- Centro de Investigación Genética y Genómica, Facultad de Ciencias de la Salud Eugenio Espejo, Universidad UTE, Quito, Ecuador
Papillary thyroid cancer accounts for 85% of thyroid cancer. The diagnosis is based on ultrasound methods and tumor biopsies (FNA). In recent years, research has revealed the importance of miRNAs, non-coding RNA molecules that regulate gene expression and are involved in many diseases. The present mini review describes upregulated and downregulated miRNAs expression in papillary thyroid cancer patient samples (tissue, serum, plasma) and the genes regulated by these non-coding molecules. In addition, a bibliographic search was performed to identify the expression of miRNAs that are common in tumor tissue and blood. The miRNAs miR-146b, miR-221-3p, miRNA 222, miR-21, miR-296-5p, and miR-145 are common in both tissue and bloodstream of PTC patient samples. Furthermore, these miRNAs regulate genes involved in biological processes such as cell differentiation, proliferation, migration, invasion, and apoptosis. In conclusion, miRNAs could potentially become valuable biomarkers, which could help in the early diagnosis and prognosis of papillary thyroid cancer.
Introduction
Thyroid cancer is one of the most frequent neoplasms, the incidence of which increases yearly. It is estimated that 586,202 new cases of thyroid cancer were diagnosed until 2020 worldwide (1). Papillary thyroid cancer (PTC) is the most common type and accounts for 85% of these cases (2). Patients diagnosed with PTC have a good prognosis; however, 10% develop metastases. The number of patients diagnosed with PTC is increasing due to improvements in diagnostic methods (3).
One of the conventional diagnostic methods is ultrasound, which detects thyroid nodules in approximately 68% of PTC patients (4). Another diagnostic method is an ultrasound-guided fine needle aspiration biopsy (FNA), which has limitations and may lead to inconclusive results (5). Therefore, establishing biomarkers that could contribute to an accurate diagnosis is essential.
microRNAs (miRNAs) are small non-coding molecules of approximately 19 to 24 nucleotides related to gene regulation. Alteration of some of these miRNAs has been associated with the expression of pathological features in PTC and gene dysregulation in PTC tumor-promoting cells (4). Hence, obtaining specific miRNA expression patterns may improve the patient’s diagnosis using less invasive methods (6).
This review aims to describe the miRNAs involved in PTC from different samples, such as blood and tumor tissue, identifying common miRNAs in both samples that could be used as biomarkers.
The role of microRNAs in the oncogenesis of PTC
The miRNAs are small molecules involved in post-transcriptional gene expression processes (7). These molecules can bind and regulate several target messenger RNAs (mRNAs); one mRNA molecule can be regulated by different miRNAs (8). miRNAs are involved in fundamental processes such as cell signaling and homeostasis; control of these processes prevents uncontrolled cell proliferation, modulates cell differentiation, and regulates mRNAs in response to physiological processes (9).
In vivo and in vitro analyses have revealed the role of miRNAs in cancer. The miRNAs can be described as oncomiRNAs, decreasing the expression of tumor suppressor genes (10), or as suppressor miRNAs, reducing the expression of oncogenes or inducing apoptosis (8).
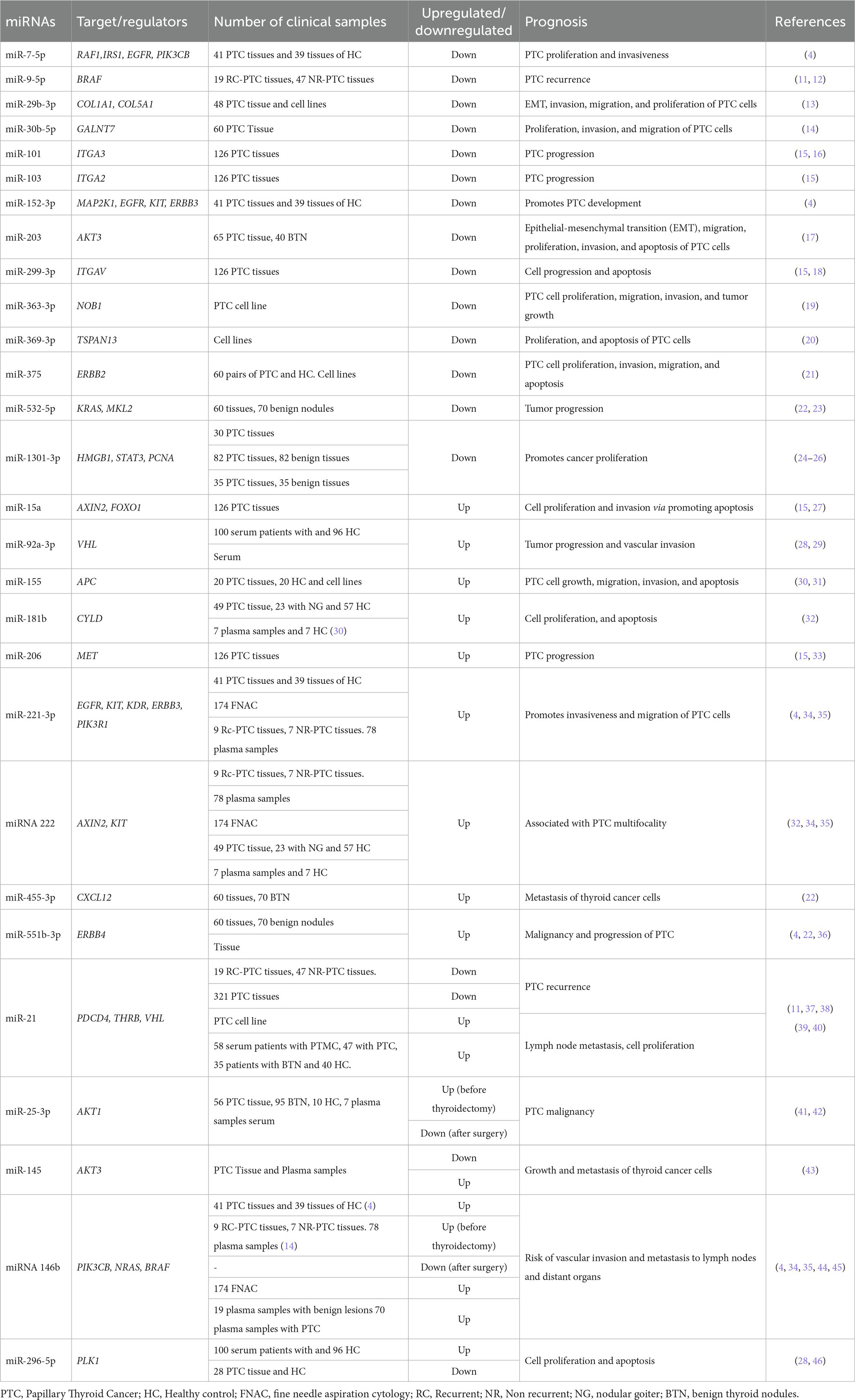
In papillary thyroid cancer, oncogenic and suppressor miRNAs have been described as being upregulated and downregulated. Moreover, different miRNAs may regulate the same genes related to biological processes such as angiogenesis, proliferation, and apoptosis in PTC (Table 1).
Analysis of microRNAs in tumoral tissue of PTC
