- 1University of Bordeaux, Interdisciplinary Institute for Neuroscience, UMR 5297, Bordeaux, France
- 2CNRS, Interdisciplinary Institute for Neuroscience, UMR 5297, Bordeaux, France
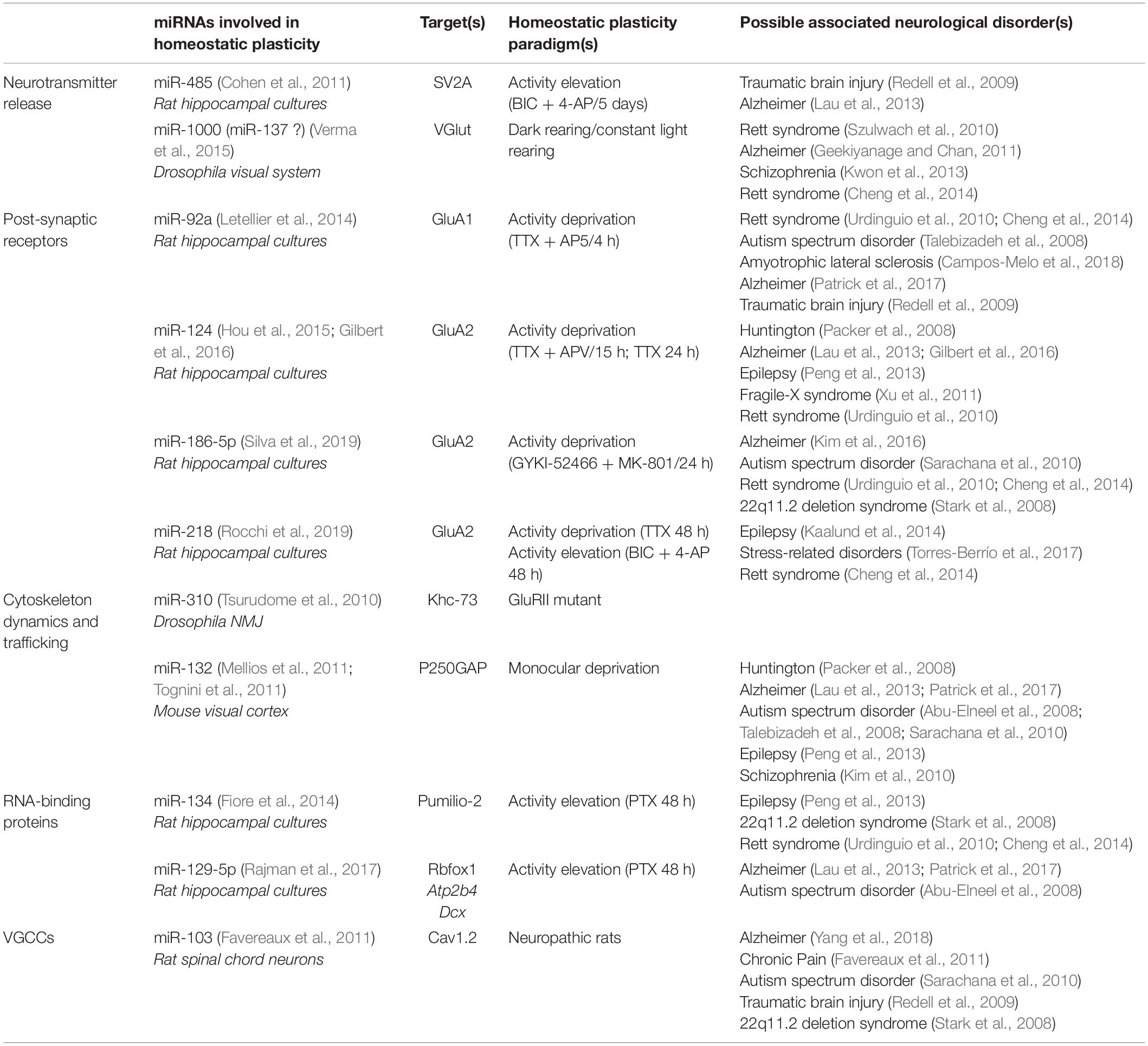
Homeostatic plasticity is a form of plasticity in which neurons compensate for changes in neuronal activity through the control of key physiological parameters such as the number and the strength of their synaptic inputs and intrinsic excitability. Recent studies revealed that miRNAs, which are small non-coding RNAs repressing mRNA translation, participate in this process by controlling the translation of multiple effectors such as glutamate transporters, receptors, signaling molecules and voltage-gated ion channels. In this review, we present and discuss the role of miRNAs in both cell-wide and compartmentalized forms of homeostatic plasticity as well as their implication in pathological processes associated with homeostatic failure.
Introduction
Neurons employ a variety of homeostatic mechanisms to maintain network activity within physiological ranges in response to a wide range of remodeling events. These include for instance the assembly of synaptic circuits during development, learning and memory, or the pathological loss of synapses associated with neurodegenerative disorders (Hengen et al., 2013; Keck et al., 2013; Vitureira and Goda, 2013; Fernandes and Carvalho, 2016; Turrigiano, 2017). Depending on the situation, such homeostatic mechanisms may involve different signaling pathways, act on various physiological parameters, and operate on multiple time and space scales (reviewed in Turrigiano, 2012; Vitureira et al., 2012; Fernandes and Carvalho, 2016). It is therefore not surprising that the failure of neuronal homeostasis can impact physiological processes such as memory consolidation and synaptic circuit refinement (Mrsic-Flogel et al., 2007; Hengen et al., 2016; Diering et al., 2017), can contribute to epilepsy (Swann and Rho, 2014) and to various neurological disorders (Ramocki and Zoghbi, 2008; Dickman and Davis, 2009; Wondolowski and Dickman, 2013; Nelson and Valakh, 2015; Penn et al., 2017).
One important feature shared among the multiple forms of homeostatic plasticity is that they are slow as compared to Hebbian forms of plasticity, i.e., long-term potentiation (LTP) or depression (LTD), in which synaptic strengths are rapidly and durably potentiated or depressed, respectively. Homeostatic plasticity usually develops over the course of several hours, and up to several days, and relies on the synthesis of new proteins which regulate key physiological parameters (Turrigiano, 2012; Fernandes and Carvalho, 2016). Proteins as diverse as glutamate receptors (e.g., AMPARs), scaffolding proteins (e.g., PSD-95, PICK1), voltage-gated ion channels (e.g., P/Q-type calcium channels), kinases (e.g., CAMKIIβ, PKA), cell-adhesion molecules (e.g., β3-integrins) or soluble factors (e.g., TNFα, retinoic acid, BDNF) contribute to homeostatic plasticity through the regulation of synaptic efficacy, synapse number, and/or membrane excitability (reviewed in Turrigiano, 2012; Fernandes and Carvalho, 2016). So far, several studies have uncovered the role of activity-dependent mRNA transcription of immediate early genes like Plk2, Homer1a, Arc, and Narp (Shepherd et al., 2006; Seeburg et al., 2008; Chang et al., 2010; Gao et al., 2010; Diering et al., 2017) and the contribution of transcription regulators such as MSK1, MeCP2, and CaMKIV (Ibata et al., 2008; Blackman et al., 2012; Correa et al., 2012; Qiu et al., 2012). In contrast to transcriptional studies, a potential contribution of mechanisms regulating de novo protein synthesis at the post-transcriptional level such as mRNA translation and/or stability is just emerging (Fernandez-Moya et al., 2014; Kosik, 2016).
Among the actors that may be involved in these processes, microRNAs (miRNAs) appear as important regulators of homeostatic plasticity in the nervous system. These small non-coding RNAs are highly enriched in the brain where they regulate a very large number of genes and shape transcriptomic diversity across regions (Filipowicz et al., 2008; Friedman et al., 2008; Soula et al., 2018). miRNAs are first synthesized in the nucleus then loaded in the RNA induced silencing complex (RISC), where they hybridize to the 3′ UTR of target mRNAs and inhibit protein synthesis through translational repression or destabilization of the transcript (Figure 1). The sequence involved in miRNA–mRNA interaction is called the “seed” region and is composed of the nucleotides 2–8 of the 5′ region of the miRNA (Bartel, 2009). Due to the small size of the “seed” region and the length of 3′ UTRs, the translation of a given mRNA is often under the control of multiple miRNAs while individual miRNAs can regulate the expression of dozens, if not 100s, of genes (Friedman et al., 2008). Loss of function approaches targeting individual miRNAs or their maturation through the endoribonuclease Dicer (Giraldez, 2005; Kim et al., 2007; Cuellar et al., 2008; Störchel et al., 2015; Fiorenza et al., 2016) have unveiled a contribution of the miRNA system in most aspects of neuronal development and plasticity, including neuronal differentiation and survival, neurite growth, synapse development, and plasticity (Kosik, 2006; Fineberg et al., 2009; Follert et al., 2014; Hu and Li, 2017; Tien and Kerschensteiner, 2018). In comparison with the regulation of transcription, which is spatially restricted to the nucleus, miRNAs provide an additional layer of regulations to finely tune in time and space protein synthesis in remote subcellular compartments such as synapses, and help cells adapt to their complex environment (Figure 1).
