- 1Department of Organ Transplantation, First Affiliated Hospital of Sun Yat-sen University, Guangzhou, China
- 2Department of Hepatology Surgery, First Affiliated Hospital of Sun Yat-sen University, Guangzhou, China
- 3Department of Gastroenterology, First Affiliated Hospital of Sun Yat-sen University, Guangzhou, China
Hepatocellular carcinoma (HCC) is the second leading cause of cancer-related death worldwide despite the availability of diverse treatment strategies. Much research progress has been made regarding immunotherapy but the effects remain unsatisfactory, highlighting the urgent need for novel immune-related therapy targets. In recent years, more and more studies have pointed out the associations between certain transmembrane (TMEM) family proteins and tumor progression, but the role of TMEM205 remains unclear. In this study, we analyzed the RNA-seq and clinical data of 371 patients from The Cancer Genome Atlas (TCGA) and found significant differential expression of TMEM205 between normal and tumor tissues (P < 0.001). Low TMEM205 expression was also found to be independently associated with poor overall survival (OS; p = 0.032) and poor disease-specific survival (DSS; p = 0.002) in multivariate Cox regression analyses. RNA-seq and clinical data from hepatocellular carcinoma patients in the International Cancer Genome Consortium (ICGC) also showed significant differential expression of TMEM205 (P < 0.001) and association between low TMEM205 expression and poor survival (P < 0.001). We also used the Estimate the Proportion of Immune and Cancer cells (EPIC) tool to estimate the proportions of various immune cells in the tumor tissues. A correlation analysis was conducted, and TMEM205 expression in tumor tissues was found to be significantly associated with the proportion of macrophages (Pearson r = 0.45, p < 0.0001). A negative correlation was found between TMEM205 expression and M2 macrophage markers (CD163, EGR2, and MS4A4A) and between TMEM205 expression and regulatory T cell (Treg) markers (CCR8, STAT5B, and IL2RA), while a positive correlation was found between TMEM205 expression and the proportion of CD8+ T cells (Pearson r = 0.26, p < 0.0001). In conclusion, TMEM205 might improve HCC patients’ prognosis by reducing the levels of immunosuppressive cells (M2 macrophages and Tregs) and facilitating the infiltration of cytotoxic T cells into the tumor microenvironment. Therefore, TMEM205 has potential as a prognostic biomarker and immunotherapy agent in combination therapy regimens for HCC.
Introduction
Hepatocellular carcinoma (HCC) is the second leading cause of cancer-related deaths worldwide, causing nearly 800,000 deaths every year (Ferlay et al., 2015). Most cases develop from cirrhosis, which is mainly associated with chronic infection with hepatitis B virus and hepatitis C virus.
Nowadays, diverse strategies are used to treat HCC, including surgery, liver transplantation, ablation, transhepatic arterial chemotherapy and embolization, and molecularly targeted therapy. Unfortunately, the prognosis of advanced HCC patients remains poor (Bruix et al., 2017; Finn et al., 2018). In the past few decades, increasing knowledge of immune-related mechanisms has led to the development of immunotherapy for HCC. Programmed death-1 (PD-1) is expressed on CD8+ and CD4+ T cells and, upon binding to its ligand PD-L1, which is expressed on many cancer cells, it inhibits T cell activation; it has therefore been considered an ideal target for immunotherapy. Although nivolumab (anti-PD-1) has shown some therapeutic effect as a second line treatment in advanced HCC (El-Khoueiry et al., 2017), its variable clinical efficacy and limited applications (in terms of the suitable patients) make its future use in HCC unclear (Zhu et al., 2018). Therefore, a novel immune-related therapy target in HCC is urgently needed.
In the tumor microenvironment (TME), the fate of tumor cells is often decided by the interactions with immune cells and stromal cells. Tumor-associated macrophages (TAMs) surprisingly occupy up to 50% of the tumor tissue and dominate the immune cells that infiltrate the TME (Tu et al., 2014; Van Overmeire et al., 2014). The M2-like phagocytic TAMs are induced by interleukin (IL)-4, IL-6, and IL-10 (Mantovani et al., 2004; Wang et al., 2010) and Notch. They can recruit regulatory T cells (Treg) to cancer sites via the expression of CCL17, CCL18, and CCL22, thereby blocking the activation of cytotoxic T cells (Mamrot et al., 2019; Wang et al., 2019). A positive feedback loop may exist between M2-like macrophages and Tregs, which can lead to severe immunosuppressive effects, affecting cancer progression. Additionally, mounting evidence has indicated that the presence of functionally exhausted T cells in tumor tissues may be associated with resistance to immunotherapy (Anderson et al., 2017; Chen and Mellman, 2017). The limited clinical effect of immunotherapy (e.g., anti-PD-1) may be related to disturbances in the immunosuppressive cells (such as increased M2-like macrophage levels) in the TME, and there is an urgent need for a detailed understanding of the TAM-mediated inhibition of cytotoxic T cells (Noy and Pollard, 2014).
The transmembrane (TMEM) family of proteins are proteins that span biological membranes, most of which extend through the lipid bilayer of the plasma membrane, while others are located in the membranes of organelles. Although the functions of most TMEM proteins remain unknown, increasing attention has been paid to the associations between TMEM and tumor initiation and progression. TMEM176A expression inhibited the tumor cell growth of esophageal squamous cell carcinoma both in vitro and in vivo (Wang et al., 2017), and a very similar study revealed that the TMEM176 promoter was methylated in nearly 50% of primary colorectal cancers (Gao et al., 2017). Additionally, TMEM48 overexpression in lung cancer was associated with lung cancer cell proliferation, migration, and invasion (Qiao et al., 2016). Furthermore, TMEM48 suppression led to increased expression of apoptotic and tumor suppressor proteins such as caspase 3, PTEN, and p53 (Akkafa et al., 2018).
TMEM205 has been found to be highly expressed in the liver (Shen et al., 2010), but its role in HCC has not previously been explored. Additionally, TMEM205 was found to be in the top five differentially expressed genes at 1 week and 1 month after kidney transplantation, which suggests that it may have a role in the immune response (Bueno et al., 2015). Using RNA-seq data and clinical data from The Cancer Genome Atlas (TCGA) and International Cancer Genome Consortium (ICGC), this study explored whether TMEM205 is a prognostic marker in HCC patients and the associations between TMEM205 expression and the proportions of different types of immune cells in the TME. We hope to provide new understanding of the immune microenvironment and insights into a potential immunotherapy target for future treatment of HCC.
Materials and Methods
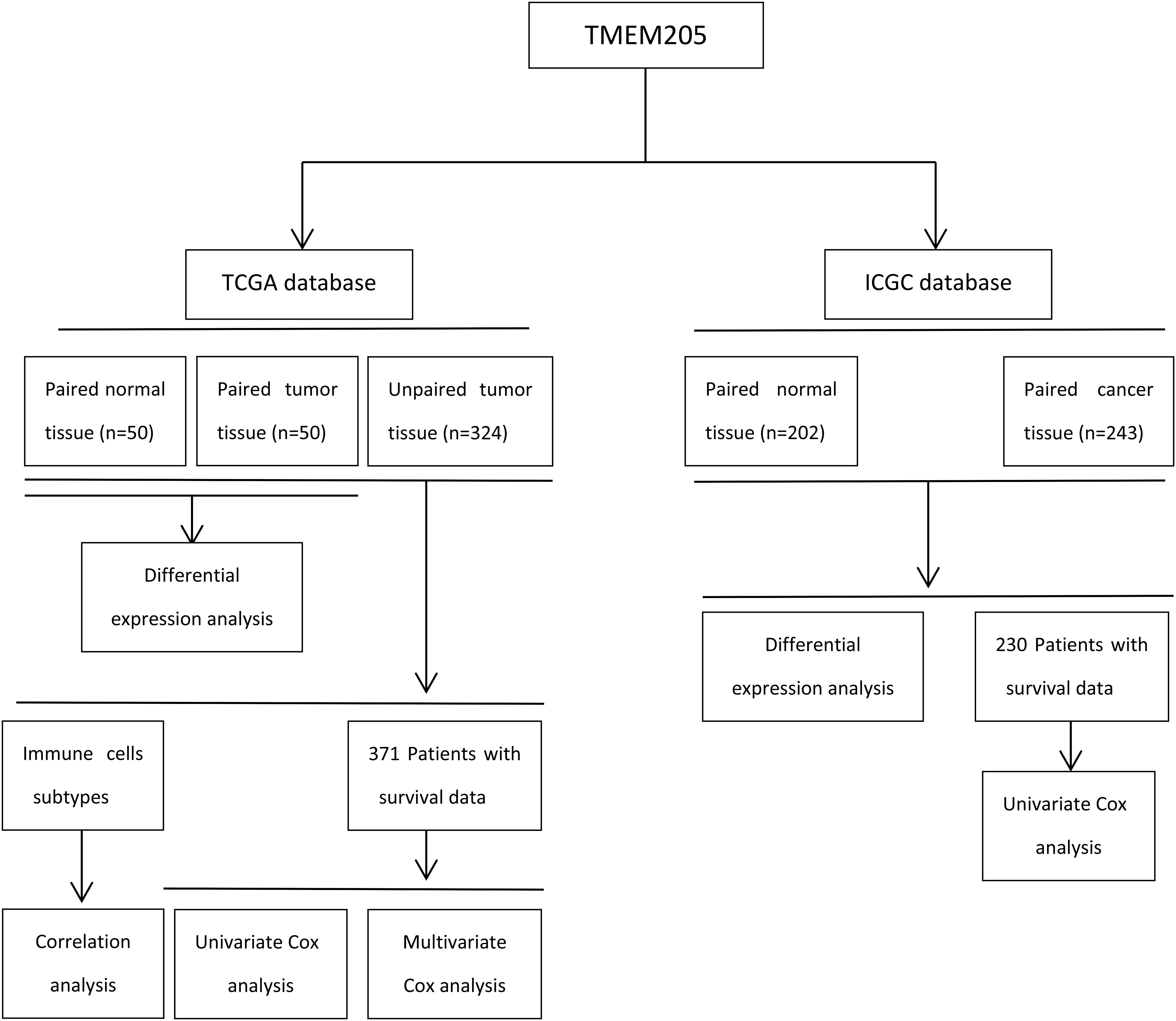
Publicly available RNA-seq data (samples = 424) and complete clinical data from HCC patients (n = 371) were downloaded from TCGA. Six patients were excluded in TCGA because their survival time was 0. We also collected available RNA-seq data (samples = 445) and clinical data from HCC patients (n = 230) in ICGC. The proportions of various types of immune and stromal cells in the tumor tissues were estimated using RNA-seq data and the Estimate the Proportion of Immune and Cancer cells (EPIC) tool, which integrates novel marker expression profiles from non-malignant cell types found in tumors and involves renormalization based on the cell-type-specific mRNA levels1. Details of the steps are summarized in Figure 1 through a flowchart.
