitochondrial disease. Proc. Natl. Acad. Sci. U.S.A. 102, 7127–7132. doi: 10.1073/pnas.0500563102
Kitamura, A., Nishimoto, M., Sengoku, T., Shibata, R., Jäger, G., Björk, G. R., et al. (2012). Characterization and structure of the Aquifex aeolicus protein DUF752: a bacterial tRNA-methyltransferase (MnmC2) functioning without the usually fused oxidase domain (MnmC1). J. Biol. Chem. 287, 43950–43960. doi: 10.1074/jbc.M112.409300
Kitamura, A., Sengoku, T., Nishimoto, M., Yokoyama, S., and Bessho, Y. (2011). Crystal structure of the bifunctional tRNA modification enzyme MnmC from Escherichia coli. Protein Sci. 20, 1105–1113. doi: 10.1002/pro.659
Klassen, R., Paluszynski, J. P., Wemhoff, S., Pfeiffer, A., Fricke, J., and Meinhardt, F. (2008). The primary target of the killer toxin from Pichia acaciae is tRNA(Gln). Mol. Microbiol. 69, 681–697. doi: 10.1111/j.1365-2958.2008.06319.x
Kowalak, J. A., Dalluge, J. J., McCloskey, J. A., and Stetter, K. O. (1994). The role of posttranscriptional modification in stabilization of transfer RNA from hyperthermophiles. Biochemistry 33, 7869–7876. doi: 10.1021/bi00191a014
Kuchino, Y., and Borek, E. (1978). Tumour-specific phenylalanine tRNA contains two supernumerary methylated bases. Nature 271, 126–129. doi: 10.1038/271126a0
Kuchino, Y., Shindo-Okada, N., Ando, N., Watanabe, S., and Nishimura, S. (1981). Nucleotide sequences of two aspartic acid tRNAs from rat liver and rat ascites hepatoma. J. Biol. Chem. 256, 9059–9062.
Kurata, S., Weixlbaumer, A., Ohtsuki, T., Shimazaki, T., Wada, T., Kirino, Y., et al. (2008). Modified uridines with C5-methylene substituents at the first position of the tRNA anticodon stabilize U.G wobble pairing during decoding. J. Biol. Chem. 283, 18801–18811. doi: 10.1074/jbc.M800233200
Kuratani, M., Bessho, Y., Nishimoto, M., Grosjean, H., and Yokoyama, S. (2008). Crystal structure and mutational study of a unique SpoU family archaeal methylase that forms 2'-O-methylcytidine at position 56 of tRNA. J. Mol. Biol. 375, 1064–1075. doi: 10.1016/j.jmb.2007.11.023
Kuratani, M., Hirano, M., Goto-Ito, S., Itoh, Y., Hikida, Y., Nishimoto, M., et al. (2010). Crystal structure of Methanocaldococcus jannaschii Trm4 complexed with sinefungin. J. Mol. Biol. 401, 323–333. doi: 10.1016/j.jmb.2010.06.046
Lai, T. P., Stauffer, K. A., Murthi, A., Shaheen, H. H., Peng, G., Martin, N. C., et al. (2009). Mechanism and a peptide motif for targeting peripheral proteins to the yeast inner nuclear membrane. Traffic 10, 1243–1256. doi: 10.1111/j.1600-0854.2009.00956.x
Laxman, S., Sutter, B. M., Wu, X., Kumar, S., Guo, X., Trudgian, D. C., et al. (2013). Sulfur amino acids regulate translational capacity and metabolic homeostasis through modulation of tRNA thiolation. Cell 154, 416–429. doi: 10.1016/j.cell.2013.06.043
Lee, C., Kramer, G., Graham, D. E., and Appling, D. R. (2007). Yeast mitochondrial initiator tRNA is methylated at guanosine 37 by the Trm5-encoded tRNA (guanine-N1-)-methyltransferase. J. Biol. Chem. 282, 27744–27753. doi: 10.1074/jbc.M704572200
Leihne, V., Kirpekar, F., Vågbø, C. B., van den Born, E., Krokan, H. E., Grini, P. E., et al. (2011). Roles of Trm9- and ALKBH8-like proteins in the formation of modified wobble uridines in Arabidopsis tRNA. Nucleic Acids Res. 39, 7688–7701. doi: 10.1093/nar/gkr406
Leipuviene, R., and Björk, G. R. (2005). A reduced level of charged tRNAArgmnm5UCU triggers the wild-type peptidyl-tRNA to frameshift. RNA 11, 796–807. doi: 10.1261/rna.7256705
Leulliot, N., Chaillet, M., Durand, D., Ulryck, N., Blondeau, K., and van Tilbeurgh, H. (2008). Structure of the yeast tRNA m7G methylation complex. Structure 16, 52–61. doi: 10.1016/j.str.2007.10.025
Li, J., Esberg, B., Curran, J. F., and Björk, G. R. (1997). Three modified nucleosides present in the anticodon stem and loop influence the in vivo aa-tRNA selection in a tRNA-dependent manner. J. Mol. Biol. 271, 209–221. doi: 10.1006/jmbi.1997.1176
Liger, D., Mora, L., Lazar, N., Figaro, S., Henri, J., Scrima, N., et al. (2011). Mechanism of activation of methyltransferases involved in translation by the Trm112 ‘hub’ protein. Nucleic Acids Res. 39, 6249–6259. doi: 10.1093/nar/gkr176
Lim, K., Zhang, H., Tempczyk, A., Krajewski, W., Bonander, N., Toedt, J., et al. (2003). Structure of the YibK methyltransferase from Haemophilus influenzae (HI0766): a cofactor bound at a site formed by a knot. Proteins 51, 56–67. doi: 10.1002/prot.10323
