Abstract
Background:
Aeromonas dhakensis is a gram-negative bacterium. In recent years, Aeromonas dhakensis has gradually attracted increasing attention due to its strong virulence and poor prognosis. Clinical reports of pulmonary infection caused by Aeromonas dhakensis are rare.
Case presentation:
A patient with acute T lymphoblastic leukemia experienced myelosuppression after chemotherapy, developed a secondary pulmonary infection with Aeromonas dhakensis and was hospitalized due to fever. The patient underwent testing for inflammatory markers, chest imaging, blood culture, bronchoalveolar lavage, pleural drainage, and metagenomic next-generation sequencing of alveolar lavage fluid and pleural fluid to obtain evidence of Aeromonas dhakensis infection, and was treated with four generations of cephalosporin combined with fluoroquinolone antibiotics. The patient’s condition significantly improved.
Discussion:
Among pulmonary infectious pathogens, Aeromonas dhakensis is relatively rare. Once an Aeromonas strain is cultured in the clinical work, pathogenic sequencing should be performed on the detected samples for early accurate diagnosis and effective anti-infection treatment.
1 Introduction
Aeromonas dhakensis is an emerging opportunistic pathogen. It was first found in the feces of children with diarrhea in Dhaka, Bangladesh, and described in 2002 (1). It is present in polluted water, fish and food. It often occurs in people with poor immune function, malignant tumors or biliary tract infections and can also be seen in healthy people who are in contact with polluted water. Aeromonas dhakensis can cause severe infection in the gastrointestinal tract, skin and soft tissues, even throughout the global body. Due to the complex classification of Aeromonas, routine laboratory tests cannot accurately identify Aeromonas species, and the epidemiology of Aeromonas dhakensis is underestimated in the clinic (2). Currently, Aeromonas dhakensis is considered the second most common pathogen of Aeromonas species and the most virulent (3–5). We report a case of pulmonary infection with Aeromonas dhakensis secondary to chemotherapy in a patient with acute T lymphoblastic leukemia. This infection was cured with standard antibiotics. This paper also reviews the treatment and prognosis of pulmonary infection with Aeromonas dhakensis and provides experience for clinicians in diagnosing and treating this disease.
2 Case report
An 18-year-old female patient was diagnosed with acute T-lymphoblastic leukemia more than 8 months earlier. She was started on vincristine, daunorubicin, cyclophosphamide, prednisone, and asparaginase (VDCLP) chemotherapy and leucocyte elevation therapy. Two days later, the patient developed fever, with a fluctuating body temperature and a highest temperature of 39°C. She was referred to our hospital the next day.
The admission test results were as follows: hemoglobin, 93 g/L (reference range: 115–150 g/L); platelet count, 74 × 109/L (reference range: 125–350 × 109/L); white blood cell count, 0.06 × 109/L (reference range: 3.5–9.5 × 109/L); neutrophils, not detected (reference range: 1.8–6.3 × 109/L); lymphocytes, not detected (reference range: 1.1–3.2 × 109/L); C-reactive protein (CRP), 25.32 mg/L (reference range: 0–10 mg/L); procalcitonin (PCT), 15 ng/L (reference range: < 0.05 ng/L); serum galactomannan (GM) test, negative; fungal detection (G) test, 86.2 pg/mL (reference range: < 60 pg/mL: negative, > 100 pg/mL: positive); cytokines: interleukin-6: 29.73 pg/mL (reference range: ≤5.4 pg/mL); interleukin-4: 9.03 pg/mL (reference range: ≤8.56 pg/mL); interferon-α: 49.3 pg/mL (reference range: ≤23.1 pg/mL); Lymphocyte immunoassay: CD19 + B lymphocytes: 0.27% (reference range: 6.58–24.52%); CD3 + T lymphocytes: 90.47% (reference range: 55.62–84.84%); CD3+/CD8+ toxic T lymphocytes: 49.69% (reference range: 13.27–40.63%); (CD3-CD16+/56+) NK cells: 4.31% (reference range: 5.15–27.08%); absolute number of lymphocytes: 37.6/μL (reference range: 1530–3,700/μL); absolute number of total T lymphocytes: 33.6/μL (reference range: 955–2,860/μL); absolute number of CD4+ helper T lymphocytes: 11.9/μL (reference range: 550–1,440/μL); absolute number of CD8+ toxic T lymphocytes: 17.3/μL (reference range: 320–1,250/μL); absolute number of B lymphocytes: 0.101/μL (reference range: 90–560/μL); and absolute number of NK cells: 1.72/μL (reference range: 150–1,100/μL). Acid-fast staining of the sputum smear was negative; fungal examination of the sputum smear was negative. Gram staining and microscopic examination of the sputum smear showed gram-positive cocci and gram-negative bacilli. Combined with the fever symptoms and laboratory results, the patient’s immune status was low at the time. Considering the diagnosis of “bone marrow suppression period after chemotherapy and coinfection,” anti-infection treatment made up of Biapenem 300 mg q8h as an intravenous infusion, vancomycin 0.5 g q8h as an intravenous infusion combined with posaconazole oral suspension 200 mg tid po was given for 6 days.