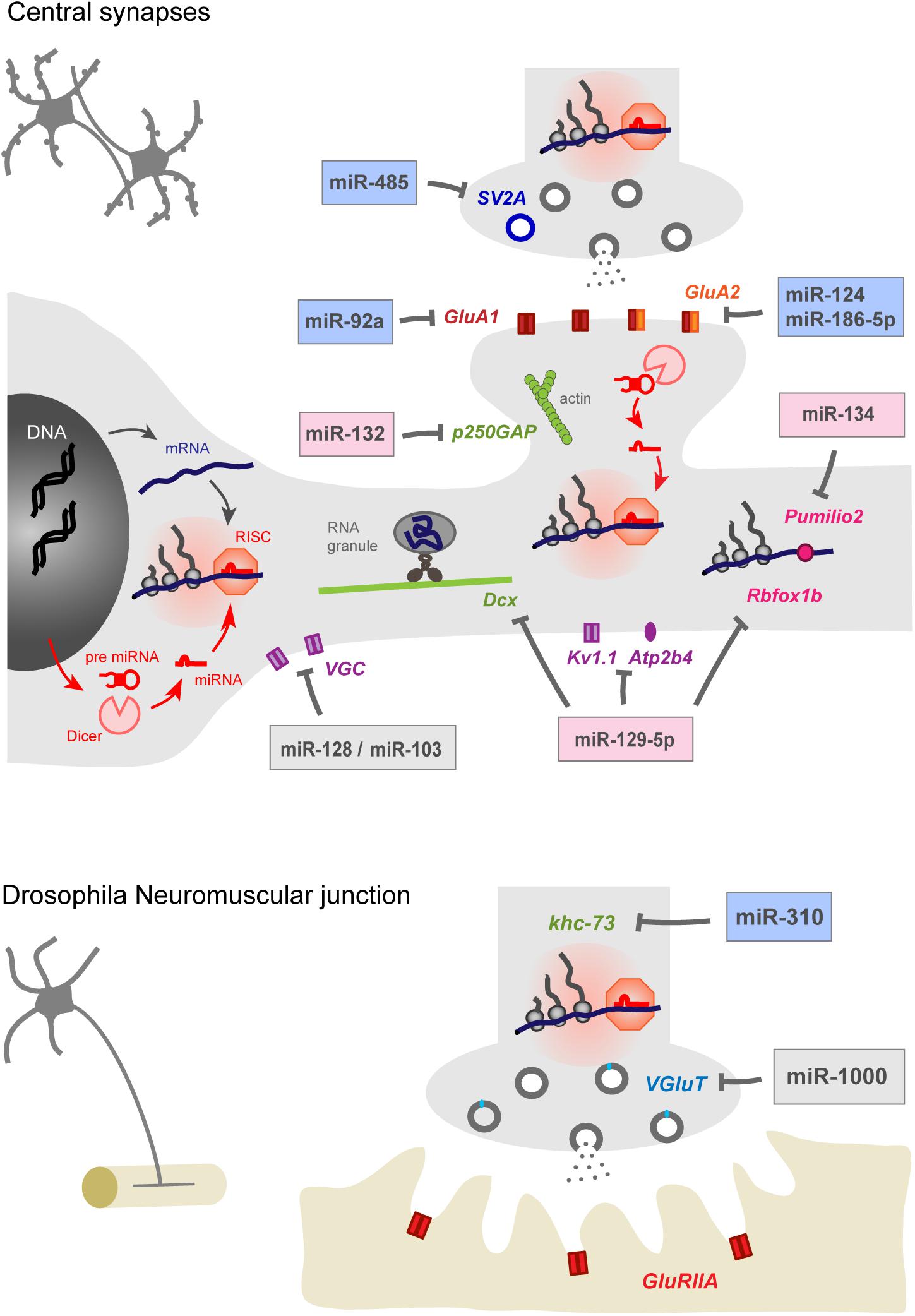
Figure 1. miRNAs contribute to homeostatic plasticity by controlling multiple effectors at central synapses of rodent models (top) and at the drosophila neuromuscular junction (bottom). Identified miRNAs target presynaptic proteins regulating neurotransmitter release, post-synaptic AMPAR subunits, cytoskeleton-related proteins, voltage gated ion channels, calcium pumps, and RNA-binding proteins. miRNAs may repress protein translation at the cell body or in pre- or post-synaptic compartments, thereby providing autonomy to subcellular compartments and regulating appropriate physiological parameters, such as pre- and post-synaptic strengths and membrane excitability. miRNAs involved in the homeostatic up- or down-regulation of neuronal activity are highlighted in blue and pink while those showing bidirectional regulations are indicated in gray. Note that the schematic for rodent central synapses summarizes collective data from several neuronal types (see Table 1 for details).
In this review, we present recent advances showing the contribution of several miRNAs in both cell-wide and compartmentalized forms of homeostatic plasticity through the regulation of the translation of multiple effectors (Figure 1 and Table 1). We first focus on homeostatic plasticity mechanisms that are regulated by miRNAs at the pre and post-synaptic levels, then discuss the impact of miRNAs on experience-dependent homeostatic synaptic plasticity (HSP) and neuronal excitability. Finally, we discuss several important questions that remain to be addressed, including the local versus global miRNA regulation and the implication of miRNAs in neuronal diseases.
miRNA-Dependent Control of Post-Synaptic Function During Homeostatic Synaptic Plasticity
One parameter that is commonly regulated to maintain synaptic homeostasis is the abundance of post-synaptic receptors. At excitatory synapses, the accumulation or depletion of synaptic AMPA-type glutamate receptors (AMPARs) has been well-characterized, mostly in primary neuronal cultures, following prolonged deprivation or elevation of neuronal activity, respectively. Depending on how neuronal activity is altered, this plasticity can be cell-wide or synapse-specific, and can engage different signaling pathways and combinations of AMPAR subunits (Vitureira and Goda, 2013; Fernandes and Carvalho, 2016). Interestingly, global pharmacological manipulations of neuronal activity known to induce HSP (Turrigiano et al., 1998; Thiagarajan et al., 2005; Sutton et al., 2006) alter the expression of several miRNAs in primary hippocampal cultures, which likely contribute to the proteome remodeling observed upon such conditions (Schanzenbächer et al., 2016, 2018).
Homeostatic Increase of Post-synaptic Strength in Response to Activity Deprivation
In rat cultured hippocampal neurons, the blockade for > 4 h of action potentials (APs) and NMDA receptors (NMDARs) with tetrodotoxin (TTX) and APV, respectively, leads to the local synthesis and synaptic insertion of AMPARs likely formed of GluA1 homomers (Sutton et al., 2006). This process is mediated by a decrease of miR-92a targeting the AMPAR subunit GluA1 in dendrites (Letellier et al., 2014). As a result, GluA1 translation is de-repressed and new AMPARs are targeted to synapses to support the increase in synaptic strength (Letellier et al., 2014). Importantly, this form of HSP is maintained in dendrites disconnected from the cell body (Sutton et al., 2006; Letellier et al., 2014), suggesting that transcription is not required and that the miR-92a-dependent GluA1 translation occurs locally. Intriguingly, incubating hippocampal neurons with TTX/APV for longer periods (>12 h) increases the expression of another miRNA, miR-124, which targets the GluA2 AMPAR subunit (Ho et al., 2014; Hou et al., 2015). While both miR-92a downregulation and miR-124 upregulation promote the expression of GluA2-lacking, calcium permeable AMPARs, the TTX/APV-induced elevation of miR-124 seems to rely on transcription-dependent mechanisms (Hou et al., 2015) and therefore may affect synaptic strengths more widely and uniformly as compared to miR-92a.
Interestingly, a 24 h activity-deprivation paradigm in cultured hippocampal neurons using non-competitive antagonists of AMPARs and NMDARs (GYKI-52466 and MK-801, respectively) does not affect miR-92a or miR-124 levels but rather downregulates miR-186-5p, a miRNA which also targets GluA2, thereby leading to the synaptic insertion of GluA2-containing AMPARs which are not permeable to calcium (Silva et al., 2019). Finally, a 24–48 h treatment with TTX alone induces the insertion of GluA2-containing AMPARs (Sutton et al., 2006; Gainey et al., 2009), but a specific regulation of this process by miRs has not been reported yet. Together, these studies suggest that neurons engage different miR-dependent pathways depending on the activity-deprivation paradigm, to produce a selective homeostatic compensation with regards to the AMPAR subunit composition that confers specific plastic properties to synapses (Diering and Huganir, 2018). A key point will be to determine the functional significance of these multiple miRNA-dependent regulations, and whether they extend to more physiological –in vivo- systems and to other brain regions.
Homeostatic Decrease of Post-synaptic Strengths in Response to Activity Elevation
The expression of some specific miRNAs is also altered following pharmacological manipulations to elevate network activity (Fiore et al., 2014; Rajman et al., 2017; Rocchi et al., 2019), suggesting that miRNAs bi-directionally adapt synaptic strengths across dendrites depending on network activity. In cultured hippocampal neurons, miR-134 elevation induced by the chronic (>24 h) pharmacological blockade of GABAA receptors (GABAARs) using picrotoxin (PTX) contributes to homeostatic synaptic downscaling by decreasing GluA2 surface expression and by promoting the elimination of excitatory synapses (Fiore et al., 2014). Specifically, miR-134 downregulates the RNA-binding protein Pumilio 2 which normally inhibits the polo-like kinase 2 (Plk2) pathway that promotes homeostatic downscaling through the degradation of the spine-associated protein RapGAP SPAR (SPAR) and the sequestration of the GluA2-interacting N-ethylmaleimide-sensitive fusion (NSF) protein (Seeburg et al., 2008; Evers et al., 2010). Curiously, other known targets of miR-134 including the protein kinase Limk1 which promotes spine development by regulating actin dynamics (Schratt et al., 2006) are not affected by the PTX treatment (Fiore et al., 2014), suggesting a selective effect. Interestingly, miR-134 is also upregulated in the temporal lobe neocortex of patients with epilepsy (Jimenez-Mateos et al., 2012). While it is currently unknown whether Pumilio 2 is downregulated in this condition, Limk1 expression level is decreased, which could result in smaller dendritic spines to dampen hyperactivity and may represent some homeostatic adaptation (Jimenez-Mateos et al., 2012). Surprisingly, however, silencing miR-134 in mice using antagomirs suppresses seizures and has a neuroprotective action (Jimenez-Mateos et al., 2012), suggesting that abnormal increased levels of miR-134 may rather promote epilepsy. Therefore, despite the therapeutical potential of miR-134 antagomirs in the context of epilepsy, more investigations are required to understand the exact mode of action of miR-134 in vivo.