Papillary thyroid cancer can be diagnosed with an ultrasound, FNA, and cytology testing. However, it is sometimes impossible to differentiate between malignant and benign tissue at a preoperative stage (4). Some investigations have revealed that the expression of specific miRNAs can be associated with characteristics such as tumor size, capsular and vascular invasion, and tumor aggressiveness (47). Rogucki and colleagues have shown that a panel of four miRNAs (miR-152-3p, miR-221-3p, miR-551b-3p, and miR-7-5p) could be used for the diagnosis of PTC. Furthermore, miR-152-3p, miR-221-3p, and miR-7-5p target EGFR gene regulation (4). The EGFR gene is a tyrosine kinase involved in cell proliferation, and mutations in this gene are associated with PTC tumor cell development (48). Qiao et al. revealed that miR-1301-3p and miR-532-5p are downregulated, whereas miR-551b-3p and miR-455-3p are upregulated in PTC patients compared with benign nodules and suggested that the four miRNAs have diagnostic value (22). The decrease in the expression of miR-1301-3p has been associated with the overexpression of the PCNA gene. The PCNA gene regulates the cell proliferation cycle (DNA transcription, synthesis, and repair), and studies reveal that overexpression of this gene promotes the differentiation and progression of various types of tumors (24). In another study, Wang et al. showed that overexpression of miR-551b, a regulator of the gene ERBB4, may predict poor prognosis in patients with PTC (36). For instance, ERBB4 could be downregulated in thyroid tumors; hence, this miRNA could be considered a novel biomarker (49).
In a group of 126 tissue samples from patients with PTC, it was observed that miR-15a was overexpressed, and its target gene FOX01 decreased in expression (15). FOXO1 is a transcription factor that regulates proliferation, differentiation, and response to cellular oxidative stress (50). The difference in its expression could trigger the development of tumor cells.
Integrin proteins are molecules involved in cell–cell and cell-extracellular matrix binding. Some integrin genes ITGAV, ITGA3, and ITGA2 are regulated by miR-299-3p, miR-101, and miR-103, respectively (Table 1). Studies mention that integrins play an important role in tumor progression and metastasis (15). Mautone et al. mentioned that higher ITGA3 expression is associated with a higher risk of tumor recurrence, advanced stage of disease, and worse prognosis. In contrast, ITGA2 and ITGAV expression was associated with an intermediate risk of recurrence (51).
Moreover, in an analysis of PTC tumor samples from recurrent and non-recurrent patients, miR-9-5p was downregulated, and one of the target genes of miR-9-5p is the BRAF gene. The BRAF gene triggers processes such as extrathyroidal invasion, metastasis, and recurrent disease (52). Similarly, in another study, the authors mentioned that low expression of miR-30b-5p is associated with worse prognostic features of PTC. Furthermore, miR-30b-5p is associated with GALNT7 gene regulation, and the microRNA may trigger cell proliferation and invasion via EGFR/PI3K/AKT pathway (14).
Likewise, miR-363-3 p has been associated with tumor suppression in several types of cancer (53). In vivo and in vitro investigations have associated the low expression of miR-363-3p with the overexpression levels of NOB1 (Nin-one binding protein). NOB1 silencing modifies the cell cycle, increasing the G0/G1 phase and decreasing the S phase, inhibiting PTC growth, proliferation, and invasion (19).
miRNAs like miR-375, miR-203, miR-29b-3p, and miR-369-3p were downregulated, whereas their target genes ERBB2, AKT3, COL1A1, and TSPAN13 were overexpressed. Tumor development, metastasis, apoptosis, and cell invasion are some of the processes in which these genes are involved (13, 17, 20, 21).
Analysis of microRNAs in blood (serum/plasma) of PTC
The expression of circulating miRNAs in some types of cancer has contributed to the development of noninvasive diagnostic tests (54). Furthermore, research reveals that different circulating miRNAs expression are released by cancer cells, accumulate in the bloodstream, could transmit cell-to-cell signals, and regulate target genes (55).