However, she still had fever and developed right chest pain on the 7th day after admission. Auscultation of the right lung indicated reduced breathing sounds. Complete chest computed tomography (CT) showed streaks and clumps of high-density shadows in the middle and lower lobes of the right lung with cavity formation and a small amount of fluid in the right chest cavity and interlobar fissure (Figure 1A). Blood in the sputum appeared on the 8th day. Blood examination showed that the white blood cell count was 6.24 × 109/L, the neutrophil count was 5.98 × 109/L, the lymphocyte count was 0.07 × 109/L, CRP was 152.89 mg/L, and PCT was 1.96 ng/mL. The BD PhoenixTM Automated Microbiology System (BD Diagnostic Systems, Sparks, United States) was used for bacterial identification and drug sensitivity tests. Blood culture showed the growth of Aeromonas hydrophila. The drug sensitivity test results were as follows (Table 1): Aeromonas hydrophila was sensitive to amikacin, aztreonam, cefepime, cefotaxime, ceftazidime, chloramphenicol, ciprofloxacin, gentamicin, imipenem, levofloxacin, meropenem, piperacillin/tazobactam, and the compound sulfamide but resistant to tetracycline. Therefore, the antibiotic regimen was adjusted to Ceftazidime 2 g q8h as an intravenous infusion combined with moxifloxacin 0.4 g qd as an intravenous infusion for 13 days.
Figure 1

The picture shows the changes in the chest CT images during treatment. (A) the first CT scan in the hospital taken on 29 June 2023. The patient underwent a CT scan of the chest to observe the therapeutic effect on (B) 12 July 2023 and (C) 31 July 2023. Outpatient re-examinations using a repeat CT scan on (D) 25 August 2023, (E) 3 October 2023, and (F) 14 November 2023. The red arrow represents the inflammatory lesion, the black arrow shows the cavity, the blue arrow indicates fluid accumulation, and the green arrow represents residual striped shadows.
Table 1
| Antibiotic | MIC (ug/mL) | Sensitivity |
|---|---|---|
| Amikacin | ≤8 | Sensitive |
| Aztreonam | ≤2 | Sensitive |
| Cefepime | ≤2 | Sensitive |
| Cefotaxime | ≤1 | Sensitive |
| Ceftazidime | ≤1 | Sensitive |
| Chloramphenicol | ≤4 | Sensitive |
| Ciprofloxacin | ≤0.5 | Sensitive |
| Gentamicin | 4 | Sensitive |
| Imipenem | ≤1 | Sensitive |
| Levofloxacin | ≤1 | Sensitive |
| Meropenem | ≤1 | Sensitive |
| Piperacillin/tazobactam | ≤4/4 | Sensitive |
| Tetracycline | >8 | Resistant |
| Compound sulfamide | ≤0.5/9.5 | Sensitive |
Antibiotic susceptibility test for Aeromonas dhakensis.
