- 1Ecological Genetics Laboratory, Department of Biological Sciences, Middle East Technical University, Ankara, Turkey
- 2Bee Research Laboratory, United States Department of Agriculture-Agricultural Research Service, Beltsville, MD, USA
Honey bees face numerous biotic threats from viruses to bacteria, fungi, protists, and mites. Here we describe a thorough analysis of microbes harbored by worker honey bees collected from field colonies in geographically distinct regions of Turkey. Turkey is one of the World's most important centers of apiculture, harboring five subspecies of Apis mellifera L., approximately 20% of the honey bee subspecies in the world. We use deep ILLUMINA-based RNA sequencing to capture RNA species for the honey bee and a sampling of all non-endogenous species carried by bees. After trimming and mapping these reads to the honey bee genome, approximately 10% of the sequences (9–10 million reads per library) remained. These were then mapped to a curated set of public sequences containing ca. Sixty megabase-pairs of sequence representing known microbial species associated with honey bees. Levels of key honey bee pathogens were confirmed using quantitative PCR screens. We contrast microbial matches across different sites in Turkey, showing new country recordings of Lake Sinai virus, two Spiroplasma bacterium species, symbionts Candidatus Schmidhempelia bombi, Frischella perrara, Snodgrassella alvi, Gilliamella apicola, Lactobacillus spp.), neogregarines, and a trypanosome species. By using metagenomic analysis, this study also reveals deep molecular evidence for the presence of bacterial pathogens (Melissococcus plutonius, Paenibacillus larvae), Varroa destructor-1 virus, Sacbrood virus, and fungi. Despite this effort we did not detect KBV, SBPV, Tobacco ringspot virus, VdMLV (Varroa Macula like virus), Acarapis spp., Tropilaeleps spp. and Apocephalus (phorid fly). We discuss possible impacts of management practices and honey bee subspecies on microbial retinues. The described workflow and curated microbial database will be generally useful for microbial surveys of healthy and declining honey bees.
Introduction
The honey bee (Apis mellifera L.) has ecological importance as a natural pollinator of wild flora and crops. Moreover, managed honey bees have economical importance with hive products including honey, pollen, wax, propolis, and royal jelly (Maheshwari, 2003). Recently, declines of managed colonies have been noted on many continents. Several causes of these large-scale losses have been reported, including honey bee parasites (Varroa destructor, Acarapis woodi); pathogens (Nosema spp. and bee viruses); pesticides, contaminated water, use of antibiotics, poor nutrition, and migratory beekeeping practices (Kevan et al., 2007; Higes et al., 2008; Naug, 2009; vanEngelsdorp et al., 2009; Bacandritsos et al., 2010; vanEngelsdorp and Meixner, 2010). The high density of individuals and the exchange of food among A. mellifera colony members create a favorable environment for bacterial, viral, fungal, and protist pathogens and several studies have noted an increase in diversity and infection rates of pathogens in failing bee colonies. Here we will focus on honey bee pathogens and parasites and the use of modern sequencing techniques to identify these agents in healthy and declining colonies.
Among the honey bee pathogens, viruses are of special concern. Viruses are widespread in honey bees although most often without noticeable symptoms (Ball and Bailey, 1997). Multiple viral infections have been diagnosed in many bee colonies (Chen et al., 2004). At least 18 different viruses exist in honey bees (Bailey and Ball, 1991) with six of them; Sacbrood virus (SBV), Deformed wing virus (DWV), Acute bee paralysis virus (ABPV), Black queen cell virus (BQCV), Chronic bee paralysis virus (CBPV), and Kashmir bee virus (KBV) most commonly linked to bee disease. These viruses have strong impacts on managed bee populations, pollination services, and honey production on several continents (Allen and Ball, 1996; Nordstrom et al., 1999; Ellis and Munn, 2005; De Miranda et al., 2010). DWV and ABPV have been linked to parasitic mite loads, while Chronic bee paralysis virus (CBPV) and the related Lake Sinai viruses are also widespread and tied to significant losses in honey bee colonies (Runckel et al., 2011; Ravoet et al., 2013). Sacbrood virus is the only common virus of developing bees, and this virus is not generally implicated in adult bee mortality or morbidity (Anderson and Gibbs, 1989).
Two species of Microsporidia, Nosema apis and Nosema ceranae, are widespread parasites of adult honey bees. N. apis is a long-standing infection agent of the European honey bee, A. mellifera, (Zander, 1909) that causes nosemosis, a disease with mild virulence. N. ceranae was first found as a parasite of the Asian honey bee, Apis cerana (Fries et al., 1996) although this species is now widespread throughout the range of A. mellifera (Fries et al., 1996, 2006; Paxton et al., 2007) arguably thanks to worldwide trade in bees (Klee et al., 2007) and perhaps pollen supplements. Honey bees can be co-infected with both Nosema species (Fries, 2010). Additionally, N. ceranae seems to be replacing N. apis worldwide (Klee et al., 2007). In association with Nosema infections, several viruses can significantly affect the apparent virulence of Nosema (Bailey and Ball, 1991).
Trypanosomes are a varied group of parasites that infect insects (Merzlyak et al., 2001). Crithidia bombi, a trypanosome that infects bumble bees, has effects on behavior (Gegear et al., 2005) and longevity during stressful conditions (Brown et al., 2003). Trypanosomatid parasites parasitize the mid- and hind-guts of their hosts (Lange and Lord, 2012) and can be widespread (Langridge and McGhee, 1967; Schmid-Hempel and Tognazzo, 2010). The current role of trypanosomes in honey bee health is not clear (Schwarz and Evans, 2013), although they have been recognized as possible correlates with bee declines in two field surveys (Runckel et al., 2011; Ravoet et al., 2013). The most common honey bee trypanosomatid currently is distinct from the species Crithidia mellificae described by Langridge and McGhee (1967), and has recently been named as Lotmaria passim (Schwarz et al., 2015).
Spiroplasmas are particularly virulent pathogens that are found in various environments and implicated as pathogens of plants, vertebrates and insects. They have a seasonal occurrence associated with the nectarines and surfaces of flowers (Markham and Townsend, 1981; Williamson et al., 1989). Adult honey bees are parasitized by two species of bacteria, Spiroplasma apis (Mouches et al., 1983) and Spiroplasma melliferum (Clark et al., 1985). Upon invading the hemolymph, these bacteria can cause a fatal disease called spiroplasmosis or May disease. Gut microbiota of animals living in social communities may influence their health with their functions related to nutrition, immune responses and resistance against pathogens (Dillon and Dillon, 2004; Round and Mazmanian, 2009). Surveys of 16S rDNA sequences from the honey bee indicate the presence of eight predominant species (defined as strains sharing >97% 16S rRNA identity) which account for 95% of the resident bacteria (Moran et al., 2012). These species include the beta-proteobacterium Snodgrassella alvi (family Neisseriaceae) and the gamma-proteobacterium Gilliamella apicola (family Orbaceae), the dominant Gram-negative members of the gut community, with each comprising up to 30–39% of the microbiota (Moran et al., 2012).
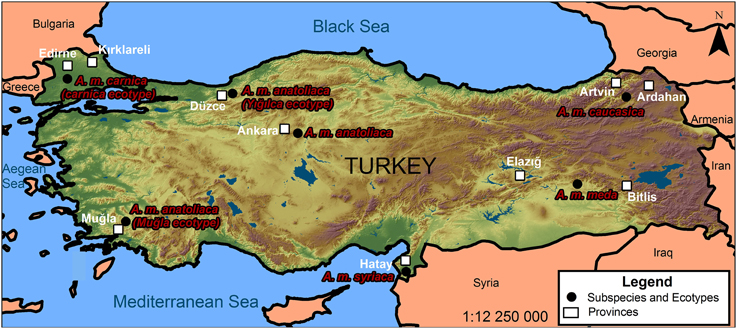
The surveillance and discovery of novel pathogens via high-throughput sequencing can provide a relatively unbiased view of pathogens and microbes associated with insects and other arthropods (Bishop-Lilly et al., 2010; Ma et al., 2011; Vayssier-Taussat et al., 2013). Recent efforts based on these technologies have uncovered novel and unexpected taxa associated with honey bees (e.g., Cox-Foster et al., 2007; Runckel et al., 2011; Cornman et al., 2013) and have provided estimates of normal microbial levels vs. those of diseased colonies. Turkey is one of the World's most important centers of apiculture, with managed and wild populations of five subspecies of Apis mellifera L. including A. m. caucasica, A. m. syriaca, A. m. anatoliaca, A. m. meda and an ecotype in Thrace belonging to the carnica subspecies group which is distinctly different from the subspecies found in Anatolia (Figure 1) (Kandemir et al., 2000; Bodur et al., 2007). These subspecies cover approximately 20% of honey bee subspecies in the world. Our purpose here was to determine the regional prevalence of bacterial pathogens, viruses, fungi, parasites, protists, and symbionts and to compare their loads in areas where migratory and stationary beekeeping is practiced in Turkey. Monitoring viruses and other disease agents can help solve problems related to the health of stationary and migratory honey bee colonies and limit or avoid their spread. Although commercial migratory beekeeping practices are necessary for pollination and crop production, their effects on honey bee colony health and pathogen transmission should be addressed. We used transcriptome analyses to survey a wide range of pathogens and to detect unexpected or rare taxa. RNA-seq technology allows the precise detection of rare transcripts by mapping reads against inclusive sequence databases, reducing the repetitive effort and possible biases of conventional molecular diagnostics (Minoche et al., 2011). Identification and confirmation of select virus and pathogen loads was confirmed by quantitative RT-PCR. Our survey was aligned with the European Honey bee colony loss network (www.COLOSS.org) survey, and was conducted to determine the colony losses in many regions representing honey bee diversity in Turkey. By using metagenomic analysis, our results provide new incidence records of the virome and microbiome in search of etiologically unexpected or previously unknown agents among 10 distinct provinces in Turkey and suggest a higher viral prevalence, and increased losses, in migratory beekeeping operations. Apis mellifera is an economically and ecologically important model organism and identification of pathogens and other microbes can have extensive implications for current practices in apiculture and agriculture.
