- 1Department of Biochemical Engineering and Biotechnology, Indian Institute of Technology Delhi, New Delhi, India
- 2Department of Molecular Biology, Ariel University, Ariel, Israel
Apoptosis, a form of programmed cell death, is a highly regulated process, the deregulation of which has been associated with the tumor initiation, progression, and metastasis in various cancers including breast cancer. Induction of apoptosis is a popular target of various therapies currently being tested or used for breast cancer treatment. Thus, identifying apoptotic mediators and regulators is imperative for molecular biologists and clinicians for benefit of patients. The regulation of apoptosis is complex and involves a tight equilibrium between the pro- and anti-apoptotic factors. Recent studies have highlighted the role of miRNAs in the control of apoptosis and their interplay with p53, the master guardian of apoptosis. Here, we summarize and integrate the data on the role of miRNAs in apoptosis in breast cancer and the clinical advantage it may offer for the prognosis or treatment of breast cancer patients.
Introduction
Breast cancer is the most common malignancy in women in western nations with a scary current estimate of one in eight US women likely to develop invasive breast cancer once in her lifetime (American Cancer Society, 2015; Ferlay et al., 2015). The lowest incidences of breast cancer are seen in Asia and Africa but the cases are steadily rising and poor survival rates remain a cause of grave concern (American Cancer Society, 2015; Ferlay et al., 2015). The various risk factors are age, density of the breast, family and personal health history, germline mutations in the tumor suppressor genes-BRCA1, BRCA2, and other breast cancer susceptibility genes, alcohol intake, endogenous estrogen levels, use of hormone replacement therapy, radiation exposure to the breast, obesity, and race/ethnicity (American Cancer Society, 2015; Ferlay et al., 2015). Breast cancer is a heterogeneous disease characterized by up to 21 distinct histological subtypes and at least four different molecular subtypes each with distinct presentations, response to treatment, and clinical outcomes (American Cancer Society, 2015; Ferlay et al., 2015).
There has been marked improvement in the breast cancer survival rates and decline in deaths owing to early detection, better understanding of the disease, and personalized treatment (Ferlay et al., 2015). Current breast cancer treatment modality involves surgery (both curative and preventive), radiation therapy, systemic chemotherapy, endocrine therapy, and targeted therapy that interfere with the cellular functions thereby inducing cell cycle arrest and eventually apoptosis, a form of programmed cell death (Gonzalez-Angulo et al., 2007; Florea and Busselberg, 2013). However, breast cancer still remains the leading cause of death from cancer among women. The most significant reasons for therapy failure are tumor metastases to distant sites and development of treatment resistance (Gonzalez-Angulo et al., 2007; Florea and Busselberg, 2013). It is often seen that metastatic and treatment resistant cells lose control over apoptosis and thus fail to die in response to various therapies. Identification of the mediators and regulators of the apoptotic pathway may help to tailor treatments for maximization of treatment efficacy leading to complete tumor remission and prevention of metastases (Kasibhatla and Tseng, 2003).
Recently, microRNAs (miRNAs) have emerged as critical regulators of apoptosis. miRNAs are small non-coding RNAs (~18–25 bases) regulating gene expression at the post-transcriptional and/or translational levels. Traditionally, they are known to bind to the 3′ untranslated region of the target transcript to bring about transcript degradation or inhibition of translation depending on the extent of complementarity (Bartel, 2004). However, recent research suggests that miRNA may bind anywhere in the transcript to regulate its levels and can also bring about increase in the transcript levels by various mechanisms (Antonio et al., 2006). It is increasingly becoming clear that miRNAs generate a complex network with mRNAs, snoRNAs, long non-coding RNAs, and RNA binding proteins to establish cellular transcriptome (Mattick and Makunin, 2006).
The generation of miRNA profiles of normal or patient samples using microarray, qPCR, or deep sequencing technology and its analyses has helped in identifying useful clinical correlations of miRNAs with potential applications in cancer diagnosis, prognosis or therapy (Hammond, 2006). Several human miRNAs have been reported to be significantly deregulated in at least one cancer type and have been shown to function as oncomiRs or tumor suppressor miRs. Such deregulation of miRNA expressions across various tumor types suggests that these miRNAs may be involved in crucial cellular pathways that are commonly deregulated in cancer development. In this review, we will focus on the recent progress of research on miRNA-mediated regulation of apoptosis in breast cancer, discuss pro- and anti-apoptotic miRNAs (ApoptomiRs), miRNA targets, and their interaction with the p53 pathway. Finally, we will discuss current/future cancer prognostics and treatment targets based on these apoptotic regulators.
Breast Cancer and Apoptosis
A critical balance between cell proliferation and apoptosis is essential for normal breast development. Accumulation of genetic or epigenetic mutations by various factors brings about disturbance in this balance leading to the development of breast tumors. Breast tumors are heterogeneous in their genetic composition and are histologically diverse based on receptor status, proliferation and differentiation markers that correlate to disease prognosis (Vakkala et al., 1999). The histological analyses guide clinicians in selecting adjuvant therapies for breast cancer treatment. Several studies have made attempts to correlate apoptosis and its associated molecular markers with known histologic and prognostic factors in breast cancer (Lipponen et al., 1994). A high apoptotic index has been correlated with increased tumor grade, aneuploidy, high mitotic index, negative status for estrogen receptor (ER), and progesterone receptor (PR), tumor necrosis and increased lymphocyte infiltration (Lipponen, 1999). The tumor suppressor gene, p53, and the BCL2 family have been widely studied in breast cancer. The mutant p53 correlates with negative ER status, high mitotic index, and the tumor grade, while BCL2 expression correlates to ER positive status, wild type p53, low mitotic index and low grade (Megha et al., 2002). Strikingly, expression of BCL2 is seen in ~80% of breast cancer patients. Contrary to expectations, high apoptotic index is significantly associated to poor prognosis while high BCL2 expression correlates to good prognosis of breast cancer patients (Silvestrini et al., 1996). Possible explanations may be that neither apoptotic index nor mutant p53 or BCL2 is independent predictor of survival in breast cancer patients. High apoptosis index also correlates to high proliferative index and negative ER status, which are poor prognostic factors. High apoptotic index may also put selective pressure on tumor cells leading to development of apoptosis resistant cells leading to tumor progression and decreased survival in patients (Frenzel et al., 2009). Similarly, BCL2, an estrogen regulated gene correlates with positive ER status, a good prognostic feature. Nevertheless, in vitro experiments with BCL2 overexpression or silencing in breast cancer cells confirm its pro-survival effects accounting for current pre-clinical and clinical trials involving BCL2 silencing for breast cancer treatment (Honma et al., 2015). Breast cancer cells show activation of the upstream apoptotic signaling such as an increase in Fas-L expression and presence of active caspases in cancer cells (Hengartner, 2001; Fulda and Debatin, 2006). However, upregulation of Inhibitors of apoptosis proteins (IAPs), X-linked inhibitor of apoptosis protein (XIAP), and Survivin inhibits activity of the caspases and blocks apoptotic signaling (Deveraux and Reed, 1999). Overall, the balance between the concentrations of pro- and anti-apoptotic proteins is an important regulatory factor for apoptotic regulation (Quail and Joyce, 2013).
Pro- and Anti-Apoptotic miRNAs in Breast Cancer
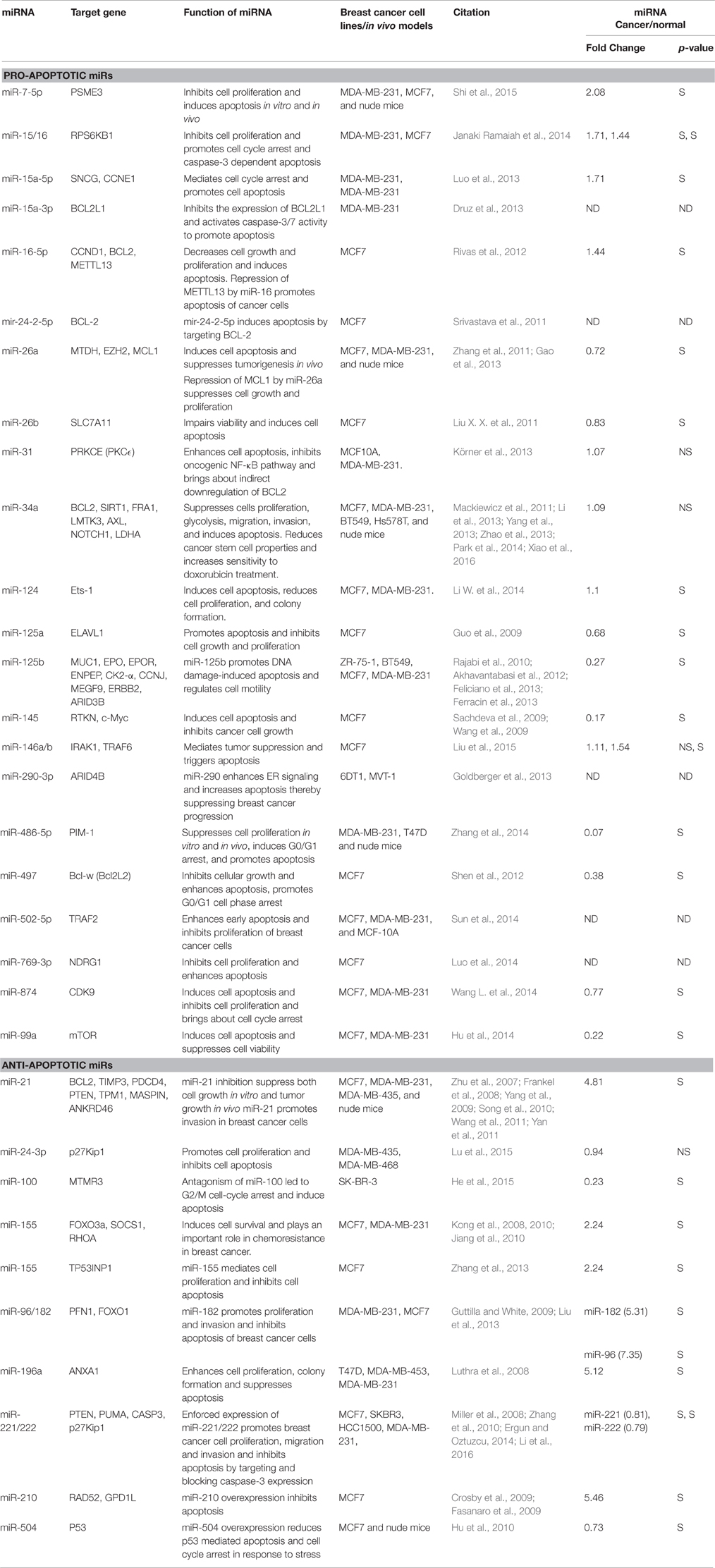
Deregulation of apoptosis is an important step in cancer as it allows the genetically unstable cells to survive and accumulate further mutations that eventually lead to tumorigenesis (Elmore, 2007). One of the mechanism by which miRNAs influence tumor development is by regulation of proteins involved in the apoptotic process. miRNAs that function to promote or inhibit apoptotis are called pro- and anti-apoptotic miRNAs, respectively. Table 1 enlists various miRNAs that have been reported to function as pro- or anti-apoptotic miRNAs in breast cancer.
