- 1Animal Production and Health Division, Food and Agriculture Organization of the United Nations, Rome, Italy
- 2In Situ and On Farm Conservation, Bioversity International, Maccarese, Italy
- 3INCT Pecuaria, Universidade de Brasília, Brasilia, Brazil
- 4NordGen Farm Animals, Nordic Genetic Resource Center, Ås, Norway
- 5Parco Tecnologico Padano, Lodi, Italy
- 6Institute of Agricultural Biology and Biotechnology, National Research Council, Milan, Italy
- 7Plant Production and Protection Division, Food and Agriculture Organization of the United Nations, Rome, Italy
- 8NADEL – Center for Development and Cooperation, Swiss Federal Institute of Technology in Zürich, Zürich, Switzerland
- 9Sustainable Rural Systems, Scotland's Rural College, Edinburgh, UK
- 10UR143, Unité de Recherches Zootechniques, Institut National de la Recherche Agronomique, Petit-Bourg, Guadeloupe, France
- 11Basque Center for Climate Change, Bilbao, Spain
A commentary on
Global plan of action for animal genetic resources and the interlaken declaration
by FAO. (2007). Rome. ISBN: 978-92-5-105848-0
Introduction
Animal genetic resources (AnGR) are critical for global food security and livelihoods. Livestock products have high densities of energy, protein, and other critical nutrients, which are particularly beneficial for infants and expectant mothers. Around a billion people rely directly on livestock for their livelihoods, many of which are among the rural poor (FAO, 2009). Demand for animal products is foreseen to increase significantly in the future while competition for resources will intensify, dictating that livestock systems must increase both productivity and efficiency. Maintaining sufficient diversity of AnGR is necessary to ensure adaptation potential in times of uncertainty. In the future, climate change is expected to be a major force testing resilience of global food production systems (Thornton et al., 2009; Renaudeau et al., 2012). Ensuring that livestock systems remain productive and efficient while maintaining their flexibility will be a major challenge.
Adaptation to climate change is unlikely to be achieved with a single strategy (Hoffmann, 2010). Clearly, modifications will be needed in animals' housing, reproduction, nutrition, and health care. Genetic changes in the animals (both within and across species) will also play a role. Preparation for these transformations will require a significant research commitment and genomics will play a role in the genetic measures taken for adaptation of livestock to climate change.
Information Needs
The first step in this process will be gathering relevant information. There is currently a shortage of specific knowledge explaining why certain AnGR are adapted to a given environment and in which other environments they can survive and flourish. This deficit calls for greater effort in characterization of AnGR, including their production environments, by using the most modern tools available. To date, many livestock breeds have been genetically characterized (e.g., Groeneveld et al., 2010), but the value of these data for study of adaptation is questionable. For climate change adaptation, the breeds of greatest interest may be those reared today in harsh environments. Past studies have mostly addressed breeds from developed countries, where climate-control is widely practiced. The study of adaptation implies the use of a “landscape approach,” with detailed information describing the production system (e.g., FAO, 2012), including socio-economic information (e.g., Drucker, 2010) and indigenous knowledge about management of the breed in its environment as well as geographic coordinates to incorporate climatic data and soil, vegetation, and water resources. Collection of such detailed complementary data is a relatively recent trend. Past studies have emphasized pure breeds, whereas crossbreeding can be a valuable strategy for achieving increased productivity and adaptability, so those populations also require characterization. Finally, many studies have assayed only small numbers of selectively-neutral markers. The rapid development of genomic tools now allows analysis of functional genomic regions with potential associations with adaptation (e.g., Qian et al., 2013).
Selection Objectives and Strategies
A second research need is the elaboration of the genetic objective for adaptation. Increasing productivity and efficiency will be fundamental, but maintenance of genetic diversity will also be of importance. Having diverse AnGR will allow for more opportunities to match breeds to a changing climate or to replace populations hit by severe climatic events such as droughts and floods. Within breed, broad genetic diversity will clearly allow for greater opportunities for selection for adaptation, but there is evidence from wild populations that increased genetic diversity is also selectively advantageous on the individual level (Fourcada and Hoffman, 2014). Directional selection for adaptive traits will likely accompany maintenance of diversity, but questions remain about indicators of adaptation and resilience and hence the breeding goal. An obvious option is to breed for traits associated with superior productivity and resilience in conditions expected to be prevalent as a result of climate change, such as heat and drought tolerance and resistance to certain diseases. However, care must be taken when defining such traits. In dairy cattle, the ability to maintain high milk production with increasing ambient temperatures seems like a logical definition of heat tolerance, but research at the physiological level has shown that such cattle direct their energy toward milk production, making them vulnerable to extremely high temperatures (Dikmen et al., 2012). An alternative to breeding for specific traits is to target general robustness; the ability of animals to adjust to a range of environmental conditions. The Brown Swiss dairy cow, developed under comparatively cool, but rugged conditions in the Alps, seems to show greater heat tolerance than the Holstein (Correa-Calderon et al., 2005), which originated in more temperate lowland regions. The Domestic Animal Diversity Information system of FAO (http://dad.fao.org) lists numerous breeds, particularly from mountainous and arid areas, that are adapted to extreme ranges in temperature and such breeds may merit further research (Hoffmann, 2013). Table 1 has the numbers of local breeds by region and species for the major livestock species, as recorded in DAD-IS. Within-breed selection for robustness would likely require the development of an index involving multiple traits.
