- 1Coastal Disease Ecology Lab, Smithsonian Environmental Research Center, Edgewater, MD, United States
- 2Department of Biology, Frostburg State University, Frostburg, MD, United States
- 3Department of Earth and Planetary Sciences, John Hopkins University, Baltimore, MD, United States
- 4Department of Biological Sciences, University of the Pacific, Stockton, CA, United States
Introduction: The North American river otter (Lontra canadensis) occurs from freshwater to marine habitats across North America, yet very little information exists regarding the biology and ecology of river otters residing in the largest estuary in the United States, the Chesapeake Bay.
Methods: We characterized latrines and collected scat to non-invasively examine the diet of river otters residing along 12km of the shoreline of the Rhode River, a tidally influenced subestuary of the Chesapeake Bay.
Results: We found and assessed characteristics of 18 latrines, which were compared to habitat models of characteristics of latrines in riverine habitats. Scat was examined microscopically and genetically to assess consumed prey and parasites potentially infecting these river otters or their prey. Our results indicate high similarity of latrine characteristics in both tidally influenced and riverine habitats. Additionally, coastal otters appear to readily use manmade structures as latrines. Our results also indicate that river otters consume a wide range of terrestrial and aquatic fauna, primarily consisting of finfish and crustaceans, but also including frogs and ducks. Finally, multiple parasite species were identified, including parasites of river otters and those infecting prey, indicating that parasites likely play an important role in both prey availability and otter health.
Discussion: This study is the first to characterize river otter latrines and diet in a tidally influenced estuarine habitat within the Chesapeake Bay, a critical step in expanding the ecological research of this apex predator across the increasingly urbanized Chesapeake Bay.
Introduction
North American river otters (hereafter river otter; Lontra canadensis) are semi-aquatic mammals that were historically distributed across the North American continent (Larivière and Walton, 1998) and that reside in and rely upon aquatic and coastal-marine systems (Reid et al., 1994; Brooks and Serfass, 2019). Though previously thought to require pristine aquatic habitat, river otter populations are expanding into urbanized and degraded estuaries (Bouley et al., 2015; Carroll et al., 2020; Stubbs et al., 2020). One such example is the apparent spread of river otters in the Chesapeake Bay, particularly in highly urbanized areas (Grablick, 2021; Cape Gazette, 2022; Bay Bulletin, 2023). While prior studies recognized the Chesapeake Bay as important river otter habitat (Mowbray et al., 1976, 1979), no published studies have examined the ecology or biology of river otters residing in this area, including systematic examination of latrines or diet.
Though often considered elusive due to their nocturnal behavior, river otter activity can be monitored non-invasively at areas where river otters congregate, which are known as latrines. River otter latrines are primarily used for scent-marking through defecation, urination, and anal gland secretions (Greer, 1955; Melquist and Hornocker, 1983) though river otters also use these locations to eat, play, socialize, and rest (Melquist and Hornocker, 1983). Active latrines are often littered with scat and can be identified by a fishy odor, tracks, and other signs of habitat manipulation (Mowbray et al., 1979). As river otter latrines are often persistent landscape features that may remain active for decades (Greer, 1955), finding these latrines in a landscape can provide an opportunity for long-term, non-invasive monitoring and research of resident river otters.
To date, most research assessing the habitat characteristics of river otter latrines examined inland riverine systems in the United States or coastal marine and estuarine environments along the Pacific coast of North America (Mowbray et al., 1976; Swimley et al., 1998; Gaydos et al., 2007a; Cote et al., 2008; Barocas et al., 2016). Predictive habitat models indicated river otter latrines are characterized by steep shores, downed logs, beaver activity, and mature trees across traditional riverine systems (Swimley et al., 1998), arid riverine systems (Depue and Ben-David, 2010), wetlands (Newman and Griffin, 1994), and coastal marine systems (Gaydos et al., 2007a; Cote et al., 2008; Barocas et al., 2016). Though previous papers noted evidence consistent with latrines in the Maryland portion of the Chesapeake Bay (Mowbray et al., 1976, 1979), this study is the first to provide detailed habitat characterization of river otter latrines within this estuary.
Previous studies reported that river otter diets consist mainly of teleost fish, crayfish, and aquatic invertebrates, and occasionally include amphibians, small mammals, and birds (Greer, 1955; Speich and Pitman, 1984). River otter diets vary seasonally and spatially, likely due to prey availability (Serfass et al., 1990; Cote et al., 2008). Traditionally, river otter diets were assessed via microscopic examination of scat collected from river otter latrines (Cote et al., 2008; Stearns and Serfass, 2011) or microscopic examination of stomach contents from carcasses (Sanders et al., 2023). However, there are several major challenges with microscopic analysis of prey in scats or stomachs including, 1) the high level of taxonomic expertise required to identify the wide range of taxa consumed, 2) the difficulty of visually identifying degraded contents, and 3) the time-consuming nature of sifting through and attempting to identify degraded contents. Alternatively, new genetic methods for assessing dietary DNA (dDNA; de Sousa et al., 2019), such as amplicon-based high throughput sequencing or metabarcoding, can often provide higher taxonomic resolution for a wide-range of taxa compared to morphological approaches. For example, Quéméré et al. (2021) used a metabarcoding approach on scat to assess the diet diversity of the giant otter (Pteronura brasiliensis) and was able to identify >90% of prey to the species level, providing unparalleled resolution in the diet of this elusive predator. Similarly, Drake et al. (2023) compared microscopic and metabarcode methods to examine the diet of the Eurasian otter (Lutra lutra) and found that the genetic approach provided greater taxonomic resolution and broader depth. A few studies have utilized a metabarcode approach to examine diets of river otters, including from stomach contents (Heenkenda et al., 2024; Sanders et al., 2023) and fecal swabs from carcasses (Bakker et al., 2024). These studies illustrate the wide breadth and taxonomic scope that genetic methods can provide for diet studies and provide additional evidence for the broad dietary habits of river otters. In this study, we use a metabarcoding approach to examine the diet of wild, free-living otters through the collection and processing of scat from wild river otter latrines, providing a completely non-invasive approach to further study the ecology of river otters.
In addition to providing information about prey, a community-based genetic approach such as metabarcoding can also provide information about broader-scale trophic interactions, such as the parasites infecting the prey or those that infect river otters through parasite consumption. As apex aquatic predators, the diet of river otters may provide information about the diversity of available prey and parasites within aquatic habitats (Fleming et al., 1977; Kimber and Kollias, 2000; Ben-David et al., 2001). Additionally, as river otters increasingly reside in urban areas, they are more frequently exposed to several zoonotic parasites, such as Cryptosporidium spp. and Giardia spp (Kimber and Kollias, 2000; Gaydos et al., 2007b). Thus, river otter diets can provide information about the role of parasites in trophic linkages and the presence of parasites that present potential public health risks.
In this study, we aimed to characterize coastal river otter latrines and examine the diet of river otters residing at the Smithsonian Environmental Research Center (SERC), which is located on the Rhode River, a subestuary of the Chesapeake Bay. Our objectives were to 1) identify key habitat variables of latrines at SERC and compare to prior latrine models, 2) compare microscopic and genetic methods to identify prey from scat collected at latrines, and 3) identify parasites of prey and river otters from scat collected from these latrines. To our knowledge, this is the first study to characterize the latrines and diet of river otters residing in a tidally influenced estuarine habitat within the Chesapeake Bay, providing a foundation for expanding ecological research on river otters across the entire estuary.
Methods
Study site description
The Smithsonian Environmental Research Center (SERC) is a 1,072 hectare property in eastern Maryland situated on the Rhode River subestuary within the Chesapeake Bay. The property has 22 km of shoreline along tidally influenced aquatic habitats, with salinity across these habitats varying spatially and seasonally. We conducted three transects (Smithsonian: SERC, Corn Island: CI, Global Change Research Wetland: GCREW) on foot along 12 km of shoreline (the walkable component of the SERC shoreline) a minimum of three times to identify river otter latrines (Figure 1). All transects were located within 3 linear km of each other along tidally-influenced estuarine waters of low salinity. All three transects contained mixed-hardwood forests with varying degrees of understory density and shoreline that varied from open water to dense marsh throughout. CI and GCREW transects included patches of dense mountain laurel canopy with mossy shaded understory. Transects at CI and SERC included tidal and permanent beaches while GCREW contained several wide tidal mud flats.
