- 1Department of Integrative Biology and Physiology, University of California, Los Angeles, Los Angeles, CA, USA
- 2Molecular Biology Institute, University of California, Los Angeles, Los Angeles, CA, USA
- 3School of Dentistry, University of California, Los Angeles, Los Angeles, CA, USA
- 4Jonnson Comprehensive Cancer Center, University of California, Los Angeles, Los Angeles, CA, USA
Human saliva emerged as a research material in as early as the 17th century when investigators sought to understand the basis for salivary secretion (Garrett, 1975). Over the centuries, the focus of salivary research has evolved greatly and a wide range of topics has been examined (Garrett, 1975; Schipper et al., 2007). It is now known that the functions of saliva include at least lubrication, digestion of food, remineralization, prevention of demineralization, protection against microbial and viral infection, speech facilitation, and maintenance of oral and general health (Schipper et al., 2007; Malathi et al., 2014).
One active research area related to human saliva is the discovery of biomarker molecules for a variety of diseases. Compared to other body fluids, saliva is easily accessible in a non-invasive manner. However, it is also immediately exposed to the outside environment, thus may be confounded by a wide variety of environmental factors. Nevertheless, previous studies have established that at least some human molecules in saliva are highly stable and potential biomarkers have been examined for a number of oral and systemic diseases (Bonne and Wong, 2012; Schafer et al., 2014).
Key Components of Human Saliva
Saliva consists of both cellular and fluid contents. Epithelial cells, leukocytes, and erythrocytes are the three major human cell types, which co-exist with bacterial cells in human whole saliva (Aps et al., 2002). The fluid content of saliva is primarily generated by the salivary glands, but with additional contributions from blood, oral tissue, bacteria, viruses, and food remnants (Schipper et al., 2007). It mainly consists of water, macromolecules (such as glycoproteins, enzymes), small organic molecules, inorganic components (e.g., electrolytes), and metabolites from oral bacteria (Almstahl and Wikstrom, 2003; Aps and Martens, 2005; Schipper et al., 2007).
Many biomarker studies focused on profiling and quantification of proteins or RNA molecules in saliva. Since the diseases investigated for salivary biomarkers are often systemic, it is of great interest to identify circulating protein or RNA molecules that may have originated from disease-relevant cells (such as tumor cells). Such molecules reside outside of the cells in saliva and are often captured in cell-free saliva (CFS), the fraction of saliva with cellular contents removed (often by centrifugation). Most of the salivary RNAs appear to be highly degraded compared to full-length mRNAs in cellular compartments, possibly due to presence of RNA degradation enzymes in saliva and other body fluids (for circulating mRNAs) (Park et al., 2007). Notably, certain miRNA and mRNA molecules were shown to be highly stable, possible owing to protection by exosomes or protein complexes (Park et al., 2006, 2009; Palanisamy and Wong, 2010; Palanisamy et al., 2010).
Technologies for Salivary RNA Profiling
About a decade ago, microarrays were applied to characterize the global profile of mRNAs in saliva (Li et al., 2004; Park et al., 2007). These studies revealed that there were over one thousand distinct mRNA molecules in human CFS (Li et al., 2004). In addition to mRNAs derived from coding genes, many noncoding RNAs (ncRNAs) were also detected. Data from these studies demonstrate that there are hundreds of microRNAs (miRNAs) in human saliva, and most of them likely exist in exosomes (Michael et al., 2010; Gallo et al., 2012).
However, microarray techniques have inherent limitations, such as the dependence on gene annotation and cross-hybridization noise. In recent years, more powerful techniques based on next generation sequencing (NGS) revealed additional coding and ncRNA species in human saliva (Spielmann et al., 2012; Bahn et al., 2015). In contrast to the hybridization-based microarrays, RNA sequencing (RNA-Seq) offers single nucleotide information, high sensitivity and accuracy in transcript detection, and the capability to detect novel RNA species and transcript isoforms (Lee et al., 2011, 2013; Li et al., 2012). An increasing number of bioinformatic tools are emerging for analysis of RNA-Seq data, ranging from rapid short read aligners to detailed examination of RNA expression patterns (Oshlack et al., 2010). Owing to these improvements, the catalog of human genes, especially ncRNA genes, has been greatly expanded (refer to Sai Lakshmi and Agrawal, 2008; Kozomara and Griffiths-Jones, 2014; Xie et al., 2014 for ncRNA databases).
ncRNA Molecules in Saliva
In 2012, the Wong group reported the first global characterization of the human salivary transcriptome using high-throughput RNA-Seq (Spielmann et al., 2012). This study demonstrated that saliva harbors a wide variety of RNA species. More than 4000 distinct RNA molecules derived from coding or noncoding human RNAs were identified, including a small number of miRNAs. This study established that the RNA content in saliva is very diverse, which should be fully explored in future biomarker studies.
