- Department of Molecular Genetics, Faculty of Biological Sciences, Tarbiat Modares University, Tehran, Iran
microRNAs (miRNAs) are a novel class of non-coding RNAs which found their way into the clinic due to their fundamental roles in cellular processes such as differentiation, proliferation, and apoptosis. Recently, miRNAs have been known as micromodulators in cellular communications being involved in cell signaling and microenvironment remodeling. In this review, we will focus on the role of miRNAs in cardiovascular diseases (CVDs) and their reliability as diagnostic and therapeutic biomarkers in these conditions. CVDs comprise a variety of blood vessels and heart disorders with a high rate of morbidity and mortality worldwide. This necessitates introduction of novel molecular biomarkers for early detection, prevention, or treatment of these diseases. miRNAs, due to their stability, tissue-specific expression pattern and secretion to the corresponding body fluids, are attractive targets for cardiovascular-associated therapeutics. Explaining the challenges ahead of miRNA-based therapies, we will discuss the exosomes as delivery packages for miRNA drugs and promising novel strategies for the future of miRNA-based therapeutics. These approaches provide insights to the future of personalized medicine for the treatment of CVDs.
Introduction
Among multifactorial diseases, cardiovascular diseases (CVDs) are significant due to their variable symptoms and high mortality rate accounting for one third of global deaths (Santulli, 2013). They include a wide range of disorders connected to blood vessels and heart ranging from coronary artery disease (CAD), pulmonary arterial hypertension (PAH), and congenital heart disease to deep vein thrombosis and cerebrovascular disease. Major CVD risk factors include family history, obesity, hypertension, diabetes mellitus, and hypercholesterolemia (Mendis et al., 2011). Since CVDs are hard-to-cure, several investigations have focused on different mechanisms underlying CVD in order to manage the symptoms. microRNAs have emerged as one of the most favorable molecular targets in this regard. microRNAs are a class of non-coding RNAs with a short length of 18–24 nucleotides. They mainly act as post-translational repressors of gene expression. By regulating the fundamental cellular mechanisms such as cell differentiation, proliferation, growth, and apoptosis, miRNAs have received enormous attention for therapeutic applications. Huge investigations and investments have been made to bring these molecules into the clinic. In this review, we will explain the novel diagnostic, therapeutic and prognostic approaches based on miRNAs in the field of CVDs (cardio-miRs) and the problems ahead of this research area; where we are and where we expect to be in the future.
miRNA Mechanisms of Action
Calin et al. (2002) were first to link miRNAs with cancer progression. Since then several studies have focused on the role of these small molecules in the pathogenesis of different human diseases. The primary findings emphasized on the post-transcriptional regulatory role of miRNAs through base pairing with the 3′ untranslated region (UTR) of their target mRNAs. This leads to the mRNA degradation or translation inhibition and in both cases the miRNA binding results in the suppression of their target mRNAs. This is the main mechanism reported for the miRNAs regulatory role; however, many variations have also been reported (Ha and Kim, 2014; Lin and Gregory, 2015). miRNA binding to the 5′ UTR of transcripts has also been demonstrated to be able to activate or suppress target genes (Lee et al., 2009). Some miRNAs can also bind to the open reading frame (ORF) of mRNA transcripts and repress translation. This mechanism was first reported by Tay et al. (2008). They demonstrated that miRNA binding to the coding region of pluripotency genes can regulate the embryonic stem cell differentiation. miRNA binding might also happen at the promoter of target gene which causes repression of gene translation. On the other hand, some miRNAs modulate their target expression by binding to RNA-binding proteins that regulate the expression of mRNA transcripts (Eiring et al., 2010). Salmena et al. (2011) proposed the competing endogenous RNA (ceRNA) hypothesis according to which, miRNAs, long non-coding RNAs (lncRNAs) and target mRNAs are in a finely tuned interaction through miRNA response elements (MREs) and their competition for target binding based on their total concentration in the cytoplasm defines a higher level of regulation. Recent studies have indicated another fascinating aspect of miRNA regulatory network. Fabbri et al. (2012) proved that miRNAs excreted from cancer cells, can directly bind the toll-like receptors (TLRs) at the surface of neighboring immune cells and activate the relevant signaling pathways in the recipient cells. The complexity of miRNA-mediated regulatory systems highlights the importance of these small molecules in the clinic and needs further proceeding technologies to get closer to clinical therapies.
miRNA Diagnostics in CVD
microRNAs show tissue-specific and time-dependent expression patterns turning them into a leading fact in using these small molecules as diagnostic and therapeutic targets (Lu et al., 2005). Similar to other developmental phenomena in embryogenesis, miRNAs play critical roles in cardiovascular development and also their expression profile changes according to different pathological conditions. Cardiac resynchronization therapy (CRT) after heart failure helps improve the heart arrhythmia and it also affects the myocardial miRNA expression. In responder patients, CRT affects cardiac processes including cardiac fibrosis, apoptosis, angiogenesis, and channel alterations and consequently alters the expression levels of miR-29 (implicated in cardiac fibrosis), miR-30, miR-92, and miR-145 (involved with cardiac angiogenesis), miR-30 (modulated in cardiac apoptosis), and miR-26 (affected by modified ionic channel function; Sardu et al., 2014).
