- Department of Pharmacology, School of Pharmacy, Second Military Medical University, Shanghai, China
MicroRNA-124 (miR-124) is the most abundant miRNA in the brain. Biogenesis of miR-124 displays specific temporal and spatial profiles in various cell and tissue types and affects a broad spectrum of biological functions in the central nervous system (CNS). Recently, the link between dysregulation of miR-124 and CNS disorders, such as neurodegeneration, CNS stress, neuroimmune disorders, stroke, and brain tumors, has become evident. Here, we provide an overview of the specific molecular function of miR-124 in the CNS and a revealing insight for the therapeutic potential of miR-124 in the treatment of human CNS diseases.
Introduction
The discovery of microRNAs (miRNAs) introduced an understanding of the novel type of regulatory control over gene expression during plant and animal development (Ambros, 2004; Bartel, 2004). Mature miRNAs are non-coding transcripts of 18–25 nucleotides, which imperfectly bind to complementary sequences in the 3′-untranslated regions (UTRs) of target mRNAs to negatively regulate target gene expression. It is predicted that one third of human genes are miRNA targets (Lewis et al., 2005).
MicroRNA-124 (miR-124) is highly and specifically expressed in all brain regions except for the pituitary gland, and at 100 times lower expression in other tissues (Mishima et al., 2007; Baroukh and Van Obberghen, 2009). It was first identified in mice (Lagos-Quintana et al., 2002), and mature miR-124 is wholly homologous in mice, rats, and human. miR-124 has been reported to participate in chronic stress, neurodegeneration, alcohol/cocaine neuroadaptation, synapse morphology, neurotransmission long-term potentiation, neurodevelopment myeloid cell function, and hematopoiesis (Soreq and Wolf, 2011). In mammalian neurons, miR-124 suppresses the levels of 100s of non-neural genes, which contributes to the acquisition and maintenance of neuronal identity (Lim et al., 2005; Conaco et al., 2006). Furthermore, when miR-124 is aberrantly expressed, it contributes to pathological conditions involving the central nervous system (CNS) system. It has also been shown to be useful as a diagnostic and prognostic indicator of CNS disorders, such as brain tumor and stroke. Here we review the role and potential therapy of miR-124 in CNS development and disorders.
miR-124 in CNS Development
Central nervous system development refers to the processes that generate, shape, and reshape the nervous system, from the earliest stages of embryogenesis to death. There has been particular attention on miRNAs in the context of mammalian nervous system development. Smirnova et al. (2005) compared miRNAs expression in embryonic neuron and astrocyte cultures, and found miR-124, a brain-enriched miRNA, preferentially expressed in neurons. miR-124 expression gradually increased and accumulated in parallel to neuronal maturation (Smirnova et al., 2005) during CNS development (Deo et al., 2006; Krichevsky et al., 2006). Blocking miR-124 activity in mature neurons leads selectively to increased levels of non-neuronal transcripts (Conaco et al., 2006), while increasing miR-124 activity in non-neuronal HeLa cells showed a shift of expression profile toward that of neuronal phenotype (Lim et al., 2005). Makeyev et al. (2007) demonstrated that miR-124 promoted this shift by triggering brain-specific alternative pre-mRNA splicing. In mouse embryonic development, miR-124 directly targets polypyrimidine-tract-binding protein (PTBP) mRNA, which encodes a global repressor of alternative pre-mRNA splicing in non-neuronal cells (Makeyev et al., 2007). During neuronal differentiation, miR-124 reduces PTBP levels, leading to the transition from non-nervous system to nervous system specific alternative splicing patterns. Interestingly, repression of PTBP is sufficient to induce trans-differentiation of fibroblasts into functional neurons (Xue et al., 2013).
Moreover, microRNA-124 is a subventricular zone (SVZ) neuronal fate determinant. The SVZ is the largest neurogenic niche in the adult mammalian brain. Knockdown of endogenous miR-124 maintained purified SVZ stem cells as dividing precursors, whereas ectopic expression led to increased neuron formation. Sox9 was demonstrated to be the physiological target of miR-124 responsible for this role (Cheng et al., 2009; Akerblom et al., 2012). Neurogenesis in embryonic neuroepithelial cells in the spinal cord is also regulated by miR-124 and its target Sox9 (Farrell et al., 2011). Several other targets of miR-124 have been identified in mediating the process of neurogenesis. For example, miR-124 in SVZ progenitor cells mediates stroke-induced neurogenesis by targeting the JAG-Notch signaling pathway in adult rats (Liu et al., 2011b). miR-124 also ensures the transition from neural progenitors to neurons by repressing two endogenous targets, LAMC1 and ITGB1, which are highly expressed by neural progenitors, but are repressed upon neuronal differentiation in the chick embryos (Cao et al., 2007). SCP1 down-regulation is another critical factor for inducing neurogenesis during embryonic CNS development in both chick and mouse embryos, and miR-124 contributes to this process in part by down-regulating SCP1 expression (Visvanathan et al., 2007).
Besides effects on neuron fate, miR-124 also contributes to the control of neurite outgrowth during neuronal differentiation possibly by cytoskeleton regulation in mouse P19 cells (Yu et al., 2008), and affects dendritic differentiation by regulating RhoG (Schumacher and Franke, 2013). Overexpression of miR-124 could lead to reduced astrocytic lineage differentiation by inhibiting STAT3 signaling (Krichevsky et al., 2006). miR-124 is essential for hippocampal axogenesis and retinal cone survival, as it represses Lhx2 translation in mice (Sanuki et al., 2011). In post-mitotic neurons, miR-124 represses BAF53a, which is essential for an evolutionarily conserved program of post-mitotic neural development and dendritic morphogenesis in mouse embryos (Yoo et al., 2009).