Recently, another in-depth analysis of human salivary extracellular ncRNA revealed novel insights regarding its RNA content and provided a comparative view of salivary ncRNAs relative to those of other body fluids (Bahn et al., 2015). Using human CFS, this study confirmed previous findings that miRNAs are stably and abundantly present in saliva (Patel et al., 2011), often harbored within exosomes (Gallo et al., 2012). miRNA expression profiles of healthy individuals were quantified and compared. Highly concordant miRNA expression was observed across individuals. Furthermore, considerable similarity was observed between miRNA expression levels of saliva and other body fluids (blood, cerebral spinal fluid (CSF)). Thus, these data suggest that salivary miRNAs could serve as candidate biomarkers, at least with equivalent promise as those derived from more invasive fluids.
A surprising observation from this study was the relative abundance of human piwi-interacting RNAs (piRNAs) in saliva. piRNAs are small ncRNAs typically ~26–32 nt in length observed in germ cells of both vertebrates and invertebrates (Aravin et al., 2006; Girard et al., 2006; Grivna et al., 2006; Lau et al., 2006; Watanabe et al., 2006; Das et al., 2008). piRNAs are known to target transposons and repress their mobility (Das et al., 2008; Malone and Hannon, 2009). The number of abundant piRNAs is less than that of miRNAs in saliva, despite the large number of annotated piRNAs in various databases (Sai Lakshmi and Agrawal, 2008; Bahn et al., 2015). Nevertheless, piRNA expression levels were highly concordant between healthy individuals, similarly as miRNA levels. However, in contrast to the consistent expression profile of miRNAs across body fluids, piRNAs were highly exclusive to saliva with very low abundance in blood or CSF. These observations indicate that salivary piRNAs may have originated from cells in the oral mucosa or salivary glands, rather than circulating from systemic organs via blood. Nevertheless, salivary piRNAs may impose systemic functional impact, which needs to be further investigated.
Another novel finding in this study was the discovery of circular RNAs (circRNAs) in CFS, which is the first report of the presence of circRNAs in an extracellular fluid (Bahn et al., 2015). CircRNAs were originally identified in RNA viruses (Sanger et al., 1976; Kos et al., 1986). Later, intracellular circRNAs generated from specific exons of coding genes were reported (Nigro et al., 1991; Cocquerelle et al., 1992; Capel et al., 1993). Recent studies demonstrated that circRNAs exist in many different cell types and species (Salzman et al., 2012, 2013; Jeck et al., 2013; Memczak et al., 2013). Some circRNAs are likely noncoding (Capel et al., 1993; Memczak et al., 2013; Guo et al., 2014), but others may code for proteins (Wang and Wang, 2015). The function of most circRNAs remains unknown. Two circRNAs were shown to function as miRNA sponges (Hansen et al., 2013; Memczak et al., 2013). However, this function may not apply to the majority of other circRNAs as they lack bioinformatic evidence of significant miRNA complementarity (Guo et al., 2014). The discovery of circRNAs in CFS indicates that this type of molecule may have extracellular function and should be considered as a type of candidate biomarker (Li et al., 2015).
Although mRNAs are highly degraded in saliva and other body fluids, small ncRNAs are often stable with reproducible expression across individuals. Indeed, miRNAs have been extensively studied in blood and other body fluids as potential disease biomarkers (Chen et al., 2008; Gilad et al., 2008; Mitchell et al., 2008; Wang et al., 2009, 2010; Fichtlscherer et al., 2010; Li et al., 2010; Liu et al., 2011). The similarity between miRNA profiles of saliva and other body fluids (Weber et al., 2010; Bahn et al., 2015) strongly supports the potential of using miRNAs (and possibly other ncRNAs) from human CFS as biomarkers for various human diseases.