Materials and Methods
Sampling
Colony loss surveys were carried out in eight regions and 158 beekeeping operations in 2010. Among those, 98 were migratory beekeepers and 60 were stationary. In 2011, surveys of 221 beekeepers from seven different regions were evaluated for this study. Samples analyzed for microbial loads comprised adult honey bees collected from field colonies in Turkey during summer and fall of 2010 and 2011. Efforts were made to collect from different regions of Turkey, and from a diverse set of beekeepers (Figure 1). Adult bees were collected from 134 colonies from different regions of Turkey from 38, 48, and 51 beekeepers in 2010-Fall, 2010-Spring, and 2011 respectfully and the samples were kept frozen until molecular diagnosis.
RNA Isolation
RNA was extracted from a pooled sample of 50 bees from each sampled colony, using an acid-phenol RNA extraction method (Evans et al., 2013).
cDNA Synthesis and Real Time qPCR
RNA extracts were used to generate first and second-strand cDNA's using random hexamer primers and the reverse transcriptase Superscript II® (Invitrogen™), as described in vanEngelsdorp et al. (2009). Pathogen loads were estimated using real-time quantitative-PCR (qPCR) and a Bio-Rad CFX-96™ thermocycler. Complementary DNA (cDNA) was generated from 1 μg RNA template and was amplified in a separate 20 μl final reaction volume of Sso-Fast™ SYBR® Green reaction mix (Bio-Rad™) for each diagnostic primer pair. We used published primers to survey for SBV, KBV, IAPV, DWV, ABPV, BQCV, trypanosomes (vanEngelsdorp et al., 2009), AFB (Evans et al., 2006), Nosema ceranae (Fries et al., 2013) and Nosema apis (Schwarz and Evans, 2013), Spiroplasma apis and Spiroplasma melliferum (Schwarz et al., 2014). Honey bee ribosomal protein S5 (RPS5) was used to normalize for cDNA content and to filter samples for degradation or experimental losses.
We used a thermal profile of 95°C for 30 s followed by 95°C for 5 s and 60°C for 30 s. Steps two and three were repeated for a total of 50 cycles and included plate reads for florescence during each 60°C step. Following the cycle program, products were denatured for 10 s at 95°C., reannealed and then a dissociation profile was measured between 69 and 95°C at an increment of 0.5°C to provide evidence for reaction fidelity (Evans et al., 2006). Annealing temperature of 55°C was used for the CytbSF (trypanosomatid L. passim) primers. Positive and negative control reactions were run on each 96-well plate. Pathogen loads (ΔΔCT) were determined as the difference between the CT of RPS5 and the CT of each target (ΔCT), scaled up from the minimal ΔCT across all samples.
PCR Purification and Sequencing
RT-PCR products were selected for sequencing to confirm the identities of products indicating the trypanosome, L. passim, and the bacteria S. apis and S. melliferum. These PCR products were purified using QIAquick PCR purification kit according to protocol recommended by manufacturer, and then were sequenced commercially by Macrogen (Rockville, MD, USA) DNA sequence similarity with trypanosomes, S. apis and S. melliferum, was confirmed using the BLAST search tool (Altschul et al., 1990) from the U.S. National Institutes of Health and searches against the National Center for Biotechnology Information (NCBI) nr database.
High Throughput Sequencing and Data Analysis
High throughput sequencing was performed with the pooled samples from stationary colonies of each region and all RNA samples were pooled before sequencing (n = 6 ILLUMINA RNA libraries). Libraries were run on paired-end 100-cycle reactions and flow cells using Illumina Hi-Seq 2000 machines at the University of Maryland Institute of Genome Sciences. Sequences for all six libraries have been deposited at the US National Institutes of Health NCBI “Honey Bee Disease Database” Bioproject (PRJNA52851).
After trimming and quality control of the generated sequences, transcriptomic analysis was done with the support of CLC Genomics Workbench 7.0.3 (CLC Bio, Aarhus, Denmark) by mapping sequencing reads and counting and distributing the reads across genes and transcripts based on annotated reference genes (Figure 2). During the alignment, two mismatches were allowed with deletion cost of three, insertion cost of three, length fraction of 0.8, similarity fraction of 0.8 and maximum 10 hits for a read. For statistical analyses, proportions-based tests were used for the comparison of the counts by considering the proportions that they comprise of the total sum of counts in each sample. Multi-group comparison was done by weighted t-type test statistic Baggerly's test (Baggerly et al., 2003). FDR (False Discovery Rate) corrected p-values were calculated according to the methods of Benjamini and Hochberg (1995) to determine the statistical significance of the pathogen load.
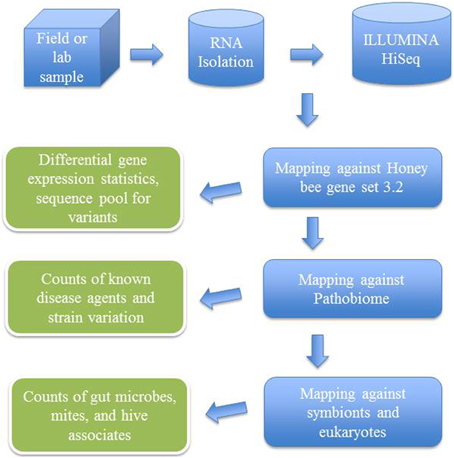
Figure 2. Flow chart showing analytical steps for mapping ILLUMINA RNA-seq reads against an annotated bee microbial dataset.
Real time q-PCR data statistics were performed by using JMP™ (SAS Institute, Cary, NC, USA, v.9). A One-Way analysis of variance was used to test differences between group means. The total variability in the response was divided into two parts; within-group variability and between-group variability. The differences between the group means were considered to be significant if the between-group variability was broader relative to the within-group variability. Multiple comparisons of group means were done by using pooled variance estimates for these means. Student's t-tests were computed for each pair of group levels and individual pairwise comparisons. Matrix of correlation coefficients that summarized the strength of the linear relationships between each pair of response variables was calculated and Pearson product-moment correlations for each pair of variables were listed. Correlations and the significance probabilities were calculated by the pairwise deletion method and the count values differed if any pair had a missing value for either variable.
Results
RT-qPCR Results
Kashmir bee virus (KBV), Israeli acute paralysis virus (IAPV), American foulbrood (AFB) and Sacbrood virus (SBV) were not detected by PCR in any of the samples, nor was the microsporidian pathogen N. apis (except for only three colonies in 2011) by RT-PCR. Present in these samples were Acute bee paralysis virus (ABPV), Deformed wing virus (DWV), Black queen cell virus (BQCV), and N. ceranae. Mixed infections of ABPV, DWV, BQCV, and N. ceranae were detected (Tables 1, 2).
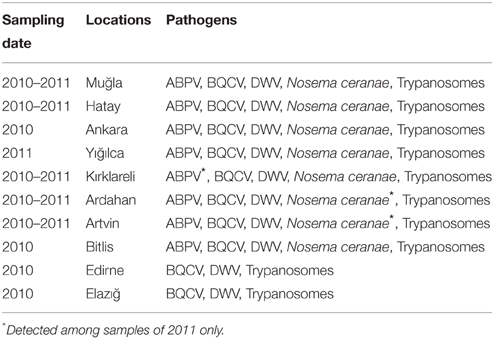
Table 1. The results of pathogen analysis of honey bee samples collected from different regions of Turkey.