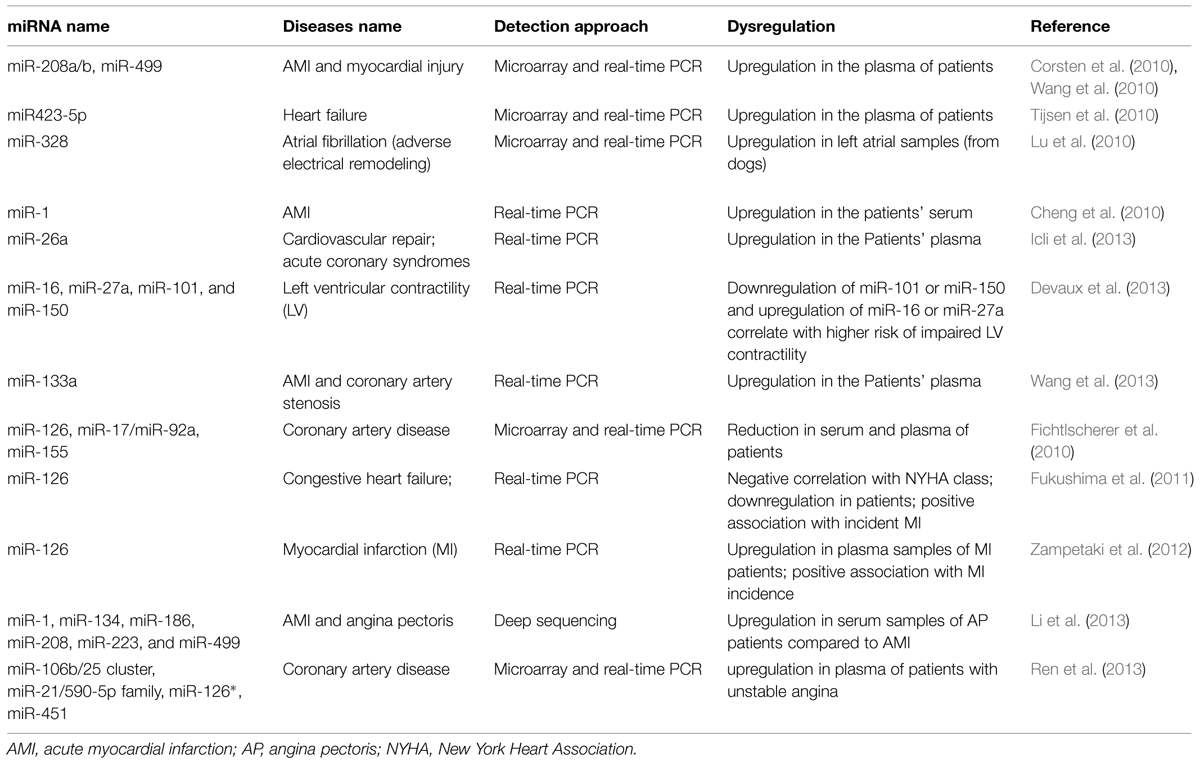
Recently circulatory miRNA expression profiling has provided strong molecular markers for detection of various diseases including CVD such as myocardial hyperthrophy, infarction, angiogenesis and fibrosis (Charan Reddy, 2014; Wronska et al., 2015). van Rooij et al. (2006) defined a miRNA signature for cardiac hypertrophy. They showed that miR-195 overexpression is sufficient to drive cardiac hypertrophy. miR-1 family, miR-133a/b and miR-208 are well-known for their implication in myogenesis both in skeletal muscles and cardiac development and their dysregulation has been detected in several CVDs including myocardial infarction, hypertrophy, and arrhythmias (Thum et al., 2008a). While miR-208 showed significant upregulation (pro-hypertrophic), miR-1 and miR-133a expression levels significantly decreased (anti-hypertrophic) in acute myocardial infarction (AMI) compared with normal individuals heart (Bostjancic et al., 2010). In CAD, miR-1 is upregulated in the left ventricular endocardium (Yang et al., 2007) and also plasma levels of miR-1, miR-133, and miR-208b have been reported to be elevated after AMI (Widera et al., 2011; Devaux et al., 2015). However, the circulatory miR-1 is not sensitive or specific enough to be known as an AMI specific biomarker since it is also affected by factors other than AMI. Moreover, by targeting Protein phosphatase 2A regulatory subunit B56 alpha (PP2A), miR-1 has been linked to the arrhythmia mainly through hyperphosphorylation of ryanodine receptor (RyR2; Terentyev et al., 2009). Following hypertension or cardiac infarction, cardiac hyperthrophy, and cardiac fibrosis are common diseases in which the excess amounts of extracellular matrix (ECM) proteins accumulate in cardiac tissue in order to adapt the system to the pathological conditions. Overexpression of miR-29 and miR-21 and downregulation of miR-133 and miR-30 have been linked to ECM remodeling and fibrosis by regulating several components of the ECM including collagen type I alpha 1 and 2 (Col1A1 and Col1A2) as targets of miR-29 (van Rooij et al., 2008), sprouty homolog 1 (SPRY1) as a target of miR-21 (Thum et al., 2008b) and connective tissue growth factor (CTGF) as target of miR-133 and miR-30c (Duisters et al., 2009). miR-133 is specifically expressed in cardiomyocytes while miR-30 is also detectable in cardiac fibroblasts as well as cardiomyocytes. These and other miRNAs (including miR-328) have been linked to atrial fibrillation which is mainly resulted from structural remodeling and fibrosis (Santulli et al., 2014a). Table 1 summarizes a list of miRNAs involved in different CVDs (cardio-miRs).
