- 1Department of Chemistry, Biotechnology and Food Science, Faculty of Veterinary Medicine and Biosciences, Norwegian University of Life Sciences, Aas, Norway
- 2School of Life Sciences, Arizona State University, Tempe, AZ, USA
Epigenetic changes enable genomes to respond to changes in the environment, such as altered nutrition, activity, or social setting. Epigenetic modifications, thereby, provide a source of phenotypic plasticity in many species. The honey bee (Apis mellifera) uses nutritionally sensitive epigenetic control mechanisms in the development of the royal caste (queens) and the workers. The workers are functionally sterile females that can take on a range of distinct physiological and/or behavioral phenotypes in response to environmental changes. Honey bees have a wide repertoire of epigenetic mechanisms which, as in mammals, include cytosine methylation, hydroxymethylated cytosines, together with the enzymatic machinery responsible for these cytosine modifications. Current data suggests that honey bees provide an excellent system for studying the “social repertoire” of the epigenome. In this review, we elucidate what is known so far about the honey bee epigenome and its mechanisms. Our discussion includes what may distinguish honey bees from other model animals, how the epigenome can influence worker behavioral task separation, and how future studies can answer central questions about the role of the epigenome in social behavior.
Introduction
Since the first honey bee methylome was sequenced in 2010, our understanding of the functional implications of DNA methylation in the honey bee has begun to unfold (Lyko et al., 2010). 5-methylcytosine (5mC) is believed to be involved in alternative splicing, caste differentiation and worker behavioral task separation (Lyko et al., 2010; Flores et al., 2012; Herb et al., 2012). Recently, several other cytosine modifications were discovered in mammalian genomes (Kriaucionis and Heintz, 2009; He et al., 2011; Ito et al., 2011). These modifications are believed to have separate functions from 5mC as they are distributed differently in the genome, and specific reader proteins for one of these modifications exist (Spruijt et al., 2013). Although studies to investigate cytosine modifications other than 5mC in bees have been performed, little is known about their functions and distributions (Cingolani et al., 2013; Wojciechowski et al., 2014). Here we review cytosine modifications and the enzymatic machinery responsible for their generation in different model organisms.
Honey Bees
Nutritional cues lead female honey bee larvae into one of two developmental trajectories. The larvae either develop into a queen or into a worker (Winston, 1991). Queens are larger, highly fecund and long-lived (years), while the smaller workers are functionally sterile and shorter lived (weeks, months). Workers show a flexible physiological and behavioral progression that typically starts with care behavior toward siblings (nursing) and culminates in food collection (foraging) weeks later. Nursing is associated with enhanced somatic maintenance and slower aging than foraging (Münch and Amdam, 2010). Yet, foragers can return to nursing tasks, and this behavioral reversion can put age-associated cognitive decline in reverse as well (Baker et al., 2012).
Honey bees, in other words, display a wide range of phenotypes that include complex social caste development and behavior, behavioral shifts, and plasticity of aging. Epigenetic mechanisms are already found to likely play major roles in queen-worker development as well as in worker behavioral progression and reversion (Kucharski et al., 2008; Spannhoff et al., 2011; Herb et al., 2012). These findings put the honey bee forward as a very interesting study organism to investigate the interplay between the social milieu and the epigenome. The use of the honey bee for complex epigenetic research is, furthermore, not diminished by the mainstream models fruit fly (Drosophila melanogaster) and nematode (Caenorhabditis elegans), since they do not have the full complement of the mammalian epigenetic machinery (Table 1).