Studies establish differences between the expression patterns of microRNAs in plasma and serum. Some miRNAs are more frequently observed in serum, whereas others are more recurrent in plasma (28, 56, 57). For instance, He et al. established that miR-92a-3p, miR-145-5p, and miR-155-5p are more observed deregulated in plasma samples, whereas miR-222-3p have been more frequently observed deregulated in serum samples, in 5 types of cancer (58). The differences are probably due to the intercellular trafficking of miRNAs during the coagulation process. Thus, a more detailed understanding of the coagulation process is essential to determine its impact on the spectrum of miRNAs in serum and plasma (59).
Zou et al. screened for miRNAs in the serum of 100 patients with PTC and 96 healthy controls. The investigation revealed that a group of miR-25-3p, miR-296-5p, and miR-92a-3p were overexpressed in PTC patient samples compared to controls. Moreover, miR-25-3p, miR-296-5p, and miR-92a-3p target the genes AKT1, PLK1, and VHL1, respectively (Table 1) (29, 41, 46). Furthermore, the diagnostic value was assessed by the areas under the curves (AUC), and the values were statistically significant, with values of 0.727, 0.771, and 0.862, respectively. Thus, these miRNAs may be used as a noninvasive diagnostic test (28).
Research suggests that circulating levels of tumor-suppressing miRNAs are downregulated, whereas those that promote oncogenic activation are upregulated in PTC patients (55). For instance, Lee et al. found that miR-155 was upregulated in the plasma of patients with PTC with an AUC of 0.695 (44). miRNA-155 acts as an oncogenic miRNA and targets the APC gene. The dysregulation of miR-155 causes low APC expression, activating the WNT/βcatenin signaling pathway and triggering increased tumor cell viability and growth (30).
Kondrotienė et al. compared plasma samples from patients before and after thyroidectomy. The authors found that some miRNAs like miR-21 and miR-181b were overexpressed in plasma samples before surgery, whereas, after surgery, the expression of these miRNAs decreased (32). Therefore, circulating levels could be useful to differentiate the stages of the disease. In addition, miR-181b directs the CYLD gene, which regulates processes of the NF-κB pathway, involved in inflammatory responses in tumorigenesis (60). Similarly, the VHL gene is regulated by miR-21 and miR-92a-3p. The VHL is a tumor suppressor gene that is downregulated in some types of cancer. This gene dysregulation is involved in the development of carcinomas and clinically manifests aggressive behavior (29).
Common miRNAs detected in tissue and blood on PTC
The expression of some tumor-deregulated miRNAs has also been found up or down regulated in blood, making them potential noninvasive markers for PTC patients (61).
Studies have described alterations in miRNA expression between tissue and blood samples (62). Zou et al. stated that the phenomenon might be partly due to the communication of miRNAs between the tumor, tumor microenvironment, and peripheral blood circulation. Additionally, different biological states of tumors, the influence of other non-tumor cells, or changing immune states could also alter the expression of miRNAs (28).
Lee et al. analyzed tissue and plasma samples from patients with PTC. The authors identified that in tissue samples, the expression of miR-222 was 10.8 times higher, and miR-146b was 8.9 times higher in tumor samples with recurrence (34). Furthermore, in the same study, the expression of miRNAs was analyzed in plasma samples from 42 PTC patients before and after surgery. It was found that miR-146b, miR-221, and miR-222 were significantly upregulated before and downregulated after surgery (34). Similarly, in a study of 100 PTC samples, it was shown that in tumors where the BRAF V600E mutation was detected, miR-146b was overexpressed, and it was determined in a subsequent study that this association could triggered shorter survival (63). Likewise, another study concluded that miR-146b, miR-221, and miR-222 are related to the regulation of the KIT gene. This gene is a receptor tyrosine kinase, which is involved in cell differentiation and growth. KIT plays the role of an oncogene in some types of cancer, and in PTC, its expression is lower than in normal tissues (64).
Furthermore, Boufraqech et al. analyzed the expression of miR-145 and revealed that it is downregulated in PTC tissues, whereas, in the plasma of PTC patients, it is upregulated. Furthermore, the authors demonstrated that miR-145 has tissue specific tumor suppressor functions by interacting with AKT3, which regulates the PI3K/Akt pathway. In vivo and in vitro studies show that the upregulation of miR-145 decreases the growth and metastasis of thyroid cancer cells (43).
Likewise, another study with tissue samples from 28 patients with PTC and adjacent non-tumor tissue from the same patients revealed that miR-296-5p expression in tissue was downregulated (46). In contrast, Zou et al. described that plasma miR-296-5p was overexpressed in PTC patients. Therefore, based on the different levels of miR-296-5p expression between plasma and tissue, miR-296-5p may be a biomarker for the diagnosis of thyroid cancer (Figure 1) (28).