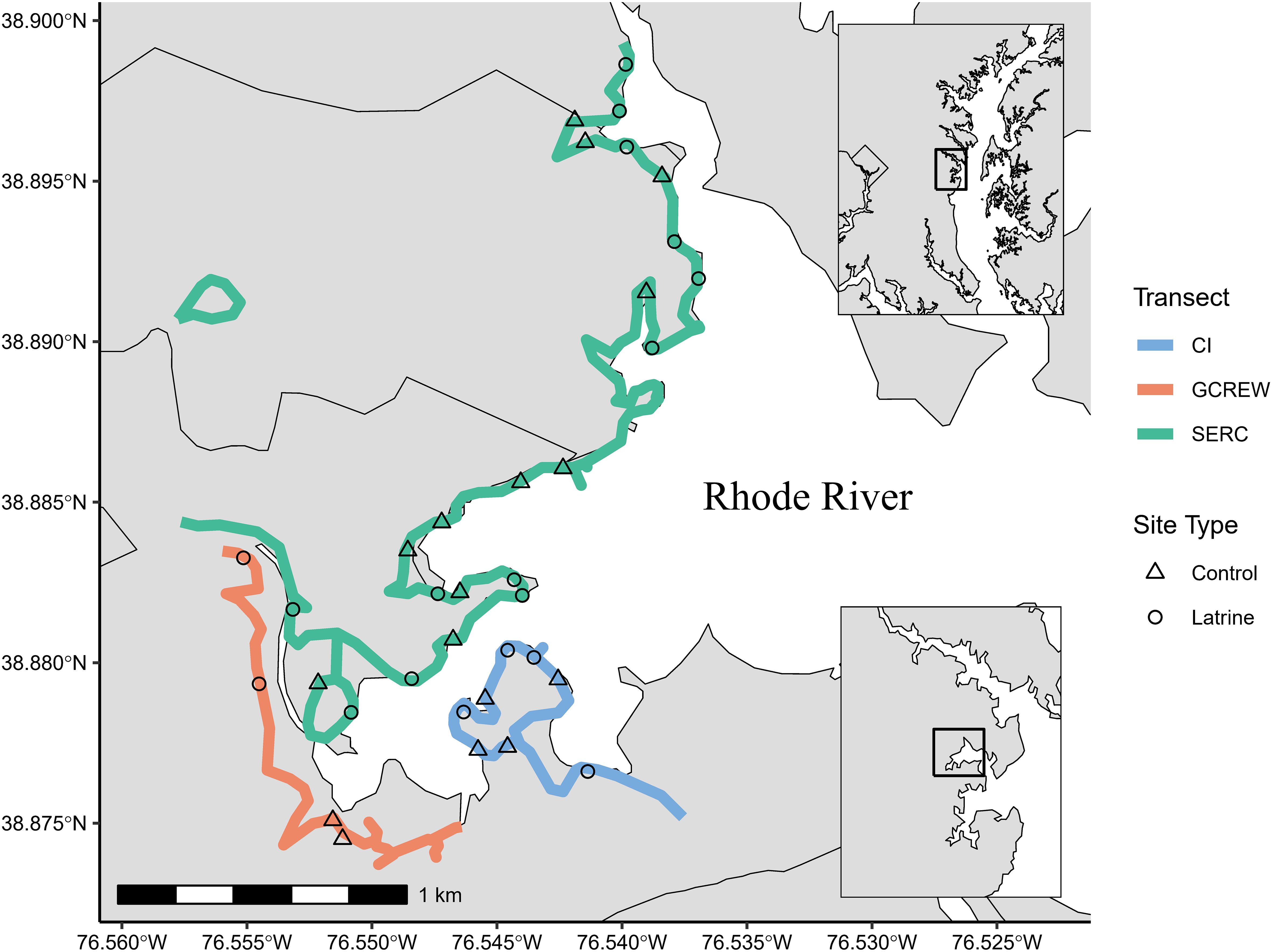
Figure 1. Shoreline surveyed for river otter activity at the Smithsonian Environmental Research Center along the Rhode River subestuary of the Chesapeake Bay. Active river otter latrines (n=18) and non-latrine control points (n=17) used for analysis are shown along three transects at the main Smithsonian Environmental Research Center (SERC), Corn Island (CI), and Global Change Research Wetland (GCREW). Bottom inset map indicates position within the broader subestuary, and top inset shows study area position within the Chesapeake Bay. Map was created using mapping files provided by the Chesapeake Bay Program (2016).
Latrines were defined by the recurrent presence of river otter scat, which was identified through its size, shape, content, and fishy odor, as well as river otter tracks and scrapes. Due to availability of researchers, our surveys occurred primarily between May and August of 2021. River otter tracks, scrapes, and scat could be obscured by tides, heavy summer rains, wind, falling leaves, and intentional clearing of recreationally utilized boardwalks and walkways, so a maximum of one river otter visit was recorded at each latrine with new activity between each observer visit. River otters probably revisited latrines multiple times between observations and we surveyed in summer when marking is generally lowest (Olson et al., 2008; Serfass et al., 2019, 2020), so we likely underestimated river otter visits and the number of latrines in the study area. River otters occasionally mark in non-latrine areas, so we established criteria to define active latrines for the purpose of analysis. We define active latrines as those with 1) at least 2 scats deposited over multiple visits during the study period, and 2) deposited scat was concentrated in a focal area. Additionally, we required that most environmental features were measurable for an active latrine to be included in statistical analyses. We used transect-specific latrine site densities and the average distance between the water and latrine focal point to establish randomized control points across all three transects. Control sites were replaced if river otter activity was found within 100 m of the focal point at any time during the study period so that control sites were then more likely to be unmodified locations along the shoreline.
To characterize river otter latrines, we recorded the presence of several terrestrial and aquatic physical characteristics at the latrines and control points, which were compiled based on prior latrine habitat models (Swimley et al., 1998; Depue and Ben-David, 2010; Newman and Griffin, 1994). For the terrestrial features, we measured the focal point area, slope, focal point substrate, and the distance between the focal point and the high tide line. The understory and overstory height above the focal point were measured using a tape measure and a Nikon Monarch LASER800 digital rangefinder. Additionally, we recorded burrows, drop-offs, undercut banks, and woody debris within 30 m of the focal point, as well as the focal material substrate type (Table 1). We defined beaches as exposed sandy flats between the focal point and water access and further identified them as permanent or intermittent (i.e., tidal). We classified each water interface into one of three categories (i.e., open water, marsh, or edge) if the focal point was at the interface of marsh and open water access (Table 1). Finally, we noted latrines found on manmade structures (e.g., docks, boardwalks), but due to their configurations, could not measure any of these habitat characteristics. We statistically compared habitat variables across latrine and control sites using chi-squared tests, two-tailed t-tests or their respective non-parametric counterparts (i.e., Fisher exact, Mann-Whitney tests) to identify trends using R, version 4.3.2 (R Core Team, 2020). For all statistical analyses, we used an α of 0.05 to determine significance.
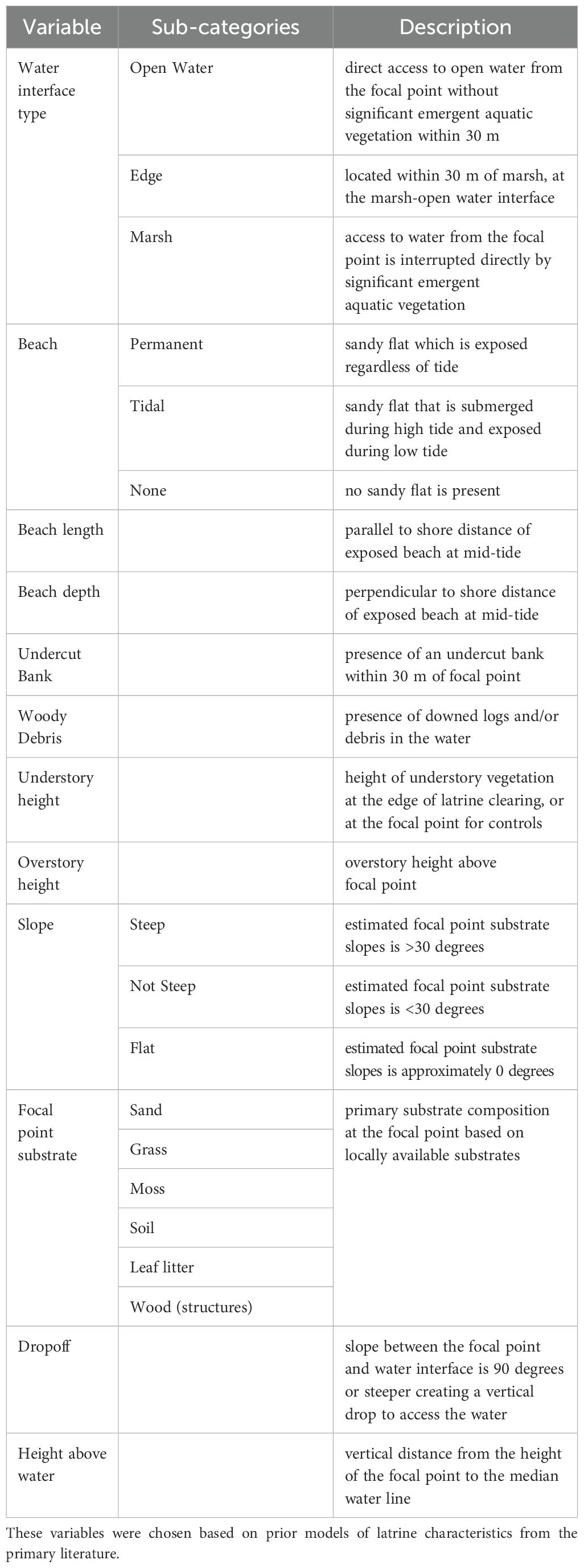
Table 1. Habitat characteristics measured at active river otter latrines and control points at the Smithsonian Environmental Research Center (SERC) in August 2021.
Collection, DNA extraction, and library preparation of genomic DNA from scat
Every time a latrine was visited, scat was collected if present. Scat was qualitatively characterized by degree of degradation (freshness), then collected following sterile protocols, homogenized, and frozen at -20°C until genetic processing. Through the course of the study period, 90 scat were collected and preserved for genetic analysis. Genomic DNA was extracted from a subset of homogenate from each frozen scat (0.2 g) using a Qiagen QIAamp Fast DNA Stool mini Kit (Qiagen, Germantown, MD) following the manufacturer’s protocol. All extractions within the same day included a blank extraction, which served as a negative extraction control for PCR. Extraction blanks were treated exactly like samples to identify contaminants that may have been introduced during genetic processing.