Pro-ApoptomiRs
A total of 22 miRNAs have been reported so far to be involved in the induction of apoptosis suggesting them to be tumor suppressors. Tumor suppressor miRNAs prevent tumor progression by negatively regulating the expression of genes that promote cell proliferation, differentiation, migration, or apoptosis. We evaluated the expression of pro-apoptomiRs in breast cancer patients using StarBase software that analyzes TCGA data (Li J. H. et al., 2014). The data set included 683 breast cancer patients and 87 normal samples. Interestingly, we found that miR-26a, -26b, -125a, -125b, -145, -486-5p, -497, -874, and -99a that are known to function as proapoptomiRs were significantly downregulated in breast cancer patients. In the following section, we have discussed some pro-apoptotic miRNAs.
miR-15/16 Cluster
miR-15/16 was shown to inhibit cell proliferation and promote cell cycle arrest and apoptosis in various breast cancer cell lines (Rivas et al., 2012; Druz et al., 2013; Luo et al., 2013; Janaki Ramaiah et al., 2014). The members of miR-15/16 cluster negatively regulate the anti-apoptotic protein BCL2, cell cycle regulators CCND1, CCNE1 and other proteins RPS6KB1, SNCG and METTL13 in breast cancer (refer Table 1).
miR-26a/b
miR-26a is downregulated in breast cancer specimens and cell lines and its transient transfection initiates apoptosis of breast cancer cell line, MCF7. Oncogenes, MTDH, and EZH2 (a chromatin regulator), were identified as direct targets of miR-26a. MCF7 xenografts with exogenous miR-26a show that a decrease in expression of both MTDH and EZH2 is accompanied by an increase in apoptosis (Zhang et al., 2011). MCL1, an anti-apoptotic member of the BCL2 family and SLC7A11, an amino acid transporter are other targets of miR-26a (Gao et al., 2013). Silencing of SLC7A11 mimics miR-26b aroused viability impairment and apoptosis in MCF7 cells.
miR-34a
Several groups have independently reported pro-apoptotic role of miR-34a in breast cancer (Mackiewicz et al., 2011; Li et al., 2013; Yang et al., 2013; Zhao et al., 2013; Park et al., 2014; Xiao et al., 2016). miR-34 overexpression suppresses cell proliferation, glycolysis, metastasis, invasion, stem cell properties, and promotes apoptosis and sensitivity to doxorubicin treatment in breast cancer (Mackiewicz et al., 2011; Li et al., 2013; Yang et al., 2013; Zhao et al., 2013; Park et al., 2014; Xiao et al., 2016). miR-34a is transcriptionally regulated by p53 and targets SIRT1 (a modulator of p53 activity) thus making a mir-34-p53 positive feedback loop (Li et al., 2013). Other targets of miR-34a are BCL2, FRA1, LMTK3, AXL, NOTCH1 and LDHA. miR-34a/AXL interaction reduces phospho-AKT expression, and impairs the motility of triple-negative breast cancer cells (Mackiewicz et al., 2011). miR-34a suppresses MCF7 cell proliferation in vitro and in vivo by targeting LMTK3 in an estrogen dependent manner (Zhao et al., 2013). Overall, miR-34a functions as a tumor suppressor in breast cancer as also reported for other cancers.
miR-125a/b
miR-125a/b promotes apoptosis and inhibits cell growth and proliferation. miR-125a functions as a tumor suppressor for breast cancer, with HuR as a direct and functional target (Guo et al., 2009). miR-125b perform its tumor suppressor function via the direct targeting of the 3′-UTRs of ENPEP, CK2-α, CCNJ, MUC1, and MEGF9 mRNAs. miR-125b regulates cell motility by targeting ERBB2 and ARID3B (Rajabi et al., 2010; Akhavantabasi et al., 2012; Feliciano et al., 2013; Ferracin et al., 2013). Overexpression of miR-125b markedly inhibits Taxol-induced cell cytotoxicity in MDA-MB-435, MDA-MB-231, and BT474 breast cancer cells by targeting 3′UTR of BAK1 mRNA, which is an effector of taxol induced apoptosis (Zhou et al., 2010).
Anti-ApoptomiRs
A total of 11 miRNAs have been shown to inhibit apoptosis suggesting them to be oncomiRs in breast cancer. Among these miR-21, -155, -96, -182, -196a, and -210 were found to be highly upregulated in breast cancer patients (Table 1).
miR-21
miRNA 21 is overexpressed in many cancers and has been shown to have oncogenic activity. Si et al., 2007 showed that miR-21 knock down inhibits MCF7 cell growth in vitro and suppresses MCF7 cell derived breast tumor growth in a murine xenograft model (Si et al., 2007). This was associated with increased apoptosis and decreased cell proliferation by downregulation of anti-apoptotic BCL2. miR-21 was also shown to promote breast cancer cell invasion. Overall, miR-21 functions as an oncomiR and modulates tumorigenesis through regulation of genes such as ANKRD46, BCL2, TIMP3, PDCD4, PTEN, TPM1, and MASPIN in breast cancer (Zhu et al., 2007; Frankel et al., 2008; Yang et al., 2009; Song et al., 2010; Wang et al., 2011; Yan et al., 2011).
miR-155
miR-155 exerts its anti-apoptotic activity by downregulating FOXO3a for cell survival (Kong et al., 2010). Active form of FOXO3a resides in the nucleus and induces cell death by up-regulation of apoptotic proteins, such as BIM, p27, BNIP3, and 24p3 and repression of anti-apoptotic proteins, FLIP, and BCL-xL (Kong et al., 2010). miR-155 also targets TP531NP1, SOCS1, and RHOA and inhibits apoptosis and chemosensitivity in MCF7 cells (Kong et al., 2008, 2010; Jiang et al., 2010; Zhang et al., 2013).
miR-210
miR-210 is a hypoxia inducible miRNA that functions as an oncomiR in breast cancer. High miR-210 expression is a poor prognostic factor in breast cancer patients. miR-210 inhibition promotes apoptosis in breast cancer. It is known to target DNA repair protein RAD52 and a protein involved in mitochondrial function regulation, GPD1L (Crosby et al., 2009; Fasanaro et al., 2009).
miR-221/222
miR-221/222 are frequently up-regulated in human epithelial cancers. Enforced expression of miR-221/222 promotes breast cancer cell proliferation, migration, and invasion via targeting PTEN/AKT pathway. In breast cancer miR-221/222 directly targets pro-apoptotic proteins PUMA and CASP3 and cell cycle inhibitor, p27Kip1 to show its oncogenic effect (Zhang et al., 2010). miR-221/222 inhibition results in an increase in mitochondrial membrane potential and caspase-3/7 activity thereby leading to apoptosis in the A549 and MCF7 cell lines (Zhang et al., 2010).
Targets of ApoptomiRs
A total of 61 genes have been validated as direct targets of apoptomiRs. Interestingly, ~36% of these targets were found to be altered at the transcript (BCL2, CCNE1, CSKN2A1, ELAVL1, ERBB2, ETS1, EZH2, FOXO1, IRAK1, MUC1, MYC, NDRG1, PDCD4, SERPINB5, RPS6B1, SNCG, TIMP3, TP53, and TP53INP1) or the protein (BCL2, CASP3, CCNE1, CDKN1B, ERBB2, MUC1, MYC, PTEN, SERPINB5, TP53) levels in breast cancer based on analyses using genes to system breast cancer (G2SBC) database (Mosca et al., 2010). Whether the effects of the miRNAs on apoptosis are mediated through these targets has not been fully elucidated. Pathway analyses of the target genes of apoptomiRs using Reactome pathway browser identified transcriptional regulation by TP53 (CCNE1, CDKN1B, mTOR, CDK9, CSKN2A1, NDRG1, PTEN, PUMA, TP53, TP53INP1); signaling by nerve growth factor, NGF (IRAK1, ERBB2, CASP3, CDKN1B, RHOA, mTOR, FOXO1, PSME3, FOXO3, PRKCE, TRAF6, PTEN); and PI3K/AKT activation (CDKN1B, ERBB2, RHOA, mTOR, FOXO1, FOXO3, PTEN) as the top three overrepresented pathways (Joshi-Tope et al., 2005). NGF is known to be overexpressed in ~80% of breast tumor biopsies and has been implicated in enhanced survival, proliferation, migration, and inhibition of apoptosis in breast cancer (Molloy et al., 2011). Similarly, the pro-proliferative and anti-apoptotic effects of PI3K/AKT pathway are well documented. A protein interaction network analyses using STRING database (Von Mering et al., 2005) reveals that a strong interaction network with a high confidence (0.7) exists between the validated targets of the apoptomiRs suggesting that this set of the genes are biologically related (Figure 1).