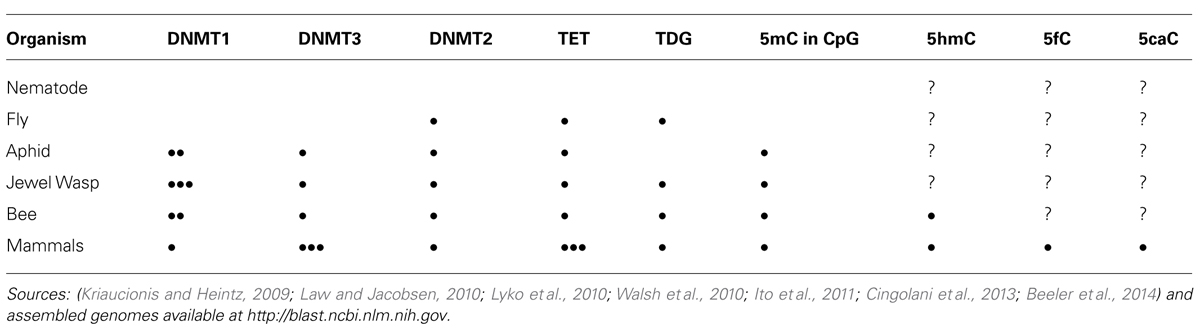

TABLE 1. Genomic copies of enzymes implicated in DNA methylation and demethylation and presence of epigenetically modified cytosines in select metazoan groups.
Epigenetic Machinery
DNA methyltransferases (DNMTs) are enzymes that add a methyl group to the 5′ carbon of the DNA base cytosine from the donor S-Adenosyl methionine (Law and Jacobsen, 2010). DNMT1 is the “maintenance” DNMT that copies the methylation pattern to the newly synthesized strand during DNA replication. DNMT3 is the de novo methyltransferase that can methylate specific loci independently of replication. DNMT2 is primarily an RNA methyltransferase that methylates t-RNAAsp (Goll et al., 2006), however, DNA activity has been shown in vivo in the fruit fly (Phalke et al., 2009). The de novo and the maintenance DNMTs are found in a range of species including honey bees, mammals, aphids, and jewel wasps (Table 1). They are catalytically active in the honey bee (Wang et al., 2006), while fruit fly and nematode only contain a single copy of DNMT2. Nevertheless, 5mC originating from DNA has been reported in the fruit fly in both embryos and adult flies (Lyko et al., 2000), suggesting that DNMT2 has some DNA methylation activity in vivo. The impact of 5mC in the fruit fly genome is still debated, however (Phalke et al., 2010; Schaefer and Lyko, 2010).
In mammals, the ten eleven translocation (TET) enzyme is responsible for further oxidizing 5mC to 5-hydroxymethylcytosine (5hmC) that again can be oxidized to 5-formylcytosine (5fC), and ultimately 5-carboxylcytosine (5caC) (Tahiliani et al., 2009; He et al., 2011; Ito et al., 2011). 5fC and 5caC are recognized by the thymine DNA glycosylase (TGD), which is a part of the base excision repair pathway of the mammalian cell (Maiti and Drohat, 2011). The TET enzyme together with TDG are probably central to the mammalian active demethylation pathway (Pastor et al., 2013). Mammalian genomes harbor multiple TET enzyme genes, while bees, fruit flies, aphids, and jewel wasps only have one (Table 1). The RNA expression levels of the different mammalian TET enzymes vary greatly between developmental stages and cell types. The honey bee TET catalytic domain is catalytically active in vitro, and active transcription of the honey bee TET gene has been shown to vary in different stages of development as well as in different adult tissues (Wojciechowski et al., 2014). Interestingly, some species (including fruit fly) that contain only DNMT2 have well conserved TET orthologs, but their activity and function have not been deciphered (Dunwell et al., 2013).
The honey bee genome encodes several core histone modifying enzymes, which are also part of the epigenetic machinery of the honey bees (The Honeybee Genome Sequencing, 2006). However, the impact of and the mechanisms behind histone modifications are beyond the scope of this review.
5-Methylcytosine
The distribution and relative abundance of 5mC vary significantly between mammals, honey bee and fruit fly (Figure 1). 5mC is primarily located in a CpG dinucleotide context within repeat sequences and in proximity of promoter areas in mammals (Law and Jacobsen, 2010), whereas in bees methylated CpGs are primarily located within genes (Lyko et al., 2010). However, 5mC can exist in a non-CpG dinucleotide context in both mammals and honey bees (Lister et al., 2009; Cingolani et al., 2013). In addition, the honey bee genome is much more sparsely methylated than mammalian genomes, thus reducing overall complexity and simplifying data analyses for studies conducted in bees. In the fruit fly genome, 5-mC is located within a non-CpG dinucleotide context and seems to be distributed randomly within the genome at an abundance 3- to 100-fold less when compared to honey bees and mammals (Mandrioli and Borsatti, 2006; Phalke et al., 2009). C. elegans, on the other hand, does not contain 5mC in its genome (Simpson et al., 1986).