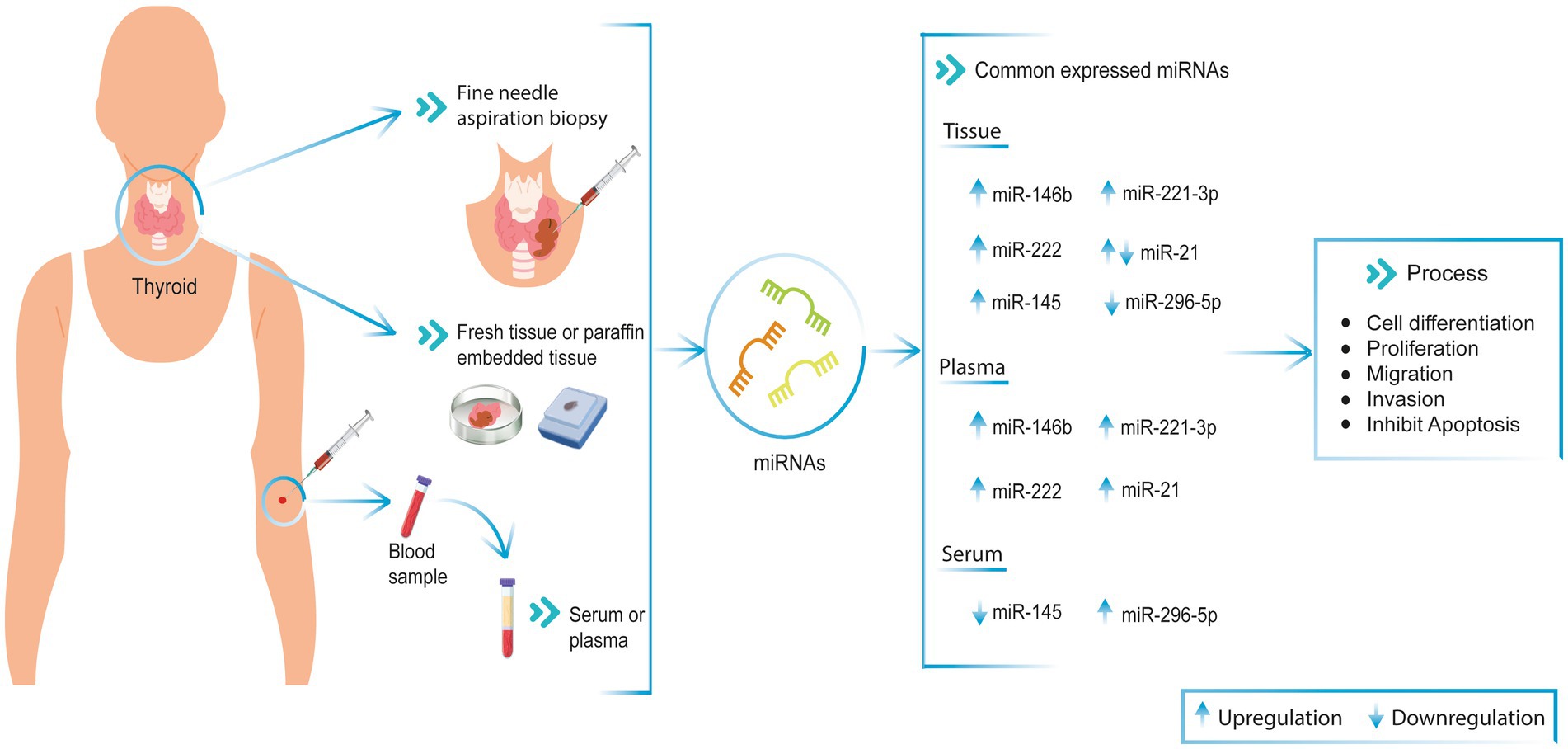
Figure 1. Commonly miRNAs expression in PTC patient’s tissue and blood (serum and plasma). These miRNAs are involved in cell differentiation, proliferation, migration, invasion, and apoptosis processes in PTC patient samples. ↑ Upregulated, ↓ Downregulated.
Diagnostic utility of microRNA in PTC
Thyroid cancer presents several tumor types with different levels of differentiation and cellular origin, such as PTC (65). Research has determined that the expression levels of miRNAs are different in normal tissues compared to tumor tissues, as the role played by miRNAs is the regulation of signaling pathways. These miRNAs have been related to cell proliferation, survival, invasion, and migration processes. In other words, miRNAs could be valuable tools for determining the cellular characteristics of tumors (66, 67).
Additionally, the expression of miRNAs can give information about the early stages of the disease by analyzing circulating miRNAs in blood (68).
The technologies used to determine the expression of miRNAs have been diverse. Early methods were based on hybridizing single or multiple miRNAs and panel development using microarray platforms. Nowadays, Next Generation Sequencing (NGS) is being applied to obtain gene expression profiles. NGS makes it possible to get information on coding and non-coding RNAs to analyze the expression of specific miRNAs (67). Despite the more accurate technologies currently available, it is still necessary to work on variables such as sample size, sample type, and sample collection conditions to obtain miRNAs. In addition, it is important to evaluate the expression levels of miRNAs in both blood and tissue that may be used as biomarkers in PTC diagnosis and prognosis.
Conclusion
The incidence of thyroid cancer has increased worldwide (69). Fine needle aspiration (FNA) biopsy is one of the conventional diagnostic methods; however, it is an invasive test, and its results could be indeterminate (70).
The use of miRNA expression has been investigated as an alternative non-invasive diagnostic method in different types of cancer. miRNA molecules are released from the tissue into the bloodstream at separate stages of the disease (10).
In this review, we collected miRNA expression data in fresh tissue, FNA samples, and blood (serum/plasma) from patients with PTC (Table 1). In addition, common miRNAs in both tissue and blood, and their relationship with genes associated with physiological processes, were described. The common miRNAs identified in this work are miR-146b, miR-221, miR-222, miR-21, miR-145, and miR-296-5p in patients with PTC. These miRNAs are involved in cell differentiation, proliferation, migration, invasion, and apoptosis processes (45, 71). Moreover, it is important to mention that some studies use miRNAs panels to differentiate between malignant and benign tissues (4, 22).