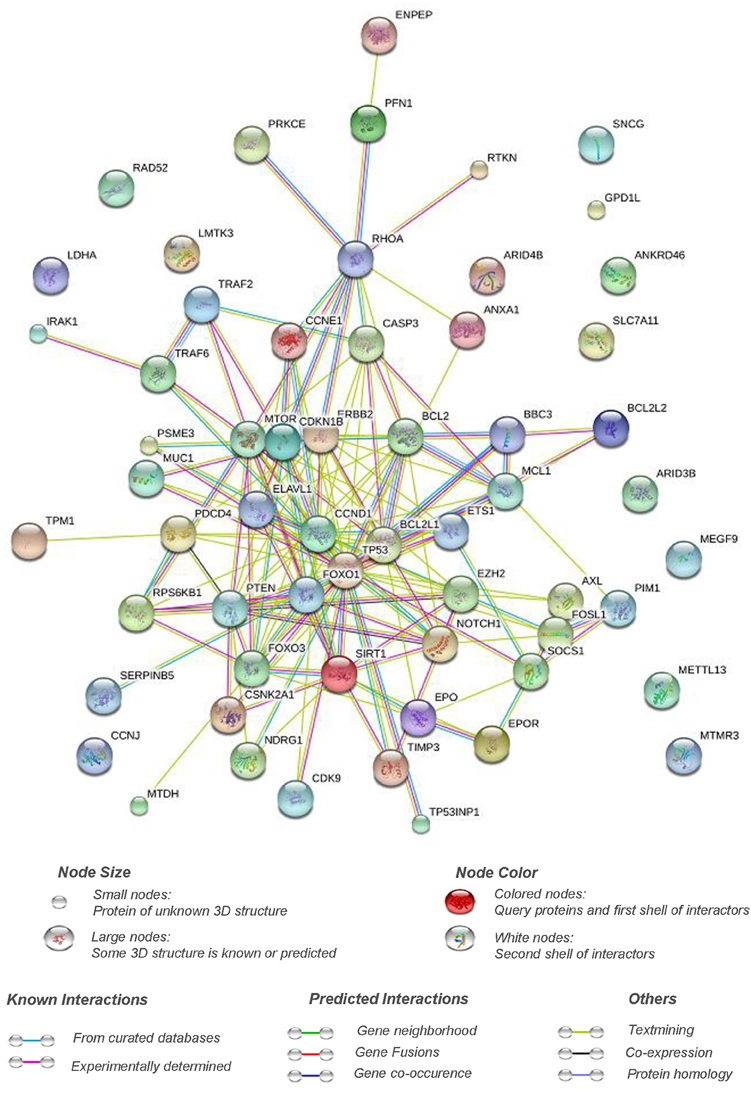
Figure 1. Pathway interaction network of targets of apoptomiRs in breast cancer. STRING interaction database (Von Mering et al., 2005) was used at setting of high confidence (0.7) to generate the interaction network of the list of the targets of apoptomiRs in breast cancer (Zhu et al., 2007; Frankel et al., 2008; Kong et al., 2008; Luthra et al., 2008; Miller et al., 2008; Crosby et al., 2009; Fasanaro et al., 2009; Guo et al., 2009; Guttilla and White, 2009; Sachdeva et al., 2009; Wang et al., 2009; Yang et al., 2009; Hu et al., 2010; Jiang et al., 2010; Kong et al., 2010; Rajabi et al., 2010; Song et al., 2010; Zhang et al., 2010; Liu X. X. et al., 2011; Mackiewicz et al., 2011; Srivastava et al., 2011; Wang et al., 2011; Yan et al., 2011; Zhang et al., 2011; Akhavantabasi et al., 2012; Rivas et al., 2012; Shen et al., 2012; Druz et al., 2013; Feliciano et al., 2013; Ferracin et al., 2013; Gao et al., 2013; Goldberger et al., 2013; Körner et al., 2013; Li et al., 2013; Liu et al., 2013; Luo et al., 2013; Yang et al., 2013; Zhang et al., 2013; Zhao et al., 2013; Ergun and Oztuzcu, 2014; Hu et al., 2014; Janaki Ramaiah et al., 2014; Luo et al., 2014; Li W. et al., 2014; Park et al., 2014; Sun et al., 2014; Wang L. et al., 2014; Zhang et al., 2014; He et al., 2015; Liu et al., 2015; Lu et al., 2015; Shi et al., 2015; Li et al., 2016; Xiao et al., 2016). The implication of the various connective lines based on different data sources are shown below the figure.
Interestingly, around 40% of the apoptomiR targets (BCL2, BCL2L1, BCl2L2, FOSL1, NOTCH1, TIMP3, TRAF2, TRAF6, ANXA1, CASP3, CDKN1B, FOXO1, FOXO3, IRAK1, MCL1, PTEN, PIM1, PSME3, PRKCE, PUMA, RHOA, SIRT1, TP53, TP53INP1, ERBB2, and ETS1) were found to be associated with regulation of apoptosis based on functional annotation clustering using DAVID microarray software (Huang et al., 2009a,b). Several of these belong to the p53 network or BCL2 family of proteins involved in control of apoptosis.
p53, is a tumor suppressor protein that mediates cellular responses to various stress signals like DNA damage, hypoxia, or aberrant oncogene expression by promoting cell-cycle checkpoints, DNA repair, cellular senescence, and apoptosis. In breast cancer, TP53 mutations have been associated with poor response to various therapies and thus poor prognosis. Several miRNAs were found to be associated with the p53 network by targeting p53 directly or indirectly (by targeting its upstream regulators or downstream mediators) to influence apoptosis in breast cancer. miR-504 was shown to directly target p53 and thus, inhibit p53 dependent apoptosis in breast cancer (Hu et al., 2010). MDM2 is an E3 ubiquitin-protein ligase that mediates ubiquitination of p53, leading to its proteasomal degradation and nuclear export. miR-143/145, and miR-1827 were found to directly target MDM2 in breast cancer and thus enhance p53-mediated stress responses, including apoptosis and senescence (Okamura et al., 2001). TP53INP1, SIRT1, and EGR1, regulators of p53 transcriptional activity, have also been shown to be direct targets of apoptomiRs- miR-155, miR-34, and miR-191, respectively, in breast cancer (Li et al., 2013; Nagpal et al., 2013; Zhang et al., 2013). Similarly, several miRNAs (miR-9, miR-17/20, 21, 125b, 155, 203, 222, and 342) affect the levels of p53 downstream genes to establish their effect on apoptosis. Additionally, p53 has been shown to induce the levels of several miRNAs involved in regulation of apoptosis in breast cancer. A total of six tumor suppressor miRNAs (miR-143, miR-145, miR-15, miR-16, miR-26a, and miR-34) have been shown to be upregulated by p53 in breast cancer (Sachdeva et al., 2009; Gao et al., 2013; Li et al., 2013; Janaki Ramaiah et al., 2014). Since miR-143/145 regulate MDM2 (regulator and transcriptional target of p53) and are also regulated by p53, it indicates existence of miR-143/145-MDM2-p53 feedback loop in breast cancer that controls cell proliferation and apoptosis (Okamura et al., 2001). Similarly, miR-34 is induced by p53 and targets SIRT1 (a modulator of p53 activity) thus making a mir-34-p53 positive feedback loop (Yamakuchi et al., 2008). Overall, apoptomiRs emerge as important regulators of the p53 network at multiple levels to regulate apoptosis in breast cancer (Figure 2).
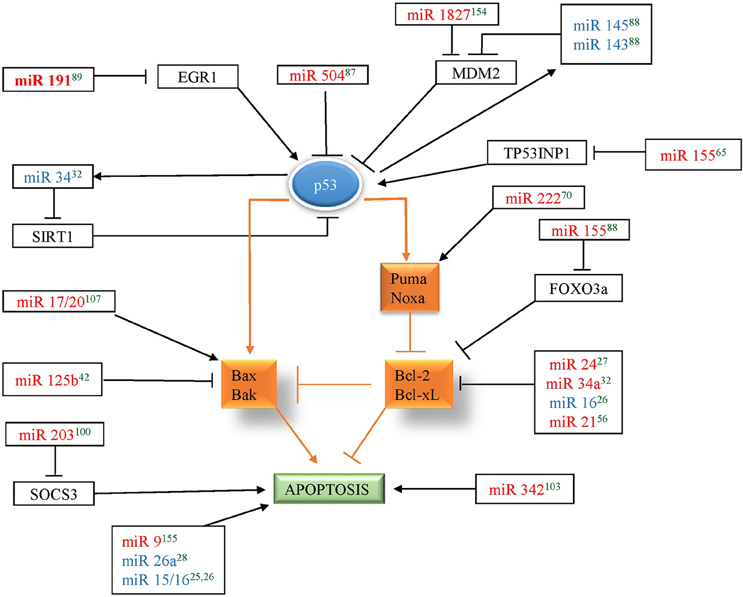
Figure 2. miRNAs in red color correspond to the miRNAs that regulate p53 in breast cancer and miRNAs in blue color corresponds to the miRNAs that are regulated by p53 in breast cancer. Arrow indicates activation and bar indicates inhibition. The references for each miRNA are marked as superscript in green color. 25(Druz et al., 2013), 26(Rivas et al., 2012), 27(Srivastava et al., 2011), 28(Zhang et al., 2011), 32(Li et al., 2013), 42(Feliciano et al., 2013), 56(Frankel et al., 2008), 65(Zhang et al., 2013), 70(Zhang et al., 2010), 87(Hu et al., 2010), 88(Okamura et al., 2001), 89(Nagpal et al., 2013), 100(Ru et al., 2011), 103(He et al., 2013), 107(Yu et al., 2014), 154(Zhang et al., 2016), 155(Zhou et al., 2015).
A number of BCL2 family members were found to be targeted by apoptomiRs by various groups. BCL2 family consists of both pro and anti-apoptotic proteins. BCL2, an antiapototic protein, was found to be targeted by four miRNAs (miR-21, -24, -34a, and -16) indicating that its levels are kept under tight control in breast tissue. Overexpression of miR-24, 34a, and 16 reduces BCL2 expression that leads to apoptosis through disruption of the mitochondrial membrane potential and the release of cytochrome C (Srivastava et al., 2011; Rivas et al., 2012; Li et al., 2013). Other anti-apoptotic members of the BCL2 family are also directly targeted by apoptomiRs- (BCL2L1- mir-15a-3p, BCL2L2- miR-497, MCL1- miR-26a; Shen et al., 2012; Druz et al., 2013; Gao et al., 2013). The level of BAK1, a pro-apoptotic BCL2 family protein, was found to be a direct target of oncomiR, miR-125b. PUMA, belonging to the BH3-only subgroup of BCL2 family protein was also found to be targeted by miR-221/222 (Zhang et al., 2010). miR-17/20 transduction of MCF7 cells has been shown to induce p53 and apoptosis characterized by activation of BAX, release of cytochrome C, and induction of BCL-X(S) in response to DNA damaging agents, like doxorubicin and tamoxifen. Overall, a multipronged attack by miRNAs seems to directly or indirectly target BCL2 family proteins or downstream targets in breast cancer.