On the 20th day after admission, chest CT showed increased density in the middle and lower lobes of the right lung with stripy and clump shapes, a larger cavity area than before, and increased pleural effusion on the right side (Figure 1B). Considering the worsening of the patient’s hemoptysis symptoms, chest CT indicated progressive inflammation and pleural effusion. Therefore, bronchoalveolar lavage and pleural drainage were performed to help find unknown pathogens. Conventional testing of the pleural effusion revealed the following results: color, orange yellow; transparency, turbidity, visible solidification; Rivalta test, positive; and white blood cell count, 10,154 × 106/L. The biochemistry of the pleural effusion showed the following: total protein 37.3 g/L, albumin 21.3 g/L, lactate dehydrogenase 6,537 U/L, and glucose 0.48 mmol/L. Ordinary bacterial culture of the pleural effusion detected no bacteria. Cytological staining of alveolar lavage fluid and pleural fluid via hematoxylin–eosin staining showed inflammatory cells (Figure 2). To identify potential pathogenic microorganisms, we performed metagenomic next-generation sequencing (mNGS) on the pleural effusion and bronchoalveolar lavage fluid. Pleural effusion mNGS indicated Aeromonas dhakensis (number of sequences: 43, identification confidence: 99%) (reference range: not found). Bronchoalveolar lavage fluid mNGS showed Aeromonas dhakensis (number of sequences: 101, identification confidence: 99%) (reference range: not found). According to the ABX POC-IT guidelines and published literature (6–12), Aeromonas is sensitive to fluoroquinolones and fourth-generation cephalosporins. Therefore, the antibiotic was adjusted to cefepime 2 g q8h as an intravenous infusion combined with levofloxacin 0.5 g qd as an intravenous infusion.
Figure 2
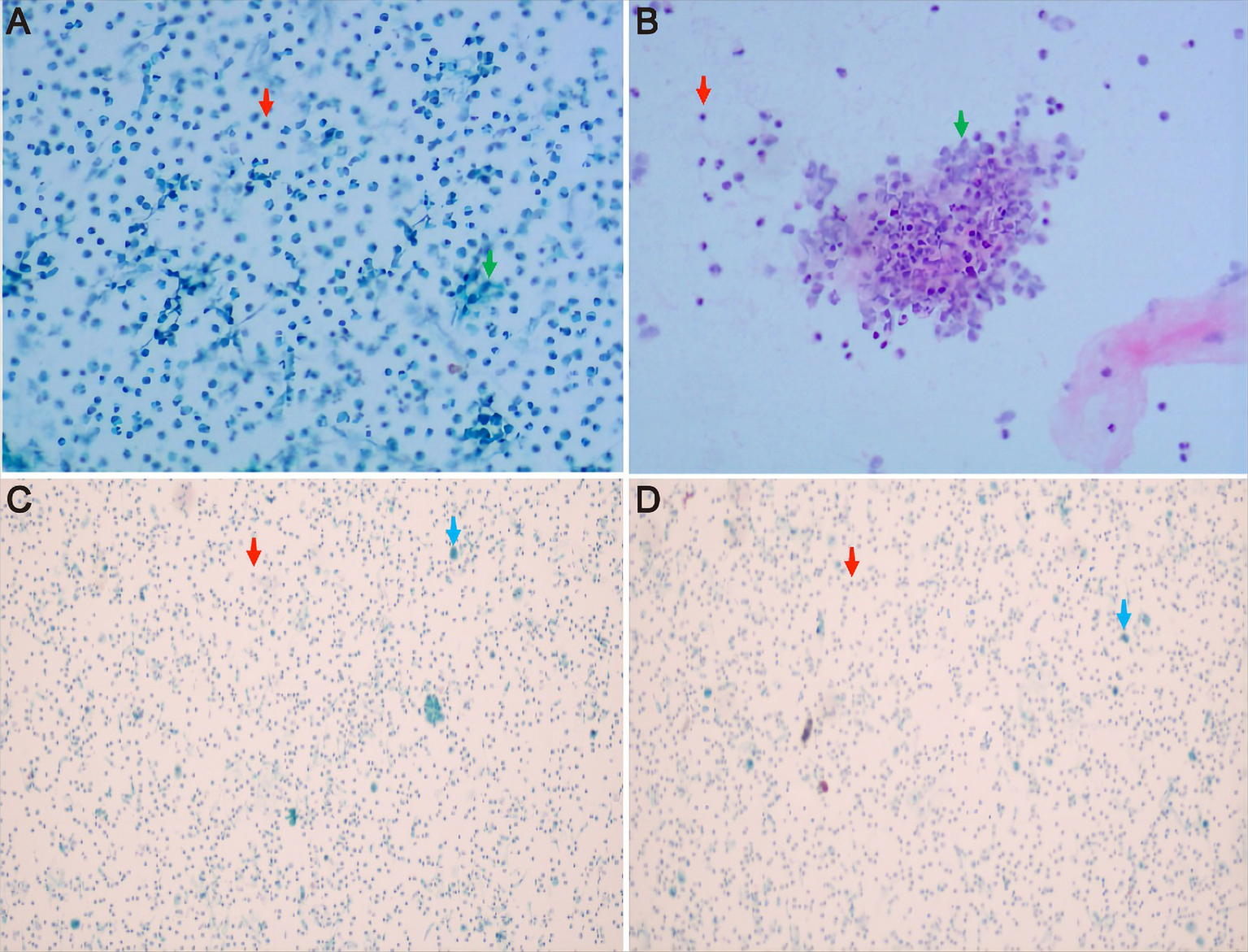
Cytological results of pleural fluid and alveolar lavage fluid by hematoxylin–eosin staining. (A,B) the pleural fluid Thinprep cytologic test showed inflammatory cells, with a small number of mesothelial cells and tissue cells, but no clear atypical tumor cells were observed. (C,D) alveolar lavage fluid under the microscope indicated ciliated columnar epithelial cells and inflammatory cells in the right middle lobe bronchial lavage fluid, but no clear atypical tumor cells were observed. The red arrow represents inflammatory cells, the green arrow shows mesothelial cells, and the blue arrow indicates ciliated columnar epithelial cells.
