- 1Department of Biology, Indiana State University, Terre Haute, IN, USA
- 2Department of Biochemistry and Molecular Medicine, The George Washington University School of Medicine and Health Science, Washington, DC, USA
Due to recent advances of bioinformatics and high throughput sequencing technology, discovery of regulatory non-coding RNAs in bacteria has been increased to a great extent. Based on this bandwagon, many studies searching for trans-acting small non-coding RNAs in streptococci have been performed intensively, especially in the important human pathogen, group A and B streptococci. However, studies for cis-encoded non-coding antisense RNAs in streptococci have been scarce. A recent study shows antisense RNAs are involved in virulence gene regulation in group B streptococcus, S. agalactiae. This suggests antisense RNAs could have important roles in the pathogenesis of streptococcal pathogens. In this review, we describe recent discoveries of chromosomal cis-encoded antisense RNAs in streptococcal pathogens and other low GC Gram (+) bacteria to provide a guide for future studies.
Introduction
Non-coding regulatory RNAs exist in all three kingdoms and confer another layer of regulation mechanism for gene expression. Generally, the regulation by non-coding RNAs occurs at a post-transcriptional level, so their regulation would be fast and effective. Bacteria produce three general groups of non-coding regulatory RNAs: (i) cis-acting 5′ element non-coding RNAs, (ii) trans-acting small non-coding RNAs, and (iii) cis-encoded antisense RNAs. A cis-acting 5′ non-coding RNA is usually attached to the 5′ side of an mRNA whose expression is regulated by the non-coding RNA. A structural change of the non-coding RNA occurs by binding to small metabolites (riboswitches), or by change of temperature (thermoregulators) or pH (pH sensors). The structural change influences transcription or translation of the downstream gene or genes in an operon. Trans-acting small non-coding RNAs are usually encoded in intergenic regions on the chromosome and control translation or degradation of their target mRNAs. Generally, each trans-acting non-coding RNA has multiple target mRNAs and binds near the ribosomal binding site of the target mRNAs. A cis-acting antisense RNA (antisense RNA) is expressed as a complementary sequence of an mRNA that becomes the sole target RNA.
Previously, these non-coding RNAs had been discovered by computational predictions coupled with expression studies, microarrays, sequencing of small sized cDNA libraries, and high throughput sequencing approaches. Due to recent technological advances of tiling microarray, RNA deep sequencing, and bioinformatics, the search for non-coding regulatory RNAs on a genome-wide scale has been actively performed. As a result, the functions and regulatory mechanisms of discovered non-coding regulatory RNAs are widely studied. However, because of technical difficulties to distinguish the source of expressed RNAs between the two DNA strands, the search for antisense RNAs using high throughput methods has been retarded, compared to the search for trans-acting small RNAs. This makes antisense RNAs the least studied non-coding RNAs in streptococci to date. Currently no systematic search for antisense RNAs has been done in S. pyogenes, and only one search has been performed in S. agalactiae.
Considerable antisense transcription has been discovered in both eukaryotes and prokaryotes. The number of cis-encoded antisense RNAs in bacteria was once considered much smaller than that of eukaryotes due to the compact organization of protein-coding genes in the chromosome. However, recent studies indicate bacteria also produce a number of cis-encoded antisense RNAs. Bacterial cis-encoded antisense RNAs were discovered several decades ago, and most antisense RNAs were expressed from mobile genetic elements such as plasmids, phages, and transposons (Brantl, 2007). Since antisense RNAs expressed from bacterial chromosomes had not been discovered, it was thought that antisense RNAs were not generally used to control chromosomal gene expression in bacteria. However, during recent decades, many RNAs antisense to chromosomal genes have been discovered in bacteria. The other kingdom of prokaryotic microorganisms, archaea, also express cis-encoded antisense transcripts. An archaeal organism, Sulfolobus solfataricus P2, expresses about 310 non-coding RNAs and among these non-coding RNAs, almost 60% (185 non-coding RNAs) are cis-encoded antisense RNAs (Wurtzel et al., 2010). Although many antisense RNAs have been discovered in prokaryotes recently, their functions and regulation mechanisms are largely not studied.