CASP3 is a mediator of apoptosis that gets activated in apoptotic cell by both death ligand (extrinsic) and mitochondrial (intrinsic) pathways. It was found to be a direct target of miR-221/222 in breast cancer (Miller et al., 2008). Several cell cycle related genes (CCND1, CCNE1, CCNJ, CDK9, CDKN1B, and PSME3) also feature in the list suggesting important roles these apoptomiRs may play in maintaining balance between apoptosis and cell cycle. Finally, apoptomiRs target crucial genes involved mainly in the apoptotic or cell cycle related pathways in breast cancer. Fishing out more targets using advanced technologies will likely widen the range of the pathways affected by these targets.
miRNAs and Drug Resistance in Breast Cancer
The commonly used drugs for breast cancer treatment are Cyclophosphamide, Docetaxel, Doxorubicin, Epirubicin, Methotrexate, Paclitaxel, Cisplatin, Carboplatin, 5 Fluorouracil, Gemcitabine, etc. The patients are also given treatment based on the receptor status. The patients are tested for ER, Her2, and PR status. ER+ve patients receive tamoxifen (antagonist of ER signaling) and Her2+ve patients receive trastuzumab (herceptin), a monoclonal antibody against Her2, included in the chemotherapy regimen (Osborne, 1998; Ariazi et al., 2006). While the response of breast cancer patients to radiation, drug, or hormonal treatment has considerably improved the patient disease free survival and overall survival, chances of recurrence still remain. Since apoptosis mediated cancer cell death remains central to all the treatment strategies, the recurrence has been attributed to the development of resistance to apoptotic death and maintenance of cell viability by undergoing viable cellular responses such as cellular senescence (generates secretomes which can directly enhance the malignant phenotype) or autophagy (a state of reversible dormancy making it possible for cancer cells to survive and regrow at a later stage; Degenhardt et al., 2006).
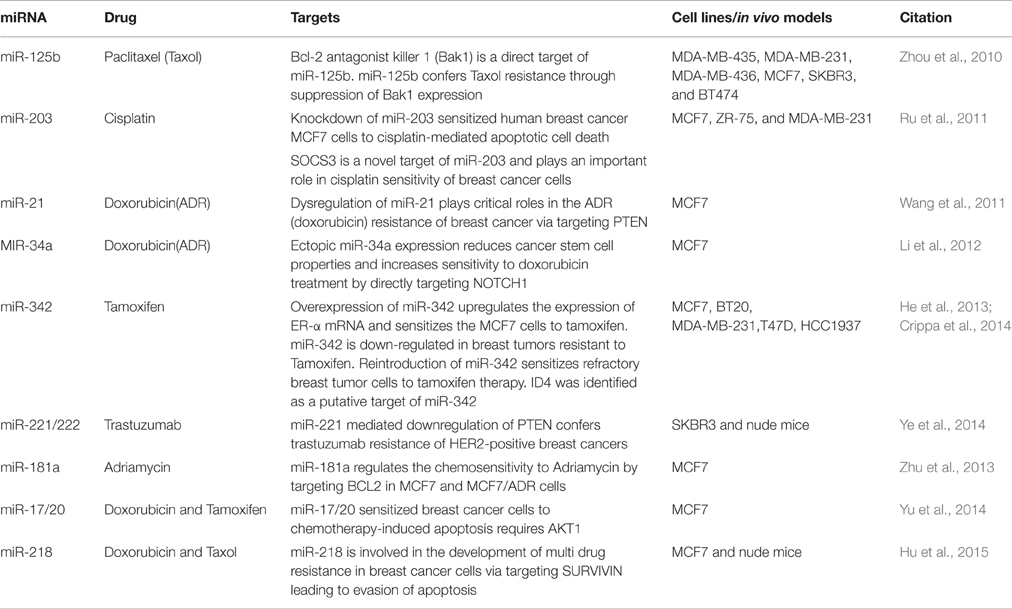
Several mechanisms of chemoresistance in breast cancer have been reported in breast cancer such as upregulation of the ATP-binding cassette (ABC) transporters, drug inactivation or detoxification, alterations in genes related to cell cycle, apoptosis or DNA repair enzymes, epigenetic modifications, and activation of signaling pathways related to the progression of cancer (Baker et al., 2005; Pogribny et al., 2010). Recently, alterations of miRNA levels too have been linked to the development of drug resistance in breast cancer (Kovalchuk et al., 2008; Baylin, 2011). Very few miRNAs (miR-17/20, -21, -34a, -125b, -181a, -203, -218, -221, -222, -342) have been reported till date to affect the sensitivity to drugs (paclitaxel, cisplatin, doxorubicin, and taxol) or hormonal (tamoxifen and trastuzumab) treatment in various breast cancer cell lines (Table 2). These miRNAs affect the drug sensitivity by interfering in apoptotic death by affecting the levels of various apoptotic regulators (TP53, BCL2, BAX, BAK1, SURVIVIN, NOTCH1), members or regulators of AKT signaling pathway (AKT1, PTEN), ID4, NOTCH1 and SOCS3. Considering that the levels of several miRNAs change in response to drug treatment or are differentially regulated between sensitive versus resistant breast cancer cell lines, it is very likely that the list of miRNAs associated with drug resistance will expand. Some of these miRNAs may also function as biomarkers in predicting and monitoring drug response in breast cancer patients (Nana-Sinkam and Croce, 2013). These miRNAs are also being evaluated for their use in combination therapies to counteract drug resistance (Nana-Sinkam and Croce, 2013).
ApoptomiRs as Biomarkers and Therapeutic Targets in Breast Cancer
Early detection of breast cancer is important to reduce the risk of disease. Current diagnostic methods for early detection of breast cancer are still limited to some procedures such as tissue biopsies and histological examination using mRNA or protein biomarkers. However, there is still need for sensitive and specific markers. As miRNAs have small size, high specificity, and greater stability, they can be used as biomarkers with diagnostic (based on their ability to discriminate between normal and breast cancer patients), predictive [which are involved in response (sensitivity/resistance) to conventional breast cancer therapeutic strategies] or prognostic (which predict patient overall or disease-free survival) potential (Krutovskikh and Herceg, 2010; Abba et al., 2012; Schooneveld et al., 2015). Several studies have shown the role of miRNAs in diagnosis and prognosis of breast cancer. Iorio et al. (2005) identified a 13 miRNA signature that could differentiate breast cancer from normal breast tissue with 100% accuracy (Iorio et al., 2005). Blenkiron et al. (2007) identified 133 miRNAs that displayed aberrant expression levels in breast tumor tissues compared with normal breast tissues (Blenkiron et al., 2007). Recently several reports suggested that circulating miRNAs are highly stable in serum/plasma and levels of some miRNAs are specifically elevated in patients with breast cancer (Heneghan et al., 2010). In a recent study, nine circulating miRNAs were shown to be capable of discriminating between early-stage breast cancer and healthy controls (Kodahl et al., 2014). Another study reported three significantly overexpressed serum miRNAs in breast cancer patients as compared to breast cancer free controls (Ng et al., 2013). Recently, Bertoli et al. (2015) and Schooneveld et al. (2015) reported circulating and non-circulating miRNAs as breast cancer biomarkers. Several of the apoptomiRs have also been shown to serve as biomarkers in breast cancer (Abba et al., 2012; Bertoli et al., 2015; Figure 3). While some apoptomiRs have only diagnostic (miR-15a/16, -17, -181a, -145, -146) or predictive (miR-26a/b, -181b, -502) or prognostic (miR-497) potential, some have been proposed to serve as biomarkers for any two (miR-7, -34, -125a/b, -155, -222, -342) or all the three purposes (miR-21). Based on their presence and high stability in patient serum, capacity to reflect tumor status, and high clinical outcome, some apoptomiRs (miR-155, -21, -16, -222, and -210) are present as circulating miRNAs and can be used as potential biomarkers for breast cancer. miR-210 is a hypoxia regulated anti-apoptotic miRNA in breast cancer and high miR-210 baseline expression has been associated with poor prognosis and resistance to trastuzumab-included chemotherapy (Jung et al., 2012) High miR-155 levels too have been reported in tumor tissues sections and the serum of breast cancer patients and have been correlated to poor prognosis (Kong et al., 2014). Similarly, miR-21 upregulation is associated with taxol resistance and poor prognosis in breast cancer cells (Mei et al., 2010). miR-34 has been proposed to serve as a predictive marker for response to radiotherapy (Stankevicins et al., 2013). Kato et al. reported that low levels of miR-34a rendered breast cancer cells more resistant to radiotherapy (Kato et al., 2009).
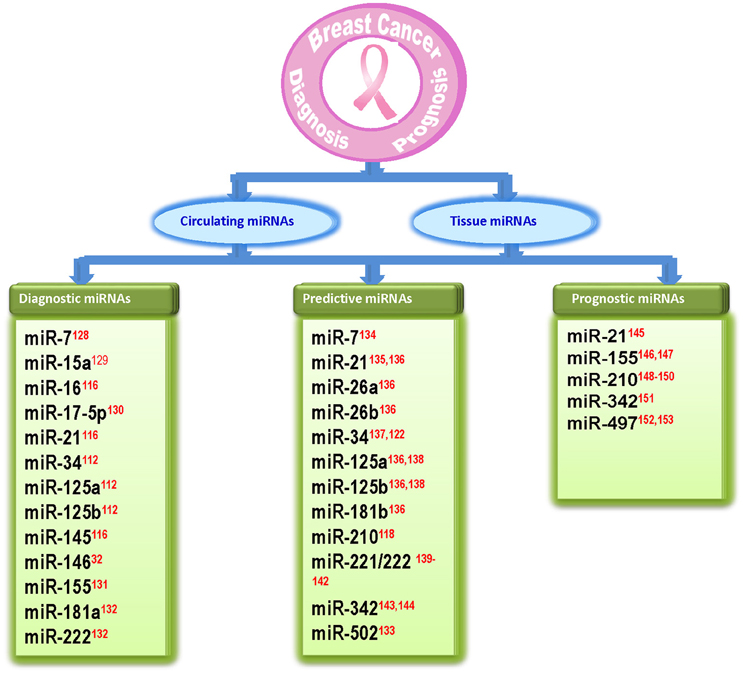
Figure 3. Diagram showing apoptosis related miRNAs that are identified as biomarkers in breast cancer. The references for each miRNA are marked as superscript in red color. 32(Li et al., 2013), 112(Iorio et al., 2005), 116(Ng et al., 2013), 118(Jung et al., 2012), 122(Kato et al., 2009), 128(Foekens et al., 2008), 129(Kodahl et al., 2014), 130(Volinia et al., 2006), 131(Wang F. et al., 2014), 132(Godfrey et al., 2013), 133(Lyng et al., 2012), 134(Chen and Bourguignon, 2014), 135(Mei et al., 2010), 136(Maillot et al., 2009), 137(Stankevicins et al., 2013), 138(Valabrega et al., 2007), 139(Zhao et al., 2008), 140(Wei et al., 2014), 141(Gan et al., 2014), 142(Rao et al., 2011), 143(He et al., 2013), 144(Cittelly et al., 2010), 145(Lee et al., 2011), 146(Song et al., 2012), 147(Kong et al., 2014), 148(Volinia et al., 2012), 149 (Li M. et al., 2014), 150(Wang J. et al., 2014), 151(Leivonen et al., 2014), 152(Shen et al., 2012), 153(Wang et al., 2013).