On the 38th day after admission, her body temperature returned to normal, and her symptoms of hemoptysis disappeared. Blood examination revealed a white blood cell count of 6.15 × 109/L, a neutrophil count of 3.55 × 109/L, a lymphocyte count of 1.81 × 109/L, CRP of 28.85 mg/L, and PCT of <0.05 ng/mL. Chest CT showed that the cord-like and flaky-like high-density shadows in the middle and lower lobes of the right lung were reduced in scope, and the pleural effusion on the right side was less than that of the previous lesions (Figure 1C). Therefore, the patient was discharged home, and the original antibiotic regimen was continued. On the 63rd day of follow-up, her white blood cell count was 3.74 × 109/L, her neutrophil count was 2.13 × 109/L, her lymphocyte count was 1.05 × 109/L, and her CRP level was 2.51 mg/L. Chest CT showed that the extent of inflammation and pleural effusion both significantly improved (Figure 1D). On the 101st day of follow-up, chest CT indicated that most of the inflammatory lesions had been absorbed (Figure 1E), so antibiotics were discontinued. On the 144th day of follow-up, chest CT showed that the inflammatory lesions had basically resolved, though a few traces remained (Figure 1F).
3 Discussion
Aeromonas infection often occurs in people who come into contact with contaminated water or food, people with liver and biliary diseases, people with blood diseases and people with weakened immunity. More than 30 phenotypes of Aeromonas have been reported (13), among which Aeromonas dhakensis can cause purulent lesions. According to the previously reported cases (Table 2), most were out-of-hospital infections, the disease progressed rapidly, the 14-day mortality rate was high, and the cause of death was mostly bacteremia or systemic infection. These reported cases were all misjudged in the early diagnostic process based on routine biochemical tests and were eventually confirmed by etiological sequencing (6, 14, 16, 18). Pulmonary infection caused by Aeromonas dhakensis has rarely been reported.