Most cis-encoded antisense RNAs are complementary to a small portion of an open reading frame (ORF) and often the complementary portion includes the ribosome-binding site (Figure 1A). These small antisense RNAs are widely expressed on the chromosomes, plasmids, and transposons. However, some antisense RNAs are longer than typical ones and even reach several kilobases. Long antisense RNAs can be complementary to an entire gene or genes (Figure 1B). Among long antisense RNAs, some contains the sequence of a neighboring ORF or ORFs on their 5′ or 3′ side (Figure 1C). These type of antisense RNAs, which were named excludons (Sesto et al., 2013), have been discovered only on the chromosomes of several bacteria such as Listeria monocytogenes (Toledo-Arana et al., 2009), Bacillus subtilis (Rasmussen et al., 2009), a cyanobacterium Synechocystis sp. (Stazic et al., 2011), and Staphylococcus aureus (Beaume et al., 2010). However, as more bacteria are searched for antisense RNAs, more excludons are expected to be discovered.
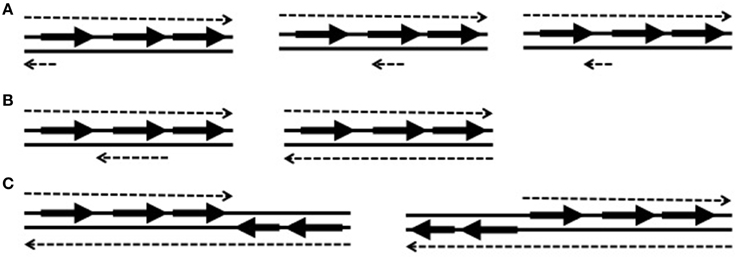
Figure 1. Examples of antisense RNA types illustrated with a three-gene operon. The solid lines depict double stranded DNA with genes (arrows). Each dotted line represents expressed RNA matching with the sequence of each DNA strand. The top dotted lines are mRNAs and the bottom dotted lines are cis-encoded antisense RNAs. (A) Small antisense RNAs complementary to the sequence of ribosome-binding site (RBS), in the middle of a gene, or of an intergenic region. (B) Long antisense RNAs complementary to an entire gene or an operon. (C) Excludons containing genes at its 5′ or 3′ side.
Cis-Encoded Antisense RNAs in Streptococci and Other Low GC Gram (+) Bacteria
S. agalactiae (Group B Streptococcus, GBS), which is an opportunistic pathogen and causative agent of bacterial sepsis, pneumonia, and meningitis in newborns, employs antisense RNAs to control virulence factors (Pichon et al., 2012). In the study of Pichon et al. they used an in silico method to find small non-coding RNAs and predicted the existence of 63 antisense RNAs (Table 1). They validated the existence of these antisense RNAs by verifying three of them through northern blotting (Table 2). The three RNAs, which have the sizes of 123 bps, 239 bps, and 243 bps, are fully or partially antisense to coding sequences (CDSs) involved in the pathogenicity of S. agalactiae. When they overexpressed two of these antisense RNAs using a multi-copy plasmid, one reduced the expression of the adjacent target gene but the other increased the expression of its target gene. This shows that antisense RNAs can carry out both negative and positive regulation.
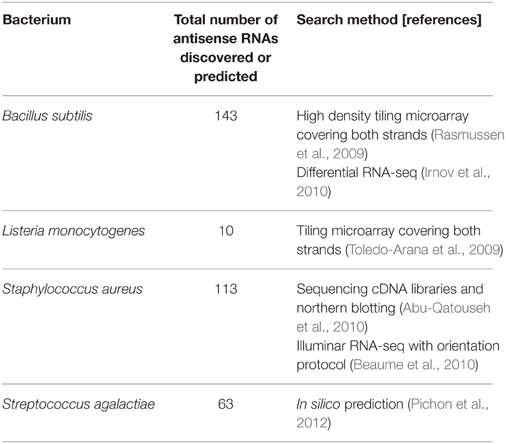
Table 1. High throughput searches for chromosomal cis-encoded antisense RNAs in low GC Gram-positive bacteria.