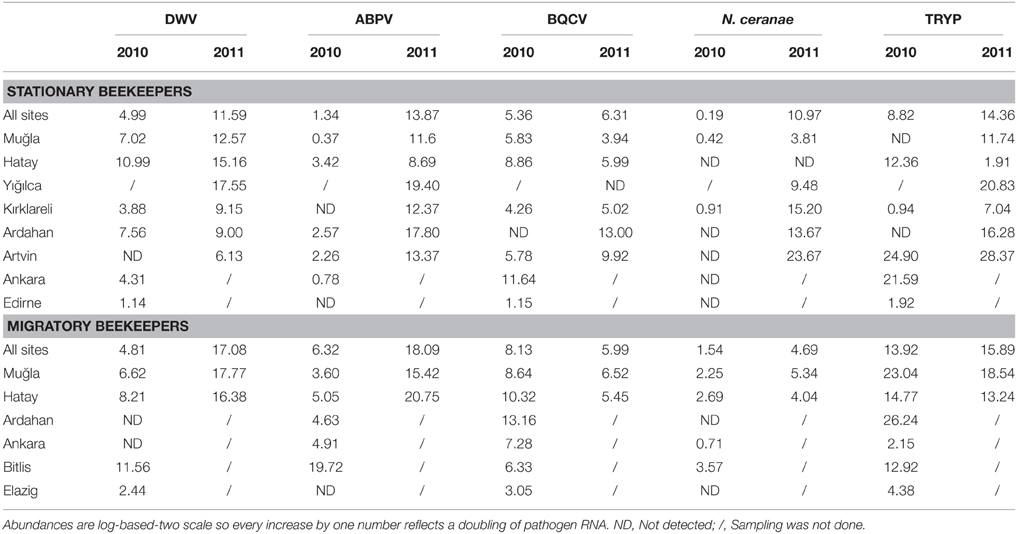
Table 2. Normalized pathogen loads across sites and between stationary and migratory beekeepers for 2010 and 2011.
The distribution of bee pathogens was significantly different among provinces and with beekeeping practices in 2010 (Table 3 in Supplementary Material). Generally, DWV loads were higher in Bitlis, Hatay, Muğla, and Ardahan than in other regions. ABPV was the most common virus in Bitlis and was especially high in the samples of migratory beekeepers. Among provinces that were sampled, BQCV loads were lowest in Edirne and highest in colonies of migratory beekeepers sampled in Ardahan, Hatay, and Muğla (Table 2). In 2011, DWV, BQCV, and N. ceranae loads were significantly different among the regions (Table 3 in Supplementary Material). DWV loads were higher in Muğla, Hatay, and Yığılca. BQCV levels were high in Ardahan and Artvin and was not detected at all in Yığılca. N. ceranae was more frequent among Artvin, Ardahan, and Kırklareli but not detected among stationary colonies of Hatay (Table 2). ABPV occurrence was the highest in samples from stationary apiaries of Yığılca and Ardahan and samples from migratory ones of Hatay and Muğla. The lowest level was in the stationary colonies of Hatay. ABPV loads showed difference between Yığılca-Kırklareli and Yığılca-Muğla (p = 0.0398 and p = 0.0478).
ABPV loads in Muğla and Ankara, BQCV loads in Ardahan, Muğla, and N. ceranae loads in Hatay were significantly higher in the samples of migratory beekeepers than the samples of stationary beekeepers in 2010. Among 2011 samples, the pathogen loads of migratory colonies were also higher in Muğla and Hatay but the results were not significant (Table 3 in Supplementary Material).
In 2010, N. ceranae was widespread in migratory colonies of Muğla, Hatay, and Bitlis and less frequent in samples from Ankara and Kırklareli. N. ceranae was not found in samples of beekeepers from Edirne, Artvin, Ardahan, and Elazığ in 2010 (Table 2). In contrast, N. ceranae showed high incidence in all sites except for the stationary colonies of Hatay in 2011 (Table 2). N. ceranae loads were higher among migratory colonies than stationary ones in both years. N. apis was not observed among the samples of 2010 and most of the samples of 2011 although the analysis was repeated twice. In 2011, N. apis was detected in three stationary colonies in total, from three beekeepers in Muğla-Bodrum, Düzce-Nas, and Ardahan-Posof.
Trypanosome loads were significantly different among regions in both years. Trypanosome abundance was higher within the samples of migratory beekeepers than those of stationary ones (Table 3 in Supplementary Material). Ten/ten positive samples were confirmed by DNA sequencing as reflecting the 28S rRNA locus of trypanosomes. In general trypanosome levels were higher among samples from Muğla, Ankara, Artvin and Ardahan compared to the samples from Kırklareli, Edirne and Elazığ, for 2010. In 2011, levels were especially high in samples from Artvin, Yığılca, and Muğla, and low in samples of stationary colonies in Hatay and Kırklareli (Table 2). Seasonal variation was observed among samples collected in 2010 (ANOVA p = 0.0023). Specifically, trypanosome loads were highest in spring as opposed to fall. In Ardahan and Hatay, trypanosomes were not detected among any of the fall samples. Among the samples with trypanosome infection, thirteen 2010 samples for which products were assayed by DNA sequencing were all confirmed as being part of the “SF” strain of L. passim (identified as C. mellificae) (Runckel et al., 2011) and 10 were positive among 51 samples in 2011, based on qPCR and melt-curve analyses (seven products were confirmed by DNA sequencing). four of the “SF” positive samples were from Muğla, six from Hatay, eight from Artvin-Ardahan, four from Yığılca, and one from Elazığ. GAPDH sequence analyses from Artvin and Yığılca all matched L. passim, haplotype A (Morimoto et al., 2013), indicating that this haplotype was more common among the samples when compared with haplotype B.
S. melliferum was detected for two samples in Artvin province in 2010 while 13 samples were positive for Artvin, Ardahan, Muğla, Yığılca, and Kırklareli during 2011. Six of the positive samples were confirmed by DNA sequencing. S. apis was detected in only two samples from Muğla and Elazığ in 2010 and in one sample from Yığılca in 2011.
Dual infections of DWV and BQCV were detected in both migratory and stationary beekeepers in 2010 (50–100% of honey bees in six provinces sampled). Dual infections with these two viruses were less commonly observed in 2011 (18–40% of honey bees in four of the provinces sampled). Triple infections of ABPV, BQCV, and DWV were seen in three regions (50–100% of honeybee samples) in 2010, whereas occurred less (18–40% of honey bee samples) at four regions in 2011. Among 2010 samples, N. ceranae loads were correlated with DWV loads (r = 0.25, p = 0.0186). ABPV loads were correlated with both DWV (r = 0.40, p = 0.0001) and N. ceranae (r = 0.35, p = 0.0009). BQCV correlated with DWV within samples from Ankara, Bitlis, Edirne, and Muğla. In Bitlis, ABPV levels were positively correlated with DWV (r = 0.89, p = 0.0061). In Hatay, BQCV correlated with N. ceranae (r = 0.47, p = 0.0443). Among 2011 samples ABPV showed correlations with DWV (r = 0.29, p = 0.0381) and in some regions with N. ceranae. A positive correlation was observed between trypanosome load and N. ceranae (r = 0.28, p = 0.0083) and between trypanosomes and BQCV (r = 0.59, p < 0.0001).
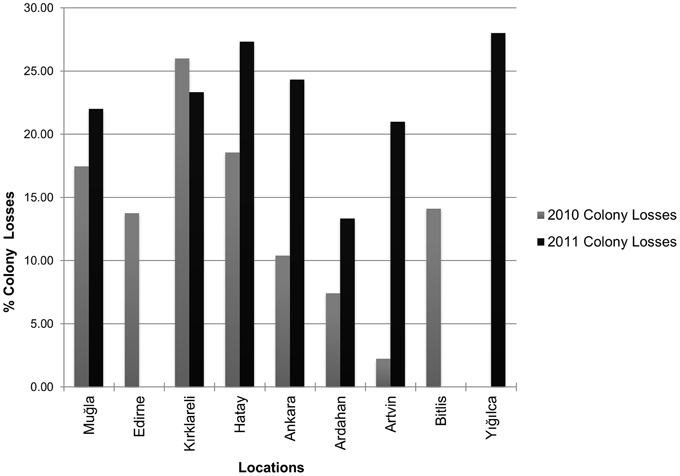
According to the survey results, there was an increase in colony losses in 2011 when compared with 2010 losses (Figure 3). Kırklareli, Muğla, Hatay, and Ankara had relatively higher colony losses in both years while Ardahan and Artvin tended to have lower losses than the other provinces. High colony losses were also observed in Edirne and Bitlis in 2010 and especially in Yığılca in 2011. Average colony losses of migratory beekeepers were significantly higher than those of the stationary beekeepers.
Read Mapping Results
DWV reads were detected in sequenced RNA libraries from all of the regions (Figure 4 and Table 4 in Supplementary Material), although they were most highly represented in Hatay, Yığılca, and Muğla, matching the results from real time q-PCR. We found strong evidence for Varroa destructor virus-1 as well, a first for these regions. Overall, the DWV/VDV group of iflaviruses was the predominant viral taxon (Figure 4). RNA sequences of BQCV were highest in the Ardahan region, although BQCV reads were also abundant in other provinces. Consistent with the fact that BQCV was not detected in Yığılca by RT-qPCR very few reads were found in this province during RNA sequencing According to the sequence analysis, ABPV contigs were common in most regions. Consistent with RT-PCR results, they were most frequently observed in Yığılca and Ardahan. Very few CBPV reads were found in Kirklareli province and this virus was not detected or was very few in other regions. In contrast, the related Lake Sinai viruses were highly represented both in diversity across this large clade, and in abundance. IAPV and KBV were notably rare or were not present across all regions. Sacbrood virus was similarly rare, although 1885 SBV reads were detected in Ardahan region. SBPV reads (Slow Bee Paralysis Virus) were not observed in any of the regions (Table 4 in Supplementary Material).
