- 1Institute for Virology and Immunobiology, University of Würzburg, Würzburg, Germany
- 2Institute for Experimental Infection Research, TWINCORE, Centre for Experimental and Clinical Infection Research, a joint venture between the Helmholtz Centre for Infection Research and the Hannover Medical School, Hannover, Germany
- 3Helmholtz Institute for RNA-Based Infection Research (HIRI), Würzburg, Germany
Dendritic cells (DCs) are key directors of tolerogenic and immunogenic immune responses. During the steady state, DCs maintain T cell tolerance to self-antigens by multiple mechanisms including inducing anergy, deletion, and Treg activity. All of these mechanisms help to prevent autoimmune diseases or other hyperreactivities. Different DC subsets contribute to pathogen recognition by expression of different subsets of pattern recognition receptors, including Toll-like receptors or C-type lectins. In addition to the triggering of immune responses in infected hosts, most pathogens have evolved mechanisms for evasion of targeted responses. One such strategy is characterized by adopting the host’s T cell tolerance mechanisms. Understanding these tolerogenic mechanisms is of utmost importance for therapeutic approaches to treat immune pathologies, tumors and infections. Transcriptional profiling has developed into a potent tool for DC subset identification. Here, we review and compile pathogen-induced tolerogenic transcriptional signatures from mRNA profiling data of currently available bacterial- or helminth-induced transcriptional signatures. We compare them with signatures of tolerogenic steady-state DC subtypes to identify common and divergent strategies of pathogen induced immune evasion. Candidate molecules are discussed in detail. Our analysis provides further insights into tolerogenic DC signatures and their exploitation by different pathogens.
Tolerogenic Dendritic Cells (DCs)
Tolerogenicity of DCs is an intrinsic functional definition for this cell type and their induction of T cell anergy, regulatory T cells and T cell deletion have been reported (1). All major DC subsets have been described to exert tolerogenic functions. Tolerogenic DCs were first described ex vivo, showing that UV-irradiated Langerhans cells induced T cell anergy (2). Spontaneous or UV-induced apoptotic cell death represents a source of self-antigens employed by DCs for tolerance induction. Steady-state mechanisms to maintain self-tolerance rely on the uptake of apoptotic material and its tolerogenic presentation (3–6). The ability to generate tolerogenic DCs in vitro facilitated their subsequent use for adoptive cell therapy in mice. However, in vitro generated immature DCs injected to protect from allo-transplant rejection matured, as indicated by their upregulation of B7-1 and B7-2 molecules, an unwanted phenomenon that was hypothesized to dampen the DCs tolerogenicity (7). Later, this hypothesis was confirmed by generating immature and maturation-resistant DCs in the same transplantation model, which dramatically extended the allograft survival time from 22 days to more than 120 days (8). Thus, maturation resistance was considered as a hallmark of tolerogenic DCs to maintain their immaturity. Several protocols have been developed to achieve maturation resistance, mostly using maturation inhibitors such as IL-10, TGF-β, dexamethasone, or vitamin D3 alone or in combinations (1). Reports on the transcriptional profiling of such DCs treated with tolerogenic substances followed and have been described elsewhere (9, 10).
Here, we analyzed transcriptional data sets deposited on public databases from steady-state migratory DCs (ssmDCs) and functionally similar spontaneously matured GM-CSF-derived bone marrow DCs (BM-DCs) as tolerogenic DC sources. Since ssmDCs act in a tolerogenic manner, despite their partial maturity, they phenotypically resemble much more mature DCs than non-migratory, immature DCs do. Therefore, they represent a more similar DC phenotype for our comparison of tolerance markers. We then analyzed transcriptional datasets of DCs treated with substances known to cause inflammation, including pathogen-derived molecules. The comparisons concentrated on bacteria or bacterial products but also included helminths, known as masters of immune evasion, but excluded protozoa and viruses. Candidate tolerogenic molecules that were highly upregulated by selected inflammatory or pathogenic stimuli in DCs are then discussed individually and compiled in tables.
Tolerogenic Markers Identified for Steady-State and Pathogen-Exposed DCs
Self-tolerance versus Microbial Immune Evasion
Dendritic cells residing in peripheral tissues at an immature stage act as immune sensors for pathogens. Pathogens, danger or inflammatory signals convert DCs into a mature/activated state which enables their migration into the draining lymph nodes. Subsequent stimulation of T cell immunity occurs by DC presentation of pathogen-derived antigens in the context of costimulation and proinflammatory cytokine production (11). In contrast, during homeostasis lymphoid organ-resident DCs and ssmDCs contribute to immune tolerance, thus controlling unwanted T cell responses against harmless or self-antigens (12).
Most microbes, especially those causing chronic infections, are evolutionarily well-adapted to their host. Such adaptation results in a balance between a pathogen-induced protective immune response and immune tolerance mechanisms that prevent microbial elimination. Infections with non-adapted microbes either kill the host rapidly or the microbe is immediately cleared by the host’s immune response. In both cases, the microbes cannot replicate and spread to another host. A successful microbe induces a chronic and preferably asymptomatic infection. This can be achieved by exploitation of the host’s immune tolerance mechanisms during pathogen–host coevolution.
Here, we analyzed public data in a comparative manner including tolerogenic and anti-inflammatory mRNA signatures of (1) steady-state DCs, (2) helminth-exposed DCs, (3) mycobacteria-exposed DCs, and (4) defined in vitro generated murine GM-CSF BM-DCs and human monocyte-derived DCs (MoDCs) treated with different inflammatory or pathogen-derived stimuli.
Transcriptional Signatures of Tolerogenic Migratory DCs under Steady-State Conditions
To identify tolerogenic DC signatures after pathogen stimulation, we first sought to identify comparative DC subsets known for their tolerogenic function as a reference dataset. While CCR7− resident DCs appear at an immature stage, CCR7+ ssmDCs undergo a homeostatic maturation process reaching a semi-mature stage, which is characterized by low expression of MHC II and costimulatory molecules, such as CD40 and CD86, and the absence of proinflammatory cytokine production (13–16). In several respects, steady-state plasmacytoid DCs (pDCs) resemble resident CD4+ or CD8α+ conventional DCs (cDCs) of cutaneous lymph nodes and spleen (Figure 1). After pathogen-induced maturation DCs upregulate MHC II, CD40 and CD86 molecules on their surface (14, 15). Depending on the stimulus, mature RelB+++, RelA+++, and cRel+++ DCs differ qualitatively in the production of the proinflammatory cytokines IL-6, TNF, IL-1β, IL-12p70, IL-23, or type-I interferon, while RelB+++, RelA+, and cRel+ ssmDCs induce Tregs by their release active TGF-β+ from its latent form of surface-bound latency-associated peptide (LAP) molecules (14–16). While tolerogenic functions of ssmDCs have been described by many authors, the demonstration of T cell tolerogenicity by immature lymph node-resident DCs is much less understood (17). Thus, due to their increased maturity, we selected the tolerance markers of ssmDCs for comparison with pathogen-induced DCs.
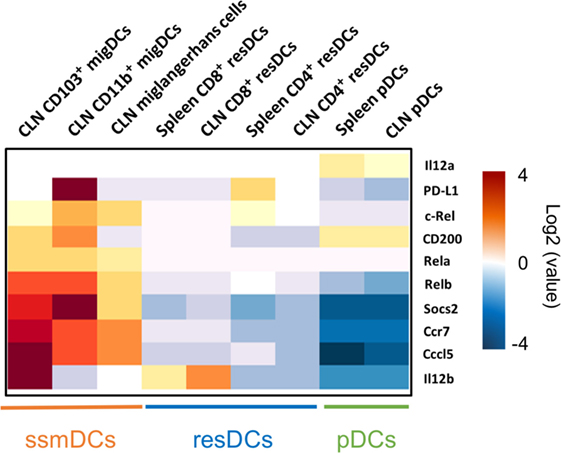
Figure 1. Major tolerogenic signatures upregulated on CCR7+ steady-state migratory dendritic cells (ssmDCs). Selected tolerogenic transcripts of IL12b, RelB, Socs2, Cd200, Cd274, and Ccl5 are upregulated in CCR7+ ssmDCs (CD103+ migDC, CD11b+ migDC, Langerhans cells) as compared to CCR7− resident DC subsets (CD8+ resDCs, CD4+ resDCs) or plasmacytoid DCs (pDCs). Data were obtained from the Immgen database (http://www.immgen.org/).
In ssmDCs increased transcription of Cd274 (PD-L1), CD200, Socs2, Relb, Ccl5, and IL12b was observed as compared with pDCs and resDCs (Figure 1). Their enhanced transcription was observed in all three subsets of semimatured CCR7+ ssmDCs but not immature resident DCs of lymph nodes (Figure 1). In addition, high levels of CD83, Cd150, Aldh1a2 (Raldh2,) Adora2a, and Itgb8 were found in ssmDCs (Table 1). Of these 11 molecules, 6 were also found in spontaneously matured BM-DCs (Table 1). The individual roles and mechanisms of tolerogenicity are explained below or referred to in Tables 1–5. Although the extent to which GM-CSF-derived BM-DCs resemble cDCs is still a matter for debate (18), the tolerogenic signatures observed in spontaneously matured BM-DCs (19) are strikingly similar to those observed in ssmDCs (14–16) (Table 1).
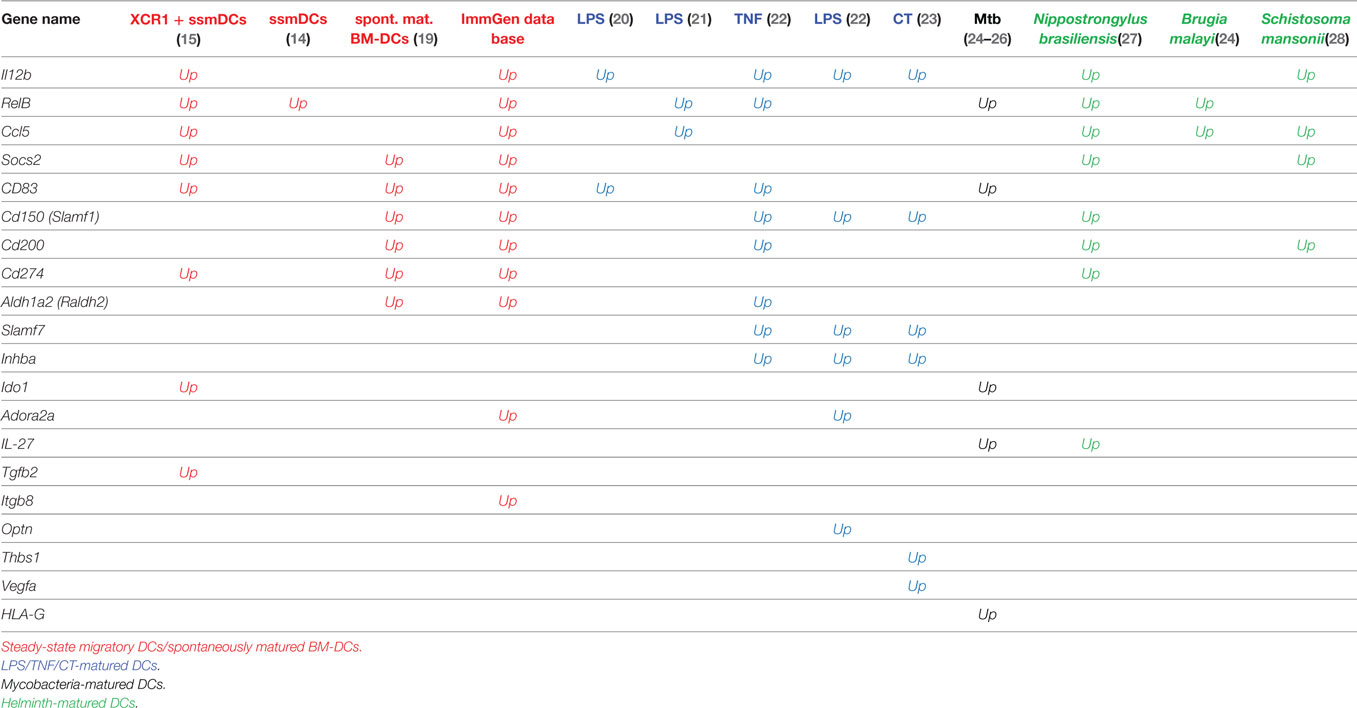
Table 1. Tolerogenic genes upregulated more than log2-fold by DCs matured during steady state, inflammation, or by pathogens.