In general, miR-124 has been shown to promote cell differentiation and repress cell proliferation. In neuroblasts, miR-124 was highly expressed in cells at G0/G1 phase (Cheng et al., 2009), which results in the repression of Cdk6, a protein mediating cell-cycle progression from G0/G1 (Silber et al., 2008). miR-124 is expressed from the beginning of eye development in Xenopus, and has been shown to repress cell proliferation in the optic cup. However, this is not true during earlier development. It was recently reported that miR-124 is both necessary and sufficient to promote cell proliferation and repress neurogenesis at the optic vesicle stage which precedes optic cup formation, showing an anti-neural role by negatively regulating the expression of the pro-neural marker NeuroD1, and revealing a novel regulatory role of miR-124 in neural development (Liu et al., 2011a).
miR-124 in Neurodegeneration
Memory storage and memory-related synaptic plasticity rely on precise spatiotemporal regulation of gene expression. miR-124 plays a critical role in the regulation of signaling molecules underlying synaptic plasticity and memory (Fischbach and Carew, 2009). miR-124 was exclusively present presynaptically in a sensory-motor synapse where it constrains serotonin-induced synaptic facilitation through regulation of the transcriptional factor CREB in Aplysia, suggesting a role for miR-124 in long-term plasticity of synapses in the mature nervous system (Rajasethupathy et al., 2009). miR-124 regulates cocaine-induced plasticity by targeting BDNF in rats, which is well implicated in synaptic plasticity and plays a central role in reward and memory (Chandrasekar and Dreyer, 2009). Thus, miR-124 may be involved in neurodegeneration diseases such as Alzheimer’s disease (AD) and Parkinson’s disease (PD).
Alzheimer’s disease is an age-related chronic neurodegenerative disease and the most prevalent type of dementia in elderly people. miR-124 level is down-regulated (human-originated, n = 5; Lukiw, 2007), while the expression of β-site APP cleaving enzyme 1 (BACE1) is up-regulated, in AD patients’ brain (Bigl et al., 2000; Sun et al., 2002; Yang et al., 2003; Lukiw, 2007; Smith et al., 2011), indicating a possible inverse relationship between them. Fang et al. (2012) further demonstrated that miR-124 overexpression or knockdown could decrease or increase the expression of BACE1, and found that miR-124 may work as an important regulating factor to alleviate cell death in the process of AD by targeting BACE1 in rat PC12 cells (Fang et al., 2012), which is considered to participate in the rate-limiting step in the production of neurotoxic Aβ (Hebert et al., 2008). Melatonin levels are decreased in the serum of AD patients, and its supplementation is able to reverse AD pathology and memory deficits. Wang et al. (2013) demonstrated that melatonin rescues the EPACs/miR-124/Egrl signaling pathway in rats, which is important in learning and memory (Yang et al., 2012).
Non-coding miRNAs are necessary for the survival of post-mitotic cells such as neurons that die in PD and other brain diseases. Kim et al. (2007) showed that mice lacking Dicer in specific dopamine neurons develop a progressive loss of neurons later in life, displaying a Parkinson’s-like disease. Thus, miRNAs are essential for maintaining dopaminergic neurons in the brain, and participate in the pathogenesis of PD. Furthermore, a recent review has shown that one–fourth (49 out of 202, MIRECORDS database) of “validated” targets of miR-124 are de-regulated in PD (Sonntag, 2010), indicating an important role for miR-124 in the regulation of this disease.
miR-124 in CNS Stress
A number of miRNAs are shown to have important roles in the regulation of stress responses, and indeed miRNAs are shown to impart robustness to stress responses. The capacity of miRNAs to inhibit 100s of transcripts that are activated by stress makes them candidates for stabilizers of the homeostatic state of the transcriptome (Manakov et al., 2012).
Mental stress modifies both cholinergic neuro-transmission and alternative splicing in the brain. Meerson et al. (2010) reported that stress changes rat brain miRNA profiles detected by microarray, and some of these stress-regulated miRNAs, including miR-124, regulate alternative splicing. Also, miR-124 is up-regulated in the rat hippocampus after acute immobilization stress (Meerson et al., 2010). Another study examined the effect of maternal separation stress on miRNA expression and found the miR-124 level is elevated in the prefrontal cortex of stressed mice at P14 (Uchida et al., 2010). Paraventricular nucleus (PVN) neurons are affected by psychological stress through immune response activation. However, miR-124 was not affected by stress in mouse PVN (Mckennirey, 2011). In vitro, miR-124 from primary cultured neurons was significantly increased in response to all four neuronal challenge response sets (transfection, KCl, kainite, aging; Manakov et al., 2012).
Hundreds of transcripts endogenously expressed in neurons with target sites for miR-124 are coordinately upregulated in a variety of neuronal stresses. Overexpression of miR-124 indeed significantly inhibits expression of 100s of stress-induced transcripts detected by microarray (Manakov et al., 2012). Glucocorticoid and its receptor (glucocorticoid receptor, GR) exert profound effects on a variety of physiological processes, including adaptation to stress. Acute or chronic stress decreased GR mRNA in the PVN (Noguchi et al., 2010). Vreugdenhil et al. (2009) found that miR-124 decreased GR protein level and GR-mediated events. miR-124 expression also varies over time during the stress hyporesponsive period, a neonatal period when GC signaling is modulated (Vreugdenhil et al., 2009). By regulating GRs, miR-124 can affect a variety of systemic stress responses.
miR-124 in Neuroimmunity
MicroRNAs have unique expression profiles in cells of the innate and adaptive immune systems and have pivotal roles in the regulation of both cell development and function (O’Connell et al., 2010). Soreq and Wolf (2011) designated miRNAs which notably affect both immune and neuronal functions as NeurimmiRs. NeurimmiRs may act as ‘negotiators’ between the nervous and immune systems (Soreq and Wolf, 2011). The cholinergic anti-inflammatory pathway is a link between the brain and the immune system of the host in response to an immune challenge. This pathway controls the inflammatory response through interaction with peripheral α7 subunit–containing nicotinic acetylcholine receptors (α7AChR) expressed on macrophages (Pavlov et al., 2003). Our group found miR-124 is necessary for the cholinergic anti-inflammatory action by inhibiting the production of pro-inflammatory cytokines (Sun et al., 2013b), and miR-124 may act as an NeurimmiRs.