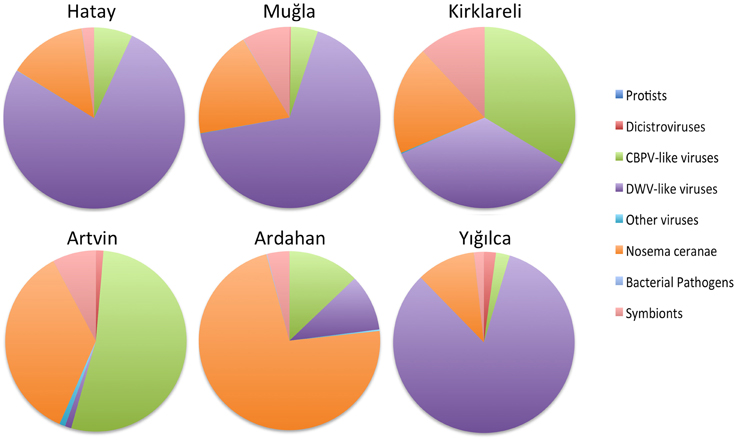
Figure 4. Relative microbial loads across the six regions, as inferred by RNA-Seq matches. Taxa are binned by taxonomic group.
Nosema ceranae was highly prevalent in our RNA-seq analysis. N. ceranae was especially common in Artvin province, again matching our RT-qPCR analyses. N. ceranae was not detected in stationary colonies of Hatay province but it was evident there by RNA-seq analysis. We found minimal, if any reads matching N. apis in the RNASeq data, those present were largely the result of regions with high sequence similarity to N. ceranae. Reads for the betaproteobacterium, Snodgrassella alvi were common among all of the provinces with highest frequency in Kırklareli, Muğla, Yığılca, and Ardahan. The gamma-proteobacterium, Gilliamella apicola was prevalent in all regions, while the related Frischella perrara was rare (Table 4 in Supplementary Material). Consistent with both RT-qPCR and DNA sequencing, the presence of Spiroplasma melliferum was confirmed in most of the regions. The abundance of congener S. apis was confirmed with the RNA-seq analysis despite being missed by PCR, yet S. apis remained the minor of the two species.
Discussion
Evaluation of Real Time q-PCR Results
The surveyed regions comprise important centers of beekeeping in Turkey. Kashmir Bee Virus (KBV) and Israel Acute Bee Paralysis Virus (IAPV) were not detected in sampled colonies by RT-qPCR, while the related virus Acute Bee Paralysis Virus (ABPV) was abundant in most of the samples and was present in the regions where colony declines were observed. Historically, KBV has been found most frequently in the United States and Australia (Allen and Ball, 1996) and this virus is thought to be an exotic bee virus in central Europe (Berenyi et al., 2006). Our discovery of just one of these three species could reflect climatic and environmental conditions (Anderson, 1991) or differential abilities of these viruses to infect different honey bee subspecies.
DWV was detected in both migratory and stationary operations in all of the regions, consistent with the high prevalence of this virus worldwide (Tentcheva et al., 2004; Berenyi et al., 2006; Kukielka et al., 2008; Nielsen et al., 2008; Welch et al., 2009). The presence of DWV has been reported in colonies of Apis mellifera L. in Ordu province of Turkey (Gülmez et al., 2009). BQCV was the second most widespread bee virus in our study. BQCV is also the second-most prevalent virus in honey bee colonies in Asia and Europe (Tentcheva et al., 2004) and its presence was confirmed in many studies (Berenyi et al., 2006; Kukielka et al., 2008; Welch et al., 2009; Choe et al., 2012). BQCV was previously reported in 21.42% of 28 bee samples from six provinces of the Black Sea Region in Turkey (Gümüşova et al., 2010). BQCV was more prevalent in migratory bee colonies, consistent with the results found for migratory bees sampled in the U.S. (Welch et al., 2009).
In most of the regions we studied, BQCV and DWV were found together more often than expected by chance. This result is consistent with the findings in a previous study of virus infections (Chen et al., 2004) for which these distantly related viruses were found coinfecting honey bees at a high frequency. Triple infections of ABPV, BQCV, and DWV were detected in some colonies. Nielsen et al. (2008) also found a high incidence of dual and triple infections. According to Chen et al. (2004), 50% of colonies in the USA had dual infections while 7% had triple infections. Simultaneous multiple infections were also common in Austria (Berenyi et al., 2006), Southwest England (Baker and Schroeder, 2008), Brazil (Teixeira et al., 2008), Jordan (Haddad et al., 2008), France (Tentcheva et al., 2004), and Hungary (Forgach et al., 2008).
In Turkey, the presence of N. apis was confirmed earlier (Aydin et al., 2005; Muz et al., 2010; Whitaker et al., 2010). Using qPCR, we detected N. apis in only three colonies, indicating the replacement of N. apis by N. ceranae in many regions included in this study. Worldwide, N. ceranae is now far more common than N. apis (Chen et al., 2009; Valera et al., 2011; Yoshiyama and Kimura, 2011).
In 2006, N. ceranae was found in the provinces of Artvin, Hatay, and Muğla (Whitaker et al., 2010). Collapsed colonies from the Hatay overwintering region and Southeastern Marmara were found to show infections of N. ceranae (Muz et al., 2010). In our 2010 samples, N. ceranae was observed in Bitlis, Hatay, and Muğla and not in Edirne, Artvin, Ardahan, and Elazığ in 2010. Interestingly, N. ceranae was widely common for all of the regions involved in the study in 2011. Detection of N. ceranae among 2011 samples of Ardahan-Artvin might be the reason for the increase in colony losses in these provinces in 2011. The incidence of N. ceranae in Kırklareli province in 2011 was much more than the previous year. N. ceranae loads of samples from migratory beekeepers were significantly higher than the stationary ones and N. ceranae loads were correlated the viruses ABPV, BQCV and DWV. We propose that the infectivity of N. ceranae expands with migratory beekeeping activities and in association with different viruses and trypanosomatids. In July, the Thrace region is a highly frequented location by migratory beekeepers seeking to harvest sunflower honey. It is thought that these beekeepers in this region had colony losses because of the application of pesticides rather than honey bee disease factors. Hatay and Ankara are also important locations for migratory beekeepers. Caucasian bees which are used by most of the migratory beekeepers, are raised in Ardahan and Artvin. Bitlis also suffered high colony losses in 2010, combined with high disease loads among samples of migratory beekeepers. In contrast, while honeybee samples in Yığılca were derived from stationary colonies, high colony losses and high pathogen loads were observed. Probably some other factors were contributing to colony losses in this province. All five viruses and pathogens were also detected in Muğla, the center of migratory beekeeping. These results could reflect the differences in pathogen exposure of local and migratory colonies, varying resistance levels, or perhaps a differential ability to handle stress. Genetic impacts of migratory beekeeping has become an important concern in Turkey and elsewhere. Population structure can be disturbed with hybridization, leading to a loss of regionally adaptive traits and perhaps decreasing colony fitness.
Trypanosomes were highly prevalent in our samples. Trypanosomes have also been reported in Australia (Langridge and McGhee, 1967), China (Yang et al., 2013), France (Dainat et al., 2012), Japan (Morimoto et al., 2013), Switzerland (Schmid-Hempel and Tognazzo, 2010), USA (vanEngelsdorp et al., 2009; Runckel et al., 2011), and Spain (Orantes-Bermejo, 1999). Trypanosome levels were higher in summer samples than the fall samples in our samples. In southern Spain, this parasite appears in July and August (Orantes-Bermejo, 1999) and L. passim (recently named as the primary trypanosomatid in lieu of C. mellificae, Schwarz et al., 2015) was common in summer colonies in Belgium (Ravoet et al., 2013). These findings are in contrast with the results of Runckel et al. (2011) which show a peak in L. passim levels in January in U.S. colonies.
Synergistic effects can make colonies more vulnerable to other pathogens (Cornman et al., 2012) and we noted a positive correlation between L. passim and N. ceranae prevalence. Similarly, this relationship was documented in field surveys from the U.S. (Runckel et al., 2011) and the co-occurence of L. passim and N. ceranae in summer was tied to higher colony mortality in Belgium (Ravoet et al., 2013). Complex dynamic immune responses of honey bees to both Nosema and trypanosomatids were recently reported (Schwarz and Evans, 2013) and it will be interesting to test further for mechanistic explanations for any synergisms between these parasites.
Metagenomic Sequencing
As in prior studies, iflaviruses related to Deformed wing virus (DWV) were especially prevalent. While DWV amounts were higher, reads for the closely related Varroa destructor-1 virus (VDV-1) were fairly abundant in Hatay, Yığılca, and Muğla. CBPV reads were relatively rare in most of the regions and were not detected in Yığılca and Ardahan. CBPV was reported in seven of 28 (25%) samples from six provinces of Black Sea region in Turkey (Gümüşova et al., 2010). CBPV was detected in four of 96 apiaries in survey study of Denmark, in 73% of the samples from Greek and 9% of apiaries in China (Nielsen et al., 2008; Bacandritsos et al., 2010; Ai et al., 2012). A recently described relative of CBPV, Lake Sinai virus, was prevalent in all regions, second only to the DWV and VDV group. In fact, this group was the most prevalent virus in Kirklareli and Ardahan provinces. The pathogenic or epidemiological significance of Lake Sinai viruses are not well-known. The LSV species complex is diverse, with members sharing between 70 and 99% sequence identity, hence this group is often missed in screenings based on PCR. LSV4 appears to be especially abundant, along with LSV1 and LSV2. There was significant variation across the regions in the specific lineages seen for this group. LSV2 was the most abundant single component of the honey bee microbiome in the study of Runckel et al. (2011). The presence of LSV was also confirmed in honey bees from Spain by high-throughput sequencing (Granberg et al., 2013). A new fourth strain of Lake Sinai Virus (LSV) was identified in the study of Ravoet et al. (2013).