Lin, J., Lai, S., Jia, R., Xu, A., Zhang, L., Lu, J., et al. (2011). Structural basis for site-specific ribose methylation by box C/D RNA protein complexes. Nature 469, 559–563. doi: 10.1038/nature09688
Liu, J., and Straby, K. B. (2000). The human tRNA(m(2)(2)G(26))dimethyltransferase: functional expression and characterization of a cloned hTRM1 gene. Nucleic Acids Res. 28, 3445–3451. doi: 10.1093/nar/28.18.3445
Liu, J., Wang, W., Shin, D. H., Yokota, H., Kim, R., and Kim, S. H. (2003). Crystal structure of tRNA (m1G37) methyltransferase from Aquifex aeolicus at 2.6 A resolution: a novel methyltransferase fold. Proteins 53, 326–328. doi: 10.1002/prot.10479
Liu, J., Zhou, G. Q., and Straby, K. B. (1999). Caenorhabditis elegans ZC376.5 encodes a tRNA (m2/2G(26))dimethyltransferance in which (246)arginine is important for the enzyme activity. Gene 226, 73–81. doi: 10.1016/S0378-1119(98)00550-2
Liu, R. J., Zhou, M., Fang, Z. P., Wang, M., Zhou, X. L., and Wang, E. D. (2013). The tRNA recognition mechanism of the minimalist SPOUT methyltransferase, TrmL. Nucleic Acids Res. 41, 7828–7842. doi: 10.1093/nar/gkt568
Lu, J., Huang, B., Esberg, A., Johansson, M. J., and Byström, A. S. (2005). The Kluyveromyces lactis gamma-toxin targets tRNA anticodons. RNA 11, 1648–1654. doi: 10.1261/rna.2172105
Machnicka, M. A., Milanowska, K., Osman Oglou, O., Purta, E., Kurkowska, M., Olchowik, A., et al. (2013). MODOMICS: a database of RNA modification pathways–2013 update. Nucleic Acids Res. 41, D262–267. doi: 10.1093/nar/gks1007
Marquet, R. (1998). “Chapter 28 Importance of modified nucleotides in replication of retrovirus, plant pararetrovirus, and retrotransposons,” in Modification and Editing of RNA, eds H. Grosjean and R. Benne (Washington, DC: ASM press), 517–533.
Martin, N. C., and Hopper, A. K. (1994). How single genes provide tRNA processing enzymes to mitochondria, nuclei and the cytosol. Biochimie 76, 1161–1167. doi: 10.1016/0300-9084(94)90045-0
Matsumoto, K., Toyooka, T., Tomikawa, C., Ochi, A., Takano, Y., Takayanagi, N., et al. (2007). RNA recognition mechanism of eukaryote tRNA (m7G46) methyltransferase (Trm8-Trm82 complex). FEBS Lett. 581, 1599–1604. doi: 10.1016/j.febslet.2007.03.023
Matsuyama, S., Ueda, T., Crain, P. F., McCloskey, J. A., and Watanabe, K. (1998). A novel wobble rule found in starfish mitochondria. Presence of 7-methylguanosine at the anticodon wobble position expands decoding capability of tRNA. J. Biol. Chem. 273, 3363–3368. doi: 10.1074/jbc.273.6.3363
Mazauric, M. H., Dirick, L., Purushothaman, S. K., Björk, G. R., and Lapeyre, B. (2010). Trm112p is a 15-kDa zinc finger protein essential for the activity of two tRNA and one protein methyltransferases in yeast. J. Biol. Chem. 285, 18505–18515. doi: 10.1074/jbc.M110.113100
McCrate, N. E., Varner, M. E., Kim, K. I., and Nagan, M. C. (2006). Molecular dynamics simulations of human tRNA Lys,3 UUU: the role of modified bases in mRNA recognition Nucleic Acids Res. 34, 5361–5368. doi: 10.1093/nar/gkl580
Menezes, S., Gaston, K. W., Krivos, K. L., Apolinario, E. E., Reich, N. O., Sowers, K. R., et al. (2011). Formation of m2G6 in Methanocaldococcus jannaschii tRNA catalyzed by the novel methyltransferase Trm14. Nucleic Acids Res. 39, 7641–7655. doi: 10.1093/nar/gkr475
Meyer, S., Scrima, A., Versées, W., and Wittinghofer, A. (2008). Crystal structures of the conserved tRNA-modifying enzyme GidA: implications for its interaction with MnmE and substrate. J. Mol. Biol. 380, 532–547. doi: 10.1016/j.jmb.2008.04.072
Meyer, S., Wittinghofer, A., and Versées, W. (2009). G-domain dimerization orchestrates the tRNA wobble modification reaction in the MnmE/GidA complex. J. Mol. Biol. 392, 910–922. doi: 10.1016/j.jmb.2009.07.004
Miyauchi, K., Kimura, S., and Suzuki, T. (2013). A cyclic form of N6-threonylcarbamoyladenosine as a widely distributed tRNA hypermodification. Nat. Chem. Biol. 9, 105–111. doi: 10.1038/nchembio.1137
Motorin, Y., and Grosjean, H. (1999). Multisite-specific tRNA:m5C-methyltransferase (Trm4) in yeast Saccharomyces cerevisiae: identification of the gene and substrate specificity of the enzyme. RNA 5, 1105–1118. doi: 10.1017/S1355838299982201
Motorin, Y., and Helm, M. (2010). tRNA stabilization by modified nucleotides. Biochemistry 49, 4934–4944. doi: 10.1021/bi100408z
Moukadiri, I., Garzón, M. J., Björk, G. R., and Armengod, M. E. (2014). The output of the tRNA modification pathways controlled by the Escherichia coli MnmEG and MnmC enzymes depends on the growth conditions and the tRNA species. Nucleic Acids Res. 42, 2603–2623. doi: 10.1093/nar/gkt1228