miRNA-Bearing Exosomes as Communicators of CVD
Recent miRNA-based studies have focused on the role of exosomes as natural delivery vehicles for some proteins, mRNA, and miRNAs which facilitate communication between cells and their neighboring stroma. Exosomes are small vesicles (40–100 nm) originating from the plasma membrane or multivesicular bodies (MVBs) and are present in almost all biological fluids. They shuttle between neighboring cells and transfer their cargoes which can be cellular components including proteins, mRNA, and non-coding RNAs. These cargoes can perform regulatory effects and control gene expression in the recipient cells (Pegtel et al., 2010). Exosomes are famous for being the communicators of microenvironment. Exosomal membrane proteins as well as other components are usually similar to their originating cell. Their membranes usually contain higher levels of sphingomyelin, cholesterol and phosphatidylserine compared to their originating cells. Their cargo also might represent the cellular composition of RNAs and proteins or they might show a separate profile. Methodologies including immunoblotting, affinity extraction into magnetic beads and flow cytometry have been used to identify the protein components of exosomes. Profiling of RNAs – miRNAs in particular – is usually performed by means of microarrays, qPCR-based arrays and most recently by next generation sequencing techniques which analyze the transcriptome – miRnome – of each sample.
Recent studies have proved targeted packaging of miRNAs and their biogenesis components including Dicer and AGO2 in exosomes (Melo et al., 2014). Exosome shuttling is implicated in a variety of disease including cancer, CVDs or viral infections. In CVD, exosomal transfer of miRNAs is a well-known mechanism through which, cells educate their adjacent environment in order to confront the pathological condition. In heart ischemia or fibrosis, paracrine or endocrine secretion of miRNA-bearing exosomes into cardiomyocytes or active fibroblasts of the heart have been demonstrated. This leads to trans-differentiation of fibroblasts into an active state with a different rate of growth factors secretion (van Rooij and Olson, 2009; Yoo et al., 2011). Bang et al. (2014) proved that following heart infarction, exosome-secretion from cardiomyocytes with a new miRNA profile, can reprogram the cardiac fibroblasts leading to cardiac hypertrophy. These mechanisms can introduce potential novel therapeutic targets. As well, recognizing tissue-specific CAF markers, the mechanisms underlying their reactivation and the miRNA signals in these processes, will potentially provide promising targets for CVD therapies. In addition to the fibroblasts in the microenvironment, exosomal transfer of signals is also used for cardiovascular protection after a disease. For example in heart ischemia, secretory exosomes with miRNA cargos from cardiomyocytes have been shown to activate and induce the bone marrow-derived stem cells. These latter cells release the second subset of exosomes that lead to myocardial regeneration or protection (Sahoo and Losordo, 2014). Several investigations have demonstrated the selective packaging of miRNAs in exosomes and their secretion to the stroma of cells by means of signal molecules. This is beyond shedding the vesicles by plasma membrane (Yang et al., 2011; Montecalvo et al., 2012; Stoorvogel, 2012). These disease-specific expression patterns of exosomal miRNAs – which can be non-invasively detected in the body fluids – provides potential molecular markers and promising therapeutic targets for treatment of CVDs. Several circulating miRNAs have been linked to some CVDs. In patients with AMI and myocardial injury, higher levels of miR-208a have been detected in the plasma of patients and they showed higher specificity and sensitivity than conventional biomarkers such as cardiac troponin I (TnI) for diagnosis of the disease (Ji et al., 2009; Wang et al., 2010). Another potential biomarker for AMI is reported by Adachi et al. (2010) who detected higher plasma levels of miR-499 in patients with AMI compared with normal individuals. Using microarray analysis of exosomes obtained from cardiac progenitor cells, Gray et al. (2015) introduced 11 miRNAs significantly up-regulated in response to hypoxic conditions while miR-292 showed the highest variations in these exosomes. Moreover, exosome profiling from cardiac fibroblasts was shown to be enriched with miR-21-3p (miR-21*). These fibroblasts affect cardiomyocytes and secrete the mediators of cardiac hypertrophy (Bang et al., 2014). Knowing the mechanisms underlying these communications and recognizing potential strong biomarkers for diagnosis and treatment of CVDs will help us in development of future molecular therapeutics.