The p38 mitogen-activated protein kinases (MAPKs) are central regulatory nodes coordinating acute stress and inflammatory responses. Lawson et al. (2013) reported that expression of the p38α protein is suppressed in the brain by two neuron-selective miRNAs, miR-124, and -128. miR-124 may influence neuroimmunity by affecting p38α -mediated signaling.
Microglial cells are macrophages that are resident in the brain and spinal cord and form the frontline defense of the innate immune system. Utilizing a mouse model of multiple sclerosis (MS) and experimental autoimmune encephalomyelitis (EAE), Ponomarev et al. (2011) showed that microglia miR-124 expression decreased by ~70% during the course of the disease, while overexpression of miR-124 could promote microglia quiescence and suppress EAE by deactivating macrophages via the C/EBP-α-PU.1 pathway (Zhang et al., 2004). Treatment of mice with miR-124 at the onset of EAE substantially ameliorated clinical symptoms and enhanced recovery in mice (Ponomarev et al., 2011).
The ability to prevent and attenuate EAE has implications for miR-124 in the treatment of neurodegenerative diseases such as MS and AD, where microglial cells are thought to play an integral role in the inflammatory process (Conrad and Dittel, 2011). Studies have already documented inflammation in AD, PD, amyotrophic lateral sclerosis (ALS), MS, and a growing number of other nervous system pathologies (Glass et al., 2010). However, a major question is whether pharmacological inhibition of inflammation pathways will be able to safely reverse or slow the course of disease, which needs further investigation.
miR-124 in Stroke
Plasma miRNAs have been investigated as biomarkers for various diseases, including stroke. Recently, microarray analyses were done to characterize the miRNA expression profile in various stroke models (Jeyaseelan et al., 2008; Dharap et al., 2009; Liu et al., 2010; Yuan et al., 2010; Weng et al., 2011). The abundance of miR-124 in the CNS has accelerated efforts to determine if it can be used as an effective stroke treatment.
Jeyaseelan et al. (2008) first reported an elevated level of miR-124 in the brain samples from rats with middle cerebral artery occlusion (MCAO) followed by 24 h reperfusion. One of its predicted targets, the VSNL1 gene, which is a neuronal calcium sensor protein identified as a specific and promising plasma biomarker of stroke patients (Laterza et al., 2006), was decreased in parallel with the increased miR-124 under the same conditions (Jeyaseelan et al., 2008). Our previous study found miR-124 is significantly increased in ischemic penumbra as compared with that in the non-ischemic area in MCAO mice. Accordingly, brain tissues from stroke-prone spontaneously hypertensive rats (SHR-SP) showed higher miR-124 levels than in spontaneously hypertensive rats (SHRs; Sun et al., 2013a). Markedly increased plasma miR-124 was also observed at 24 h after stroke for both transient and permanent occlusions in rats (Laterza et al., 2009; Weng et al., 2011). Consistently, miR-124 was predicted to suppress acetylcholinesterase (AChE; Nadorp and Soreq, 2014), whose plasma level was reduced in patients post ischemic stroke (Ben Assayag et al., 2010), indicating a physiological relevance between miR-124 and AChE. The above suggests that plasma miR-124 released from the infracted brain may be a promising candidate biomarker for stroke identification.
Although there is no correlation between the infarct size and plasma miR-124 level in rats after MCAO introduction (Weng et al., 2011), efforts have been made to explore whether miR-124 treatment is effective in stroke. Doeppner et al. (2013) showed that exogenous miR-124 reduced brain injury and functional impairment, enhanced neurovascular remodeling, and increased angioneurogenesis 8 weeks post-stroke in mice with MCAO, which is possible via the pathway involving Usp14-dependent REST degradation. Similarly, our previous study showed that miR-124 overexpression decreased the infarct area of MCAO mice (Sun et al., 2013a). The anti-apoptosis proteins Bcl-2 and Bcl-xL, key regulators in attenuating stroke-induced apoptotic cell death (Martinou et al., 1994; Wiessner et al., 1999; Graham et al., 2000), are found to be the targets of miR-124 in this protective role for stroke. miR-124 was also found to inhibit stroke-induced neurogenesis by targeting the JAG1/Notch signaling pathway (Liu et al., 2011b).
However, conflicting miR-124 stroke therapy results have also been reported. For example, Zhu et al. (2014) showed that knockdown of cerebral miR-124 reduced cell death and infarct size, and improved neurological outcomes by negatively regulating Ku70. Additionally, Liu et al. (2013) found that although a miR-124 mimic did not affect the infarct volume at 24 h after ischemia, inhibition of miR-124 effectively reduced the ischemic injury due to iASPP expression up-regulation. Therefore, the use of miR-124 as an effective stroke treatment necessitates further research.
miR-124 in Brain Tumor
Distinct patterns of miR-124 expression have been observed in many cancers including glioblastomas (Silber et al., 2008). miR-124 expression is significantly decreased in anaplastic astrocytoma and glioblastoma relative in human patients to non-neoplastic brain tissue (Fowler et al., 2011; Hua et al., 2012; Lang et al., 2012; Ho et al., 2013), and is expressed at different levels in glioblastoma compared with normal brain (Silber et al., 2008; Godlewski et al., 2010). Overexpression of miR-124 induced morphological changes and neural differentiation in mouse neural stem cells and oligodendroglioma, accompanied by reduced self-renewal and tumorigenicity (Silber et al., 2008). In addition, the ectopic expression of miR-124 in a glioblastoma cell line resulted in significant inhibition of migration and invasion, suggesting that miR-124 may be a novel inhibitor of glioblastoma invasion (Fowler et al., 2011).
Several targets of miR-124 mediating this process have been identified. It was reported that miR-124 controls self-renewal and tumorigenic competence of human glioblastoma cells by targeting SCP1 and PTPN12 phosphatases (Conti et al., 2012; Lee et al., 2013). Xia et al. (2012) demonstrated that the tumor suppressor activity of miR-124 could by partly due to its inhibitory effects on glioma stem – like traits and invasiveness through down-regulation of SNAI2 in human. Others reported that miR-124 inhibits glioma cells migration and invasion by down-regulation of ROCK1 (An et al., 2013), and induced glioma differentiation by suppressing Twist and SNAI2 (Xie et al., 2012).