In another study, miR-129-5p elevation was also shown to be required for the PTX-induced downscaling of synaptic strength, by promoting the downregulation of the calcium pump Atp2b4 and the microtubule-associated protein doublecortin (Dcx) (Rajman et al., 2017). Furthermore, the authors uncover a functional interaction between miR-129-5p and the RNA binding protein Rbfox1, which normally promotes the expression of both Atp2b4 and Dcx through their 3′ UTR. Upon PTX-induced synaptic scaling, Rbfox1 expression is downregulated in a miR-129-5p manner, thereby allowing the repression of Atp2b4, Dcx and possibly other synaptic genes. However, how miR-134 and miR-129-5p work in conjunction to drive homeostatic downscaling triggered by GABAR blockade and whether other regulated miRNAs including miR-132, miR-495, miR-543-3p, or miR-218 contribute to this process (Rajman et al., 2017; Rocchi et al., 2019) remain to be investigated.
miRNA-Dependent Control of Presynaptic Function During Homeostatic Synaptic Plasticity
While many studies have uncovered post-synaptic mechanisms for homeostatic synaptic plasticity, there is strong evidence that neurons can also regulate the number and efficacy of presynaptic release sites to compensate for prolonged perturbations in network activity (Thiagarajan et al., 2005; Jakawich et al., 2010; Lindskog et al., 2010; Vitureira et al., 2011; Davis and Müller, 2015). Several miRNAs likely contribute to this process by targeting presynaptic proteins. For instance, in cultured hippocampal neurons, miR-485 is upregulated following the chronic elevation of neuronal activity (>5 days) using bicuculline and 4-aminopyridine (4-AP) to block GABAARs and potassium channels, respectively (Cohen et al., 2011). miR-485 targets the synaptic vesicle protein 2A (SV2A) which is known to facilitate neurotransmitter release through an interaction with synaptotagmin (Custer, 2006; Yao et al., 2010), and which is downregulated following seizures in the hippocampus, thus possibly representing a homeostatic mechanism (van Vliet et al., 2009). Surprisingly, miR-485 expression does not downregulate presynaptic neurotransmitter release per se but rather decreases the number of functional synapses, as evidenced by a decreased density of PSD-95 and AMPAR clusters, suggesting a functional crosstalk between pre and post-synaptic elements. In any case, the mechanism by which miR-485 adapts the number of functional synapses in response to elevated network activity (and possibly during epilepsy) remains to be clarified, as other targets of miR-485 may also contribute to this process (Cohen et al., 2011).
One model system that has been extensively studied in the context of presynaptic homeostatic plasticity is the drosophila neuromuscular junction, where experimentally reducing the sensitivity or the expression of post-synaptic glutamate receptors is precisely balanced by an increase in glutamate release through retrograde signaling (Petersen et al., 1997; Frank et al., 2006; Frank, 2014; Davis and Müller, 2015). The miR-310-313 cluster contributes to this process most likely by targeting the kinesin family member khc-73 in motor neurons (Tsurudome et al., 2010). Specifically, overexpressing miR-310 or knocking-down khc-73 in motor neurons both inhibit the homeostatic increase in quantal content normally observed in GluRIIA mutants.
miR-1000 is another drosophila miRNA which modulates glutamate release by down-regulating the expression of the glutamate transporter VGlut (Verma et al., 2015). miR-1000 genetic deletion enhances VGlut expression, resulting in an excess of glutamate release through a higher number of active boutons, which are also bigger in size. Interestingly, miR-1000 expression level in the drosophila visual system is regulated in an homeostatic manner by visual input. Indeed, miR-1000 transcript levels are significantly reduced in dark-reared flies, raising the possibility that glutamate release is enhanced and compensates for reduced sensory input. In contrast, flies reared in constant light show increased miR-1000 expression compared to animals reared under a normal light-dark cycle, suggesting a reduction of glutamate release to compensate for a prolonged elevation of sensory input (Verma et al., 2015). Importantly, the failure of miR-1000-dependent regulation of glutamate release results in excitotoxicity and reduced neuron survival. While miR-1000 is not expressed in mammals, the seed-similar miRNA miR-137 is expressed in mouse hippocampal neurons and may similarly regulate VGluT2 (Verma et al., 2015) in addition to its post-synaptic target GluA1 (Olde Loohuis et al., 2015). Interestingly, miR-137 has been genetically associated with schizophrenia and miR-137 overexpression in the mouse dentate gyrus impairs presynaptic plasticity and hippocampus-dependent learning and memory through the regulation of the presynaptic proteins synaptotagmin-1, complexin-1, and NSF (Siegert et al., 2015).
miRNA-Dependent Control of Experience-Dependent Homeostatic Synaptic Plasticity In Vivo
Besides compensating for global perturbations of network activity, whether induced pharmacologically or genetically (see above), there is evidence that HSP also contributes to experience-dependent plasticity and refinement of developing synaptic circuits. In such situations, the strengthening of active inputs is compensated by the weakening of less active inputs on the target cell, presumably through competition-based mechanisms; this eventually leads to the selective stabilization of specific inputs at the expense of others. This activity-dependent process has been extensively studied in the mammalian visual cortex where a population of neurons respond to the two eyes. Occluding the vision of one eye during a critical developmental period (monocular deprivation paradigm), produces a loss of responsiveness of binocular neurons to the deprived eye which is precisely balanced by a corresponding homeostatic increase in response to the undeprived eye (ocular dominance shift), thus preserving the net visual drive for each neuron (Mrsic-Flogel et al., 2007; Kaneko et al., 2008; Ranson et al., 2012; Kaneko and Stryker, 2017).
Interestingly, the expression of some specific miRNAs is altered following sensory deprivation in the visual cortex, and may contribute to the homeostatic component of the ocular dominance shift. Among them, miR-132, is decreased after monocular deprivation or dark rearing (Mellios et al., 2011; Tognini et al., 2011). Inhibiting miR-132 through the injection of a miRNA-sponge-expressing lentivirus (Mellios et al., 2011) or counteracting miR-132 reduction by infusing a miR-132 mimic (Tognini et al., 2011) both prevent the ocular dominance plasticity shift induced by monocular sensory deprivation, suggesting that miR-132 drop is necessary to both weaken the deprived visual input and strengthen the undeprived input. At the cellular level, such a homeostatic balance between active and inactive inputs may involve the de-repression of the miR-132 target p250GAP, a Rho family GTPase that regulates spine morphology and remodeling through Rac1 inhibition (Wayman et al., 2008; Edbauer et al., 2010; Impey et al., 2010; Remenyi et al., 2013) and which has been implicated in epileptogenesis process (Yuan et al., 2016). In one possible mechanism, sensory-deprived synaptic inputs could depress and shrink through the GTPase p250GAP/Rac1 pathway while more active synapses would get strengthened and grow in size. Importantly, the differential regulation of active versus inactive synapses in the same post-synaptic cell suggests the existence of local signaling within dendrites (Oh et al., 2015; El-Boustani et al., 2018; Letellier et al., 2019) to which miRNAs might contribute.
miRNA-Dependent Control of Intrinsic Excitability
In addition to controlling synapse number and efficacy to compensate for local or global activity perturbations, miRNAs can directly regulate membrane excitability, thereby controlling the probability that synaptic inputs trigger action potentials in dendrites and/or axon. For instance, miR-128, which is highly abundant in the mammalian brain, regulates neuronal excitability and motor behavior in the mouse by downregulating the expression of various ion channels and signaling components of the extracellular signal-regulated kinase ERK2 network (Tan et al., 2013). Interestingly, a reduction in miR-128 expression causes increased motor activity and fatal epilepsy in mice. While it would be interesting to see to what extent variations in network activity affect miR-128 expression, these finding suggests that the level of miR-128 is tightly regulated to maintain the neuronal firing rate (Tan et al., 2013).
miRNAs have been involved in the control of intrinsic excitability through the regulation of voltage-gated calcium channel. In the context of chronic pain, miR-103 regulates the expression of the three subunits of the Cav1.2-comprising L-type calcium channel in rat spinal cord neurons, thereby modulating sensitization to pain. Moreover, miR-103 was downregulated in neuropathic rats and miR-103 intrathecal applications successfully relieved pain, thus identifying miR-103 as a possible therapeutic target in neuropathic chronic pain (Favereaux et al., 2011).
Another example is miR-129 which not only controls homeostatic downscaling by targeting Atp2b4 and Dcx (see above; Rajman et al., 2017) but also regulates the dendritic expression of the Shaker-like potassium channel Kv1.1 (Sosanya et al., 2013). Kv1.1 is a dendrotoxin-sensitive voltage gated potassium channel that is expressed in the axon but also in dendrites (Raab-Graham et al., 2006; Sosanya et al., 2013). A proposed mechanism involves miR-129 and the mRNA binding protein HuD which binds to Kv1.1 mRNA, depending on mTORC1 kinase activity to repress or enhance Kv1.1 expression, respectively (Sosanya et al., 2013). Interestingly, miR-129-mediated translational repression of Kv1.1 is enhanced 3 weeks after status epilepticus in rats, suggesting that miR-129 promotes excitability by targeting Kv1.1 and that this mechanism is tightly regulated to maintain neuronal homeostasis (Sosanya et al., 2015). However, that the same miRNA can promote both synaptic downscaling and dendritic excitability suggests the involvement of complex regulations to orchestrate the homeostatic response in time and space.