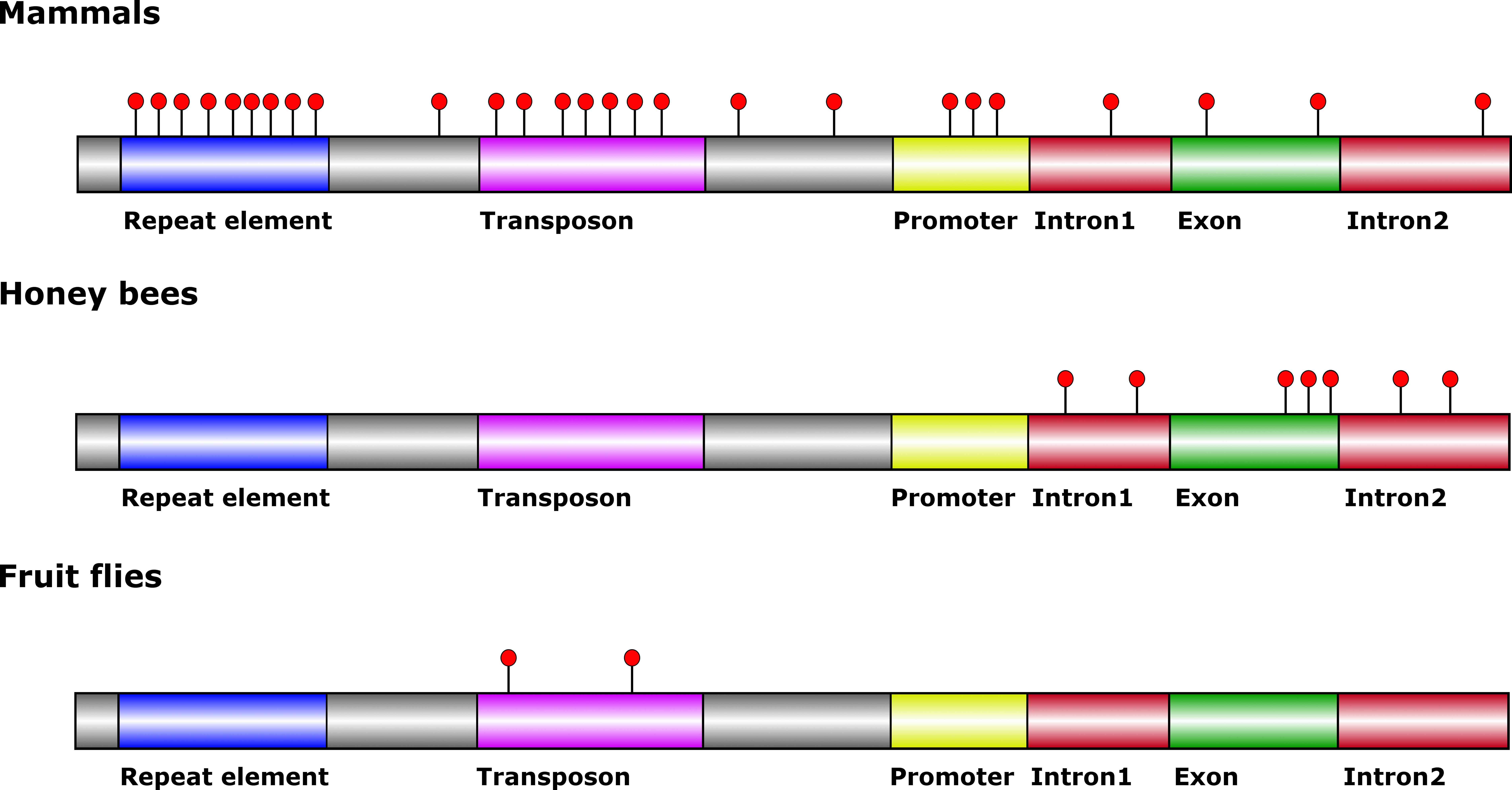
FIGURE 1. General features of the 5-methylcytosine distribution in DNA from mammals, honey bees and fruit flies. Red circles indicate 5-mC. Mammalian genomes are typically methylated in transposon and repeat elements, and at some promoter regions. Intergenic DNA methylation occurs albeit at lower levels. Honey bee genomes are typically methylated in exons close to the exon-intron borders. Non-CpG methylation occurs in introns. Methylation outside of transposons has not been mapped in fruit fly genomes.