The primers, MG2-LCO1490F and MG2-univ-R (Tournayre et al., 2020), were used to amplify and sequence ~133 bp fragment of the mitochondrial cytochrome oxidase I (COI) gene, and the primers, 1380_eukF and 1510_eukR (Amaral-Zettler et al., 2009), were used to amplify and sequence ~130 bp fragment of the V9 region of the small subunit (SSU or 18S) gene of the ribosomal gene complex. The COI gene was chosen primarily for identification of prey items, as the COI gene has a curated database with high taxonomic coverage (Chesapeake Bay Barcode Initiative; BioProjects PRJNA396533 and PRJNA498040) and has been found to provide the highest resolution for species-level assignments (Pagenkopp Lohan et al., 2023). The 18S gene was primarily used for the identification of parasites. For both primer sets, partial Nextera indices were added to the 5’ ends to act as primers for the indexing PCR and degenerate base pairs (0, 1, 2 and 3 bp) were added before the locus-specific primer to increase heterogeneity for sequencing. These primers were combined in equal concentrations for amplification. The PCR reactions for the COI gene consisted of the following final concentrations: 1x GeneAmp 10x PCR Gold Buffer (150 mM Tris–HCl, pH 8.0; 500 mM KCl; Applied Biosystems), 2 mM MgCl2, 0.2 mM of dNTP, 0.6 μM of each pooled primer, 0.4 mg ml−1 BSA (BSA; New England Biolabs), 0.03 U/ul AmpliTaq Gold polymerase, 3 μl of DNA template, and water to a final volume of 15 μl. Thermocycling was carried out using a Bio-Rad C1000 or S1000 Thermocycler with an initial denaturation of 94°C for 10 min; followed by 40 cycles of 94°C for 30 s, 45°C for 45 s, and 72°C for 60 s; and a final extension of 72°C for 5 min. The PCR reactions for the 18S gene consisted of the following final concentrations: 1x GeneAmp 10x PCR Gold Buffer (150 mM Tris–HCl, pH 8.0; 500 mM KCl; Applied Biosystems), 1.5 mM MgCl2, 0.2 mM dNTP, 0.5 μM of each pooled primer, 0.4 mg ml−1 BSA, 0.03 U/uL units AmpliTaq Gold polymerase, 3 μl of DNA template, and water to a final volume of 15 μl. Thermocycling was carried out using a Bio-Rad C1000 or S1000 Thermocycler with an initial denaturation of 94°C for 10 min; followed by 35 cycles of 94°C for 30 s, 45°C for 45 s, and 72°C for 60 s; and a final extension of 72°C for 5 min. For all PCR products, an aliquot of each PCR product (5 μl) was electrophoresed on agarose gel (2% w/v) and visualized under UV light after GelRed staining. All PCRs were conducted in triplicate, then triplicate PCR amplicons were pooled for each sample based on the brightness of the bands on the gel.
We used dual-indexing with Nextera adapters using a unique combination for each sample. Indexing PCR reagents consisted of 12.5 µL KAPA Ready Mix, 1 µL each index (i7 or i5), 4 µL amplicon (pooled product), and 6.5 µL water for a final reaction volume of 25 µL. Thermocycling was carried out with an initial denaturation of 95°C for 5 min, followed by 8 cycles of 98 °C for 20 s, 60 °C for 45 s, and 72 °C for 45 s, and a final extension of 72 °C for 5min. To verify that indexing was successful, an aliquot of indexed product and unindexed product were both electrophoresed on agarose gel (2% w/v) stained with GelRed and visualized under UV light. The indexed product was purified with AMPure XP Beads (Beckman-Coulter, USA) following the manufacturer’s instructions for 10 µL sample reaction volume and 1.5X ratio. The bead-cleaned samples were then quantified using Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific) with a Qubit 2.0 Fluorometer following the manufacturer’s instructions. Samples were pooled based on equimolar concentrations into two separate libraries (COI and 18S, respectively). The final pooled libraries were sequenced using a MiSeq v3–300 Reagent Kit (Illumina) on an Illumina MiSeq platform at the Laboratories of Analytical Biology at the Smithsonian National Museum of Natural History. Raw sequence data were deposited in the Sequence Read Archive (SRA) under BioProject PRJNA1281117.
Microscopic examination of scat
After removing a subset for genetic analyses, 89 remaining scats had sufficient material remaining (5 mL to 45 mL) for microscopic examination and were individually rinsed in a 0.5 mm sieve using tap water to remove liquid components. Sieves were rinsed thoroughly between samples. Rinsed scats were placed in individual containers and dried at room temperature for 12–24 hours. Prey remains (e.g., bones, scales, hair, feathers) were identified to the lowest taxonomic level using a dissecting scope. If < 3 scales of a finfish species were found, these were considered contamination (e.g., sieve rinsing, proximity of scats in the field) and excluded. Undigested flies, ants, and other decomposers were also excluded from analyses, as these were likely attracted to the scat after deposition. Fish scales were identified to family using a scale key (Daniels, 1996) and reference scales from fish collected in the Rhode River. When possible, fish identifications were refined to a lower taxonomic level based on a list of local fish species collected from 1980 to present in the Rhode River (A. Hines and M. Ogburn, unpublished data). As most microscopy identifications were limited to higher taxonomic assignments, comparisons between COI and microscopy were conducted at the Order level. Three scats were excluded from the comparison of genetic vs. microscopic identification of prey as they were only analyzed using one of the two methods.
Bioinformatics
After removing the primers using cutadapt (Martin, 2011), we used the DADA2 package (Callahan et al., 2016) in R Core Team (2020) to trim for quality, merge the forward and reverse sequences, and remove chimeras. Based upon the expected fragment sizes and the size distributions, we retained sequences between 168–175 bp for COI and 167–181 bp for 18S. We assigned taxonomic rankings to amplicon sequence variants (ASVs) from COI sequences using a curated library of local sequences from the Chesapeake Bay Barcode Initiative (CBBI; BioProjects PRJNA396533 and PRJNA498040) and the nr database of the National Center for Biotechnology Information (NCBI), with priority given to the local assignments. Only those with percent identities of at least 85% and percent coverages of 80% were retained throughout the analysis and were summarized by species. Given the small size of these fragments and that prior analyses showed the likely inaccuracy of species-specific identifications for metazoans with the 18S gene (Pagenkopp Lohan et al., 2023), we only used identifications at the genus or family level. All species from the genus Lepomis were combined given that members often hybridize, making identification to species with genetics difficult. For 18S, we used the PR2 database (Guillou, L. et al., 2013) with the assignTaxonomy function using default parameters in the DADA2 package to assign taxonomy to ASVs. Taxonomic assignments were carefully reviewed and compared to known taxa collected from the Rhode River over the past decade to assess accuracy (A. Hines and M. Ogburn, unpublished data). Parasitic taxa were identified based on assignments at the appropriate taxonomic level using de Meeûs and Renaud (2002); Roberts and Janovy (2009), and peer-reviewed literature searches for those taxa containing free-living and parasitic species. If the appropriate life history information could not be found, the organism was excluded from the parasite analyses. For both markers, we defined the frequency of occurrence as the proportion of scats containing the target taxa divided by the total number of collected scats. All figures were created using the packages ggplot2 (Wickham, 2016) and ggpubr (Kassambara, 2023). We expect that river otters revisited the same latrines during the study period and could not distinguish either the number of otters residing in the study area or the number of otters leaving scat at individual sites. Thus, we assess the diet of river otters living in the study area, rather than examining variation in diet among individuals.
Results
Latrine characteristics
We identified and regularly monitored 36 areas of otter activity at SERC between October 2020 and August 2021. Of those, 25 qualified as active latrines, based on specific criteria (see Methods). At the end of the study period in August 2021, we narrowed analysis to 18 active latrines (SERC: n =12; CI: n = 4; GCREW: n = 2) where habitat measurements could be recorded. The 7 areas of otter activity that were not considered “active latrines” for this study and were excluded from analysis included 3 on manmade structures that lacked sufficient habitat characteristics to measure, 2 where habitat measurements were not feasible (i.e., those on a road culvert and narrow marsh peninsula), and 2 on natural shoreline that were inactive and lacked an identifiable focal point by the end of the study period. Each site was visited 1 to 20 times (mean: 14 ± 1.4 visits) after its initial identification. On average, river otters visited active latrines 7 times (SE = 1.1; range: 1-15) and we recovered an average of 28 scats (SE = 4.8; range: 2-63) per latrine over the course of the study. Active latrines were significantly more often characterized by permanent beaches, an open water interface, drop-offs, and undercut banks than the randomized control points (Table 2). Slope incline at the shoreline and burrows (Latrine: n = 1 each) were seldom present at latrines.
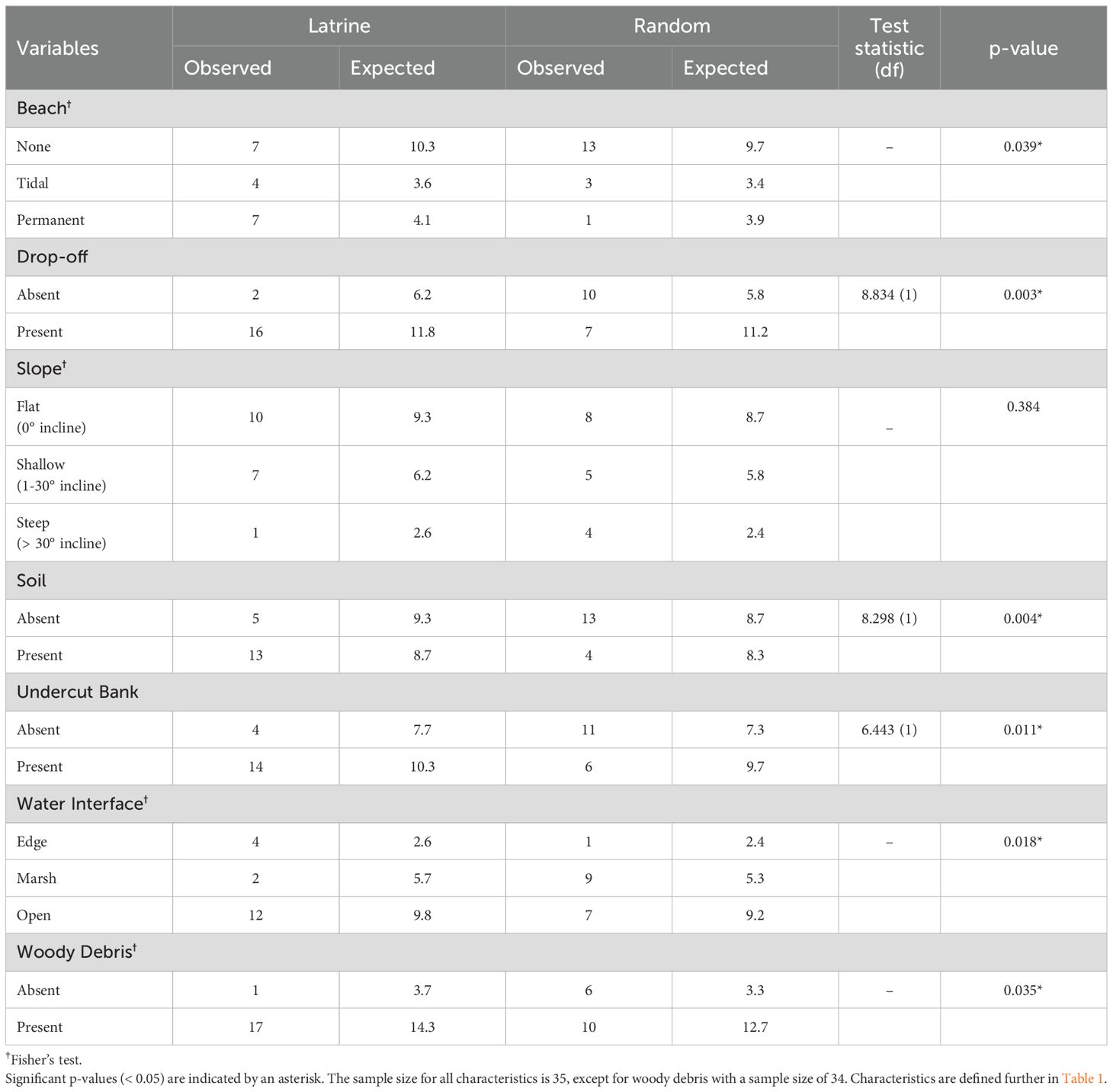
Table 2. Characteristics measured and compared between active river otter latrines and random control points using chi-squared tests of independence and Fisher’s tests.