There is also evidence that the RNA-binding protein Pumilio 2, a key miR-134 target involved in PTX-induced downscaling (see above), controls the homeostasis of membrane excitability in cultured cortical neurons. Indeed, Pumilio 2 expression at the cellular level is increased upon elevating neuronal activity and thereby suppresses translation of the voltage-gated sodium channel transcript Nav1.6 to decrease intrinsic excitability (Driscoll et al., 2013). However, this is at odds with the fact that the prolonged elevation of neuronal activity reduces Pumilio 2 expression locally in the dendritic compartment in a miR-134-dependent manner to promote downscaling (Fiore et al., 2014). Therefore, it is unclear how elevating activity can simultaneously promote the up- and down-regulation of Pumilio 2 to cause decrease in membrane excitability and miR-134-dependent synaptic downscaling, respectively. This suggests the existence of compartmentalized mechanisms, where Pumilio 2 expression might be differently regulated in the cell body versus dendrites, but this remains to be investigated experimentally.
Do miRNAs Regulate Homeostatic Plasticity Locally?
What makes miRNAs interesting candidates in the regulation of synaptic plasticity is that they potentially control protein synthesis in remote subcellular compartments such as dendrites and synapses to provide an appropriate and targeted physiological response. While this idea has not been directly tested in the context of HSP, some of the studies discussed above provide indirect evidence that miRNA-dependent homeostatic plasticity requires local regulations, supporting the concept that synapses do not always adapt uniformly and that homeostatic plasticity can operate within autonomous subcellular compartments, and down to single synapses (Thiagarajan et al., 2005; Sutton et al., 2006; Echegoyen et al., 2007; Aoto et al., 2008; Branco et al., 2008; Kim and Tsien, 2008; Maghsoodi et al., 2008; Beique et al., 2011; Wang et al., 2019).
In support of a role for miRNAs in regulating the function of local compartments like synapses, subcellular fractionation and in situ hybridization experiments revealed that several miRNAs are present in dendrites, axons or even synapses and that neuronal activity regulates both their abundance and function (Kye et al., 2007; Lugli et al., 2008; Schratt, 2009; Siegel et al., 2009; Natera-Naranjo et al., 2010). Interestingly, the distribution of miRNAs seems to parallel the distribution of their cognate target mRNAs (Kye et al., 2007); such a spatial proximity may enable the efficient regulation of local protein translation to serve a specific function at the right time and place (Kosik, 2016; Park et al., 2019).
What are the mechanisms by which neuronal activity regulates the local amount of miRNAs? While the activity-dependent expression of several miRNAs including miR-132, miR-134 and miR-124 may be regulated at the transcriptional level by transcription factors such like CREB, Mef2 or EVI1 (Fiore et al., 2009; Nudelman et al., 2010; Remenyi et al., 2010; Hou et al., 2015), there is evidence that neuronal activity directly controls the local processing of pre-miRNAs into mature miRNAs at the level of single dendritic spines. Using a fluorescent pre-miRNA sensor to probe Dicer activity, it was recently shown that the local stimulation of single spines through glutamate uncaging promotes the maturation of miR-181a in a NMDAR-dependent manner, leading to the local repression of CamKIIα synthesis (Sambandan et al., 2017). Furthermore, the local abundance of miR-134, previously implicated in PTX-induced downscaling (Fiore et al., 2014), varies depending on spine maturation and activity, while BDNF local stimulation leads to a decrease in the number of miR-134 copies present at the neck of spines (Park et al., 2019). In addition to the local control of miRNA maturation through Dicer, neuronal activity regulate the turnover of the RISC complex itself, which could possibly impact miR-dependent local protein translation in a non-specific way. In particular, the RISC component MOV10 is degraded upon NMDAR activation, which may result in the release of miRNAs from their mRNA targets and de-repress local protein translation (Chendrimada et al., 2007; Banerjee et al., 2009).
However, one important question remains: how specific activity variations can regulate the local expression and/or function of some specific miRNAs and not others in order to achieve the appropriate physiological response? Potential mechanisms involve specific interactions with cognate mRNA targets which could protect miRNAs from degradation (Pitchiaya et al., 2017), storage in P-bodies whose dendritic location is regulated by neuronal activity (Cougot et al., 2008), or interaction with circular RNAs serving as natural miRNA-sponges (Hansen et al., 2013).
Conclusion
There is strong evidence that miRNAs contribute to homeostatic plasticity and associated neurological disorders including epilepsy, neuropsychiatric, and neurodegenerative diseases (Mellios and Sur, 2012; Henshall et al., 2016; Quinlan et al., 2017; Rajman et al., 2017) (Table 1). However, despite some recent progress, important questions remain. In particular, the signaling pathways linking physiological synaptic activity variations to miRNA function, trafficking, and turnover remain largely unknown, as most of the current knowledge relies on pharmacological manipulations in culture systems. Some effort will thus be required to investigate the role of identified miRNAs in more physiologically relevant systems; the development of new probes and live-imaging tools to track individual RNAs and investigate translation dynamics (Park et al., 2014; Wang et al., 2016; Wu et al., 2016; Yan et al., 2016) should provide new insights into these mechanisms. Equally important will be to investigate whether they regulate inhibitory synapses which also undergo homeostatic plasticity (Kilman et al., 2002; Peng et al., 2010; Rannals and Kapur, 2011), and whether they contribute to the neuron-glia interactions involved in homeostatic plasticity (Stellwagen and Malenka, 2006; Letellier et al., 2016). Finally, considering that miRNAs also control LTP and depression (Hu and Li, 2017), it will be interesting to investigate whether and how miRNAs enable the integration in time and space of both Hebbian and homeostatic plasticity. A better understanding of the miRNA function in synaptic plasticity and the possible links with pathologies will be very helpful in refining promising therapeutic strategies (Wen, 2016; Angelucci et al., 2019).
Author Contributions
ML, OT, AF, and SD wrote the manuscript. SD built the figure.
Funding
This work received funding from the Centre National de la Recherche Scientifique, Agence Nationale pour la Recherche (grant «SynSpe» ANR-13-PDOC-0012-01), Conseil Régional Aquitaine («SiMoDyn»), Investissements d’Avenir Labex BRAIN ANR-10-LABX-43, and Fondation pour la Recherche Médicale («Equipe FRM» DEQ20160334916).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Abu-Elneel, K., Liu, T., Gazzaniga, F. S., Nishimura, Y., Wall, D. P., Geschwind, D. H., et al. (2008). Heterogeneous dysregulation of microRNAs across the autism spectrum. Neurogenetics 9, 153–161. doi: 10.1007/s10048-008-0133-5
Angelucci, F., Cechova, K., Valis, M., Kuca, K., Zhang, B., and Hort, J. (2019). MicroRNAs in Alzheimer’s Disease: diagnostic markers or therapeutic agents? Front. Pharmacol. 10:665. doi: 10.3389/fphar.2019.00665
Aoto, J., Nam, C. I., Poon, M. M., Ting, P., and Chen, L. (2008). Synaptic signaling by all-trans retinoic acid in homeostatic synaptic plasticity. Neuron 60, 308–320. doi: 10.1016/j.neuron.2008.08.012
Banerjee, S., Neveu, P., and Kosik, K. S. (2009). A coordinated local translational control point at the synapse involving relief from silencing and MOV10 degradation. Neuron 64, 871–884. doi: 10.1016/j.neuron.2009.11.023
Bartel, D. P. (2009). MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233. doi: 10.1016/j.cell.2009.01.002