In this study, despite tens of millions of microbial gene reads, few reads matching IAPV, KBV, and SBV were seen. IAPV, KBV, and SBV were not detected using RT-qPCR, indicating that RNA-seq sensitivity was higher than that of qPCR, or that current qPCR primers for this group must be redesigned to capture all strains. Given the RNA-Seq data, we can consider that these pathogens are quite rare in our samples. KBV has been absent in some other European surveys (e.g., Berenyi et al., 2006; Forgach et al., 2008) and rare in surveys in France, Denmark, and United Kingdom (Tentcheva et al., 2004; Ward et al., 2007; Nielsen et al., 2008). IAPV is prevalent in the Middle East and Australia (Maori et al., 2007; Palacios et al., 2008) and this species was reported in 71 samples containing 10 bees; each from 20 provinces in Turkey (Ozkirim and Schiesser, 2013). In this study IAPV wasn't confirmed by RT-qPCR in our samples and RNA sequence analysis revealed that this virus was not present in Muğla and Hatay and was incredibly rare among samples from Artvin, Yığılca, and Ardahan. SBV was similarly scarce in this study. This conflicts with results of Tentcheva et al. (2004) but overlaps with the results of Baker and Schroeder (2008) and Forgach et al. (2008). As with most surveys, we examined only adult honey bees. SBV causes a fatal disease in honey bee larvae (Bailey, 1975), thus brood samples could provide more evidence for its prevalence.
Apis mellifera filamentous virus (AmFV) was not detected within our samples, nor was VdMLV (Varroa Macula-like virus). A plant-pathogenic RNA virus, tobacco ringspot virus (TRSV) was not detected in any of the RNA samples in our study.
Recent culture-independent studies reported eight bacterial phylotypes inhabiting the gut of the honey bee, Apis mellifera from several continents (Jeyaprakash et al., 2003; Mohr and Tebbe, 2006; Babendreier et al., 2007; Cox-Foster et al., 2007; Olofsson and Vasquez, 2008). Colonies founded by swarms, interactions within the colony, intercolony interactions like robbing food in neighboring hives and mixing of colonies by beekeepers all might affect the gut microbiota (Engel et al., 2012). In our study, unique gene reads of Snodgrassella alvi and Gilliamella apicola were highly represented and showed differences among provinces because of geographic, environmental and subspecies differences of hosts. The differences in social behaviors of the subspecies, the dietary sources and exposure to varying pathogens and pesticides might influence the abundance of these bacteria among regions. One gamma-proteobacterial member of the gut microbiota Candidatus Schmidhempelia bombi, was present in 90% of bumble bee individuals in the study of Martinson et al. (2014). This symbiont was prevalent in all of our surveyed locations and widely represented in Hatay, Yığılca, and Muğla. The recently described gamma-proteobacterium Frischella perrara (Engel et al., 2013) was present in our study albeit at low levels.
Lactobacillales symbionts have been proposed as actors in both nutrition and parasite defenses of honey bees. Lactobacillales stimulate the innate immune system, arguably increasing honey bee defenses against disease agents (Evans and Lopez, 2004). Along with their impacts on immunity, the microbial symbionts have been proposed to nutritionally compete with pathogens by occupying the available niches (Crotti et al., 2012). In this study, very low number of 16S Lactobacillus reads were observed among regions. Overall bacterial loads were especially high in Hatay, Yığılca, and Artvin. The prevalence of the bacterial pathogens S. apis and S. melliferum is low among Belgian honey bee colonies (Ravoet et al., 2013), and these bacteria are present only seasonally in North and South America (Runckel et al., 2011; Schwarz et al., 2014). We found low Spiroplasma levels, with highest incidence in Artvin province. S. melliferum was the more common species, matching results from the Americas (Schwarz et al., 2014). The bacterial brood diseases European Foulbrood (EFB) caused by the bacterium Melissococcus plutonius (Bailey et al., 1983) and American Foulbrood (AFB) caused by the bacterium Paenibacillus larvae (Genersch et al., 2006) are globally important diseases of honey bees. P. larvae reads were not common among our samples. Similarly, M. plutonius was not prevalent among the regions but was more ubiquitous in Hatay, Muğla, and Yığılca. Environmental conditions at these sites can be conductive for the expression of the disease. Like AFB, EFB transmission is also linked to larval immune responses (Evans, 2004), hygienic behavior (Spivak and Reuter, 2001) as well as interaction between M. plutonius and the intestinal microbiota of the honey bee larvae (Gilliam, 1997; Olofsson and Vasquez, 2008), nutritional and stress conditions, weather and geography (Bailey, 1961).
Sequences for Ascosphaera apis, the causative agent for Chalkbrood disease, were generally rare, with the highest incidence from samples of in Hatay province. Similarly, neogregarines (nominally Apicystis bombi) persisted at extremely low levels among regions in this study. Among arthropod parasites of honey bees, we found no genetic evidence for the presence of the tarsonemid tracheal mite, Acarapis woodi within our samples. The Asian parasitic mite Tropilaelaps is considered more dangerous to A. mellifera than the parasitic mite Varroa destructor (Rath et al., 1995), and this mite is worthy of screening. Our deep sequencing analysis showed no sign of Tropilaelaps. The presence of phorid flies (Apocephalus borealis) in the study of Ravoet et al. (2013) proves their existence in Europe, but our deep sequencing did not reveal signs of this parasite.
In conclusion, we screened bee-derived RNA against the most complete sequence set for honey bee associates used to date. We assessed levels of known and novel parasites, pathogens, and symbionts. We present quantitative data for bacterial pathogens (Melissococcus plutonius, Paenibacillus larvae, S. apis, S. melliferum), protists (Apicytis, trypanosomatids), viruses (Lake Sinai virus, Chronic bee paralysis virus, Deformed wing virus, Varroa destructor virus, Sacbrood, and Dicistroviruses), symbionts (Candidatus Schmidhempelia bombi, Frischella perrara, Snodgrassella alvi, Gilliamella apicola, Lactobacillus spp., Acetobacteracea), microsporidia and fungi in A. mellifera colonies in distinct regions of Turkey. The presence of KBV, SBPV, Tobacco ringspot virus, VdMLV (Varroa Macula like), Acarapis spp., Tropilaelaps claerae and Apocephalus (phorid fly) were also examined. As in other countries, bee viruses were correlated with colony losses in Turkey. In comparison with 2010, the increase in pathogen loads in 2011 might be a factor for increased colony losses observed in this study. It seems likely that migratory beekeeping practices enable the spread of disease factors among honey bees in places where they visit and causing an important threat to the honey bee colonies. In addition, the impacts of parasites and pathogens varies between regions, perhaps reflecting different honey bee genetic traits. Migratory beekeeping was correlated with both higher disease loads and a potential risk of dispersing regional parasites and pathogens across the country. This practice also allows for greater gene flow between migratory honey bee populations and local populations. Current diversity and local genetic structure can be preserved with selection strategies and establishing broad areas of isolation to reduce the risks of migratory beekeeping practices. While experimental work and longitudinal analyses will be needed to confirm causes of bee declines, our analyses, reference sequences, and strategy will help reduce the set of likely causes.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
It is with sadness that we note the passing of COLOSS Management Committee member Dr. AK on February 1, 2014. We dedicate this paper to the memory of Professor AK, for initiating the study and for his teaching, mentoring, collegiality and friendship. The work was part of the project “The effects of the genetic differences of Turkish honey bees on colony vitality” and the work was funded by “The Science and Technological Research Council of Turkey” (TUBITAK Project Number:109T547). The work was also supported by Middle East Technical University (BAP Project number:BAP-01-08-2012-004. The molecular diagnostics part was technically and financially supported by the USDA-ARS Bee Research Laboratory. We would like to express our gratitude to Dr. Ryan Schwarz for technical advice, Dawn Lopez and Margaret Smith for their laboratory guidance, and the scientists of the BRL for their feedback. We appreciate the advice and help of Prof. Dr. Muhsin Dogaroglu, Prof.Dr. Banu Yucel and Dr. Hasan Huseyin Inal in the initiation of the project. We are also grateful to Assist. Prof. Dr. Mustafa Muz, Okan Can Arslan, Mehmet Kayim, Mert Kukrer, and Mustafa Nail Cirik for their contributions to the field work.
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fgene.2015.00100/abstract
References
Ai, H., Yan, X., and Han, R. (2012). Occurrence and prevalence of seven bee viruses in Apis melliferas and Apis cerana apiaries in China. J. Invertebr. Pathol. 109, 160–164. doi: 10.1016/j.jip.2011.10.006
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Allen, M., and Ball, B. (1996). The incidence and world distribution of honey bee viruses. Bee World 77, 141–162.