A clinical investigation also showed a negative correlation between miR-124 expression and a hypoxic gene signature in glioblastoma patient samples. Increased miR-124 expression affects the ability of tumor cells to survive under O2 and/or nutrient deprivation, while miR-124 re-expression increases cell death in vivo and enhances the survival of mice bearing intracranial xenograft tumors. miR-124 exerts this phenotype in part by directly regulating TEAD1, MAPK14/p38α, and SERP1, which are factors involved in cell proliferation and survival under stress (Mucaj et al., 2014). Besides glioblastoma, neuroblastoma is an embryonic tumor derived from the autonomic nervous system neural-crest tissues, and is the most common extracranial solid tumor in children. Huang et al. (2011) showed that miR-124 plays a pivotal role in neuroblastoma by targeting aryl hydrocarbon receptor (AHR), which may promote neuroblastoma cell differentiation. These studies indicate that miR-124 is a potential therapeutic target in brain tumor treatment.
Conclusion
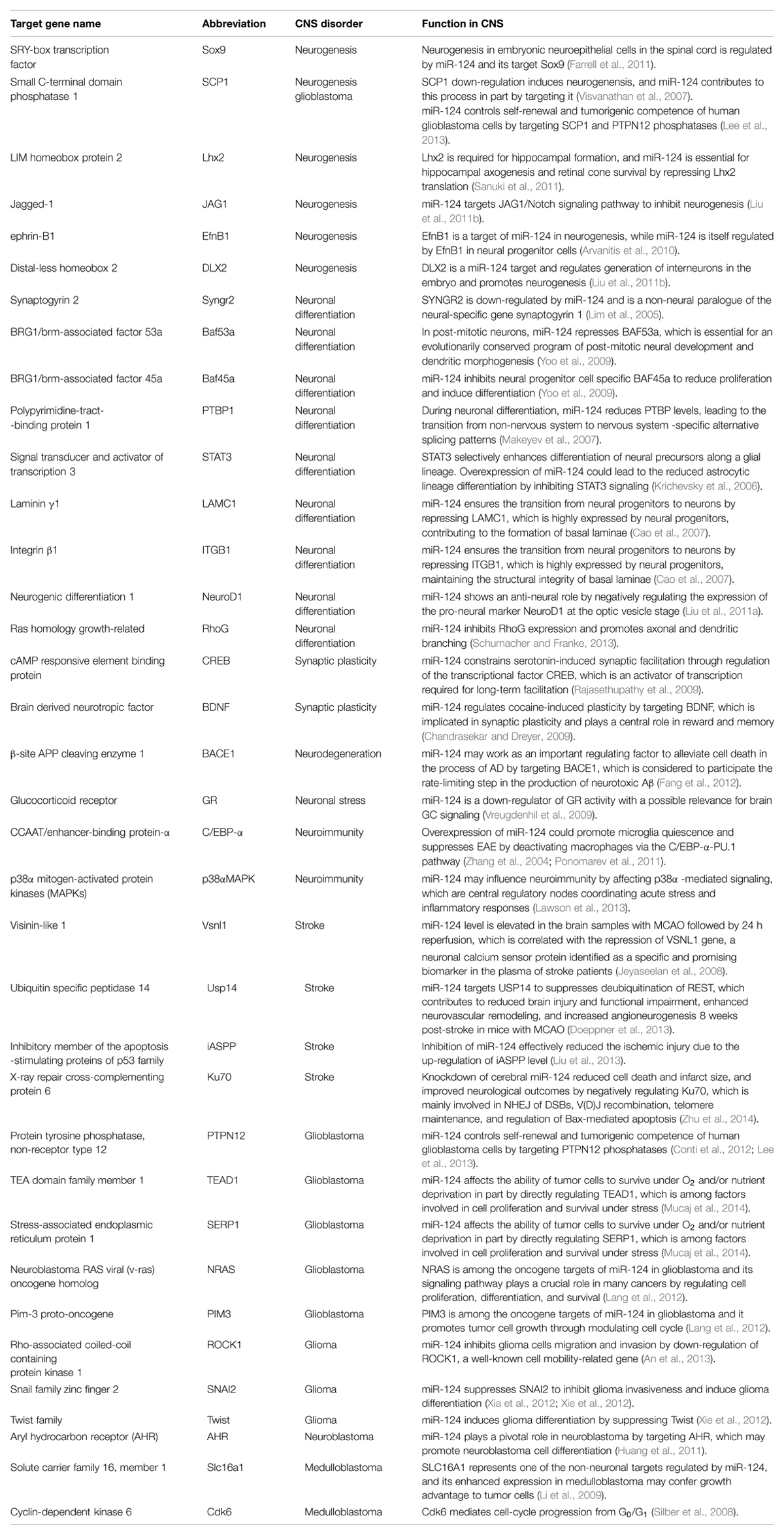
We have discussed the importance of miR-124 in neuronal development and function of the brain. Changes of miR-124 levels can serve as biomarkers that indicate the functional status of a normal brain, as well as progression of CNS diseases. Interestingly, it seems inverse effect of miR-124 in tumorogenic events and neurodegenerative processes. For example, miR-124 alleviates cell death in the process of AD by targeting BACE1, while increases cell death in glioblastoma by regulating TEAD1, MAPK14/p38α, and SERP1 (Mucaj et al., 2014). This might be relevant to different targets involved depending on the context of diseases, cells and surroundings. Moreover, its regulation of numerous targets (summarized in Table 1) classifies it as a suitable treatment for the complexity of pathophysiological events launched by multifactorial CNS diseases, which require therapies that are able to trigger a large set of orchestrated genes. Therefore, we believe miR-124 and other miRNAs that fulfill this criterion will replace single gene therapies for the treatment of these kinds of diseases in the near future.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (81273606 and 81473259 to XL, 81230083 to DFS). The authors declare no competing financial interests.