While successful miRNA delivery still remains a challenge, the prospects of using miRNAs as means and targets for breast cancer therapy remain attractive. There are two lines of action for therapeutic use of miRNAs: miRNA antagonists for miRNA inhibition and miRNA mimics for miRNA replacement therapy (Bader, 2012; Thorsen et al., 2012). Based on in vitro studies several miRNAs have reached preclinical trials and two miRNAs, have entered clinical trials. A locked nucleic acid-based antisense oligonucleotide against miR-122 (Miravirsen), successfully completed phase I clinical study and is undergoing phase II clinical trials for treatment of chronic hepatitis C virus infection (Janssen et al., 2013). In preclinical studies, Liu et al. reported that the injection of miR-34a mimic extended the survival of tumor-bearing mice (Liu C. et al., 2011). Another study demonstrated that systemic administration of miR-34 in a pancreatic xenograft cancer model significantly inhibited tumor growth and induced cancer cell apoptosis (Hu et al., 2013). A clinical trial using miR-34 mimic (MRX34) has already entered phase I clinical trial in patients with unresectable primary liver cancer or metastatic cancer with liver involvement (ClinicalTrials.gov Identifier: NCT01829971). Considering that miR-34 plays a major role in breast tumorigenesis and shows promising results in preclinical trials, miR-34 may soon enter clinical trials for treatment of breast cancer patients.
Conclusions
Mechanisms of cell death and its regulation during initiation, progression, and treatment of breast cancer are complex and still remain partially understood. The role of miRNAs as critical mediators of apoptotic cell death has been a revelation in the past decade and has given us means to manipulate cell death for breast cancer treatment or for disease monitoring. The current impediments for successful treatment are miRNA delivery issues, concerns over off-target effects and our incomplete knowledge of the functions of these miRNAs. Many of these miRNAs may have uninvestigated roles in other forms of cell death that may change the output based on the balance between the desired (apoptosis) and undesired (autophagy and necrosis) forms of cell death for breast cancer treatment. Nevertheless, miRNA delivery issues are slowly getting resolved, miRNAs are also being considered as means to target multiple aberrant pathways and certainly preclinical and clinical trials of some of these miRNAs look promising. The pool of apoptomiRs in breast cancer is likely to get bigger with advanced technologies such as large scale miRNA functional screens using 3D tumor models, thus giving us more probable candidates as novel targets for breast cancer therapy.
Author Contributions
RK conceptualized the theme of the review and has written the review. SS compiled all the data on pro- and anti-apoptotic miRNAs and their correlation to drug resistance and has contributed in writing the review. PP and SA compiled the data of miRNA and p53 regulatory loop and contributed toward writing this section.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The reviewer KT and handling editor declared their shared affiliation, and the handling editor states that the process nevertheless met the standards of a fair and objective review.
Acknowledgments
This work was supported by the grant SB/S0/BB-0088/2013 from Science and Engineering Research Board, Department of Science & Technology, Government of India to RK. SS thanks Centre for Scientific and Industrial Research for Senior Research Fellowship.
References
Abba, M., Mudduluru, G., and Allgayer, H. (2012). Micrornas in cancer: small molecules, big chances. Anticancer Agents Med. Chem. 12, 733–743. doi: 10.2174/187152012802650273
Akhavantabasi, S., Sapmaz, A., Tuna, S., and Erson-Bensan, A. E. (2012). miR-125b targets ARID3B in breast cancer cells. Cell Struct. Funct. 37, 27–38. doi: 10.1247/csf.11025
American Cancer Society (2015). Breast Cancer Facts & Figures 2015-2016. Atlanta: American Cancer Society.
Antonio, M., Sanchez, V., Liu, J., Hannon, G. J., and Parker, R. (2006). Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev. 20, 515–524. doi: 10.1101/gad.1399806
Ariazi, E. A., Ariazi, J. L., Cordera, F., and Jordan, V. C. (2006). Estrogen receptors as therapeutic targets in breast cancer. Curr. Top. Med. Chem. 6, 181–202. doi: 10.2174/156802606776173483
Bader, A. G. (2012). miR-34 - a microRNA replacement therapy is headed to the clinic. Front. Genet. 3:120. doi: 10.3389/fgene.2012.00120
Baker, E. K., Johnstone, R. W., Zalcberg, J. R., and El-Osta, A. (2005). Epigenetic changes to the MDR1 locus in response to chemotherapeutic drugs. Oncogene 24, 8061–8075. doi: 10.1038/sj.onc.1208955
Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297. doi: 10.1016/S0092-8674(04)00045-5
Baylin, S. B. (2011). Resistance, epigenetics and the cancer ecosystem. Nat. Med. 17, 288–289. doi: 10.1038/nm0311-288
Bertoli, G., Cava, C., and Castiglioni, I. (2015). MicroRNAs: new biomarkers for diagnosis, prognosis, therapy prediction and therapeutic tools for breast cancer. Theranostics 5, 1122–1143. doi: 10.7150/thno.11543
Blenkiron, C., Goldstein, L. D., Thorne, N. P., Spiteri, I., Chin, S. F., Dunning, M. J., et al. (2007). MicroRNA expression profiling of human breast cancer identifies new markers of tumor subtype. Genome Biol. 8:R214. doi: 10.1186/gb-2007-8-10-r214
Chen, L., and Bourguignon, L. Y. (2014). Hyaluronan-CD44 interaction promotes c-Jun signaling and miRNA21 expression leading to Bcl-2 expression and chemoresistance in breast cancer cells. Mol. Cancer 13:52. doi: 10.1186/1476-4598-13-52
Cittelly, D. M., Das, P. M., Spoelstra, N. S., Edgerton, S. M., Richer, J. K., et al. (2010). Downregulation of miR-342 is associated with tamoxifen resistant breast tumors. Mol. Cancer 9:317. doi: 10.1186/1476-4598-9-317
Crippa, E., Lusa, L., De Cecco, L., Marchesi, E., Calin, G. A., Radice, P., et al. (2014). miR-342 regulates BRCA1 expression through modulation of ID4 in breast cancer. PLoS ONE 9:e87039. doi: 10.1371/journal.pone.0087039
Crosby, M. E., Kulshreshtha, R., Ivan, M., and Glazer, P. M. (2009). MicroRNA regulation of DNA repair gene expression in hypoxic stress. Cancer Res. 69, 1221–1229. doi: 10.1158/0008-5472.CAN-08-2516
Degenhardt, K., Mathew, R., Beaudoin, B., Bray, K., Anderson, D., et al. (2006). Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 10, 51–64. doi: 10.1016/j.ccr.2006.06.001
Deveraux, Q. L., and Reed, J. C. (1999). IAP family proteins—suppressors of apoptosis. Genes Dev. 13, 239–252. doi: 10.1101/gad.13.3.239
Druz, A., Chen, Y. C., Guha, R., Betenbaugh, M., Martin, S. E., and Shiloach, J. (2013). Large-scale screening identifies a novel microRNA, miR-15a-3p, which induces apoptosis in human cancer cell lines. RNA Biol. 10, 287–300. doi: 10.4161/rna.23339
Elmore, S. (2007). Apoptosis: a review of programmed cell death. Toxicol. Pathol. 35, 495–516. doi: 10.1080/01926230701320337
Ergun, S., and Oztuzcu, S. (2014). miR-221: a critical player in apoptosis as a target of Caspase-3. Cancer Cell Microenviron. 1:e313. doi: 10.14800/ccm.313
Fasanaro, P., Greco, S., Lorenzi, M., Pescatori, M., Brioschi, M., Kulshreshtha, R., et al. (2009). An integrated approach for experimental target identification of hypoxia-induced miR-210. J. Biol. Chem. 284, 35134–35143. doi: 10.1074/jbc.m109.052779
Feliciano, A., Castellvi, J., Artero-Castro, A., Leal, J. A., Romagosa, C., Hernández-Losa, J., et al. (2013). miR-125b acts as a tumor suppressor in breast tumorigenesis via its novel direct targets ENPEP, CK2-α, CCNJ, and MEGF9. PLoS ONE 8:e76247. doi: 10.1371/journal.pone.0076247
Ferlay, J., Soerjomataram, I., Dikshit, R., Eser, S., Mathers, C., Rebelo, M., et al. (2015). Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, 359–386. doi: 10.1002/ijc.29210
Ferracin, M., Bassi, C., Pedriali, M., Pagotto, S., D'Abundo, L., Zagatti, B., et al. (2013). miR-125b targets erythropoietin and its receptor and their expression correlates with metastatic potential and ERBB2/HER2 expression. Mol. Cancer 12:130. doi: 10.1186/1476-4598-12-130
Florea, A.-M., and Busselberg, D. (2013). Breast cancer and possible mechanisms of therapy resistance. J. Local Glob. Health Sci. 2:9. doi: 10.5339/jlghs.2013.2
Foekens, J. A., Sieuwerts, A. M., Smid, M., et al. (2008). Four miRNAs associated with aggressiveness of lymph node-negative, estrogen receptor-positive human breast cancer. Proc. Natl. Acad. Sci. U.S.A. 105, 13021–13026. doi: 10.1073/pnas.0803304105