The effect of 5mC on transcription varies between metazoans and genomic context. In mammalian promoters, 5mC is principally a repressive mark, silencing transcription (Bird, 2002). On the other hand, 5mC within gene bodies in mammals, honey bees, and the fruit fly, does not influence transcription levels to the same extent (Mandrioli and Borsatti, 2006; Flores et al., 2012). In honey bees, 5mC within gene bodies rather plays a role in the generation of alternative splice variants on the genome-wide level (Flores et al., 2012; Foret et al., 2012; Li-Byarlay et al., 2013). This role is not clearly defined in mammalian cells, as the role of 5mC in gene bodies differs between cell types and depends on whether 5mC is in a CpG context or not (Lister et al., 2009). These findings make honey bees an attractive system for studies on how 5mC influences the generation of alternative transcripts.
5-methylcytosine is found in multiple cell types, tissues, and life stages in both honey bees and mammals (Ikeda et al., 2011; Ziller et al., 2013). In D. melanogaster, 5mC is mostly found during early embryonic stages (Lyko et al., 2000). Although adult 5mC has been reported in fruit fly, the content is too low to be robustly detected by bisulfite sequencing, the gold standard in base resolution 5mC interrogation techniques, making further studies difficult with many established methods depending on bisulfite conversion (Capuano et al., 2014).
5-Hydroxymethylcytosine
The TET oxidative products of 5mC recently became a center of attention in mammalian epigenetic research. Many questions about TET and 5hmC dynamics have been answered in embryonic stem cells (Pastor et al., 2013), although 5hmC has been detected in different tissues at different life stages (Kriaucionis and Heintz, 2009; Ivanov et al., 2013). The abundance of 5hmC compared to 5mC is much lower ranging from 2- to 100-fold times less depending on tissue (Kriaucionis and Heintz, 2009; Song et al., 2012). The distribution of 5hmC does not seem to be directly linked to 5mC, as 5hmC is found more often in promoter areas and enhancers, and much less in repetitive elements (Pastor et al., 2011; Stroud et al., 2011; Yu et al., 2012). In addition, proteins capable of specifically binding 5hmC have been discovered, fueling the theory that 5hmC exists as separate epigenetic mark and not simply just as an intermediate in an active demethylation pathway (Frauer et al., 2011; Méllen et al., 2012; Spruijt et al., 2013). In honey bees, 5hmC has been characterized in multiple tissues, and its abundance seems to be highest in germ cells and the brain (7–10% of 5mC and about 4% of 5mC, respectively), following the trend in mammalian cell types (Wojciechowski et al., 2014). Only one study has attempted to map 5hmC in honey bees at a single nucleotide resolution (Cingolani et al., 2013). This same study, surprisingly, mapped the majority of 5hmC in head tissue to non-CpG intronic sequences. Further studies seems warranted to precisely quantify and map 5hmC in bees, especially in non-brain tissue, which has received less interest so far. To date, 5hmC together with 5fC and 5caC have not been identified in the fruit fly, aphid, jewel wasp, and C. elegans genomes. However, since C. elegans has no 5mC precursor or TET homolog, the existence of 5hmC, 5fC, and 5caC seems highly unlikely.
5-Formylcytosine and 5-Carboxylcytosine
The recently identified nucleotides 5fC and 5caC have, so far, not accumulated the same level of information as their precursors 5mC and 5hmC. This situation is in part due to extremely low abundance, especially for 5caC, making robust detection difficult (in mammals 5caC is 10- to 1000-fold less abundant than 5hmC). Moreover, the molecular toolbox for investigating 5fC and 5caC is not as developed as it is for 5hmC (Song and He, 2013). Bisulfite sequencing for example, only discriminates between “methylated” and “unmethylated” cytosines, so that 5mC and 5hmC are identified as “methylated” and 5fC and 5caC as “unmethylated” (Pastor et al., 2013). Such data are therefore difficult to use as guidelines in narrowing down possible locations of 5fC and 5caC.