References
Akerblom, M., Sachdeva, R., Barde, I., Verp, S., Gentner, B., Trono, D., et al. (2012). MicroRNA-124 is a subventricular zone neuronal fate determinant. J. Neurosci. 32, 8879–8889. doi: 10.1523/JNEUROSCI.0558-12.2012
An, L., Liu, Y., Wu, A., and Guan, Y. (2013). microRNA-124 inhibits migration and invasion by down-regulating ROCK1 in glioma. PLoS ONE 8:e69478. doi: 10.1371/journal.pone.0069478
Arvanitis, D. N., Jungas, T., Behar, A., and Davy, A. (2010). Ephrin-B1 reverse signaling controls a posttranscriptional feedback mechanism via miR-124. Mol. Cell. Biol. 30, 2508–2517. doi: 10.1128/MCB.01620-09
Baroukh, N. N., and Van Obberghen, E. (2009). Function of microRNA-375 and microRNA-124a in pancreas and brain. FEBS J. 276, 6509–6521. doi: 10.1111/j.1742-4658.2009.07353.x
Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297. doi: 10.1016/S0092-8674(04)00045-5
Ben Assayag, E., Shenhar-Tsarfaty, S., Ofek, K., Soreq, L., Bova, I., Shopin, L., et al. (2010). Serum cholinesterase activities distinguish between stroke patients and controls and predict 12-month mortality. Mol. Med. 16, 278–286. doi: 10.2119/molmed.2010.00015
Bigl, M., Apelt, J., Luschekina, E. A., Lange-Dohna, C., Rossner, S., and Schliebs, R. (2000). Expression of beta-secretase mRNA in transgenic Tg2576 mouse brain with Alzheimer plaque pathology. Neurosci. Lett. 292, 107–110. doi: 10.1016/S0304-3940(00)01452-X
Cao, X., Pfaff, S. L., and Gage, F. H. (2007). A functional study of miR-124 in the developing neural tube. Genes Dev. 21, 531–536. doi: 10.1101/gad.1519207
Chandrasekar, V., and Dreyer, J. L. (2009). microRNAs miR-124, let-7d and miR-181a regulate cocaine-induced plasticity. Mol. Cell. Neurosci. 42, 350–362. doi: 10.1016/j.mcn.2009.08.009
Cheng, L. C., Pastrana, E., Tavazoie, M., and Doetsch, F. (2009). miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nat. Neurosci. 12, 399–408. doi: 10.1038/nn.2294
Conaco, C., Otto, S., Han, J. J., and Mandel, G. (2006). Reciprocal actions of REST and a microRNA promote neuronal identity. Proc. Natl. Acad. Sci. U.S.A. 103, 2422–2427. doi: 10.1073/pnas.0511041103
Conrad, A. T., and Dittel, B. N. (2011). Taming of macrophage and microglial cell activation by microRNA-124. Cell Res. 21, 213–216. doi: 10.1038/cr.2011.9
Conti, L., Crisafulli, L., Caldera, V., Tortoreto, M., Brilli, E., Conforti, P., et al. (2012). REST controls self-renewal and tumorigenic competence of human glioblastoma cells. PLoS ONE 7:e38486. doi: 10.1371/journal.pone.0038486
Deo, M., Yu, J. Y., Chung, K. H., Tippens, M., and Turner, D. L. (2006). Detection of mammalian microRNA expression by in situ hybridization with RNA oligonucleotides. Dev. Dyn. 235, 2538–2548. doi: 10.1002/dvdy.20847
Dharap, A., Bowen, K., Place, R., Li, L. C., and Vemuganti, R. (2009). Transient focal ischemia induces extensive temporal changes in rat cerebral microRNAome. J. Cereb. Blood Flow Metab. 29, 675–687. doi: 10.1038/jcbfm.2008.157
Doeppner, T. R., Doehring, M., Bretschneider, E., Zechariah, A., Kaltwasser, B., Muller, B., et al. (2013). MicroRNA-124 protects against focal cerebral ischemia via mechanisms involving Usp14-dependent REST degradation. Acta Neuropathol. 126, 251–265. doi: 10.1007/s00401-013-1142-5
Fang, M., Wang, J., Zhang, X., Geng, Y., Hu, Z., Rudd, J. A., et al. (2012). The miR-124 regulates the expression of BACE1/beta-secretase correlated with cell death in Alzheimer’s disease. Toxicol. Lett. 209, 94–105. doi: 10.1016/j.toxlet.2011.11.032
Farrell, B. C., Power, E. M., and Mc Dermott, K. W. (2011). Developmentally regulated expression of Sox9 and microRNAs 124, 128 and 23 in neuroepithelial stem cells in the developing spinal cord. Int. J. Dev. Neurosci. 29, 31–36. doi: 10.1016/j.ijdevneu.2010.10.001
Fischbach, S. J., and Carew, T. J. (2009). MicroRNAs in memory processing. Neuron 63, 714–716. doi: 10.1016/j.neuron.2009.09.007
Fowler, A., Thomson, D., Giles, K., Maleki, S., Mreich, E., Wheeler, H., et al. (2011). miR-124a is frequently down-regulated in glioblastoma and is involved in migration and invasion. Eur. J. Cancer 47, 953–963. doi: 10.1016/j.ejca.2010.11.026
Glass, C. K., Saijo, K., Winner, B., Marchetto, M. C., and Gage, F. H. (2010). Mechanisms underlying inflammation in neurodegeneration. Cell 140, 918–934. doi: 10.1016/j.cell.2010.02.016
Godlewski, J., Newton, H. B., Chiocca, E. A., and Lawler, S. E. (2010). MicroRNAs and glioblastoma; the stem cell connection. Cell Death. Differ. 17, 221–228. doi: 10.1038/cdd.2009.71
Graham, S. H., Chen, J., and Clark, R. S. (2000). Bcl-2 family gene products in cerebral ischemia and traumatic brain injury. J. Neurotrauma 17, 831–841. doi: 10.1089/neu.2000.17.831
Hebert, S. S., Horre, K., Nicolai, L., Papadopoulou, A. S., Mandemakers, W., Silahtaroglu, A. N., et al. (2008). Loss of microRNA cluster miR-29a/b-1 in sporadic Alzheimer’s disease correlates with increased BACE1/beta-secretase expression. Proc. Natl. Acad. Sci. U.S.A. 105, 6415–6420. doi: 10.1073/pnas.0710263105
Ho, C. Y., Bar, E., Giannini, C., Marchionni, L., Karajannis, M. A., Zagzag, D., et al. (2013). MicroRNA profiling in pediatric pilocytic astrocytoma reveals biologically relevant targets, including PBX3, NFIB, and METAP2. Neuro Oncol. 15, 69–82. doi: 10.1093/neuonc/nos269