The statistical analysis was performed in R 3.6.3 (R Foundation for Statistical Computing, Vienna, Austria). The differential expression of TMEM205 between normal and tumor tissues were visualized through ggboxplot function of “ggpubr” package. Kaplan–Meier analyses and Cox regression analyses (both univariate and multivariate) were performed using the “survival” and “survminer” packages in R, with p < 0.05 being considered to represent statistical significance (though not in the subsequent Pearson correlation analyses). The optimum cutpoint regarding low TMEM205 expression was determined using the “surv_cutpoint” function in the “survminer” package in R. Pearson correlation analyses were conducted using the basic “cor.test” function, with only p < 0.0001 being considered to represent statistical significance.
Most of the images in this paper were generated using R. We used the “ggsurvplot” and “ggforest” functions in the “survminer” package to generate Kaplan–Meier survival curves and forest plots, respectively. The “ggscater” function in the “ggpubr” package and the “corrplot” function in the “corrplot” package were used to visualize the correlation analyses. The estimated proportions of various types of immune and stromal cells in the tumor tissues were plotted in pie chart using Microsoft Excel 2010.
Results
Differentially Expressed TMEM205 Is an Independent Prognostic Factor in HCC Patients
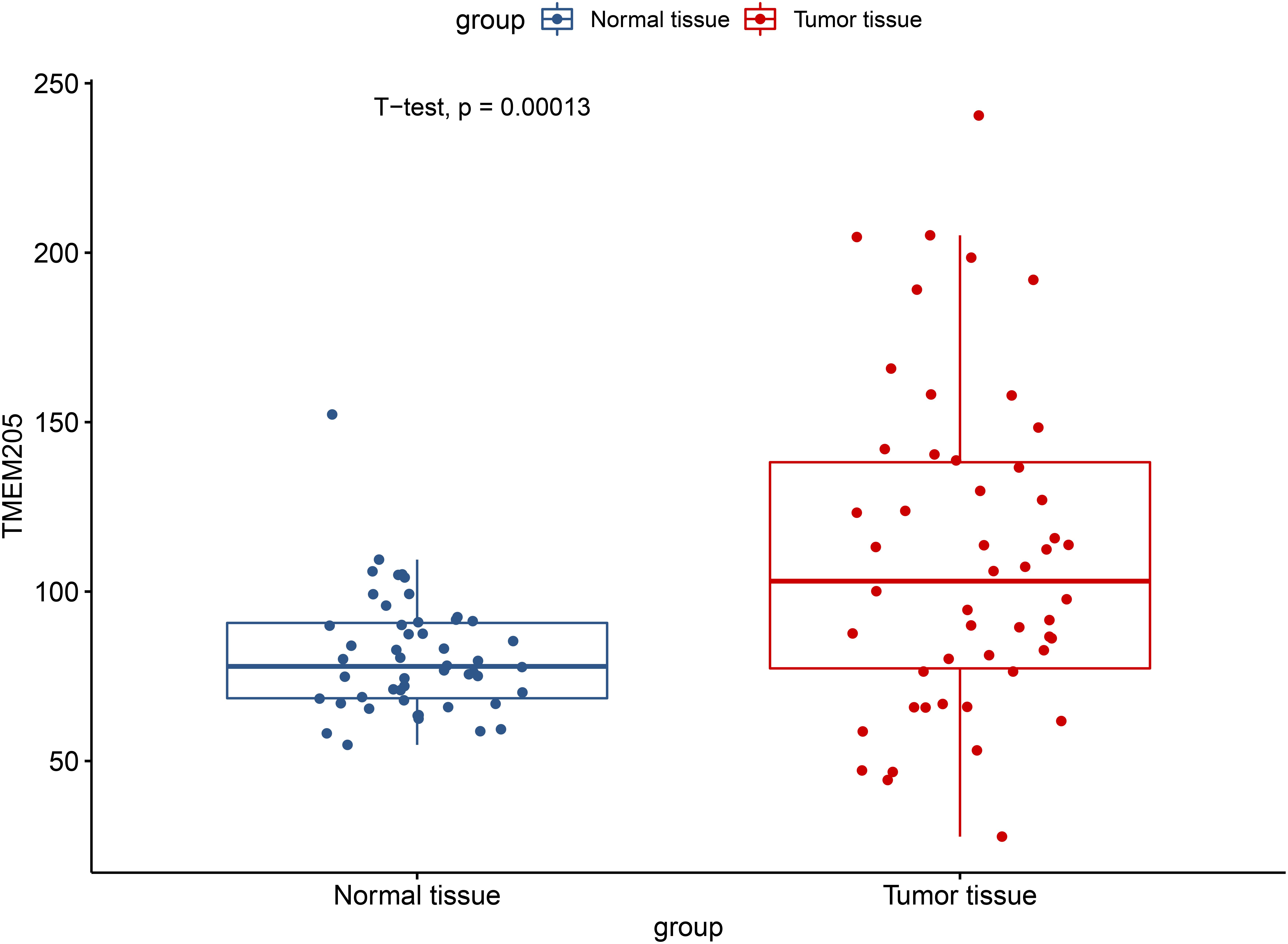
Through t-test with the TCGA RNA-seq datasets, we found differential expression of TMEM205 in 50 paired tumor and normal tissues (p < 0.001; Figure 2). Using Kaplan–Meier analysis and univariate Cox regression analysis with the TCGA RNA-seq datasets, low TMEM205 expression in tumor tissues was found to predict poor disease-specific survival (DSS; p = 0.005) in HCC patients (Figure 3). Low TMEM205 expression was also associated with poor overall survival (OS; p = 0.015; Figure 4) and poor progression-free interval (PFI; p = 0.040) in HCC patients. To control for potential confounding clinical factors (age, gender, pathologic stage, pathologic T stage, and pathologic grade), we performed multivariate Cox regression analysis and confirmed that low TMEM205 expression independently predicted poor DSS (p = 0.002; Figure 5) and poor OS (p = 0.032; Figure 6), but not poor PFI (p = 0.082).
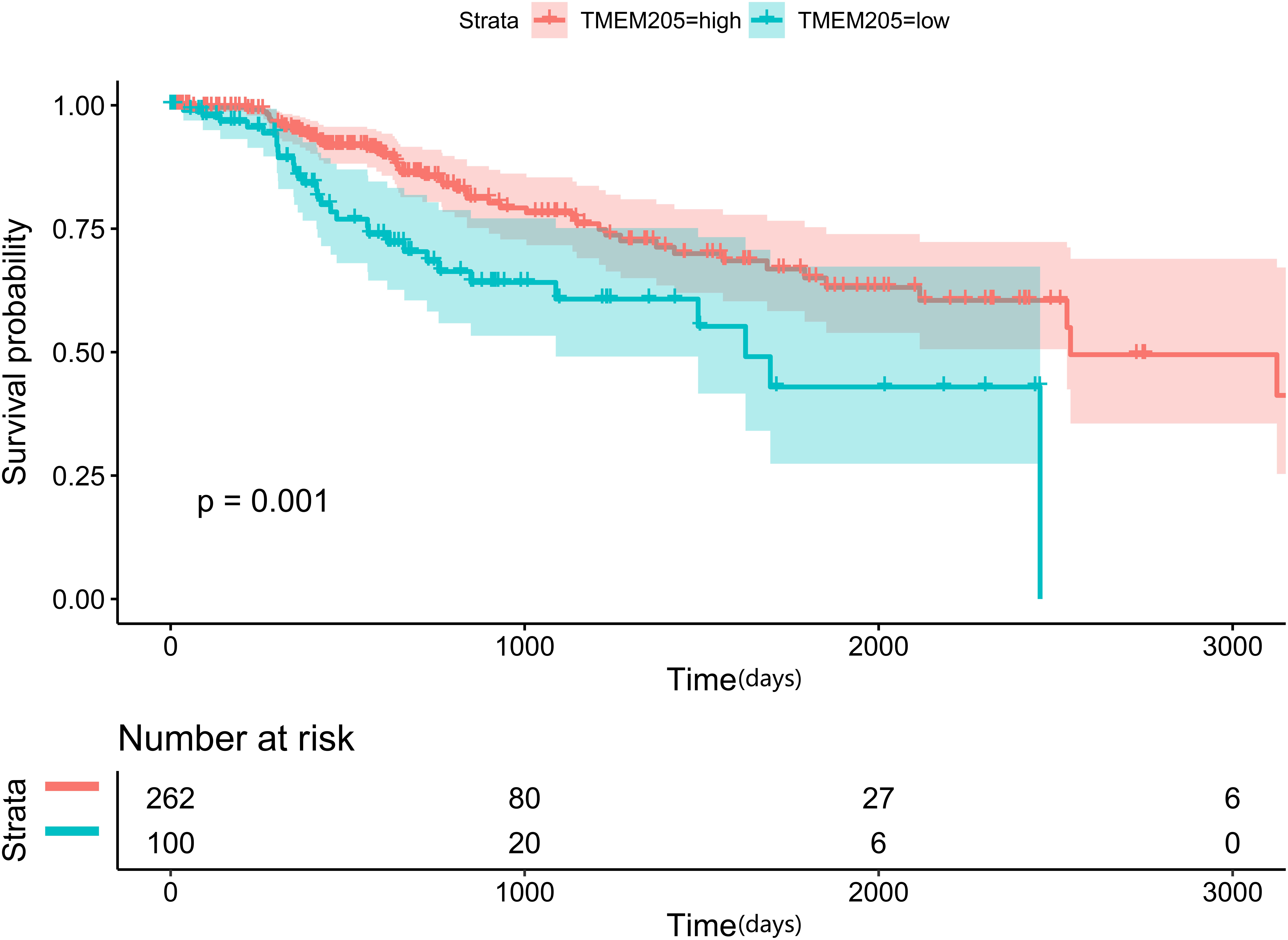
Figure 3. Low expression of TMEM205 is predictive for poor disease-specific survival (DSS) in hepatocellular carcinoma (HCC) patients from TCGA database.