Tolerogenic Signatures of DCs Induced by Helminths
Due to evolutionary pressure, phylogenetically distinct parasitic worms—collectively termed “helminths”—convergently evolved the ability to manipulate their host’s immune systems. In nearly all cases, the antihelminth type 2 immunity of M2 macrophages and T helper cell 2 (Th2) cells fails to eliminate the worms (59, 60); hence helminths persist within their hosts for years. Helminths often exploit the host’s immune regulation machinery with DCs being major targets (59, 61, 62).
Type 2 immunity, in contrast to type 1, is promoted by weaker costimulation and/or absence of proinflammatory and polarizing cytokines such as IL-12p70 and IL-23 (13, 63). Moreover, the DC potential to induce type 2 immunity can be associated with tolerogenic mechanisms such as IL-10 secretion (63). Phenotypic maturation of DCs occurs after recognition of pathogen-associated molecular patterns (PAMPs) frequently inducing canonical NF-κB signaling (involving classical IκBα, -β, and -ε, NF-κB1 p50, RelA, and c-Rel). In contrast, recognition of helminths and their products by DCs results only in partial maturation resulting in low levels of costimulatory molecules at the surface and poor release of proinflammatory cytokines (64). It is believed that the non-canonical NF-κB pathway (Nfκb2/p52, RelB) not only direct cell development (65) but also might play a role in the regulation of immune tolerance (14, 66–68). Transcriptomic analyses of human DCs treated with Brugia malayi revealed upregulation of RELB and NFκB2 (24) and RelB in DCs isolated from mice after infection with Nippostrongylus brasiliensis (27) or Schistosoma mansoni eggs (28) (Table 1). This was similar to what has been observed in ssmDCs which induced Foxp3+ Tregs from naive T cells (14). In line with this hypothesis, Lacto-N-fucopentaose III, a carbohydrate found in S. mansoni egg antigen, has been shown to activate the alternative NF-κB pathway in DCs (69). Thus, non-canonical NF-κB activation in the absence of low activity of canonical RelA and cRel may be characteristic for tolerogenic DCs in helminth infections.
The activation status and cytokine release of DCs fine-tunes the polarization of different T cell-effector and regulatory mechanisms. Suppressor of cytokine signaling (SOCS) proteins play decisive roles in innate immune cell signaling. They modify the polarization of immune responses by negative regulation of cytokine signals (70, 71). Different helminth species promote upregulation of Socs2 and Socs3 (24, 27) (Table 1), which may skew immune responses toward a Th2-biased anti-inflammatory phenotype. Indeed, it was shown that SOCS3-transduced DCs express low levels of MHC II and CD86 molecules on their cell surface and produced high levels of IL-10 but low levels of proinflammatory cytokines such as IL12p70. They thereby induced Th2-cell differentiation in mice supporting allergic Th2 responses but impairing Th1/Th17 development by means of immune deviation toward Th2 as shown in the autoimmune model EAE (72, 73). As described above, tolerogenic ssmDCs express Socs2 (Table 1). Therefore, induction of Socs2 during helminth infection might even inhibit Th2 differentiation and instead support a tolerogenic environment (27, 74). It is not clear whether helminths induce Socs expression directly or through indirect cell mechanisms such as host-derived cytokines. For example, anti-inflammatory Il27 is expressed in DCs after immunization with Nippostrongylus brasilienis (27) (Table 1). IL-27 induces expression of Socs3 in mouse and human cells leading to induction of IL-10-producing Tr1 cells (75).
Different DC populations exposed to helminths induce expression of the regulatory cytokines IL12b and IL-10 (27, 28). CD103+ migratory mature DCs from N. brasiliensis and S. mansoni infected mice significantly upregulate IL12b (27), also expressed in ssmDCs (Figure 1; Table 1).
Among others, Cd200 and Cd274 (PD-L1) were upregulated in DCs from N. brasiliensis immunized mice (Table 1). As detailed below, PD-L1 transmits inhibitory signals to PD-1 (CD279) on T cells. This interaction modifies TCR signaling, results in anergy or functional inactivation of T cells and is currently used for anticancer “checkpoint” inhibitory therapies (76, 77). PD-L1 expression would certainly support the chronicity of helminth infection. Suppression of T cell responses by PD1 during helminth infections has mainly been attributed to macrophages expressing PD-L1 and/or PD-L2 (78–80). Although the role of PD-L1 on DCs was not experimentally addressed, it may play a similar role.
Gene expression profiling using microarray or RNA sequencing technologies has been widely used to reveal cellular processes involved in host immune responses to different pathogens. Transcriptomic meta-analyses characterizing host immune responses against helminths have shown robust effects on immune gene signatures across different species (62). However, the common tolerogenic gene signature of DCs during helminth infection has not been addressed. Despite the fact that transcriptional profiling of DCs would improve our understanding of helminth effect during infection, the available helminth-related datasets are limited and further studies are required.
Tolerogenic Markers Expressed after Infection with Mycobacterium tuberculosis (Mtb)
During coevolution with the human immune system, Mtb has developed multiple immune evasion strategies (81). To address whether Mtb is able to induce tolerogenic gene signatures in DCs, we analyzed transcriptional profiles of human DCs infected with Mtb and evaluated those for known tolerogenic markers.
Monocyte-derived DCs infected with Mtb or BCG highly upregulated the two tolerogenic genes IDO-1 and IL27. IDO-1 upregulation was detected already 8 h after infection of human MoDCs, whereas IL27 transcripts were detected only upon Mtb, but not BCG, infection (25). Others showed upregulation of RELB, CD83, and HLA-G in MoDCs after 16 h of Mtb infection (24). The tolerogenic function of RELB is discussed below. CD83 might also confer a regulatory function, as indicated by inhibition of T-cell proliferation that was mediated by the soluble form of the CD83 protein (58). Finally, HLA-G has been shown to induce human MoDC tolerogenicity via its CD85b/ILT4 ligand in huILT4-transgenic mice, inducing anergy and suppressor T cells (82). Hence, expression of IDO, IL27, RELB, CD83, and HLA-G (Table 1) by DCs might promote tolerogenic responses in Mtb infection.
Tolerogenic Signatures of Murine and Human DCs Upregulated by Selected Inflammatory or Pathogenic Stimuli: TNF, Cholera Toxin, Lipopolysaccharide (LPS)
Transcriptional profiles of DCs stimulated in vitro under tolerogenic conditions have been reviewed before (10). Early transcriptional profiling work revealed that expression profiles of some cytokines are tightly regulated with time kinetic mRNA profiling revealing clear insights. IL-10 production stimulated by Escherichia coli LPS was only induced after 6 h in the DC cell line D1, but not earlier or later, whereas mRNA for TGF-β1 or IL-12p40 was detectable in time windows of more than 18–20 h after stimulation (83). DC cell line D1 showed IL-12p40 induction with LPS but not TNF (84) as reported for murine BM-DCs and human MoDCs (85). The fact that only two tolerogenic markers were identified in D1 cells may indicate a general limitation of obtaining transcriptional data from cell lines.
Of note, LPS stimulated DCs produce immunogenic Th1-polarizing IL-12p70, formed by the p35/p40 heterodimer (Il12a and Il12b genes), but the protein amounts of IL-12p40 secreted by DCs are typically 50–100 times higher than the amount measured for IL-12p70. Similarly, the IL-23 heterodimer secretion, composed of p19/p40 (Il23a Il12b genes) is much lower than p40 by cholera toxin stimulation of DCs (22, 23). This opens space for speculation on a counterbalancing and thereby tolerogenic role for excessive IL-12p40 production.
Dendritic cell maturation induced by inflammatory or microbial products triggering DAMPs or PAMPs, respectively, direct polarized Th1, Th2, or Th17 responses. Previously, we performed transcriptional profiling of murine GM-CSF generated BM-DCs and human MoDCs. Selected in vitro maturation protocols for induction of Th1 responses by LPS, Th2 by TNF and Th17 by cholera toxin (CT-DCs) were applied to both human and mouse DCs for the same time period of 6 h (22, 23) (GEO data bases GSE106080). Among the clearly immunogenic transcriptomic signatures, we also identified additional molecules at the protein level that exert tolerogenic immune functions. These include IL-10 production by LPS-DCs (86), Tr1 induction by Trypanosoma-matured or TNF-DCs after three injections (22, 87) and Foxp3+ Treg induction via TGF-β plus CTLA-2, a newly identified tolerogenic molecule from CT-DCs (23).
It remains a subject for debate whether the tolerogenic signature observed after infection has evolved as protective mechanism by the host or is actively induced by the pathogen. Pathogens aim to prevent their elimination and also the host aims to survive. If a pathogen cannot be eliminated, the host has to develop a protection strategy including the prevention of immunopathology. Excessive immune responses may be more deleterious than microbial pathogenicity in the host, as observed in sepsis. Thus, host-intrinsic negative feedback regulation of immune stimulation may be advantageous. To address this in our analyses, we included TNF as a non-pathogen-derived inflammatory stimulus. Interestingly, four tolerogenic genes showed increased mRNA transcription overlapping between TNF, CT and LPS stimulation (Figure 2) (Tables 1 and 2).