Salivary ncRNAs as Potential Biomarkers for Diseases
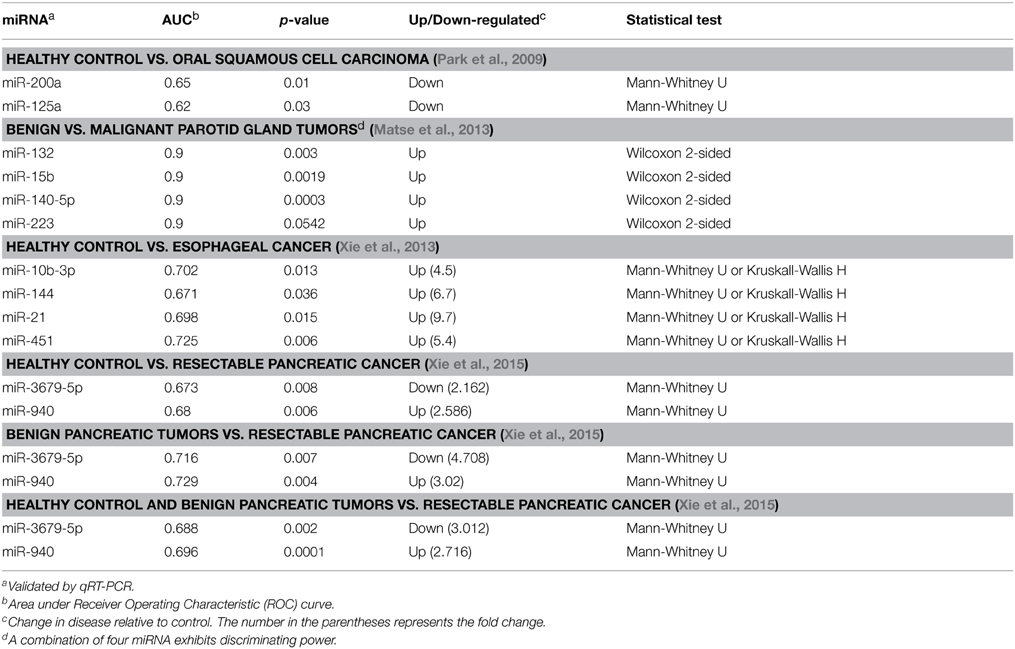
Although at an early stage, salivary ncRNA studies have revealed potential disease biomarkers. Thus, far, most studies focused on miRNA expression in saliva. Table 1 summarizes a number of studies where miRNAs were assessed as putative biomarkers for oral squamous cell carcinoma (OSCC) (Park et al., 2009), parotid gland tumors (Matse et al., 2013), and esophageal cancer (Xie et al., 2013). In addition to oral and esophageal diseases, salivary ncRNAs were also examined as potential biomarkers for systemic diseases. In a clinical study focusing on Sjögren's Syndrome, a chronic autoimmune disease, the authors observed different miRNA expression patterns in minor salivary glands of Sjögren's Syndrome patient compared to healthy individuals (Alevizos et al., 2011). The disease group can be clearly distinguished from the normal group using the miRNA expression profile by principal components and hierarchical clustering analyses. A very recent study focused on pancreatic cancer, using samples of patients with pancreatic cancer, benign pancreatic tumor or healthy controls (Xie et al., 2015). The authors observed significant down-regulation of miR-3679-5p and up-regulation of miR-940 in the cancer group compared to the other groups (Table 1), suggesting salivary miRNA may potentially be used for early detection of pancreatic cancer.
In addition to human ncRNAs, exogenous ncRNAs in saliva may also serve as potential disease biomarkers. The human-associated microbial communities have profound impact on the individual's physiological outcome (Human Microbiome Project Consortium, 2012). In human saliva, over 1500 bacteria have been identified and completely sequenced [Human Oral Microbiome Database; http://www.homd.org/]. Some studies have shown that saliva can be used to detect microbial infection (Schafer et al., 2014). In addition, both DNA and RNA viruses were detected in human saliva from viral infected hosts (Liou et al., 1992; Chen et al., 1997; Vieira et al., 1997; Shugars et al., 2001; Hermida et al., 2002; Mackiewicz et al., 2004; Goncalves et al., 2005; Balamane et al., 2010; Pride et al., 2012), thus could serve as biomarkers of viral infection. Thus, far, little is known regarding the landscape and function of exogenous ncRNAs in saliva.