The extremely low abundance of 5caC suggest that this nucleotide is merely an intermediate step in complete demethylation (Song and He, 2013). Although 5fC is a more prominent epigenetic mark than 5caC, its function is still not fully understood. It is possible that 5fC might regulate transcription through stalling of RNA pol II (Kellinger et al., 2012), but further research is needed to elucidate the role of 5fC and 5caC in both vertebrates and invertebrates. In honey bees, 5fC and 5caC have not been investigated yet, though their precursors and catalytic enzyme have been reported (Lyko et al., 2010; Wojciechowski et al., 2014).
Future Work: Epigenetics and Worker Behavior
Epigenetic mechanisms have been linked to the queen-worker differentiation of honey bees (Kucharski et al., 2008), as well as to worker behavioral progression and reversion (Herb et al., 2012). Herb et al. (2012) bisulfite sequenced brains of age-matched nurses, foragers, and reverted workers (previous foragers now involved in care behavior). Their data revealed differentially methylated regions (DMRs) between the behavioral groups indicating that DNA methylation can play a role in regulation of social behavior. These DMRs are associated with genes involved in development, nuclear pore formation, and ATP binding. RNA sequencing revealed that these same DMRs were connected to alternative splicing events. It is also very likely that the “behaviorally sensitive” DMRs of honey bees are hydroxymethylated at some point during either transition from nurse to forager, or reversion from forager to nurse. Since the study was conducted in adult brain tissue, which has no neurogenesis (Fahrbach et al., 1995), dilution by replication would be unlikely or would only display minor effects. This situation makes these DMRs excellent candidates for investigating if 5hmC is associated with worker behavioral transitions, and if these hydroxymethylated regions are differentially hydroxymethylated between nurses, foragers, and reverted worker bees. Such a study could be the first to establish a putative link between hydroxymethylation and behavior.
Future studies should also dissect the role of TET in worker transitions from nurse to foragers, and back. Other candidate tissues than brain should include the fat body. This tissue is functionally homologous to liver and white adipose tissue and undergoes major remodeling during honey bee behavioral change (Chan et al., 2011). Functional implications of an RNA interference-mediated TET knockdown should provide insight into TET function. Studies can be conducted in honey bee larvae to investigate if TET knockdowns are capable of both queen and worker development. Similarly, consequences for behavioral plasticity can be studied in adult honey bee workers and perhaps link TET and its products with behavior for the first time.
Finally, a possible link between 5hmC and alternative splicing can be investigated by combining 5hmC sequencing at single nucleotide resolution with RNA sequencing of honey bee tissue samples. 5mC is reportedly implicated in the generation of alternative transcripts in the bee, but using methods not able to distinguish 5mC from 5hmC (Flores et al., 2012; Herb et al., 2012). Therefore, further studies that can map 5hmC alongside RNA sequencing data seems warranted, and could potentially give 5hmC a novel function in gene regulation.
Conclusion
The honey bee offers a system where the interplay between DNA methylation and social behavior can be studied in great detail. Published studies of the honey bee epigenome are dominated by questions surrounding queen and worker development, while the epigenetic dynamics of worker behavioral castes have only more recently gained attention. The readily identifiable social behaviors of worker honey bees make setting up precise, large scale experiments feasible (Münch et al., 2013). Better knowledge about honey bee epigenetics also has a dual purpose; increasing the understanding of epigenetic machineries in general, and gaining specific information about gene regulatory mechanisms in an economically important beneficial insect.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank the two reviewers for their helpful comments. This study was supported by funds from the Research Council of Norway, grant no. 191699.