Moukadiri, I., Prado, S., Piera, J., Velázquez-Campoy, A., Björk, G. R., and Armengod, M. E. (2009). Evolutionarily conserved proteins MnmE and GidA catalyze the formation of two methyluridine derivatives at tRNA wobble positions. Nucleic Acids Res. 37, 7177–7193. doi: 10.1093/nar/gkp762
Nasvall, S. J., Chen, P., and Björk, G. R. (2004). The modified wobble nucleoside uridine-5-oxyacetic acid in tRNAPro(cmo5UGG) promotes reading of all four proline codons in vivo. RNA 10, 1662–1673. doi: 10.1261/rna.7106404
Niederberger, C., Graub, R., Costa, A., Desgres, J., and Schweingruber, M. E. (1999). The tRNA N2,N2-dimethylguanosine-26 methyltransferase encoded by gene trm1 increases efficiency of suppression of an ochre codon in Schizosaccharomyces pombe. FEBS Lett. 464, 67–70. doi: 10.1016/S0014-5793(99)01679-8
Nishimasu, H., Ishitani, R., Yamashita, K., Iwashita, C., Hirata, A., Hori, H., et al. (2009). Atomic structure of a folate/FAD-dependent tRNA T54 methyltransferase. Proc. Natl. Acad. Sci. U.S.A. 106, 8180–8185. doi: 10.1073/pnas.0901330106
Noma, A., Ishitani, R., Kato, M., Nagao, A., Nureki, O., and Suzuki, T. (2010). Expanding role of the jumonji C domain as an RNA hydroxylase. J. Biol. Chem. 285, 34503–34507. doi: 10.1074/jbc.M110.156398
Noma, A., Kirino, Y., Ikeuchi, Y., and Suzuki, T. (2006). Biosynthesis of wybutosine, a hyper-modified nucleoside in eukaryotic phenylalanine tRNA. EMBO J. 25, 2142–2154. doi: 10.1038/sj.emboj.7601105
Noma, A., Yi, S., Katoh, T., Takai, Y., Suzuki, T., and Suzuki, T. (2011). Actin-binding protein ABP140 is a methyltransferase for 3-methylcytidine at position 32 of tRNAs in Saccharomyces cerevisiae. RNA 17, 1111–1119. doi: 10.1261/rna.2653411
Nordlund, M. E., Johansson, J. O., von Pawel-Rammingen, U., and Byström, A. S. (2000). Identification of the TRM2 gene encoding the tRNA(m5U54)methyltransferase of Saccharomyces cerevisiae. RNA 6, 844–860. doi: 10.1017/S1355838200992422
Novoa, E. M., Pavon-Eternod, M., Pan, T., and de Pouplana, L. R. (2012). A role for tRNA modifications in genome structure and codon usage. Cell 149, 202–213. doi: 10.1016/j.cell.2012.01.050
Nureki, O., Watanabe, K., Fukai, S., Ishii, R., Endo, Y., Hori, H., et al. (2004). Deep knot structure for construction of active site and cofactor binding site of tRNA modification enzyme. Structure 12, 593–604. doi: 10.1016/j.str.2004.03.003
Ny, T., and Bjork, G. R. (1980). Cloning and restriction mapping of the trmA gene coding for transfer ribonucleic acid (5-methyluridine)-methyltransferase in Escherichia coli K-12. J. Bacteriol. 142, 371–379.
Ny, T., Lindstrom, P. H. R., Hagervall, T. G., and Bjork, G. R. (1988). Purification of transfer RNA (m5U54)-methyltransferase from Escherichia coli association with RNA. Eur. J. Biochem. 177, 467–475. doi: 10.1111/j.1432-1033.1988.tb14396.x
Ochi, A., Makabe, K., Kuwajima, K., and Hori, H. (2010). Flexible recognition of the tRNA G18 methylation target site by TrmH methyltransferase through first binding and induced fit processes. J. Biol. Chem. 285, 9018–9029. doi: 10.1074/jbc.M109.065698
Ochi, A., Makabe, K., Yamagami, R., Hirata, A., Sakaguchi, R., Hou, Y. M., et al. (2013). The catalytic domain of topological knot tRNA methyltransferase (TrmH) discriminates between substrate tRNA and nonsubstrate tRNA via an induced-fit process. J. Biol. Chem. 288, 25562–25574. doi: 10.1074/jbc.M113.485128
O'Dwyer, K., Watts, J. M., Biswas, S., Ambrad, J., Barber, M., and Holmes, W. M. (2004). Characterization of Streptococcus pneumoniae TrmD, a tRNA methyltransferase essential for growth. J. Bacteriol. 186, 2346–2354. doi: 10.1128/JB.186.8.2346-2354.2004
Ohira, T., and Suzuki, T. (2011). Retrograde nuclear import of tRNA precursors is required for modified base biogenesis in yeast. Proc. Natl. Acad. Sci. U.S.A. 108, 10502–10507. doi: 10.1073/pnas.1105645108
Ohira, T., Suzuki, T., Miyauchi, K., Suzuki, T., Yokobori, S., Yamagishi, A., et al. (2013). Decoding mechanism of non-universal genetic codes in Loligo bleekeri mitochondria. J. Biol. Chem. 288, 7645–7652. doi: 10.1074/jbc.M112.439554
Okamoto, H., Watanabe, K., Ikeuchi, Y., Suzuki, T., Endo, Y., and Hori, H. (2004). Substrate tRNA recognition mechanism of tRNA (m7G46) methyltransferase from Aquifex aeolicus. J. Biol. Chem. 279, 49151–49159. doi: 10.1074/jbc.M408209200
Osawa, S., Jukes, T. H., Watanabe, K., and Muto, A. (1992). Recent evidence for evolution of the genetic code. Microbiol. Rev. 56, 229–264.