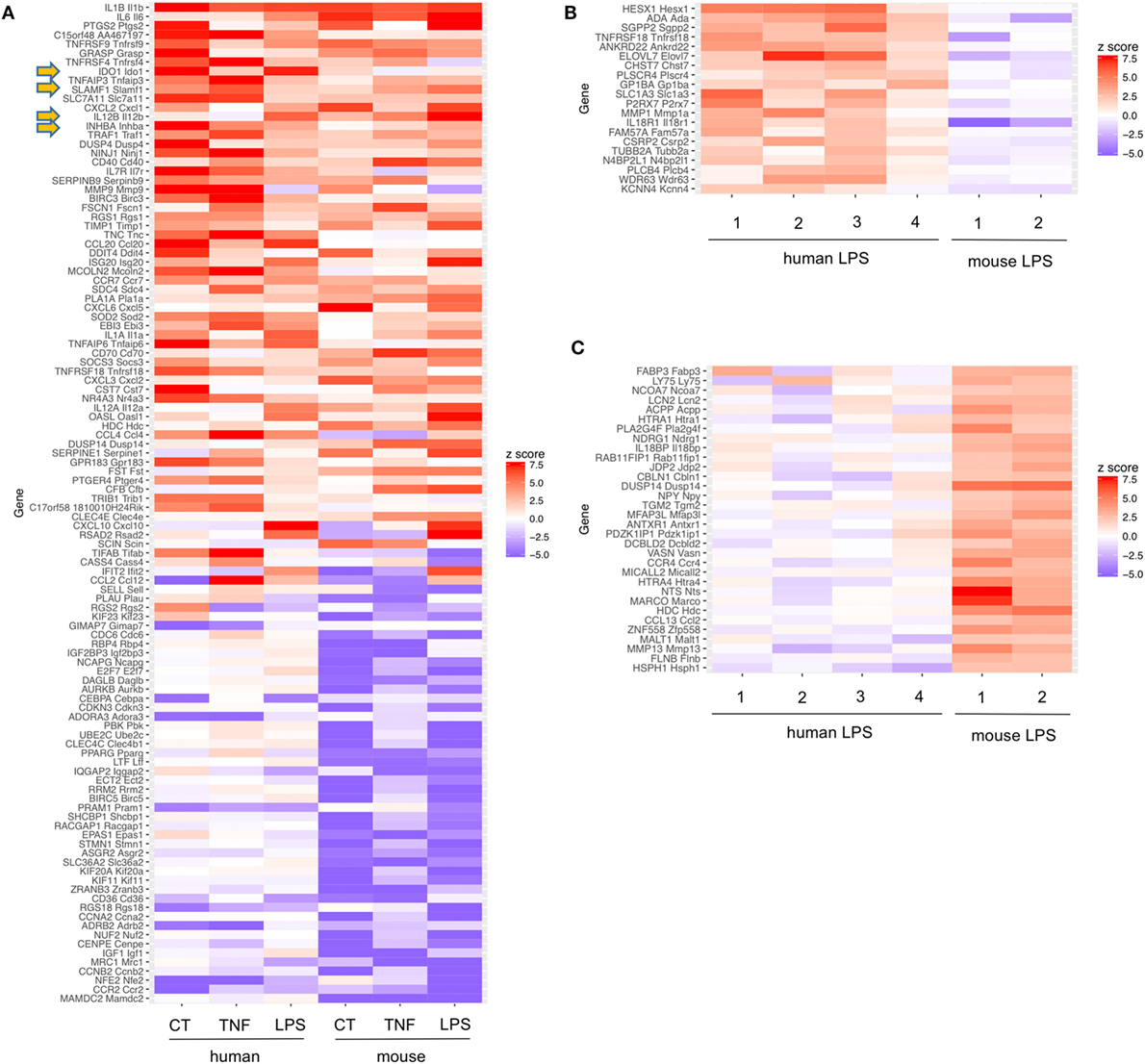
Figure 2. Common tolerance signatures of human monocyte-derived dendritic cells (MoDCs) or mouse bone marrow DCs (BM-DCs) stimulated with cholera toxin (CT), TNF, or lipopolysaccharide (LPS). (A) Among the top 14 genes upregulated under these conditions in DC of two different species, the 4 tolerogenic molecules IDO, SLAM, Inhibin A, and IL12b were found commonly upregulated (arrows). Expression signatures of strongly regulated genes in human and mouse DCs stimulated with CT, TNF, or LPS. Only genes with 1:1 ortholog mappings between human and mouse (obtained from MGI) were retained. z-scores were computed from the log2 fold changes for each experiment. Only genes having a z–score >4 or <−4 in at least two experiments are shown. Of note, for generation of human MoDCs additional IL-4 was added. Murine data are from Ref. (22, 23), human data obtained from GEO data bases (GSE106080). (B,C) Expression signatures of LPS stimulated human DCs [1 = (85); 2 = (88); 3 = (89); 4 = GSE106080] or mouse DCs [1 = (90); 2 = (22)]. Only genes with probe sets on each of the microarrays used were retained and z-scores were computed as in Panel (A). In Panel (B), genes with z-score > 2 in at least two human experiments and z-score < 0 in both mouse experiments are shown. Panel (C) depicts genes with z-score > 2 in both mouse experiments and <0 in at least two human experiments.
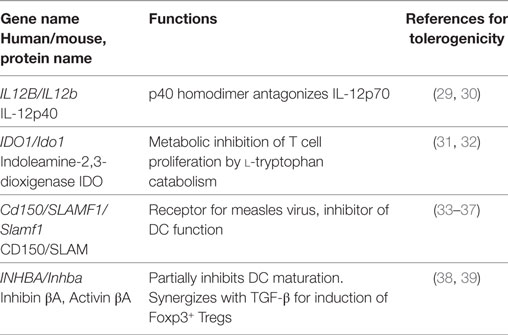
Table 2. Common transcripts induced under all six conditions (TNF, CT, LPS, each human and mouse data from Figure 2) and for which anti-inflammatory or tolerogenic functions have been reported.
Pathogens and inflammatory mediators induce numerous mechanisms of immunity in DCs. Additionally, molecules with tolerogenic or anti-inflammatory functions are induced. Mouse BM-DCs and human MoDC generated with GM-CSF (±IL-4) result from conversion of classical human CD14+ or mouse Ly-6Chigh monocytes into DCs, in a human-mouse interspecies comparison. As expected, common proinflammatory genes such as Il-1β, Il-6, and Cox2 (Ptgs2) were upregulated under all six conditions. Furthermore, four gene transcripts: Il12b, Ido-1, Cd150 (Slamf1), and Inhba (coding for Inhibin/Activin) with reported anti-inflammatory or tolerogenic function were upregulated under all 6 conditions by stimulation with TNF, CT, or LPS of both human MoDCs or mouse BM-DCs (Figure 2 arrows, Table 2). Taken together, as these four genes were also upregulated by TNF, this tolerogenic DC response may reflect a host-initiated protection mechanism to avoid immunopathology rather than a purely pathogen-driven strategy.
Besides the common tolerogenic genes upregulated by all three stimuli, additional tolerogenic transcripts were found by the individual stimuli LPS (Table 3), CT (Table 4), and TNF (Table 5). These data indicate that microbial adaptation to the host and induction of tolerogenic signatures by LPS and CT not only share mechanisms of tolerogenicity but also differ in their strategies of immune evasion. Therfore, LPS selectively upregulates mRNA for adenosine A2a receptor, optineurin, and Slamf7/CD319, while CT induces higher transcription of thrombospondin-1 (TSP1) and Vegfa indicating divergent tolerance strategies (Tables 3 and 4).
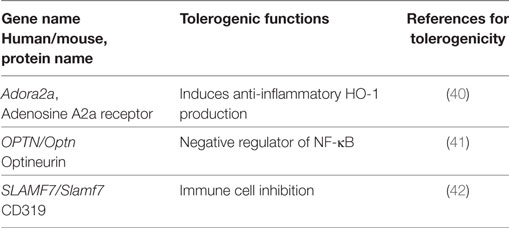
Table 3. Tolerogenic transcripts induced specifically by LPS (human and mouse data from Figure 2) and for which anti-inflammatory or tolerogenic functions have been reported.
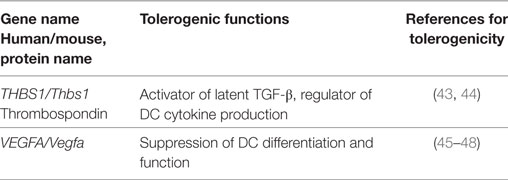

Table 4. Tolerogenic transcripts induced specifically by CT (human and mouse data from Figure 2) and for which anti-inflammatory or tolerogenic functions have been reported.
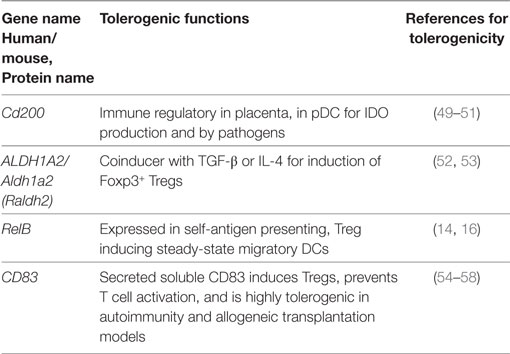
Table 5. Tolerogenic transcripts induced specifically by TNF (human and mouse data from Figure 2) and for which anti-inflammatory or tolerogenic functions have been reported.
Since tolerogenic signatures of differentially stimulated human MoDCs and mouse BM-DCs were strikingly similar (Table 2), we asked whether also distinct differences exist between DC from the two species. Surprisingly, very few genes were selectively upregulated by human MoDCs but remained unaltered or downregulated in murine BM-DCs and vice versa (Figures 2B,C). Among those, no tolerogenic genes appeared. Interestingly, differences in the expression of Gitr (Tnfrsf18) were found, confirming known differences in expression and function of GITR in mice and humans on DCs (91). Thus, with respect to LPS sensing and transcriptional responses, human MoDCs and murine BM-DCs are remarkably similar.
The Role of Selected Tolerogenic Molecules in Homeostasis and Immune Evasion
Il12b
Il12b codes for IL-12p40 protein forming homo- and heterodimers. Two heterodimers can be formed with p40: p35/p40 that are linked via a disulfide bond to form IL-12p70 and p19/p40 to form IL-23. The release of IL-12p70 by DCs plays a pivotal role in the induction of Th1 responses (92, 93) while IL-23 supports Th17 generation (94, 95). However, the p40 monomer and especially the homodimer (p40)2 have been shown to strongly inhibit IL-12-dependent T cell or Th1 responses in vitro and in vivo (29, 30, 96), mainly by competing with IL-12p70. Interestingly, the total serum IL-12, and the ratio of IL-12p40/IL-12p70 increased with age in healthy individuals compared to IL-12p70 levels (97). This observation likely contributes to impaired immunity in the elderly. The expression of IL12b by ssmDCs is observed only in the CD103+ Langerin+ CD11blow subset (15), and is significantly higher on ssmDCs when compared to lymphoid organ-resident DCs (Figure 1) (Table 1). Since IL12a mRNA coding for IL-12p35, is undetectable or at very low levels in any of the subsets under steady-state conditions, this may point to a tolerogenic role of p40 homodimers as described.