Future Challenges and Perspectives
A comprehensive ncRNA expression profile is emerging for human saliva including the presence of miRNAs, piRNAs, and circular RNAs (Ogawa et al., 2013; Bahn et al., 2015). More RNA species may be discovered in the future given the rapid evolution of new technologies and powerful bioinformatic methods. The value of saliva as a body fluid for biomarker discovery is just becoming widely recognized. However, there are a number of challenges in this field, most of which are general to usage of any body fluid in biomarker discoveries. One challenge lies in the unbiased isolation of short and long RNA molecules from saliva samples. Although this topic is under intensive investigation, improved methods that can retain most RNA species unbiasedly in an operator-independent manner are highly desired.
Another challenge is accurate quantification of ncRNA abundance, which is key to biomarker assessment. RNA yield from different samples may vary greatly, which calls for effective experimental and bioinformatic methods for normalization of RNA expression. Most RNA-Seq studies discussed above calculated RNA expression levels by normalizing the number of reads of a particular RNA molecule against the total number of mapped reads (i.e., the RPKM measure Mortazavi et al., 2008). However, to estimate the absolute concentration of an RNA molecule in a sample, synthetic spike-in RNAs with known concentration should be added to the RNA sample before library generation. This approach necessitates accurate measurement of RNA concentration of the sample and synthesis of a large number of spike-in RNAs with varying sequence contents and concentrations (see Williams et al., 2013 for a demonstration of this approach). This challenging approach, though highly desirable and necessary for clinical usage of a biomarker, has not been widely adopted.
A third major challenge is a better understanding of the biogenesis pathways of human ncRNAs in saliva, which constitutes the basis to assess whether and to what extent ncRNA expression can reflect a person's health condition. Salivary RNAs could be derived from systemic organs or local cells of the oral cavity. Cellular origins of candidate biomarkers for various diseases should be further examined to substantiate our understanding of the validity of the biomarkers. Indeed, the presence, origin, and functional roles of disease biomarkers are all essential questions general to studies of different types of biomarkers and diseases. A valid disease biomarker should be directly involved in disease mechanisms or indirectly associated/correlated with key pathways driving the pathogenesis of disease. The ultimate question is how knowledge gained in biomarker studies could be utilized to develop effective strategies for disease prevention and treatment, which closely relies on a clear understanding of disease mechanisms.
Conflict of Interest Statement
David T. W. Wong is co-founder of RNAmeTRIX Inc., a molecular diagnostic company. He holds equity in RNAmeTRIX, and serves as a company Director and Scientific Advisor. The University of California also holds equity in RNAmeTRIX. Intellectual property that David Wong invented and which was patented by the University of California has been licensed to RNAmeTRIX. Additionally, he is a consultant to PeriRx. The other authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by NIH grants R01HG006264 (to XX) and UH2 TR000923 (DTWW).
References
Alevizos, I., Alexander, S., Turner, R. J., and Illei, G. G. (2011). MicroRNA expression profiles as biomarkers of minor salivary gland inflammation and dysfunction in Sjogren's syndrome. Arthritis Rheum. 63, 535–544. doi: 10.1002/art.30131
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Almstahl, A., and Wikstrom, M. (2003). Electrolytes in stimulated whole saliva in individuals with hyposalivation of different origins. Arch. Oral Biol. 48, 337–344. doi: 10.1016/S0003-9969(02)00200-5
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Aps, J. K., and Martens, L. C. (2005). Review: the physiology of saliva and transfer of drugs into saliva. Forensic Sci. Int. 150, 119–131. doi: 10.1016/j.forsciint.2004.10.026
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Aps, J. K., Van Den Maagdenberg, K., Delanghe, J. R., and Martens, L. C. (2002). Flow cytometry as a new method to quantify the cellular content of human saliva and its relation to gingivitis. Clin. Chim. Acta 321, 35–41. doi: 10.1016/S0009-8981(02)00062-1
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Aravin, A., Gaidatzis, D., Pfeffer, S., Lagos-Quintana, M., Landgraf, P., Iovino, N., et al. (2006). A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207. doi: 10.1038/nature04916
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bahn, J. H., Zhang, Q., Li, F., Chan, T. M., Lin, X., Kim, Y., et al. (2015). The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 61, 221–230. doi: 10.1373/clinchem.2014.230433
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Balamane, M., Winters, M. A., Dalai, S. C., Freeman, A. H., Traves, M. W., Israelski, D. M., et al. (2010). Detection of HIV-1 in Saliva: implications for case-identification, clinical monitoring and surveillance for drug resistance. Open Virol. J. 4, 88–93. doi: 10.2174/1874357901004010088
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bonne, N. J., and Wong, D. T. (2012). Salivary biomarker development using genomic, proteomic and metabolomic approaches. Genome Med. 4, 82. doi: 10.1186/gm383
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Capel, B., Swain, A., Nicolis, S., Hacker, A., Walter, M., Koopman, P., et al. (1993). Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell 73, 1019–1030. doi: 10.1016/0092-8674(93)90279-Y
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Chen, M., Sonnerborg, A., Johansson, B., and Sallberg, M. (1997). Detection of hepatitis G virus (GB virus C) RNA in human saliva. J. Clin. Microbiol. 35, 973–975.