Osawa, T., Ito, K., Inanaga, H., Nureki, O., Tomita, K., and Numata, T. (2009). Conserved cysteine residues of GidA are essential for biogenesis of 5-carboxymethylaminomethyluridine at tRNA anticodon. Structure 17, 713–724. doi: 10.1016/j.str.2009.03.013
Osorio-Almeida, M. L., Guillemaut, P., Keith, G., Canaday, J., and Weil, J. H. (1980). Primary structure of three leucine transfer RNAs from bean chloroplast. Biochem. Biophys. Res. Commun. 92, 102–108. doi: 10.1016/0006-291X(80)91525-9
Ote, T., Hashimoto, M., Ikeuchi, Y., Su'etsugu, M., Suzuki, T., Katayama, T., et al. (2006). Involvement of the Escherichia coli folate-binding protein YgfZ in RNA modification and regulation of chromosomal replication initiation. Mol. Microbol. 59, 265–275. doi: 10.1111/j.1365-2958.2005.04932.x
Ozanick, S. G., Bujnicki, J. M., Sem, D. S., and Anderson, J. T. (2007). Conserved amino acids in each subunit of the heteroligomeric tRNA m1A58 Mtase from Saccharomyces cerevisiae contribute to tRNA binding. Nucleic Acids Res. 35, 6808–6819. doi: 10.1093/nar/gkm574
Paris, Z., Horáková, E., Rubio, M. A., Sample, P., Fleming, I. M., Armocida, S., et al. (2013). The T. brucei TRM5 methyltransferase plays an essential role in mitochondrial protein synthesis and function. RNA 19, 649–658. doi: 10.1261/rna.036665.112
Pastore, C., Topalidou, I., Forouhar, F., Yan, A. C., Levy, M., and Hunt, J. F. (2012). Crystal structure and RNA binding properties of the RNA recognition motif (RRM) and AlkB domains in human AlkB homolog 8 (ABH8), an enzyme catalyzing tRNA hypermodification. J. Biol. Chem. 287, 2130–2143. doi: 10.1074/jbc.M111.286187
Patil, A., Chan, C. T., Dyavaiah, M., Rooney, J. P., Dedon, P. C., and Begley, T. J. (2012a). Translational infidelity-induced protein stress results from a deficiency in Trm9-catalyzed tRNA modifications. RNA Biol. 9, 990–1001. doi: 10.4161/rna.20531
Patil, A., Dyavaiah, M., Joseph, F., Rooney, J. P., Chan, C. T., Dedon, P. C., et al. (2012b). Increased tRNA modification and gene-specific codon usage regulate cell cycle progression during the DNA damage response. Cell Cycle 11, 3656–3665. doi: 10.4161/cc.21919
Pearson, D., and Carell, T. (2011). Assay of both activities of the bifunctional tRNA-modifying enzyme MnmC reveals a kinetic basis for selective full modification of cmnm5s2U to mnm5s2U. Nucleic Acids Res. 39, 4818–4826. doi: 10.1093/nar/gkr071
Perche-Letuvée, P., Kathirvelu, V., Berggren, G., Clemancey, M., Latour, J. M., Maurel, V., et al. (2012). 4-Demethylwyosine synthase from Pyrococcus abyssi is a radical-S-adenosyl-L-methionine enzyme with an additional [4Fe-4S](+2) cluster that interacts with the pyruvate co-substrate. J. Biol. Chem. 287, 41174–41185. doi: 10.1074/jbc.M112.405019
Perret, V., Garcia, A., Grosjean, H., Ebel, J. P., Florentz, C., and Giegé, R. (1990). Relaxation of a transfer RNA specificity by removal of modified nucleotides. Nature 344, 787–789. doi: 10.1038/344787a0
Perrochia, L., Crozat, E., Hecker, A., Zhang, W., Bareille, J., Collinet, B., et al. (2013). In vitro biosynthesis of a universal t6A tRNA modification in Archaea and Eukarya. Nucleic Acids Res. 41, 1953–1964. doi: 10.1093/nar/gks1287
Persson, B. C., Jäger, G., and Gustafsson, C. (1997). The spoU gene of Escherichia coli, the fourth gene of the spoT operon, is essential for tRNA (Gm18) 2'-O-methyltransferase activity. Nucleic Acid Res. 25, 3969–3973. doi: 10.1093/nar/25.20.4093
Persson, B. C., Olafsson, O., Lundgren, H. K., Hederstedt, L., and Björk, G. R. (1998). The ms2io6A37 modification of tRNA in Salmonella typhimurium regulates growth on citric acid cycle intermediates. J. Bacteriol. 180, 3144–3151.
Phillips, G., Swairjo, M. A., Gaston, K. W., Bailly, M., Limbach, P. A., Iwata-Reuyl, D., et al. (2012). Diversity of archaeosine synthesis in crenarchaeota. ACS Chem. Biol. 7, 300–305. doi: 10.1021/cb200361w
Phillips, J. H., and Kjellin-Straby, K. (1967). Studies on microbial ribonucleic acid. IV. Two mutants of Saccharomyces cerevisiae lacking N-2-dimethylguanine in soluble ribonucleic acid. J. Mol. Biol. 26, 509–518. doi: 10.1016/0022-2836(67)90318-X
Phizicky, E. M., and Hopper, A. K. (2010). tRNA biology charges to the front. Genes Dev. 24, 1832–1860. doi: 10.1101/gad.1956510
Pierre, A., Berneman, A., Vedel, M., Robert-Gero, M., and Vigier, P. (1978). Avian oncornavirus associated N2-methylguanine transferase, location and origin. Biochem. Biophys. Res. Commun. 81, 315–321. doi: 10.1016/0006-291X(78)91535-8
Pierrel, F., Hernandez, H. L., Johnson, M. K., Fontecave, M., and Atta, M. (2003). MiaB protein from Thermotoga maritima. characterization of an extremely thermophilic tRNA-methylthiotransferase. J. Biol. Chem. 278, 29515–29524. doi: 10.1074/jbc.M301518200
Pintard, L., Lecointe, F., Bujnicki, J. M., Bonnerot, C., Grosjean, H., and Lapeyre, B. (2002). Trm7p catalyses the formation of two 2'-O-methylriboses in yeast tRNA anticodon loop. EMBO J. 21, 1811–1820. doi: 10.1093/emboj/21.7.1811
Pleshe, E., Truesdell, J., and Batey, R. T. (2005). Structure of a class II TrmH tRNA-modifying enzyme from Aquifex aeolicus. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 61, 722–728. doi: 10.1107/S1744309105022980
Pope, W. T., Brown, A., and Reeves, R. H. (1978). The identification of the tRNA substrates for the supK tRNA methylase. Nucleic Acids Res. 5, 1041–1057. doi: 10.1093/nar/5.3.1041
Preston, M. A., D'Silva, S., Kon, Y., and Phizicky, E. M. (2013). tRNAHis 5-methylcytidine levels increase in response to several growth arrest conditions in Saccharomyces cerevisiae. RNA 19, 243–256. doi: 10.1261/rna.035808.112
Purta, E., van Vliet, F., Tkaczuk, K. L., Dunin-Horkawicz, S., Mori, H., Droogmans, L., et al. (2006). The yfhQ gene of Escherichia coli encodes a tRNA:Cm32/Um32 methyltransferase. BMC Mol. Biol. 7:23. doi: 10.1186/1471-2199-7-23
Purushothaman, S. K., Bujnicki, J. M., Grosjean, H., and Lapeyre, B. (2005). Trm11p and Trm112p are both required for the formation of 2-methylguanosine at position 10 in yeast tRNA. Mol. Cell Biol. 25, 4359–4370. doi: 10.1128/MCB.25.11.4359-4370.2005
Qian, Q., Curran, J. F., and Björk, G. R. (1998). The methyl group of the N6-methyl-N6-threonylcarbamoyladenosine in tRNA of Escherichia coli modestly improves the efficiency of the tRNA. J. Bacteriol. 180, 1808–1813.