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local alignment search tool. J. Mol. Biol. 215, 403–410. doi: 10.1016/S0022-2836(05)80360-2
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Anderson, D. L. (1991). Pests and pathogens of the honeybee (Apis mellifera L.) in Fiji. J. Apic. Res. 29, 53–59.
Anderson, D. L., and Gibbs, A. J. (1989). Transpuparial transmiion of Kashmir bee virus and sacbrood virus in the honey bee (Apis mellifera). Ann. Appl. Biol. 114, 1–7. doi: 10.1111/j.1744-7348.1989.tb06781.x
Aydin, L., Çakmak, Y., Güleğen, E., and Wells, H. (2005). Honey bee nosema disease in the Republic of Turkey. J. Apic. Res. 44, 196–197. doi: 10.3896/IBRA.1.44.4.13
Babendreier, D., Joller, D., Romeis, J., Bigler, F., and Widmer, F. (2007). Bacterial community structures in honeybee intestines and their response to two insecticidal proteins. FEMS. Microbiol. Ecol. 59, 600–610. doi: 10.1111/j.1574-6941.2006.00249.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bacandritsos, N., Granato, A., Budge, G., Papanastasiou, I., Roinioti, E., Caldon, M., et al. (2010). Sudden deaths and colony population decline in Greek honey bee colonies. J. Invertebr. Pathol. 105, 335–340. doi: 10.1016/j.jip.2010.08.004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Baggerly, K., Deng, L., Morris, J., and Aldaz, C. (2003). Differential expression in SAGE: accounting for normal between-library variation. Bioinformatics 19, 1477–1483. doi: 10.1093/bioinformatics/btg173
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bailey, L., Ball, B. V., and Perry, J. N. (1983). Association of viruses with 2 protozoal pathogens of the honeybee. Ann. Appl. Biol. 103, 13–20. doi: 10.1111/j.1744-7348.1983.tb02735.x
Baker, A., and Schroeder, D. (2008). Occurrence and genetic analysis of picorna-like viruses infecting worker bees of Apis mellifera L. populations in Devon, South West England. J. Invertebr. Pathol. 98, 239–242. doi: 10.1016/j.jip.2008.02.010
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ball, B. V., and Bailey, L. (1997). “Viruses,” in Honey Bee Pests Predators and Diseases, eds R. A. Morse and K. Flottum (Medina: AI Root Co), 11–31.
Benjamini, Y., and Hochberg, Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B. Stat. Methodol. 57, 289–289.
Berenyi, O., Bakonyi, T., Derakhshifar, I., Koglberger, H., and Nowotny, N. (2006). Occurrence of six honeybee viruses in diseased Austrian apiaries. Appl. Environ. Microbiol. 72, 2414–2420. doi: 10.1128/AEM.72.4.2414-2420.2006
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bishop-Lilly, K. A., Turell, M. J., Willner, K. M., Butani, A., Nolan, N. M., Lentz, S. M., et al. (2010). Arbovirus detection in insect vectors by rapid, high-throughput pyrosequencing. PLoS Negl. Trop. Dis. 4:e878. doi: 10.1371/journal.pntd.0000878
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bodur, C., Kence, M., and Kence, A. (2007). Genetic structure of honeybee, Apis mellifera L. (Hymenoptera: Apidae) populations of Turkey inferred from microsatellite analysis. J. Apic. Res. 46, 50–56. doi: 10.3896/IBRA.1.46.1.09
Brown, M. J. F., Schmid-Hempel, R., and Schmid-Hempel, P. (2003). Strong context-dependent virulence in a hostparasite system: reconciling genetic evidence with theory. J. Anim. Ecol. 72, 994–1002. doi: 10.1046/j.1365-2656.2003.00770.x
Chen, Y., Zhao, Y., Hammond, J., Hsu, H. T., Evans, J., and Feldlaufer, M. (2004). Multiple virus infections in the honey bee and genome divergence of honey bee viruses. J. Invertebr. Pathol. 87, 84–93. doi: 10.1016/j.jip.2004.07.005
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Chen, Y. P., Evans, J. D., Zhou, L., Boncristiani, H., Kimura, K., Xiao, T., et al. (2009). Asymmetrical coexistence of Nosema ceranae and Nosema apis in honey bees. J. Invertebr. Pathol. 101, 204–209. doi: 10.1016/j.jip.2009.05.012
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Choe, S. E., Nguyen, L. T. K., Noh, J. H., Koh, H. B., Jean, Y. H., Kweon, C. H., et al. (2012). Prevalence and distribution of six bee viruses in Korean Apis cerana populations. J. Invertebr. Pathol. 109, 330–333. doi: 10.1016/j.jip.2012.01.003
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Clark, T. B., Whitcomb, R. F., Tully, J. G., Mouches, C., Saillard, C., Bove, J. M., et al. (1985). Spiroplasma melliferum, a new species from the honey bee (Apis mellifera). Int. J. Syst. Bacteriol. 35, 296–308. doi: 10.1099/00207713-35-3-296
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cornman, R. S., Boncristiani, H., Dainat, B., Chen, Y. P., vanEngelsdorp, D., Weaver, D., et al. (2013). Population genomic variation within RNA viruses of the Western honey bee, Apis mellifera, inferred from deep sequencing. BMC Genomics 14:154. doi: 10.1186/1471-2164-14-154
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cornman, R. S., Tarpy, D. R., Chen, Y., Jeffreys, L., Lopez, D., Pettis, J. S., et al. (2012). Pathogen webs in collapsing honey bee colonies. PLoS ONE 7:e43562. doi: 10.1371/journal.pone.0043562
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cox-Foster, D. L., Conlan, S., Holmes, E. C., Palacios, G., Evans, J. D., Moran, N. A., et al. (2007). A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318, 283–287. doi: 10.1126/science.1146498
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Crotti, E., Balloi, A., Hamdi, C., Sansonno, L., Marzorati, M., Gonella, E., et al. (2012). Microbial symbionts: a resource for the management of insect-related problems. Microb. Biotechnol. 5, 307–317. doi: 10.1111/j.1751-7915.2011.00312.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dainat, B., Evans, J. D., Gauthier, L., and Neumann, P. (2012). “Crithidia mellificae is widespread in Europe and can be used as a predictive marker of honeybee colony losses,” in Proceedings of the 5th European Conference of Apidology (Halle an der Saale).
De Miranda, J. R., Cordoni, G., and Budge, G. (2010). The Acute bee paralysis virus-Kashmir bee virus-Israeli acute paralysis virus complex. J. Invert. Pathol. 103, 30–47. doi: 10.1016/j.jip.2009.06.014
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dillon, R. J., and Dillon, V. M. (2004). The gut bacteria of insects: nonpathogenic interactions. Annu. Rev. Entomol. 49, 71–92. doi: 10.1146/annurev.ento.49.061802.123416
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ellis, J. D., and Munn, P. A. (2005). The worldwide health status of honey bees. Bee World 86, 88–101. Available online at: http://www.ibra.org.uk/articles/20080611_27
Engel, P., Kwong, W. K., and Moran, N. A. (2013). Frischella perrara gen. nov., sp. nov., a gammaproteobacterium isolated from the gut of the honeybee, Apis mellifera. Int. J. Syst. Evol. Microbiol. 63, 3646–3651. doi: 10.1099/ijs.0.049569-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Engel, P., Martinson, V. G., and Moran, N. A. (2012). Functional diversity within the simple gut microbiota of the honey bee. Proc. Natl. Acad. Sci. U.S.A. 109, 11002–11007. doi: 10.1073/pnas.1202970109
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Evans, J., Aronstein, K. A., Chen, Y., Hetru, C., Imler, J.-L., Jiang, H., et al. (2006). Immune pathways and defence mechanisms in honey bees Apis mellifera. Insect. Mol. Biol. 15, 645–656. doi: 10.1111/j.1365-2583.2006.00682.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Evans, J. D. (2004). Transcriptional immune responses by honey bee larvae during invasion by the bacterial pathogen, Paenibacillus larvae. J. Invertebr. Pathol. 85, 105–111. doi: 10.1016/j.jip.2004.02.004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Evans, J. D., Chen, Y. P., Cornman, R. S., De La Rua, P., Foret, S., Foster, L., et al. (2013). “Standard methodologies for molecular research in Apis mellifera,” in The COLOSS BEEBOOK, Volume I: Standard Methods for Apis Mellifera Research, eds V. Dietemann, J. D. Ellis, and P. Neumann (Treforest, UK: International Bee Research Association), 52.