In conclusion, this literature research provided data on miRNA expression in tissue and blood (serum/plasma) that could be used as biomarkers for diagnosis, and prognosis in patients with PTC.
Author contributions
VR-P, SC-U, PG-R, and AZ: conceptualization and writing—review and editing. EP-C and RT-T: research. AZ: supervision and conceptualization. All authors contributed to the article and approved the submitted version.
Funding
The publication fee will be funded by Universidad UTE.
Acknowledgments
We are grateful to the Universidad UTE for supporting the researchers.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
1. Sung, H, Ferlay, J, Siegel, RL, Laversanne, M, Soerjomataram, I, Jemal, A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. (2021) 71:209–49. doi: 10.3322/caac.21660
2. Zhang, D, Tao, L, Xu, N, Lu, X, Wang, J, He, G, et al. CircRNA circTIAM1 promotes papillary thyroid cancer progression through the miR-646/HNRNPA1 signaling pathway. Cell Death Dis. (2022) 8:21. doi: 10.1038/s41420-021-00798-1
3. Celano, M, Rosignolo, F, Maggisano, V, Pecce, V, Iannone, M, Russo, D, et al. MicroRNAs as biomarkers in thyroid carcinoma. Int J Genomics. (2017) 2017:1–11. doi: 10.1155/2017/6496570
4. Rogucki, M, Sidorkiewicz, I, Niemira, M, Dzięcioł, JB, Buczyńska, A, Adamska, A, et al. Expression profile and diagnostic significance of MicroRNAs in papillary thyroid cancer. Cancers. (2022) 14:2679. doi: 10.3390/cancers14112679
5. Papaioannou, M, Chorti, AG, Chatzikyriakidou, A, Giannoulis, K, Bakkar, S, and Papavramidis, TS. MicroRNAs in papillary thyroid cancer: What is new in diagnosis and treatment. Front Oncol. (2022) 11:755097. doi: 10.3389/fonc.2021.755097
6. Toraih, EA, Fawzy, MS, Ning, B, Zerfaoui, M, Errami, Y, Ruiz, EM, et al. A miRNA-based prognostic model to trace thyroid cancer recurrence. Cancers. (2022) 14:4128. doi: 10.3390/cancers14174128
7. Vieira Geraldo, M, Imoto Nakaya, H, and Teruko, KE. Down-regulation of 14q32-encoded miRNAs and tumor suppressor role for miR-654-3p in papillary thyroid cancer [internet]. Oncotarget. (2017) 8:9597–607. doi: 10.18632/oncotarget.14162
8. Smolarz, B, Durczyński, A, Romanowicz, H, Szyłło, K, and Hogendorf, P. miRNAs in cancer (review of literature). Int J Mol Sci. (2022) 23:2805. doi: 10.3390/ijms23052805
9. Galvão-Lima, LJ, Morais, AHF, Valentim, RAM, and Barreto, EJSS. miRNAs as biomarkers for early cancer detection and their application in the development of new diagnostic tools. Biomed Eng Online. (2021) 20:21–07. doi: 10.1186/s12938-021-00857-9
10. Ghafouri-Fard, S, Shirvani-Farsani, Z, and Taheri, M. The role of microRNAs in the pathogenesis of thyroid cancer. Noncoding RNA Res. (2020) 5:88–98. doi: 10.1016/j.ncrna.2020.06.001
11. Sondermann, A, Andreghetto, FM, Moulatlet, ACB, da Silva, VE, de Castro, MG, Nunes, FD, et al. MiR-9 and miR-21 as prognostic biomarkers for recurrence in papillary thyroid cancer. Clin Exp Metastasis. (2015) 32:521–30. doi: 10.1007/s10585-015-9724-3
12. Yang, F, Zhang, J, Li, B, Zhao, Z, Liu, Y, Zhao, Z, et al. Identification of potential lncRNAs and miRNAs as diagnostic biomarkers for papillary thyroid carcinoma based on machine learning. Int J Endocrinol. (2021) 2021:1–15. doi: 10.1155/2021/3984463
13. Wang, C, Wang, Y, Fu, Z, Huang, W, Yu, Z, Wang, J, et al. MiR-29b-3p inhibits migration and invasion of papillary thyroid carcinoma by downregulating COL1A1 and COL5A1. Front Oncol. (2022) 12:1–14. doi: 10.3389/fonc.2022.837581
14. Wang, Y, Wang, C, Fu, Z, Zhang, S, and Chen, J. miR-30b-5p inhibits proliferation, invasion, and migration of papillary thyroid cancer by targeting GALNT7 via the EGFR/PI3K/AKT pathway. Cancer Cell Int. (2021) 21:618–7. doi: 10.1186/s12935-021-02323-x
15. Liu, X, He, M, Hou, Y, Liang, B, Zhao, L, Ma, S, et al. Expression profiles of microRNAs and their target genes in papillary thyroid carcinoma. Oncol Rep. (2013) 29:1415–20. doi: 10.3892/or.2013.2263
16. Lin, X, Guan, H, Li, HAI, Liu, L, and Liu, J. miR—01 inhibits cell proliferation by targeting Rac1 in papillary thyroid carcinoma. Biomed Rep. (2014) 2:122–6. doi: 10.3892/br.2013.192
17. You, A, Fu, L, Li, Y, Li, X, and You, B. MicroRNA-203 restrains epithelial–mesenchymal transition, invasion and migration of papillary thyroid cancer by downregulating AKT3. Cell Cycle. (2020) 19:1105–21. doi: 10.1080/15384101.2020.1746490
18. Wang, Z, Liu, Q, Huang, P, and Cai, G. miR-299-3p suppresses cell progression and induces apoptosis by downregulating PAX3 in gastric cancer. Open Life Sci. (2021) 16:266–76. doi: 10.1515/biol-2021-0022