References
Baker, N., Wolschin, F., and Amdam, G. V. (2012). Age-related learning deficits can be reversible in honeybees Apis mellifera. Exp. Gerontol. 47, 764–772. doi: 10.1016/j.exger.2012.05.011
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Beeler, S. M., Wong, G. T., Zheng, J. M., Bush, E. C., Remnant, E. J., Oldroyd, B. P.,et al. (2014). Whole-genome DNA methylation profile of the jewel wasp (Nasonia vitripennis). G3 (Bethesda) 4, 383–388. doi: 10.1534/g3.113.008953
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bird, A. (2002). DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21. doi: 10.1101/gad.947102
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Capuano, F., Mulleder, M., Kok, R., Blom, H. J., and Ralser, M. (2014). Cytosine DNA methylation is found in Drosophila melanogaster but absent in Saccharomyces cerevisiae, Schizosaccharomyces pombe, and other yeast species. Anal. Chem. 86, 3697–3702. doi: 10.1021/ac500447w
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chan, Q. W., Mutti, N. S., Foster, L. J., Kocher, S. D., Amdam, G. V., and Wolschin, F. (2011). The worker honeybee fat body proteome is extensively remodeled preceding a major life-history transition. PLoS ONE 6:e24794. doi: 10.1371/journal.pone.0024794
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cingolani, P., Cao, X., Khetani, R. S., Chen, C. C., Coon, M., Sammak, A. A.,et al. (2013). Intronic Non-CG DNA hydroxymethylation and alternative mRNA splicing in honey bees. BMC Genomics 14:666. doi: 10.1186/1471-2164-14-666
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Dunwell, T. L., Mcguffin, L. J., Dunwell, J. M., and Pfeifer, G. P. (2013). The mysterious presence of a 5-methylcytosine oxidase in the Drosophila genome: possible explanations. Cell Cycle 12, 3357–3365. doi: 10.4161/cc.26540
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fahrbach, S. E., Strande, J. L., and Robinson, G. E. (1995). Neurogenesis is absent in the brains of adult honey bees and does not explain behavioral neuroplasticity. Neurosci. Lett. 197, 145–148. doi: 10.1016/0304-3940(95)11913-H
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Flores, K. B., Wolschin, F., Allen, A. N., Corneveaux, J. J., Huentelman, M., and Amdam, G. V. (2012). Genome-wide association between DNA methylation and alternative splicing in an invertebrate. BMC Genomics 13:480. doi: 10.1186/1471-2164-13-480
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Foret, S., Kucharski, R., Pellegrini, M., Feng, S., Jacobsen, S. E., Robinson, G. E.,et al. (2012). DNA methylation dynamics, metabolic fluxes, gene splicing, and alternative phenotypes in honey bees. Proc. Natl. Acad. Sci. U.S.A. 109, 4968–4973. doi: 10.1073/pnas.1202392109
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Frauer, C., Hoffmann, T., Bultmann, S., Casa, V., Cardoso, M. C., Antes, I.,et al. (2011). Recognition of 5-hydroxymethylcytosine by the Uhrf1 SRA domain. PLoS ONE 6:e21306. doi: 10.1371/journal.pone.0021306
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Goll, M. G., Kirpekar, F., Maggert, K. A., Yoder, J. A., Hsieh, C.-L., Zhang, X.,et al. (2006). Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398. doi: 10.1126/science.1120976
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
He, Y. F., Li, B. Z., Li, Z., Liu, P., Wang, Y., Tang, Q.,et al. (2011). Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307. doi: 10.1126/science.1210944
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Herb, B. R., Wolschin, F., Hansen, K. D., Aryee, M. J., Langmead, B., Irizarry, R.,et al. (2012). Reversible switching between epigenetic states in honeybee behavioral subcastes. Nat. Neurosci. 15, 1371–1373. doi: 10.1038/nn.3218
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ikeda, T., Furukawa, S., Nakamura, J., Sasaki, M., and Sasaki, T. (2011). CpG methylation in the hexamerin 110 gene in the European honeybee, Apis mellifera. J. Insect. Sci. 11:74. doi: 10.1673/031.011.7401
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ito, S., Shen, L., Dai, Q., Wu, S. C., Collins, L. B., Swenberg, J. A.,et al. (2011). Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303. doi: 10.1126/science.1210597