Frankel, L. B., Christoffersen, N. R., Jacobsen, A., Lindow, M., Krogh, A., and Lund, A. H. (2008). Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J. Biol. Chem. 283, 1026–1033. doi: 10.1074/jbc.M707224200
Frenzel, A., Grespi, F., Chmelewskij, W., and Villunger, A. (2009). Bcl2 family proteins in carcinogenesis and the treatment of cancer. Apoptosis 14, 584–596. doi: 10.1007/s10495-008-0300-z
Fulda, S., and Debatin, K.-M. (2006). Extrinsic versus intrinsic apoptosis pathways in anticancer chemotherapy. Oncogene 25, 4798–4811. doi: 10.1038/sj.onc.1209608
Gan, R., Yang, Y., Yang, X., Zhao, L., Lu, J., and Meng, Q. H. (2014). Downregulation of miR-221/222 enhances sensitivity of breast cancer cells to tamoxifen through upregulation of TIMP3. Cancer Gene Ther. 21, 290–296. doi: 10.1038/cgt.2014.29
Gao, J., Li, L., Wu, M., Liu, M., Xie, X., Guo, J., et al. (2013). MiR-26a inhibits proliferation and migration of breast cancer through repression of MCL-1. PLoS ONE 8:e65138. doi: 10.1371/journal.pone.0065138
Godfrey, A. C., Xu, Z., Weinberg, C. R., Getts, R. C., Wade, P. A., DeRoo, L. A., et al. (2013). Serum microRNA expression as an early marker for breast cancer risk in prospectively collected samples from the Sister Study cohort. Breast Cancer Res. 15, R42. doi: 10.1186/bcr3428
Goldberger, N., Walker, R. C., Kim, C. H., Winter, S., and Hunter, K. W. (2013). Inherited variation in miR-290 expression suppresses breast cancer progression by targeting the metastasis susceptibility gene Arid4b. Cancer Res. 73, 2671–2681. doi: 10.1158/0008-5472.CAN-12-3513
Gonzalez-Angulo, A. M., Morales-Vasquez, F., and Hortobagyi, G. N. (2007). Overview of resistance to systemic therapy in patients with breast cancer. Adv. Exp. Med. Biol. 608, 1–22. doi: 10.1007/978-0-387-74039-3_1
Guo, X., Wu, Y., and Hartley, R. S. (2009). MicroRNA-125a represses cell growth by targeting HuR in breast cancer. RNA Biol. 6, 575–583. doi: 10.4161/rna.6.5.10079
Guttilla, I. K., and White, B. A. (2009). Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. J. Biol. Chem. 284, 23204–23216. doi: 10.1074/jbc.M109.031427
Hammond, S. M. (2006). microRNA detection comes of age. Nat. Methods 3, 12–13. doi: 10.1038/nmeth0106-12
He, T. Y. G., Yang, L., Yang, G., Chen, Y., and Zhang, X. (2015). The role of miR-100 in regulating apoptosis of breast cancer cells. Sci. Rep. 5:11650. doi: 10.1038/srep11650
He, Y. J., Wu, J. Z., Ji, M. H., Ma, T., Qiao, E. Q., Ma, R., et al. (2013). miR-342 is associated with estrogen receptor-α expression and response to tamoxifen in breast cancer. Exp. Ther. Med. 5, 813–818. doi: 10.3892/etm.2013.915
Heneghan, H. M., Miller, N., Lowery, A. J., Sweeney, K. J., Newell, J., et al. (2010). Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann. Surg. 251, 499–505. doi: 10.1097/SLA.0b013e3181cc939f
Hengartner, M. O. (2001). Apoptosis: corralling the corpses. Cell 104, 325–328. doi: 10.1016/S0092-8674(01)00219-7
Honma, N., Horii, R., Ito, Y., Saji, S., Younes, M., et al. (2015). Differences in clinical importance of Bcl-2 in breast cancer according to hormone receptors status or adjuvant endocrine therapy. BMC Cancer. 15:698. doi: 10.1186/s12885-015-1686-y
Hu, Q. L., Jiang, Q. Y., Jin, X., Shen, J., Wang, K., Li, Y. B., et al. (2013). Cationic microRNA-delivering nanovectors with bifunctional peptides for efficient treatment of PANC-1 xenograft model. Biomaterials 34, 2265–2276. doi: 10.1016/j.biomaterials.2012.12.016
Hu, W., Chan, C. S., Wu, R., Zhang, C., Sun, Y., Song, J. S., et al. (2010). Negative regulation of tumor suppressor p53 by microRNA miR-504. Mol. Cell 38, 689–699. doi: 10.1016/j.molcel.2010.05.027
Hu, Y., Xu, K., and Yagüe, E. (2015). miR-218 targets survivin and regulates resistance to chemotherapeutics in breast cancer. Breast Cancer Res. Treat. 151, 269–280. doi: 10.1007/s10549-015-3372-9
Hu, Y., Zhu, Q., and Tang, L. (2014). MiR-99a antitumor activity in human breast cancer cells through targeting of mTOR expression. PLoS ONE 9:e92099. doi: 10.1371/journal.pone.0092099
Huang, D. W., Sherman, B. T., and Lempicki, R. A. (2009a). Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protoc. 4, 44–57. doi: 10.1038/nprot.2008.211
Huang, D. W., Sherman, B. T., and Lempicki, R. A. (2009b). Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13. doi: 10.1093/nar/gkn923
Iorio, M. V., Ferracin, M., Liu, C. G., et al. (2005). MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 65, 7065–7070. doi: 10.1158/0008-5472.CAN-05-1783
Janaki Ramaiah, M., Lavanya, A., Honarpisheh, M., Zarea, M., Bhadra, U., and Bhadra, M. P. (2014). MiR-15/16 complex targets p70S6 kinase 1 and controls cell proliferation in MDA-MB-231 breast cancer cells. Gene 552, 255–264. doi: 10.1016/j.gene.2014.09.052
Janssen, H. L., Reesink, H. W., Lawitz, E. J., Zeuzem, S., Rodriguez-Torres, M., Patel, K., et al. (2013). Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 368, 1685–1694. doi: 10.1056/NEJMoa1209026
Jiang, S., Zhang, H. W., Lu, M. H., He, X. H., et al. (2010). MicroRNA-155 functions as an OncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer Res. 70, 3119–3127. doi: 10.1158/0008-5472.CAN-09-4250
Joshi-Tope, G., Gillespie, M., Vastrik, I., D'Eustachio, P., Schmidt, E., et al. (2005). Reactome: a knowledgebase of biological pathways. Nucleic Acids Res. 33, D428–D432. doi: 10.1093/nar/gki072
Jung, E. J., Santarpia, L., Kim, J., Esteva, F. J., Moretti, E., et al. (2012). Plasma microRNA 210 levels correlate with sensitivity to trastuzumab and tumor presence in breast cancer patients. Cancer 118, 2603–2614. doi: 10.1002/cncr.26565
Kasibhatla, S., and Tseng, B. (2003). Why target apoptosis in cancer treatment? Mol. Cancer Ther. 2, 573–580.
Kato, M., Paranjape, T., Müller, R. U., Nallur, S., Gillespie, E., Keane, K., et al. (2009). The mir-34 microRNA is required for the DNA damage response in vivo in C. elegans and in vitro in human breast cancer cells. Oncogene 28, 2419–2424. doi: 10.1038/onc.2009.106
Kodahl, A. R., Lyng, M. B., Binder, H., Cold, S., Gravgaard, K., Knoop, A. S., et al. (2014). Novel circulating microRNA signature as a potential non-invasive multi-marker test in ER-positive early-stage breast cancer: a case control study. Mol. Oncol. 8, 874–883. doi: 10.1016/j.molonc.2014.03.002
Kong, W., He, L., Coppola, M., Guo, J., Esposito, N. N., Coppola, D., et al. (2010). MicroRNA-155 regulates cell survival, growth, and chemosensitivity by targeting FOXO3a in breast cancer. J. Biol. Chem. 285, 17869–17879. doi: 10.1074/jbc.M110.101055
Kong, W., He, L., Richards, E. J., Challa, S., Xu, C. X., Permuth-Wey, J., et al. (2014). Upregulation of miRNA-155 promotes tumour angiogenesis by targeting VHL and is associated with poor prognosis and triple-negative breast cancer. Oncogene 33, 679–689. doi: 10.1038/onc.2012.636
Kong, W., Yang, H., He, L., Zhao, J. J., Coppola, D., et al. (2008). MicroRNA-155 is regulated by the transforming growth factor β/smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol. Cell Biol. 28, 6773–6784. doi: 10.1128/MCB.00941-08
Körner, C., Keklikoglou, I., Bender, C., Wörner, A., Münstermann, E., and Wiemann, S. (2013). MicroRNA-31 sensitizes human breast cells to apoptosis by direct targeting of protein kinase C epsilon (PKCepsilon). J. Biol. Chem. 288, 8750–8761. doi: 10.1074/jbc.M112.414128
Kovalchuk, O., Filkowski, J., Meservy, J., Ilnytskyy, Y., et al. (2008). Involvement of microRNA-451 in resistance of the MCF7 breast cancer cells to chemotherapeutic drug doxorubicin. Mol. Cancer Ther. 7, 2152–2159. doi: 10.1158/1535-7163.MCT-08-0021
Krutovskikh, V. A., and Herceg, Z. (2010). Oncogenic microRNAs (OncomiRs) as a new class of cancer biomarkers. Bioessays 32, 894–904. doi: 10.1002/bies.201000040
Lee, J. A., Lee, H. Y., Lee, E. S., Kim, I., and Bae, J. W. (2011). Prognostic implications of microRNA-21 overexpression in invasive ductal carcinomas of the breast. J. Breast Cancer 14:269. doi: 10.4048/jbc.2011.14.4.269
Leivonen, S. K., Sahlberg, K. K., Mäkelä, R., Due, E. U., et al. (2014). High-throughput screens identify microRNAs essential for HER2 positive breast cancer cell growth. Mol. Oncol. 8, 93–104. doi: 10.1016/j.molonc.2013.10.001
Li, B., Lu, Y., Wang, H., Han, X., Mao, J., Li, J., et al. (2016). miR-221/222 enhance the tumorigenicity of human breast cancer stem cells via modulation of PTEN/Akt pathway. Biomed. Pharmacother. 79, 93–101. doi: 10.1016/j.biopha.2016.01.045
Li, J. H., Liu, S., Zhou, H., Qu, L. H., and Yang, J. H. (2014). starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42, D92–D97. doi: 10.1093/nar/gkt1248
Li, L., Yuan, L., Luo, J., Gao, J., Guo, J., and Xie, X. (2013). MiR-34a inhibits proliferation and migration of breast cancer through down-regulation of Bcl-2 and SIRT1. Clin. Exp. Med. 13, 109–117. doi: 10.1007/s10238-012-0186-5