miRNA Therapeutics in CVD
In the past decade, outstanding researches in the field of miRNA drugs have changed the face of molecular medicine. miRNA-based therapeutics mainly focus on reinstating the miRNA expression levels. Two main approaches include overexpression of downregulated miRNAs and suppression of overexpressed ones. These approaches have been subject to different modifications in order to improve the efficiency of delivery and less off-target effects. Promising tools in this regard are the oligonucleotides that mimic the endogenous miRNA or suppress the mature miRNA by sequence complementarity. Addition of locked nucleic acids (LNAs) or 2′-O-methylation of the antisense oligonucleotides increases the binding specificity while cholesterol conjugation enhances the circulation time, serum stability, and cellular uptake. Expression vectors bearing the pre-miRNA sequences or tandem miRNA target sites (sponges) sound promising for in vitro overexpression or suppression of miRNAs, respectively. Another issue to overcome is the efficient delivery of each of the miRNA drugs. Naked oligonucleotides are less efficient due to their instability in vitro or in vivo which subject them to different nucleases. Lipid-based vehicles, viral systems, and cationic polymers are among the main delivery tools for miRNA-based therapeutics. Each of these strategies has its own challenges and still needs improvements to address problems such as cytotoxicity, immunogenicity, and low efficiency (van Rooij and Olson, 2012). Fiedler et al. (2011) compared the effect of different doses of cholesterol-based anti-miR-24 (antagomirs) on miR-24 expression in cardiomyocytes and cardiac endothelial cells. They showed a cell-type specific tendency or mechanism for antagomir uptake by these tissues. Cholesterol-based antagomirs for silencing miR-21 were also proved to inhibit cardiac fibrosis and dysfunction (Kumarswamy et al., 2012). van Rooij et al. (2008) used anti-miR-29b oligonucleotides (cholesterol modified) after myocardial infarction and observed upregulation of ECM proteins leading to cardiac fibrosis. Also, Bonauer et al. (2009) demonstrated that intravenous injection of miR-92a antagomir, improved the function of damaged tissue in models of myocardial infarction. Inhibition of miR-92a results in neoangiogenic effects and functional recovery of ischemic tissues (Bonauer et al., 2009). Another promising miRNA in CVD therapeutics is miR-208 which is implicated in cardiac remodeling. LNA-modified anti-miR-208 oligonucleotides have successfully prevented pathologically associated cardiac remodeling during diastolic heart disease (van Rooij et al., 2007). A common issue in these approaches is in vivo instability and the low homing efficiency of oligonucleotides which results in modest changes in the expression levels of the target protein. Accordingly, improved stabilization or efficient delivery vehicles are required. Examples of such vehicles are adeno-associated viruses (AAVs). AAV9, a specific serotype of AAVs, has tropism for myocardiocytes and enriches in the heart (Bish et al., 2008). In another study Santulli et al. (2014b). proved that miR-126-3p down-regulation using an adeno-viral vector containing miR-126-3p target sites inhibits proliferative vascular smooth muscle cells and prevents restenosis in animal models. miR-126 is an endothelial specific miRNA that regulates vascular integrity and angiogenesis. These strategies provide an efficient tool for cardiac gene transfer and targeted delivery of miRNA-based therapeutics.