Chen, X., Ba, Y., Ma, L., Cai, X., Yin, Y., Wang, K., et al. (2008). Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 18, 997–1006. doi: 10.1038/cr.2008.282
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cocquerelle, C., Daubersies, P., Majerus, M. A., Kerckaert, J. P., and Bailleul, B. (1992). Splicing with inverted order of exons occurs proximal to large introns. EMBO J. 11, 1095–1098.
Das, P. P., Bagijn, M. P., Goldstein, L. D., Woolford, J. R., Lehrbach, N. J., Sapetschnig, A., et al. (2008). Piwi and piRNAs act upstream of an endogenous siRNA pathway to suppress Tc3 transposon mobility in the Caenorhabditis elegans germline. Mol. Cell 31, 79–90. doi: 10.1016/j.molcel.2008.06.003
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fichtlscherer, S., De Rosa, S., Fox, H., Schwietz, T., Fischer, A., Liebetrau, C., et al. (2010). Circulating microRNAs in patients with coronary artery disease. Circ. Res. 107, 677–684. doi: 10.1161/CIRCRESAHA.109.215566
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gallo, A., Tandon, M., Alevizos, I., and Illei, G. G. (2012). The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 7:e30679. doi: 10.1371/journal.pone.0030679
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Garrett, J. R. (1975). Changing attitudes on salivary secretion—a short history on spit. Proc. R. Soc. Med. 68, 553–560.
Gilad, S., Meiri, E., Yogev, Y., Benjamin, S., Lebanony, D., Yerushalmi, N., et al. (2008). Serum microRNAs are promising novel biomarkers. PLoS ONE 3:e3148. doi: 10.1371/journal.pone.0003148
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Girard, A., Sachidanandam, R., Hannon, G. J., and Carmell, M. A. (2006). A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202. doi: 10.1038/nature04917
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Goncalves, P. L., Cunha, C. B., Busek, S. C., Oliveira, G. C., Ribeiro-Rodrigues, R., and Pereira, F. E. (2005). Detection of hepatitis C virus RNA in saliva samples from patients with seric anti-HCV antibodies. Braz. J. Infect. Dis. 9, 28–34. doi: 10.1590/S1413-86702005000100006
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Grivna, S. T., Beyret, E., Wang, Z., and Lin, H. (2006). A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 20, 1709–1714. doi: 10.1101/gad.1434406
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Guo, J. U., Agarwal, V., Guo, H., and Bartel, D. P. (2014). Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 15:409. doi: 10.1186/s13059-014-0409-z
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hansen, T. B., Jensen, T. I., Clausen, B. H., Bramsen, J. B., Finsen, B., Damgaard, C. K., et al. (2013). Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388. doi: 10.1038/nature11993
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hermida, M., Ferreiro, M. C., Barral, S., Laredo, R., Castro, A., and Diz Dios, P. (2002). Detection of HCV RNA in saliva of patients with hepatitis C virus infection by using a highly sensitive test. J. Virol. Methods 101, 29–35. doi: 10.1016/S0166-0934(01)00417-7
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Human Microbiome Project Consortium. (2012). Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214. doi: 10.1038/nature11234
Jeck, W. R., Sorrentino, J. A., Wang, K., Slevin, M. K., Burd, C. E., Liu, J., et al. (2013). Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157. doi: 10.1261/rna.035667.112
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kos, A., Dijkema, R., Arnberg, A. C., Van der Meide, P. H., and Schellekens, H. (1986). The hepatitis delta (delta) virus possesses a circular RNA. Nature 323, 558–560. doi: 10.1038/323558a0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kozomara, A., and Griffiths-Jones, S. (2014). miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–D73. doi: 10.1093/nar/gkt1181
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lau, N. C., Seto, A. G., Kim, J., Kuramochi-Miyagawa, S., Nakano, T., Bartel, D. P., et al. (2006). Characterization of the piRNA complex from rat testes. Science 313, 363–367. doi: 10.1126/science.1130164
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, J. H., Ang, J. K., and Xiao, X. (2013). Analysis and design of RNA sequencing experiments for identifying RNA editing and other single-nucleotide variants. RNA 19, 725–732. doi: 10.1261/rna.037903.112