Qiu, X., Huang, K., Ma, J., and Gao, Y. (2011). Crystallization and preliminary X-ray diffraction crystallographic study of tRNA m(1)A58 methyltransferase from Saccharomyces cerevisiae. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 67, 1448–1450. doi: 10.1107/S174430911103733X
Raba, M., Limburg, K., Burghagen, M., Katze, J. R., Simsek, M., Heckman, J. E., et al. (1979). Nucleotide sequence of three isoaccepting lysine tRNAs from rabbit liver and SV40-transformed mouse fibroblasts. Eur. J. Biochem. 97, 305–318. doi: 10.1111/j.1432-1033.1979.tb13115.x
Reinhart, M. P., Lewis, J. M., and Leboy, P. S. (1986). A single tRNA (guanine)-methyltransferase from tetrahymena with both mono- and di-methylating activity. Nucleic Acids Res. 14, 1131–1148. doi: 10.1093/nar/14.3.1131
Renalier, M. H., Joseph, N., Gaspin, C., Thebault, P., and Mougin, A. (2005). The Cm56 tRNA modification in archaea is catalyzed either by a specific 2'-O-methylase or a C/D sRNP. RNA 11, 1051–1063. doi: 10.1261/rna.2110805
Roovers, M., Kaminska, K. H., Tkaczuk, K. L., Gigot, D., Droogmans, L., and Bujnicki, J. M. (2008a). The YqfN protein of Bacillus subtilis is the tRNA: m1A22 methyltransferase (TrmK). Nucleic Acids Res. 36, 3252–3262. doi: 10.1093/nar/gkn169
Roovers, M., Oudjama, Y., Fislage, M., Bujnicki, J. M., Versees, W., and Droogmans, L. (2012). The open reading frame TTC1157 of Thermus thermophilus HB27 encodes the methyltransferase forming N2-methylguanosine at position 6 in tRNA. RNA 18, 815–824. doi: 10.1261/rna.030411.111
Roovers, M., Oudjama, Y., Kaminska, K. H., Purta, E., Caillet, J., Droogmans, L., et al. (2008b). Sequence-structure-function analysis of the bifunctional enzyme MnmC that catalyses the last two steps in the biosynthesis of hypermodified nucleoside mnm5s2U in tRNA. Proteins 71, 2076–2085. doi: 10.1002/prot.21918
Roovers, M., Wouters, J., Bujnicki, J. M., Tricot, C., Stalon, V., Grosjean, H., et al. (2004). A primordial RNA modification enzyme: the case of tRNA (m1A) methyltransferase. Nucleic Acids Res. 32, 465–476. doi: 10.1093/nar/gkh191
Saadatmand, J., and Kleiman, L. (2012). Aspects of HIV-1 assembly that promote primer tRNA(Lys3) annealing to viral RNA. Virus Res. 169, 340–348. doi: 10.1016/j.virusres.2012.06.001
Saga, A. E., Vasil, A. I., and Vasil, M. L. (1997). Molecular characterization of mutants affected in the osmoprotectant-dependent induction of phspholipase C in Pseudomonas aeruginosa PAO1. Mol. Microbiol. 23, 43–56. doi: 10.1046/j.1365-2958.1997.1681542.x
Sakaguchi, R., Giessing, A., Dai, Q., Lahoud, G., Liutkeviciute, Z., Klimasauskas, S., et al. (2012). Recognition of guanosine by dissimilar tRNA methyltransferases. RNA 18, 1687–1701. doi: 10.1261/rna.032029.111
Sakurai, M., Ohtsuki, T., Suzuki, T., and Watanabe, K. (2005b). Unusual usage of wobble modifications in mitochondrial tRNAs of the nematode Ascaris suum. FEBS Lett. 579, 2767–2772. doi: 10.1016/j.febslet.2005.04.009
Sakurai, M., Otsuki, T., and Watanabe, K. (2005a). Modification at position 9 with 1-methyladenosine is crucial for structure and function of nematode mitochondrial tRNAs lacking the entire T-arm. Nucleic Acids Res. 33, 1653–1661. doi: 10.1093/nar/gki309
Schaub, M., and Keller, W. (2002). RNA editing by adenosine deaminases generates RNA and protein diversity. Biochimie 84, 791–803. doi: 10.1016/S0300-9084(02)01446-3
Schubert, H. G., Blumenthal, R. M., and Cheng, X. (2003). Many paths to methyltransfer: a chronicle of convergence. Trends Biochem. Sci. 28, 329–332. doi: 10.1016/S0968-0004(03)00090-2
Shao, Z., Yan, W., Peng, J., Zuo, X., Zou, Y., Li, F., et al. (2013). Crystal structure of tRNA m1G9 methyltransferase Trm10: insight into the catalytic mechanism and recognition of tRNA substrate. Nucleic Acids Res. 42, 509–525. doi: 10.1093/nar/gkt869
Shi, R., Villarroya, M., Ruiz-Partida, R., Li, Y., Proteau, A., Prado, S., et al. (2009). Structure-function analysis of Escherichia coli MnmG (GidA), a highly conserved tRNA-modifying enzyme. J. Bacteriol. 191, 7614–7619. doi: 10.1128/JB.00650-09
Shigi, N., Sakaguchi, Y., Asai, S., Suzuki, T., and Watanabe, K. (2008). Common thiolation mechanism in the biosynthesis of tRNA thiouridine and sulphur-containing cofactors. EMBO J. 27, 3267–3278. doi: 10.1038/emboj.2008.246
Shigi, N., Suzuki, T., Tamakoshi, M., Oshima, T., and Watanabe, K. (2002). Conserved bases in the TPsi C loop of tRNA are determinants for thermophile-specific 2-thiouridylation at position 54. J. Biol. Chem. 277, 39128–39135. doi: 10.1074/jbc.M207323200