Evans, J. D., and Lopez, D. L. (2004). Bacterial probiotics induce an immune response in the honey bee (Hymenoptera: Apidae). J. Econ. Entomol. 97, 752–756. doi: 10.1093/jee/97.3.752
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Forgach, P., Bakonyi, T., Tapaszti, Z., Nowotny, N., and Rusvai, M. (2008). Prevalence of pathogenic bee viruses in Hungarian apiaries: situation before joining the European Union. J. Invertebr. Pathol. 98, 235–238. doi: 10.1016/j.jip.2007.11.002
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fries, I. (2010). Nosema ceranae in European honey bees (Apis mellifera). J. Invertebr. Pathol. 103, 73–79. doi: 10.1016/j.jip.2009.06.017
Fries, I., Chauzat, M. P., Chen, Y. P., Doublet, V., Genersch, E., Gisder, S., et al. (2013). Standard methods for Nosema research. J. Apic. Res. 52, 1–28. doi: 10.3896/IBRA.1.52.1.14
Fries, I., Feng, F., daSilva, A., Slemenda, S. B., and Pieniazek, N. J. (1996). Nosema ceranae n. sp. (Microspora, Nosematidae), morphological and molecular characterization of a microsporidian parasite of the Asian honey bee Apis cerana (Hymenoptera, Apidae). Eur. J. Protistol. 32, 356–365. doi: 10.1016/S0932-4739(96)80059-9
Fries, I., Martin, R., Meana, A., Garcia-Palencia, P., and Higes, M. (2006). Natural infections of Nosema ceranae in European honey bees. J. Apic. Res. 45, 230–233. doi: 10.3896/IBRA.1.45.4.13
Gegear, R. J., Otterstatter, M. C., and Thomson, J. D. (2005). Does parasitic infection impair the ability of bumblebees to learn flower-handling techniques? Anim. Behav. 70, 209–215. doi: 10.1016/j.anbehav.2004.09.025
Genersch, E., Forsgren, E., Pentikainen, J., Ashiralieva, A., Rauch, S., Kilwinski, J., et al. (2006). Reclassification of Paenibacillus larvae subsp. pulvifaciens and Paenibacillus larvae subsp larvae as Paenibacillus larvae without subspecies differentiation. Int. J. Syst. Evol. Microbiol. 56, 501–511. doi: 10.1099/ijs.0.63928-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gilliam, M. (1997). Identification and roles of non-pathogenic microflora associated with honey bees. FEMS Microbiol. Lett. 155, 1–10. doi: 10.1016/S0378-1097(97)00337-6
Granberg, F., Vicente-Rubiano, M., Rubio-Guerri, C., Karlsson, O. E., Kukielka, D., Belák, S., et al. (2013). Metagenomic detection of viral pathogens in spanish honeybees: coinfection by aphid lethal paralysis, Israel acute paralysis and lake sinai viruses. PLoS ONE 8:e57459. doi: 10.1371/journal.pone.0057459
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gülmez, Y., Bursali, A., and Tekin, S. (2009). First molecular detection and characterization of deformed wing virus (DWV) in honeybees (Apis mellifera L.) and mites (Varroa destructor) in Turkey. Afr. J. Biotechnol. 8, 3698–3702. Available online at: http://www.ajol.info/index.php/ajb/article/viewFile/61919/49978
Gümüşova, S. O., Albayrak, H., Kurt, M., and Yazici, Z. (2010). Prevelance of three honey bee viruses in Turkey. Veterinarski Arhiv. 80, 779–785. Available online at: http://www.vef.unizg.hr/vetarhiv/papers/2010-80-6-10.pdf
Haddad, N., Brake, M., Migdadi, H., and de Miranda, J. R. (2008). First detection of honey bee viruses in Jordan by RT-PCR. Jordan J. Agricul. Sci. 4, 242–246. Available online at: https://journals.ju.edu.jo/JJAS/article/view/1003/996
Higes, M., Martín-Hernández, R., Botías, C., Bailón, E., González-Porto, A., Barrios, L., et al. (2008). How natural infection by Nosema ceranae causes honey bee colony collapse. Environ. Microbiol. 10, 2659–2669. doi: 10.1111/j.1462-2920.2008.01687.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jeyaprakash, A., Hoy, M. A., and Allsopp, M. H. (2003). Bacterial diversity in worker adults of Apis mellifera capensis and Apis mellifera scutellata (Insecta: Hymenoptera) assessed using 16S rRNA sequences. J. Invertebr. Pathol. 84, 96–103. doi: 10.1016/j.jip.2003.08.007
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kandemir, Ý., Kence, M., and Kence, A. (2000). Genetic and morphometric variation in honeybee (Apis mellifera) populations of Turkey. Apidologie 31, 343–356. doi: 10.1051/apido:2000126
Kevan, P. G., Guzman, E., Skinner, A., and van Englesdorp, D. (2007). Colony Collapse Disorder (CCD) in Canada: do we have a problem? Hivelights 20, 15–18. Available online at: http://www.honeycouncil.ca/hivelights.php?hlID=21
Klee, J., Besana, A. M., Genersch, E., Gisder, S., Nanetti, A., Tam, D. Q., et al. (2007). Widespread dispersal of the microsporidian Nosema ceranae, an emergent pathogen of the western honey bee, Apis mellifera. J. Invertebr. Pathol. 96, 1–10. doi: 10.1016/j.jip.2007.02.014
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kukielka, D., Esperon, F., Higes, M., and Sanchez-Vizcaino, J. M. (2008). A sensitive one-step real-time RT-PCR method for detection of deformed wing virus and black queen cell virus in honeybee Apis mellifera. J. Virol. Methods 147, 275–281. doi: 10.1016/j.jviromet.2007.09.008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lange, C. E., and Lord, J. (2012). “Protistan entomopathogens,” in Insect Pathology, 2nd Edn., eds B. Vega and H. Kaya (Amsterdam: Elsevier), 367–394. doi: 10.1016/B978-0-12-384984-7.00010-5
Langridge, D. F., and McGhee, R. B. (1967). Crithidia mellificae n. sp. an acidophilic trypanosomatid of the honey bee Apis mellifera. J. Protozool. 14, 485–487. doi: 10.1111/j.1550-7408.1967.tb02033.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ma, M., Huang, Y., Gong, Z., Zhuang, L., Li, C., Yang, H., et al. (2011). Discovery of DNA viruses in wild-caught mosquitoes using small RNA high throughput sequencing. PLoS ONE 6:e24758. doi: 10.1371/journal.pone.0024758
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Maori, E., Lavi, S., Mozes-Koch, R., Gantman, Y., Peretz, Y., Edelbaum, O., et al. (2007). Isolation and characterization of Israeli acute paralysis virus, a dicistrovirus affecting honeybees in Israel: evidence for diversity due to intra- and interspecies recombination. J. Gen. Virol. 88, 3428–3438. doi: 10.1099/vir.0.83284-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Martinson, V. G., Magoc, T., Koch, H., Salzberg, S. L., and Moran, N. A. (2014). Genomic features of a bumble bee symbiont reflect its host environment. Appl. Environ. Microbiol. 80, 3793–3803. doi: 10.1128/AEM.00322-14
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Merzlyak, E., Yurchenko, V., Kolesnikov, A. A., Alexandrov, K., Podlipaev, S. A., and Maslov, D. A. (2001). Diversity and phylogeny of insect trypanosomatids based on small subunit rRNA genes: polyphyly of Leptomonas and Blastocrithidia. J. Eukaryot. Microbiol. 48, 161–169. doi: 10.1111/j.1550-7408.2001.tb00298.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Minoche, A. E., Dohm, J. C., and Himmelbauer, H. (2011). Evaluation of genomic high-throughput sequencing data generated on Illumina HiSeq and genome analyzer systems. Genome Biol. 12, R112. doi: 10.1186/gb-2011-12-11-r112
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Mohr, K. I., and Tebbe, C. C. (2006). Diversity and phylotype consistency of bacteria in the guts of three bee species (Apoidea) at an oilseed rape field. Environ. Microbiol. 8, 258–272. doi: 10.1111/j.1462-2920.2005.00893.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Moran, N. A., Hansen, A. K., Powell, J. E., and Sabree, Z. L. (2012). Distinctive gut microbiota of honey bees assessed using deep sampling from individual worker bees. PLoS ONE 7:e36393. doi: 10.1371/journal.pone.0036393
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Morimoto, T., Kojima, Y., Yoshiyama, M., Kimura, K., Yang, B., Peng, G., et al. (2013). Molecular detection of protozoan parasites infecting Apis mellifera colonies in Japan. Environ. Microbiol. Rep. 5, 74–77. doi: 10.1111/j.1758-2229.2012.00385.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Mouches, C., Bove, J., Tully, J., Rose, D., McCoy, R., Carle-Junca, P., et al. (1983). Spiroplasma apis, a new species from the honey-bee (Apis mellifera). Ann. Inst. Pasteur. Microbiol. 134, 383–397.