On the other hand, the discovery of antisense RNAs in another important streptococcal pathogen Streptococcus pyogenes (Group A Streptococcus, GAS) has not been reported. Many studies have been done to search for trans-acting small non-coding RNAs, but no systematic study has been done so far to search for antisense RNAs. Thus, it is not known if antisense RNAs in this pathogen have an important role in controlling gene expression and/or virulence.
An RNA-based toxin-antitoxin system was discovered on the chromosome of Streptococcus mutans, an oral streptococcal pathogen (Table 2) (Koyanagi and Levesque, 2013). This is an unusual case because most toxin-antitoxin systems in bacteria are encoded in plasmids. The S. mutans antitoxin is an antisense RNA (srSm) converging toward the end of the gene of Fst-like toxin (Fst-Sm), so the expression of the antitoxin antisense RNA inhibits the production of Fst-like toxin.
High throughput searches for non-coding regulatory RNAs in Bacillus subtilis have been performed to gain more knowledge on the regulation of gene expression by non-coding RNAs in this low GC Gram (+) model organism (Rasmussen et al., 2009; Irnov et al., 2010). In these searches, Rasmussen et al. discovered 127 antisense RNAs through a high density tiling array (Rasmussen et al., 2009), and then Irnov et al. discovered 16 novel antisense RNAs using a differential RNA-seq analysis (Table 1) (Irnov et al., 2010). The results from these studies reveal that target genes of antisense RNAs are involved in stress response, sporulation, and expression of SigA, the principal sigma factor during vegetative growth (Table 2). Therefore, antisense RNAs in B. subtilis appear to influence a variety of important regulations to adapt diverse environmental conditions.
Staphylococcus aureus is a remarkable opportunistic pathogen causing a broad spectrum of diseases like S. pyogenes, which range from superficial skin diseases to fatal systemic infections including sepsis, pneumonia, and bone infections. Since the emergence and spread of drug-resistant and community-acquired strains, S. aureus infections have drawn great attention. The most intensively studied non-coding RNA in S. aureus is RNAIII that is a regulatory RNA controlling many virulence factors as the effector of the agr quorum sensing system. Even though RNAIII controls translation and degradation of target mRNAs with an antisense mechanism, its action is trans, not cis, thus RNAIII is not discussed here because of the narrow scope of this review (for a review on RNAIII, see Novick and Geisinger, 2008).
Previously, several studies have been performed to discover non-coding regulatory RNAs in S. aureus through computational methods, sequencing of small sized cDNAs, and high throughput strand-specific RNA sequencing technology (Table 1) (Pichon and Felden, 2005; Geissmann et al., 2009; Abu-Qatouseh et al., 2010; Beaume et al., 2010; Bohn et al., 2010). From these studies, about 100 cis-encoded antisense RNAs have been discovered, some of which were experimentally detected by northern blotting, Rapid Amplification of cDNA Ends (RACE) mapping, or reverse transcriptase quantitative PCR (RT-qPCR) (Table 2). Many of these antisense RNAs are expressed from pathogenicity islands and mobile elements such as plasmids and transposons. Interestingly, existence of some antisense RNAs was unique in a strain, suggesting that gene regulation by cis-encoded antisense RNA could be strain specific. Long antisense RNAs are also present in S. aureus. The antisense RNA complementary to the gene encoding a secretory antigen (SA0620) is bigger than 1 kb (Beaume et al., 2010).
In the study by Beaume et al., 10 cis-encoded antisense RNAs out of total discovered 35 were expressed in pathogenicity islands or in the chromosome mec cassette, which is a mobile genetic element conferring methicillin resistance (Beaume et al., 2010). This indicates that antisense RNAs could play a key role in S. aureus infections. These antisense RNAs are particularly abundant in genes involved in cell wall and cell envelope biogenesis and in replication, recombination, and repair. Interestingly, two of these antisense RNAs are complementary to the small non-coding RNAs, SprA1, and AprG. These two antisense RNA-small non-coding RNA pairs are predicted to form type I toxin-antitoxin modules. The study of S. aureus small colony variants identified 78 antisense RNA candidates (Abu-Qatouseh et al., 2010). Some antisense RNAs in S. aureus are involved in the differential expression of genes in the same operon. An example is antisense RNAs complementary to a part of each capF and capM transcript of the same capsular polysaccharide synthesis operon (cap operon) (Abu-Qatouseh et al., 2010; Beaume et al., 2010). Even though they are expressed as one mRNA, the two genes are differentially translated by the antisense RNAs.