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, J. H., Gao, C., Peng, G., Greer, C., Ren, S., Wang, Y., et al. (2011). Analysis of transcriptome complexity through RNA sequencing in normal and failing murine hearts. Circ. Res. 109, 1332–1341. doi: 10.1161/CIRCRESAHA.111.249433
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, G., Bahn, J. H., Lee, J. H., Peng, G., Chen, Z., Nelson, S. F., et al. (2012). Identification of allele-specific alternative mRNA processing via transcriptome sequencing. Nucleic Acids Res. 40, e104. doi: 10.1093/nar/gks280
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, L. M., Hu, Z. B., Zhou, Z. X., Chen, X., Liu, F. Y., Zhang, J. F., et al. (2010). Serum microRNA profiles serve as novel biomarkers for HBV infection and diagnosis of HBV-positive hepatocarcinoma. Cancer Res. 70, 9798–9807. doi: 10.1158/0008-5472.CAN-10-1001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, P., Chen, S., Chen, H., Mo, X., Li, T., Shao, Y., et al. (2015). Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin. Chim. Acta 444C, 132–136. doi: 10.1016/j.cca.2015.02.018
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, Y., Zhou, X., St John, M. A., and Wong, D. T. (2004). RNA profiling of cell-free saliva using microarray technology. J. Dent. Res. 83, 199–203. doi: 10.1177/154405910408300303
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Liou, T. C., Chang, T. T., Young, K. C., Lin, X. Z., Lin, C. Y., and Wu, H. L. (1992). Detection of HCV RNA in saliva, urine, seminal fluid, and ascites. J. Med. Virol. 37, 197–202. doi: 10.1002/jmv.1890370309
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Liu, R., Zhang, C., Hu, Z., Li, G., Wang, C., Yang, C., et al. (2011). A five-microRNA signature identified from genome-wide serum microRNA expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur. J. Cancer 47, 784–791. doi: 10.1016/j.ejca.2010.10.025
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Mackiewicz, V., Dussaix, E., Le Petitcorps, M. F., and Roque-Afonso, A. M. (2004). Detection of hepatitis A virus RNA in saliva. J. Clin. Microbiol. 42, 4329–4331. doi: 10.1128/JCM.42.9.4329-4331.2004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Malathi, N., Mythili, S., and Vasanthi, H. R. (2014). Salivary diagnostics: a brief review. ISRN Dent. 2014:158786. doi: 10.1155/2014/158786
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Malone, C. D., and Hannon, G. J. (2009). Molecular evolution of piRNA and transposon control pathways in Drosophila. Cold Spring Harb. Symp. Quant. Biol. 74, 225–234. doi: 10.1101/sqb.2009.74.052
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Matse, J. H., Yoshizawa, J., Wang, X., Elashoff, D., Bolscher, J. G., Veerman, E. C., et al. (2013). Discovery and prevalidation of salivary extracellular microRNA biomarkers panel for the noninvasive detection of benign and malignant parotid gland tumors. Clin. Cancer Res. 19, 3032–3038. doi: 10.1158/1078-0432.CCR-12-3505
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Memczak, S., Jens, M., Elefsinioti, A., Torti, F., Krueger, J., Rybak, A., et al. (2013). Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338. doi: 10.1038/nature11928
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Michael, A., Bajracharya, S. D., Yuen, P. S., Zhou, H., Star, R. A., Illei, G. G., et al. (2010). Exosomes from human saliva as a source of microRNA biomarkers. Oral Dis. 16, 34–38. doi: 10.1111/j.1601-0825.2009.01604.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Mitchell, P. S., Parkin, R. K., Kroh, E. M., Fritz, B. R., Wyman, S. K., Pogosova-Agadjanyan, E. L., et al. (2008). Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. U.S.A. 105, 10513–10518. doi: 10.1073/pnas.0804549105
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L., and Wold, B. (2008). Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628. doi: 10.1038/nmeth.1226
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nigro, J. M., Cho, K. R., Fearon, E. R., Kern, S. E., Ruppert, J. M., Oliner, J. D., et al. (1991). Scrambled exons. Cell 64, 607–613. doi: 10.1016/0092-8674(91)90244-S
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ogawa, Y., Taketomi, Y., Murakami, M., Tsujimoto, M., and Yanoshita, R. (2013). Small RNA transcriptomes of two types of exosomes in human whole saliva determined by next generation sequencing. Biol. Pharm. Bull. 36, 66–75. doi: 10.1248/bpb.b12-00607