Focal point clearings were approximately 4-m long and 4.5-m deep, with an average area of 19 m2 (SE = 5.7). The focal points of active latrines were significantly more likely to have soil-based sediment (p = 0.011; Table 2) and woody debris (p = 0.035; Table 2) compared to randomized control sites. Additionally, the focal points of the active latrines had significantly lower understory (p = 0.012; Table 3) and overstory (p = 0.001; Table 3) vegetation height compared to randomized control sites, while focal point height above the shoreline was not significantly different.
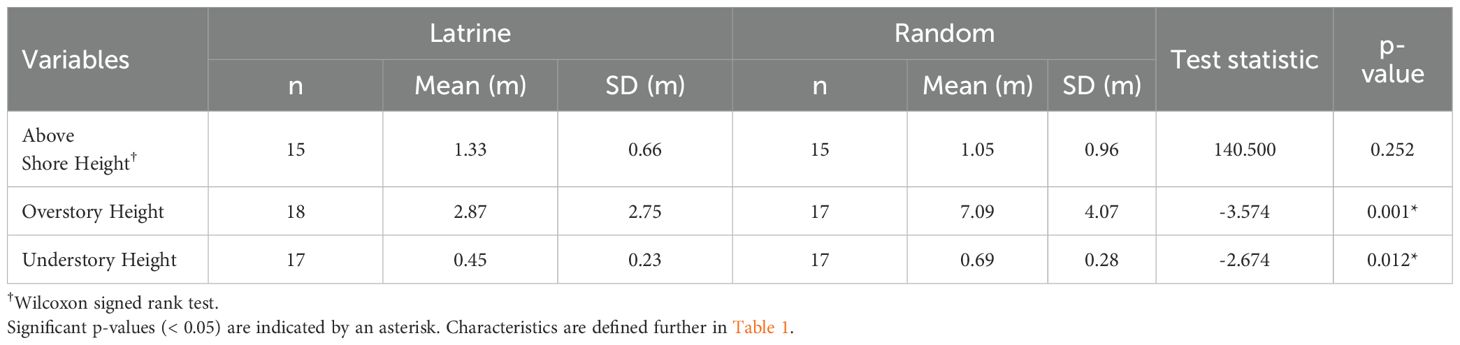
Table 3. Habitat characteristic comparisons between active river otter latrines and random control points using Student’s t and Wilcoxon signed rank tests.
We identified 5 latrines on manmade structures including floating and fixed docks, elevated boardwalks, and a shoreline staircase in the study area. Dock and boardwalk latrines had some of the highest river otter activity in the study area and were used for scat collection, but were excluded from habitat analysis due to their unique structures and our inability to measure habitat features comparable to those found in latrines along natural shoreline.
Metabarcoding of scat contents: COI
We successfully amplified and sequenced the COI gene from 90 river otter scats. In total, 8,981,256 raw reads were generated (including 4 negative extraction controls); this total was reduced to 1,246,975 reads after initial filtering, merging, and chimera removal. With the removal of (1) negative control samples, (2) L. canadensis sequences, and (3) unassigned sequences, 696,115 sequences remained in 5,922 ASVs. After careful review of taxa names, we determined that 4,604 ASVs (596,199 sequences) were likely primary prey items. When ASVs were collapsed to the lowest identifiable taxonomic level for a better approximation of species-level richness, 27 (of the original 64 taxa identified) were categorized as potential primary prey items with all but one assigned to either genus or species (Figure 2). These 27 taxa were found 333 times across all the scat collected. On average, we detected 3.7 ± 2.2 SD (range: 1-11) prey items per scat with metabarcoding, with 57.0% (190/333) identified as finfish and 36.6% (122/333) identified as crustaceans. Despite lower occurrence in scat, ~75% (432,159/596,199) of the reads generated were identified as crustaceans, while only 25.5% (152,166/596,199) of reads were identified as finfish. Other prey items occurring per scat included 11 (3.3%) annelids, 8 (2.4%) amphibians, and 2 (0.6%) birds. Two prey items identified, Cyprinus carpio and Procambarus zonangulus, are non-native species that have established within the Maryland waters of the Chesapeake Bay.
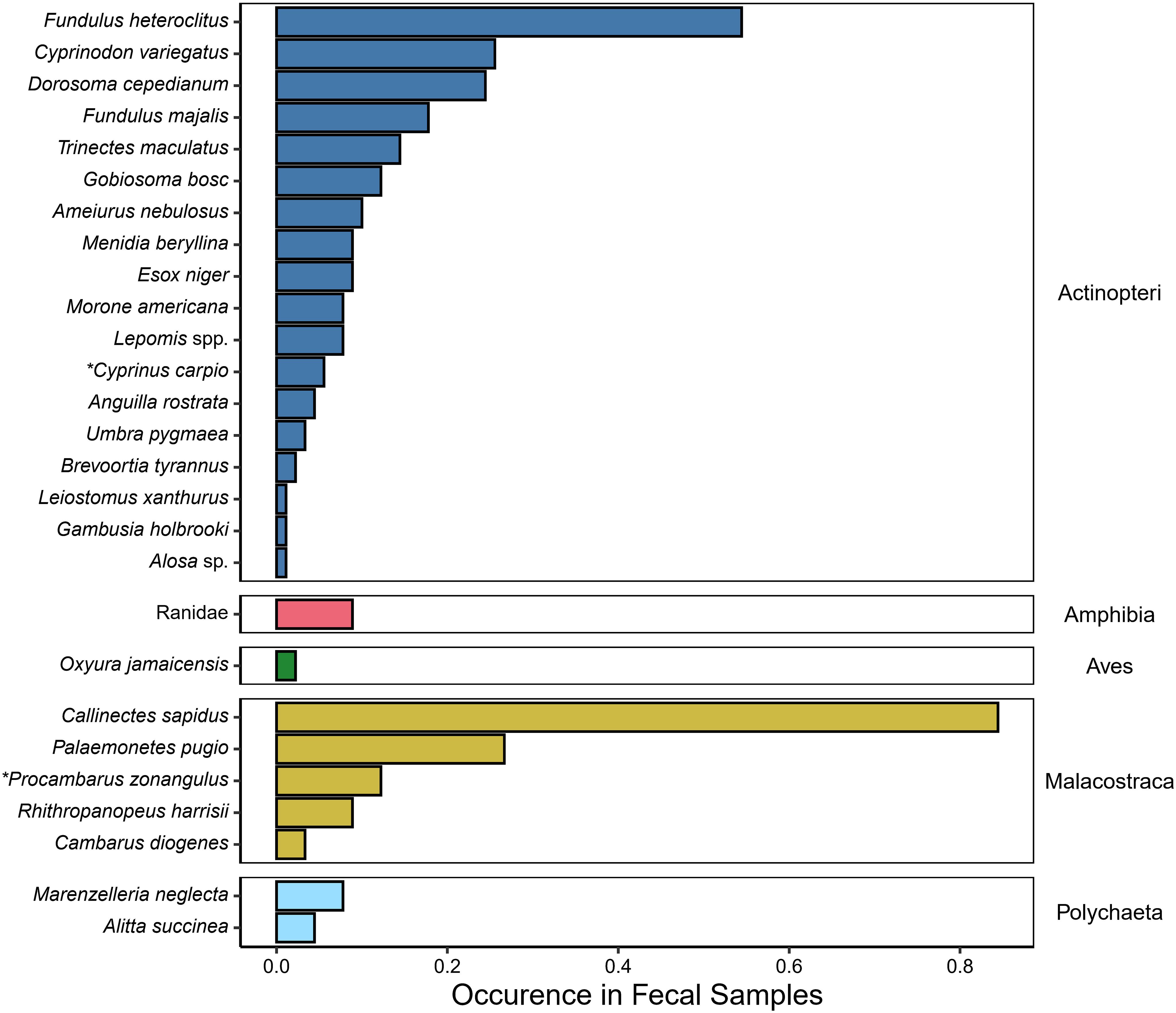
Figure 2. Occurrence of primary prey taxa detected using COI metabarcoding across all 90 fecal samples examined. The lowest taxonomic level identified is on the left, while the class for each group of taxa is listed on the right. Scientific names with an asterisk denote non-native species detected.
Microscopic examination of scat contents
From the 89 scats examined microscopically, we identified 199 individual prey items, which constituted 18 taxa across 15 orders, with an average of 2.2 ± 0.9 (mean ± SD, range: 1-5) diet items identified per scat. Generally, we could identify taxa to order or family. Remains from Actinopteri and Malacostraca were most frequently recovered, occurring in 92.1% (82/89) and 52.8% (47/89) scats examined and comprising 73.3% (146/199) and 23.6% (47/199) of the total number of prey items identified, respectively. We also identified two occurrences (1.0%) of Aves and Anura each, one Cicadoidea (1.0%), and one Amblyomma americanum (1.0%).