Beique, J.-C., Na, Y., Kuhl, D., Worley, P. F., and Huganir, R. L. (2011). Arc-dependent synapse-specific homeostatic plasticity. Proc. Natl. Acad. Sci. U.S.A. 108, 816–821. doi: 10.1073/pnas.1017914108
Blackman, M. P., Djukic, B., Nelson, S. B., and Turrigiano, G. G. (2012). A critical and cell-autonomous role for MeCP2 in synaptic scaling up. J. Neurosci. 32, 13529–13536. doi: 10.1523/JNEUROSCI.3077-12.2012
Branco, T., Staras, K., Darcy, K. J., and Goda, Y. (2008). Local dendritic activity sets release probability at hippocampal synapses. Neuron 59, 475–485. doi: 10.1016/j.neuron.2008.07.006
Campos-Melo, D., Hawley, Z. C. E., and Strong, M. J. (2018). Dysregulation of human NEFM and NEFH mRNA stability by ALS-linked miRNAs. Mol. Brain 11:43. doi: 10.1186/s13041-018-0386-3
Chang, M. C., Park, J. M., Pelkey, K. A., Grabenstatter, H. L., Xu, D., Linden, D. J., et al. (2010). Narp regulates homeostatic scaling of excitatory synapses on parvalbumin-expressing interneurons. Nat. Neurosci. 13, 1090–1097. doi: 10.1038/nn.2621
Chendrimada, T. P., Finn, K. J., Ji, X., Baillat, D., Gregory, R. I., Liebhaber, S. A., et al. (2007). MicroRNA silencing through RISC recruitment of eIF6. Nature 447, 823–828. doi: 10.1038/nature05841
Cheng, T. L., Wang, Z., Liao, Q., Zhu, Y., Zhou, W. H., Xu, W., et al. (2014). MeCP2 suppresses nuclear MicroRNA processing and dendritic growth by regulating the DGCR8/drosha complex. Dev. Cell. 28, 547–560. doi: 10.1016/j.devcel.2014.01.032
Cohen, J. E., Lee, P. R., Chen, S., Li, W., and Fields, R. D. (2011). MicroRNA regulation of homeostatic synaptic plasticity. Proc. Natl. Acad. Sci. U.S.A. 108, 11650–11655. doi: 10.1073/pnas.1017576108
Correa, S. A. L., Hunter, C. J., Palygin, O., Wauters, S. C., Martin, K. J., McKenzie, C., et al. (2012). MSK1 regulates homeostatic and experience-dependent synaptic plasticity. J. Neurosci. 32, 13039–13051. doi: 10.1523/JNEUROSCI.0930-12.2012
Cougot, N., Bhattacharyya, S. N., Tapia-Arancibia, L., Bordonné, R., Filipowicz, W., Bertrand, E., et al. (2008). Dendrites of mammalian neurons contain specialized P-body-like structures that respond to neuronal activation. J. Neurosci. 28, 13793–13804. doi: 10.1523/JNEUROSCI.4155-08.2008
Cuellar, T. L., Davis, T. H., Nelson, P. T., Loeb, G. B., Harfe, B. D., Ullian, E., et al. (2008). Dicer loss in striatal neurons produces behavioral and neuroanatomical phenotypes in the absence of neurodegeneration. Proc. Natl. Acad. Sci. U.S.A. 105, 5614–5619. doi: 10.1073/pnas.0801689105
Custer, K. L. (2006). Synaptic vesicle protein 2 enhances release probability at quiescent synapses. J. Neurosci. 26, 1303–1313. doi: 10.1523/JNEUROSCI.2699-05.2006
Davis, G. W., and Müller, M. (2015). Homeostatic control of presynaptic neurotransmitter release. Annu. Rev. Physiol. 77, 251–270. doi: 10.1146/annurev-physiol-021014-071740
Dickman, D. K., and Davis, G. W. (2009). The schizophrenia susceptibility gene dysbindin controls synaptic homeostasis. Science 326, 1127–1130. doi: 10.1126/science.1179685
Diering, G. H., and Huganir, R. L. (2018). The AMPA receptor code of synaptic plasticity. Neuron 100, 314–329. doi: 10.1016/j.neuron.2018.10.018
Diering, G. H., Nirujogi, R. S., Roth, R. H., Worley, P. F., Pandey, A., and Huganir, R. L. (2017). Homer1a drives homeostatic scaling-down of excitatory synapses during sleep. Science 355, 511–515. doi: 10.1126/science.aai8355
Driscoll, H. E., Muraro, N. I., He, M., and Baines, R. A. (2013). Pumilio-2 Regulates translation of Nav1.6 to mediate homeostasis of membrane excitability. J. Neurosci. 33, 9644–9654. doi: 10.1523/JNEUROSCI.0921-13.2013
Echegoyen, J., Neu, A., Graber, K. D., and Soltesz, I. (2007). Homeostatic plasticity studied using in vivo hippocampal activity-blockade: synaptic scaling, intrinsic plasticity and age-dependence. PLoS One 2:e700. doi: 10.1371/journal.pone.0000700
Edbauer, D., Neilson, J. R., Foster, K. A., Wang, C.-F., Seeburg, D. P., Batterton, M. N., et al. (2010). Regulation of synaptic structure and function by FMRP-associated microRNAs miR-125b and miR-132. Neuron 65, 373–384. doi: 10.1016/j.neuron.2010.01.005
El-Boustani, S., Ip, J. P. K. K., Breton-Provencher, V., Knott, G. W., Okuno, H., Bito, H., et al. (2018). Locally coordinated synaptic plasticity of visual cortex neurons in vivo. Science 360, 1349–1354. doi: 10.1126/science.aao0862
Evers, D. M., Matta, J. A., Hoe, H. S., Zarkowsky, D., Lee, S. H., Isaac, J. T., et al. (2010). Plk2 attachment to NSF induces homeostatic removal of GluA2 during chronic overexcitation. Nat. Neurosci. 13, 1199–1207. doi: 10.1038/nn.2624
Favereaux, A., Thoumine, O., Bouali-Benazzouz, R., Roques, V., Papon, M.-A., Salam, S. A., et al. (2011). Bidirectional integrative regulation of Cav1.2 calcium channel by microRNA miR-103: role in pain. EMBO J. 30, 3830–3841. doi: 10.1038/emboj.2011.249
Fernandes, D., and Carvalho, A. L. (2016). Mechanisms of homeostatic plasticity in the excitatory synapse. J. Neurochem. 139, 973–996. doi: 10.1111/jnc.13687
Fernandez-Moya, S. M., Bauer, K. E., and Kiebler, M. A. (2014). Meet the players: local translation at the synapse. Front. Mol. Neurosci. 7:84. doi: 10.3389/fnmol.2014.00084
Filipowicz, W., Bhattacharyya, S. N., and Sonenberg, N. (2008). Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat. Rev. Genet. 9, 102–114. doi: 10.1038/nrg2290
Fineberg, S. K., Kosik, K. S., and Davidson, B. L. (2009). MicroRNAs Potentiate Neural Development. Neuron 64, 303–309. doi: 10.1016/j.neuron.2009.10.020
Fiore, R., Khudayberdiev, S., Christensen, M., Siegel, G., Flavell, S. W., Kim, T., et al. (2009). Mef2-mediated transcription of the miR379–410 cluster regulates activity-dependent dendritogenesis by fine-tuning Pumilio2 protein levels. EMBO J. 28, 697–710. doi: 10.1038/emboj.2009.10
Fiore, R., Rajman, M., Schwale, C., Bicker, S., Antoniou, A., Bruehl, C., et al. (2014). MiR -134-dependent regulation of pumilio-2 is necessary for homeostatic synaptic depression. EMBO J. 33, 2231–2246. doi: 10.15252/embj.201487921
Fiorenza, A., Lopez-Atalaya, J. P., Rovira, V., Scandaglia, M., Geijo-Barrientos, E., and Barco, A. (2016). Blocking miRNA biogenesis in adult forebrain neurons enhances seizure susceptibility, fear memory, and food intake by increasing neuronal responsiveness. Cereb. Cortex 26, 1619–1633. doi: 10.1093/cercor/bhu332
Follert, P., Cremer, H., and Béclin, C. (2014). MicroRNAs in brain development and function: a matter of flexibility and stability. Front. Mol. Neurosci. 7:5. doi: 10.3389/fnmol.2014.00005
Frank, C. A. (2014). Homeostatic plasticity at the drosophila neuromuscular junction. Neuropharmacology 78, 63–74. doi: 10.1016/j.neuropharm.2013.06.015
Frank, C. A., Kennedy, M. J., Goold, C. P., Marek, K. W., and Davis, G. W. (2006). Mechanisms underlying the rapid induction and sustained expression of synaptic homeostasis. Neuron 52, 663–677. doi: 10.1016/j.neuron.2006.09.029
Friedman, R. C., Farh, K. K.-H., Burge, C. B., and Bartel, D. P. (2008). Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105. doi: 10.1101/gr.082701.108
Gainey, M. A., Hurvitz-Wolff, J. R., Lambo, M. E., and Turrigiano, G. G. (2009). Synaptic scaling requires the GluR2 subunit of the AMPA receptor. J. Neurosci. 29, 6479–6489. doi: 10.1523/JNEUROSCI.3753-08.2009
Gao, M., Sossa, K., Song, L., Errington, L., Cummings, L., Hwang, H., et al. (2010). A Specific requirement of Arc/Arg3.1 for visual experience-induced homeostatic synaptic plasticity in mouse primary visual cortex. J. Neurosci. 30, 7168–7178. doi: 10.1523/jneurosci.1067-10.2010
Geekiyanage, H., and Chan, C. (2011). Micro RNA-137/181c regulates serine palmitoyltransferase and in turn amyloid β novel targets in sporadic Alzheimer’s disease. J. Neurosci. 31, 14820–14830. doi: 10.1523/JNEUROSCI.3883-11.2011
Gilbert, J., Shu, S., Yang, X., Lu, Y., Zhu, L.-Q., and Man, H.-Y. (2016). β-Amyloid triggers aberrant over-scaling of homeostatic synaptic plasticity. Acta Neuropathol. Commun. 4:131. doi: 10.1186/s40478-017-0423-y
Giraldez, A. J. (2005). MicroRNAs regulate brain morphogenesis in zebrafish. Science 308, 833–838. doi: 10.1126/science.1109020