Relb
RelB is an NF-κB/Rel transcription factor family member associated with both tolerogenic and immunogenic functions (98). The RelB-p50 heterodimer has been associated with inflammatory and immunogenic responses (68). In this case, it functions through the RelA-NF-κB canonical pathway and cooperates with the cRel-p50 heterodimer (65). cRel is specifically required for IL-12p70 production (99). On the other hand, the RelB-p52 heterodimer, which functions through the NF-κB non-canonical pathway, was shown to be important for organogenesis of lymphoid organs (100), for normal development of splenic CD4+ and CD8+ (101, 102) and ssmDCs (14). RelB, but absence of (or extremely low levels) of RelA or cRel, is expressed by migratory DCs both under steady-state conditions and upon immune activation (14, 15) (Figure 1). In the peripheral lymph nodes of p52−/− mice, the ssmDC subsets were severely reduced while the resident DCs were not affected. In contrast, p50−/− mice did not show a specific preference for migratory or resident DCs and both were equally reduced (14). Additionally, RelB-deficient animals show a severe pathological phenotype characterized by inflammatory infiltrates into multiple organs, which is caused by hyper activity of conventional T cells (100). RelB+ ssmDCs have been shown to be either critical for conversion of naive T cells into Foxp3+ iTreg (14, 103), or for maintaining the homeostatic Foxp3+ natural Treg pool (16). Together, the available data indicate that moderate RelB expression in DCs alone is associated with lymphoid organogenesis and tolerogenic functions, whereas coexpression of RelB with RelA and cRel at high levels in DCs marks immunogenic functions.
CC Chemokine Ligand 5 (Ccl5)
The Ccl5 gene encodes CCL5, also known as RANTES, has been described as a gene expressed by activated T cells, macrophages, eosinophils, fibroblasts, epithelial cells as well as certain types of tumor cells. CCL5 plays an important role in the migration of different leukocytes toward inflammatory sites where it acts through its binding to CCR1, CCR3, or CCR5 (104). One interesting observation is that certain types of tumors express high levels of CCL5, which is a predictor of a poor prognosis (105, 106). Blocking of CCL5 can redirect myeloid-derived suppressor cells (MDSCs) and thereby improve antitumor immunity (107). CCL5 has been shown to be important for the generation of CD11b+/Gr-1+ MDSCs and its absence alters their differentiation and their immunosuppressive capacity (108). CCL5 release by NKT cells was required for the recruitment of antigen-specific CD8+ regulatory T cells and TGFβ-dependent tolerogenic antigen-presenting cells in order to mediate tolerance in the immune-privileged anterior eye chamber (109). Given the higher Ccl5 expression by ssmDCs relative to resident DCs it will be interesting to uncover its precise function in these cell types (Figure 1) (15).
IL-10
Several TLR ligands, including LPS, induce IL-12p70 release from DCs to induce Th1 immunity and, in parallel, release of IL-10 (110). Listeria infection in neonates induces CD8α+ DCs to release IL-10 (111). The suppressive effect of IL-10 on Th1 responses is indirect via DCs or macrophages (112) and seems to control IFN-γ release but not proliferation of Th1 clones in vitro (113). This IL-10 production has been suggested to serve as a self-control mechanism to avoid Th1-mediated immunopathology (114) but also as a means of microbial immune evasion (115, 116). IL-10 can inhibit the differentiation of monocytes into Mo-DCs (117). Others found DC-to-DC effects by observing CpG-activated cDC-derived IL-10 blocked pDC release of type I interferons (118). Persistent production of IL-10 may then facilitate the conversion of Th1 (or Th2) responses into a IL-10+ Foxp3− regulatory T cell response of the Tr1 type (119), similar to what had been observed for harmless antigen application and steady-state transport and Tr1 induction by lung DCs (120). The detailed regulation of IL-10 production (121) or its role of IL-10 for Tr1 cell induction has been reviewed elsewhere (122). However, although all this indicates an important role of IL-10 in immune tolerance, remarkably in none of the data sets analyzed herein (Figure 2; Table 1) was IL-10 identified as part of the tolerogenic transcriptional signature in DCs. The reasons for this may depend on delayed gene transcription kinetics or epigenetic regulation, thus identifying a limitation of tolerogenic transcriptional profiling.
TGF-β/Itgb8
Foxp3 is the major transcription factor directing functions of thymus-derived natural Foxp3+ Tregs, but also peripherally induced Foxp3+ iTregs (123). Therefore, the production or employment of TGF-β by tolerogenic DCs for Treg generation or maintenance is of interest. TGF-β inhibits the maturation of BM-DC (124). However, murine BM-DCs produce soluble TGF-β when stimulated by Lactobacilli (125) and its release may be under the control of GITR (91). GM-CSF cultured BM-DCs lack the surface expression of LAP which can bind TGF-β in a latent form before it can be released for Treg induction (126). Therefore, they are unable to mediate iTreg conversion from naive CD4+ T cells in vitro without addition of exogenous TGF-β (23). In contrast, lymph node DCs express LAP and the partially matured ssmDCs do so at even higher levels when compared with immature resident DCs (14).
The release of active TGF-β from its latent form is the critical event in TGF-β biological activity. The integrins αVβ6 (Itgav, Itgb6) (127), αVβ8 (Itgav, Itgb8) (128), and TSP1 (43) have been described to mediate non-proteolytic release of TGF-β, while metalloproteinase 9 (MMP9) performs proteolytic release (129). The activity of integrin αVβ8 has been shown as a key mechanism to prevent autoimmunity by maintaining Treg activity (130). Thus, these genes might be better markers for transcriptional signatures of TGF-β activity, although not identified in any of the RNA profiling data sets analyzed here (Figure 1). This indicates that not all important tolerogenic molecules are transcriptionally regulated and can be identified in such studies. A broader tolerogenic transcriptional signature was also identified for the subset of incompletely matured XCR1+ ssmDCs ex vivo, including the upregulation of TGF-β2 (15).
Cd150/Slamf1
Cd150 is upregulated on activated lymphoid and myeloid cells and acts via homotypic interaction (131). It represents the main human receptor for measles virus has been shown to inhibit DC functions (Table 2). Interestingly, the SH2D1A gene encoding for the SLAM-associated adapter protein to mediate SLAM signaling is mutated on X-linked immunodeficiency patients and responsible for the observed uncontrolled T and B lymphocyte proliferation after an EBV infection (132, 133). These data indicate that intact SLAM acts as an immune control molecule to prevent over activation of adaptive immunity during EBV infection.
Indoleamine 2,3-dioxygenase (Ido)
IDO is an enzyme catabolizing l-tryptophan. Deprivation of this essential amino acid in the environment of proliferating T cells results in metabolic starvation, apoptosis and thus inhibition of the T cell responses (134). Interestingly, in pDCs a TGF-β-dependent tolerogenic function of IDO has been reported that is independent of its enzymatic activity (135). IDO also plays a decisive role in establishment of LPS tolerance via control of the aryl hydrocarbon receptor signaling (136).
Inhba
The genes INHBA/Inhba encode for the Inhibin-βA or Activin-βA protein. Inhibin-βA forms homo- or heterodimers with other inhibin/activin family members to form the protein complexes Activin A (βA/βA homodimer), Inhibin B (α/βA heterodimer), or Inhibin AB (βA/βB heterodimer). They all belong to the TGFβ family (126) and many of the TGF-β family members influence DC development and function (137). Inhibition of DC maturation has been reported for Activin A and Inhibin A (38). Activin A may cooperate with TGF-β to increase generation of Foxp3+ induced regulatory T cells (iTregs) (39). Why Inhibin/Activin and not directly TGF-β are targets of immune evasion at the transcriptional level requires further investigation.
Il27
IL-27 protein belongs to the IL-12 cytokine family and is a heterodimeric protein consisting of IL-27p28 and the Epstein-Barr virus-induced gene 3 (EBI3) (138). This cytokine is expressed early upon activation of antigen presenting cells. It has been shown to induce the initial step in Th-1 differentiation of naive CD4 T-cells by STAT1 dependent induction of T-bet (139). Besides this immunogenic function, several studies have analyzed the regulatory function of IL-27 during infection with various different pathogens (140). Infection with Mtb IL-27 was described to suppress T-cell responses by the reduction of TNF, IL-12p40, and IFN-γ expression and to inhibit T-cell recruitment and proliferation (141). Furthermore, IL-27 can induce the expression of IL-10 in activated CD4+ effector T-cells and thus reduce antimycobacterial activity (116).
Socs2
Suppressor of cytokine signaling proteins play important roles in both the maintenance of homeostasis and the resolution of inflammation (71). Recent evidence suggests that SOCS2 plays a role in immune regulation. Similar to SOCS1 and SOCS3, also SOCS2 regulates pattern recognition receptor signaling in both human and murine DCs by counterregulating their activation (142). Socs2−/− mice showed uncontrolled Th1 responses to Toxoplasma gondii, due to generalized proinflammatory responses to the infection (143). Besides innate immunity, SOCS proteins balance T helper cell polarization. SOCS1 and SOCS3 support Th17 cell generation by inhibiting Th1 differentiation while Th2 differentiation is regulated by SOCS3 (72, 73). SOCS2 was recently shown to play a major role in inhibiting the development of Th2 cells and Th2-associated allergic responses (74). However, whether SOCS expression in DCs is responsible for observed effects in T cells was not investigated by these studies. Here, we identified Socs2 transcript elevation in all ssmDCs and spontaneously matured BM-DCs (Table 1) and upon in vitro exposure of DCs with N. brasiliensis and S. mansoni, further suggesting its important role in immune regulation (27, 28).
Cd274
Cd274 encodes programmed death ligand-1 (PD-L1) which delivers inhibitory signals via PD1 into T cells to regulate the delicate balance between immune defense and tissue-damage. PD-L1 is constitutively expressed or upregulated after activation on wide hematopoietic and non-hematopoietic cells and affect the responses against self and foreign antigens (76). Unsurprisingly, to evade immunity, microbes and tumors exploit the PD1/PD-L pathway which may act in concert with other immunosuppressive signals to establish chronic infection and tumor survival (76). Evidence that PD1/PD-L1 pathway is one of the main factors of tumor immune escape in humans is provided by the strategy of PD1/PD-L1 blockade. In addition to PD-L1 expression by tumors, myeloid DCs infiltrating tumors also express PD-L1. PD-L1 blockade improves myeloid DC-mediated antitumor immunity in several types of cancer (144). The blockade of this so called “check-point” has already been applied to clinical cancer therapy (145).
Discussion and Future Perspectives by Single-Cell RNA-seq
The identification and the definition of DCs based on morphology, functional studies and surface markers have been subjected to many controversies and transcriptional studies have played a pivotal role in characterizing DC ontology (18). Disentangling DCs from monocytes and macrophages and understanding how DCs plasticity is shaped after stimulation or pathogen sensing remain technologically challenging because transcriptomics applied to a population of cells assumes a strict homogeneity among the cells, which does not reflect the biological reality. Genome-wide transcriptomics at the single-cell level (single-cell RNA-seq) is emerging as a powerful tool to phenotype cells and is elevating biased bulk approaches and profiling methods restricted to selected surface markers (146, 147). The revolution of single-cell RNA-seq lies in that cellular identities are no longer bounded by a restricted number of signals, but instead are inferred in an unbiased manner from an array of expressed genes. Single-cell RNA-seq can capture thousands of transcripts (148) to assess a cellular identity and enables profiling how a single-cell responds to stimulus. The response of in vitro differentiated DCs stimulated with three pathogenic components at the single-cell level (149, 150) revealed a dramatic difference between individual cells. The analysis demonstrated the existence of “gene modules” indicating the differential activation of gene circuits between cells where some cells are prone to mounting a precocious response, acting as “leaders” of an antiviral response. Furthermore, combining genome editing with CRISPR/Cas9-based technologies and single-cell RNA-seq helped to uncover the regulatory network controlling DC response to LPS (151). As a proof-of-concept the perturbation of Rela, Irf9, and Cebpb facilitated the decoupling of antiviral and inflammatory pathways. Such approaches, termed CRISPR-seq or Peturb-seq, are not only restricted to in vitro cultures, but can uncover the complexity of DC regulatory circuits in vivo. Notably, this approach has been used to resolve the contribution of STAT-1/2-dependent antiviral genes to myeloid cell function (151). Future applications of single-cell RNA-seq technologies should include in-depth studies of DCs exposed to pathogens, revealing their immunogenic and tolerogenic signatures.