While over half of the potential prey items were found using both COI and microscopy (Figure 3), our metabarcode analysis recovered more unique taxa compared to microscopy. Of the 20 orders that were detected, 45% (n=9) were detected with both methods, while 35% (n=7) were only detected with metabarcoding and 20% (n=4) were only detected with microscopy. Specifically, soft-bodied organisms such as polychaetes (e.g., Spionia, Phyllodocida), fish with small scales (e.g., Anguilla rostrata), and scaleless fish (e.g., Ameiurus nebulosus, Gobiosoma bosc) were only identified genetically (Figure 3). In contrast, the only truly unique order we detected using microscopy was Hemiptera, including both Ambyloma americanum and Cicadoidea. The other unique taxa to this method could only be identified to class. Most of the scaled Actinopteri were similarly detected by both methods (e.g., Cyprinodontiformes, Cypriniformes). Likewise, though decapods were detected with both methods, they were detected more frequently with genetic data as compared to microscopy (Figure 3).
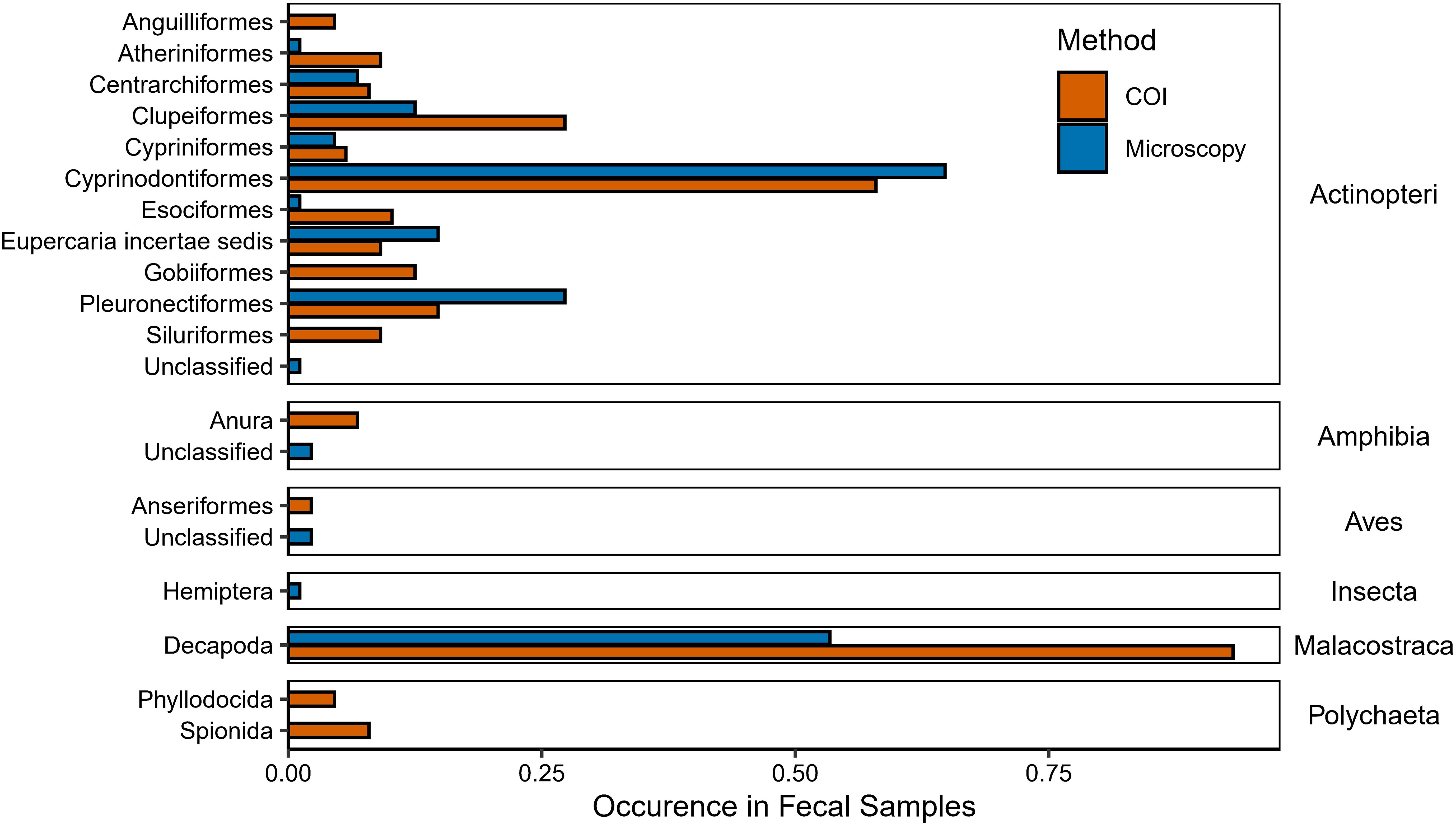
Figure 3. Comparison of the occurrence of primary prey taxa detected by COI metabarcoding and/or microscopy. The lowest taxonomic level identified is on the left, while the class for each group of taxa is listed on the right. Note that unclassified prey items could not be identified below class and are thus listed as unclassified as the lowest taxonomic level.
Metabarcoding of scat contents for parasites: 18S
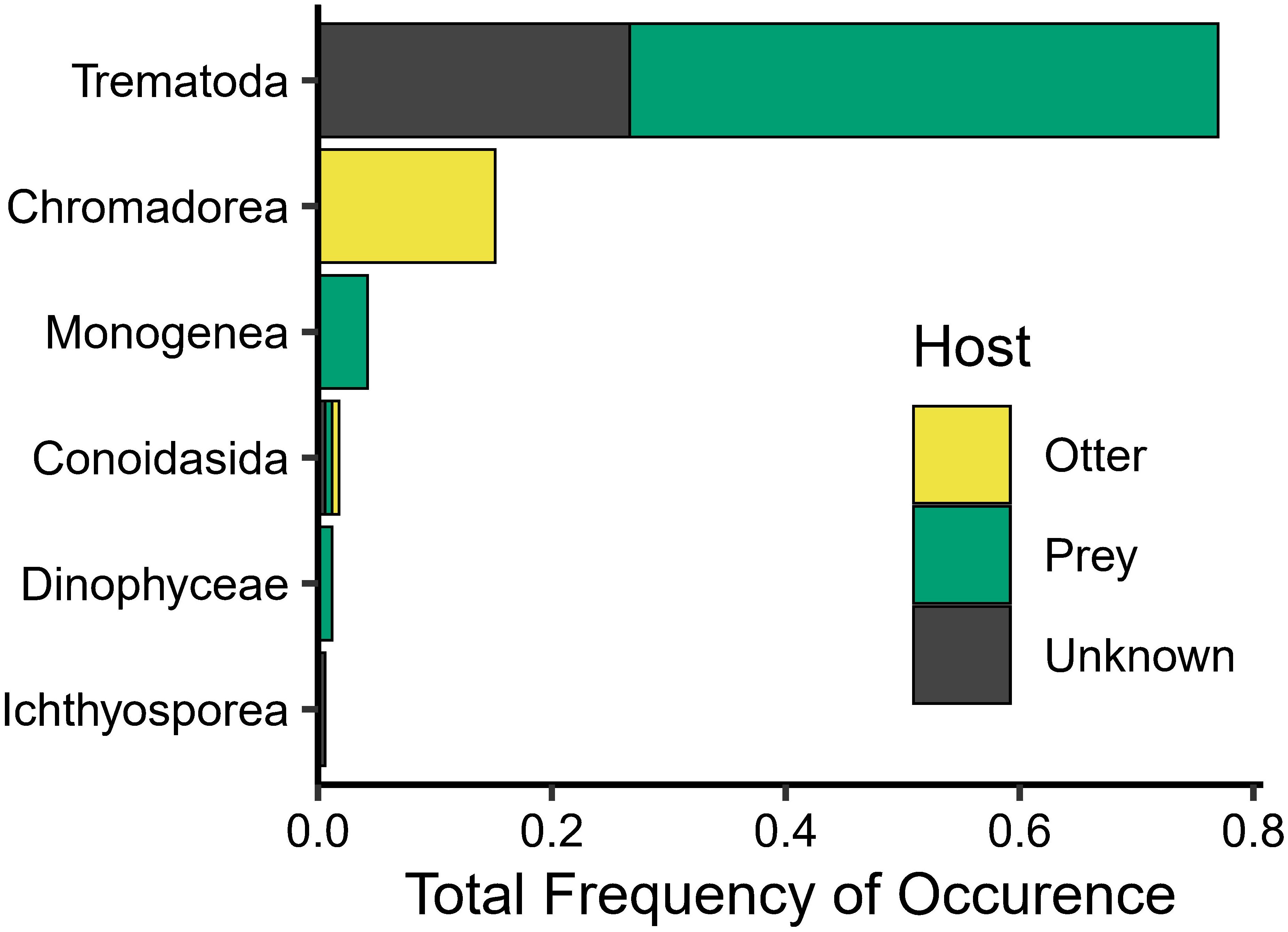
We successfully amplified and sequenced the 18S gene from 90 river otter scats. In total, 6,923,816 raw reads were generated (including 4 negative extraction controls); this total was reduced to 3,138,908 reads after initial filtering, merging, and chimera removal. Upon removing (1) negative control samples, (2) L. canadensis sequences, (3) unassigned sequences, and (4) narrowing the dataset to only parasitic taxa, this dataset included 591,412 sequences and 138 ASVs from 10 taxa from six classes. On average, parasite taxa occurred 1.8 ± 1.0 SD times (range: 0-4) per scat with trematodes being the most frequently detected (Figure 4). Trematodes also had the highest relative abundance with 99.5% (588,553/591,412) of sequences, 81.3% (481,030/591,412) of which were Plagiorchiids. While the nematodes (e.g., Strongylida) and some Apicomplexans (e.g., Cystoisospora) identified most likely parasitized the river otters, most of the parasitic taxa identified (Figure 4), such as the monogeneans (e.g., Gyrodactylideans), myxozoans (e.g., Amyloodinium), and some trematodes (e.g., Plagiorchiida) most likely parasitized the primary prey items.