Hansen, T. B., Jensen, T. I., Clausen, B. H., Bramsen, J. B., Finsen, B., Damgaard, C. K., et al. (2013). Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388. doi: 10.1038/nature11993
Hengen, K. B., Lambo, M. E., Van Hooser, S. D., Katz, D. B., and Turrigiano, G. G. (2013). Firing rate homeostasis in visual cortex of freely behaving rodents. Neuron 80, 335–342. doi: 10.1016/j.neuron.2013.08.038
Hengen, K. B., Torrado Pacheco, A., McGregor, J. N., Van Hooser, S. D., and Turrigiano, G. G. (2016). Neuronal firing rate homeostasis is inhibited by sleep and promoted by wake. Cell 165, 180–191. doi: 10.1016/j.cell.2016.01.046
Henshall, D. C., Hamer, H. M., Pasterkamp, R. J., Goldstein, D. B., Kjems, J., Prehn, J. H. M., et al. (2016). MicroRNAs in epilepsy: pathophysiology and clinical utility. Lancet Neurol. 15, 1368–1376. doi: 10.1016/S1474-4422(16)30246-0
Ho, V. M., Dallalzadeh, L. O., Karathanasis, N., Keles, M. F., Vangala, S., Grogan, T., et al. (2014). GluA2 mRNA distribution and regulation by miR-124 in hippocampal neurons. Mol. Cell. Neurosci. 61, 1–12. doi: 10.1016/j.mcn.2014.04.006
Hou, Q., Ruan, H., Gilbert, J., Wang, G., Ma, Q., Yao, W.-D., et al. (2015). MicroRNA miR124 is required for the expression of homeostatic synaptic plasticity. Nat. Commun. 6:10045. doi: 10.1038/ncomms10045
Hu, Z., and Li, Z. (2017). miRNAs in synapse development and synaptic plasticity. Curr. Opin Neurobiol. 45, 24–31. doi: 10.1016/j.conb.2017.02.014
Ibata, K., Sun, Q., and Turrigiano, G. G. (2008). Rapid synaptic scaling induced by changes in postsynaptic firing. Neuron 57, 819–826. doi: 10.1016/j.neuron.2008.02.031
Impey, S., Davare, M., Lasiek, A., Fortin, D., Ando, H., Varlamova, O., et al. (2010). An activity-induced microRNA controls dendritic spine formation by regulating Rac1-PAK signaling. Mol. Cell. Neurosci. 43, 146–156. doi: 10.1016/j.mcn.2009.10.005
Jakawich, S. K., Nasser, H. B., Strong, M. J., McCartney, A. J., Perez, A. S., Rakesh, N., et al. (2010). Local presynaptic activity gates homeostatic changes in presynaptic function driven by dendritic BDNF synthesis. Neuron 68, 1143–1158. doi: 10.1016/j.neuron.2010.11.034
Jimenez-Mateos, E. M., Engel, T., Merino-Serrais, P., McKiernan, R. C., Tanaka, K., Mouri, G., et al. (2012). Silencing microRNA-134 produces neuroprotective and prolonged seizure-suppressive effects. Nat. Med. 18, 1087–1094. doi: 10.1038/nm.2834
Kaalund, S. S., Venø, M. T., Bak, M., Møller, R. S., Laursen, H., Madsen, F., et al. (2014). Aberrant expression of miR-218 and miR-204 in human mesial temporal lobe epilepsy and hippocampal sclerosis-convergence on axonal guidance. Epilepsia 55, 2017–2027. doi: 10.1111/epi.12839
Kaneko, M., Stellwagen, D., Malenka, R. C., and Stryker, M. P. (2008). Tumor necrosis factor-alpha mediates one component of competitive, experience-dependent plasticity in developing visual cortex. Neuron 58, 673–680. doi: 10.1016/j.neuron.2008.04.023
Kaneko, M., and Stryker, M. P. (2017). Homeostatic plasticity mechanisms in mouse V1. Philos. Trans. R. Soc. B Biol. Sci. 372:20160504. doi: 10.1098/rstb.2016.0504
Keck, T., Keller, G. B., Jacobsen, R. I., Eysel, U. T., Bonhoeffer, T., and Hübener, M. (2013). Synaptic scaling and homeostatic plasticity in the mouse visual cortex in vivo. Neuron 80, 327–334. doi: 10.1016/j.neuron.2013.08.018
Kilman, V., van Rossum, M. C. W., and Turrigiano, G. G. (2002). Activity deprivation reduces miniature IPSC amplitude by decreasing the number of postsynaptic GABA(A) receptors clustered at neocortical synapses. J. Neurosci. 22, 1328–1337. doi: 10.1523/jneurosci.22-04-01328.2002
Kim, A. H., Reimers, M., Maher, B., Williamson, V., McMichael, O., McClay, J. L., et al. (2010). MicroRNA expression profiling in the prefrontal cortex of individuals affected with schizophrenia and bipolar disorders. Schizophr. Res. 124, 183–189. doi: 10.1016/j.schres.2010.07.002
Kim, J., Inoue, K., Ishii, J., Vanti, W. B., Voronov, S. V., Murchison, E., et al. (2007). A MicroRNA feedback circuit in midbrain dopamine neurons. Science 317, 1220–1224. doi: 10.1126/science.1140481
Kim, J., and Tsien, R. W. (2008). Synapse-specific adaptations to inactivity in hippocampal circuits achieve homeostatic gain control while dampening network reverberation. Neuron 58, 925–937. doi: 10.1016/j.neuron.2008.05.009
Kim, J., Yoon, H., Chung, D., Brown, J. L., Belmonte, K. C., and Kim, J. (2016). miR-186 is decreased in aged brain and suppresses BACE1 expression. J. Neurochem. 137, 436–445. doi: 10.1111/jnc.13507
Kosik, K. S. (2006). The neuronal microRNA system. Nat. Rev. Neurosci. 7, 911–920. doi: 10.1038/nrn2037
Kosik, K. S. (2016). Life at low copy number: how dendrites manage with so few mRNAs. Neuron 92, 1168–1180. doi: 10.1016/j.neuron.2016.11.002
Kwon, E., Wang, W., and Tsai, L. H. (2013). Validation of schizophrenia-associated genes CSMD1, C10orf26, CACNA1C and TCF4 as miR-137 targets. Mol. Psychiatry 18, 11–12. doi: 10.1038/mp.2011.170
Kye, M.-J., Liu, T., Levy, S. F., Xu, N. L., Groves, B. B., Bonneau, R., et al. (2007). Somatodendritic microRNAs identified by laser capture and multiplex RT-PCR. RNA 13, 1224–1234. doi: 10.1261/rna.480407
Lau, P., Bossers, K., Janky, R., Salta, E., Frigerio, C. S., Barbash, S., et al. (2013). Alteration of the microRNA network during the progression of Alzheimer’s disease. EMBO Mol. Med. 5, 1613–1634. doi: 10.1002/emmm.201201974
Letellier, M., Elramah, S., Mondin, M., Soula, A., Penn, A., Choquet, D., et al. (2014). miR-92a regulates expression of synaptic GluA1-containing AMPA receptors during homeostatic scaling. Nat. Neurosci. 17, 1040–1042. doi: 10.1038/nn.3762
Letellier, M., Levet, F., Thoumine, O., and Goda, Y. (2019). Differential role of pre- and postsynaptic neurons in the activity-dependent control of synaptic strengths across dendrites. PLoS Biol. 17:e2006223. doi: 10.1371/journal.pbio.2006223
Letellier, M., Park, Y. K., Chater, T. E., Chipman, P. H., Gautam, S. G., Oshima-Takago, T., et al. (2016). Astrocytes regulate heterogeneity of presynaptic strengths in hippocampal networks. Proc. Natl. Acad. Sci. U.S.A. 113, E2685–E2694. doi: 10.1073/pnas.1523717113
Lindskog, M., Li, L., Groth, R. D., Poburko, D., Thiagarajan, T. C., Han, X., et al. (2010). Postsynaptic GluA1 enables acute retrograde enhancement of presynaptic function to coordinate adaptation to synaptic inactivity. Proc. Natl. Acad. Sci. U.S.A. 107, 21806–21811. doi: 10.1073/pnas.1016399107
Lugli, G., Torvik, V. I., Larson, J., and Smalheiser, N. R. (2008). Expression of microRNAs and their precursors in synaptic fractions of adult mouse forebrain. J. Neurochem. 106, 650–661. doi: 10.1111/j.1471-4159.2008.05413.x
Maghsoodi, B., Poon, M. M., Nam, C. I., Aoto, J., Ting, P., and Chen, L. (2008). Retinoic acid regulates RARalpha-mediated control of translation in dendritic RNA granules during homeostatic synaptic plasticity. Proc. Natl. Acad. Sci. U.S.A. 105, 16015–16020. doi: 10.1073/pnas.0804801105
Mellios, N., Sugihara, H., Castro, J., Banerjee, A., Le, C., Kumar, A., et al. (2011). miR-132, an experience-dependent microRNA, is essential for visual cortex plasticity. Nat. Neurosci. 14, 1240–1242. doi: 10.1038/nn.2909
Mellios, N., and Sur, M. (2012). The emerging role of microRNAs in schizophrenia and autism spectrum disorders. Front. Psychiatry 3:39. doi: 10.3389/fpsyt.2012.00039
Mrsic-Flogel, T. D., Hofer, S. B., Ohki, K., Reid, R. C., Bonhoeffer, T., and Hübener, M. (2007). Homeostatic regulation of eye-specific responses in visual cortex during ocular dominance plasticity. Neuron 54, 961–972. doi: 10.1016/j.neuron.2007.05.028
Natera-Naranjo, O., Aschrafi, A., Gioio, A. E., and Kaplan, B. B. (2010). Identification and quantitative analyses of microRNAs located in the distal axons of sympathetic neurons. RNA 16, 1516–1529. doi: 10.1261/rna.1833310
Nelson, S. B., and Valakh, V. (2015). Excitatory/inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron 87, 684–698. doi: 10.1016/j.neuron.2015.07.033