Table 2
| First author | Year | Location | Number | Age | Gender | Risk factor/comorbidity | Etiology | Position | Treatment | Mic | Prognosis |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Chen et al. (3) | 2013 | China | 7 | Exposure to environmental water | rpoD and gyrB genes sequencing: Aeromonas dhakensis | / | |||||
| 33 | Male | Flame burn | Multiple | Cefazolin, Gentamicin | / | Survival | |||||
| 45 | Male | Necrotizing fasciitis | Foot | Cefotaxime, Doxycycline | / | Survival | |||||
| 47 | Female | Trauma | Multiple | Nil | / | Survival | |||||
| 62 | Male | Necrotizing fasciitis | Foot | Cefazolin, Gentamicin | / | Survival | |||||
| 67 | Female | Surgical wound infection | Upper limb | Cephalexin | / | Survival | |||||
| 74 | Male | Necrotizing fasciitis | Lower limb | Ceftriaxone, Doxycycline | / | Survival | |||||
| 75 | Male | Cellulitis | Lower limb | Amoxicillin-clavulanic acid | / | Survival | |||||
| Chang et al. (14) | 2018 | China | 2 | Blood or/and pus culture: Aeromonas Hydrophila. rpoD gene sequencing: Aeromonas dhakensis |
/ | ||||||
| 65 | Female | Dengue, Necrotizing Fasciitis | Bloodstream, left leg | Ceftazidime | / | Death | |||||
| 75 | Male | Dengue, Necrotizing Fasciitis | Bloodstream, lower limb | / | / | Death | |||||
| Melo-Bolivar et al. (15) | 2019 | Australia | 1 | / | / | / | rpoB and gyrB genes sequencing: Aeromonas dhakensis | Necrotizing Fasciitis | / | / | Death |
| Huang et al. (16) | 2019 | China | 1 | 29 | Male | Snake headed fish, HBV | Blood culture and biochemical tests: Aeromonas hydrophila. The whole-genome sequencing: Aeromonas dhakensis | Bloodstream | Ceftriaxone Meropenem | ≤1 ≤0.25 |
Death |
| Kitagawa et al. (6) | 2019 | Japan | 4 | 57 to 90 | 2 males and 2 females | Solid cancer/Obstructive Biliary disease/Surgical History | Blood culture and biochemical tests: Aeromonas hydrophila, Aeromonas caviae, and Aeromonas jandaei. rpoD and gyrB genes sequencing: Aeromonas dhakensis | Bloodstream | / | / | 3 survived and 1 died |
| Sun et al. (17) | 2021 | China | 26 | / | / | / | Blood culture and biochemical tests: Aeromonas hydrophila. gyrB gene sequencing: Aeromonas dhakensis | Bloodstream | / | / | 12 died or treatment failed |
Previous reports of Aeromonas dhakensis infection.
The common pathogens of community-acquired pneumonia in patients with hematological malignancies were fungi, Mycobacterium tuberculosis, Pseudomonas aeruginosa and Staphylococcus aureus. Identification of pathogenic microorganisms occurs at least 24–48 h after admission, and empirical antibiotic treatment is one of the factors that leads to prolonged disease and then worsening infection (19, 20). Aeromonas is not a common opportunistic pathogen in patients with hematological malignancies. The patient in this case had no history of contact with fish or sewage, any history of travel, or any skin injuries before the onset of the disease. No significant therapeutic effect was achieved by the early empirical anti-infection treatment. After blood culture indicated the presence of Aeromonas hydrophila, we adjusted the antibiotic regimen to ceftazidime combined with moxifloxacin according to the drug sensitivity test results. Unfortunately, re-examination via chest CT indicated that the pulmonary infection had progressed. We wondered whether there was an undetected pathogen in the lungs or whether the pathogens were not fully covered by the original antibiotic regimen. To better identify the pathogen, the patient underwent bronchoalveolar lavage and pleural drainage, and the respiratory tract samples were subjected to mNGS by Vision Medicals (Guangzhou, China). Only Aeromonas dhakensis was detected in both bronchoalveolar lavage fluid and pleural effusion by mNGS, which was not consistent with the blood culture results. Traditional mNGS requires clinical presets to select DNA and/or RNA assay procedures, which may cause pathogen omission. We did mNGS with IDseq™ Ultra, which is a bidirectional enrichment technique (forward enrichment: multiple primers were designed to amplify the target pathogen; reverse enrichment: reducing the proportion of human cells or nucleic acids to the host) based on the combination of pathogen metagenomics and probe capture. It has the advantages of being comprehensive, sensitive and in depth. Some studies have shown that Aeromonas dhakensis can be misidentified as Aeromonas hydrophila by blood culture and subsequent phenotypic identification. According to the above data, the possibility of Aeromonas hydrophila and Aeromonas dhakensis coinfection cannot be ruled out. In addition, Aeromonas was not cultured from the pleural effusion, which may be due to differences in the in vitro and in vivo bacterial culture environments of the pleural effusion. Therefore, in clinical practice, when the anti-infective treatment regimen based on blood culture and corresponding drug sensitivity tests has not effectively