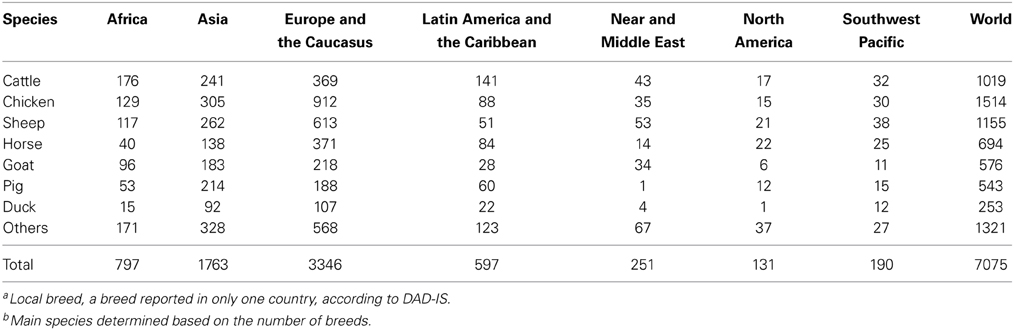
Table 1. Numbers of locala breeds recorded per species and region for the six main livestock speciesb for food and agriculture, according to the Domestic Animal Diversity Information System (DAD-IS) of FAO (http://dad.fao.org – November 2014).
A third category of research involves the genetic strategy for adaptation. Options comprise purebreeding, crossbreeding (including introgression), and breed or species substitution. Among the key influential factors is the expected rate of climate change and the speed with which genetic change can realistically occur with the various strategies. Substitution and crossbreeding expedite genetic change, but their implementation may be more complex than purebreeding and thus involve additional research needs (e.g., on genotype-by-environment interaction).
Tools for the Genetics of Adaptation
A final research topic will be the development of tools required for the aforementioned topics. Genomics will surely play a role in all three of these areas, as well as in implementation of the results obtained. Increased characterization with high-throughput single nucleotide polymorphism (SNP) assays or genome sequencing will be necessary for unraveling the physiological basis for adaptation. Species-wide HapMap studies (e.g., Gibbs et al., 2009; Jiang et al., 2014) and multi-species studies (e.g., Stella, 2014) have represented a valuable first step in understanding the genome and its function in adaptation, but must be expanded to more breeds and geographical areas and augmented with more information on production environments. Metagenomics can provide insight regarding the co-adaptation of AnGR with other organisms in their production environments. Genomic selection has the potential to expedite both pure- and crossbreeding programmes for adaptation, assuming phenotypes are available (Hayes et al., 2012); programs for performance recording in developing countries are thus needed. Given the importance of the landscape approach, tools and methods for improved integration of geographical information will be critical. The information gathered in all of these processes will be of little value if they are not properly organized, stored, and disseminated to stakeholders through new and improved databases and information systems. Finally, cooperative efforts between all stakeholders will be needed to achieve the final goal, the optimal utilization of the genetic and genomic resources for the adaptation of livestock to climate change.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This report comprises conclusions from the Expert Workshop on Crop and Livestock Diversity for Climate Change Adaptation held in Rome from 8 to 12 October 2013. The workshop was a collaborative effort among FAO, Bioversity International, the Basque Center for Climate Change and DIVERSITAS and was financially supported by the Government of Sweden through FAO project GCP/GLO/287/MUL and by the CGIAR program on Climate Change, Agriculture, and Food Security (CCAFS).
References
Correa-Calderon, A., Armstrong, D. V., Ray, D. E., DeNise, S. K., Enns, R. M., and Howison, C. M. (2005). Productive and reproductive response of Holstein and Brown Swiss heat stressed dairy cows to two different cooling systems. J. Anim. Vet. Adv. 4, 572–578.