19. Dong, S, Xue, S, Sun, Y, Han, Z, Sun, L, Xu, J, et al. MicroRNA-363-3p downregulation in papillary thyroid cancer inhibits tumor progression by targeting NOB1. J Investig Med. (2021) 69:66–74. doi: 10.1136/jim-2020-001562
20. Li, P, Dong, M, and Wang, Z. Downregulation of TSPAN13 by miR-369-3p inhibits cell proliferation in papillary thyroid cancer (PTC). Bosn J Basic Med Sci. (2019) 19:146–54. doi: 10.17305/bjbms.2018.2865
21. Wang, XZ, Hang, YK, Liu, JB, Hou, YQ, Wang, N, and Wang, MJ. Over-expression of microRNA-375 inhibits papillary thyroid carcinoma cell proliferation and induces cell apoptosis by targeting ERBB2. J Pharmacol Sci. (2016) 130:78–84. doi: 10.1016/j.jphs.2015.12.001
22. Qiao, D, Hui, H, Mei, X, Deng, X, Chi, JY, Yang, H, et al. Aberrant expression of five miRNAs in papillary thyroid carcinomas. J Clin Lab Anal. (2021) 35:e23907. doi: 10.1002/jcla.23907
23. Griesing, S, Kajino, T, Tai, MC, Liu, Z, Nakatochi, M, Shimada, Y, et al. Thyroid transcription factor-1-regulated microRNA-532-5p targets KRAS and MKL2 oncogenes and induces apoptosis in lung adenocarcinoma cells. Cancer Sci. (2017) 108:1394–404. doi: 10.1111/cas.13271
24. Qiao, D, He, X, Yang, H, Zhou, Y, Deng, X, Cheng, L, et al. miR-1301-3p suppresses tumor growth by downregulating PCNA in thyroid papillary cancer. Am J Otolaryngol Head Neck Med Surg. (2021) 42:102920. doi: 10.1016/j.amjoto.2021.102920
25. Dong, LP, Chen, LY, Bai, B, Qi, XF, Liu, JN, and Qin, S. circ_0067934 promotes the progression of papillary thyroid carcinoma cells through miR-1301-3p/HMGB1 axis. Neoplasma. (2022) 69:1–15. doi: 10.4149/neo_2021_210608N771
26. Wen, J, Wang, H, Dong, T, Gan, P, Fang, H, Wu, S, et al. STAT3-induced upregulation of lncRNA ABHD11-AS1 promotes tumour progression in papillary thyroid carcinoma by regulating miR-1301-3p/STAT3 axis and PI3K/AKT signalling pathway. Cell Prolif. (2019) 52:e12569. doi: 10.1111/cpr.12569
27. Jin, J, Zhang, J, Xue, Y, Luo, L, Wang, S, and Tian, H. MiRNA-15a regulates the proliferation and apoptosis of papillary thyroid carcinoma via regulating AKT pathway. Onco Targets Ther. (2019) 12:6217–26. doi: 10.2147/OTT.S213210
28. Zou, X, Gao, F, Wang, ZY, Zhang, H, Liu, QX, Jiang, L, et al. A three-microRNA panel in serum as novel biomarker for papillary thyroid carcinoma diagnosis. Chin Med J. (2020) 133:2543–51. doi: 10.1097/CM9.0000000000001107
29. Todorović, L, and Stanojević, B. VHL tumor suppressor as a novel potential candidate biomarker in papillary thyroid carcinoma. Bosn J Basic Med Sci. (2022) 1:26–36. doi: 10.17305/bjbms.2022.7850
30. Zhang, X, Li, M, Zuo, K, Li, D, Ye, M, Ding, L, et al. Upregulated miR-155 in papillary thyroid carcinoma promotes tumor growth by targeting APC and activating Wnt/β-catenin signaling. J Clin Endocrinol Metab. (2013) 98:E1305–13. doi: 10.1210/jc.2012-3602
31. Li, Y, Zeng, QG, Qiu, JL, Pang, T, Wang, H, and Zhang, XX. Sevoflurane inhibits the progression of PTC by downregulating miR-155. Eur Rev Med Pharmacol Sci. (2019) 12:6579–87. doi: 10.26355/eurrev_201908_18544
32. Kondrotienė, A, Daukša, A, Pamedytytė, D, Kazokaitė, M, Žvirblienė, A, Daukšienė, D, et al. Plasma-derived miRNA-222 as a candidate marker for papillary thyroid cancer. Int J Mol Sci. (2020) 21:1–16. doi: 10.3390/ijms21176445
33. Liu, F, Yin, R, Chen, X, Chen, W, Qian, Y, and Zhao, Y. Biomedicine & pharmacotherapy over-expression of miR-206 decreases the Euthyrox-resistance by targeting MAP4K3 in papillary thyroid carcinoma. Biomed Pharmacother. (2019) 114:108605. doi: 10.1016/j.biopha.2019.108605
34. Lee, JC, Zhao, JT, Clifton-Bligh, RJ, Gill, A, Gundara, JS, Ip, JC, et al. MicroRNA-222 and MicroRNA-146b are tissue and circulating biomarkers of recurrent papillary thyroid cancer. Cancer. (2013) 119:4358–65. doi: 10.1002/cncr.28254
35. Castagna, MG, Marzocchi, C, Pilli, T, Forleo, R, Pacini, F, and Cantara, S. MicroRNA expression profile of thyroid nodules in fine-needle aspiration cytology: a confirmatory series. J Endocrinol Investig. (2019) 42:97–100. doi: 10.1007/s40618-018-0880-6
36. Wang, J, and Liu, H. miR-551b is associated with the poor prognosis and malignant development of papillary thyroid cancer through regulating ERBB4. Horm Metab. (2022) 54:113–8. doi: 10.1055/a-1735-3318
37. Stokowy, T, Gawel, D, and Wojtas, B. Differences in miRNA and mRNA profile of papillary thyroid cancer variants. Int. J Endocrinol. (2016) 2016:1–10. doi: 10.1155/2016/1427042
38. Zhang, J, Yang, Y, Liu, Y, Fan, Y, Liu, Z, Wang, X, et al. MicroRNA-21 regulates biological behaviors in papillary thyroid carcinoma by targeting programmed cell death 4. J Surg Res. (2014) 189:68–74. doi: 10.1016/j.jss.2014.02.012