Listeria monocytogenes is a Gram (+) pathogenic bacterium causing food-borne infection, listeriosis, which can lead to meningitis in newborns. This pathogen has a well-defined virulence mechanism to inhibit phagolysosome formation and proliferate inside host cells, so has been extensively used as a model organism for the study of pathogen-host interaction (Hamon et al., 2006). Previously, the Cossart group examined the transcription profile of this pathogen using tiling microarrays that covered both strands of the chromosome, and discovered many non-coding RNAs including 10 cis-encoded antisense RNAs (Table 1). Three of them were already classified as small RNAs and seven were newly discovered (Toledo-Arana et al., 2009). Most cis-encoded antisense RNAs cover a small portion of an open reading frame (ORF), but three antisense RNAs are large enough to cover more than one ORF. Interestingly, all of these long antisense RNAs are expressed with a shorter antisense RNA. Both shorter and longer antisense RNAs are expressed at the same start site but they have different termination sites. The importance of these two different size antisense transcripts has not been determined yet.
Regulation Mechanisms by Cis-Encoded Antisense RNAs
Antisense RNAs can control gene expression by binding to their cognate sense RNAs. The binding occurs at the 5′ end, 3′ end, or in the middle of mRNAs depending on the location they are expressed (Figure 1A). Also, long antisense RNAs can overlap an entire mRNA encoding a protein or proteins (Figure 1B). The different binding locations confer different control mechanisms. Based on their binding locations on sense RNAs, antisense RNAs may act in three ways: (i) transcription terminators in the mechanism of transcription attenuation or transcription interference, (ii) potential inhibitors of translation initiation, or (iii) modulators of mRNA degradation. Antisense RNAs influence gene expression at the transcriptional or post-transcriptional level. Transcription interference and transcription attenuation occur at the transcriptional levels, and translation inhibition and mRNA degradation occur at post-transcriptional levels. The degree of control by antisense RNAs can be achieved by their differential expression level at different conditions. The expression ratio between a sense RNA and the antisense RNA will influence the expression of the sense gene.
In transcription interference, two promoters of an antisense RNA and its target sense RNA present very close in cis-position and their transcriptions occur in the convergent direction, and then the transcription rate from one promoter becomes suppressed by the other promoter (Callen et al., 2004). In this case, the transcription of the weaker promoter seems suppressed more. Another regulation mechanism at the transcriptional level by antisense RNAs is transcription attenuation. In transcription attenuation, an antisense RNA binds to the region in front of the Shine-Dalgano sequence of the target mRNA, and this binding induces the formation of transcription terminator structure. Hence, when the antisense RNA binds near or at the 5′ end of the cognate sense RNA, the transcription of the sense RNA is terminated (Brantl, 2002; Stork et al., 2007). In this regulation, if an antisense RNA binds an intergenic region in a polycistronic mRNA, then it can create differential gene expression between the genes located upstream and downstream of the intergenic region, and the upstream gene is more expressed than the downstream gene (Stork et al., 2007).
A common post-transcriptional level regulation by antisense RNAs is modulating translation resulting in translation inhibition or activation. In translation inhibition, antisense RNAs bind directly to the Shine-Dalgano sequence (SD sequence) of mRNAs, and inhibit ribosome-binding (Greenfield et al., 2001; Hernandez et al., 2006; Kawano et al., 2007). This inhibition of translation might increase or decrease the degradation of mRNAs by ribonuclease. In translation activation, an antisense RNA bind near the SD sequence whose access by ribosomes are blocked by a preformed stem and loop structure, then the binding of the antisense RNA frees the SD sequence (Asano et al., 1998).