Shigi, N., Suzuki, T., Terada, T., Shirouzu, M., Yokoyama, S., and Watanabe, K. (2006). Temperature-dependent biosynthesis of 2-thioribothymidine of Thermus thermophilus tRNA. J. Biol. Chem. 281, 2104–2113. doi: 10.1074/jbc.M510771200
Shimada, K., Nakamura, M., Anai, S., De Velasco, M., Tanaka, M., Tsujikawa, K., et al. (2009). A novel human AlkB homologue, ALKBH8, contributes to human bladder cancer progression. Cancer Res. 69, 3157–3164. doi: 10.1158/0008-5472.CAN-08-3530
Shindo-Okada, N., Kuchino, Y., Harada, F., Okada, N., and Nishimura, S. (1981). Biological and structural differences between tRNAVal species isolated from rat ascites hepatoma cells and normal rat liver. J. Biochem. 90, 535–544.
Singh, S. K., Gurha, P., Tran, E. J., Maxwell, E. S., and Gupta, R. (2004). Sequential 2'-O-methylation of archaeal pre-tRNATrp nucleotides is guided by the intron-encoded but trans-acting box C/D ribonucleoprotein of pre-tRNA. J. Biol. Chem. 279, 47661–47671. doi: 10.1074/jbc.M408868200
Sleiman, D., Goldschmidt, V., Barraud, P., Marquet, R., Paillart, J. C., and Tisné, C. (2012). Initiation of HIV-1 reverse transcription and functional role of nucleocapsid-mediated tRNA/viral genome interactions. Virus Res. 169, 324–339. doi: 10.1016/j.virusres.2012.06.006
Songe-Møller, L., van den Born, E., Leihne, V., Vågbø, C. B., Kristoffersen, T., Krokan, H. E., et al. (2010). Mammalian ALKBH8 possesses tRNA methyltransferase activity required for the biogenesis of multiple wobble uridine modifications implicated in translational decoding. Mol. Cell Biol. 30, 1814–1827. doi: 10.1128/MCB.01602-09
Suzuki, T., and Miyauchi, K. (2010). Discovery and characterization of tRNAIle lysidine synthetase (TilS). FEBS Lett. 584, 272–277. doi: 10.1016/j.febslet.2009.11.085
Suzuki, T., Miyauchi, K., Suzuki, T., Yokobori, S., Shigi, N., Kondow, A., et al. (2011a). Taurine-containing uridine modifications in tRNA anticodons are required to decipher non-universal genetic codes in ascidian mitochondria. J. Biol. Chem. 286, 35494–35498. doi: 10.1074/jbc.M111.279810
Suzuki, T., Nagao, A., and Suzuki, T. (2011b). Human mitochondrial diseases caused by lack of taurine modification in mitochondrial tRNAs. Willey Interdiscip. Rev. RNA 2, 376–386. doi: 10.1002/wrna.65
Suzuki, T., Suzuki, T., Wada, T., Saigo, K., and Watanabe, K. (2002). Taurine as a constituent of mitochondrial tRNAs: new insights into the functions of taurine and human mitochondrial diseases. EMBO J. 21, 6581–6589. doi: 10.1093/emboj/cdf656
Suzuki, Y., Noma, A., Suzuki, T., Senda, M., Senda, T., Ishitani, R., et al. (2007). Crystal structure of the radical SAM enzyme catalyzing tricyclic modified base formation in tRNA. J. Mol. Biol. 372, 1204–1214. doi: 10.1016/j.jmb.2007.07.024
Swinehart, W. E., Henderson, J. C., and Jackman, J. E. (2013). Unexpected expansion of tRNA substrate recognition by the yeast m1G9 methyltransferase Trm10. RNA 19, 1137–1146. doi: 10.1261/rna.039651.113
Takai, K., and Yokoyama, S. (2003). Roles of 5-substituents of tRNA wobble uridines in the recognition of purine-ending codons. Nucleic Acids Res. 31, 6383–6391. doi: 10.1093/nar/gkg839
Takano, A., Endo, T., and Yoshihisa, T. (2005). tRNA actively shuttles between the nucleus and cytosol in yeast. Science 309, 140–142. doi: 10.1126/science.1113346
Takano, Y., Takayanagi, N., Hori, H., Ikeuchi, Y., Suzuki, T., Kimura, A., et al. (2006). A gene involved in modifying transfer RNA is required for fungal pathogenicity and stress tolerance of Colletotrichum lagenarium. Mol. Microbiol. 60, 81–92. doi: 10.1111/j.1365-2958.2006.05080.x
Takeda, H., Toyooka, T., Ikeuchi, Y., Yokobori, S., Okadome, K., Takano, F., et al. (2006). The substrate specificity of tRNA (m1G37) methyltransferase (TrmD) from Aquifex aeolicus. Genes Cells 11, 1353–1365. doi: 10.1111/j.1365-2443.2006.01022.x
Taya, Y., and Nishimura, S. (1973). Biosynthesis of 5-methylaminomethyl-2-thiouridylate. I. Isolation of a new tRNA-methylase specific for 5-methylaminomethyl-2-thiouridylate. Biochem. Biophys. Res. Commun. 51, 1062–1068. doi: 10.1016/0006-291X(73)90035-1
Tkaczuk, K. L. (2010). Trm13p, the tRNA:Xm4 modification enzyme from Saccharomyces cerevisiae is a member of the Rossmann-fold MTase superfamily: prediction of structure and active site. J. Mol. Model. 16, 599–606. doi: 10.1007/s00894-009-0570-6