Dikmen, S., Cole, J. B., Null, D. J., and Hansen, P. J. (2012). Heritability of rectal temperature and genetic correlations with production and reproduction traits in dairy cattle. J. Dairy Sci. 95, 3401–3405. doi: 10.3168/jds.2011-4306
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Drucker, A. G. (2010). Where's the beef? The economics of AnGR conservation and its influence on policy design and implementation. Anim. Genetic Resour. 47, 85–90. doi: 10.1017/S2078633610000913
FAO. (2009). The State of Food and Agriculture – Livestock in the Balance. Rome. Available online at: http://www.fao.org/docrep/012/i0680e/i0680e.pdf
FAO. (2012). Phenotypic Characterization of Animal Genetic Resources. FAO Animal Production and Health Guidelines No. 11. Rome.
Fourcada, J., and Hoffman, J. I. (2014). Climate change selects for heterozygosity in a declining fur seal population. Nature 511, 462–465. doi: 10.1038/nature13542
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Gibbs, R. A., Taylor, J. F., Van Tassell, C. P., Barendse, W., Eversole, K. A., Gill, C. A., et al. (2009). Genome-wide survey of SNP variation uncovers the genetic structure of cattle breeds. Science 324, 528–532. doi: 10.1126/science.1167936
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Groeneveld, L. F., Lenstra, J. A., Eding, H., Toro, M. A., Scherf, B., Pilling, D., et al. (2010). Genetic diversity in farm animals–a review. Anim. Genet. 41, 6–31. doi: 10.1111/j.1365-2052.2010.02038.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hayes, B. J., Lewin, H. A., and Goddard, M. E. (2012). The future of livestock breeding: genomic selection for efficiency, reduced emissions intensity, and adaptation. Trends Genet. 29, 206–214. doi: 10.1016/j.tig.2012.11.009
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hoffmann, I. (2010). Climate change and the characterization, breeding and conservation of animal genetic resources. Anim. Genet. 41, 32–46. doi: 10.1111/j.1365-2052.2010.02043.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hoffmann, I. (2013). Adaptation to climate change – exploring the potential of locally adapted breeds. Animal 7, 346–362. doi: 10.1017/S1751731113000815
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Jiang, Y., Xie, M., Chen, W., Talbot, R., Maddox, J. F., Faraut, T., et al. (2014). The sheep genome illuminates biology of the rumen and lipid metabolism. Science 344, 1168–1173. doi: 10.1126/science.1252806
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Qian, W., Deng, L., Lu, D., and Xu, S. (2013). Genome-wide landscapes of human local adaptation in Asia. PLoS ONE 8:e54224. doi: 10.1371/journal.pone.0054224
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Renaudeau, D., Collin, A., Yahav, S., de Basilio, V., Gourdine, J. L., and Collier, R. J. (2012). Adaptation to hot climate and strategies to alleviate heat stress in livestock production. Animal 6, 707–728. doi: 10.1017/S1751731111002448
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Stella, A. (2014). “Insights into the interaction of goat breeds and their environment,” in Proceedings of the 10th World Congress on Genetics Applied to Livestock Production (Vancouver, BC).
Keywords: animal genetic resources, adaptation, biological, genetic diversity, livestock genomics, climate change
Citation: Boettcher PJ, Hoffmann I, Baumung R, Drucker AG, McManus C, Berg P, Stella A, Nilsen LB, Moran D, Naves M and Thompson MC (2015) Genetic resources and genomics for adaptation of livestock to climate change. Front. Genet. 5:461. doi: 10.3389/fgene.2014.00461
Received: 12 November 2014; Paper pending published: 26 November 2014;
Accepted: 15 December 2014; Published online: 19 January 2015.
Edited by:
Juha Kantanen, MTT Agrifood Research Finland, FinlandReviewed by:
Dirk-Jan De Koning, Swedish University of Agricultural Sciences, SwedenJarmo Juga, University of Helsinki, Finland
Copyright © 2015 Boettcher, Hoffmann, Baumung, Drucker, McManus, Berg, Stella, Nilsen, Moran, Naves and Thompson. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence:cGF1bC5ib2V0dGNoZXJAZmFvLm9yZw==
 Paul J. Boettcher
Paul J. Boettcher Irene Hoffmann
Irene Hoffmann Roswitha Baumung1
Roswitha Baumung1 Concepta McManus
Concepta McManus Peer Berg
Peer Berg Alessandra Stella
Alessandra Stella Michel Naves
Michel Naves Mary C. Thompson
Mary C. Thompson