As mentioned, mRNA degradation can be influenced by a bound antisense RNA. The pairs of antisense RNA–target mRNA can be substrates of RNase III, which is a double strand specific endoribonuclease. RNase III is conserved in all the three kingdoms. A previous study of S. aureus showed that the deletion of RNase III increased the amount of antisense transcripts, indicating that target mRNAs bound by antisense RNAs are degraded by RNase III in vivo (Lasa et al., 2011). Deep sequencing analysis in the same study showed that RNase III generates 22 nt long RNA fragments with 2 nucleotide 3′ overhang from the pairs of sense-antisense transcripts. Surprisingly, 75% of mRNAs are processed by RNase III, implying that antisense regulation occurs more extensively than previously thought. Studies on other bacteria also indicate that antisense transcription occurs extensively throughout the chromosome (For a review, see Georg and Hess, 2011).
Another RNase shown to be involved in degradation of sense-antisense RNA pairs is RNase E, an endoribonuclease degrading 5′ monophosphorylated mRNAs. RNase E degrades mgtC mRNA in Salmonella enterica with an unknown mechanism when the sense RNA is bound by the antisense RNA, AmgR (Lee and Groisman, 2010). RNase E is a member of the RNA degradosome in Gram (−) bacteria, a multicomponent complex that also includes an RNA helicase, RhlB, a glycolytic enzyme, enolase, and the exoribonuclease polynucleotide phosphorylase (PNPase) (Carpousis, 2007). The main function of the RNA degradosome is known to control mRNA turnover. Most Gram (+) bacteria including streptococci, bacilli, and staphylococci do not possess an RNase E homolog. However, these bacteria possess the RNA degradosome. The Gram (+) RNA degradosome contains similar kinds of components but more members, compared to the Gram (−) counterpart: four ribonuclases, RNase Y, RNase J1, J2, and PNPase; an RNA helicase, CshA; two glycolytic enzymes, phosphofructokinase (PfkA) and enolase (Lehnik-Habrink et al., 2012). RNase E is a membrane bound protein providing the major structural scaffold interacting with other components in the Gram (−) degradosome. The structure of Gram (+) RNA degradosome has not been resolved, but protein interaction studies revealed that the endoribonuclease RNase Y, a membrane anchored protein, interacts with most other components in the degradosome, so RNase Y might be the functional homolog of RNase E (Kang et al., 2010). No study has been done yet if RNase Y is also involved in the degradation of some sense-antisense RNA pairs in Gram (+) bacteria.
In Gram (−) bacteria, most small non-coding regulatory RNAs work with the RNA chaperone protein Hfq. Generally, the presence of the Hfq protein increases the stability of small non-coding RNAs and facilitates the interaction to their target mRNAs (Gottesman and Storz, 2011). However, the role of Hfq does not seem critical in Gram (+) bacteria. The role of Hfq is dispensable in S. aureus (Bohn et al., 2007). There have not been many studies of Hfq in terms of cis-encoded antisense RNAs so far, but previous studies show that some antisense RNAs interact with Hfq (Sittka et al., 2008; Lorenz et al., 2010), and Hfq is required for the function of a cis-encoded antisense RNA (Ross et al., 2010). Streptococci and lactobacilli do not possess any Hfq homologs, and it has not been studied if some other protein or proteins replace the role of Hfq in trans-acting small RNA- or cis-acting antisense RNA-mediated regulation. It has been suggested that the role of Hfq might be dispensable in low GC Gram (+) bacteria because non-coding RNAs in these bacteria are longer than higher GC Gram (−) bacteria to compensate for the low GC content of the pairings (Jousselin et al., 2009).
One advantage of regulation by antisense RNAs is to confer an additional layer of gene regulation like other non-coding regulatory RNAs. In concert with protein regulators, antisense RNAs can provide more precise regulation or regulation responding to different signals. Compared to trans-acting small non-coding RNAs, the regulation by antisense RNAs are generally more specific. Usually trans-acting non-coding small RNAs have multiple target mRNAs with imperfect base-pairs, but antisense RNAs usually have just one target mRNA with the complete complementary sequence. Even though we cannot completely rule out the possibility that some antisense RNAs have several targets with partial base matches by acting in trans, multiple targets of an antisense RNA have not been discovered yet. Another advantage of regulation by cis-encoded antisense RNAs is regulation speed. Like other non-coding regulatory RNAs, most antisense RNAs act at the post-transcriptional level, so the result of the action by antisense RNAs would be faster than protein transcriptional regulators.