improved the patient’s condition, it is particularly important to combine mNGS to further discover evidence of etiological infection to guide diagnosis and treatment. Both Aeromonas dhakensis and Aeromonas hydrophila are Aeromonas strains, but the former is more virulent. It is important to use antibiotics that can cover Aeromonas dhakensis and Aeromonas hydrophila. We consulted the ABX guidelines and found that Aeromonas was sensitive to aminoglycosides, fluoroquinolones, cabapenem, aztreonam, and third-and fourth-generation cephalosporins (11). Related studies have shown that second-and third-generation cephalosporins (cefuroxime, ceftriaxone, and ceftazidime) and carbapenem antibiotics (imipenem) have higher minimum inhibitory concentrations and drug resistance rates against Aeromonas dhakensis than against Aeromonas hydrophila, while both bacteria are sensitive to fourth-generation cephalosporins (cefepime) and fluoroquinolone antibiotics (levofloxacin) (3, 8, 9). With the overuse of antibiotics in agriculture, fish culture and clinical environments, the resistance of Aeromonas to drugs has increased in recent years, and the prevalence of multidrug-resistant Aeromonas has been on the rise (7, 17). Furthermore, pathogenic Aeromonas species are reported to be resistant to carbapenems due to the presence of the CphA gene, which encodes a metallo-β-lactamases that hydrolyzes carbapenems but not fourth-generation cephalosporins (9, 12, 21). Chromosomal AmpC β-lactamases which species-specifically distributes in Aeromonas strains and included AsbA1 (Aeromonas jandaei), CepH and CepS (Aeromonas hydrophila), CAV1 and MOX (Aeromonas caviae), TRU-1 (Aeromonas enteropelogenes), and AQU-1 (Aeromonas dhakensis) (9, 19, 22–26), can hydrolyze cephamycins and third-generation cephalosporins (27, 28). Based on the above data, we selected a fourth-generation cephalosporin (cefepime) combined with a fluoroquinolone drug (levofloxacin), and the patient’s symptoms and signs improved significantly. Compared with the deaths reported by Chang et al. and Huang et al. (14, 16), in which all patients received third-generation cephalosporin treatment immediately after admission but still died quickly, timely adjustment of the antibiotic to a fourth-generation cephalosporin in this case may have been critical to reduce the risk of death and improve the condition of the patient.
4 Conclusion
Aeromonas dhakensis is virulent and has potential drug resistance, which causes rapid progression of pulmonary infection. Because it is not common in the clinic, the diagnostic specificity is poor when the evidence is based only on the characteristics of symptoms and blood culture. Once the Aeromonas strain is cultured in the clinical work, the detected sample can be further subjected to mNGS for accurate diagnosis and improvement of the antibiotic regimen. Therefore, when specific antibiotics are selected for anti-infection treatment based on biological sample culture and the corresponding drug sensitivity tests, but the patient’s condition still does not improve much, adding mNGS may identify the etiology and guide the choice of anti-infection therapy.
Statements
Data availability statement
The original contributions presented in the study are included in the article/supplementary material, further inquiries can be directed to the corresponding author.
Ethics statement
The studies involving humans were approved by the Ethics Committee of Zhengzhou University People’s Hospital. The studies were conducted in accordance with the local legislation and institutional requirements. The participants provided their written informed consent to participate in this study. Written informed consent was obtained from the individual(s) for the publication of any potentially identifiable images or data included in this article.
Author contributions
CW: Conceptualization, Data curation, Formal analysis, Writing – original draft. NW: Data curation, Investigation, Writing – review & editing. MZ: Data curation, Methodology, Writing – original draft. XZ: Conceptualization, Writing – review & editing.
Funding
The authors declare that no financial support was received for the research, authorship, and/or publication of this article.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
1.
Huys G Kämpfer P Albert MJ Kühn I Denys R Swings J . Aeromonas hydrophila subsp. dhakensis subsp. nov., isolated from children with diarrhoea in Bangladesh, and extended description of Aeromonas hydrophila subsp. hydrophila (Chester 1901) Stanier 1943 (approved lists 1980). hydrophila (Chester 1901) Stanier 1943 (approved lists 1980). Int J Syst Evol Microbiol. (2002) 52:705–12. doi: 10.1099/00207713-52-3-705
2.
Reclassification of Aeromonas hydrophila subsp. dhakensis Huys et al. 2002 and Aeromonas aquariorum Martínez-Murcia et al. 2008 as Aeromonas dhakensis sp. nov. comb nov. and emendation of the species Aeromonas hydrophila
3.