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ivanov, M., Kals, M., Kacevska, M., Barragan, I., Kasuga, K., Rane, A.,et al. (2013). Ontogeny, distribution and potential roles of 5-hydroxymethylcytosine in human liver function. Genome Biol. 14:R83. doi: 10.1186/gb-2013-14-8-r83
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kellinger, M. W., Song, C. X., Chong, J., Lu, X. Y., He, C., and Wang, D. (2012). 5-formylcytosine and 5-carboxylcytosine reduce the rate and substrate specificity of RNA polymerase II transcription. Nat. Struct. Mol. Biol. 19, 831–833. doi: 10.1038/nsmb.2346
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kriaucionis, S., and Heintz, N. (2009). The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930. doi: 10.1126/science.1169786
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kucharski, R., Maleszka, J., Foret, S., and Maleszka, R. (2008). Nutritional control of reproductive status in honeybees via DNA methylation. Science 319, 1827–1830. doi: 10.1126/science.1153069
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Law, J. A., and Jacobsen, S. E. (2010). Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220. doi: 10.1038/nrg2719
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Li-Byarlay, H., Li, Y., Stroud, H., Feng, S., Newman, T. C., Kaneda, M.,et al. (2013). RNA interference knockdown of DNA methyl-transferase 3 affects gene alternative splicing in the honey bee. Proc. Natl. Acad. Sci. U.S.A. 110, 12750–12755. doi: 10.1073/pnas.1310735110
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lister, R., Pelizzola, M., Dowen, R. H., Hawkins, R. D., Hon, G., Tonti-Filippini, J.,et al. (2009). Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322. doi: 10.1038/nature08514
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lyko, F., Foret, S., Kucharski, R., Wolf, S., Falckenhayn, C., and Maleszka, R. (2010). The honey bee epigenomes: differential methylation of brain DNA in queens and workers. PLoS Biol. 8:e1000506. doi: 10.1371/journal.pbio.1000506
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lyko, F., Ramsahoye, B. H., and Jaenisch, R. (2000). DNA methylation in Drosophila melanogaster. Nature 408, 538–540. doi: 10.1038/35046205
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Maiti, A., and Drohat, A. C. (2011). Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–35338. doi: 10.1074/jbc.C111.284620
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mandrioli, M., and Borsatti, F. (2006). DNA methylation of fly genes and transposons. Cell. Mol. Life Sci. 63, 1933–1936. doi: 10.1007/s00018-006-6039-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Méllen, M., Ayata, P., Dewell, S., Kriaucionis, S., and Heintz, N. (2012). MeCP2 binds to 5hmC enriched within active genes and accessible chromatin in the nervous system. Cell 151, 1417–1430. doi: 10.1016/j.cell.2012.11.022
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Münch, D., and Amdam, G. V. (2010). The curious case of aging plasticity in honey bees. FEBS Lett. 584, 2496–2503. doi: 10.1016/j.febslet.2010.04.007
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Münch, D., Baker, N., Rasmussen, E. M. K., Shah, A. K., Kreibich, C. D., Heidem, L. E.,et al. (2013). Obtaining specimens with slowed, accelerated and reversed aging in the honey bee model. J. Vis. Exp. 8, e50550. doi: 10.3791/50550
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Pastor, W. A., Aravind, L., and Rao, A. (2013). TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat. Rev. Mol. Cell Biol. 14, 341–356. doi: 10.1038/nrm3589
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Pastor, W. A., Pape, U. J., Huang, Y., Henderson, H. R., Lister, R., Ko, M.,et al. (2011). Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 473, 394–397. doi: 10.1038/nature10102
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Phalke, S., Nickel, O., and Reuter, G. (2010). Reply to “Lack of evidence for DNA methylation of Invader4 retroelements in Drosophila and implications for Dnmt2-mediated epigenetic regulation.” Nat. Genet. 42, 921–921. doi: 10.1038/ng1110-921
Phalke, S., Nickel, O., Walluscheck, D., Hortig, F., Onorati, M. C., and Reuter, G. (2009). Retrotransposon silencing and telomere integrity in somatic cells of Drosophila depends on the cytosine-5 methyltransferase DNMT2. Nat. Genet. 41, 696–702. doi: 10.1038/ng.360