Perspectives
Compared to small non-coding trans-acting RNAs, bacterial cis-encoded antisense RNAs had not been studied in the genome-wide scale because of technical difficulties. However, due to the recent development of strand specific RNA sequencing and tiling microarrays covering both strands, cis-encoded antisense RNAs have been subjected under the genome-wide search in many bacteria. Already hundreds of bacterial antisense RNAs have been discovered and changed the concept of regulation by antisense RNAs. So far few streptococcal antisense RNAs have been discovered, but further genome-wide search would definitely find a number of antisense RNAs in this group of bacteria and promote studies to investigate the function and molecular mechanism of regulation by antisense RNAs.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study was supported in part by American Heart Association Grant 11SDG7440083 and National Institutes of Health Grant R15GM101603-01 to KHC.
References
Abu-Qatouseh, L. F., Chinni, S. V., Seggewiss, J., Proctor, R. A., Brosius, J., Rozhdestvensky, T. S., et al. (2010). Identification of differentially expressed small non-protein-coding RNAs in Staphylococcus aureus displaying both the normal and the small-colony variant phenotype. J. Mol. Med. 88, 565–575. doi: 10.1007/s00109-010-0597-2
Asano, K., Niimi, T., Yokoyama, S., and Mizobuchi, K. (1998). Structural basis for binding of the plasmid ColIb-P9 antisense Inc RNA to its target RNA with the 5′-rUUGGCG-3′ motif in the loop sequence. J. Biol. Chem. 273, 11826–11838. doi: 10.1074/jbc.273.19.11826
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Beaume, M., Hernandez, D., Farinelli, L., Deluen, C., Linder, P., Gaspin, C., et al. (2010). Cartography of methicillin-resistant S. aureus transcripts: detection, orientation and temporal expression during growth phase and stress conditions. PLoS ONE 5:e10725. doi: 10.1371/journal.pone.0010725
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bohn, C., Rigoulay, C., and Bouloc, P. (2007). No detectable effect of RNA-binding protein Hfq absence in Staphylococcus aureus. BMC Microbiol. 7:10. doi: 10.1186/1471-2180-7-10
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bohn, C., Rigoulay, C., Chabelskaya, S., Sharma, C. M., Marchais, A., Skorski, P., et al. (2010). Experimental discovery of small RNAs in Staphylococcus aureus reveals a riboregulator of central metabolism. Nucleic Acids Res. 38, 6620–6636. doi: 10.1093/nar/gkq462
Brantl, S. (2002). Antisense-RNA regulation and RNA interference. Biochim. Biophys. Acta 1575, 15–25. doi: 10.1016/S0167-4781(02)00280-4
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Brantl, S. (2007). Regulatory mechanisms employed by cis-encoded antisense RNAs. Curr. Opin. Microbiol. 10, 102–109. doi: 10.1016/j.mib.2007.03.012
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Callen, B. P., Shearwin, K. E., and Egan, J. B. (2004). Transcriptional interference between convergent promoters caused by elongation over the promoter. Mol. Cell 14, 647–656. doi: 10.1016/j.molcel.2004.05.010
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Carpousis, A. J. (2007). The RNA degradosome of Escherichia coli: an mRNA-degrading machine assembled on RNase E. Annu. Rev. Microbiol. 61, 71–87. doi: 10.1146/annurev.micro.61.080706.093440
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Geissmann, T., Chevalier, C., Cros, M. J., Boisset, S., Fechter, P., Noirot, C., et al. (2009). A search for small noncoding RNAs in Staphylococcus aureus reveals a conserved sequence motif for regulation. Nucleic Acids Res. 37, 7239–7257. doi: 10.1093/nar/gkp668
Georg, J., and Hess, W. R. (2011). cis-antisense RNA, another level of gene regulation in bacteria. Microbiol. Mol. Biol. Rev. 75, 286–300. doi: 10.1128/MMBR.00032-10