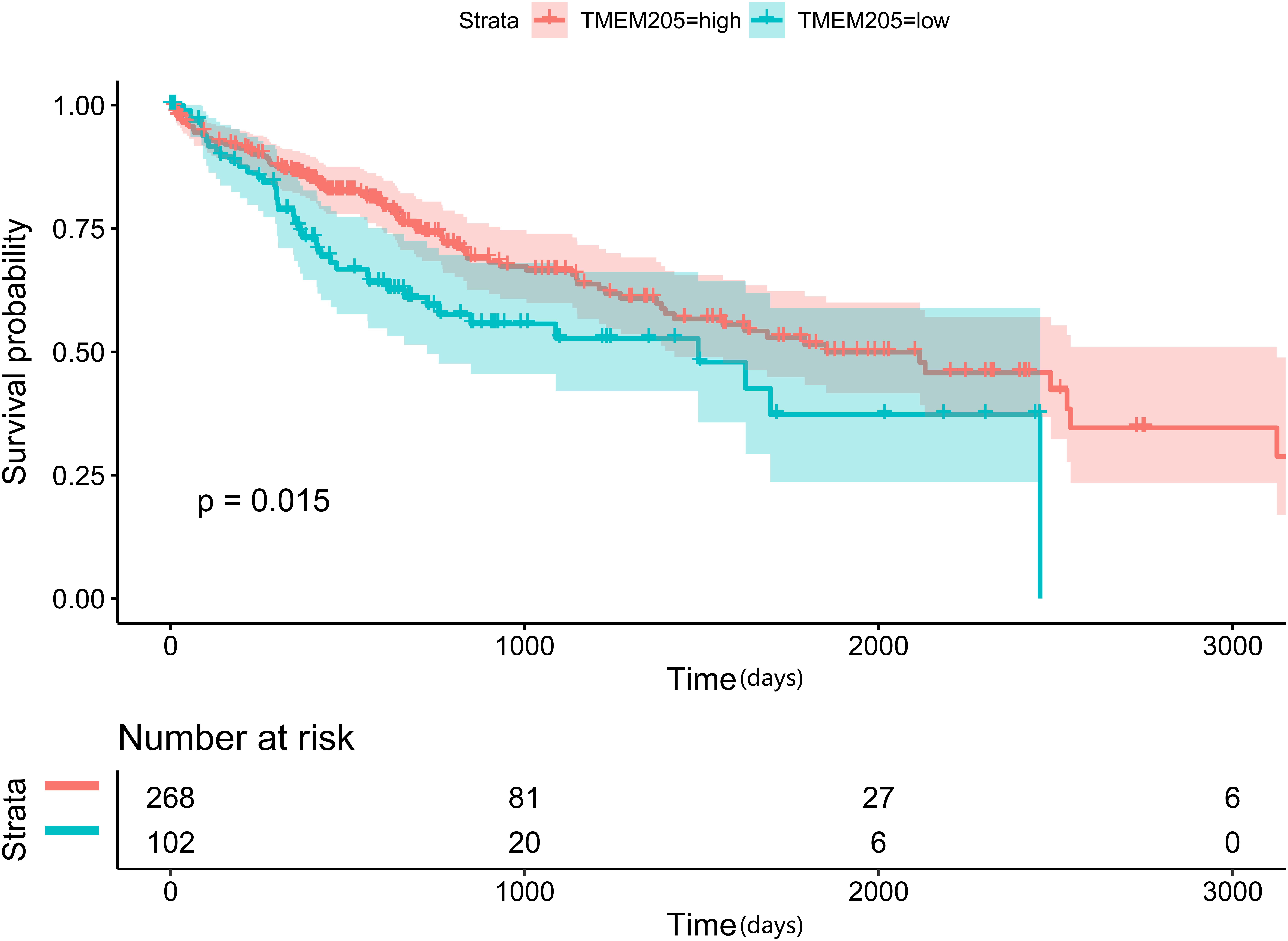
Figure 4. Low expression of TMEM205 is predictive for overall survival (OS) in hepatocellular carcinoma (HCC) patients from TCGA database.
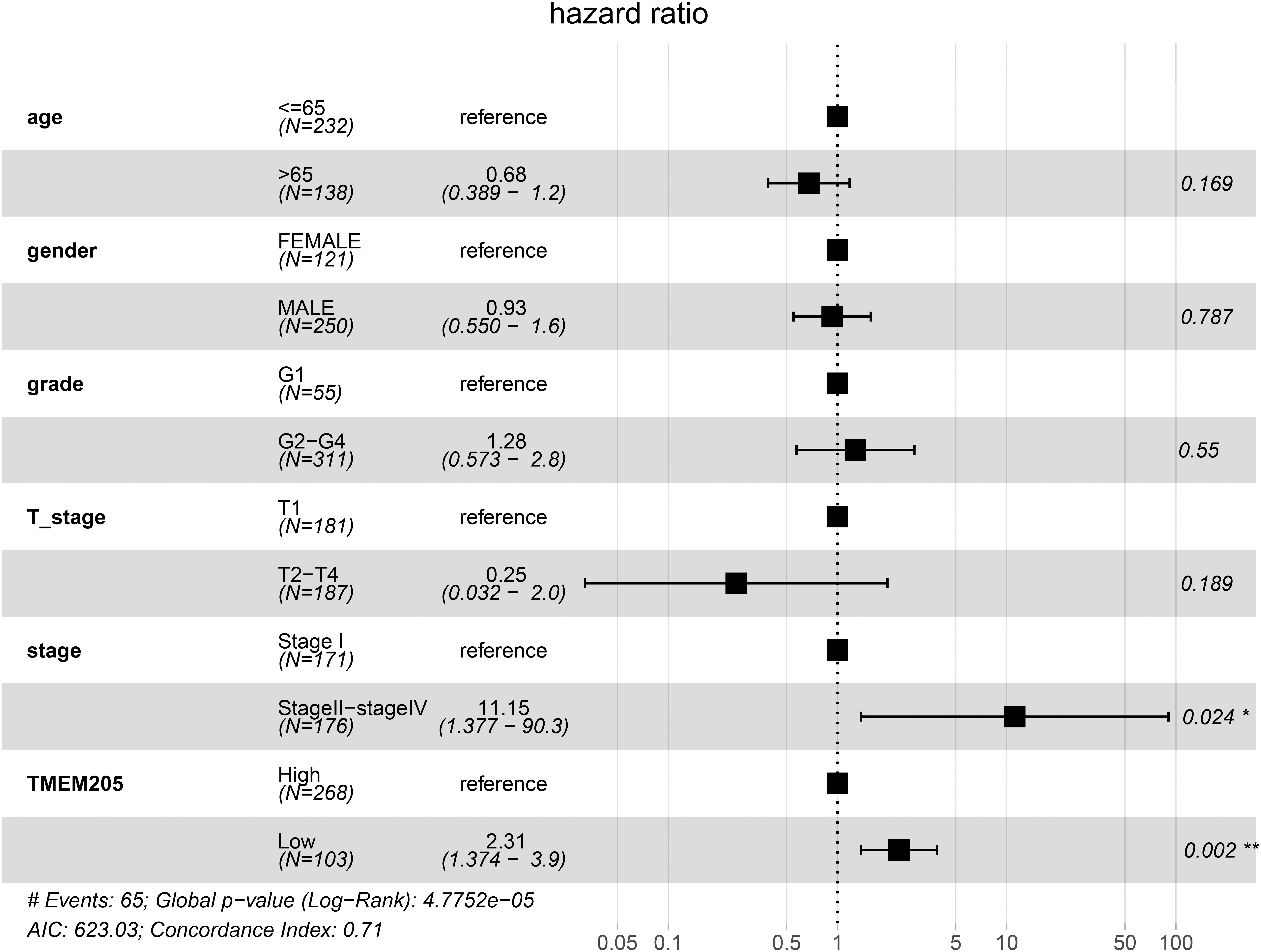
Figure 5. Low expression of TMEM205 is an independent prognostic factor for disease-specific survival (DSS) in HCC patients from TCGA database.
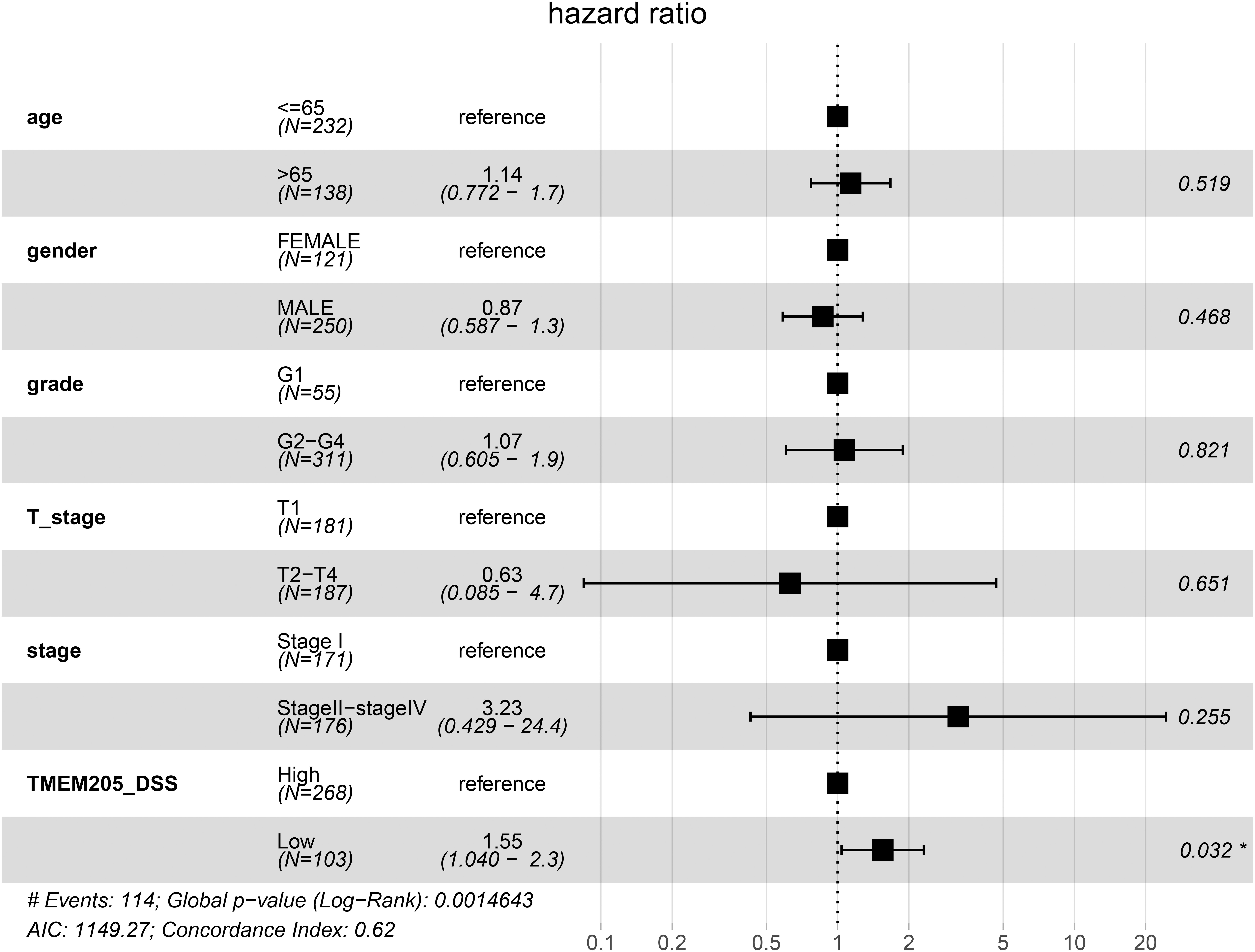
Figure 6. Low expression of TMEM205 is an independent prognostic factor for overall survival (OS) in HCC patients from TCGA database.