Li, M., Ma, X., Li, M., Zhang, B., et al. (2014). Prognostic role of microRNA-210 in various carcinomas: a syst ematic review and meta-analysis. Dis. Markers. 2014:106197. doi: 10.1155/2014/106197
Li, W., Zang, W., Liu, P., Wang, Y., Du, Y., Chen, X., et al. (2014). MicroRNA-124 inhibits cellular proliferation and invasion by targeting Ets-1 in breast cancer. Tumour Biol. 35, 10897–10904. doi: 10.1007/s13277-014-2402-2
Li, X. J., Ji, M. H., Zhong, S. L., Zha, Q. B., et al. (2012). MicroRNA-34a modulates chemosensitivity of breast cancer cells to adriamycin by targeting Notch1. Arch. Med. Res. 43, 514–521. doi: 10.1016/j.arcmed.2012.09.007
Lipponen, P. (1999). Apoptosis in breast cancer: relationship with other pathological parameters. Endocr. Relat. Cancer 6, 13–16. doi: 10.1677/erc.0.0060013
Lipponen, P., Aaltomaa, S., Kosma, V.-M., and Syrjänen, K. (1994). Apoptosis in breast cancer as related to histopathological characteristics and prognosis. Eur. J. Cancer 30A, 2068–2073. doi: 10.1016/0959-8049(94)00342-3
Liu, C., Kelnar, K., Liu, B., Chen, X., Calhoun-Davis, T., Li, H., et al. (2011). The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat. Med. 17, 211–215. doi: 10.1038/nm.2284
Liu, H., Wang, Y., Li, X., Zhang, Y. J., Li, J., Zheng, Y. Q., et al. (2013). Expression and regulatory function of miRNA-182 in triple-negative breast cancer cells through its targeting of profilin 1. Tumour Biol. 34, 1713–1722. doi: 10.1007/s13277-013-0708-0
Liu, R., Liu, C., Chen, D., Yang, W. H., Liu, X., Liu, C. G., et al. (2015). FOXP3 controls an miR-146/NF-κB negative feedback loop that inhibits apoptosis in breast cancer cells. Cancer Res. 75, 1703–1713. doi: 10.1158/0008-5472.CAN-14-2108
Liu, X. X., Li, X. J., Zhang, B., Liang, Y. J., Zhou, C. X., Cao, D. X., et al. (2011). MicroRNA-26b is underexpressed in human breast cancer and induces cell apoptosis by targeting SLC7A11. FEBS Lett. 585, 1363–1367. doi: 10.1016/j.febslet.2011.04.018
Lu, K., Wang, J., Song, Y., Zhao, S., Liu, H., et al. (2015). miRNA-24-3p promotes cell proliferation and inhibits apoptosis in human breast cancer by targeting p27Kip1. Oncol. Rep. 34, 995–1002. doi: 10.3892/or.2015.4025
Luo, E. C., Chang, Y. C., Sher, Y. P., Huang, W. Y., Chuang, L. L., et al. (2014). MicroRNA-769-3p down-regulates NDRG1 and enhances apoptosis in MCF7 cells during reoxygenation. Sci. Rep. 4:e5908. doi: 10.1038/srep05908
Luo, Q., Li, X., Li, J., Kong, X., Zhang, J., Chen, L., et al. (2013). MiR-15a is underexpressed and inhibits the cell cycle by targeting CCNE1 in breast cancer. Int. J. Oncol. 43, 1212–1218. doi: 10.3892/ijo.2013.2034
Luthra, R., Singh, R. R., Luthra, M. G., Li, Y. X., Hannah, C., Romans, A. M., et al. (2008). MicroRNA-196a targets annexin A1: a microRNA-mediated mechanism of annexin A1 downregulation in cancers. Oncogene 27, 6667–6678. doi: 10.1038/onc.2008.256
Lyng, M. B., Laenkholm, A. V., Sokilde, R., et al. (2012). Global microRNA expression profiling of high-risk ER+ breast cancers from patients receiving adjuvant tamoxifen mono-therapy: a DBCG study. PLoS ONE 7:e36170. doi: 10.1371/journal.pone.0036170
Mackiewicz, M., Huppi, K., Pitt, J. J., Dorsey, T. H., Ambs, S., and Caplen, N. J. (2011). Identification of the receptor tyrosine kinase AXL in breast cancer as a target for the human miR-34a microRNA. Breast Cancer Res. Treat. 130, 663–679. doi: 10.1007/s10549-011-1690-0
Maillot, G., Lacroix-Triki, M., Pierredon, S., et al. (2009). Widespread estrogen-dependent repression of micrornas involved in breast tumor cell growth. Cancer Res. 69, 8332–8340. doi: 10.1158/0008-5472.CAN-09-2206
Mattick, J. S., and Makunin, I. V. (2006). Non-coding RNA. Hum. Mol. Genet. 15, R17–R29. doi: 10.1093/hmg/ddl046
Megha, T., Ferrari, F., Benvenuto, A., Bellan, C., Lalinga, A. V., Lazzi, S., et al. (2002). p53 mutation in breast cancer. Correlation with cell kinetics and cell of origin. J. Clin. Pathol. 55, 461–466. doi: 10.1136/jcp.55.6.461
Mei, M., Ren, Y., Zhou, X., Yuan, X. B., Han, L., Wang, G. X., et al. (2010). Downregulation of miR-21 enhances chemotherapeutic effect of taxol in breast carcinoma cells. Technol. Cancer Res. Treat. 9, 77–86. doi: 10.1177/153303461000900109
Miller, T. E., Ghoshal, K., Ramaswamy, B., Roy, S., Datta, J., et al. (2008). MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J. Biol. Chem. 283, 29897–29903. doi: 10.1074/jbc.M804612200
Molloy, N. H., Read, D. E., and Gorman, A. M. (2011). Nerve growth factor in cancer cell death and survival. Cancers 3, 510–530. doi: 10.3390/cancers3010510
Mosca, E., Alfieri, R., Merelli, I., Viti, F., Calabria, A., and Milanesi, L. (2010). A multilevel data integration resource for breast cancer study. BMC Syst. Biol. 4:76. doi: 10.1186/1752-0509-4-76
Nagpal, N., Ahmad, H. M., Molparia, B., and Kulshreshtha, R. (2013). MicroRNA- 191, an estrogen-responsive microRNA, functions as an oncogenic regulator in human breast cancer. Carcinogenesis 34, 1889–1899. doi: 10.1093/carcin/bgt107
Nana-Sinkam, S. P., and Croce, C. M. (2013). Clinical applications for microRNAs in cancer. Clin. Pharmacol. Ther. 93, 98–104. doi: 10.1038/clpt.2012.192
Ng, E. K., Li, R., Shin, V. Y., Jin, H. C., Leung, C. P., Ma, E. S., et al. (2013). Circulating microRNAs as specific biomarkers for breast cancer detection. PLoS ONE 8:e53141. doi: 10.1371/journal.pone.0053141
Okamura, S., Arakawa, H., Tanaka, T., Nakanishi, H., et al. (2001). p53DINP1, a p53-inducible gene, regulates p53-dependent apoptosis. Mol. Cell 8, 85–94. doi: 10.1016/S1097-2765(01)00284-2
Osborne, C. K. (1998). Tamoxifen in the treatment of breast cancer. N. Engl. J. Med. 339, 1609–1618. doi: 10.1056/NEJM199811263392207
Park, E. Y., Chang, E., Lee, E. J., Lee, H. W., Kang, H. G., Chun, K. H., et al. (2014). Targeting of miR34a-NOTCH1 axis reduced breast cancer stemness and chemoresistance. Cancer Res. 74, 7573–7582. doi: 10.1158/0008-5472.CAN-14-1140
Pogribny, I. P., Filkowski, J. N., Tryndyak, V. P., Golubov, A., Shpyleva, S. I., and Kovalchuk, O. (2010). Alterations of microRNAs and their targets are associated with acquired resistance of MCF7 breast cancer cells to cisplatin. Int. J. Cancer 127, 1785–1794. doi: 10.1002/ijc.25191
Quail, D. F., and Joyce, J. A. (2013). Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 11, 1423–1437. doi: 10.1038/nm.3394
Rajabi, H., Jin, C., Ahmad, R., McClary, A. C., Joshi, M. D., and Kufe, D. (2010). Mucin 1 oncoprotein expression is suppressed by the miR-125b oncomir. Genes Cancer 1, 62–68. doi: 10.1177/1947601909357933
Rao, X., Di Leva, G., Li, M., Fang, F., Devlin, C., et al. (2011). MicroRNA-221/222 confers breast cancer fulvestrant resistance by regulating multiple signaling pathways. Oncogene 30, 1082–1097. doi: 10.1038/onc.2010.487
Rivas, M. A., Venturutti, L., Huang, Y. W., Schillaci, R., Huang, T. H., and Elizalde, P. V. (2012). Downregulation of the tumor-suppressor miR-16 via progestin-mediated oncogenic signaling contributes to breast cancer development. Breast Cancer Res. 14, R77. doi: 10.1186/bcr3187
Ru, P., Steele, R., Hsueh, E. C., and Ray, R. B. (2011). Anti-miR-203 upregulates SOCS3 expression in breast cancer cells and enhances cisplatin chemosensitivity. Genes Cancer 2, 720–727. doi: 10.1177/1947601911425832
Sachdeva, M., Zhu, S., Wu, F., Wu, H., Walia, V., Kumar, S., et al. (2009). p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc. Natl. Acad. Sci. U.S.A. 106, 3207–3212. doi: 10.1073/pnas.0808042106
Schooneveld, E. V., Wildiers, H., Vergote, I., Vermeulen, P. B., Dirix, L. Y., and Van Laere, S. J. (2015). Dysregulation of microRNAs in breast cancer and their potential role as prognostic and predictive biomarkers in patient management. Breast Cancer Res. 17, 21. doi: 10.1186/s13058-015-0526-y
Shen, L., Li, J., Xu, L., Ma, J., Li, H., Xiao, X., et al. (2012). miR-497 induces apoptosis of breast cancer cells by targeting Bcl-w. Exp. Ther. Med. 3, 475–480. doi: 10.3892/etm.2011.428
Shi, Y., Luo, X., Li, P., Tan, J., Wang, X., et al. (2015). miR-7-5p suppresses cell proliferation and induces apoptosis of breast cancer cells mainly by targeting REGγ. Cancer Lett. 358, 27–36. doi: 10.1016/j.canlet.2014.12.014
Si, M.-L., Zhu, S., Wu, H., Lu, Z., Wu, F., and Mo, Y.-Y. (2007). miR-21-mediated tumor growth. Oncogene 26, 2799–2803. doi: 10.1038/sj.onc.1210083
Silvestrini, R., Benini, E., Veneroni, S., Daidone, M. G., Tomasic, G., et al. (1996). p53 and bcl-2 expression correlates with clinical outcome in a series of node-positive breast cancer patients. J. Clin. Oncol. 14, 1604–1610.