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gottesman, S., and Storz, G. (2011). Bacterial small RNA regulators: versatile roles and rapidly evolving variations. Cold Spring Harb. Perspect. Biol. 3:a003798. doi: 10.1101/cshperspect.a003798
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Greenfield, T. J., Franch, T., Gerdes, K., and Weaver, K. E. (2001). Antisense RNA regulation of the par post-segregational killing system: structural analysis and mechanism of binding of the antisense RNA, RNAII and its target, RNAI. Mol. Microbiol. 42, 527–537. doi: 10.1046/j.1365-2958.2001.02663.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hamon, M., Bierne, H., and Cossart, P. (2006). Listeria monocytogenes: a multifaceted model. Nat. Rev. Microbiol. 4, 423–434. doi: 10.1038/nrmicro1413
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hernandez, J. A., Muro-Pastor, A. M., Flores, E., Bes, M. T., Peleato, M. L., and Fillat, M. F. (2006). Identification of a furA cis antisense RNA in the cyanobacterium Anabaena sp. PCC 7120. J. Mol. Biol. 355, 325–334. doi: 10.1016/j.jmb.2005.10.079
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Irnov, I., Sharma, C. M., Vogel, J., and Winkler, W. C. (2010). Identification of regulatory RNAs in Bacillus subtilis. Nucleic Acids Res. 38, 6637–6651. doi: 10.1093/nar/gkq454
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jousselin, A., Metzinger, L., and Felden, B. (2009). On the facultative requirement of the bacterial RNA chaperone, Hfq. Trends Microbiol. 17, 399–405. doi: 10.1016/j.tim.2009.06.003
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kang, S. O., Caparon, M. G., and Cho, K. H. (2010). Virulence gene regulation by CvfA, a putative RNase: the CvfA-enolase complex in Streptococcus pyogenes links nutritional stress, growth-phase control, and virulence gene expression. Infect. Immun. 78, 2754–2767. doi: 10.1128/IAI.01370-09
Kawano, M., Aravind, L., and Storz, G. (2007). An antisense RNA controls synthesis of an SOS-induced toxin evolved from an antitoxin. Mol. Microbiol. 64, 738–754. doi: 10.1111/j.1365-2958.2007.05688.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Koyanagi, S., and Levesque, C. M. (2013). Characterization of a Streptococcus mutans intergenic region containing a small toxic peptide and its cis-encoded antisense small RNA antitoxin. PLoS ONE 8:e54291. doi: 10.1371/journal.pone.0054291
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lasa, I., Toledo-Arana, A., Dobin, A., Villanueva, M., de los Mozos, I. R., Vergara-Irigaray, M., et al. (2011). Genome-wide antisense transcription drives mRNA processing in bacteria. Proc. Natl. Acad. Sci. U.S.A. 108, 20172–20177. doi: 10.1073/pnas.1113521108
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, E. J., and Groisman, E. A. (2010). An antisense RNA that governs the expression kinetics of a multifunctional virulence gene. Mol. Microbiol. 76, 1020–1033. doi: 10.1111/j.1365-2958.2010.07161.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lehnik-Habrink, M., Lewis, R. J., Mader, U., and Stulke, J. (2012). RNA degradation in Bacillus subtilis: an interplay of essential endo- and exoribonucleases. Mol. Microbiol. 84, 1005–1017. doi: 10.1111/j.1365-2958.2012.08072.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lorenz, C., Gesell, T., Zimmermann, B., Schoeberl, U., Bilusic, I., Rajkowitsch, L., et al. (2010). Genomic SELEX for Hfq-binding RNAs identifies genomic aptamers predominantly in antisense transcripts. Nucleic Acids Res. 38, 3794–3808. doi: 10.1093/nar/gkq032
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Novick, R. P., and Geisinger, E. (2008). Quorum sensing in staphylococci. Annu. Rev. Genet. 42, 541–564. doi: 10.1146/annurev.genet.42.110807.091640