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Oshlack, A., Robinson, M. D., and Young, M. D. (2010). From RNA-seq reads to differential expression results. Genome Biol. 11:220. doi: 10.1186/gb-2010-11-12-220
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Palanisamy, V., Sharma, S., Deshpande, A., Zhou, H., Gimzewski, J., and Wong, D. T. (2010). Nanostructural and transcriptomic analyses of human saliva derived exosomes. PLoS ONE 5:e8577. doi: 10.1371/journal.pone.0008577
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Palanisamy, V., and Wong, D. T. (2010). Transcriptomic analyses of saliva. Methods Mol. Biol. 666, 43–51. doi: 10.1007/978-1-60761-820-1_4
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Park, N. J., Li, Y., Yu, T., Brinkman, B. M., and Wong, D. T. (2006). Characterization of RNA in saliva. Clin. Chem. 52, 988–994. doi: 10.1373/clinchem.2005.063206
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Park, N. J., Zhou, H., Elashoff, D., Henson, B. S., Kastratovic, D. A., Abemayor, E., et al. (2009). Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 15, 5473–5477. doi: 10.1158/1078-0432.CCR-09-0736
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Park, N. J., Zhou, X., Yu, T., Brinkman, B. M., Zimmermann, B. G., Palanisamy, V., et al. (2007). Characterization of salivary RNA by cDNA library analysis. Arch. Oral Biol. 52, 30–35. doi: 10.1016/j.archoralbio.2006.08.014
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Patel, R. S., Jakymiw, A., Yao, B., Pauley, B. A., Carcamo, W. C., Katz, J., et al. (2011). High resolution of microRNA signatures in human whole saliva. Arch. Oral Biol. 56, 1506–1513. doi: 10.1016/j.archoralbio.2011.05.015
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Pride, D. T., Salzman, J., Haynes, M., Rohwer, F., Davis-Long, C., White, R. A. III, et al. (2012). Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J. 6, 915–926. doi: 10.1038/ismej.2011.169
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sai Lakshmi, S., and Agrawal, S. (2008). piRNABank: a web resource on classified and clustered Piwi-interacting RNAs. Nucleic Acids Res. 36, D173–D177. doi: 10.1093/nar/gkm696
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L., and Brown, P. O. (2013). Cell-type specific features of circular RNA expression. PLoS Genet. 9:e1003777. doi: 10.1371/journal.pgen.1003777
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N., and Brown, P. O. (2012). Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7:e30733. doi: 10.1371/journal.pone.0030733
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sanger, H. L., Klotz, G., Riesner, D., Gross, H. J., and Kleinschmidt, A. K. (1976). Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc. Natl. Acad. Sci. U.S.A. 73, 3852–3856. doi: 10.1073/pnas.73.11.3852
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schafer, C. A., Schafer, J. J., Yakob, M., Lima, P., Camargo, P., and Wong, D. T. (2014). Saliva diagnostics: utilizing oral fluids to determine health status. Monogr. Oral Sci. 24, 88–98. doi: 10.1159/000358791
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schipper, R. G., Silletti, E., and Vingerhoeds, M. H. (2007). Saliva as research material: biochemical, physicochemical and practical aspects. Arch. Oral Biol. 52, 1114–1135. doi: 10.1016/j.archoralbio.2007.06.009
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Shugars, D. C., Patton, L. L., Freel, S. A., Gray, L. R., Vollmer, R. T., Eron, J. J. Jr., et al. (2001). Hyper-excretion of human immunodeficiency virus type 1 RNA in saliva. J. Dent. Res. 80, 414–420. doi: 10.1177/00220345010800020301
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Spielmann, N., Ilsley, D., Gu, J., Lea, K., Brockman, J., Heater, S., et al. (2012). The human salivary RNA transcriptome revealed by massively parallel sequencing. Clin. Chem. 58, 1314–1321. doi: 10.1373/clinchem.2011.176941
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Vieira, J., Huang, M. L., Koelle, D. M., and Corey, L. (1997). Transmissible Kaposi's sarcoma-associated herpesvirus (human herpesvirus 8) in saliva of men with a history of Kaposi's sarcoma. J. Virol. 71, 7083–7087.