Song, B., Wang, C., Liu, J., Wang, X., Lv, L., Wei, L., et al. (2010). MicroRNA-21 regulates breast cancer invasion partly by targeting tissue inhibitor of metalloproteinase 3 expression. J. Exp. Clin. Cancer Res. 29:29. doi: 10.1186/1756-9966-29-29
Song, C. G., Wu, X. Y., Wang, C., Fu, F. M., and Shao, Z. M. (2012). Correlation of miR-155 on formalin-fixed paraffin embedded tissues with invasiveness and prognosis of breast cancer. Zhonghua Wai Ke Za Zhi 50, 1011–1014. doi: 10.3760/cma.j.issn.0529-5815.2012.11.012
Srivastava, N., Manvati, S., Srivastava, A., Pal, R., Kalaiarasan, P., Chattopadhyay, S., et al. (2011). miR-24-2 controls H2AFX expression regardless of gene copy number alteration and induces apoptosis by targeting antiapoptotic gene BCL-2: a potential for therapeutic intervention. Breast Cancer Res. 13, R39. doi: 10.1186/bcr2861
Stankevicins, L., Almeida da Silva, A. P., Ventura Dos Passos, F., Dos Santos Ferreira, E., Menks Ribeiro, M. C. G., David, M., et al. (2013). MiR-34a is up-regulated in response to low dose, low energy X-ray induced DNA damage in breast cells. Radiat. Oncol. 8:231. doi: 10.1186/1748-717X-8-231
Sun, L. L., Wang, J., Zhao, Z. J., Liu, N., Wang, A. L., et al. (2014). Suppressive role of miR-502-5p in breast cancer via downregulation of TRAF2. Oncol. Rep. 31, 2085–2092. doi: 10.3892/or.2014.3105
Thorsen, S. B., Obad, S., Jensen, N. F., Stenvang, J., and Kauppinen, S. (2012). The therapeutic potential of microRNAs in cancer. Cancer J. 18, 275–284. doi: 10.1097/PPO.0b013e318258b5d6
Vakkala, M., Lahteenmaki, K., Raunio, H., Paakko, P., and Soini, Y. (1999). Apoptosis during breast carcinoma progression. Clin. Cancer Res. 5, 319–324.
Valabrega, G., Montemurro, F., and Aglietta, M. (2007). Trastuzumab: mechanism of action, resistance and future perspectives in HER2-overexpressing breast cancer. Ann. Oncol. 18, 977–984. doi: 10.1093/annonc/mdl475
Volinia, S., Calin, G. A., Liu, C. G., Ambs, S., Cimmino, A., Petrocca, F., et al. (2006). A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. U.S.A. 103, 2257–2261. doi: 10.1073/pnas.0510565103
Volinia, S., Galasso, M., Sana, M. E., Wise, T. F., et al. (2012). Breast cancer signatures for invasiveness and prognosis defined by deep sequencing of microRNA. Proc. Natl. Acad. Sci. U.S.A. 109, 1–6. doi: 10.1073/pnas.1200010109
Von Mering, C., Jensen, L. J., Snel, B., Hooper, S. D., Krupp, M., et al. (2005). STRING: known and predicted protein–protein associations integrated and transferred across organisms. Nucleic Acids Res. 33, D433–D437. doi: 10.1093/nar/gki005
Wang, F., Hou, J., Jin, W., Li, J., Yue, Y., Jin, H., et al. (2014). Increased circulating microRNA-155 as a potential biomarker for breast cancer screening: a meta-analysis. Molecules 19, 6282–6293. doi: 10.3390/molecules19056282
Wang, J., Zhao, J., Shi, M., Ding, Y., et al. (2014). Elevated expression of miR-210 predicts poor survival of cancer patients: a systematic review and meta-analysis. PLoS ONE 9:e89223. doi: 10.1371/journal.pone.0089223
Wang, L., Gao, W., Hu, F., Xu, Z., and Wang, F. (2014). MicroRNA-874 inhibits cell proliferation and induces apoptosis in human breast cancer by targeting CDK9. FEBS Lett. 588, 4527–4535. doi: 10.1016/j.febslet.2014.09.035
Wang, S., Bian, C., Yang, Z., Bo, Y., Li, J., Zeng, L., et al. (2009). miR-145 inhibits breast cancer cell growth through RTKN. Int. J. Oncol. 34, 1461–1466. doi: 10.3892/ijo_00000275
Wang, S., Li, H., Wang, J., and Wang, D. (2013). Expression of microRNA-497 and its prognostic significance in human breast cancer. Diagn Pathol. 8:172. doi: 10.1186/1746-1596-8-172
Wang, Z. X., Lu, B. B., Wang, H., Cheng, Z. X., and Yin, Y. M. (2011). MicroRNA-21 modulates chemosensitivity of breast cancer cells to doxorubicin by targeting PTEN. Arch. Med. Res. 42, 281–290. doi: 10.1016/j.arcmed.2011.06.008
Wei, Y., Lai, X., Yu, S., Chen, S., Ma, Y., Zhang, Y., et al. (2014). Exosomal miR-221/222 enhances tamoxifen resistance in recipient ER-positive breast cancer cells. Breast Cancer Res. Treat. 147, 423–431. doi: 10.1007/s10549-014-3037-0
Xiao, X., Huang, X., Ye, F., Chen, B., Song, C., Wen, J., et al. (2016). The miR-34a-LDHA axis regulates glucose metabolism and tumor growth in breast cancer. Sci. Rep. 6:21735. doi: 10.1038/srep21735
Yamakuchi, M., Ferlito, M., and Lowenstein, C. J. (2008). miR-34a repression of SIRT1 regulates apoptosis. Proc. Natl. Acad. Sci. U.S.A. 105, 13421–13426. doi: 10.1073/pnas.0801613105
Yan, L. X., Wu, Q. N., Zhang, Y., Li, Y. Y., Liao, D. Z., et al. (2011). Knockdown of miR-21 in human breast cancer cell lines inhibits proliferation, in vitro migration and in vivo tumor growth. Breast Cancer Res. 13:e2803. doi: 10.1186/bcr2803
Yang, S., Li, Y., Gao, J., Zhang, T., Li, S., Luo, A., et al. (2013). MicroRNA-34 suppresses breast cancer invasion and metastasis by directly targeting Fra-1. Oncogene 32, 4294–4303. doi: 10.1038/onc.2012.432
Yang, Y., Chaerkady, R., Beer, M. A., Mendell, J. T., and Pandey, A. (2009). Identification of miR-21 targets in breast cancer cells using a quantitative proteomic approach. Proteomics 9, 1374–1384. doi: 10.1002/pmic.200800551
Ye, X., Bai, W., Zhu, H., Zhang, X., Chen, Y., Wang, L., et al. (2014). MiR-221 promotes trastuzumab-resistance and metastasis in HER2-positive breast cancers by targeting PTEN. BMB Rep. 47, 268–273. doi: 10.5483/BMBRep.2014.47.5.165
Yu, Z., Xu, Z., Disante, G., Wright, J., et al. (2014). miR-17/20 sensitization of breast cancer cells to chemotherapy-induced apoptosis requires Akt1. Oncotarget 5, 1083–1090. doi: 10.18632/oncotarget.1804
Zhang, B., Liu, X. X., He, J. R., Zhou, C. X., Guo, M., He, M., et al. (2011). Pathologically decreased miR-26a antagonizes apoptosis and facilitates carcinogenesis by targeting MTDH and EZH2 in breast cancer. Carcinogenesis 32, 2–9. doi: 10.1093/carcin/bgq209
Zhang, C., Liu, J., Tan, C., Yue, X., Zhao, Y., Peng, J., et al. (2016). microRNA-1827 represses MDM2 to to positively regulate tumor suppressor p53 and suppress tumorigenesis. Oncotarget. 7, 8783–8796. doi: 10.18632/oncotarget.7088
Zhang, C., Zhang, J., Zhang, A., Wang, Y., Han, L., et al. (2010). PUMA is a novel target of miR-221/222 in human epithelial cancers. Int. J. Oncol. 37, 1621–1626. doi: 10.3892/ijo_00000816
Zhang, C. M., Zhao, J., and Deng, H. (2013). MiR-155 promotes proliferation of human breast cancer MCF7 cells through targeting tumor protein 53-induced nuclear protein 1. J. Biomed. Sci. 20:79. doi: 10.1186/1423-0127-20-79
Zhang, G., Liu, Z., Cui, G., Wang, X., and Yang, Z. (2014). MicroRNA-486-5p targeting PIM-1 suppresses cell proliferation in breast cancer cells. Tumour Biol. 35, 11137–11145. doi: 10.1007/s13277-014-2412-0
Zhao, G., Guo, J., Li, D., Jia, C., Yin, W., Sun, R., et al. (2013). MicroRNA-34a suppresses cell proliferation by targeting LMTK3 in human breast cancer MCF7 cell line. DNA Cell Biol. 32, 699–707. doi: 10.1089/dna.2013.2130
Zhao, J. J., Lin, J., Yang, H., Kong, W., He, L., Ma, X., et al. (2008). MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J. Biol. Chem. 283, 31079–31086. doi: 10.1074/jbc.M806041200
Zhou, E., Hui, N., Shu, M., Wu, B., and Zhou, J. (2015). Systematic analysis of the p53-related microRNAs in breast cancer revealing their essential roles in the cell cycle. Oncol Lett. 10, 3488–3494. doi: 10.3892/ol.2015.3751
Zhou, M., Liu, Z., Zhao, Y., Ding, Y., Liu, H., et al. (2010). MicroRNA-125b confers the resistance of breast cancer cells to paclitaxel through suppression of pro-apoptotic Bcl-2 antagonist killer 1 (Bak1) expression. J. Biol. Chem. 285, 21496–21507. doi: 10.1074/jbc.M109.083337
Zhu, S., Si, M. L., Wu, H., and Mo, Y. Y. (2007). MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J. Biol. Chem. 282, 14328–14336. doi: 10.1074/jbc.M611393200
Keywords: breast cancer, apoptosis, microRNA, apoptomiR, biomarker
Citation: Sharma S, Patnaik PK, Aronov S and Kulshreshtha R (2016) ApoptomiRs of Breast Cancer: Basics to Clinics. Front. Genet. 7:175. doi: 10.3389/fgene.2016.00175
Received: 15 July 2016; Accepted: 15 September 2016;
Published: 29 September 2016.
Edited by:
Sanjeev Kumar Srivastava, Mitchell Cancer Institute, USAReviewed by:
Vijay Kumar Prajapati, Central University of Rajasthan, IndiaUdayan Bhattacharya, University of South Alabama, USA
Kaushlendra Tripathi, Mitchell Cancer Institute, USA
Copyright © 2016 Sharma, Patnaik, Aronov and Kulshreshtha. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ritu Kulshreshtha, cml0dUBkYmViLmlpdGQuYWMuaW4=; ZHJyaXR1a3Vsc2hyZXNodGhhQGdtYWlsLmNvbQ==
 Shivani Sharma
Shivani Sharma Praveen K. Patnaik
Praveen K. Patnaik Stella Aronov
Stella Aronov Ritu Kulshreshtha
Ritu Kulshreshtha