Tkaczuk, K. L., Dunin-Horkawicz, S., Purta, E., and Bujnicki, J. M. (2007). Structural and evolutionary bioinformatics of the SPOUT superfamily of methyltransferases. BMC Bioinformatics 8:73. doi: 10.1186/1471-2105-8-73
Tomikawa, C., Ochi, A., and Hori, H. (2008). The C-terminal region of thermophilic tRNA (m7G46) methyltransferase (TrmB) stabilizes the dimer structure and enhances fidelity of methylation. Proteins 71, 1400–1408. doi: 10.1002/prot.21827
Tomikawa, C., Ohira, T., Inoue, Y., Kawamura, T., Yamagishi, A., Suzuki, T., et al. (2013). Distinct tRNA modifications in the thermo-acidophilic archaeon, Thermoplasma acidophilum. FEBS Lett. 587, 3575–3580. doi: 10.1016/j.febslet.2013.09.021
Tomikawa, C., Yokogawa, T., Kanai, T., and Hori, H. (2010). N7-Methylguanine at position 46 (m7G46) in tRNA from Thermus thermophilus is required for cell viability through a tRNA modification network. Nucleic Acids Res. 38, 942–957. doi: 10.1093/nar/gkp1059
Tomita, K., Ueda, T., and Watanabe, K. (1998). 7-Methylguanosine at the anticodon wobble position of squid mitochondrial tRNA(Ser)GCU: molecular basis for assignment of AGA/AGG codons as serine in invertebrate mitochondria. Biochim. Biophys. Acta 1399, 78–82. doi: 10.1016/S0167-4781(98)00099-2
Towns, W. L., and Begley, T. J. (2012). Transfer RNA methytransferases and their corresponding modifications in budding yeast and humans: activities, predications, and potential roles in human health. DNA Cell Biol. 31, 434–454. doi: 10.1089/dna.2011.1437
Toyooka, T., Awai, T., Kanai, T., Imanaka, T., and Hori H. (2008). Stabilization of tRNA (m1G37) methyltransferase [TrmD] from Aquifex aeolicus by an intersubunit disulfide bond formation. Genes Cells 13, 807–816. doi: 10.1111/j.1365-2443.2008.01207.x
Tsutsumi, S., Sugiura, R., Ma, Y., Tokuoka, H., Ohta, K., Ohte, R., et al. (2007). Wobble inosine tRNA modification is essential to cell cycle progression in G(1)/S and G(2)/M transitions in fission yeast. J. Biol. Chem. 282, 33459–33465. doi: 10.1074/jbc.M706869200
Umeda, N., Suzuki, T., Yukawa, M., Ohya, Y., Shindo, H., Watanabe, K., et al. (2005). Mitochondria-specific RNA-modifying enzymes responsible for the biosynthesis of the wobble base in mitochondrial tRNAs. Implications for the molecular pathogenesis of human mitochondrial diseases. J. Biol. Chem. 280, 1613–1624. doi: 10.1074/jbc.M409306200
Umitsu, M., Nishimasu, H., Noma, A., Suzuki, T., Ishitani, R., and Nureki, O. (2009). Structural basis of AdoMet-dependent aminocarboxypropyl transfer reaction catalyzed by tRNA-wybutosine synthesizing enzyme, TYW2. Proc. Natl. Acad. Sci. U.S.A. 106, 15616–15621. doi: 10.1073/pnas.0905270106
Urbonavicius, J., Armengaud, J., and Grosjean, H. (2006). Identity elements required for enzymatic formation of N2,N2-dimethylguanosine from N2-monomethylated derivative and its possible role in avoiding alternative conformations in archaeal tRNA. J. Mol. Biol. 357, 387–399. doi: 10.1016/j.jmb.2005.12.087
Urbonavicius, J., Durand, J. M., and Björk, G. R. (2002). Three modifications in the D and T arms of tRNA influence translation in Escherichia coli and expression of virulence genes in Shigella flexneri. J. Bacteriol. 184, 5384–5357. doi: 10.1128/JB.184.19.5348-5357.2002
Urbonavicius, J., Qian, Q., Durand, J. M., Hagervall, T. G., and Björk, G. R. (2001). Improvement of reading frame maintenance is a common function for several tRNA modifications. EMBO J. 20, 4863–4873. doi: 10.1093/emboj/20.17.4863
Urbonavicius, J., Skouloubris, S., Myllykallio, H., and Grosjean, H. (2005). Identification of a novel gene encoding a flavin-dependent tRNA:m5U methyltransferase in bacteria–evolutionary implications. Nucleic Acids Res. 33, 3955–3964. doi: 10.1093/nar/gki703
Urbonavicius, J., Stahl, G., Durand, J. M., Ben Salem, S. N., Qian, Q., Farabaugh, P. J., et al. (2003). Transfer RNA modifications that alter +1 frameshifting in general fail to affect -1 frameshifting. RNA 9, 760–768. doi: 10.1261/rna.5210803
van den Born, E., Vågbø, C. B., Songe-Møller, L., Leihne, V., Lien, G. F., Leszczynska, G., et al. (2011). ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA. Nat Commun. 2, 172. doi: 10.1038/ncomms1173
van Tol, H., Stange, N., Gross, H. J., and Beier, H. (1987). A human and a plant intron-containing tRNATyr gene are both transcribed in a HeLa cell extract but spliced along different pathways. EMBO J. 6, 35–41.