TMEM205 Expression Significantly Influences the Distribution of Immune Cell Subsets in the TME
Cancer cells interact with various types of immune cells and stromal cells in the TME throughout all stages, from early carcinogenesis to tumor progression and metastasis (Hanahan and Weinberg, 2011). We used the efficient EPIC tool to estimate the proportions of cancer and immune cell types based on tumor mRNA expression data (Racle et al., 2017; Figure 7). Macrophages made up approximately 24.99%, while CD4+ and CD8+ T cells comprised 5.81 and 0.34%, respectively. Cancer-related fibroblasts accounted for 2.38%, endothelial cells accounted for 2.17%, and natural killer cells accounted for 0.03%. A total of 63.78% of the tumor tissues comprised other cells, which mostly consisted of cancer cells and other immune cells. Using the Kaplan–Meier survival curve and univariate Cox regression analysis, we found that a low proportion of macrophages was associated with poor DSS (p < 0.001) in HCC patients (Figure 8).
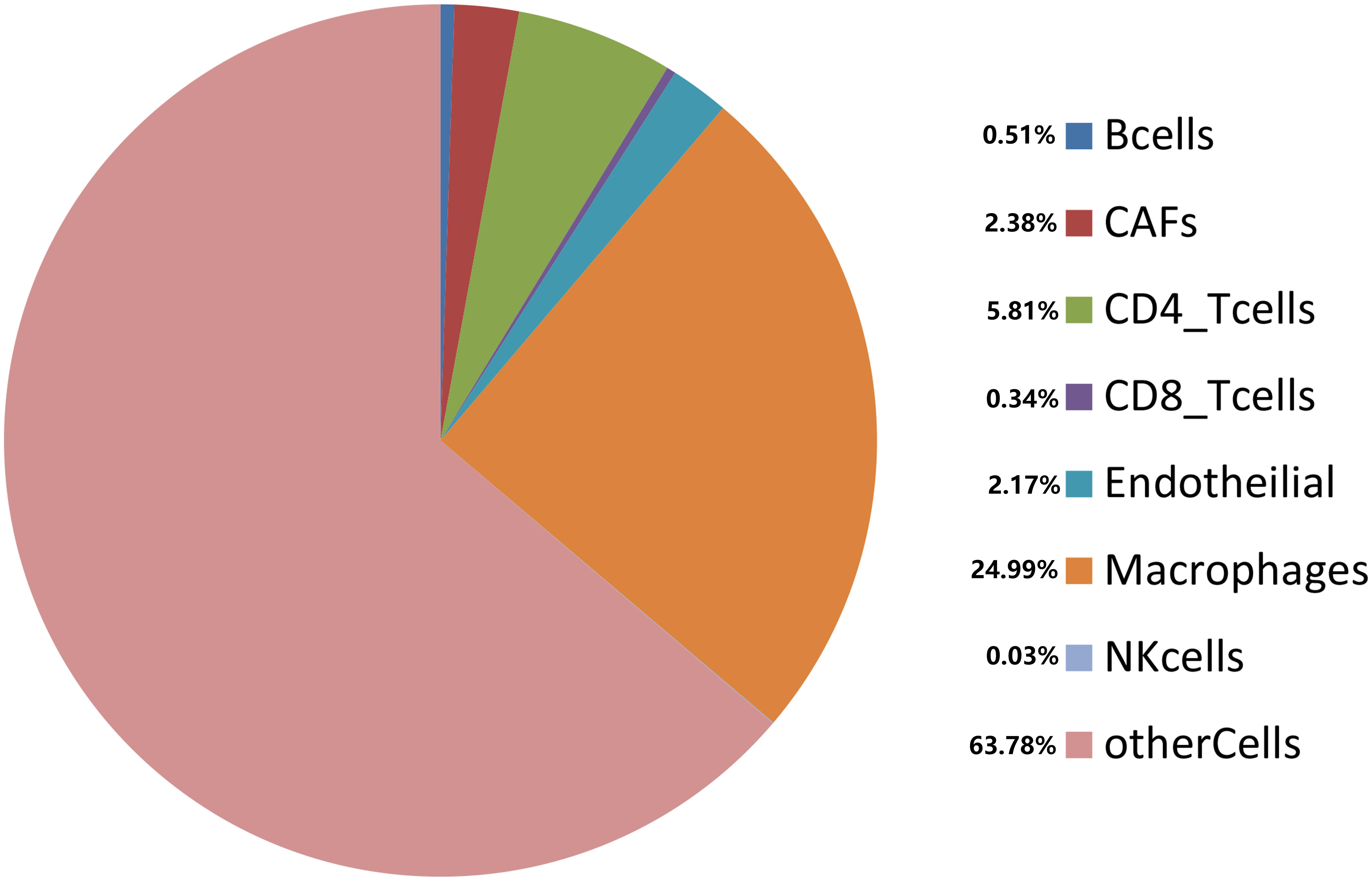
Figure 7. Estimated proportions of different types of immune cells in tumor tissues from TCGA database.
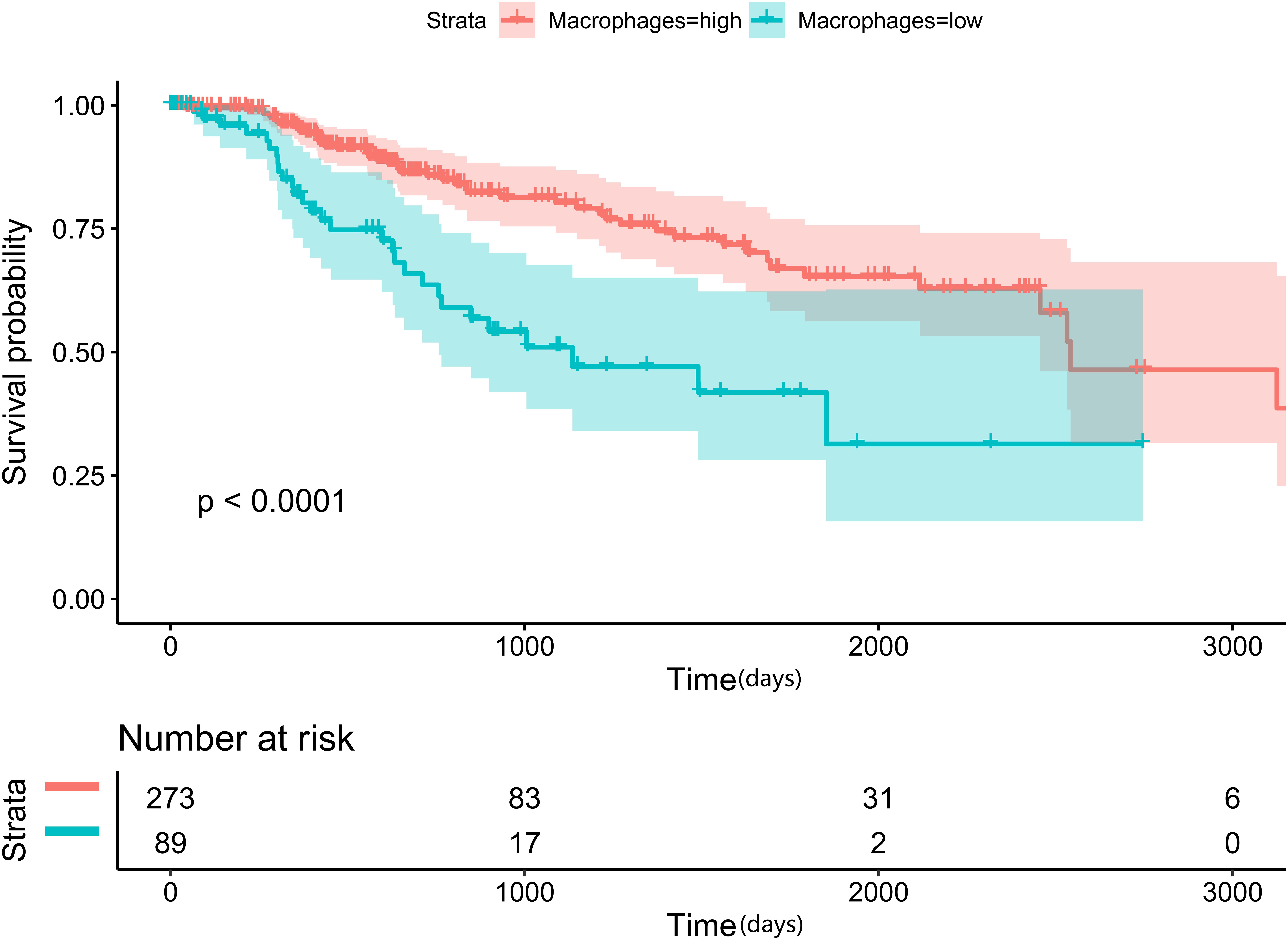
Figure 8. Low proportion of macrophages in tumor tissues is an independent prognostic factor for disease-specific survival (DSS) in HCC patients from TCGA database.
Additionally, TMEM205 expression in tumor tissues was significantly associated with the proportion of macrophages (Pearson r = 0.45, p < 0.0001; Figure 9). Combined with the above results, this indicates that the prognostic role of TMEM205 expression might be explained by alterations in the distribution of immune cell subsets in the TME.