Chen PL Wu CJ Chen CS Tsai PJ Tang HJ Ko WC . A comparative study of clinical Aeromonas dhakensis and Aeromonas hydrophila isolates in southern Taiwan: A.dhakensis is more predominant and virulent. Clin Microbiol Infect. (2014) 20:O428–34. doi: 10.1111/1469-0691.12456
4.
Fernández-Bravo A Fort-Gallifa I Ballester F Pujol I Gomez-Bertomeu F et al . A case of Aeromonas trota in an immunocompromised patient with Diarrhea. Microorganisms. (2020) 8:399. doi: 10.3390/microorganisms8030399
5.
Pérez L Abarca ML Latif-Eugenín F Beaz-Hidalgo R Figueras MJ Domingo M . Aeromonas dhakensis pneumonia and sepsis in a neonate Risso's dolphin Grampus griseus from the Mediterranean Sea. Dis Aquat Org. (2015) 116:69–74. doi: 10.3354/dao02899
6.
Kitagawa H Ohge H Yu L Kayama S Hara T Kashiyama S et al . Aeromonas dhakensis is not a rare cause of Aeromonas bacteremia in Hiroshima, Japan. J Infect Chemother. (2020) 26:316–20. doi: 10.1016/j.jiac.2019.08.020
7.
Moreno-Sanchez F Gomez-Gomez B . Antibiotic Management of Patients with hematologic malignancies: from prophylaxis to unusual infections. Curr Oncol Rep. (2022) 24:835–42. doi: 10.1007/s11912-022-01226-y
8.
Xu C Lin Q Zhao Y Zhu G Jiang E Li S et al . Clinical characteristics and risk factors of Aeromonas bloodstream infections in patients with hematological diseases. BMC Infect Dis. (2022) 22:303. doi: 10.1186/s12879-022-07277-7
9.
Chen PL Lamy B Ko WC . Aeromonas dhakensis, an increasingly recognized human pathogen. Front Microbiol. (2016) 7:793. doi: 10.3389/fmicb.2016.00793
10.
Puah SM Khor WC Aung KT Lau TTV Puthucheary SD Chua KH . Aeromonas dhakensis: clinical isolates with high carbapenem resistance. Pathogens. (2022) 11:833. doi: 10.3390/pathogens11080833
11.
Johns Hopkins Division of Infectious Diseases Antibiotic Guide . Available at: http://www.hopkins-abxguide.org (Accessed 16 September 2002) (2002).
12.
Xu C Zhang T Cai J Yu Z Qiu J Yin J . Progress in regulatory mechanism for inducing β-lactamase in gram-negative bacteria. Sheng Wu Gong Cheng Xue Bao. (2018) 34:1288–96. doi: 10.13345/j.cjb.180187
13.
Puah SM Khor WC Kee BP Tan JAMA Puthucheary SD Chua KH . Development of a species-specific PCR-RFLP targeting rpo D gene fragment for discrimination of Aeromonas species. J Med Microbiol. (2018) 67:1271–8. doi: 10.1099/jmm.0.000796
14.
Chang HL Chen PL Lin SY Chen TC Chang K Ko WC et al . Two fatal cases of Aeromonas dhakensis bacteremia and necrotizing fasciitis in severe dengue patients. J Microbiol Immunol Infect. (2018) 51:692–4. doi: 10.1016/j.jmii.2018.03.003
15.
Melo-Bolivar JF Sinclair HA Sidjabat HE . Draft genome sequence of Aeromonas dhakensis, isolated from a patient with fatal necrotizing fasciitis. Microbiol Resour Announc. (2019) 8:e00009–19. doi: 10.1128/MRA.00009-19
16.
Huang M Chen H Li C Liu Y Gan C el-Sayed Ahmed MAEG et al . Rapid fulminant progression and mortality secondary to Aeromonas dhakensis Septicemia with hepatitis B virus infection following the ingestion of snakehead fish in mainland China: a case report. Foodborne Pathog Dis. (2020) 17:743–9. doi: 10.1089/fpd.2019.2780
17.
Sun Y Zhao Y Xu W Fang R Wu Q He H et al . Taxonomy, virulence determinants and antimicrobial susceptibility of Aeromonas spp. isolated from bacteremia in southeastern China. Antimicrob Resist Infect Control. (2021) 10:43. doi: 10.1186/s13756-021-00911-0
18.