Discussion
We found high similarity in the characteristics of latrines in both tidally influenced and riverine habitats, indicating that river otters appear to seek similar features across these aquatic habitats, with river otters frequently using man-made structures as latrines in urbanized coastal habitats. Our analyses of river otter diets at SERC indicate they consume both terrestrial and aquatic fauna, with finfish and crustaceans making up the bulk of their diet. Additionally, the sensitivity of the genetic approach for scat contents allowed us an unparalleled examination of the role of parasites in the food web. While we identified some parasites that are likely infecting river otters, most of the parasites identified were parasites of their prey, adding to an already large number of publications highlighting the importance of parasites in coastal food webs (e.g., Lafferty et al., 2008; Johnson et al., 2010; Preston et al., 2020; Thongthaisong et al., 2022).
Latrine characteristics
Similar to previous research examining characteristics of river otter latrines in riverine systems (Swimley et al., 1998; Crait and Ben-David, 2007), river otter latrines at SERC were characterized by woody debris, undercut banks, beaches, access to an open water interface, soil as the focal point substrate, and lower understory and overstory vegetation height. Latrines at SERC were unlikely to be found near marshes, which were characterized by sloped shorelines and vegetation obscuring the view of water that may limit river otter’s timely access to water consistent with previous latrine studies (Swimley et al., 1998). While marshes comprise much of the SERC shoreline, the river otters instead chose areas of shoreline with woody debris and access to open water, characteristics that are broadly associated with river otter latrines across riverine, lacustrine, and coastal systems (Swimley et al., 1998). Woody debris may serve as a river otter rest area, provide access to land, and create aquatic habitat for prey (Greer, 1955; Melquist and Hornocker, 1983). Drop-offs may provide easier access to water than a gradually sloped shoreline (Dronkert-Egnew, 1991; Bowyer et al., 1995; Swimley et al., 1998).
Some habitat characteristics of latrines, such as overstory cover, understory cover, and understory density, are hypothesized to provide protective cover from terrestrial and aerial predators (Dronkert-Egnew, 1991; Depue and Ben-David, 2010). Despite the fact the overall canopy height was consistent across the study area, we found that focal point overstory canopy height was shorter at latrines than control points, likely indicating that river otters concentrate marking activity under low hanging branches. While open understory may improve a river otter’s ability to detect terrestrial predators, dense understory vegetation and visual obscurity could alternatively provide cover for river otters (Crowley et al., 2012). Additionally, river otter latrines are important sites for olfactory communication, so increased cover may protect scat from degradation and thus extend the length of the scent, but further research is needed to assess this impact. For example, we found that the scat deposited at a latrine on a manmade dock (not included in habitat analysis) was concentrated under a permanent canopy, rather than along the larger exposed areas. Future research is needed to investigate the trade-offs in latrine characteristics in coastal areas, particularly between those on man-made versus natural substrates.
Tidally influenced waters of the Chesapeake Bay contain many non-natural structures, such as permanent or floating docks, benches, picnic tables, and even some floating wetlands (e.g., floating wetland in Baltimore’s Inner Harbor). We found that the docks and boardwalk on the SERC campus were frequently visited and used as latrines by river otters. Latrines on manmade structures like docks were likely easier to access than most natural shoreline latrines (for us and the river otters) and showed signs of regular river otter activity, despite boat use and proximity to actively used walking trails. Though not often examined for latrine usage in river systems (Swimley et al., 1998; Depue and Ben-David, 2010), manmade structures and developed shorelines may be a significant component of river otter habitat in increasingly urban landscapes.
Due to the timing of staff availability, these surveys were conducted during the time of year when latrine usage is generally low (i.e., spring and summer). Thus, we expect that our study underestimates the number of active latrines along the shoreline, as we expect increases in number and usage of latrines during fall and winter months. However, while more latrines might be used during different seasons, we do not expect that the characteristics of the latrine sites would not change due to the high consistency found in our study and compared to previous studies.
Diet: metabarcoding vs. microscopy
Our findings are consistent with previous diet studies that river otters are generalist predators known to eat native and non-native prey species, including teleost fish, crayfish, and aquatic invertebrates, and occasionally amphibians, small mammals, and birds (Greer, 1955; Wengeler et al., 2010; Stearns and Serfass, 2011; Day et al., 2015; Juarez-Sanchez et al., 2019). Additionally, as generalist predators, river otters will consume available prey, which varies seasonally and spatially (Serfass et al., 1990; Cote et al., 2008). For example, river otters across Humboldt Bay consumed more waterfowl during peak migration times and when in areas where more birds were located Penland and Black, 2009). Our examination of scat through both methods revealed that the diet of river otters was being dominated by fish and crustaceans of various sizes, consistent with these prior findings that river otters consume those prey that are most abundant.
While river otters are known to consume teleost fish year round, multiple studies indicate that their summer diets may be more varied, with the proportion of fish decreasing and being replaced by other seasonally available prey. For example, river otters in Humboldt Bay were found to consume more crustaceans during summer months (Penland and Black, 2009). River otters in North Dakota primarily consumed teleost fish year-round (commonly cyprinids, ictalurids, and catostomids), but increasingly consumed crayfish, insects, birds, and amphibians during the summer months (Stearns and Serfass, 2011). Similarly, river otters in Utah were also found to consume fish year-round, but higher occurrences of crustaceans, reptiles, and birds were found in spring and summer months (Day et al., 2015). Our results are consistent with what these prior studies report for diets during summer months, indicating that our study period likely captured the diet of local river otters when it is expected to be most diverse. We found river otters consumed primarily teleost fish, with species in the Cyprinodontidae and Clupeidae being the most frequently consumed. However, the most abundant crustacean identified (based on number of sequences) and the prey item found in the greatest number of scats was the American blue crab (Callinectes sapidus), which is abundant and widespread in Chesapeake Bay and a commercially important species. We also found that river otters consumed anadromous species, such as Striped Bass (Morone saxatilis), Gizzard Shad (Dorosoma cepedianum), and American Eel (Anguila rostrata). Additionally, these river otters were found to opportunistically consume freshwater species (e.g., devil crayfish, Cambarus diogenes), amphibians, and ducks. While most prey identified from scat were native, two non-native species commonly found in the area, common carp (Cyprinus carpio) and white river crayfish (Procambarus zonangulus), were identified in scats. Additional research is needed to examine how seasonality impacts river otter diets in the Chesapeake Bay, as we expect that teleost fish may be more important prey items for river otters during fall and winter months, which would be consist with prior studies.
While traditional studies on river otter diets have used only microscopic examination of river otter scat, we found that metabarcoding provided lower-level taxonomic resolution and identified a wider breadth of prey taxa. For example, with metabarcoding, we identified scaleless and finely scaled species that were not found using microscopy including catfish (Siluriformes), gobies (Gobiformes), annelids (Phyllodocida, Spionida), and eels (Anguilliformes). Additionally, pickerel (Esociformes), silversides (Atheriniformes), herring/anchovies (Clupeiformes), frogs (anura), and crabs (Decapoda) were found more frequently using metabarcoding, while killifish (Cyprinodontiformes) and flatfish (Pleuronectiformes) were identified more frequently using microscopy. Thus, given that summer diet diversity tends to be higher for river otters, it is possible that prior studies that used only microscopic methods underestimated the increase in variability during summer months. The high sensitivity of metabarcoding meant that we detected both primary and secondary prey, so we had to assess which taxa were more likely primary prey for our comparisons. As a result, true bugs (Hemiptera) were only considered diet items using microscopy, because we could visualize that these were prey items (rather than insects that colonized the scat after deposition). This affected a single scat, where cicadas were identified as an abundant prey item, likely resulting from their consumption during the summer 2021 brood X emergence. In contrast, metabarcoding cannot provide information on the size of fish prey (e.g., Stearns and Serfass, 2011), or the volume of prey item that was consumed, though information about volume consumed and size of prey could potentially be obtained from microscopic examination of prey. Our findings indicate that metabarcoding from river otter scat is an effective, efficient, and non-invasive method to examine river otter diets, though a combined morphological and genetic approach is required to get a more holistic assessment of prey consumed.
Parasites of river otters and prey
Kimber and Kollias (2000) wrote the last major review of parasites and infectious diseases of river otters, in which they note how little is known on this topic. Most previous studies on parasites of river otters involved dissections of carcasses, usually obtained from trappers, to identify parasites and potential diseases impacting river otters at the time of their capture (Fleming et al., 1977; McAllister et al., 2016). In this study, we tested a non-invasive approach to examining parasites infecting river otters, which limited us to identify only those parasites that would be present in scat. Nonetheless, our results indicated that this method was able to detect parasites of the river otters and provide a unique examination of the role of parasites more broadly, particularly those likely infecting the teleost fish prey, in the diet of river otters.
Multiple previous studies have discussed the importance of parasites in food webs, as the parasites can be important as prey and can increase the likelihood of some prey being consumed through behavior modifications (Johnson et al., 2010; Preston et al., 2020; Thongthaisong et al., 2022). Most parasites identified from the scat do not infect mammals, so they most likely infected the river otters’ primary prey, resulting in concomitant predation of the parasites (Johnson et al., 2010). The most frequently occurring parasitic taxa was trematodes, which are parasitic worms (or flukes) that require multiple hosts throughout their life cycle, and many are trophically transmitted. For example, we identified trematodes in the Order Diplostomida which typically use a fish as a second intermediate, then a bird or mammal as final or definitive host (Blasco-Costa and Locke, 2017). We also identified trematodes in the Order Plagiorchiida, which typically use an arthropod as a second intermediate host and then a fish as a final host (Cribb et al., 2003). In addition to the trematodes, we recovered monogeneans, including Gyrodactylidea, which commonly infect fish gills (Bakke et al., 2007). Finally, we detected multiple myxozoan taxa, many of which parasitize fish (Okamura et al., 2015). The relatively high occurrence of parasites that primarily infect prey taxa in the scat indicate that these parasites are most likely consumed along with the prey.
While most of the parasites detected were likely infecting the prey, we also detected parasites likely infecting the river otters. For example, we identified nematodes in the Order Strongylida, which are primarily known to infect terrestrial vertebrates and marine mammals, with only one species known to infect fish (Fleming et al., 1977; Chilton et al., 2006). This finding is also supported by our observations of large red nematodes present in fresh scat. Additionally, we detected multiple apicomplexans, some of which were most likely infecting the river otters, including parasites in Order Sarcocystidae, which infect a wide range of mammalian species (Seeber and Steinfelder, 2016; Cotey et al., 2022).