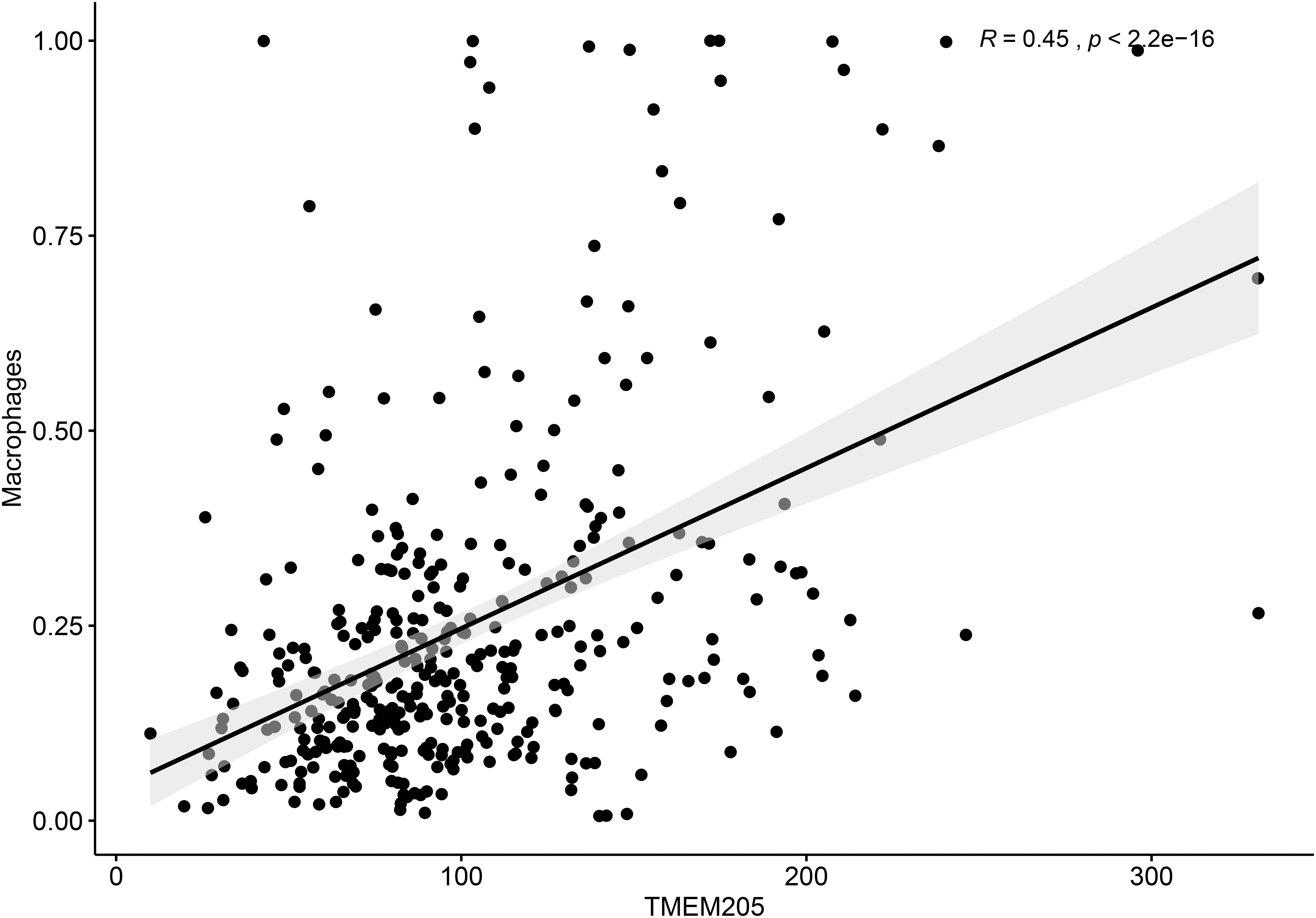
Figure 9. Expression of TMEM205 shows significant correlation with proportion of macrophages in tumor tissues from TCGA database.
TMEM205 Expression Is Associated With Decreased Numbers of Immunosuppressive Cells (M2 Macrophages and Tregs) and Infiltration of CD8+ T Cells Into the TME
A recent study showed that the transformation of monocytes into specific CD163+ M2-like macrophages was induced by colony-stimulating factor 1 (CSF1) (Papin et al., 2019). Inhibition of CSF1 receptor (CSF1R) on TAMs did not kill the TAMs but caused them to repolarize to a tumoricidal state (Quail and Joyce, 2013). We found that TMEM205 expression was negatively correlated with CSF1 expression (Pearson r = −0.27, p < 0.0001) and CSF1R expression (Pearson r = −0.23, p < 0.0001). TMEM205 expression was also negatively correlated with the expression of M2 macrophage markers (CD163, EGR2, and MS4A4A) (Jablonski et al., 2015; Sanyal et al., 2017; Terai et al., 2018; Figure 10). Additionally, TMEM205 expression was negatively correlated not only with the expression of the chemokine receptors CCR4 (Pearson r = −0.20, p < 0.0001) and CCR5 (Pearson r = −0.25, p < 0.0001) (Kurose et al., 2015; Jiao et al., 2019), which are responsible for Treg migration to the TME, but also with the Treg markers (CCR8, STAT5B, and IL2RA) (Burchill et al., 2008; Nishikawa and Sakaguchi, 2014; Villarreal et al., 2018; Figure 10). Finally, we found that TMEM205 expression appeared to inhibit IL-10 expression (Pearson r = −0.22, p < 0.0001), while TMEM205 expression was positively correlated with the proportion of CD8+ T cells in tumor tissues (Pearson r = 0.26, p < 0.0001). Together, these data suggest that TMEM205 expression might inhibit M2 macrophage polarization, inhibit Treg recruitment, and facilitate CD8+ T cell infiltration into tumor tissues, thereby improving the patients’ prognosis, which is summarized in Figure 11.
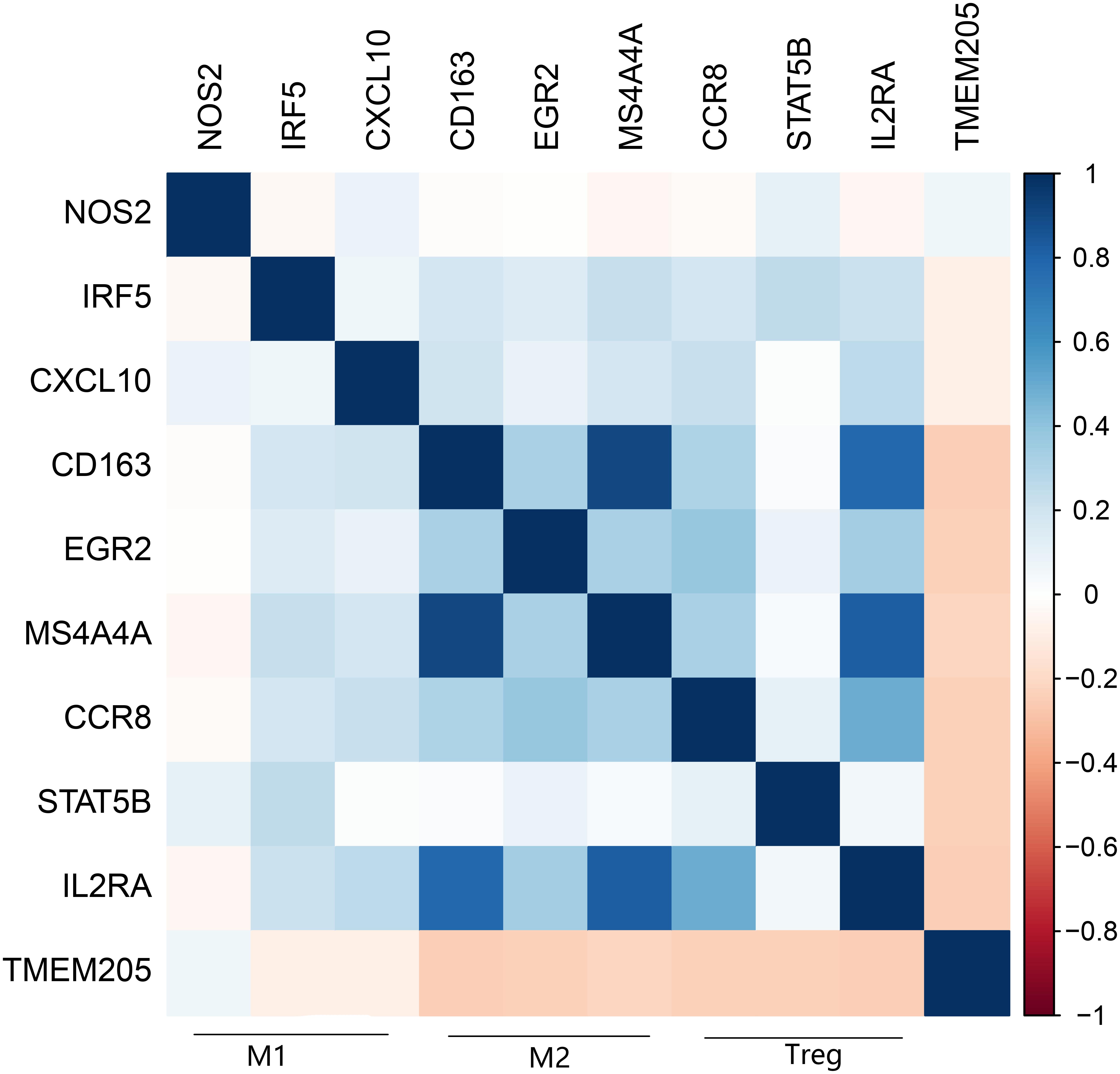
Figure 10. Expression of TMEM205 is significantly negatively correlated with markers of M2 macrophages (CD163, EGR2, and MS4A4A) and Treg (CCR8, STATA5B, and IL2RA) but is not correlated with markers of M1 macrophages (NOS2, IRF5, and CXCL10) from TCGA database. The color in the square reflects the correlation coefficient between square’s abscissa and ordinate.
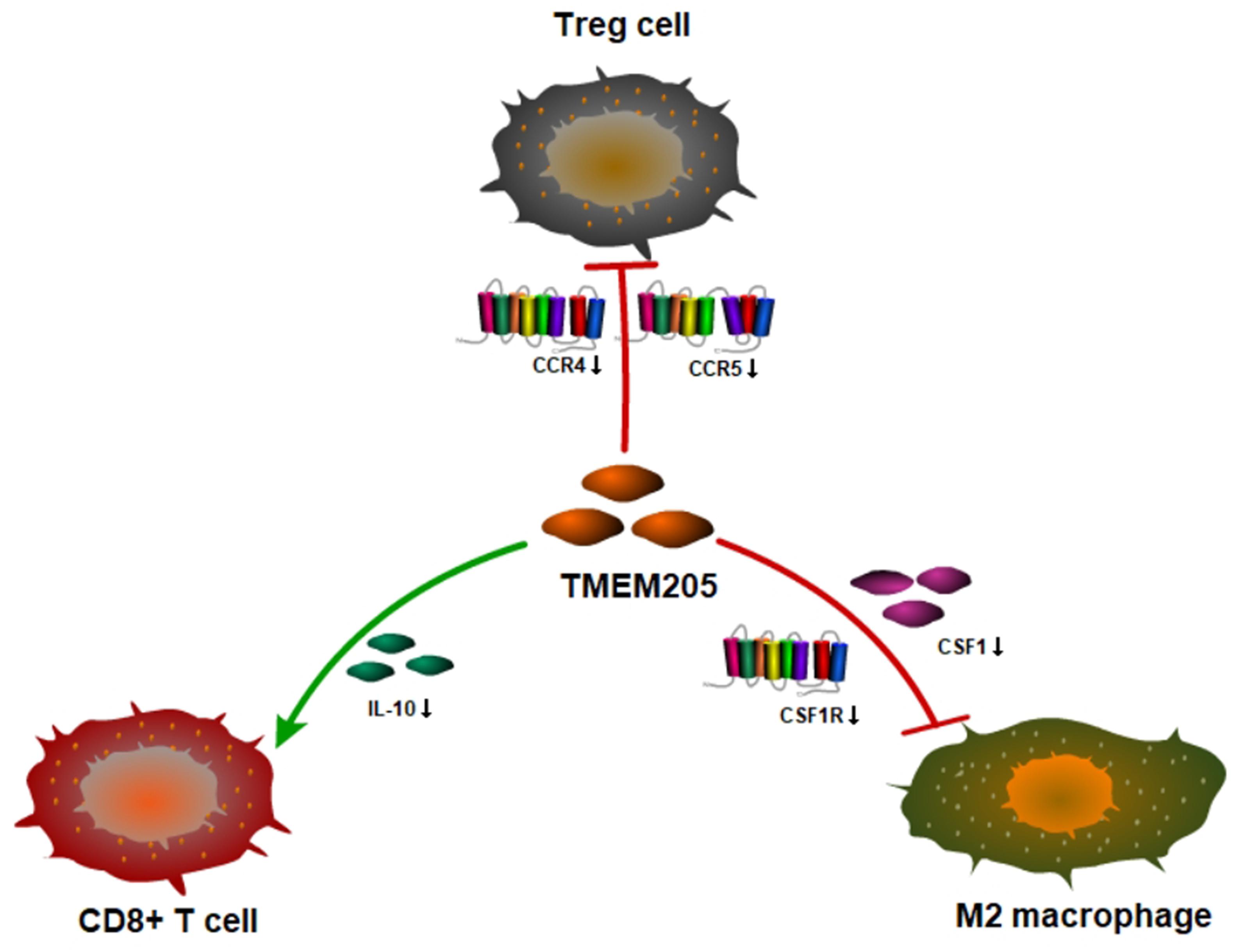
Figure 11. Hypothesis figure for the correlation between TMEM205 expression and immune cell distribution.