Murata M Morinaga Y Akamatsu N Matsuda J Uno N Kosai K et al . The rapid induction of carbapenem-resistance in an Aeromonas dhakensis blood isolate. Jpn J Infect Dis. (2016) 69:439–41. doi: 10.7883/yoken.JJID.2015.098
19.
Wu CJ Chen PL Hsueh PR Chang MC Tsai PJ Shih HI et al . Clinical implications of species identification in monomicrobial Aeromonas bacteremia. PLoS One. (2015) 10:e0117821. doi: 10.1371/journal.pone.0117821
20.
di Pasquale MF Sotgiu G Gramegna A Radovanovic D Terraneo S Reyes LF et al . Prevalence and Etiology of community-acquired pneumonia in immunocompromised patients. Clin Infect Dis. (2019) 68:1482–93. doi: 10.1093/cid/ciy723
21.
Wu CJ Ko WC Lee NY Su SL Li CW Li MC et al . AeromonasIsolates from fish and patients in Tainan City, Taiwan: genotypic and phenotypic characteristics. Appl Environ Microbiol. (2019) 85:e01360–19. Published 2019 Oct 16. doi: 10.1128/AEM.01360-19
22.
Alksne LE Rasmussen BA . Expression of the AsbA1, OXA-12, and AsbM1 beta-lactamases in Aeromonas jandaei AER 14 is coordinated by a two-component regulon. J Bacteriol. (1997) 179:2006–13. doi: 10.1128/jb.179.6.2006-2013.1997
23.
Avison MB Niumsup P Walsh TR Bennett PM . Aeromonas hydrophila AmpH and CepH beta-lactamases: derepressed expression in mutants of Escherichia coli lacking creB. J Antimicrob Chemother. (2000) 46:695–02. doi: 10.1093/jac/46.5.695
24.
de Luca F Giraud-Morin C Rossolini GM Docquier JD Fosse T . Genetic and biochemical characterization of TRU-1, the endogenous class C beta-lactamase from Aeromonas enteropelogenes. Antimicrob Agents Chemother. (2010) 54:1547–54. doi: 10.1128/AAC.01252-09
25.
Fosse T Giraud-Morin C Madinier I Labia R . Sequence analysis and biochemical characterisation of chromosomal CAV-1 (Aeromonas caviae), the parental cephalosporinase of plasmid-mediated AmpC 'FOX' cluster. FEMS Microbiol Lett. (2003) 222:93–8. doi: 10.1016/S0378-1097(03)00253-2
26.
Wu CJ Wang HC Chen PL Chang MC Sunny Sun H Chou PH et al . AQU-1, a chromosomal class C β-lactamase, among clinical Aeromonas dhakensis isolates: distribution and clinical significance. Int J Antimicrob Agents. (2013) 42:456–61. doi: 10.1016/j.ijantimicag.2013.08.002
27.
Bush K Jacoby GA Medeiros AA . A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrob Agents Chemother. (1995) 39:1211–33. doi: 10.1128/AAC.39.6.1211
28.
Spiliopoulou I Kazmierczak K Stone GG . In vitro activity of ceftazidime/avibactam against isolates of carbapenem-non-susceptible Enterobacteriaceae collected during the INFORM global surveillance programme (2015-17). J Antimicrob Chemother. (2020) 75:384–91. doi: 10.1093/jac/dkz456
Summary
Keywords
Aeromonas dhakensis , Aeromonas , pulmonary infection, mNGS, antibiotic
Citation
Wang C, Wei N, Zhang M and Zhang X (2024) Pulmonary infection with Aeromonas dhakensis in a patient with acute T lymphoblastic leukemia: a case report and review of the literature. Front. Med. 11:1357714. doi: 10.3389/fmed.2024.1357714
Received
18 December 2023
Accepted
05 April 2024
Published
17 April 2024
Volume
11 - 2024
Edited by
Santi Nolasco, University of Catania, Italy
Reviewed by
Claudio Farina, ASST Papa Giovanni XXIII, Italy
Po Lin Chen, National Cheng Kung University, Taiwan
Updates
Copyright
© 2024 Wang, Wei, Zhang and Zhang.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiaoju Zhang, zhangxiaoju@zzu.edu.cn
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.