Due to their natural role in miRNA secretion and shuttling between different cells, exosomes are of great interest in miRNA therapeutics. Exosomes are flexible in size and cargo type and their non-synthetic nature potentiates them for more efficient and non-immunogenic delivery of cargo while they maintain the cargo integrity and stability. Exosomal membranes contain certain proteins which have binding affinity to specific receptors on the surface of recipient cells. So, they can selectively target cell types of interest and manipulating their miRNA components – as well as other molecular cargoes – will provide promising tools for the future of personalized medicine. In CVD, exosomes can be used as therapeutic agents, as protein delivery carriers or as gene therapy devices. Mesenchymal stem cell- derived exosomes have been investigated in the field of cardiac regenerative medicine especially in myocardial ischemia/reperfusion injuries (Lai et al., 2010). Martinez et al. (2006) have demonstrated that exosomes bearing differentiation signals can affect neovascularization and they are promising for treatment of angiogenic defects. Moreover, several exosomal miRNAs are implicated in angiogenesis and vascular repair. Zhang et al. (2010) showed that exosomes enriched with pre-miR-150 enhance endothelial cell migration. They transfected pre-miR-150/anti-miR-150 oligonucleotides to the THP-1 cell line and collected the conditioned media of these cells which was enriched with the exogenous oligonucleotides-bearing exosomes (Zhang et al., 2010). This conditioned medium affects the migration ability of endothelial cells. Exosome are novel promising elements for the future of CVD treatment due to their targeted delivery capacity and their microenvironment-dependent nature. The latter property triggers their activation in relation to the pathological microenvironment such as pH or substrate concentration. However, further studies would be necessary to overcome obstacles such as engineering and purifying exosomes, cargo loading into them and optimizing their quality and characterization for targeted delivery.
Conclusion
Cardiovascular diseases are the one of the leading causes of death worldwide. Most of CVDs can be prevented by controlling behavioral and environmental risk factors and early diagnosis and management play an important role in this regard. In this review, we focused on miRNAs as small non-coding RNAs involved in a variety of key cellular processes. Several miRNA biomarkers have been introduced for different CV situations. miRNAs have emerged as attractive novel therapeutics and they have several advantages over other molecular therapeutics due to their small size, conserved sequences and their stability in the body fluids. First anti-cancer miRNA-based drug, MRX-34 (a liposome-based miR-34 mimic) developed by Mirna Therapeutics came to the clinic in 2013 for the treatment of hepatocellular carcinoma (mirnarx.com; NCT01829971). Current achievements have portrayed a promising future for miRNA-based therapeutics although still several obstacles including their stability, renal clearance, off-target effects, inefficient endocytosis by target cells or the immunogenicity of delivery vehicles, need to be overcome. In CVD, some miRNA-based strategies have resulted in promising findings, including anti-miR-126 approaches that resulted in vascular smooth muscle cells and restenosis inhibition (Santulli et al., 2014b). Also, novel methodologies such as exosome-based delivery of miRNA drugs provide reliable evidence to overcome impediments such as inefficient, unspecific delivery, and immunogenic reactions. But these drugs still need investigations on different aspects of miRNA biology, their long term effects, subsequent biochemical and off-target effects and miRNA pathway analysis.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
NN is supported by a grant (#92002177) from Iran National Research Foundation (INSF).
References
Adachi, T., Nakanishi, M., Otsuka, Y., Nishimura, K., Hirokawa, G., Goto, Y., et al. (2010). Plasma microRNA 499 as a biomarker of acute myocardial infarction. Clin. Chem. 56, 1183–1185. doi: 10.1373/clinchem.2010.144121
Bang, C., Batkai, S., Dangwal, S., Gupta, S., Foinquinos, A., Holzmann, A., et al. (2014). Cardiac fibroblast–derived microRNA passenger strand-enriched exosomes mediate cardiomyocyte hypertrophy. J. Clin. Invest. 124, 2136–2146. doi: 10.1172/JCI70577
Bish, L., Morine, K., Sleeper, M., Sanmiguel, J., Wu, D., Gao, G., et al. (2008). Adeno-associated virus (AAV) serotype 9 provides global cardiac gene transfer superior to AAV1, AAV6, AAV7, and AAV8 in the mouse and rat. Hum. Gene Ther. 19, 1359–1368. doi: 10.1089/hum.2008.123
Bonauer, A., Carmona, G., Iwasaki, M., Mione, M., Koyanagi, M., Fischer, A., et al. (2009). MicroRNA-92a controls angiogenesis and functional recovery of ischemic tissues in mice. Science 324, 1710–1713. doi: 10.1126/science.1174381
Bostjancic, E., Zidar, N., Stajer, D., and Glavac, D. (2010). MicroRNAs miR-1, miR-133a, miR-133b and miR-208 are dysregulated in human myocardial infarction. Cardiology 115, 163–169. doi: 10.1159/000268088
Calin, G., Dumitru, C., Shimizu, M., Bichi, R., Zupo, S., Noch, E., et al. (2002). Frequent deletions and down-regulation of microRNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. U.S.A. 99, 15524–15529. doi: 10.1073/pnas.242606799
Charan Reddy, K. (2014). Regulatory noncoding RNAs in cardiovascular disease: shedding light on ‘Dark Matter.’ J. Cardiovasc. Dis. 3, 1–7.