Validation of TMEM205 in the ICGC Cohort
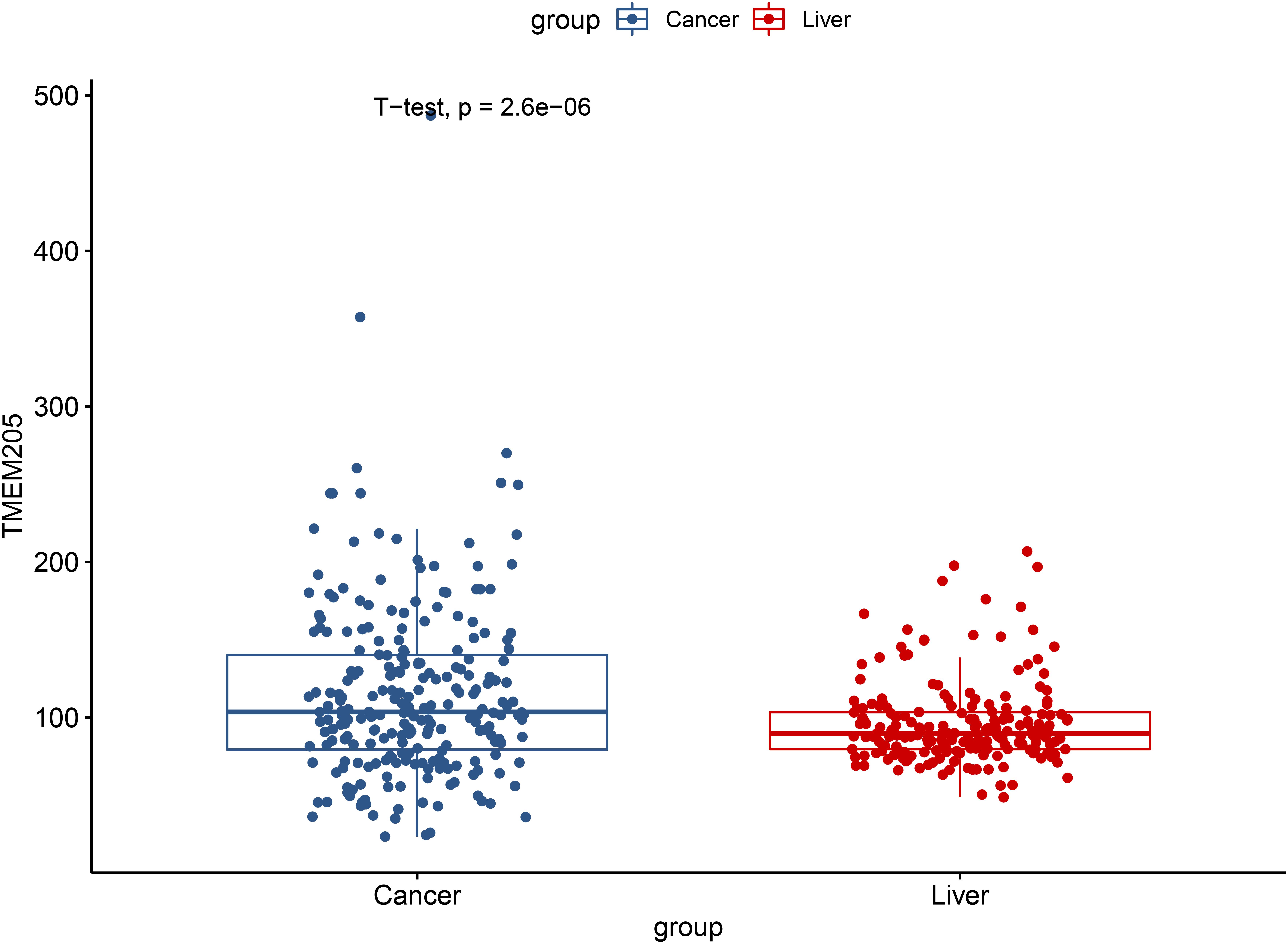
We validated the significant difference of TMEM205 expression between normal tissues and tumor tissues in the ICGC cohort (p < 0.001; Figure 12). Low expression of TMEM205 was also found to be associated with poor survival (p < 0.001; Figure 13).
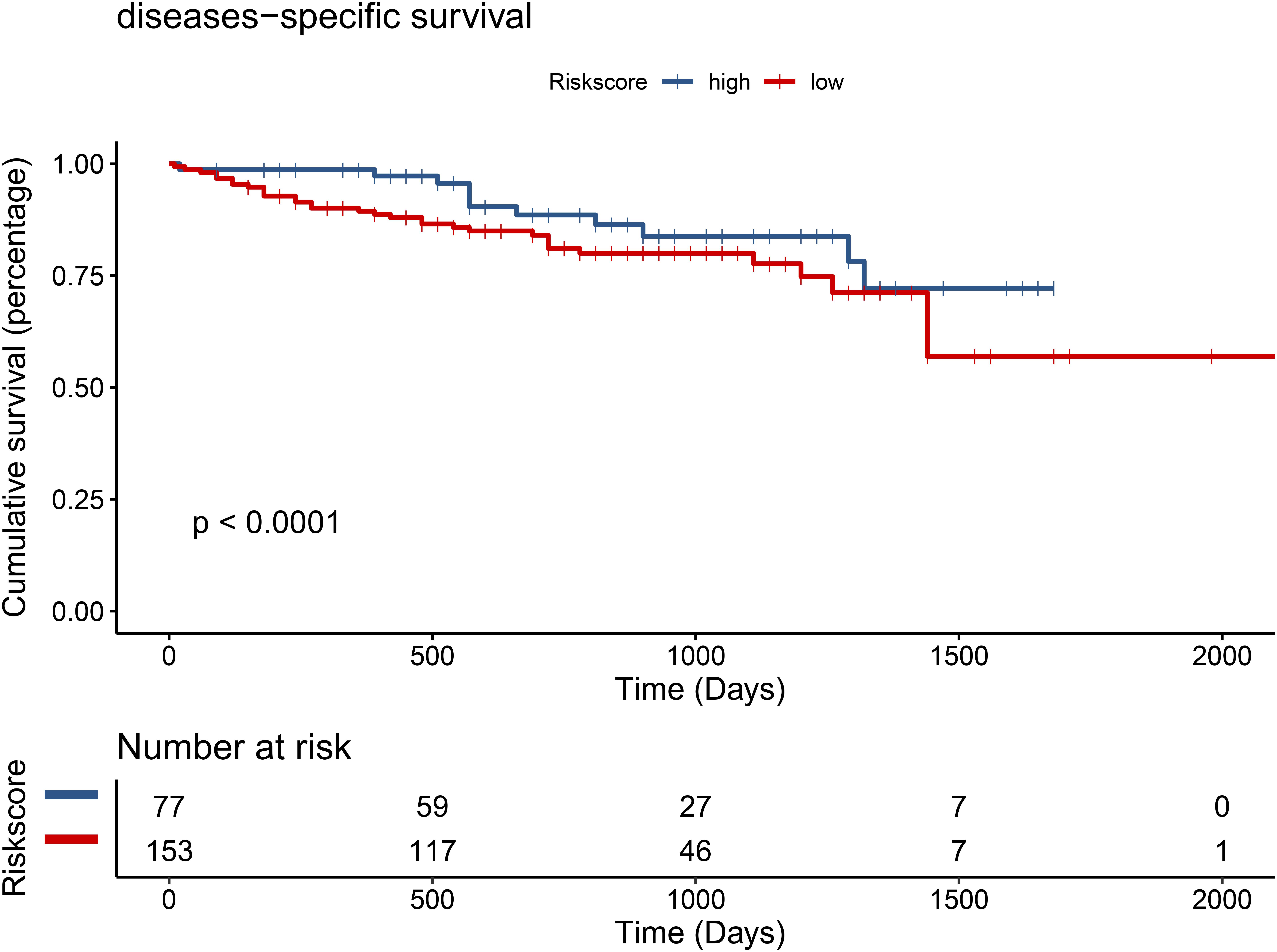
Figure 13. Low expression of TMEM205 is predictive for survival in hepatocellular carcinoma (HCC) patients from ICGC database.
Discussion
Previous studies have reported associations between many members of the TMEM family and tumor progression (Qiao et al., 2016; Gao et al., 2017; Wang et al., 2017; Akkafa et al., 2018; Song et al., 2018), but the role of TMEM205 was not previously established. In the current study, we not only found significant difference in TMEM205 expression between normal tissue and tumor tissue (p < 0.001) but also demonstrated that low TMEM205 expression was independently associated with poor OS (p = 0.032) and DSS (p = 0.002) in HCC patients from the TCGA cohort. A significant differential expression of TMEM205 was also observed (p < 0.001) and low TMEM205 expression still predicted poor survival (p < 0.001) in HCC patients from the ICGC cohort, which validated that TMEM205 expression was an independent prognostic factor in HCC.