Hua, D., Mo, F., Ding, D., Li, L., Han, X., Zhao, N., et al. (2012). A catalogue of glioblastoma and brain MicroRNAs identified by deep sequencing. OMICS 16, 690–699. doi: 10.1089/omi.2012.0069
Huang, T. C., Chang, H. Y., Chen, C. Y., Wu, P. Y., Lee, H., Liao, Y. F., et al. (2011). Silencing of miR-124 induces neuroblastoma SK-N-SH cell differentiation, cell cycle arrest and apoptosis through promoting AHR. FEBS Lett. 585, 3582–3586. doi: 10.1016/j.febslet.2011.10.025
Jeyaseelan, K., Lim, K. Y., and Armugam, A. (2008). MicroRNA expression in the blood and brain of rats subjected to transient focal ischemia by middle cerebral artery occlusion. Stroke 39, 959–966. doi: 10.1161/STROKEAHA.107.500736
Kim, J., Inoue, K., Ishii, J., Vanti, W. B., Voronov, S. V., Murchison, E., et al. (2007). A MicroRNA feedback circuit in midbrain dopamine neurons. Science 317, 1220–1224. doi: 10.1126/science.1140481
Krichevsky, A. M., Sonntag, K. C., Isacson, O., and Kosik, K. S. (2006). Specific microRNAs modulate embryonic stem cell-derived neurogenesis. Stem Cells 24, 857–864. doi: 10.1634/stemcells.2005-0441
Lagos-Quintana, M., Rauhut, R., Yalcin, A., Meyer, J., Lendeckel, W., and Tuschl, T. (2002). Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739. doi: 10.1016/S0960-9822(02)00809-6
Lang, M. F., Yang, S., Zhao, C., Sun, G., Murai, K., Wu, X., et al. (2012). Genome-wide profiling identified a set of miRNAs that are differentially expressed in glioblastoma stem cells and normal neural stem cells. PLoS ONE 7:e36248. doi: 10.1371/journal.pone.0036248
Laterza, O. F., Lim, L., Garrett-Engele, P. W., Vlasakova, K., Muniappa, N., Tanaka, W. K., et al. (2009). Plasma MicroRNAs as sensitive and specific biomarkers of tissue injury. Clin. Chem. 55, 1977–1983. doi: 10.1373/clinchem.2009.131797
Laterza, O. F., Modur, V. R., Crimmins, D. L., Olander, J. V., Landt, Y., Lee, J. M., et al. (2006). Identification of novel brain biomarkers. Clin. Chem. 52, 1713–1721. doi: 10.1373/clinchem.2006.070912
Lawson, S. K., Dobrikova, E. Y., Shveygert, M., and Gromeier, M. (2013). p38alpha mitogen-activated protein kinase depletion and repression of signal transduction to translation machinery by miR-124 and -128 in neurons. Mol. Cell. Biol. 33, 127–135. doi: 10.1128/MCB.00695-12
Lee, H. K., Finniss, S., Cazacu, S., Bucris, E., Ziv-Av, A., Xiang, C., et al. (2013). Mesenchymal stem cells deliver synthetic microRNA mimics to glioma cells and glioma stem cells and inhibit their cell migration and self-renewal. Oncotarget 4, 346–361.
Lewis, B. P., Burge, C. B., and Bartel, D. P. (2005). Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20. doi: 10.1016/j.cell.2004.12.035
Li, K. K., Pang, J. C., Ching, A. K., Wong, C. K., Kong, X., Wang, Y., et al. (2009). miR-124 is frequently down-regulated in medulloblastoma and is a negative regulator of SLC16A1. Hum. Pathol. 40, 1234–1243. doi: 10.1016/j.humpath.2009.02.003
Lim, L. P., Lau, N. C., Garrett-Engele, P., Grimson, A., Schelter, J. M., Castle, J., et al. (2005). Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769–773. doi: 10.1038/nature03315
Liu, D. Z., Tian, Y., Ander, B. P., Xu, H., Stamova, B. S., Zhan, X., et al. (2010). Brain and blood microRNA expression profiling of ischemic stroke, intracerebral hemorrhage, and kainate seizures. J. Cereb. Blood Flow Metab. 30, 92–101. doi: 10.1038/jcbfm.2009.186
Liu, K., Liu, Y., Mo, W., Qiu, R., Wang, X., Wu, J. Y., et al. (2011a). MiR-124 regulates early neurogenesis in the optic vesicle and forebrain, targeting NeuroD1. Nucleic Acids Res. 39, 2869–2879. doi: 10.1093/nar/gkq904
Liu, X. S., Chopp, M., Zhang, R. L., Tao, T., Wang, X. L., Kassis, H., et al. (2011b). MicroRNA profiling in subventricular zone after stroke: miR-124a regulates proliferation of neural progenitor cells through Notch signaling pathway. PLoS ONE 6:e23461. doi: 10.1371/journal.pone.0023461
Liu, X., Li, F., Zhao, S., Luo, Y., Kang, J., Zhao, H., et al. (2013). MicroRNA-124-mediated regulation of inhibitory member of apoptosis-stimulating protein of p53 family in experimental stroke. Stroke 44, 1973–1980. doi: 10.1161/STROKEAHA.111.000613
Lukiw, W. J. (2007). Micro-RNA speciation in fetal, adult and Alzheimer’s disease hippocampus. Neuroreport 18, 297–300. doi: 10.1097/WNR.0b013e3280148e8b
Makeyev, E. V., Zhang, J., Carrasco, M. A., and Maniatis, T. (2007). The MicroRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Mol. Cell. 27, 435–448. doi: 10.1016/j.molcel.2007.07.015
Manakov, S. A., Morton, A., Enright, A. J., and Grant, S. G. (2012). A Neuronal Transcriptome Response Involving Stress Pathways is Buffered by Neuronal microRNAs. Front. Neurosci. 6:156. doi: 10.3389/fnins.2012.00156
Martinou, J. C., Dubois-Dauphin, M., Staple, J. K., Rodriguez, I., Frankowski, H., Missotten, M., et al. (1994). Overexpression of BCL-2 in transgenic mice protects neurons from naturally occurring cell death and experimental ischemia. Neuron 13, 1017–1030. doi: 10.1016/0896-6273(94)90266-6
Mckennirey, P. (2011). The Effects of Stress on Mouse Prefrontal Cortex microRNA Expression. Ottawa: Carleton University.