Cheng, Y., Tan, N., Yang, J., Liu, X., Cao, X., He, P., et al. (2010). A translational study of circulating cell-free microRNA-1 in acute myocardial infarction. Clin. Sci. (Lond.) 119, 87–95. doi: 10.1042/CS20090645
Corsten, M., Dennert, R., Jochems, S., Kuznetsova, T., Devaux, Y., Hofstra, L., et al. (2010). Circulating MicroRNA-208b and MicroRNA-499 reflect myocardial damage in cardiovascular disease. Circ. Cardiovasc. Genet. 3, 499–506. doi: 10.1161/CIRCGENETICS.110.957415
Devaux, Y., Mueller, M., Haaf, P., Goretti, E., Twerenbold, R., Zangrando, J., et al. (2015). Diagnostic and prognostic value of circulating microRNAs in patients with acute chest pain. J. Intern. Med. 277, 260–271. doi: 10.1111/joim.12183
Devaux, Y., Vausort, M., Mccann, G., Kelly, D., Collignon, O., Ng, L., et al. (2013). A panel of 4 microRNAs facilitates the prediction of left ventricular contractility after acute myocardial infarction. PLoS ONE 8:e70644. doi: 10.1371/journal.pone.0070644
Duisters, R., Tijsen, A., Schroen, B., Leenders, J., Lentink, V., Van Der Made, I., et al. (2009). miR-133 and miR-30 regulate connective tissue growth factor: implications for a role of microRNAs in myocardial matrix remodeling. Circ. Res. 104, 170–178. doi: 10.1161/CIRCRESAHA.108.182535
Eiring, A., Harb, J., Neviani, P., Garton, C., Oaks, J., Spizzo, R., et al. (2010). miR-328 functions as an RNA decoy to modulate hnRNP E2 regulation of mRNA translation in leukemic blasts. Cell 140, 652–665. doi: 10.1016/j.cell.2010.01.007
Fabbri, M., Paone, A., Calore, F., Galli, R., Gaudio, E., Santhanam, R., et al. (2012). MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. PNAS 109, E2110–E2116. doi: 10.1073/pnas.1209414109
Fichtlscherer, S., De Rosa, S., Fox, H., Schwietz, T., Fischer, A., Liebetrau, C., et al. (2010). Circulating microRNAs in patients with coronary artery disease. Circ. Res. 107, 677–684. doi: 10.1161/CIRCRESAHA.109.215566
Fiedler, J., Jazbutyte, V., Kirchmaier, B., Gupta, S., Lorenzen, J., Hartmann, D., et al. (2011). MicroRNA-24 regulates vascularity after myocardial infarction. Circulation 124, 720–730. doi: 10.1161/CIRCULATIONAHA.111.039008
Fukushima, Y., Nakanishi, M., Nonogi, H., Goto, Y., and Iwai, N. (2011). Assessment of plasma miRNAs in congestive heart failure. Circ. J. 75, 336–340. doi: 10.1253/circj.CJ-10-0457
Gray, W., French, K., Ghosh-Choudhary, S., Maxwell, J., Brown, M., Platt, M., et al. (2015). Identification of therapeutic covariant microRNA clusters in hypoxia-treated cardiac progenitor cell exosomes using systems biology. Circ. Res. 116, 255–263. doi: 10.1161/CIRCRESAHA.116.304360
Ha, M., and Kim, V. (2014). Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 15, 509–524. doi: 10.1038/nrm3838
Icli, B., Wara, A., Moslehi, J., Sun, X., Plovie, E., Cahill, M., et al. (2013). MicroRNA-26a regulates pathological and physiological angiogenesis by targeting BMP/SMAD1 signaling. Circ. Res. 113, 1231–1241. doi: 10.1161/CIRCRESAHA.113.301780
Ji, X., Takahashi, R., Hiura, Y., Hirokawa, G., Fukushima, Y., and Iwai, N. (2009). Plasma miR-208 as a biomarker of myocardial injury. Clin. Chem. 55, 1944–1949. doi: 10.1373/clinchem.2009.125310
Kumarswamy, R., Volkmann, I., Jazbutyte, V., Dangwal, S., Park, D., and Thum, T. (2012). Transforming growth factor-β-induced endothelial-to-mesenchymal transition is partly mediated by microRNA-21. Arterioscler. Thromb. Vasc. Biol. 32, 361–369. doi: 10.1161/ATVBAHA.111.234286
Lai, R., Arslan, F., Lee, M., Sze, N., Choo, A., Chen, T., et al. (2010). Exosome secreted by MSC reduces myocardial ischemia/reperfusion injury. Stem Cell Res. 4, 214–222. doi: 10.1016/j.scr.2009.12.003
Lee, I., Ajay, S., Yook, J., Kim, H., Hong, S., Kim, N., et al. (2009). New class of microRNA targets containing simultaneous 5′-UTR and 3′-UTR interaction sites. Genome Res. 19, 1175–1183. doi: 10.1101/gr.089367.108