Conclusion
Activation-associated changes enabling DCs to activate adaptive immune responses are well understood. More recently, the scientific community has given greater attention to the counterregulation of these activation processes due to the clinical success of the checkpoint inhibitors, especially to the PD-1/PD-L1 molecules. Understanding of the tolerogenic mechanisms limiting inflammation is of utmost importance for therapeutic approaches that target immune pathologies, tumors and infections. As such, transcriptional profiling of tolerogenic DCs may provide insights into strategies allowing homeostasis and exploitation of own regulatory machinery by tumors and microbes.
Here, we aimed to uncover tolerogenic signatures of infla-matory or pathogen-matured DCs that included known tolerogenic markers of non-inflammatory ssmDCs. The present study addresses mainly transcriptomic studies as performed by microarray technologies of inflammatory or candidate bacteria- or helminth-induced DC signatures. This offered only a limited ability to fully identify all tolerance-associated mRNA species. However, our analysis revealed tolerogenic and anti-inflammatory genes among the expected expression of inflammatory genes. We reviewed the tolerogenic signatures of DCs exposed to different stimuli from both in vitro and in vivo studies across different host tissues and DCs subsets of man or mouse. Surprisingly, all pathogens analyzed here seem to use a rather restricted pool of target molecules for immune evasion. In the future, the possibility to quantify minute amounts of RNA species from single cells will enable analysis of much more complex regulatory networks in a wide variety of DC subsets.
Author Contributions
All authors contributed by writing parts of the text and edited the final version of the text. Figures and tables were generated by EV, DA, FE, and ML.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The Helmholtz Institute for RNA-based Infection Research (HIRI) supported this work with a seed grant through funds from the Bavarian Ministry of Economic Affairs and Media, Energy and Technology (grant allocation nos. 0703/68674/5/2017 and 0703/89374/3/2017). Furthermore, this publication was funded by the German Research Foundation (DFG) and the University of Wuerzburg in the funding programme Open Access Publishing. We are grateful to Richard Brown for text editing.
References
1. Lutz MB. Differentiation stages and subsets of tolerogenic dendritic cells. In: Lutz MB, Romani N, Steinkasserer A, editors. Handbook of Dendritic Cells. Biology, Diseases and Therapy. Weinheim: VCH-Wiley (2006). p. 517–43.
2. Simon JC, Tigelaar RE, Bergstresser PR, Edelbaum D, Cruz PD Jr. Ultraviolet B radiation converts Langerhans cells from immunogenic to tolerogenic antigen-presenting cells. Induction of specific clonal anergy in CD4+ T helper 1 cells. J Immunol (1991) 146:485–91.
3. Huang FP, Platt N, Wykes M, Major JR, Powell TJ, Jenkins CD, et al. A discrete subpopulation of dendritic cells transports apoptotic intestinal epithelial cells to T cell areas of mesenteric lymph nodes. J Exp Med (2000) 191:435–44. doi:10.1084/jem.191.3.435
4. Hemmi H, Yoshino M, Yamazaki H, Naito M, Iyoda T, Omatsu Y, et al. Skin antigens in the steady state are trafficked to regional lymph nodes by transforming growth factor-beta1-dependent cells. Int Immunol (2001) 13:695–704. doi:10.1093/intimm/13.5.695
5. Huang FP, MacPherson GG. Continuing education of the immune system – dendritic cells, immune regulation and tolerance. Curr Mol Med (2001) 1:457–68. doi:10.2174/1566524013363573
6. Liu K, Iyoda T, Saternus M, Kimura Y, Inaba K, Steinman RM. Immune tolerance after delivery of dying cells to dendritic cells in situ. J Exp Med (2002) 196:1091–7. doi:10.1084/jem.20021215
7. Fu F, Li Y, Qian S, Lu L, Chambers F, Starzl TE, et al. Costimulatory molecule-deficient dendritic cell progenitors (MHC class II+, CD80dim, CD86-) prolong cardiac allograft survival in nonimmunosuppressed recipients. Transplantation (1996) 62:659–65. doi:10.1097/00007890-199609150-00021
8. Lutz MB, Suri RM, Niimi M, Ogilvie AL, Kukutsch NA, Rossner S, et al. Immature dendritic cells generated with low doses of GM-CSF in the absence of IL-4 are maturation resistant and prolong allograft survival in vivo. Eur J Immunol (2000) 30:1813–22. doi:10.1002/1521-4141(200007)30:7<1813::AID-IMMU1813>3.0.CO;2-8
9. Torres-Aguilar H, Aguilar-Ruiz SR, Gonzalez-Perez G, Munguia R, Bajana S, Meraz-Rios MA, et al. Tolerogenic dendritic cells generated with different immunosuppressive cytokines induce antigen-specific anergy and regulatory properties in memory CD4+ T cells. J Immunol (2010) 184:1765–75. doi:10.4049/jimmunol.0902133
10. Schinnerling K, Garcia-Gonzalez P, Aguillon JC. Gene expression profiling of human monocyte-derived dendritic cells – searching for molecular regulators of tolerogenicity. Front Immunol (2015) 6:528. doi:10.3389/fimmu.2015.00528
11. Steinman RM. Dendritic cells: understanding immunogenicity. Eur J Immunol (2007) 37(Suppl 1):S53–60. doi:10.1002/eji.200737400
12. Steinman RM, Nussenzweig MC. Avoiding horror autotoxicus: the importance of dendritic cells in peripheral T cell tolerance. Proc Natl Acad Sci U S A (2002) 99:351–8. doi:10.1073/pnas.231606698
13. Lutz MB, Schuler G. Immature, semi-mature and fully mature dendritic cells: which signals induce tolerance or immunity? Trends Immunol (2002) 23:445–9. doi:10.1016/S1471-4906(02)02281-0
14. Azukizawa H, Dohler A, Kanazawa N, Nayak A, Lipp M, Malissen B, et al. Steady state migratory RelB+ langerin+ dermal dendritic cells mediate peripheral induction of antigen-specific CD4+ CD25+ Foxp3+ regulatory T cells. Eur J Immunol (2011) 41:1420–34. doi:10.1002/eji.201040930
15. Ardouin L, Luche H, Chelbi R, Carpentier S, Shawket A, Montanana Sanchis F, et al. Broad and largely concordant molecular changes characterize tolerogenic and immunogenic dendritic cell maturation in thymus and periphery. Immunity (2016) 45:305–18. doi:10.1016/j.immuni.2016.07.019
16. Dohler A, Schneider T, Eckert I, Ribechini E, Andreas N, Riemann M, et al. RelB+ steady-state migratory dendritic cells control the peripheral pool of the natural Foxp3+ regulatory T cells. Front Immunol (2017) 8:726. doi:10.3389/fimmu.2017.00726
17. Vitali C, Mingozzi F, Broggi A, Barresi S, Zolezzi F, Bayry J, et al. Migratory, and not lymphoid-resident, dendritic cells maintain peripheral self-tolerance and prevent autoimmunity via induction of iTreg cells. Blood (2012) 120:1237–45. doi:10.1182/blood-2011-09-379776
18. Lutz MB, Strobl H, Schuler G, Romani N. GM-CSF monocyte-derived cells and Langerhans cells as part of the dendritic cell family. Front Immunol (2017) 8:935. doi:10.3389/fimmu.2017.01388
19. Vander Lugt B, Riddell J, Khan AA, Hackney JA, Lesch J, Devoss J, et al. Transcriptional determinants of tolerogenic and immunogenic states during dendritic cell maturation. J Cell Biol (2017) 216(3):779–92. doi:10.1083/jcb.201512012
20. Torri A, Beretta O, Ranghetti A, Granucci F, Ricciardi-Castagnoli P, Foti M. Gene expression profiles identify inflammatory signatures in dendritic cells. PLoS One (2010) 5:e9404. doi:10.1371/journal.pone.0009404
21. Jaitin DA, Kenigsberg E, Keren-Shaul H, Elefant N, Paul F, Zaretsky I, et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science (2014) 343:776–9. doi:10.1126/science.1247651
22. Pletinckx K, Stijlemans B, Pavlovic V, Laube R, Brandl C, Kneitz S, et al. Similar inflammatory DC maturation signatures induced by TNF or Try-panosoma brucei antigens instruct default Th2-cell responses. Eur J Immunol (2011) 41:3479–94. doi:10.1002/eji.201141631
23. Silva-Vilches C, Pletinckx K, Lohnert M, Pavlovic V, Ashour D, John V, et al. Low doses of cholera toxin and its mediator cAMP induce CTLA-2 secretion by dendritic cells to enhance regulatory T cell conversion. PLoS One (2017) 12:e0178114. doi:10.1371/journal.pone.0178114
24. Chaussabel D, Semnani RT, Mcdowell MA, Sacks D, Sher A, Nutman TB. Unique gene expression profiles of human macrophages and dendritic cells to phylogenetically distinct parasites. Blood (2003) 102:672–81. doi:10.1182/blood-2002-10-3232
25. Etna MP, Giacomini E, Pardini M, Severa M, Bottai D, Cruciani M, et al. Impact of Mycobacterium tuberculosis RD1-locus on human primary dendritic cell immune functions. Sci Rep (2015) 5:17078. doi:10.1038/srep17078
26. Troegeler A, Mercier I, Cougoule C, Pietretti D, Colom A, Duval C, et al. C-type lectin receptor DCIR modulates immunity to tuberculosis by sustaining type I interferon signaling in dendritic cells. Proc Natl Acad Sci U S A (2017) 114:E540–9. doi:10.1073/pnas.1613254114
27. Connor LM, Tang SC, Cognard E, Ochiai S, Hilligan KL, Old SI, et al. Th2 responses are primed by skin dendritic cells with distinct transcriptional profiles. J Exp Med (2017) 214:125–42. doi:10.1084/jem.20160470
28. Mayer JU, Demiri M, Agace WW, Macdonald AS, Svensson-Frej M, Milling SW. Different populations of CD11b+ dendritic cells drive Th2 responses in the small intestine and colon. Nat Commun (2017) 8:15820. doi:10.1038/ncomms15820
29. Gillessen S, Carvajal D, Ling P, Podlaski FJ, Stremlo DL, Familletti PC, et al. Mouse interleukin-12 (IL-12) p40 homodimer: a potent IL-12 antagonist. Eur J Immunol (1995) 25:200–6. doi:10.1002/eji.1830250133