Nudelman, A. S., DiRocco, D. P., Lambert, T. J., Garelick, M. G., Le, J., Nathanson, N. M., et al. (2010). Neuronal activity rapidly induces transcription of the CREB-regulated microRNA-132, in vivo. Hippocampus 20, 492–498. doi: 10.1002/hipo.20646
Oh, W. C., Parajuli, L. K., and Zito, K. (2015). Heterosynaptic structural plasticity on local dendritic segments of hippocampal CA1 neurons. Cell Rep. 10, 162–169. doi: 10.1016/j.celrep.2014.12.016
Olde Loohuis, N. F. M., Ba, W., Stoerchel, P. H., Kos, A., Jager, A., Schratt, G., et al. (2015). MicroRNA-137 controls AMPA-receptor-mediated transmission and mGluR-dependent LTD. Cell Rep. 11, 1876–1884. doi: 10.1016/j.celrep.2015.05.040
Packer, A. N., Xing, Y., Harper, S. Q., Jones, L., and Davidson, B. L. (2008). The Bifunctional microRNA miR-9/miR-9∗ regulates REST and CoREST and is downregulated in Huntington’s Disease. J. Neurosci. 28, 14341–14346. doi: 10.1523/JNEUROSCI.2390-08.2008
Park, H. Y., Lim, H., Yoon, Y. J., Follenzi, A., Nwokafor, C., Lopez-Jones, M., et al. (2014). Visualization of dynamics of single endogenous mRNA labeled in live mouse. Science 343, 422–424. doi: 10.1126/science.1239200
Park, I., Kim, H. J., Kim, Y., Hwang, H. S., Kasai, H., Kim, J.-H., et al. (2019). Nanoscale imaging reveals miRNA-mediated control of functional states of dendritic spines. Proc. Natl. Acad. Sci. U.S.A. 116, 9616–9621. doi: 10.1073/pnas.1819374116
Patrick, E., Rajagopal, S., Wong, H. K. A., McCabe, C., Xu, J., Tang, A., et al. (2017). Dissecting the role of non-coding RNAs in the accumulation of amyloid and tau neuropathologies in Alzheimer’s disease. Mol. Neurodegener. 12:51. doi: 10.1186/s13024-017-0191-y
Peng, J., Omran, A., Ashhab, M. U., Kong, H., Gan, N., He, F., et al. (2013). Expression patterns of miR-124, miR-134, miR-132, and miR-21 in an immature rat model and children with mesial temporal lobe epilepsy. J. Mol. Neurosci. 50, 291–297. doi: 10.1007/s12031-013-9953-3
Peng, Y.-R., Zeng, S.-Y., Song, H.-L., Li, M.-Y., Yamada, M. K., and Yu, X. (2010). Postsynaptic spiking homeostatically induces cell-autonomous regulation of inhibitory inputs via retrograde signaling. J. Neurosci. 30, 16220–16231. doi: 10.1523/JNEUROSCI.3085-10.2010
Penn, A. C., Zhang, C. L., Georges, F., Royer, L., Breillat, C., Hosy, E., et al. (2017). Hippocampal LTP and contextual learning require surface diffusion of AMPA receptors. Nature 549, 384–388. doi: 10.1038/nature23658
Petersen, S. A., Fetter, R. D., Noordermeer, J. N., Goodman, C. S., and DiAntonio, A. (1997). Genetic analysis of glutamate receptors in drosophila reveals a retrograde signal regulating presynaptic transmitter release. Neuron 19, 1237–1248. doi: 10.1016/S0896-6273(00)80415-8
Pitchiaya, S., Heinicke, L. A., Park, J. I., Cameron, E. L., and Walter, N. G. (2017). Resolving Subcellular miRNA Trafficking and turnover at single-molecule resolution. Cell Rep. 19, 630–642. doi: 10.1016/j.celrep.2017.03.075
Qiu, Z., Sylwestrak, E. L., Lieberman, D. N., Zhang, Y., Liu, X.-Y., and Ghosh, A. (2012). The rett syndrome protein MeCP2 regulates synaptic scaling. J. Neurosci. 32, 989–994. doi: 10.1523/JNEUROSCI.0175-11.2012
Quinlan, S., Kenny, A., Medina, M., Engel, T., and Jimenez-Mateos, E. M. (2017). MicroRNAs in neurodegenerative Diseases. Int. Rev. Cell Mol. Biol. 334, 309–343. doi: 10.1016/bs.ircmb.2017.04.002
Raab-Graham, K. F., Haddick, P. C. G., Jan, Y. N., and Jan, L. Y. (2006). Activity- and mTOR-dependent suppression of Kv1.1 channel mRNA translation in dendrites. Science 314, 144–148. doi: 10.1126/science.1131693
Rajman, M., Metge, F., Fiore, R., Khudayberdiev, S., Aksoy-Aksel, A., Bicker, S., et al. (2017). A microRNA-129-5p/Rbfox crosstalk coordinates homeostatic downscaling of excitatory synapses. EMBO J. 36, 1770–1787. doi: 10.15252/embj.201695748
Ramocki, M. B., and Zoghbi, H. Y. (2008). Failure of neuronal homeostasis results in common neuropsychiatric phenotypes. Nature 455, 912–918. doi: 10.1038/nature07457
Rannals, M. D., and Kapur, J. (2011). Homeostatic strengthening of inhibitory synapses is mediated by the accumulation of GABAA receptors. J. Neurosci. 31, 17701–17712. doi: 10.1523/JNEUROSCI.4476-11.2011
Ranson, A., Cheetham, C. E. J., Fox, K., and Sengpiel, F. (2012). Homeostatic plasticity mechanisms are required for juvenile, but not adult, ocular dominance plasticity. Proc. Natl. Acad. Sci. U.S.A. 109, 1311–1316. doi: 10.1073/pnas.1112204109
Redell, J. B., Liu, Y., and Dash, P. K. (2009). Traumatic brain injury alters expression of hippocampal microRNAs: potential regulators of multiple pathophysiological processes. J. Neurosci. Res. 87, 1435–1448. doi: 10.1002/jnr.21945
Remenyi, J., Hunter, C. J., Cole, C., Ando, H., Impey, S., Monk, C. E., et al. (2010). Regulation of the miR-212/132 locus by MSK1 and CREB in response to neurotrophins. Biochem. J. 428, 281–291. doi: 10.1042/BJ20100024
Remenyi, J., van den Bosch, M. W. M., Palygin, O., Mistry, R. B., McKenzie, C., Macdonald, A., et al. (2013). miR-132/212 knockout mice reveal roles for these miRNAs in regulating cortical synaptic transmission and plasticity. PLoS One 8:e62509. doi: 10.1371/journal.pone.0062509
Rocchi, A., Moretti, D., Lignani, G., Colombo, E., Scholz-Starke, J., Baldelli, P., et al. (2019). Neurite-enriched MicroRNA-218 stimulates translation of the GluA2 subunit and increases excitatory synaptic strength. Mol. Neurobiol. 56, 5701–5714. doi: 10.1007/s12035-019-1492-7
Sambandan, S., Akbalik, G., Kochen, L., Rinne, J., Kahlstatt, J., Glock, C., et al. (2017). Activity-dependent spatially localized miRNA maturation in neuronal dendrites. Science 355, 634–637. doi: 10.1126/science.aaf8995
Sarachana, T., Zhou, R., Chen, G., Manji, H. K., and Hu, V. W. (2010). Investigation of post-transcriptional gene regulatory networks associated with autism spectrum disorders by microRNA expression profiling of lymphoblastoid cell lines. Genome Med. 2:23. doi: 10.1186/gm144
Schanzenbächer, C. T., Langer, J. D., and Schuman, E. M. (2018). Time- and polarity-dependent proteomic changes associated with homeostatic scaling at central synapses. eLife 7:e33322. doi: 10.7554/eLife.33322
Schanzenbächer, C. T., Sambandan, S., Langer, J. D., and Schuman, E. M. (2016). Nascent proteome remodeling following homeostatic scaling at hippocampal synapses. Neuron 92, 358–371. doi: 10.1016/j.neuron.2016.09.058
Schratt, G. M., Tuebing, F., Nigh, E. A., Kane, C. G., Sabatini, M. E., Kiebler, M., et al. (2006). A brain-specific microRNA regulates dendritic spine development. Nature 439, 283–289. doi: 10.1038/nature04367
Seeburg, D. P., Feliu-Mojer, M., Gaiottino, J., Pak, D. T. S., and Sheng, M. (2008). Critical role of CDK5 and polo-like kinase 2 in homeostatic synaptic plasticity during elevated activity. Neuron 58, 571–583. doi: 10.1016/j.neuron.2008.03.021
Shepherd, J. D., Rumbaugh, G., Wu, J., Chowdhury, S., Plath, N., Kuhl, D., et al. (2006). Arc/Arg3.1 mediates homeostatic synaptic scaling of AMPA receptors. Neuron 52, 475–484. doi: 10.1016/j.neuron.2006.08.034
Siegel, G., Obernosterer, G., Fiore, R., Oehmen, M., Bicker, S., Christensen, M., et al. (2009). A functional screen implicates microRNA-138-dependent regulation of the depalmitoylation enzyme APT1 in dendritic spine morphogenesis. Nat. Cell Biol. 11, 705–716. doi: 10.1038/ncb1876
Siegert, S., Seo, J., Kwon, E. J., Rudenko, A., Cho, S., Wang, W., et al. (2015). The schizophrenia risk gene product miR-137 alters presynaptic plasticity. Nat. Neurosci. 18, 1008–1016. doi: 10.1038/nn.4023
Silva, M. M., Rodrigues, B., Fernandes, J., Santos, S. D., Carreto, L., Santos, M. A. S., et al. (2019). MicroRNA-186-5p controls GluA2 surface expression and synaptic scaling in hippocampal neurons. Proc. Natl. Acad. Sci. U.S.A. 116, 5727–5736. doi: 10.1073/pnas.1900338116
Sosanya, N. M., Brager, D. H., Wolfe, S., Niere, F., and Raab-Graham, K. F. (2015). Rapamycin reveals an mTOR-independent repression of Kv1.1 expression during epileptogenesis. Neurobiol. Dis. 73, 96–105. doi: 10.1016/j.nbd.2014.09.011