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Schaefer, M., and Lyko, F. (2010). Lack of evidence for DNA methylation of Invader4 retroelements in Drosophila and implications for Dnmt2-mediated epigenetic regulation. Nat. Genet. 42, 920–921; author reply 921. doi: 10.1038/ng1110-920
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Simpson, V. J., Johnson, T. E., and Hammen, R. F. (1986). Caenorhabditis elegans DNA does not contain 5-methylcytosine at any time during development or aging. Nucleic Acids Res. 14, 6711–6719. doi: 10.1093/nar/14.16.6711
Song, C. X., and He, C. (2013). Potential functional roles of DNA demethylation intermediates. Trends Biochem. Sci. 38, 480–484. doi: 10.1016/j.tibs.2013.07.003
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Song, C. X., Yi, C., and He, C. (2012). Mapping recently identified nucleotide variants in the genome and transcriptome. Nat. Biotechnol. 30, 1107–1116. doi: 10.1038/nbt.2398
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Spannhoff, A., Kim, Y. K., Raynal, N. J., Gharibyan, V., Su, M. B., Zhou, Y. Y.,et al. (2011). Histone deacetylase inhibitor activity in royal jelly might facilitate caste switching in bees. EMBO Rep. 12, 238–243. doi: 10.1038/embor.2011.9
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Spruijt, C. G., Gnerlich, F., Smits, A. H., Pfaffeneder, T., Jansen, P. W., Bauer, C.,et al. (2013). Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152, 1146–1159. doi: 10.1016/j.cell.2013.02.004
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Stroud, H., Feng, S., Morey Kinney, S., Pradhan, S., and Jacobsen, S. E. (2011). 5-hydroxymethylcytosine is associated with enhancers and gene bodies in human embryonic stem cells. Genome Biol. 12:R54. doi: 10.1186/gb-2011-12-6-r54
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Tahiliani, M., Koh, K. P., Shen, Y., Pastor, W. A., Bandukwala, H., Brudno, Y.,et al. (2009). Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935. doi: 10.1126/science.1170116
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
The Honeybee Genome Sequencing, C. (2006). Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949. doi: 10.1038/nature05260
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Walsh, T. K., Brisson, J. A., Robertson, H. M., Gordon, K., Jaubert-Possamai, S., Tagu, D.,et al. (2010). A functional DNA methylation system in the pea aphid, Acyrthosiphon pisum. Insect Mol. Biol. 19(Suppl. 2), 215–228. doi: 10.1111/j.1365-2583.2009.00974.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Wang, Y., Jorda, M., Jones, P. L., Maleszka, R., Ling, X., Robertson, H. M.,et al. (2006). Functional CpG methylation system in a social insect. Science 314, 645–647. doi: 10.1126/science.1135213
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Wojciechowski, M., Rafalski, D., Kucharski, R., Misztal, K., Maleszka, J., Bochtler, M.,et al. (2014). Insights into DNA hydroxymethylation in the honeybee from in-depth analyses of TET dioxygenase. Open Biol. 4:140110. doi: 10.1098/rsob.140110
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yu, M., Hon, G. C., Szulwach, K. E., Song, C. X., Zhang, L., Kim, A.,et al. (2012). Base-resolution analysis of 5-hydroxymethylcytosine in the mammalian genome. Cell 149, 1368–1380. doi: 10.1016/j.cell.2012.04.027
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ziller, M. J., Gu, H., Müller, F., Donaghey, J., Tsai, L. T., Kohlbacher, O.,et al. (2013). Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481. doi: 10.1038/nature12433
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: honey bee, methylation, demethylation, 5-hydroxymethylcytosine, social behavior
Citation: Rasmussen EMK and Amdam GV (2015) Cytosine modifications in the honey bee (Apis mellifera) worker genome. Front. Genet. 6:8. doi: 10.3389/fgene.2015.
Received: 29 September 2014; Accepted: 09 January 2015;
Published online: 06 February 2015.
Edited by:
Greg J. Hunt, Purdue University, USACopyright © 2015 Rasmussen and Amdam. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Erik M. K. Rasmussen, Department of Chemistry, Biotechnology and Food Science, Faculty of Veterinary Medicine and Biosciences, Norwegian University of Life Sciences, Christian Magnus Falsens vei 1, N-1432 Aas, Norway e-mail:ZXJpay5yYXNtdXNzZW5Abm1idS5ubw==
 Erik M. K. Rasmussen
Erik M. K. Rasmussen Gro V. Amdam1,2
Gro V. Amdam1,2