Meerson, A., Cacheaux, L., Goosens, K. A., Sapolsky, R. M., Soreq, H., and Kaufer, D. (2010). Changes in brain MicroRNAs contribute to cholinergic stress reactions. J. Mol. Neurosci. 40, 47–55. doi: 10.1007/s12031-009-9252-1
Mishima, T., Mizuguchi, Y., Kawahigashi, Y., Takizawa, T., and Takizawa, T. (2007). RT-PCR-based analysis of microRNA (miR-1 and -124) expression in mouse CNS. Brain Res. 1131, 37–43. doi: 10.1016/j.brainres.2006.11.035
Mucaj, V., Lee, S. S., Skuli, N., Giannoukos, D. N., Qiu, B., Eisinger-Mathason, T. S., et al. (2014). MicroRNA-124 expression counteracts pro-survival stress responses in glioblastoma. Oncogene 34, 2204–2214. doi: 10.1038/onc.2014.168
Nadorp, B., and Soreq, H. (2014). Predicted overlapping microRNA regulators of acetylcholine packaging and degradation in neuroinflammation-related disorders. Front. Mol. Neurosci. 7:9. doi: 10.3389/fnmol.2014.00009
Noguchi, T., Makino, S., Matsumoto, R., Nakayama, S., Nishiyama, M., Terada, Y., et al. (2010). Regulation of glucocorticoid receptor transcription and nuclear translocation during single and repeated immobilization stress. Endocrinology 151, 4344–4355. doi: 10.1210/en.2010-0266
O’Connell, R. M., Rao, D. S., Chaudhuri, A. A., and Baltimore, D. (2010). Physiological and pathological roles for microRNAs in the immune system. Nat. Rev. Immunol. 10, 111–122. doi: 10.1038/nri2708
Pavlov, V. A., Wang, H., Czura, C. J., Friedman, S. G., and Tracey, K. J. (2003). The cholinergic anti-inflammatory pathway: a missing link in neuroimmunomodulation. Mol. Med. 9, 125–134.
Ponomarev, E. D., Veremeyko, T., Barteneva, N., Krichevsky, A. M., and Weiner, H. L. (2011). MicroRNA-124 promotes microglia quiescence and suppresses EAE by deactivating macrophages via the C/EBP-alpha-PU.1 pathway. Nat. Med. 17, 64–70. doi: 10.1038/nm.2266
Rajasethupathy, P., Fiumara, F., Sheridan, R., Betel, D., Puthanveettil, S. V., Russo, J. J., et al. (2009). Characterization of small RNAs in Aplysia reveals a role for miR-124 in constraining synaptic plasticity through CREB. Neuron 63, 803–817. doi: 10.1016/j.neuron.2009.05.029
Sanuki, R., Onishi, A., Koike, C., Muramatsu, R., Watanabe, S., Muranishi, Y., et al. (2011). miR-124a is required for hippocampal axogenesis and retinal cone survival through Lhx2 suppression. Nat. Neurosci. 14, 1125–1134. doi: 10.1038/nn.2897
Schumacher, S., and Franke, K. (2013). miR-124-regulated RhoG: a conductor of neuronal process complexity. Small GTPases 4, 42–46. doi: 10.4161/sgtp.22922
Silber, J., Lim, D. A., Petritsch, C., Persson, A. I., Maunakea, A. K., Yu, M., et al. (2008). miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med. 6:14. doi: 10.1186/1741-7015-6-14
Smirnova, L., Grafe, A., Seiler, A., Schumacher, S., Nitsch, R., and Wulczyn, F. G. (2005). Regulation of miRNA expression during neural cell specification. Eur. J. Neurosci. 21, 1469–1477. doi: 10.1111/j.1460-9568.2005.03978.x
Smith, P., Al Hashimi, A., Girard, J., Delay, C., and Hebert, S. S. (2011). In vivo regulation of amyloid precursor protein neuronal splicing by microRNAs. J. Neurochem. 116, 240–247. doi: 10.1111/j.1471-4159.2010.07097.x
Sonntag, K. C. (2010). MicroRNAs and deregulated gene expression networks in neurodegeneration. Brain Res. 1338, 48–57. doi: 10.1016/j.brainres.2010.03.106
Soreq, H., and Wolf, Y. (2011). NeurimmiRs: microRNAs in the neuroimmune interface. Trends Mol. Med. 17, 548–555. doi: 10.1016/j.molmed.2011.06.009
Sun, A., Koelsch, G., Tang, J., and Bing, G. (2002). Localization of beta-secretase memapsin 2 in the brain of Alzheimer’s patients and normal aged controls. Exp. Neurol. 175, 10–22. doi: 10.1006/exnr.2002.7875
Sun, Y., Gui, H., Li, Q., Luo, Z. M., Zheng, M. J., Duan, J. L., et al. (2013a). MicroRNA-124 protects neurons against apoptosis in cerebral ischemic stroke. CNS Neurosci. Ther. 19, 813–819. doi: 10.1111/cns.12142
Sun, Y., Li, Q., Gui, H., Xu, D. P., Yang, Y. L., Su, D. F., et al. (2013b). MicroRNA-124 mediates the cholinergic anti-inflammatory action through inhibiting the production of pro-inflammatory cytokines. Cell Res. 23, 1270–1283. doi: 10.1038/cr.2013.116
Uchida, S., Hara, K., Kobayashi, A., Funato, H., Hobara, T., Otsuki, K., et al. (2010). Early life stress enhances behavioral vulnerability to stress through the activation of REST4-mediated gene transcription in the medial prefrontal cortex of rodents. J. Neurosci. 30, 15007–15018. doi: 10.1523/JNEUROSCI.1436-10.2010