Wang, G. K., Zhu, J. Q., Zhang, J. T., Li, Q., Li, Y., He, J., et al. (2010). Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur. Heart J. 31, 659–666. doi: 10.1093/eurheartj/ehq013
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, K., Zhang, S., Marzolf, B., Troisch, P., Brightman, A., Hu, Z., et al. (2009). Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc. Natl. Acad. Sci. U.S.A. 106, 4402–4407. doi: 10.1073/pnas.0813371106
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, Y., and Wang, Z. (2015). Efficient backsplicing produces translatable circular mRNAs. RNA 21, 172–179. doi: 10.1261/rna.048272.114
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Watanabe, T., Takeda, A., Tsukiyama, T., Mise, K., Okuno, T., Sasaki, H., et al. (2006). Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 20, 1732–1743. doi: 10.1101/gad.1425706
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Weber, J. A., Baxter, D. H., Zhang, S., Huang, D. Y., Huang, K. H., Lee, M. J., et al. (2010). The microRNA spectrum in 12 body fluids. Clin. Chem. 56, 1733–1741. doi: 10.1373/clinchem.2010.147405
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Williams, Z., Ben-Dov, I. Z., Elias, R., Mihailovic, A., Brown, M., Rosenwaks, Z., et al. (2013). Comprehensive profiling of circulating microRNA via small RNA sequencing of cDNA libraries reveals biomarker potential and limitations. Proc. Natl. Acad. Sci. U.S.A. 110, 4255–4260. doi: 10.1073/pnas.1214046110
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Xie, C., Yuan, J., Li, H., Li, M., Zhao, G., Bu, D., et al. (2014). NONCODEv4: exploring the world of long non-coding RNA genes. Nucleic Acids Res. 42, D98–D103. doi: 10.1093/nar/gkt1222
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Xie, Z., Chen, G., Zhang, X., Li, D., Huang, J., Yang, C., et al. (2013). Salivary microRNAs as promising biomarkers for detection of esophageal cancer. PLoS ONE 8:e57502. doi: 10.1371/journal.pone.0057502
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: RNA-Seq, miRNA, piRNA, circular RNA, extracellular RNA, biomarker, saliva
Citation: Lin X, Lo H-C, Wong DTW and Xiao X (2015) Noncoding RNAs in human saliva as potential disease biomarkers. Front. Genet. 6:175. doi: 10.3389/fgene.2015.00175
Received: 03 March 2015; Paper pending published: 27 March 2015;
Accepted: 22 April 2015; Published: 07 May 2015.
Edited by:
Mohammadreza Hajjari, Shahid Chamran University of Ahvaz, IranReviewed by:
Edward K. L. Chan, University of Florida, USADaniel Wai Hung Ho, The University of Hong Kong, China
Jun-An Chen, Academia Sinica, Taiwan
Helen K. W. Law, The Hong Kong Polytechnic University, China
Copyright © 2015 Lin, Lo, Wong and Xiao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xinshu Xiao,Z3h4aWFvQHVjbGEuZWR1
†These authors have contributed equally to this work.
 Xianzhi Lin
Xianzhi Lin Hsien-Chun Lo
Hsien-Chun Lo David T. W. Wong
David T. W. Wong Xinshu Xiao
Xinshu Xiao