30. Gately MK, Carvajal DM, Connaughton SE, Gillessen S, Warrier RR, Kolinsky KD, et al. Interleukin-12 antagonist activity of mouse interleukin-12 p40 homodimer in vitro and in vivo. Ann N Y Acad Sci (1996) 795:1–12. doi:10.1111/j.1749-6632.1996.tb52650.x
31. Mellor AL, Munn DH. IDO expression by dendritic cells: tolerance and tryptophan catabolism. Nat Rev Immunol (2004) 4:762–74. doi:10.1038/nri1457
32. Munn DH, Mellor AL. Indoleamine 2,3 dioxygenase and metabolic control of immune responses. Trends Immunol (2013) 34:137–43. doi:10.1016/j.it.2012.10.001
33. Fugier-Vivier I, Servet-Delprat C, Rivailler P, Rissoan MC, Liu YJ, Rabourdin-Combe C. Measles virus suppresses cell-mediated immunity by interfering with the survival and functions of dendritic and T cells. J Exp Med (1997) 186:813–23. doi:10.1084/jem.186.6.813
34. Grosjean I, Caux C, Bella C, Berger I, Wild F, Banchereau J, et al. Measles virus infects human dendritic cells and blocks their allostimulatory properties for CD4+ T cells. J Exp Med (1997) 186:801–12. doi:10.1084/jem.186.6.801
35. Schnorr JJ, Xanthakos S, Keikavoussi P, Kampgen E, Ter Meulen V, Schneider-Schaulies S. Induction of maturation of human blood dendritic cell precursors by measles virus is associated with immunosuppression. Proc Natl Acad Sci U S A (1997) 94:5326–31. doi:10.1073/pnas.94.10.5326
36. Réthi B, Gogolák P, Szatmari I, Veres A, Erdôs E, Nagy L, et al. SLAM/SLAM interactions inhibit CD40-induced production of inflammatory cytokines in monocyte-derived dendritic cells. Blood (2006) 107:2821–9. doi:10.1182/blood-2005-06-2265
37. Fekete T, Szabo A, Beltrame L, Vivar N, Pivarcsi A, Lanyi A, et al. Constraints for monocyte-derived dendritic cell functions under inflammatory conditions. Eur J Immunol (2012) 42:458–69. doi:10.1002/eji.201141924
38. Segerer SE, Muller N, Brandt J, Kapp M, Dietl J, Reichardt HM, et al. The glycoprotein-hormones activin A and inhibin A interfere with dendritic cell maturation. Reprod Biol Endocrinol (2008) 6:17. doi:10.1186/1477-7827-6-17
39. Huber S, Schramm C. Role of activin A in the induction of Foxp3+ and Foxp3- CD4+ regulatory T cells. Crit Rev Immunol (2011) 31:53–60. doi:10.1615/CritRevImmunol.v31.i1.50
40. Haschemi A, Wagner O, Marculescu R, Wegiel B, Robson SC, Gagliani N, et al. Cross-regulation of carbon monoxide and the adenosine A2a receptor in macrophages. J Immunol (2007) 178:5921–9. doi:10.4049/jimmunol.178.9.5921
41. Sudhakar C, Nagabhushana A, Jain N, Swarup G. NF-kappaB mediates tumor necrosis factor alpha-induced expression of optineurin, a negative regulator of NF-kappaB. PLoS One (2009) 4:e5114. doi:10.1371/journal.pone.0005114
42. Guo H, Cruz-Munoz ME, Wu N, Robbins M, Veillette A. Immune cell inhibition by SLAMF7 is mediated by a mechanism requiring src kinases, CD45, and SHIP-1 that is defective in multiple myeloma cells. Mol Cell Biol (2015) 35:41–51. doi:10.1128/MCB.01107-14
43. Crawford SE, Stellmach V, Murphy-Ullrich JE, Ribeiro SM, Lawler J, Hynes RO, et al. Thrombospondin-1 is a major activator of TGF-beta1 in vivo. Cell (1998) 93:1159–70. doi:10.1016/S0092-8674(00)81460-9
44. Doyen V, Rubio M, Braun D, Nakajima T, Abe J, Saito H, et al. Thrombospondin 1 is an autocrine negative regulator of human dendritic cell activation. J Exp Med (2003) 198:1277–83. doi:10.1084/jem.20030705
45. Gallucci S, Lolkema M, Matzinger P. Natural adjuvants: endogenous activators of dendritic cells. Nat Med (1999) 5:1249–55. doi:10.1038/15200
46. Fricke I, Mirza N, Dupont J, Lockhart C, Jackson A, Lee JH, et al. Vascular endothelial growth factor-trap overcomes defects in dendritic cell differentiation but does not improve antigen-specific immune responses. Clin Cancer Res (2007) 13:4840–8. doi:10.1158/1078-0432.CCR-07-0409
47. Oussa NA, Dahmani A, Gomis M, Richaud M, Andreev E, Navab-Daneshmand AR, et al. VEGF requires the receptor NRP-1 to inhibit lipopolysaccharide-dependent dendritic cell maturation. J Immunol (2016) 197:3927–35. doi:10.4049/jimmunol.1601116
48. Zhang W, Shou WD, Xu YJ, Bai WK, Hu B. Low-frequency ultrasound-induced VEGF suppression and synergy with dendritic cell-mediated anti-tumor immunity in murine prostate cancer cells in vitro. Sci Rep (2017) 7:5778. doi:10.1038/s41598-017-06242-8
49. Clark DA, Yu G, Levy GA, Gorczynski RM. Procoagulants in fetus rejection: the role of the OX-2 (CD200) tolerance signal. Semin Immunol (2001) 13:255–63. doi:10.1006/smim.2001.0315
50. Fallarino F, Asselin-Paturel C, Vacca C, Bianchi R, Gizzi S, Fioretti MC, et al. Murine plasmacytoid dendritic cells initiate the immunosuppressive pathway of tryptophan catabolism in response to CD200 receptor engagement. J Immunol (2004) 173:3748–54. doi:10.4049/jimmunol.173.6.3748
51. Vaine CA, Soberman RJ. The CD200-CD200R1 inhibitory signaling pathway: immune regulation and host-pathogen interactions. Adv Immunol (2014) 121:191–211. doi:10.1016/B978-0-12-800100-4.00005-2
52. Laffont S, Siddiqui KR, Powrie F. Intestinal inflammation abrogates the tolerogenic properties of MLN CD103+ dendritic cells. Eur J Immunol (2010) 40:1877–83. doi:10.1002/eji.200939957
53. Zhu B, Buttrick T, Bassil R, Zhu C, Olah M, Wu C, et al. IL-4 and retinoic acid synergistically induce regulatory dendritic cells expressing Aldh1a2. J Immunol (2013) 191:3139–51. doi:10.4049/jimmunol.1300329
54. Zinser E, Lechmann M, Golka A, Lutz MB, Steinkasserer A. Prevention and treatment of experimental autoimmune encephalomyelitis by soluble CD83. J Exp Med (2004) 200:345–51. doi:10.1084/jem.20030973
55. Lan Z, Lian D, Liu W, Arp J, Charlton B, Ge W, et al. Prevention of chronic renal allograft rejection by soluble CD83. Transplantation (2010) 90:1278–85. doi:10.1097/TP.0b013e318200005c
56. Bock F, Rossner S, Onderka J, Lechmann M, Pallotta MT, Fallarino F, et al. Topical application of soluble CD83 induces IDO-mediated immune modulation, increases Foxp3+ T cells, and prolongs allogeneic corneal graft survival. J Immunol (2013) 191:1965–75. doi:10.4049/jimmunol.1201531
57. Horvatinovich JM, Grogan EW, Norris M, Steinkasserer A, Lemos H, Mellor AL, et al. Soluble CD83 inhibits T cell activation by binding to the TLR4/MD-2 complex on CD14+ monocytes. J Immunol (2017) 198:2286–301. doi:10.4049/jimmunol.1600802
58. Cramer SO, Trumpfheller C, Mehlhoop U, More S, Fleischer B, Von Bonin A. Activation-induced expression of murine CD83 on T cells and identification of a specific CD83 ligand on murine B cells. Int Immunol (2000) 12:1347–51. doi:10.1093/intimm/12.9.1347
59. Everts B, Smits HH, Hokke CH, Yazdanbakhsh M. Helminths and dendritic cells: sensing and regulating via pattern recognition receptors, Th2 and Treg responses. Eur J Immunol (2010) 40:1525–37. doi:10.1002/eji.200940109
60. Maizels RM, McSorley HJ. Regulation of the host immune system by helminth parasites. J Allergy Clin Immunol (2016) 138:666–75. doi:10.1016/j.jaci.2016.07.007
61. Vendelova E, Camargo De Lima J, Lorenzatto KR, Monteiro KM, Mueller T, Veepaschit J, et al. Proteomic analysis of excretory-secretory products of Mesocestoides corti metacestodes reveals potential suppressors of dendritic cell functions. PLoS Negl Trop Dis (2016) 10:e0005061. doi:10.1371/journal.pntd.0005061
62. Zhou G, Stevenson MM, Geary TG, Xia J. Comprehensive transcriptome meta-analysis to characterize host immune responses in helminth infections. PLoS Negl Trop Dis (2016) 10:e0004624. doi:10.1371/journal.pntd.0004624
63. Lutz MB. Induction of CD4(+) regulatory and polarized effector/helper T cells by dendritic cells. Immune Netw (2016) 16:13–25. doi:10.4110/in.2016.16.1.13
64. MacDonald AS, Maizels RM. Alarming dendritic cells for Th2 induction. J Exp Med (2008) 205:13–7. doi:10.1084/jem.20072665
65. Shih VF, Davis-Turak J, Macal M, Huang JQ, Ponomarenko J, Kearns JD, et al. Control of RelB during dendritic cell activation integrates canonical and noncanonical NF-kappaB pathways. Nat Immunol (2012) 13:1162–70. doi:10.1038/ni.2446
66. Manches O, Fernandez MV, Plumas J, Chaperot L, Bhardwaj N. Activation of the noncanonical NF-kappaB pathway by HIV controls a dendritic cell immunoregulatory phenotype. Proc Natl Acad Sci U S A (2012) 109:14122–7. doi:10.1073/pnas.1204032109
67. Kim NS, Mbongue JC, Nicholas DA, Esebanmen GE, Unternaehrer JJ, Firek AF, et al. Chimeric vaccine stimulation of human dendritic cell indoleamine 2, 3-dioxygenase occurs via the non-canonical NF-kappaB pathway. PLoS One (2016) 11:e0147509. doi:10.1371/journal.pone.0147509
68. Sun SC. The non-canonical NF-kappaB pathway in immunity and inflammation. Nat Rev Immunol (2017) 17:545–58. doi:10.1038/nri.2017.52
69. Thomas PG, Carter MR, Da’dara AA, Desimone TM, Harn DA. A helminth glycan induces APC maturation via alternative NF-kappa B activation independent of I kappa B alpha degradation. J Immunol (2005) 175:2082–90. doi:10.4049/jimmunol.175.4.2082
70. Baetz A, Frey M, Heeg K, Dalpke AH. Suppressor of cytokine signaling (SOCS) proteins indirectly regulate toll-like receptor signaling in innate immune cells. J Biol Chem (2004) 279:54708–15. doi:10.1074/jbc.M410992200
71. Yoshimura A, Naka T, Kubo M. SOCS proteins, cytokine signalling and immune regulation. Nat Rev Immunol (2007) 7:454–65. doi:10.1038/nri2093
72. Seki Y, Inoue H, Nagata N, Hayashi K, Fukuyama S, Matsumoto K, et al. SOCS-3 regulates onset and maintenance of T(H)2-mediated allergic responses. Nat Med (2003) 9:1047–54. doi:10.1038/nm896