Visvanathan, J., Lee, S., Lee, B., Lee, J. W., and Lee, S. K. (2007). The microRNA miR-124 antagonizes the anti-neural REST/SCP1 pathway during embryonic CNS development. Genes Dev. 21, 744–749. doi: 10.1101/gad.1519107
Vreugdenhil, E., Verissimo, C. S., Mariman, R., Kamphorst, J. T., Barbosa, J. S., Zweers, T., et al. (2009). MicroRNA 18 and 124a down-regulate the glucocorticoid receptor: implications for glucocorticoid responsiveness in the brain. Endocrinology 150, 2220–2228. doi: 10.1210/en.2008-1335
Wang, X., Wang, Z. H., Wu, Y. Y., Tang, H., Tan, L., Wang, X., et al. (2013). Melatonin attenuates scopolamine-induced memory/synaptic disorder by rescuing EPACs/miR-124/Egr1 pathway. Mol. Neurobiol. 47, 373–381. doi: 10.1007/s12035-012-8355-9
Weng, H., Shen, C., Hirokawa, G., Ji, X., Takahashi, R., Shimada, K., et al. (2011). Plasma miR-124 as a biomarker for cerebral infarction. Biomed. Res. 32, 135–141. doi: 10.2220/biomedres.32.135
Wiessner, C., Allegrini, P. R., Rupalla, K., Sauer, D., Oltersdorf, T., Mcgregor, A. L., et al. (1999). Neuron-specific transgene expression of Bcl-XL but not Bcl-2 genes reduced lesion size after permanent middle cerebral artery occlusion in mice. Neurosci. Lett. 268, 119–122. doi: 10.1016/S0304-3940(99)00392-4
Xia, H., Cheung, W. K., Ng, S. S., Jiang, X., Jiang, S., Sze, J., et al. (2012). Loss of brain-enriched miR-124 microRNA enhances stem-like traits and invasiveness of glioma cells. J. Biol. Chem. 287, 9962–9971. doi: 10.1074/jbc.M111.332627
Xie, Y. K., Huo, S. F., Zhang, G., Zhang, F., Lian, Z. P., Tang, X. L., et al. (2012). CDA-2 induces cell differentiation through suppressing Twist/SLUG signaling via miR-124 in glioma. J. Neurooncol. 110, 179–186. doi: 10.1007/s11060-012-0961-x
Xue, Y., Ouyang, K., Huang, J., Zhou, Y., Ouyang, H., Li, H., et al. (2013). Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell 152, 82–96. doi: 10.1016/j.cell.2012.11.045
Yang, L. B., Lindholm, K., Yan, R., Citron, M., Xia, W., Yang, X. L., et al. (2003). Elevated beta-secretase expression and enzymatic activity detected in sporadic Alzheimer disease. Nat. Med. 9, 3–4. doi: 10.1038/nm0103-3
Yang, Y., Shu, X., Liu, D., Shang, Y., Wu, Y., Pei, L., et al. (2012). EPAC null mutation impairs learning and social interactions via aberrant regulation of miR-124 and Zif268 translation. Neuron 73, 774–788. doi: 10.1016/j.neuron.2012.02.003
Yoo, A. S., Staahl, B. T., Chen, L., and Crabtree, G. R. (2009). MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 460, 642–646. doi: 10.1038/nature08139
Yu, J. Y., Chung, K. H., Deo, M., Thompson, R. C., and Turner, D. L. (2008). MicroRNA miR-124 regulates neurite outgrowth during neuronal differentiation. Exp. Cell Res. 314, 2618–2633. doi: 10.1016/j.yexcr.2008.06.002
Yuan, Y., Wang, J. Y., Xu, L. Y., Cai, R., Chen, Z., and Luo, B. Y. (2010). MicroRNA expression changes in the hippocampi of rats subjected to global ischemia. J. Clin. Neurosci. 17, 774–778. doi: 10.1016/j.jocn.2009.10.009
Zhang, P., Iwasaki-Arai, J., Iwasaki, H., Fenyus, M. L., Dayaram, T., Owens, B. M., et al. (2004). Enhancement of hematopoietic stem cell repopulating capacity and self-renewal in the absence of the transcription factor C/EBP alpha. Immunity 21, 853–863. doi: 10.1016/j.immuni.2004.11.006
Keywords: microRNA-124, CNS disorders, brain development, neurodegradation, CNS stress, neuroimmunity, brain tumor, stroke
Citation: Sun Y, Luo Z-M, Guo X-M, Su D-F and Liu X (2015) An updated role of microRNA-124 in central nervous system disorders: a review. Front. Cell. Neurosci. 9:193. doi: 10.3389/fncel.2015.00193
Received: 25 March 2015; Accepted: 04 May 2015;
Published online: 20 May 2015.
Edited by:
Tommaso Pizzorusso, University of Florence and Institute of Neuroscience of the National Research Council, ItalyReviewed by:
Hermona Soreq, The Hebrew University of Jerusalem, IsraelLei Liu, University of Florida College of Medicine, USA
Copyright © 2015 Sun, Luo, Guo, Su and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xia Liu, Department of Pharmacology, School of Pharmacy, Second Military Medical University, 325 Guo-He Road, Shanghai 200433, China,bHhmbHlpbmdAYWxpeXVuLmNvbQ==
 Yang Sun
Yang Sun Zhu-Min Luo
Zhu-Min Luo Xiu-Ming Guo
Xiu-Ming Guo Ding-Feng Su
Ding-Feng Su Xia Liu
Xia Liu