Next, we found that having a low proportion of macrophages in the TME predicted poor DSS (p < 0.001) in HCC patients. This result is similar to the finding by Li et al. that a high proportion of intratumoral CD68+ TAMs in HCC patients predicted improved survival (Li et al., 2009). We also found a positive correlation between TMEM205 expression and the proportion of macrophages in tumor tissues (Pearson r = 0.45, p < 0.0001). However, the role of TAMs in tumor evolution seems to be complex. Classical M1 macrophages and alternatively activated M2 macrophages are two major functional phenotypes of TAMs, and they promote tumoricidal responses and increase tumor progression, respectively (Gordon and Taylor, 2005; Wang et al., 2010). Confirming this, inhibiting M2 macrophage polarization was recently shown to lead to impaired tumor growth and impaired immune evasion (Ye et al., 2018). In the current study, we could not determine the proportions of more precise cell subtypes such as M2 macrophages, due to the restricted functions of the EPIC tool, but we identified negative correlations with M2 macrophage markers (CD163, EGR2, and MS4A4A). CD163 is a type of hemoglobin scavenger receptor that indirectly contributes to the anti-inflammatory response (Etzerodt and Moestrup, 2013), and it is well known as a specific M2 marker, based on in vitro and in vivo evidence (Ino et al., 2013; Erlandsson et al., 2019). EGR2 expression on macrophages increases after stimulation with M2-like stimuli (IL-4 or IL-13) (Veremeyko et al., 2018). MS4A4A may serve as a novel cell-surface marker for M2 macrophages (Sanyal et al., 2017). We also observed that TMEM205 expression was negatively correlated with CSF1 expression (Pearson r = −0.27, p < 0.0001) and CSF1R expression (Pearson r = −0.23, p < 0.0001). A previous study showed that the downstream signaling pathway of CSF1R was responsible for the polarization of TAMs to an immunosuppressive M2 phenotype (Ramesh et al., 2019). Thus, there might also be a negative correlation between TMEM205 expression and the proportion of M2 macrophages in tumor tissues.
Additionally, we found a negative correlation between TMEM205 expression and Treg markers (CCR8, STAT5B, and IL2RA). CCR8 is a chemokine receptor mainly expressed on Tregs and it plays a critical role in Treg-mediated immunosuppression (Villarreal et al., 2018). STAT5B deficiency is associated with decreased FOXP3 expression and Treg levels (Kanai et al., 2012). IL2RA (also known as CD25) is also a marker that is used to quantify Treg levels in individuals (Yu et al., 2012). Furthermore, we found a negative correlation between TMEM205 expression and the chemokine receptors CCR4 (Pearson r = −0.20, p < 0.0001) and CCR5 (Pearson r = −0.25, p < 0.0001), which are responsible for Treg migration to the TME (Kurose et al., 2015; Jiao et al., 2019). These results suggested a negative correlation between TMEM205 expression and the proportion of Tregs in tumor tissues. It is well known that Tregs inhibit antitumor immunity and are involved in tumor development (Ohue and Nishikawa, 2019).
We also found a negative correlation between IL-10 expression and TMEM205 expression (Pearson r = −0.22, p < 0.0001). High IL-10 levels predict poor breast cancer prognosis and the IL-10 secreted by TAMs can inhibit T cell effector functions and induce regulatory functions that recruit Tregs (Noy and Pollard, 2014; Ahmad et al., 2018). Therefore, it was not difficult to find a positive correlation between TMEM205 expression and the proportion of CD8+ T cells (Pearson r = 0.26, p < 0.0001).
In the last few decades, great progress has been made in the understanding of HCC biology and immunotherapy, but the therapeutic effects of anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA4) and anti-PD-1 immunotherapy remain unsatisfactory (Sangro et al., 2013; El-Khoueiry et al., 2017; Zhu et al., 2018). Additionally, a significant percentage of patients eventually develop resistance to immunotherapy and many patients are only partial responders (Topalian et al., 2012; Hamid et al., 2013; Zou et al., 2016). Among all the possible reasons, the disturbance of local immunosuppressive cells in the TME might be one of the most important ones. Kato et al. showed that a decreased M2 population and an increased M1/M2 ratio tended to be associated with a better therapeutic effect regarding anti-PD-1 therapy (Kato et al., 2019). High Tregs levels also impaired the response to anti-PD-1/PD-L1 therapy (causing a poor immunologic response against the tumor) (Bettelli et al., 2006), and strategies involving targeting Tregs lead to improved responsiveness to anti-PD-1/PD-L1 therapy (Arce Vargas et al., 2017; Taylor et al., 2017). Nowadays, combination therapy is the major tumor therapy approach, and a recent study found that an anti PD-1 drug (atezolizumab) combined with an anti-vascular endothelial growth factor drug (bevacizumab) led to encouraging antitumor activity and safety in patients with unresectable HCC (Finn et al., 2020). In this study, we found that TMEM205 expression was not only associated with HCC prognosis but also with decreased levels of immunosuppressive cells (M2 macrophages and Tregs) and infiltration of CD8+ T cells into the TME. This suggests that TMEM205 may be useful as part of combination therapy regimens.
Several limitations in our study should be recognized. First, due to the restricted functions of the EPIC tool, we could not determine the proportions of more specific subtypes of immune cells (such as M2 macrophages or Tregs). Instead, we could only assess associations with corresponding cell markers, which might have produced bias. Also, the clinical data on the immunotherapy administered to HCC patients were unclear, which prevented a more detailed analysis from being conducted.
In summary, we first confirmed that TMEM205 expression is an independent prognostic factor in HCC patients, and we then found that the antitumor effect might be explained by reducing the numbers of immunosuppressive cells (M2 macrophages and Tregs) and facilitation of CD8+ T cell infiltration into the TME. These results suggest that TMEM205 may be useful as a therapeutic agent.
Data Availability Statement
The RNA sequencing data for this study can be found in the TCGA data portal (https://portal.gdc.cancer.gov/). The clinical data can be found in datasets of UCSC Xena (https://xenabrowser.net/datapages/).
Author Contributions
YM conceptualized the study. JR, XW, and XZ were responsible for investigation and statistical analysis. JR and XW wrote the original manuscript. JR and RD reviewed and edited the manuscript. YM was responsible for funding acquisition. All authors contributed to the article and approved the submitted version.
Funding
This study was funded by the National Natural Science Foundation of China (81873591 and 81670591), Guangdong Natural Science Foundation, China (2016A030311028), and The Science and Technology Planning Project of Guangdong Province, China (2018A050506030).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
References
Ahmad, N., Ammar, A., Storr, S. J., Green, A. R., Rakha, E., Ellis, I. O., et al. (2018). IL-6 and IL-10 are associated with good prognosis in early stage invasive breast cancer patients. Cancer Immunol. Immunother. 67, 537–549. doi: 10.1007/s00262-017-2106-8
Akkafa, F., Koyuncu, I., Temiz, E., Dagli, H., Dilmec, F., and Akbas, H. (2018). miRNA-mediated apoptosis activation through TMEM 48 inhibition in A549 cell line. Biochem. Biophys. Res. Commun. 503, 323–329.
Anderson, K. G., Stromnes, I. M., and Greenberg, P. D. (2017). Obstacles posed by the tumor microenvironment to t cell activity: a case for synergistic therapies. Cancer Cell 31, 311–325. doi: 10.1016/j.ccell.2017.02.008
Arce Vargas, F., Furness, A. J. S., Solomon, I., Joshi, K., Mekkaoui, L., Lesko, M. H., et al. (2017). Fc-Optimized Anti-CD25 depletes tumor-infiltrating regulatory T cells and synergizes with PD-1 blockade to eradicate established tumors. Immunity 46, 577–586. doi: 10.1016/j.immuni.2017.03.013
Bettelli, E., Carrier, Y., Gao, W., Korn, T., Strom, T. B., Oukka, M., et al. (2006). Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238. doi: 10.1038/nature04753
Bruix, J., Merie, P., Granito, A., Huang, Y.-H., Bodoky, G., Pracht, M., et al. (2017). Survival by pattern of tumor progression during prior sorafenib (SOR) treatment in patients with hepatocellular carcinoma (HCC) in the phase III RESORCE trial comparing second-line treatment with regorafenib (REG) or placebo. J. Clin. Oncol. 35:229.
Bueno, V., Dorr, C., Wu, B., Guan, W., Muthusamy, A., Sanghavi, K., et al. (2015). Differentially expressed gene transcripts using RNA sequencing from the blood of immunosuppressed kidney allograft recipients. PLoS One 10:e0125045. doi: 10.1371/journal.pone.0125045
Burchill, M. A., Yang, J., Vang, K. B., Moon, J. J., Chu, H. H., Lio, C. W., et al. (2008). Linked T cell receptor and cytokine signaling govern the development of the regulatory T cell repertoire. Immunity 28, 112–121. doi: 10.1016/j.immuni.2007.11.022
Chen, D. S., and Mellman, I. (2017). Elements of cancer immunity and the cancer-immune set point. Nature 541, 321–330. doi: 10.1038/nature21349
El-Khoueiry, A. B., Sangro, B., Yau, T., Crocenzi, T. S., Kudo, M., Hsu, C., et al. (2017). Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet 389, 2492–2502. doi: 10.1016/S0140-6736(17)31046-2
Erlandsson, A., Carlsson, J., Lundholm, M., Falt, A., Andersson, S. O., Andren, O., et al. (2019). M2 macrophages and regulatory T cells in lethal prostate cancer. Prostate 79, 363–369. doi: 10.1002/pros.23742
Etzerodt, A., and Moestrup, S. K. (2013). CD163 and inflammation: biological, diagnostic, and therapeutic aspects. Antioxid. Redox. Signal. 18, 2352–2363. doi: 10.1089/ars.2012.4834
Ferlay, J., Soerjomataram, I., Dikshit, R., Eser, S., Mathers, C., Rebelo, M., et al. (2015). Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359–E386. doi: 10.1002/ijc.29210
Finn, R. S., Merle, P., Granito, A., Huang, Y. H., Bodoky, G., and Pracht, M. (2018). Outcomes of sequential treatment with sorafenib followed by regorafenib for HCC: additional analyses from the phase III RESORCE trial. J. Hepatol. 69, 353–358.