Vilardo, E., Nachbagauer, C., Buzet, A., Taschner, A., Holzmann, J., and Rossmanith, W. (2012). A subcomplex of human mitochondrial RNase P is a bifunctional methyltransferase–extensive moonlighting in mitochondrial tRNA biogenesis. Nucleic Acids Res. 40, 11583–11593. doi: 10.1093/nar/gks910
Waas, W. F., de Crécy-Lagard, V., and Schimmel, P. (2005). Discovery of a gene family critical to wyosine base formation in a subset of phenylalanine-specific transfer RNAs. J. Biol. Chem. 280, 37616–37622. doi: 10.1074/jbc.M506939200
Waas, W. F., Druzina, Z., Hanan, M., and Schimmel, P. (2007). Role of a tRNA base modification and its precursors in frameshifting in eukaryotes. J. Biol. Chem. 282, 26026–26034. doi: 10.1074/jbc.M703391200
Walbott, H., Auxilien, S., Grosjean, H., and Golinelli-Pimpaneau, B. (2007). The carboxyl-terminal extension of yeast tRNA m5C methyltransferase enhances the catalytic efficiency of the amino-terminal domain. J. Biol. Chem. 282, 23663–23671. doi: 10.1074/jbc.M703818200
Walker, R. T. (1983). Mycoplasma evolution: a review of the use of ribosomal and transfer RNA Nucleotide sequences in the determination of phylogenetic relationships, yale. J. Biol. Med. 56, 367–372.
Watanabe, K., Nureki, O., Fukai, S., Endo, Y., and Hori, H. (2006). Functional categorization of the conserved basic amino acid residues in TrmH (tRNA (Gm18) methyltransferase) enzymes. J. Biol. Chem. 281, 34630–34639. doi: 10.1074/jbc.M606141200
Watanabe, K., Nureki, O., Fukai, S., Ishii, R., Okamoto, H., Yokoyama, S., et al. (2005). Roles of conserved amino acid sequence motifs in the SpoU (TrmH) RNA methyltransferase family. J. Biol. Chem. 280, 10368–10377. doi: 10.1074/jbc.M411209200
Watanabe, K., Shinma, M., Oshima, T., and Nishimura, S. (1976). Heat-induced stability of tRNA from an extreme thermophile, Thermus thermophilus. Biochem. Biophys. Res. Commun. 72, 1137–1144. doi: 10.1016/S0006-291X(76)80250-1
Wei, F. Y., Suzuki, T., Watanabe, S., Kimura, S., Kaitsuka, T., Fujimura, A., et al. (2011). Deficit of tRNA(Lys) modification by Cdkal1 causes the development of type 2 diabetes in mice. J. Clin. Invest. 121, 3598–3608. doi: 10.1172/JCI58056
Wilkinson, M. L., Crary, S. M., Jackman, J. E., Grayhack, E. J., and Phizicky, E. M. (2007). The 2'-O-methyltransferase responsible for modification of yeast tRNA at position 4. RNA 13, 404–413. doi: 10.1261/rna.399607
Wurm, J. P., Griese, M., Bahr, U., Held, M., Heckel, A., Karas, M., et al. (2012). Identification of the enzyme responsible for N1-methylation of pseudouridine 54 in archaeal tRNAs. RNA 18, 412–420. doi: 10.1261/rna.028498.111
Yamagami, R., Yamashita, K., Nishimasu, H., Tomikawa, C., Ochi, A., Iwashita, C., et al. (2012). The tRNA recognition mechanism of folate/FAD-dependent tRNA methyltransferase (TrmFO). J. Biol. Chem. 287, 42480–42494. doi: 10.1074/jbc.M112.390112
Yaniv, M., and Folk, W. R. (1975). The nucleotide sequences of the two glutamine transfer ribonucleic acids from Escherichia coli. J. Biol. Chem. 250, 3243–3253.
Yasukawa, T., Suzuki, T., Ishii, N., Ohta, S., and Watanabe, K. (2001). Wobble modification defect in tRNA disturbs codon-anticodon interaction in a mitochondrial disease. EMBO J. 20, 4794–9802. doi: 10.1093/emboj/20.17.4794
Ye, K., Jia, R., Lin, J., Ju, M., Peng, J., Xu, A., et al. (2009). Structural organization of box C/D RNA-guided RNA methyltransferase. Proc. Natl. Acad. Sci. U.S.A. 106, 13803–13813. doi: 10.1073/pnas.0905128106
Yim, L., Moukadiri, I., Björk, G. R., and Armengod, M. E. (2006). Further insights into the tRNA modification process controlled by proteins MnmE and GidA of Escherichia coli. Nucleic Acids Res. 34, 5892–5905. doi: 10.1093/nar/gkl752
Yokoyama, S., Watanabe, K., and Miyazawa, T. (1987). Dynamic structures and functions of transfer ribonucleic acids from extreme thermophiles. Adv. Biophys. 23, 115–147. doi: 10.1016/0065-227X(87)90006-2
Zegers, I., Gigot, D., van Vliet, F., Tricot, C., Aymerich, S., Bujnicki, J. M., et al. (2006). Crystal structure of Bacillus subtilis TrmB, the tRNA (m7G46) methyltransferase. Nucleic Acids Res. 34, 1925–1934. doi: 10.1093/nar/gkl116
Keywords: RNA modification, RNA methylation, RNA maturation
Citation: Hori H (2014) Methylated nucleosides in tRNA and tRNA methyltransferases. Front. Genet. 5:144. doi: 10.3389/fgene.2014.00144
Received: 28 March 2014; Accepted: 04 May 2014;
Published online: 23 May 2014.
Edited by:
Akio Kanai, Keio University, JapanCopyright © 2014 Hori. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hiroyuki Hori, Department of Materials Science and Biotechnology, Graduate School of Science and Engineering, Ehime University, Bunkyo 3, Matsuyama, Ehime 790-8577, Japan e-mail:aG9yaUBlbmcuZWhpbWUtdS5hYy5qcA==
 Hiroyuki Hori
Hiroyuki Hori