73. Li Y, Chu N, Rostami A, Zhang GX. Dendritic cells transduced with SOCS-3 exhibit a tolerogenic/DC2 phenotype that directs type 2 Th cell differentiation in vitro and in vivo. J Immunol (2006) 177:1679–88. doi:10.4049/jimmunol.177.3.1679
74. Knosp CA, Carroll HP, Elliott J, Saunders SP, Nel HJ, Amu S, et al. SOCS2 regulates T helper type 2 differentiation and the generation of type 2 allergic responses. J Exp Med (2011) 208:1523–31. doi:10.1084/jem.20101167
75. Pot C, Apetoh L, Awasthi A, Kuchroo VK. Molecular pathways in the induction of interleukin-27-driven regulatory type 1 cells. J Interferon Cytokine Res (2010) 30:381–8. doi:10.1089/jir.2010.0047
76. Keir ME, Butte MJ, Freeman GJ, Sharpe AH. PD-1 and its ligands in tolerance and immunity. Annu Rev Immunol (2008) 26:677–704. doi:10.1146/annurev.immunol.26.021607.090331
77. Sharpe AH, Pauken KE. The diverse functions of the PD1 inhibitory pathway. Nat Rev Immunol (2017). doi:10.1038/nri.2017.108
78. Smith P, Walsh CM, Mangan NE, Fallon RE, Sayers JR, Mckenzie AN, et al. Schistosoma mansoni worms induce anergy of T cells via selective up-regulation of programmed death ligand 1 on macrophages. J Immunol (2004) 173:1240–8. doi:10.4049/jimmunol.173.2.1240
79. Reyes JL, Lopes F, Leung G, Mancini NL, Matisz CE, Wang A, et al. Treatment with cestode parasite antigens results in recruitment of CCR2+ myeloid cells, the adoptive transfer of which ameliorates colitis. Infect Immun (2016) 84:3471–83. doi:10.1128/IAI.00681-16
80. Stempin CC, Motran CC, Aoki MP, Falcon CR, Cerban FM, Cervi L. PD-L2 negatively regulates Th1-mediated immunopathology during Fasciola hepatica infection. Oncotarget (2016) 7:77721–31. doi:10.18632/oncotarget.12790
81. Goldberg MF, Saini NK, Porcelli SA. Evasion of innate and adaptive immunity by Mycobacterium tuberculosis. Microbiol Spectr (2014) 2:1–24. doi:10.1128/microbiolspec.MGM2-0005-2013
82. Ristich V, Liang S, Zhang W, Wu J, Horuzsko A. Tolerization of dendritic cells by HLA-G. Eur J Immunol (2005) 35:1133–42. doi:10.1002/eji.200425741
83. Granucci F, Vizzardelli C, Pavelka N, Feau S, Persico M, Virzi E, et al. Inducible IL-2 production by dendritic cells revealed by global gene expression analysis. Nat Immunol (2001) 2:882–8. doi:10.1038/ni0901-882
84. Granucci F, Vizzardelli C, Virzi E, Rescigno M, Ricciardi-Castagnoli P. Transcriptional reprogramming of dendritic cells by differentiation stimuli. Eur J Immunol (2001) 31:2539–46. doi:10.1002/1521-4141(200109)31:9<2539::AID-IMMU2539>3.0.CO;2-9
85. Cavalieri D, Rivero D, Beltrame L, Buschow SI, Calura E, Rizzetto L, et al. DC-ATLAS: a systems biology resource to dissect receptor specific signal transduction in dendritic cells. Immunome Res (2010) 6:10. doi:10.1186/1745-7580-6-10
86. Theiner G, Rößner S, Dalpke A, Bode K, Berger T, Gessner A, et al. TLR9 cooperates with TLR4 to increase IL-12 release by murine dendritic cells. Mol Immunol (2008) 45:244–52. doi:10.1016/j.molimm.2007.02.021
87. Menges M, Rossner S, Voigtlander C, Schindler H, Kukutsch NA, Bogdan C, et al. Repetitive injections of dendritic cells matured with tumor necrosis factor alpha induce antigen-specific protection of mice from autoimmunity. J Exp Med (2002) 195:15–21. doi:10.1084/jem.20011341
88. Fulcher JA, Hashimi ST, Levroney EL, Pang M, Gurney KB, Baum LG, et al. Galectin-1-matured human monocyte-derived dendritic cells have enhanced migration through extracellular matrix. J Immunol (2006) 177:216–26. doi:10.4049/jimmunol.177.1.216
89. Napolitani G, Rinaldi A, Bertoni F, Sallusto F, Lanzavecchia A. Selected toll-like receptor agonist combinations synergistically trigger a T helper type 1–polarizing program in dendritic cells. Nat Immunol (2005) 6:769–76. doi:10.1038/ni1223
90. Hamasaki T, Uto T, Akagi T, Akashi M, Baba M. Modulation of gene expression related to toll-like receptor signaling in dendritic cells by poly(gamma-glutamic acid) nanoparticles. Clin Vaccine Immunol (2010) 17:748–56. doi:10.1128/CVI.00505-09
91. Ronchetti S, Nocentini G, Petrillo MG, Bianchini R, Sportoletti P, Bastianelli A, et al. Glucocorticoid-Induced TNFR family related gene (GITR) enhances dendritic cell activity. Immunol Lett (2011) 135:24–33. doi:10.1016/j.imlet.2010.09.008
92. Manetti R, Parronchi P, Giudizi MG, Piccinni MP, Maggi E, Trinchieri G, et al. Natural killer cell stimulatory factor (interleukin 12 [IL-12]) induces T helper type 1 (Th1)-specific immune responses and inhibits the development of IL-4-producing Th cells. J Exp Med (1993) 177:1199–204. doi:10.1084/jem.177.4.1199
93. Heufler C, Koch F, Stanzl U, Topar G, Wysocka M, Trinchieri G, et al. Interleukin-12 is produced by dendritic cells and mediates T helper 1 development as well as interferon-gamma production by T helper 1 cells. Eur J Immunol (1996) 26:659–68. doi:10.1002/eji.1830260323
94. Bettelli E, Carrier Y, Gao W, Korn T, Strom TB, Oukka M, et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature (2006) 441:235–8. doi:10.1038/nature04753
95. Goriely S, Neurath MF, Goldman M. How microorganisms tip the balance between interleukin-12 family members. Nat Rev Immunol (2008) 8:81–6. doi:10.1038/nri2225
96. Kato K, Shimozato O, Hoshi K, Wakimoto H, Hamada H, Yagita H, et al. Local production of the p40 subunit of interleukin 12 suppresses T-helper 1-mediated immune responses and prevents allogeneic myoblast rejection. Proc Natl Acad Sci U S A (1996) 93:9085–9. doi:10.1073/pnas.93.17.9085
97. Rea IM, Mcmaster D, Woodside JV, Young IS, Archbold GP, Linton T, et al. Community-living nonagenarians in northern ireland have lower plasma homocysteine but similar methylenetetrahydrofolate reductase thermolabile genotype prevalence compared to 70-89-year-old subjects. Atherosclerosis (2000) 149:207–14. doi:10.1016/S0021-9150(99)00417-7
98. Millet P, Mccall C, Yoza B. RelB: an outlier in leukocyte biology. J Leukoc Biol (2013) 94:941–51. doi:10.1189/jlb.0513305
99. Grumont R, Hochrein H, O’keeffe M, Gugasyan R, White C, Caminschi I, et al. c-Rel regulates interleukin 12 p70 expression in CD8(+) dendritic cells by specifically inducing p35 gene transcription. J Exp Med (2001) 194:1021–32. doi:10.1084/jem.194.8.1021
100. Weih F, Carrasco D, Durham SK, Barton DS, Rizzo CA, Ryseck RP, et al. Multiorgan inflammation and hematopoietic abnormalities in mice with a targeted disruption of RelB, a member of the NF-kappa B/Rel family. Cell (1995) 80:331–40. doi:10.1016/0092-8674(95)90416-6
101. Wu L, D’amico A, Winkel KD, Suter M, Lo D, Shortman K. RelB is essential for the development of myeloid-related CD8alpha- dendritic cells but not of lymphoid-related CD8alpha+ dendritic cells. Immunity (1998) 9:839–47. doi:10.1016/S1074-7613(00)80649-4
102. Briseno CG, Gargaro M, Durai V, Davidson JT, Theisen DJ, Anderson DA III, et al. Deficiency of transcription factor RelB perturbs myeloid and DC development by hematopoietic-extrinsic mechanisms. Proc Natl Acad Sci U S A (2017) 114:3957–62. doi:10.1073/pnas.1619863114
103. Kretschmer K, Apostolou I, Hawiger D, Khazaie K, Nussenzweig MC, Von Boehmer H. Inducing and expanding regulatory T cell populations by foreign antigen. Nat Immunol (2005) 6:1219–27. doi:10.1038/ni1265
104. Griffith JW, Sokol CL, Luster AD. Chemokines and chemokine receptors: positioning cells for host defense and immunity. Annu Rev Immunol (2014) 32:659–702. doi:10.1146/annurev-immunol-032713-120145
105. Yaal-Hahoshen N, Shina S, Leider-Trejo L, Barnea I, Shabtai EL, Azenshtein E, et al. The chemokine CCL5 as a potential prognostic factor predicting disease progression in stage II breast cancer patients. Clin Cancer Res (2006) 12:4474–80. doi:10.1158/1078-0432.CCR-06-0074
106. Soria G, Yaal-Hahoshen N, Azenshtein E, Shina S, Leider-Trejo L, Ryvo L, et al. Concomitant expression of the chemokines RANTES and MCP-1 in human breast cancer: a basis for tumor-promoting interactions. Cytokine (2008) 44:191–200. doi:10.1016/j.cyto.2008.08.002
107. Ban Y, Mai J, Li X, Mitchell-Flack M, Zhang T, Zhang L, et al. Targeting autocrine CCL5-CCR5 axis reprograms immunosuppressive myeloid cells and reinvigorates antitumor immunity. Cancer Res (2017) 77:2857–68. doi:10.1158/0008-5472.CAN-16-2913
108. Zhang Y, Lv D, Kim HJ, Kurt RA, Bu W, Li Y, et al. A novel role of hematopoietic CCL5 in promoting triple-negative mammary tumor progression by regulating generation of myeloid-derived suppressor cells. Cell Res (2013) 23:394–408. doi:10.1038/cr.2012.178
109. Faunce DE, Stein-Streilein J. NKT cell-derived RANTES recruits APCs and CD8+ T cells to the spleen during the generation of regulatory T cells in tolerance. J Immunol (2002) 169:31–8. doi:10.4049/jimmunol.169.1.31
110. Re F, Strominger JL. IL-10 released by concomitant TLR2 stimulation blocks the induction of a subset of Th1 cytokines that are specifically induced by TLR4 or TLR3 in human dendritic cells. J Immunol (2004) 173:7548–55. doi:10.4049/jimmunol.173.12.7548
111. Torres D, Kohler A, Delbauve S, Caminschi I, Lahoud MH, Shortman K, et al. IL-12p40/IL-10 producing preCD8alpha/Clec9A+ dendritic cells are induced in neonates upon Listeria monocytogenes infection. PLoS Pathog (2016) 12:e1005561. doi:10.1371/journal.ppat.1005561
112. Fiorentino DF, Zlotnik A, Vieira P, Mosmann TR, Howard M, Moore KW, et al. IL-10 acts on the antigen-presenting cell to inhibit cytokine production by Th1 cells. J Immunol (1991) 146:3444–51.