Li, C., Fang, Z., Jiang, T., Zhang, Q., Liu, C., Zhang, C., et al. (2013). Serum microRNAs profile from genome-wide serves as a fingerprint for diagnosis of acute myocardial infarction and angina pectoris. BMC Med. Genomics 6:16. doi: 10.1186/1755-8794-6-16
Lin, S., and Gregory, R. (2015). MicroRNA biogenesis pathways in cancer. Nat. Rev. Cancer 15, 321–333. doi: 10.1038/nrc3932
Lu, J., Getz, G., Miska, E., Alvarez-Saavedra, E., Lamb, J., Peck, D., et al. (2005). MicroRNA expression profiles classify human cancers. Nature 435, 834–838. doi: 10.1038/nature03702
Lu, Y., Zhang, Y., Wang, N., Pan, Z., Gao, X., Zhang, F., et al. (2010). MicroRNA-328 contributes to adverse electrical remodeling in atrial fibrillation. Circulation 122, 2378–2387. doi: 10.1161/CIRCULATIONAHA.110.958967
Martinez, M., Larbret, F., Zobairi, F., Coulombe, J., Debili, N., Vainchenker, W., et al. (2006). Transfer of differentiation signal by membrane microvesicles harboring hedgehog morphogens. Blood 108, 3012–3020. doi: 10.1182/blood-2006-04-019109
Melo, S., Sugimoto, H., O’connell, J., Kato, N., Villanueva, A., Vidal, A., et al. (2014). Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 26, 707–721. doi: 10.1016/j.ccell.2014.09.005
Mendis, S., Puska, P., and Norrving, B. (eds). (2011). Global Atlas on Cardiovascular Disease Prevention and Control. Geneva: World Health Organization in collaboration with the World Heart Federation and the World Stroke Organization.
Montecalvo, A., Larregina, A., Shufesky, W., Stolz, D., Sullivan, M., Karlsson, J., et al. (2012). Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood 119, 756–766. doi: 10.1182/blood-2011-02-338004
Pegtel, D., Cosmopoulos, K., Thorley-Lawson, D., Van Eijndhoven, M., Hopmans, E., Lindenberg, J., et al. (2010). Functional delivery of viral miRNAs via exosomes. Proc. Natl. Acad. Sci. U.S.A. 107, 6328–6333. doi: 10.1073/pnas.0914843107
Ren, J., Zhang, J., Xu, N., Han, G., Geng, Q., Song, J., et al. (2013). Signature of circulating microRNAs as potential biomarkers in vulnerable coronary artery disease. PLoS ONE 8:e80738. doi: 10.1371/journal.pone.0080738
Sahoo, S., and Losordo, D. (2014). Exosomes and cardiac repair after myocardial infarction. Circ. Res. 114, 333–344. doi: 10.1161/CIRCRESAHA.114.300639
Salmena, L., Poliseno, L., Tay, Y., Kats, L., and Pandolfi, P. (2011). A ceRNA hypothesis: the rosetta stone of a hidden rna language. Cell 146, 353–358. doi: 10.1016/j.cell.2011.07.014
Santulli, G. (2013). Epidemiology of cardiovascular disease in the 21st century: updated numbers and updated facts. J. Cardiovasc. Dis. 1, 1–2.
Santulli, G., Iaccarino, G., De Luca, N., Trimarco, B., and Condorelli, G. (2014a). Atrial fibrillation and microRNAs. Front. Physiol. 5:15. doi: 10.3389/fphys.2014.00015
Santulli, G., Wronska, A., Uryu, K., Diacovo, T., Gao, M., Marx, S., et al. (2014b). A selective microRNA-based strategy inhibits restenosis while preserving endothelial function. J. Clin. Invest. 124, 4102–4114. doi: 10.1172/JCI76069
Sardu, C., Marfella, R., Santulli, G., and Paolisso, G. (2014). Functional role of miRNA in cardiac resynchronization therapy. Pharmacogenomics 15, 1159–1168. doi: 10.2217/pgs.14.76
Stoorvogel, W. (2012). Functional transfer of microRNA by exosomes. Blood 119, 646–648. doi: 10.1182/blood-2011-11-389478
Tay, Y., Zhang, J., Thomson, A., Lim, B., and Rigoutsos, I. (2008). MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455, 1124–1128. doi: 10.1038/nature07299
Terentyev, D., Belevych, A., Terentyeva, R., Martin, M., Malana, G., Kuhn, D., et al. (2009). miR-1 overexpression enhances Ca(2+) release and promotes cardiac arrhythmogenesis by targeting PP2A regulatory subunit B56alpha and causing CaMKII-dependent hyperphosphorylation of RyR2. Circ. Res. 104, 514–521. doi: 10.1161/CIRCRESAHA.108.181651
Thum, T., Catalucci, D., and Bauersachs, J. (2008a). MicroRNAs: novel regulators in cardiac development and disease. Cardiovasc. Res. 79, 562–570. doi: 10.1093/cvr/cvn137
