A Corrigendum on
Heat Shock Improves Random Spore Analysis in Diverse Strains of Saccharomyces cerevisiae
by Burke, M. K., McHugh, K. M., and Kutch, I. C. (2020). Front. Genet. 11:597482. doi: 10.3389/fgene.2020.597482
In the original article, there was a mistake in Figure 2 as published. The originally published figure had a panel out of order. The corrected Figure 2 appears below.
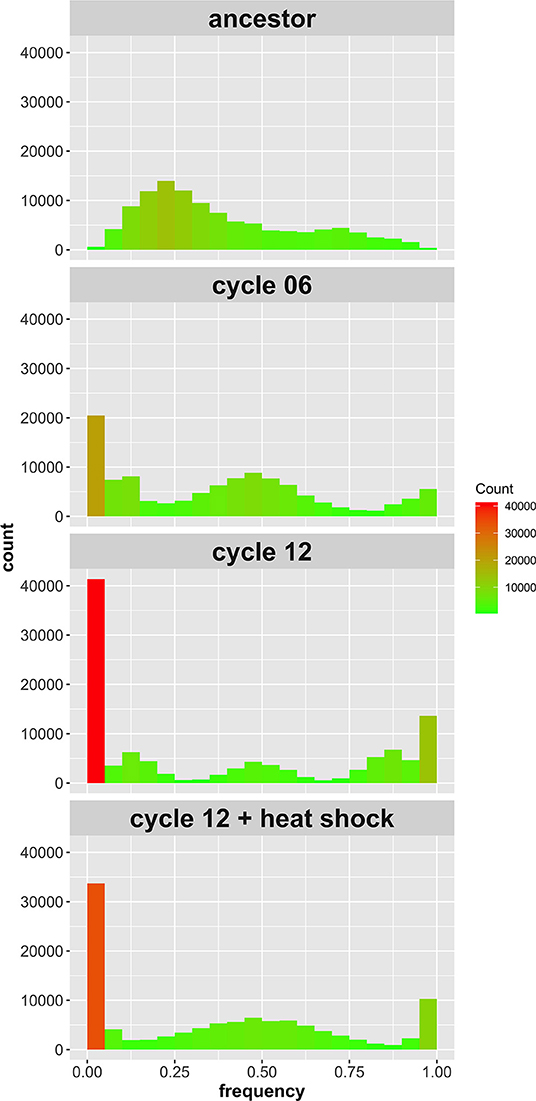
Figure 2. Site frequency spectrum of SNPs identified in the 4-parent population initially, after six outcrossing events, after 12 outcrossing events, and after 12 outcrossing events + a heat shock of 55°C for 20 min. Color in included in the histograms to emphasize the bins with high counts (red) compared to those with low counts (green); notice the reduction in sites with frequency = 0 or 1 between cycle 12 and cycle 12 + heat shock.
The authors apologize for this error and state that this does not change the scientific conclusions of the article in any way. The original article has been updated.
Keywords: random spore analysis, heat shock, sporulation, genomics, multiparent mapping population
Citation: Burke MK, McHugh KM and Kutch IC (2021) Corrigendum: Heat Shock Improves Random Spore Analysis in Diverse Strains of Saccharomyces cerevisiae. Front. Genet. 12:654893. doi: 10.3389/fgene.2021.654893
Received: 17 January 2021; Accepted: 19 January 2021;
Published: 05 February 2021.
Approved by:
Frontiers Editorial Office, Frontiers Media SA, SwitzerlandCopyright © 2021 Burke, McHugh and Kutch. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Molly K. Burke, bW9sbHkuYnVya2VAb3JlZ29uc3RhdGUuZWR1
 Molly K. Burke
Molly K. Burke Kaitlin M. McHugh
Kaitlin M. McHugh