113. Macatonia SE, Doherty TM, Knight SC, O’garra A. Differential effect of IL-10 on dendritic cell-induced T cell proliferation and IFN-gamma production. J Immunol (1993) 150:3755–65.
114. O’Garra A, Vieira P. T(H)1 cells control themselves by producing interleukin-10. Nat Rev Immunol (2007) 7:425–8. doi:10.1038/nri2097
115. Redford PS, Murray PJ, O’garra A. The role of IL-10 in immune regulation during M. tuberculosis infection. Mucosal Immunol (2011) 4:261–70. doi:10.1038/mi.2011.7
116. Moreira-Teixeira L, Redford PS, Stavropoulos E, Ghilardi N, Maynard CL, Weaver CT, et al. Cell-derived IL-10 impairs host resistance to Mycobacterium tuberculosis infection. J Immunol (2017) 199:613–23. doi:10.4049/jimmunol.1601340
117. Allavena P, Piemonti L, Longoni D, Bernasconi S, Stoppacciaro A, Ruco L, et al. IL-10 prevents the differentiation of monocytes to dendritic cells but promotes their maturation to macrophages. Eur J Immunol (1998) 28:359–69. doi:10.1002/(SICI)1521-4141(199801)28:01<359::AID-IMMU359>3.0.CO;2-4
118. Waibler Z, Anzaghe M, Konur A, Akira S, Müller W, Kalinke U. Excessive CpG 1668 stimulation triggers IL-10 production by cDC that inhibits IFN-alpha responses by pDC. Eur J Immunol (2008) 38:3127–37. doi:10.1002/eji.200838184
119. Levings MK. Differentiation of Tr1 cells by immature dendritic cells requires IL-10 but not CD25+CD4+ Tr cells. Blood (2004) 105:1162–9. doi:10.1182/blood-2004-03-1211
120. Akbari O, Dekruyff RH, Umetsu DT. Pulmonary dendritic cells producing IL-10 mediate tolerance induced by respiratory exposure to antigen. Nat Immunol (2001) 2:725–31. doi:10.1038/90667
121. Gabrysova L, Howes A, Saraiva M, O’garra A. The regulation of IL-10 expression. Curr Top Microbiol Immunol (2014) 380:157–90. doi:10.1007/978-3-662-43492-5_8
122. Pletinckx K, Dohler A, Pavlovic V, Lutz MB. Role of dendritic cell maturity/costimulation for generation, homeostasis, and suppressive activity of regulatory T cells. Front Immunol (2011) 2:39. doi:10.3389/fimmu.2011.00039
123. Huehn J, Beyer M. Epigenetic and transcriptional control of Foxp3+ regulatory T cells. Semin Immunol (2015) 27:10–8. doi:10.1016/j.smim.2015.02.002
124. Yamaguchi Y. Developmental regulation by cytokines of bone marrow-derived dendritic cells and epidermal Langerhans cells. Microbiol Immunol (1998) 42:639–50. doi:10.1111/j.1348-0421.1998.tb02334.x
125. Sakai F, Hosoya T, Ono-Ohmachi A, Ukibe K, Ogawa A, Moriya T, et al. Lactobacillus gasseri SBT2055 induces TGF-beta expression in dendritic cells and activates TLR2 signal to produce IgA in the small intestine. PLoS One (2014) 9:e105370. doi:10.1371/journal.pone.0105370
126. Shi M, Zhu J, Wang R, Chen X, Mi L, Walz T, et al. Latent TGF-β structure and activation. Nature (2011) 474:343–9. doi:10.1038/nature10152
127. Munger JS, Huang X, Kawakatsu H, Griffiths MJ, Dalton SL, Wu J, et al. The integrin alpha v beta 6 binds and activates latent TGF beta 1: a mechanism for regulating pulmonary inflammation and fibrosis. Cell (1999) 96:319–28. doi:10.1016/S0092-8674(00)80545-0
128. Mu D, Cambier S, Fjellbirkeland L, Baron JL, Munger JS, Kawakatsu H, et al. The integrin alpha(v)beta8 mediates epithelial homeostasis through MT1-MMP-dependent activation of TGF-beta1. J Cell Biol (2002) 157:493–507. doi:10.1083/jcb.200109100
129. Yu Q, Stamenkovic I. Cell surface-localized matrix metalloproteinase-9 proteolytically activates TGF-beta and promotes tumor invasion and angiogenesis. Genes Dev (2000) 14:163–76.
130. Travis MA, Reizis B, Melton AC, Masteller E, Tang Q, Proctor JM, et al. Loss of integrin alpha(v)beta8 on dendritic cells causes autoimmunity and colitis in mice. Nature (2007) 449:361–5. doi:10.1038/nature06110
131. Mavaddat N, Mason DW, Atkinson PD, Evans EJ, Gilbert RJ, Stuart DI, et al. Signaling lymphocytic activation molecule (CDw150) is homophilic but self-associates with very low affinity. J Biol Chem (2000) 275:28100–9. doi:10.1074/jbc.M004117200
132. Sayos J, Wu C, Morra M, Wang N, Zhang X, Allen D, et al. The X-linked lymphoproliferative-disease gene product SAP regulates signals induced through the co-receptor SLAM. Nature (1998) 395:462–9. doi:10.1038/26683
133. Sumegi J, Huang D, Lanyi A, Davis JD, Seemayer TA, Maeda A, et al. Correlation of mutations of the SH2D1A gene and Epstein-Barr virus infection with clinical phenotype and outcome in X-linked lymphoproliferative disease. Blood (2000) 96:3118–25.
134. Grohmann U, Mondanelli G, Belladonna ML, Orabona C, Pallotta MT, Iacono A, et al. Amino-acid sensing and degrading pathways in immune regulation. Cytokine Growth Factor Rev (2017) 35:37–45. doi:10.1016/j.cytogfr.2017.05.004
135. Pallotta MT, Orabona C, Volpi C, Vacca C, Belladonna ML, Bianchi R, et al. Indoleamine 2,3-dioxygenase is a signaling protein in long-term tolerance by dendritic cells. Nat Immunol (2011) 12:870–8. doi:10.1038/ni.2077
136. Bessede A, Gargaro M, Pallotta MT, Matino D, Servillo G, Brunacci C, et al. Aryl hydrocarbon receptor control of a disease tolerance defence pathway. Nature (2014) 511:184–90. doi:10.1038/nature13323
137. Seeger P, Musso T, Sozzani S. The TGF-beta superfamily in dendritic cell biology. Cytokine Growth Factor Rev (2015) 26:647–57. doi:10.1016/j.cytogfr.2015.06.002
138. Pflanz S, Timans JC, Cheung J, Rosales R, Kanzler H, Gilbert J, et al. IL-27, a heterodimeric cytokine composed of EBI3 and p28 protein, induces proliferation of naive CD4+ T cells. Immunity (2002) 16:779–90. doi:10.1016/S1074-7613(02)00324-2
139. Takeda A, Hamano S, Yamanaka A, Hanada T, Ishibashi T, Mak TW, et al. Cutting edge: role of IL-27/WSX-1 signaling for induction of T-bet through activation of STAT1 during initial Th1 commitment. J Immunol (2003) 170:4886–90.
140. Yoshida H, Nakaya M, Miyazaki Y. Interleukin 27: a double-edged sword for offense and defense. J Leukoc Biol (2009) 86:1295–303. doi:10.1189/jlb.0609445
141. Holscher C, Holscher A, Ruckerl D, Yoshimoto T, Yoshida H, Mak T, et al. The IL-27 receptor chain WSX-1 differentially regulates antibacterial immunity and survival during experimental tuberculosis. J Immunol (2005) 174:3534–44. doi:10.4049/jimmunol.174.6.3534
142. Posselt G, Schwarz H, Duschl A, Horejs-Hoeck J. Suppressor of cytokine signaling 2 is a feedback inhibitor of TLR-induced activation in human monocyte-derived dendritic cells. J Immunol (2011) 187:2875–84. doi:10.4049/jimmunol.1003348
143. Machado FS, Johndrow JE, Esper L, Dias A, Bafica A, Serhan CN, et al. Anti-inflammatory actions of lipoxin A4 and aspirin-triggered lipoxin are SOCS-2 dependent. Nat Med (2006) 12:330–4. doi:10.1038/nm1355
144. Curiel TJ, Wei S, Dong H, Alvarez X, Cheng P, Mottram P, et al. Blockade of B7-H1 improves myeloid dendritic cell-mediated antitumor immunity. Nat Med (2003) 9:562–7. doi:10.1038/nm863
145. Zou W, Wolchok JD, Chen L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci Transl Med (2016) 8:328rv324. doi:10.1126/scitranslmed.aad7118
146. Papalexi E, Satija R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat Rev Immunol (2017) 18:35–45. doi:10.1038/nri.2017.76
147. Tanay A, Regev A. Scaling single-cell genomics from phenomenology to mechanism. Nature (2017) 541:331–8. doi:10.1038/nature21350
148. Picelli S, Bjorklund AK, Faridani OR, Sagasser S, Winberg G, Sandberg R. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat Methods (2013) 10:1096–8. doi:10.1038/nmeth.2639
149. Shalek AK, Satija R, Adiconis X, Gertner RS, Gaublomme JT, Raychowdhury R, et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature (2013) 498:236–40. doi:10.1038/nature12172
150. Shalek AK, Satija R, Shuga J, Trombetta JJ, Gennert D, Lu D, et al. Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature (2014) 510:363–9. doi:10.1038/nature13437
Keywords: tolerogenic dendritic cells, transcriptional profiling, steady-state dendritic cells, bacteria, mycobacteria, helminths, immune evasion
Citation: Vendelova E, Ashour D, Blank P, Erhard F, Saliba A-E, Kalinke U and Lutz MB (2018) Tolerogenic Transcriptional Signatures of Steady-State and Pathogen-Induced Dendritic Cells. Front. Immunol. 9:333. doi: 10.3389/fimmu.2018.00333
Received: 24 October 2017; Accepted: 06 February 2018;
Published: 28 February 2018
Edited by:
Catharien Hilkens, Newcastle University, United KingdomReviewed by:
Eva Martinez Caceres, Universitat Autònoma de Barcelona, SpainNathalie Cools, University of Antwerp, Belgium
Copyright: © 2018 Vendelova, Ashour, Blank, Erhard, Saliba, Kalinke and Lutz. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Manfred B. Lutz, m.lutz@vim.uni-wuerzburg.de
 Emilia Vendelova1
Emilia Vendelova1 Diyaaeldin Ashour
Diyaaeldin Ashour Patrick Blank
Patrick Blank Florian Erhard
Florian Erhard Ulrich Kalinke
Ulrich Kalinke Manfred B. Lutz
Manfred B. Lutz