Thum, T., Gross, C., Fiedler, J., Fischer, T., Kissler, S., Bussen, M., et al. (2008b). MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 456, 980–984. doi: 10.1038/nature07511
Tijsen, A., Creemers, E., Moerland, P., De Windt, L., Van Der Wal, A., Kok, W., et al. (2010). MiR423-5p as a circulating biomarker for heart failure. Circ. Res. 106, 1035–1039. doi: 10.1161/CIRCRESAHA.110.218297
van Rooij, E., and Olson, E. (2009). Searching for MiR-acles in Cardiac Fibrosis. Circ. Res. 104, 138–140. doi: 10.1161/CIRCRESAHA.108.192492
van Rooij, E., and Olson, E. (2012). MicroRNA therapeutics for cardiovascular disease: opportunities and obstacles. Nat. Rev. Drug Discov. 11, 860–872. doi: 10.1038/nrd3864
van Rooij, E., Sutherland, L., Liu, N., Williams, A., Mcanally, J., Gerard, R., et al. (2006). A signature pattern of stress-responsive microRNAs that can evoke cardiac hypertrophy and heart failure. Proc. Natl. Acad. Sci. U.S.A. 103, 18255–18260. doi: 10.1073/pnas.0608791103
van Rooij, E., Sutherland, L., Qi, X., Richardson, J., Hill, J., and Olson, E. (2007). Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316, 575–579. doi: 10.1126/science.1139089
van Rooij, E., Sutherland, L., Thatcher, J., Dimaio, J., Naseem, R., Marshall, W., et al. (2008). Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc. Natl. Acad. Sci. U.S.A. 105, 13027–13032. doi: 10.1073/pnas.0805038105
Wang, F., Long, G., Zhao, C., Li, H., Chaugai, S., Wang, Y., et al. (2013). Plasma microRNA-133a is a new marker for both acute myocardial infarction and underlying coronary artery stenosis. J. Trans. Med. 11, 222. doi: 10.1186/1479-5876-11-222
Wang, G., Zhu, J., Zhang, J., Li, Q., Li, Y., He, J., et al. (2010). Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur. Heart J. 31, 659–666. doi: 10.1093/eurheartj/ehq013
Widera, C., Gupta, S., Lorenzen, J., Bang, C., Bauersachs, J., Bethmann, K., et al. (2011). Diagnostic and prognostic impact of six circulating microRNAs in acute coronary syndrome. J. Mol. Cell Cardiol. 51, 872–875. doi: 10.1016/j.yjmcc.2011.07.011
Wronska, A., Kurkowska-Jastrzebska, I., and Santulli, G. (2015). Application of microRNAs in diagnosis and treatment of cardiovascular disease. Acta Physiol. (Oxf.) 213, 60–83. doi: 10.1111/apha.12416
Yang, B., Lin, H., Xiao, J., Lu, Y., Luo, X., Li, B., et al. (2007). The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat. Med. 13, 486–891. doi: 10.1038/nm1569
Yang, M., Chen, J., Su, F., Yu, B., Su, F., Lin, L., et al. (2011). Microvesicles secreted by macrophages shuttle invasion-potentiating microRNAs into breast cancer cells. Mol. Cancer 10, 117. doi: 10.1186/1476-4598-10-117
Yoo, A., Sun, A., Li, L., Shcheglovitov, A., Portmann, T., Li, Y., et al. (2011). MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476, 228–231. doi: 10.1038/nature10323
Zampetaki, A., Willeit, P., Tilling, L., Drozdov, I., Prokopi, M., Renard, J., et al. (2012). Prospective study on circulating MicroRNAs and risk of myocardial infarction. J. Am. Coll. Cardiol. 60, 290–294. doi: 10.1016/j.jacc.2012.03.056
Keywords: miRNA, cardiovascular disease, cardio-miR, exosome, cell communication, delivery vehicle, secretory miRNA, miRNA therapeutics
Citation: Nouraee N and Mowla SJ (2015) miRNA therapeutics in cardiovascular diseases: promises and problems. Front. Genet. 6:232. doi: 10.3389/fgene.2015.00232
Received: 27 February 2015; Accepted: 17 June 2015;
Published: 30 June 2015.
Edited by:
Narasaiah Kolliputi, University of South Florida, USAReviewed by:
Gaetano Santulli, University of Naples Federico II, ItalyMichael Teng, University of South Florida, USA
Copyright © 2015 Nouraee and Mowla. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Seyed J. Mowla, Department of Molecular Genetics, Faculty of Biological Sciences, Tarbiat Modares University, P.O.Box 14115-175, Tehran, Iran,c2ptb3dsYUBtb2RhcmVzLmFjLmly
 Nazila Nouraee
Nazila Nouraee Seyed J. Mowla
Seyed J. Mowla