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Pichon, C., du Merle, L., Caliot, M. E., Trieu-Cuot, P., and Le Bouguenec, C. (2012). An in silico model for identification of small RNAs in whole bacterial genomes: characterization of antisense RNAs in pathogenic Escherichia coli and Streptococcus agalactiae strains. Nucleic Acids Res. 40, 2846–2861. doi: 10.1093/nar/gkr1141
Pichon, C., and Felden, B. (2005). Small RNA genes expressed from Staphylococcus aureus genomic and pathogenicity islands with specific expression among pathogenic strains. Proc. Natl. Acad. Sci. U.S.A. 102, 14249–14254. doi: 10.1073/pnas.0503838102
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Rasmussen, S., Nielsen, H. B., and Jarmer, H. (2009). The transcriptionally active regions in the genome of Bacillus subtilis. Mol. Microbiol. 73, 1043–1057. doi: 10.1111/j.1365-2958.2009.06830.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ross, J. A., Wardle, S. J., and Haniford, D. B. (2010). Tn10/IS10 transposition is downregulated at the level of transposase expression by the RNA-binding protein Hfq. Mol. Microbiol. 78, 607–621. doi: 10.1111/j.1365-2958.2010.07359.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sesto, N., Wurtzel, O., Archambaud, C., Sorek, R., and Cossart, P. (2013). The excludon: a new concept in bacterial antisense RNA-mediated gene regulation. Nat. Rev. Microbiol. 11, 75–82. doi: 10.1038/nrmicro2934
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sittka, A., Lucchini, S., Papenfort, K., Sharma, C. M., Rolle, K., Binnewies, T. T., et al. (2008). Deep sequencing analysis of small noncoding RNA and mRNA targets of the global post-transcriptional regulator, Hfq. PLoS Genet. 4:e1000163. doi: 10.1371/journal.pgen.1000163
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Stazic, D., Lindell, D., and Steglich, C. (2011). Antisense RNA protects mRNA from RNase E degradation by RNA-RNA duplex formation during phage infection. Nucleic Acids Res. 39, 4890–4899. doi: 10.1093/nar/gkr037
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Stork, M., Di Lorenzo, M., Welch, T. J., and Crosa, J. H. (2007). Transcription termination within the iron transport-biosynthesis operon of Vibrio anguillarum requires an antisense RNA. J. Bacteriol. 189, 3479–3488. doi: 10.1128/JB.00619-06
Toledo-Arana, A., Dussurget, O., Nikitas, G., Sesto, N., Guet-Revillet, H., Balestrino, D., et al. (2009). The Listeria transcriptional landscape from saprophytism to virulence. Nature 459, 950–956. doi: 10.1038/nature08080
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wurtzel, O., Sapra, R., Chen, F., Zhu, Y., Simmons, B. A. and Sorek, R. (2010). A single-base resolution map of an archaeal transcriptome. Genome Res. 20, 133–141. doi: 10.1101/gr.100396.109
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: non-coding RNAs, regulatory RNAs, antisense RNAs, streptococci, Gram (+) pathogens
Citation: Cho KH and Kim J-H (2015) Cis-encoded non-coding antisense RNAs in streptococci and other low GC Gram (+) bacterial pathogens. Front. Genet. 6:110. doi: 10.3389/fgene.2015.00110
Received: 12 January 2015; Accepted: 03 March 2015;
Published: 26 March 2015.
Edited by:
Mohamed Amine Zorgani, School of Medicine, Université François Rabelais/INRA, FranceReviewed by:
Erin R. Murphy, Ohio University Heritage College of Osteopathic Medicine, USAClayton Caswell, Virginia Tech, USA
Copyright © 2015 Cho and Kim. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Kyu Hong Cho, Department of Biology, Indiana State University, 600 Chestnut Street S224, Terre Haute, IN 47809, USAa3l1aG9uZy5jaG9AaW5kc3RhdGUuZWR1
 Kyu Hong Cho
Kyu Hong Cho Jeong-Ho Kim2
Jeong-Ho Kim2