39. Zang, C, Sun, J, Liu, W, Chu, C, Jiang, L, and Ge, R. miRNA-21 promotes cell proliferation and invasion via VHL/PI3K/AKT in papillary thyroid carcinoma. Hum Cell. (2019) 32:428–36. doi: 10.1007/s13577-019-00254-4
40. Zhang, Y, Pan, J, Xu, D, Yang, Z, Sun, J, Sun, L, et al. Combination of serum microRNAs and ultrasound profile as predictive biomarkers of diagnosis and prognosis for papillary thyroid microcarcinoma. Oncol Rep. (2018) 40:3611–24. doi: 10.3892/or.2018.6776
41. Zhang, F, Chen, K, Tao, H, Kang, T, Xiong, Q, Zeng, Q, et al. miR-25-3p, positively regulated by transcription factor AP-2α, regulates the metabolism of C2C12 cells by targeting Akt1. Int J Mol Sci. (2018) 19:1–13. doi: 10.3390/ijms19030773
42. Wojcicka, A, Kolanowska, M, and Jazdzewski, K. MicroRNA in diagnostics and therapy of thyroid cancer. Eur J Endocrinol. (2016) 174:R89–98. doi: 10.1530/EJE-15-0647
43. Boufraqech, M, Zhang, L, Jain, M, Patel, D, Ellis, R, Xiong, Y, et al. MiR-145 suppresses thyroid cancer growth and metastasis and targets AKT3. Endocr Relat Cancer. (2014) 21:517–31. doi: 10.1530/ERC-14-0077
44. Lee, YS, Lim, YS, Lee, JC, Wang, SG, Park, HY, Kim, SY, et al. Differential expression levels of plasma-derived miR-146b and miR-155 in papillary thyroid cancer. Oral Oncol. (2015) 51:77–83. doi: 10.1016/j.oraloncology.2014.10.006
45. Santiago, K, Chen Wongworawat, Y, and Khan, S. Differential MicroRNA-signatures in thyroid cancer subtypes. J Oncol. (2020) 2020:1–14. doi: 10.1155/2020/2052396
46. Zhou, S.-L., Tang, Q.-L, and Zhou, S.-X. MiR-296-5p suppresses papillary thyroid carcinoma cell growth via targeting PLK1. (2019). Available at: http://pictar.mdc-berlin.de/
47. Zembska, A, Jawiarczyk-Przybyłowska, A, Wojtczak, B, and Bolanowski, M. MicroRNA expression in the progression and aggressiveness of papillary thyroid carcinoma. Anticancer Res. (2019) 39:33–40. doi: 10.21873/anticanres.13077
48. Masago, K, Asato, R, Fujita, S, Hirano, S, Tamura, Y, Kanda, T, et al. Epidermal growth factor receptor gene mutations in papillary thyroid carcinoma. Int J Cancer. (2009) 124:2744–9. doi: 10.1002/ijc.24250
49. Hu, X, Xu, H, Xue, Q, Wen, R, Jiao, W, and Tian, K. The role of ERBB4 mutations in the prognosis of advanced non-small cell lung cancer treated with immune checkpoint inhibitors. Mol Med. (2021) 27:126. doi: 10.1186/s10020-021-00387-z
50. Shantanam, S, and Mueller, A. FOXO1: a potential target for human diseases. Physiol Behav. (2018) 176:139–48. doi: 10.2174/138945011796150280
51. Mautone, L, Ferravante, C, Tortora, A, Tarallo, R, Giurato, G, and Weisz, A. Higher integrin alpha 3 beta1 expression in papillary thyroid cancer is associated with worst outcome. Cancers. (2021) 13:1–15. doi: 10.3390/cancers13122937
52. Braun, J, Hoang-Vu, C, Dralle, H, and Hüttelmaier, S. Downregulation of microRNAs directs the EMT and invasive potential of anaplastic thyroid carcinomas. Oncogene. (2010) 29:4237–44. doi: 10.1038/onc.2010.169
53. Zhang, R, Li, Y, and Dong, X. MiR-363 sensitizes cisplatin-induced apoptosis targeting in Mcl-1 in breast cancer. Med Oncol. (2014) 31:347. doi: 10.1007/s12032-014-0347-3
54. Filipów, S, and Łaczmański, Ł. Blood circulating miRNAs as cancer biomarkers for diagnosis and surgical treatment response. Front Neurosci. (2019) 10:169. doi: 10.3389/fgene.2019.00169
55. Chen, G, Wang, J, and Cui, Q. Could circulating miRNAs contribute to cancer therapy? Trends Mol Med. (2013) 19:71–3. doi: 10.1016/j.molmed.2012.10.006
56. Dufourd, T, Robil, N, Mallet, D, Carcenac, C, Boulet, S, Brishoual, S, et al. Plasma or serum? A qualitative study on rodents and humans using high-throughput microRNA sequencing for circulating biomarkers. Biol Methods Protoc. (2019) 4:1–10. doi: 10.1093/biomethods/bpz006
57. Foye, C, Yan, IK, David, W, Shukla, N, Habboush, Y, Chase, L, et al. Comparison of miRNA quantitation by Nanostring in serum and plasma samples. PLoS One. (2017) 12:1–13. doi: 10.1371/journal.pone.0189165
58. He, Y, Lin, J, Kong, D, Huang, M, Xu, C, Kim, TK, et al. Current state of circulating microRNAs as cancer biomarkers. Clin Chem. (2015) 61:1138–55. doi: 10.1373/clinchem.2015.241190
59. Wang, K, Yuan, Y, Cho, JH, McClarty, S, Baxter, D, and Galas, DJ. Comparing the MicroRNA spectrum between serum and plasma. PLoS One. (2012) 7:e41561. doi: 10.1371/journal.pone.0041561
60. Li, D, Jian, W, Wei, C, Song, H, Gu, Y, Luo, Y, et al. Down-regulation of miR-181b promotes apoptosis by targeting CYLD in thyroid papillary cancer. Int J Clin Exp Pathol. (2014) 7:7672–80.