While the high sensitivity of metabarcoding can detect low level infections of a wide range of parasites of both the host and prey, this method cannot be used to determine the life stage of the parasite. Additionally, while the local database that we used for assigning identifications to COI sequences provided high-level resolution of animal prey, our ability to identify parasites is limited by the short length of the fragment that we could amplify from the scat and the lack of available reference sequences. Future studies could combine metabarcoding with a traditional approach to assess parasites in feces, such as flotations or sedimentation methods to identify parasite eggs, which could provide a wider breadth of parasites impacting river otters.
Conclusions
This study is the first to assess latrine characteristics, prey, and parasites of North America river otters in a tidally influenced subestuary of the Chesapeake Bay. As river otter populations continue to expand and move into more urbanized areas of this watershed, these latrine characteristics can be used to identify latrines from which non-invasive scat studies can be conducted to examine river otter ecology and health across these land- and seascapes. Additionally, as river otters are generalist predators, examining their diet across seasons and years would provide a broader indication of what food is consumed and how their diet changes with seasonal migrations of prey, particularly anadromous species. While our study focused on a single subestuary within the Chesapeake Bay, future studies should include additional subestuaries to provide a broader regional representation of river otter ecology, particularly regarding their diet and latrine use.
Data availability statement
The original contributions presented in the study are publicly available. This data can be found here: NCBI Sequence Read Archive, accession PRJNA1281117.
Ethics statement
This study was reviewed by the IACUC committee at the Smithsonian Environmental Research Center, which determined that an IACUC was not required. The study was conducted in accordance with the local legislation and institutional requirements.
Author contributions
CW: Methodology, Formal Analysis, Writing – original draft, Investigation, Data curation, Conceptualization, Project administration, Writing – review & editing, Funding acquisition. TS: Writing – original draft, Writing – review & editing, Conceptualization, Supervision, Methodology. EP: Formal Analysis, Visualization, Writing – original draft, Data curation, Writing – review & editing. LY: Writing – review & editing, Writing – original draft, Investigation. KB: Writing – original draft, Formal Analysis, Writing – review & editing. KP: Writing – review & editing, Conceptualization, Project administration, Methodology, Writing – original draft, Supervision, Resources, Investigation, Data curation, Funding acquisition.
Funding
The author(s) declare that financial support was received for the research and/or publication of this article. C. Wise received funding through a Smithsonian Graduate Fellowship and Frostburg State University. This project was also funded by Hunterdon funds from the Smithsonian Institution to KMP Lohan.
Acknowledgments
Thank you to the many SERC volunteers who assisted with fieldwork for this study.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Amaral-Zettler L. A., McCliment E. A., Ducklow H. W., and Huse S. M. (2009). A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS One 4, 1–9. doi: 10.1371/journal.pone.0006372
Bakke T. A., Cable J., and Harris P. D. (2007). The biology of Gyrodactylid Monogeneans: the “Russian-doll killers. Adv. Parasitol. 64, 161–460. doi: 10.1016/S0065-308X(06)64003-7
Bakker J., van Zinnicq Bergmann M. P. M., Stolen M., and Kiszka J. J. (2024). Assessing the diet of the elusive North American river otter (Lontra canadensis) in Florida using faecal DNA metabarcoding. Aquat. Conservation: Mar. Freshw. Ecosyst. 34, 1–7. doi: 10.1002/aqc.4257
Barocas A., Golden H. N., Harrington M. W., Mcdonald D. B., and Ben-David M. (2016). Coastal latrine sites as social information hubs and drivers of river otter fission-fusion dynamics. Anim. Behav. 120, 103–114. doi: 10.1016/j.anbehav.2016.07.016
Bay Bulletin (2023). “VIDEO: otters spotted in Baltimore’s often-polluted jones falls,” in Chesapeake Bay Magazine. (Annapolis, MD: Chesapeake Bay Media). Available online at: https://www.chesapeakebaymagazine.com/video-otters-spotted-in-baltimores-often-polluted-jones-falls/ (Accessed May 15, 2023).
Ben-David M., Duffy L. K., Blundell G. M., and Bowyer R. T. (2001). Natural exposure of coastal river otters to mercury: relation to age, diet, and survival. Environ. Toxicol. Chem. 20, 1986–1992. doi: 10.1002/etc.5620200917
Blasco-Costa I. and Locke S. A. (2017). Life history, systematics and evolution of the Diplostomoidea Poirier 1886 : progress, promises and challenges emerging from molecular studies. Adv. Parasitol. 98, 167–225. doi: 10.1016/BS.APAR.2017.05.001
Bouley P., Isadore M., and Carroll T. (2015). Return of North American river otters (Lontra canadensis) to coastal habitats of the San Francisco Bay Area, California. Northwestern Nat. 96, 1–12. doi: 10.1898/NWN14-09.1
Bowyer R. T., Testa J. W., and Faro J. B. (1995). Habitat selection and home ranges of river otters in a marine environment: effects of the Exxon Valdez Oil Spill. J. Mammalogy 76 (1), 1–11.
Brooks R. P. and Serfass T. L. (2019). Aquatic habitat classifications: implications for otter conservation. IUCN Otter Specialist Group Bull. 36, 3–12.
Callahan B. J., McMurdie P. J., Rosen M. J., Han A. W., Johnson A. J. A., and Holmes S. P. (2016). DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13 (7), 581–583. doi: 10.1038/nmeth.3869
Cape Gazette (2022).Otters devour Lewes garden’s prized koi. Available online at: https://www.capegazette.com/article/otters-devour-lewes-garden%E2%80%99s-prized-koi/240808google_vignette (Accessed January 12, 2022).
Carroll T., Hellwig E., and Isadore M. (2020). An approach for long-term monitoring of recovering populations of Nearctic river otters (Lontra canadensis) in the San Francisco Bay area, California. Northwestern Nat. 10, 77–91. doi: 10.1898/1051-1733-101.2.77
Chesapeake Bay Program. (2016). Map of Maryland's Chesapeake Bay Land-River Segments - Phase 5. Available online at: https://mde.maryland.gov/programs/Water/TMDL/DataCenter/Pages/CBLandRiverSegmentsP5.aspx (Accessed June 16, 2025).
Chilton N. B., Huby-Chilton F., Gasser R. B., and Beveridge I. (2006). The evolutionary origins of nematodes within the order Strongylida are related to predilection sites within hosts. Mol. Phylogenet. Evol. 40, 118–128. doi: 10.1016/J.YMPEV.2006.01.003
Cote D., Stewart H. M. J., Gregory R. S., Gosse J., Reynolds J. J., Stenson G. B., et al. (2008). Prey selection by marine-coastal river otters (Lontra canadensis) in Newfoundland, Canada. J. Mammalogy 89, 1001–1011. Available online at: https://academic.oup.com/jmammal/article/89/4/1001/867028 (Accessed January 14, 2021).
Cotey S. R., Scimeca R., Chang L., Carpenter A. L., Will E. E., Ott-Conn C., et al. (2022). Toxoplasma gondii prevalence, partial genotypes, and spatial variation in North American river otters (Lontra canadensis) in the upper peninsula of Michigan, USA. J. Wildlife Dis. 58, 869–881. doi: 10.7589/JWD-D-22-00021
Crait J. R. and Ben-David M. (2007). Effects of river otter activity on terrestrial plants in trophically altered Yellowstone Lake. Ecology 88, 1040–1052. doi: 10.1890/06-0078
Cribb T. H., Bray R. A., Olson P. D., Timothy D., and Littlewood J. (2003). Life cycle evolution in the Digenea: a new perspective from phylogeny. Adv. Parasitol. 54, 197–254. doi: 10.1016/S0065-308X(03)54004-0
Crowley S., Johnson C. J., and Hodder D. (2012). Spatial and behavioral scales of habitat selection and activity by river otters at latrine sites. J. Mammalogy 93, 170–182. doi: 10.1644/10-MAMM-A-362.1
Daniels R. A. (1996). Guide to the Identification of Scales of Inland Fishes of Northeastern North America. No. 448 (Albany, NY: University of State of New York).
Day C. C., Mcmillan B. R., and Westover M. D. (2015). Seasonal diet of the northern river otter (Lontra canadensis): what drives prey selection? Can. J. Zoology. 93, 197–205. doi: 10.1139/cjz-2014-0218
de Meeûs T. and Renaud F. (2002). Parasites within the new phylogeny of eukaryotes. Trends Parasitol. 18, 247–250. doi: 10.1016/S1471-4922(02)02269-9
Depue J. E. and Ben-David M. (2010). River otter latrine site selection in arid habitats of western Colorado, USA. J. Wildlife Manage. 74, 1763–1767. doi: 10.2193/2008-065
de Sousa L. L., Silva S. M., and Xavier R. (2019). DNA metabarcoding in diet studies: unveiling ecological aspects in aquatic and terrestrial ecosystems. Environ. DNA. 1, 199–214. doi: 10.1002/edn3.27
Drake L. E., Cuff J. P., Bedmar S., McDonald R., Symondson W. O. C., and Chadwick E. A. (2023). Otterly delicious: spatiotemporal variation in the diet of a recovering population of Eurasian otters (Lutra lutra) revealed through DNA metabarcoding and morphological analysis of prey remains. Ecol. Evol. 13, 1–18. doi: 10.1002/ece3.10038
Dronkert-Egnew A. E. (1991). River otter population status and habitat use in northwestern Montana (Ann Arbor, MI: ProQuest LLC).
Fleming W. J., Dixon C. F., and Lovett J. W. (1977). Helminth parasites of river otters (Lutra canadensis) from southeastern Alabama. Proc. Helminthological Soc. 44, 131–135.
Gaydos J. K., Miller W. A., Gilardi K. V., Melli A., Schwantje H., Engelstoft C., et al (2007a). Does human proximity affect antibody prevalence in marine-foraging river otters (Lontra canadensis)? J. Wildlife Dis. 43, 116–123. doi: 10.7589/0090-3558-43.1.116
Gaydos J. K., Miller W. A., Gilardi K. V. K., Melli A., Schwantje H., et al. (2007b). Cryptosporidium and Giardia in marine-foraging river otters (Lontra canadensis) from the Puget Sound Georgia Basin ecosystem. J. Parasitol. 93, 198–202. doi: 10.1645/GE-928R.1
Grablick C. (2021). Extremely Cute River Otter Spotted On The National Mall (Washington, DC: WAMU 88.5). Available online at: https://dcist.com/story/21/05/24/very-cute-otter-spotted-at-the-tidal-basin/ (Accessed May 24, 2025).