Finn, R. S., Qin, S., Ikeda, M., Galle, P. R., Ducreux, M., Kim, T. Y., et al. (2020). Atezolizumab plus bevacizumab in unresectable hepatocellular carcinoma. N. Engl. J. Med. 382, 1894–1905. doi: 10.1056/NEJMoa1915745
Gao, D., Han, Y., Yang, Y., Herman, J. G., Linghu, E., Zhan, Q., et al. (2017). Methylation of TMEM176A is an independent prognostic marker and is involved in human colorectal cancer development. Epigenetics 12, 575–583. doi: 10.1080/15592294.2017.1341027
Gordon, S., and Taylor, P. R. (2005). Monocyte and macrophage heterogeneity. Nat. Rev. Immunol. 5, 953–964. doi: 10.1038/nri1733
Hamid, O., Robert, C., Daud, A., Hodi, F. S., Hwu, W. J., Kefford, R., et al. (2013). Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N. Engl. J. Med. 369, 134–144. doi: 10.1056/NEJMoa1305133
Hanahan, D., and Weinberg, R. A. (2011). Hallmarks of cancer: the next generation. Cell 144, 646–674. doi: 10.1016/j.cell.2011.02.013
Ino, Y., Yamazaki-Itoh, R., Shimada, K., Iwasaki, M., Kosuge, T., Kanai, Y., et al. (2013). Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. Br. J. Cancer 108, 914–923. doi: 10.1038/bjc.2013.32
Jablonski, K. A., Amici, S. A., Webb, L. M., Ruiz-Rosado Jde, D., Popovich, P. G., Partida-Sanchez, S., et al. (2015). Novel markers to delineate murine M1 and M2 macrophages. PLoS One 10:e0145342. doi: 10.1371/journal.pone.0145342
Jiao, X., Nawab, O., Patel, T., Kossenkov, A. V., Halama, N., Jaeger, D., et al. (2019). Recent advances targeting CCR5 for cancer and its role in immuno-oncology. Cancer Res. 79, 4801–4807. doi: 10.1158/0008-5472.CAN-19-1167
Kanai, T., Jenks, J., and Nadeau, K. C. (2012). The STAT5b pathway defect and autoimmunity. Front. Immunol. 3:234. doi: 10.3389/fimmu.2012.00234
Kato, Y., Tabata, K., Kimura, T., Yachie-Kinoshita, A., Ozawa, Y., Yamada, K., et al. (2019). Lenvatinib plus anti-PD-1 antibody combination treatment activates CD8+ T cells through reduction of tumor-associated macrophage and activation of the interferon pathway. PLoS One 14:e0212513. doi: 10.1371/journal.pone.0212513
Kurose, K., Ohue, Y., Sato, E., Yamauchi, A., Eikawa, S., Isobe, M., et al. (2015). Increase in activated Treg in TIL in lung cancer and in vitro depletion of Treg by ADCC using an antihuman CCR28 mAb (KM2760). J. Thorac. Oncol. 10, 74–83.
Li, Y. W., Qiu, S. J., Fan, J., Gao, Q., Zhou, J., Xiao, Y. S., et al. (2009). Tumor-infiltrating macrophages can predict favorable prognosis in hepatocellular carcinoma after resection. J. Cancer Res. Clin. Oncol. 135, 439–449. doi: 10.1007/s00432-008-0469-0
Mamrot, J., Balachandran, S., Steele, E. J., and Lindley, R. A. (2019). Molecular model linking Th2 polarized M2 tumour-associated macrophages with deaminase-mediated cancer progression mutation signatures. Scand. J. Immunol. 89:e12760. doi: 10.1111/sji.12760
Mantovani, A., Sica, A., Sozzani, S., Allavena, P., Vecchi, A., and Locati, M. (2004). The chemokine system in diverse forms of macrophage activation and polarization. Trends Immunol. 25, 677–686. doi: 10.1016/j.it.2004.09.015
Nishikawa, H., and Sakaguchi, S. (2014). Regulatory T cells in cancer immunotherapy. Curr. Opin. Immunol. 27, 1–7. doi: 10.1016/j.coi.2013.12.005
Noy, R., and Pollard, J. W. (2014). Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61. doi: 10.1016/j.immuni.2014.06.010
Ohue, Y., and Nishikawa, H. (2019). Regulatory T (Treg) cells in cancer: can Treg cells be a new therapeutic target? Cancer Sci. 110, 2080–2089. doi: 10.1111/cas.14069
Papin, A., Tessoulin, B., Bellanger, C., Moreau, A., Le Bris, Y., Maisonneuve, H., et al. (2019). CSF1R and BTK inhibitions as novel strategies to disrupt the dialog between mantle cell lymphoma and macrophages. Leukemia 33, 2442–2453. doi: 10.1038/s41375-019-0463-3
Qiao, W., Han, Y., Jin, W., Tian, M., Chen, P., Min, J., et al. (2016). Overexpression and biological function of TMEM48 in non-small cell lung carcinoma. Tumour Biol. 37, 2575–2586. doi: 10.1007/s13277-015-4014-x
Quail, D. F., and Joyce, J. A. (2013). Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437. doi: 10.1038/nm.3394
Racle, J., de Jonge, K., Baumgaertner, P., Speiser, D. E., and Gfeller, D. (2017). Simultaneous enumeration of cancer and immune cell types from bulk tumor gene expression data. eLife 6:e026476. doi: 10.7554/eLife.26476
Ramesh, A., Kumar, S., Nandi, D., and Kulkarni, A. (2019). CSF1R- and SHP2-inhibitor-loaded nanoparticles enhance cytotoxic activity and phagocytosis in tumor-associated macrophages. Adv. Mater. 31:e1904364. doi: 10.1002/adma.201904364
Sangro, B., Gomez-Martin, C., de la Mata, M., Inarrairaegui, M., Garralda, E., Barrera, P., et al. (2013). A clinical trial of CTLA-4 blockade with tremelimumab in patients with hepatocellular carcinoma and chronic hepatitis C. J. Hepatol. 59, 81–88. doi: 10.1016/j.jhep.2013.02.022
Sanyal, R., Polyak, M. J., Zuccolo, J., Puri, M., Deng, L., Roberts, L., et al. (2017). MS4A4A: a novel cell surface marker for M2 macrophages and plasma cells. Immunol. Cell Biol. 95, 611–619. doi: 10.1038/icb.2017.18
Shen, D. W., Ma, J., Okabe, M., Zhang, G., Xia, D., and Gottesman, M. M. (2010). Elevated expression of TMEM205, a hypothetical membrane protein, is associated with cisplatin resistance. J. Cell. Physiol. 225, 822–828. doi: 10.1002/jcp.22287
Song, Y., Gao, J., Guan, L., Chen, X., Gao, J., and Wang, K. (2018). Inhibition of ANO1/TMEM16A induces apoptosis in human prostate carcinoma cells by activating TNF-alpha signaling. Cell Death Dis. 9:703. doi: 10.1038/s41419-018-0735-2
Taylor, N. A., Vick, S. C., Iglesia, M. D., Brickey, W. J., Midkiff, B. R., McKinnon, K. P., et al. (2017). Treg depletion potentiates checkpoint inhibition in claudin-low breast cancer. J. Clin. Invest. 127, 3472–3483. doi: 10.1172/JCI90499
Terai, S., Ueda-Hayakawa, I., Nguyen, C. T. H., Ly, N. T. M., Yamazaki, F., Kambe, N., et al. (2018). Palisaded neutrophilic and granulomatous dermatitis associated with systemic lupus erythematosus: possible involvement of CD163(+) M2 macrophages in two cases, and a review of published works. Lupus 27, 2220–2227. doi: 10.1177/0961203318809892
Topalian, S. L., Hodi, F. S., Brahmer, J. R., Gettinger, S. N., Smith, D. C., McDermott, D. F., et al. (2012). Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454. doi: 10.1056/NEJMoa1200690
Tu, E., Chia, P. Z., and Chen, W. (2014). TGFbeta in T cell biology and tumor immunity: angel or devil? Cytokine Growth Fact. Rev. 25, 423–435. doi: 10.1016/j.cytogfr.2014.07.014
Van Overmeire, E., Laoui, D., Keirsse, J., Van Ginderachter, J. A., and Sarukhan, A. (2014). Mechanisms driving macrophage diversity and specialization in distinct tumor microenvironments and parallelisms with other tissues. Front. Immunol. 5:127. doi: 10.3389/fimmu.2012.00127
Veremeyko, T., Yung, A. W. Y., Anthony, D. C., Strekalova, T., and Ponomarev, E. D. (2018). Early growth response gene-2 is essential for M1 and M2 macrophage activation and plasticity by modulation of the transcription factor CEBPbeta. Front. Immunol. 9:2515. doi: 10.3389/fimmu.2018.02515
Villarreal, D. O., L’Huillier, A., Armington, S., Mottershead, C., Filippova, E. V., Coder, B. D., et al. (2018). Targeting CCR8 induces protective antitumor immunity and enhances vaccine-induced responses in colon cancer. Cancer Res. 78, 5340–5348. doi: 10.1158/0008-5472.CAN-18-1119
Wang, D., Yang, L., Yue, D., Cao, L., Li, L., Wang, D., et al. (2019). Macrophage-derived CCL22 promotes an immunosuppressive tumor microenvironment via IL-8 in malignant pleural effusion. Cancer Lett. 452, 244–253. doi: 10.1016/j.canlet.2019.03.040
Wang, Y., Zhang, Y., Herman, J. G., Linghu, E., and Guo, M. (2017). Epigenetic silencing of TMEM176A promotes esophageal squamous cell cancer development. Oncotarget 8, 70035–70048. doi: 10.18632/oncotarget.19550
Wang, Y. C., He, F., Feng, F., Liu, X. W., Dong, G. Y., Qin, H. Y., et al. (2010). Notch signaling determines the M1 versus M2 polarization of macrophages in antitumor immune responses. Cancer Res. 70, 4840–4849. doi: 10.1158/0008-5472.CAN-10-0269
Ye, Y., Xu, Y., Lai, Y., He, W., Li, Y., Wang, R., et al. (2018). Long non-coding RNA cox-2 prevents immune evasion and metastasis of hepatocellular carcinoma by altering M1/M2 macrophage polarization. J. Cell. Biochem. 119, 2951–2963. doi: 10.1002/jcb.26509
Yu, N., Li, X., Song, W., Li, D., Yu, D., Zeng, X., et al. (2012). CD4(+)CD25 (+)CD127 (low/-) T cells: a more specific Treg population in human peripheral blood. Inflammation 35, 1773–1780. doi: 10.1007/s10753-012-9496-8
Zhu, A. X., Finn, R. S., Edeline, J., Cattan, S., Ogasawara, S., Palmer, D., et al. (2018). Pembrolizumab in patients with advanced hepatocellular carcinoma previously treated with sorafenib (KEYNOTE-224): a non-randomised, open-label phase 2 trial. Lancet Oncol. 19, 940–952. doi: 10.1016/S1470-2045(18)30351-6
Keywords: tumor immunity, hepatocellular carcinoma (HCC), TMEM205, prognostic factor, the Cancer Genome Atlas (TCGA)
Citation: Rao J, Wu X, Zhou X, Deng R and Ma Y (2020) TMEM205 Is an Independent Prognostic Factor and Is Associated With Immune Cell Infiltrates in Hepatocellular Carcinoma. Front. Genet. 11:575776. doi: 10.3389/fgene.2020.575776
Received: 24 June 2020; Accepted: 11 September 2020;
Published: 14 October 2020.
Edited by:
Shibiao Wan, St. Jude Children’s Research Hospital, United StatesReviewed by:
Ravikanth Nanduri, National Cancer Institute (NCI), United StatesTanima Bose, Ludwig Maximilian University of Munich, Germany
Copyright © 2020 Rao, Wu, Zhou, Deng and Ma. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ronghai Deng, bWRkcmg4MUAxNjMuY29t; Yi Ma, YW5odWltYXlpQDIwMDJAMTYzLmNvbQ==
†These authors have contributed equally to this work and share first authorship
 Jiawei Rao
Jiawei Rao Xukun Wu
Xukun Wu Xiaozhuan Zhou
Xiaozhuan Zhou Ronghai Deng
Ronghai Deng Yi Ma
Yi Ma