61. Yu, S, Liu, Y, Wang, J, Guo, Z, and Zhang, Q. Circulating MicroRNA profiles as potential biomarkers for diagnosis of papillary thyroid carcinoma. J Clin Endocrinol Metab. (2012) 97:2084–92. doi: 10.1210/jc.2011-3059
62. Nagy, ZB, Barták, BK, Kalmár, A, Galamb, O, Wichmann, B, Dank, M, et al. Comparison of circulating miRNAs expression alterations in matched tissue and plasma samples during colorectal cancer progression. Pathol Oncol Res. (2019) 25:97–105. doi: 10.1007/s12253-017-0308-1
63. Lee, JC, Gundara, JS, Glover, A, Serpell, J, and Sidhu, SB. MicroRNA expression profiles in the Management of Papillary Thyroid Cancer. Oncologist. (2014) 19:1141–7. doi: 10.1634/theoncologist.2014-0135
64. He, H, Jazdzewski, K, Li, W, Liyanarachchi, S, Nagy, R, Volinia, S, et al. The role of microRNA genes in papillary thyroid carcinoma. (2005). Available at: www.pnas.orgcgidoi10.1073pnas.0509603102
65. Hu, J, Yuan, IJ, Mirshahidi, S, Simental, A, Lee, SC, and Yuan, X. Thyroid carcinoma: phenotypic features, underlying biology and potential relevance for targeting therapy. Int J Mol Sci. (2021) 22:1–25. doi: 10.3390/ijms22041950
66. Nikiforova, MN, Tseng, GC, Steward, D, Diorio, D, and Nikiforov, YE. MicroRNA expression profiling of thyroid tumors: biological significance and diagnostic utility. J Clin Endocrinol Metab. (2008) 93:1600–8. doi: 10.1210/jc.2007-2696
67. Park, JL, Kim, SK, Jeon, S, Jung, CK, and Kim, YS. Microrna profile for diagnostic and prognostic biomarkers in thyroid cancer. Cancers. (2021) 13:1–19. doi: 10.3390/cancers13040632
68. De La Chapelle, A, and Jazdzewski, K. MicroRNAs in thyroid cancer. J Clin Endocrinol Metab. (2011) 96:3326–36. doi: 10.1210/jc.2011-1004
69. Medina, EGJ, Viúdez, JCSA, Porras, EGI, Ramón, T, and Trigo, CJ. SEOM clinical guideline thyroid cancer (2019). Clin Transl Oncol. (2020) 22:223–35. doi: 10.1007/s12094-019-02284-8
70. Pantanowitz, L, Thompson, LDR, Jing, X, and Rossi, ED. Is thyroid core needle biopsy a valid compliment to fine-needle aspiration? J Am Soc Cytopathol. (2020) 9:383–8. doi: 10.1016/j.jasc.2020.06.003
Keywords: papillary thyroid cancer, miRNAs, diagnosis, biomarkers, tumorigenesis
Citation: Ruiz-Pozo VA, Cadena-Ullauri S, Guevara-Ramírez P, Paz-Cruz E, Tamayo-Trujillo R and Zambrano AK (2023) Differential microRNA expression for diagnosis and prognosis of papillary thyroid cancer. Front. Med. 10:1139362. doi: 10.3389/fmed.2023.1139362
Edited by:
Shanthi Sabarimurugan, University of Western Australia, AustraliaReviewed by:
Dorota Pastuszak-Lewandoska, Medical University of Lodz, PolandCopyright © 2023 Ruiz-Pozo, Cadena-Ullauri, Guevara-Ramírez, Paz-Cruz, Tamayo-Trujillo and Zambrano. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ana Karina Zambrano, YW5hemFtYnJhbm8xN0Bob3RtYWlsLmNvbQ==
 Viviana A. Ruiz-Pozo
Viviana A. Ruiz-Pozo Santiago Cadena-Ullauri
Santiago Cadena-Ullauri Patricia Guevara-Ramírez
Patricia Guevara-Ramírez Elius Paz-Cruz
Elius Paz-Cruz Rafael Tamayo-Trujillo
Rafael Tamayo-Trujillo Ana Karina Zambrano
Ana Karina Zambrano