Greer K. R. (1955). Yearly food habits of the river otter in the Thompson Lakes region, northwestern Montana, as indicated by scat analyses. Am. Midland Nat. 54, 299–313. doi: 10.2307/2422569
Guillou L., Bachar D., Audic S., Bass D., Berney C., Bittner L., et al. (2013). The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res. 41, D597–D604. doi: 10.1093/NAR/GKS1160
Heenkenda E. J., Heniff A. C., Fisher B. E., Albers G., and DeWoody J. A. (2024). A DNA-based evaluation of North American river otter diet with respect to fishes: implications for the conservation of aquatic predators and their prey. Aquat. Conservation: Mar. Freshw. Ecosyst. 34, 1–7. doi: 10.1002/aqc.4078
Johnson P. T. J., Dobson A., Lafferty K. D., Marcogliese D. J., Memmott J., Orlofske S. A., et al. (2010). When parasites become prey: ecological and epidemiological significance of eating parasites. Trends Ecol. Evol. 25, 362–371. doi: 10.1016/J.TREE.2010.01.005
Juarez-Sanchez D., Blake J. G., and Hellgren E. C. (2019). Variation in Neotropical river otter (Lontra longicaudis) diet: effects of an invasive prey species. PLoS One 14, 1–21. doi: 10.1371/journal.pone.0217727
Kassambara A. (2023). ggpubr: ‘ggplot2’ Based Publication Ready Plots. R package version 0.6.0. Available online at: https://rpkgs.datanovia.com/ggpubr/ (Accessed February 21, 2025).
Kimber K. R. and Kollias G. V. (2000). Infectious and parasitic diseases and contaminant-related problems of North American river otters (Lontra canadensis): a review. J. Zoo Wildlife Med. 31, 452–472. doi: 10.1638/1042-7260(2000)031[0452:IAPDAC]2.0.CO;2
Lafferty K. D., Allesina S., Arim M., Briggs C. J., De Leo G., Dobson A. P., et al. (2008). Parasites in food webs: the ultimate missing links. Ecol. Lett. 11, 533–546. doi: 10.1111/j.1461-0248.2008.01174.x
Larivière S. and Walton L. R. (1998). Lontra canadensis. Mamm. Species 587, 1–8. doi: 10.2307/3504417
Martin M. (2011). Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12. doi: 10.14806/ej.17.1.200
McAllister C. T., Surf A., Tumlison R., Bursey C. R., and Fayton T. J. (2016). Gastrointestinal parasites of the northern river otter, Lontra canadensis (Carnivora: Mustelidae), from Arkansas. J. Arkansas Acad. Sci. 70, 284–286. doi: 10.54119/jaas.2016.7041
Melquist W. E. and Hornocker M. G. (1983). Ecology of river otters in west central Idaho. Wildlife Monogr. 83, 3–60.
Mowbray E. E., Chapman J. A., and Goldsberry J. R. (1976). Preliminary observations on otter distribution and habitat preferences in Maryland with descriptions of otter field sign. Trans. Northeast Section Wildlife Soc. 33, 125–131.
Mowbray E. E., Pursley D., and Chapman J. A. (1979). The status, population characteristics and harvest of the river otter in Maryland (Waverly Press Inc).
Newman D. G. and Griffin C. R. (1994). Wetland use by river otters in Massachusetts. J. Wildlife Manage. 58, 18–23. doi: 10.2307/3809544
Okamura B., Gruhl A., and Bartholomew J. L. (2015). An Introduction to Myxozoan Evolution, Ecology and Development (Springer, Cham).
Olson Z. H., Serfass T. L., and Rhodes O. E. (2008). Seasonal variation in latrine site visitation and scent marking by Nearctic river otters (Lontra canadensis). IUCN Otter Specialist Group Bull. 25, 108–120.
Pagenkopp Lohan K. M., Aguilar R., DiMaria R., Heggie K., Tuckey T. D., Fabrizio M. C., et al. (2023). Juvenile striped bass consume diverse prey in Chesapeake Bay tributaries. Mar. Coast. Fisheries: Dynamics Management Ecosystem Scinece 15, 1–18. doi: 10.1002/mcf2.10259
Penland T. F. and Black J. M. (2009). Seasonal variation in river otter diet in coastal northern California. Northwestern Nat. 9, 233–237. doi: 10.1898/NWN08-21.1
Preston D. L., Layden T. J., Segui L. M., Falke L. P., Brant S. V., and Novak M. (2020). Trematode parasites exceed aquatic insect biomass in Oregon stream food webs. J. Anim. Ecol. 90, 766–775. doi: 10.1111/1365-2656.13409
Quéméré E., Aucourd M., Troispoux V., Brosse S., Murienne J., Covain R., et al. (2021). Unraveling the dietary diversity of neotropical top predators using scat DNA metabarcoding: A case study on the elusive giant otter. Environ. DNA. 1–12. doi: 10.1002/edn3.195
R Core Team (2020). R: A language and environment for statistical computing (Version 4.3.2) (Vienna, Austria: R Foundation for Statistical Computing). Available online at: https://www.R-project.org/ (Accessed April 22, 2021).
Reid D. G., Code T. E., Reid A. C. H., and Herrero S. M. (1994). Spacing, movements, and habitat selection of the river otter in boreal Alberta. Can. J. Zoology 72, 1314–1324. doi: 10.1139/z94-175
Roberts L. S. and Janovy J. Jr. (2009). Foundations of Parasitology. 8th ed (McGraw-Hill, Boston: The McGraw-Hill Companies).
Sanders C. W., Spear S. F., Black K., Olfenbuttel C., and DePerno C. S. (2023). Diet of the North American river otter (Lontra canadensis) in North Carolina using 2 methods. Wildlife Soc. Bull. 47, 1–12. doi: 10.1002/wsb.1502
Seeber F. and Steinfelder S. (2016). Recent advances in understanding apicomplexan parasites. F1000Research 5, F1000 Faculty Rev. doi: 10.12688/F1000RESEARCH.7924.1
Serfass T. L., Brooks R. P., and Pearce K. J. (2020). Monthly patterns of scat marking by river otters (Lontra canadensis) at latrines along Pine Creek in Northcentral Pennsylvania, USA. Otter Specialist Group Bull. 36, 21–27.
Serfass T. L., Carpenter C. P., and Triska M. D. (2019). Monthly variation in scat marking by river otters along Tionesta Creek in Northwestern Pennsylvania. Can. Wildlife Biol. Manage. 8, 36–44.
Serfass T. L., Rymon L. M., and Brooks R. P. (1990). Feeding relationships of river otters in northeastern Pennsylvania. Trans. Northeast Section Wildlife Soc. 47, 43–53.
Speich S. M. and Pitman R. L. (1984). River otter occurrence and predation on nesting marine birds in the Washington Islands Wilderness. Murrelet 65, 25–27. doi: 10.2307/3534208
Stearns C. R. and Serfass T. L. (2011). Food habits and fish prey size selection of a newly colonizing population of river otters (Lontra canadensis) in Eastern North Dakota. Am. Midland Nat. 165, 169–184. doi: 10.1674/0003-0031-165.1.169
Stubbs Q., Yeo I., Lang M., Townshend J., Sun L., Prestegaard K., et al. (2020). Assessment of wetland change on the Delmarva peninsula from 1984 to 2010. J. Coast. Res. 36, 575–589. doi: 10.2112/JCOASTRES-D-16-00038.1
Swimley T. J., Serfass T. L., Brooks R. P., and Tzilkowski W. M. (1998). Predicting river otter latrine sites in Pennsylvania. Wildlife Soc. Bull. 26, 836–845.
Thongthaisong P., Kasada M., Grossart H., and Wollrab S. (2022). Critical role of parasite-mediated energy pathway on community response to nutrient enrichment. Ecol. Evol. 12, 1–12. doi: 10.1002/ece3.9622
Tournayre O., Leuchtmann M., Filippi-Codaccioni O., Trillat M., Piry S., Pontier D., et al. (2020). In silico and empirical evaluation of twelve metabarcoding primer sets for insectivorous diet analyses. Ecol. Evol. 10, 6310–6332. doi: 10.1002/ece3.6362
Wengeler W. R., Kelt D. A., and Johnson M. L. (2010). Ecological consequences of invasive lake trout on river otters in Yellowstone National Park. Biol. Conserv. 143, 1144–1153. doi: 10.1016/j.biocon.2010.02.012
Wickham H. (2016). ggplot2: elegant graphics for data analysis (Cham, Switzerland: Springer-Verlag New York). Available online at: https://ggplot2.tidyverse.org, ISBN (Accessed February 21, 2025).
Keywords: diet, trophic interactions, habitat, metabarcode, scat, non-invasive sampling
Citation: Wise CM, Serfass TL, Palmer EM, Yisrael L, Burton-Reeder K and Pagenkopp Lohan KM (2025) North American river otters consume diverse prey and parasites in a subestuary of the Chesapeake Bay. Front. Mamm. Sci. 4:1620318. doi: 10.3389/fmamm.2025.1620318
Received: 29 April 2025; Accepted: 30 June 2025;
Published: 14 August 2025.
Edited by:
Kyle Shanebeck, University of California, Santa Barbara, United StatesReviewed by:
J. Andrew DeWoody, Purdue University, United StatesRobert Klaver, United States Department of the Interior, United States
Copyright © 2025 Wise, Serfass, Palmer, Yisrael, Burton-Reeder and Pagenkopp Lohan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Katrina M. Pagenkopp Lohan, bG9oYW5rQHNpLmVkdQ==
 Calli M. Wise
Calli M. Wise Thomas L. Serfass
Thomas L. Serfass Emma M. Palmer
Emma M. Palmer Leone Yisrael
Leone Yisrael Karson Burton-Reeder
Karson Burton-Reeder Katrina M. Pagenkopp Lohan
Katrina M. Pagenkopp Lohan