Sosanya, N. M., Huang, P. P. C., Cacheaux, L. P., Chen, C. J., Nguyen, K., Perrone-Bizzozero, N. I., et al. (2013). Degradation of high affinity HuD targets releases Kv1.1 mRNA from miR-129 repression by mTORC1. J. Cell Biol. 202, 53–69. doi: 10.1083/jcb.201212089
Soula, A., Valere, M., López-González, M.-J., Ury-Thiery, V., Groppi, A., Landry, M., et al. (2018). Small RNA-Seq reveals novel miRNAs shaping the transcriptomic identity of rat brain structures. Life Sci. Alliance 1:e201800018. doi: 10.26508/lsa.201800018
Stark, K. L., Xu, B., Bagchi, A., Lai, W. S., Liu, H., Hsu, R., et al. (2008). Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat. Genet. 40, 751–760. doi: 10.1038/ng.138
Stellwagen, D., and Malenka, R. C. (2006). Synaptic scaling mediated by glial TNF-alpha. Nature 440, 1054–1059. doi: 10.1038/nature04671
Störchel, P. H., Thümmler, J., Siegel, G., Aksoy-Aksel, A., Zampa, F., Sumer, S., et al. (2015). A large-scale functional screen identifies Nova1 and Ncoa3 as regulators of neuronal miRNA function. EMBO J. 34, 2237–2254. doi: 10.15252/embj.201490643
Sutton, M. A., Ito, H. T., Cressy, P., Kempf, C., Woo, J. C., and Schuman, E. M. (2006). Miniature neurotransmission stabilizes synaptic function via tonic suppression of local dendritic protein synthesis. Cell 125, 785–799. doi: 10.1016/j.cell.2006.03.040
Swann, J. W., and Rho, J. M. (2014). How is homeostatic plasticity important in epilepsy? Adv. Exp. Med. Biol. 813, 123–131. doi: 10.1007/978-94-017-8914-1-10
Szulwach, K. E., Li, X., Smrt, R. D., Li, Y., Luo, Y., Lin, L., et al. (2010). Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J. Cell Biol. 189, 127–141. doi: 10.1083/jcb.200908151
Talebizadeh, Z., Butler, M. G., and Theodoro, M. F. (2008). Feasibility and relevance of examining lymphoblastoid cell lines to study role of microRNAs in autism. Autism Res. 1, 240–250. doi: 10.1002/aur.33
Tan, C. L., Plotkin, J. L., Veno, M. T., von Schimmelmann, M., Feinberg, P., Mann, S., et al. (2013). MicroRNA-128 governs neuronal excitability and motor behavior in mice. Science 342, 1254–1258. doi: 10.1126/science.1244193
Thiagarajan, T. C., Lindskog, M., and Tsien, R. W. (2005). Adaptation to synaptic inactivity in hippocampal neurons. Neuron 47, 725–737. doi: 10.1016/j.neuron.2005.06.037
Tien, N.-W., and Kerschensteiner, D. (2018). Homeostatic plasticity in neural development. Neural Dev. 13:9. doi: 10.1186/s13064-018-0105-x
Tognini, P., Putignano, E., Coatti, A., and Pizzorusso, T. (2011). Experience-dependent expression of miR-132 regulates ocular dominance plasticity. Nat. Neurosci. 14, 1237–1239. doi: 10.1038/nn.2920
Torres-Berrío, A., Lopez, J. P., Bagot, R. C., Nouel, D., Dal Bo, G., Cuesta, S., et al. (2017). DCC confers susceptibility to depression-like behaviors in humans and mice and is regulated by miR-218. Biol. Psychiatry 81, 306–315. doi: 10.1016/j.biopsych.2016.08.017
Tsurudome, K., Tsang, K., Liao, E. H., Ball, R., Penney, J., Yang, J. S., et al. (2010). The drosophila miR-310 cluster negatively regulates synaptic strength at the neuromuscular junction. Neuron 68, 879–893. doi: 10.1016/j.neuron.2010.11.016
Turrigiano, G. (2012). Homeostatic synaptic plasticity: local and global mechanisms for stabilizing neuronal function. Cold Spring Harb. Perspect. Biol. 4:a005736. doi: 10.1101/cshperspect.a005736
Turrigiano, G. G. (2017). The dialectic of Hebb and homeostasis. Philos. Trans. R. Soc. B Biol. Sci. 372:20160258. doi: 10.1098/rstb.2016.0258
Turrigiano, G. G., Leslie, K. R., Desai, N. S., Rutherford, L. C., and Nelson, S. B. (1998). Activity-dependent scaling of quantal amplitude in neocortical neurons. Nature 391, 892–896. doi: 10.1038/36103
Urdinguio, R. G., Fernandez, A. F., Lopez-Nieva, P., Rossi, S., Huertas, D., Kulis, M., et al. (2010). Disrupted microrna expression caused by Mecp2 loss in a mouse model of Rett syndrome. Epigenetics 5, 656–663. doi: 10.4161/epi.5.7.13055
van Vliet, E. A., Aronica, E., Redeker, S., Boer, K., and Gorter, J. A. (2009). Decreased expression of synaptic vesicle protein 2A, the binding site for levetiracetam, during epileptogenesis and chronic epilepsy. Epilepsia 50, 422–433. doi: 10.1111/j.1528-1167.2008.01727.x
Verma, P., Augustine, G. J., Ammar, M. R., Tashiro, A., and Cohen, S. M. (2015). A neuroprotective role for microRNA miR-1000 mediated by limiting glutamate excitotoxicity. Nat. Neurosci. 18, 379–387. doi: 10.1038/nn.3935
Vitureira, N., and Goda, Y. (2013). Cell biology in neuroscience: the interplay between Hebbian and homeostatic synaptic plasticity. J. Cell Biol. 203, 175–186. doi: 10.1083/jcb.201306030
Vitureira, N., Letellier, M., and Goda, Y. (2012). Homeostatic synaptic plasticity: from single synapses to neural circuits. Curr. Opin. Neurobiol. 22, 516–521. doi: 10.1016/j.conb.2011.09.006
Vitureira, N., Letellier, M., White, I. J., and Goda, Y. (2011). Differential control of presynaptic efficacy by postsynaptic N-cadherin and β-catenin. Nat. Neurosci. 15, 81–89. doi: 10.1038/nn.2995
Wang, C., Han, B., Zhou, R., and Zhuang, X. (2016). Real-time imaging of translation on single mRNA transcripts in live cells. Cell 165, 990–1001. doi: 10.1016/j.cell.2016.04.040
Wang, G., Zhong, J., Guttieres, D., and Man, H. Y. (2019). Non-scaling regulation of AMPA receptors in homeostatic synaptic plasticity. Neuropharmacology 158:107700. doi: 10.1016/j.neuropharm.2019.107700
Wayman, G. A., Davare, M., Ando, H., Fortin, D., Varlamova, O., Cheng, H.-Y. M., et al. (2008). An activity-regulated microRNA controls dendritic plasticity by down-regulating p250GAP. Proc. Natl. Acad. Sci. U.S.A. 105, 9093–9098. doi: 10.1073/pnas.0803072105
Wen, M. M. (2016). Getting miRNA therapeutics into the target cells for neurodegenerative diseases: a mini-review. Front. Mol. Neurosci. 9:129. doi: 10.3389/fnmol.2016.00129
Wondolowski, J., and Dickman, D. (2013). Emerging links between homeostatic synaptic plasticity and neurological disease. Front. Cell. Neurosci. 7:223. doi: 10.3389/fncel.2013.00223
Wu, B., Eliscovich, C., Yoon, Y. J., and Singer, R. H. (2016). Translation dynamics of single mRNAs in live cells and neurons. Science 352, 1430–1435. doi: 10.1126/science.aaf1084
Xu, X. L., Zong, R., Li, Z., Biswas, M. H. U., Fang, Z., Nelson, D. L., et al. (2011). Fxr1p but not fmrp regulates the levels of mammalian brain-specific microrna-9 and microrna-124. J. Neurosci. 31, 13705–13709. doi: 10.1523/JNEUROSCI.2827-11.2011
Yan, X., Hoek, T. A., Vale, R. D., and Tanenbaum, M. E. (2016). Dynamics of translation of single mRNA molecules In Vivo. Cell 165, 976–989. doi: 10.1016/j.cell.2016.04.034
Yang, H., Wang, H., Shu, Y., and Li, X. (2018). miR-103 promotes neurite outgrowth and suppresses cells apoptosis by targeting prostaglandin-endoperoxide synthase 2 in cellular models of Alzheimer’s disease. Front. Cell. Neurosci. 12:91. doi: 10.3389/fncel.2018.00091
Yao, J., Nowack, A., Kensel-Hammes, P., Gardner, R. G., and Bajjalieh, S. M. (2010). Cotrafficking of SV2 and synaptotagmin at the synapse. J. Neurosci. 30, 5569–5578. doi: 10.1523/JNEUROSCI.4781-09.2010
Keywords: homeostatic plasticity, miRNA–microRNA, synaptic scaling, protein translation, membrane excitability, synaptic strength, synaptic plasticity
Citation: Dubes S, Favereaux A, Thoumine O and Letellier M (2019) miRNA-Dependent Control of Homeostatic Plasticity in Neurons. Front. Cell. Neurosci. 13:536. doi: 10.3389/fncel.2019.00536
Received: 05 September 2019; Accepted: 19 November 2019;
Published: 05 December 2019.
Edited by:
Annalisa Scimemi, University at Albany, United StatesReviewed by:
C. Andrew Frank, The University of Iowa, United StatesAna Luisa Carvalho, University of Coimbra, Portugal
Andreas Vlachos, University of Freiburg, Germany
Mariline Silva, University of Coimbra, Portugal, in collaboration with reviewer AC
Copyright © 2019 Dubes, Favereaux, Thoumine and Letellier. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mathieu Letellier, bWF0aGlldS5sZXRlbGxpZXJAdS1ib3JkZWF1eC5mcg==
 Sandra Dubes1,2
Sandra Dubes1,2 Alexandre Favereaux
Alexandre Favereaux Olivier Thoumine
Olivier Thoumine Mathieu Letellier
Mathieu Letellier