Muz, M. N., Girisgin, A. O., Muz, D., and Aydin, L. (2010). Molecular detection of Nosema ceranae and Nosema apis infections in Turkish apiaries with collapsed colonies. J. Apic. Res. 49, 342. doi: 10.3896/IBRA.1.49.4.09
Naug, D. (2009). Nutritional stress due to habitat loss may explain recent honey bee colony collapses. Biol. Conserv. 142, 2369–2372. doi: 10.1016/j.biocon.2009.04.007
Nielsen, S. L., Nicolaisen, M., and Kryger, P. (2008). Incidence of acute bee paralysis virus, black queen cell virus, chronic bee paralysis virus, deformed wing virus, Kashmir bee virus and sacbrood virus in honey bees (Apis mellifera) in Denmark. Apidologie 39, 310–314. doi: 10.1051/apido:2008007
Nordstrom, S., Fries, A., Aarhus, H., Hansen, H., and Korpela, S. (1999). Virus infections in Nordic honey bee colonies with no, low or severe Varroa jacobsoni infestation. Apidologie 30, 457–466. doi: 10.1051/apido:19990602
Olofsson, T. C., and Vasquez, A. (2008). Detection and identification of a novel lactic acid bacterial flora within the honey stomach of the honeybee Apis mellifera. Curr. Microbiol. 57, 356–363. doi: 10.1007/s00284-008-9202-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Orantes-Bermejo, F. (1999). A scientific note on the prevalence of trypanosomatid parasites of honeybees (Apis mellifera L.) in southern Spain. Apidologie 30, 77–78. doi: 10.1051/apido:19990110
Ozkirim, A., and Schiesser, A. (2013). Israeil acute paralysis virus (IAPV) in Turkish bees. J. Apic. Res. 52, 56–57. doi: 10.3896/IBRA.1.52.2.09
Palacios, G., Hui, J., Quan, P. L., Kalkstein, A., Honkavuori, K. S., Busetti, A. V., et al. (2008). Genetic analysis of Israel acute paralysis virus: distinct clusters are circulating in the United States. J. Virol. 82, 6209–6217. doi: 10.1128/JVI.00251-08
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Paxton, R. J., Klee, J., Korpela, S., and Fries, I. (2007). Nosema ceranae has infected Apis mellifera in Europe since at least 1998 and may be more virulent than Nosema apis. Apidologie 38, 558–565. doi: 10.1051/apido:2007037
Rath, W., Boecking, O., and Drescher, W. (1995). The phenomena of simultaneous infestation of Apis mellifera in Asia with the parasitic mites Varroa jacobsoni Oud. and Tropilaelaps clareae Delfinado-Baker. Am. Bee J. 135, 125–127
Ravoet, J., Maharramov, J., Meeus, I., De Smet, L., Wenseleers, T., Smagghe, G., et al. (2013). Comprehensive bee pathogen screening in Belgium reveals Crithidia melli?cae as a new contributory factor to winter mortality. PLoS ONE 8:e72443. doi: 10.1371/journal.pone.0072443
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Round, J. L., and Mazmanian, S. K. (2009). The gut microbiota shapes intestinal immune responses during health and disease. Nat. Rev. Immunol. 9, 313–323. doi: 10.1038/nri2515
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Runckel, C., Flenniken, M. L., Engel, J. C., Ruby, J. G., Ganem, D., Andino, R., et al. (2011). Temporal analysis of the honey bee microbiome reveals four novel viruses and seasonal prevalence of known viruses, Nosema, and Crithidia. PLoS ONE 6:e20656. doi: 10.1371/journal.pone.0020656
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schmid-Hempel, R., and Tognazzo, M. (2010). Molecular divergence defines two distinct lineages of Crithidia bombi (Trypanosomatidae), parasites of bumblebees. J. Eukaryot. Microbiol. 57, 337–345. doi: 10.1111/j.1550-7408.2010.00480.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schwarz, R. S., Bauchan, G. R., Murphy, C. A., Ravoet, J., de Graaf, D. C., and Evans, J. D. (2015). Characterization of two species of trypanosomatidae from the honey bee Apis mellifera: Crithidia mellificae Langridge and McGhee, 1967 and Lotmaria passim n. gen., n. sp. J. Euk. Microbiol. doi: 10.1111/jeu.12209. [Epub ahead of print].
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schwarz, R. S., and Evans, J. D. (2013). Single and mixed-species trypanosome and microsporidia infections elicit distinct, ephemeral cellular and humoral immune responses in honey bees. Dev. Comp. Immunol. 40, 300–310. doi: 10.1016/j.dci.2013.03.010
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schwarz, R. S., Teixeira, E. W., Tauber, J. P., Birke, J. M., Martins, M., Fonseca, M., et al. (2014). Honey bee colonies act as reservoirs for two symbiotic Spiroplasma species and incur complex, multiyear infection dynamics. Microbiologyopen 3, 341–355. doi: 10.1002/mbo3.172
Spivak, M., and Reuter, G. S. (2001). Resistance to American foulbrood disease by honey bee colonies, Apis mellifera, bred for hygienic behavior. Apidologie 32, 555–565. doi: 10.1051/apido:2001103
Teixeira, E. W., Chen, Y., Message, D., Pettis, J., and Evans, J. D. (2008). Virus infections in Brazilian honey bees. J. Invertebr. Pathol. 99, 117–119. doi: 10.1016/j.jip.2008.03.014
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Tentcheva, D., Gauthier, L., Zappulla, N., Dainat, B., Cousserans, F., Colin, M. E., et al. (2004). Prevalence and seasonal variations of six bee viruses in Apis mellifera L. and Varroa destructor mite populations in France. Appl. Environ. Microbiol. 70, 7185–7191. doi: 10.1128/AEM.70.12.7185-7191.2004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Valera, F., Martín-Hernández, R., and Higes, M. (2011). Evaluation of large-scale dissemination of Nosema ceranae spores by European bee-eaters Merops apiaster. Environ. Microbiol. Rep. 3, 47–53. doi: 10.1111/j.1758-2229.2010.00186.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
vanEngelsdorp, D., Evans, J. D., Saegerman, C., Mullin, C., Haubruge, E., Nguyen, B. K., et al. (2009). Colony collapse disorder: a descriptive study. PLoS ONE 3:e6481. doi: 10.1371/journal.pone.0006481
vanEngelsdorp, D., and Meixner, M. D. (2010). A historical review of managed honey bee populations in Europe and the United States and the factors that may affect them. J. Invertebr. Pathol. 103, 80–95. doi: 10.1016/j.jip.2009.06.011
Vayssier-Taussat, M., Moutailler, S., Michelet, L., Devillers, E., Bonnet, S., Cheval, J., et al. (2013). Next generation sequencing uncovers unexpected bacterial pathogens in ticks in western Europe. PLoS ONE 8:e81439. doi: 10.1371/journal.pone.0081439
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ward, L., Waite, R., Boonham, N., Fisher, T., Pescod, K., Thompson, H., et al. (2007). First detection of Kashmir bee virus in the UK using real-time PCR. Apidologie 38, 181–190. doi: 10.1051/apido:2006072
Welch, A., Drummond, F., Tewari, S., Averill, A., and Burand, J. P. (2009). Presence and prevalence of viruses in local and migratory honeybees (Apis mellifera) in Massachusetts. Appl. Environ. Microbiol. 75, 7862–7865. doi: 10.1128/AEM.01319-09
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Whitaker, J., Szalanski, A. L., and Kence, M. (2010). Molecular detection of Nosema ceranae and N. apis from Turkish honey bees. Apidologie 42, 174–180. doi: 10.1051/apido/2010045
Williamson, D. L., Tully, J. G., and Whitcomb, R. F. (1989). “The genus Spiroplasma,” in The Mycoplasma V: Spiroplasmas, Acholeplasmas, and Mycoplasmas of Plants and Arthropods, eds R. F. Whitcomb and J. G. Tully (San Diego, CA: Academic Press), 71–111. doi: 10.1016/B978-0-12-078405-9.50011-2
Yang, B., Peng, G. D., Li, T. B., and Kadowaki, T. (2013). Molecular and phylogenetic characterization of honey bee viruses, Nosema microsporidia, protozoan parasites, and parasitic mites in China. Ecol. Evol. 3, 298–311. doi: 10.1002/ece3.464
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Yoshiyama, M., and Kimura, K. (2011). Distribution of Nosema ceranae in the European honeybee, Apis mellifera in Japan. J. Invertebr. Pathol. 106, 263–267. doi: 10.1016/j.jip.2010.10.010
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: Apis mellifera, pollination, colony collapse disorder, RNA sequencing, bioinformatics, honey bee viruses, trypanosomes
Citation: Tozkar CÖ, Kence M, Kence A, Huang Q and Evans JD (2015) Metatranscriptomic analyses of honey bee colonies. Front. Genet. 6:100. doi: 10.3389/fgene.2015.00100
Received: 16 November 2014; Accepted: 25 February 2015;
Published: 19 March 2015.
Edited by:
Juergen Rudolf Gadau, Arizona State University, USAReviewed by:
Marcial Escudero, Doñana Biological Station–Consejo Superior de Investigaciones Científicas, SpainRobert Brucker, Harvard University, USA
Copyright © 2015 Tozkar, Kence, Kence, Huang and Evans. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Cansu Ö. Tozkar, Ecological Genetics Laboratory, Department of Biological Sciences, Middle East Technical University, Dumlupınar Bulvarı No: 1, Cankaya, Ankara 06800, TurkeydG96a2FyQG1ldHUuZWR1LnRy;
Jay D. Evans, Bee Research Laboratory, USDA-ARS, 10300 Baltimore Avenue Bldg. 306 BARC-E, Beltsville, MD 20705-0000, USAamF5LmV2YW5zQGFycy51c2RhLmdvdg==
†Deceased.
 Cansu Ö. Tozkar
Cansu Ö. Tozkar Meral Kence
Meral Kence Aykut Kence1†
Aykut Kence1† Qiang Huang
Qiang Huang Jay D. Evans
Jay D. Evans