- Department of Biology, Centre for Advanced Research in Environmental Genomics, University of Ottawa, Ottawa, ON, Canada
Aromatase cytochrome P450arom (cyp19) is the only enzyme that has the ability to convert androgens into estrogens. Estrogens, which are produced locally in the vertebrate brain play many fundamental roles in neuroendocrine functions, reproductive functions, socio-sexual behaviors, and neurogenesis. Radial glial cells (RGCs) are neuronal progenitor cells that are abundant in fish brains and are the exclusive site of aromatase B expression and neuroestrogen synthesis. Using a novel in vitro RGC culture preparation we studied the regulation of aromatase B by 17β-estradiol (E2) and dopamine (DA). We have established that activation of the dopamine D1 receptor (D1R) by SKF 38393 up-regulates aromatase B gene expression most likely through the phosphorylation of cyclic AMP response element binding protein (CREB). This up-regulation can be enhanced by low concentration of E2 (100 nM) through increasing the expression of D1R and the level of p-CREB protein. However, a high concentration of E2 (1 μM) and D1R agonist together failed to up-regulate aromatase B, potentially due to attenuation of esr2b expression and p-CREB levels. Furthermore, we found the up-regulation of aromatase B by E2 and DA both requires the involvement of esr1 and esr2a. The combined effect of E2 and DA agonist indicates that aromatase B in the adult teleost brain is under tight control by both steroids and neurotransmitters to precisely regulate neuroestrogen levels.
Introduction
Aromatase cytochrome P450arom (cyp19) is the only enzyme performing the conversion of androgens into estrogens (Lephart, 1996; Garcia-Segura et al., 2003). In all species of vertebrates, including mammals, birds, and teleost fish, aromatase expression can be found in the brain in addition to testes and ovaries (Balthazart and Ball, 1998; Forlano et al., 2001). The distribution of aromatase varies between species and regions in the central nervous system (CNS), which has been detected at multiple levels, including mRNA, protein, and enzyme activity (Lephart, 1996; Balthazart and Ball, 1998; Forlano et al., 2006; Garcia-Segura, 2008; Azcoitia et al., 2011). In mammals, aromatase is expressed in both neuronal and glial cell bodies, processes and synaptic terminals of specific areas, especially the pial surface and ventricular surface in the brain under normal conditions (Martínez-Cerdeno et al., 2006; Yague et al., 2006). Aromatase may also be expressed in reactive astrocytes under conditions of cellular stress in mammalian brains (Azcoitia et al., 2003). In contrast to mammals, teleosts express two structurally and functionally different aromatase genes, aromatase A and aromatase B, which are a result of a gene duplication event (Diotel et al., 2010). The expression of aromatase B (cyp19a1b) in the teleost brain is exclusive to one particular cell type, the radial glial cell (RGC) (Forlano et al., 2001; Diotel et al., 2010; Xing et al., 2014), which allows direct in vivo study of glial aromatase contributions without interference from neuronal compartments. Callard et al. demonstrated for the first time that goldfish exhibit the highest brain aromatase activity compared to other vertebrates (Callard et al., 1981). More recently, Diotel et al. have documented the distribution cyp11a1, 3β-hsd, cyp17, and cyp19a1b mRNAs and the ability of de novo synthesis of several steroids including estrone and 17β-estradiol (E2) in the adult zebrafish brain (Diotel et al., 2011). Although it is very difficult to compare aromatase activity among species, it is clear that activity in the forebrain of teleost fish is much higher (100–1000 times) than in birds and mammals. All these data indicate that teleost fish are excellent models to study function and regulation of glial aromatase. Better understanding the regulation of glial aromatase will widen our knowledge of the control of estrogens in the brain. Local formation of estrogens are extremely important to maintain the homeostasis of cell proliferation and cell death, since estrogens in the brain are proven to be neuroprotective in mammalian brains and contributing to the generation of new neurons and brain repair after injury (Azcoitia et al., 2003; Arevalo et al., 2015). More importantly, recent studies indicate that RGCs are the progenitors of both neurons and glia during development, and are the source of new neurons in adult vertebrate brains (Alvarez-Buylla et al., 2001; Zupanc and Clint, 2003; Pellegrini et al., 2007). All these lines of evidence suggest that both RGC progenitor and steroidogenic functions contribute to neurogenesis in adult vertebrate brains.
Brain aromatization plays numerous important roles including brain differentiation, neural plasticity, neural regeneration, neuroendocrine functions, reproductive functions, and socio-sexual behaviors. It is possible that these processes are in part regulated by aromatization of androgen and thus local estrogens formation (Lephart, 1996; Garcia-Segura et al., 1999, 2003; Azcoitia et al., 2003; Ubuka et al., 2014). There are many response elements located at the promoter region of teleost cyp19a1b (Callard et al., 2001; Tchoudakova et al., 2001), including estrogen response element (ERE) and cyclic adenosine monophosphate (cAMP) response element (CRE). Through differential response element recruitment, cyp19a1b expression can be regulated differently, which suggests there should be multiple factors implicated in the regulation of cyp19a1b expression (Kato et al., 1997). It is well known that estrogens can up-regulate cyp19a1b expression in fish brain, which is mediated by estrogen receptors (ERs), some of which act as ligand-activated transcription factors (Belcher and Zsarnovszky, 2001). Upon binding to classical nuclear ERs, activated ER homo-dimerizes, and binds to ERE at the promoter of cyp19a1b, and then mediates transcription (Marlatt et al., 2008; Strobl-Mazzulla et al., 2008). Due to duplication of the estrogen receptor beta gene, teleost have three ERs (esr1/ERα, esr2a/ERβ, and esr2b/ERγ), all of which are expressed in the brain of fish with an expression pattern similar to that of aromatase (Forlano et al., 2005; Pellegrini et al., 2005; Strobl-Mazzulla et al., 2008; Fergus and Bass, 2013). There is evidence showing that moderate up-regulation of ERα by estrogens precedes the dramatic up-regulation of aromatase by its own products (Denslow et al., 2001; Bowman et al., 2002; McEwen, 2002; Marlatt et al., 2008). The possibility that cyp19a1b expression is up-regulated by estrogens is intriguing and suggests the presence of a positive feedback loop within the aromatase and ER-positive cells, whereby estrogens enhance aromatase expression and therefore the subsequent products, estrogens.
In addition to estrogenic regulation of the cyp19a1b gene, our previous neuroanatomical and pharmacological studies in goldfish reveal that there is a close relationship between catecholaminergic neurons and aromatase B- and DA D1 receptor (D1R)-positive RGCs in goldfish forebrain (Xing et al., 2015). Activation of D1R but not D2R up-regulates cyp19a1b mRNA in cultured RGCs (Xing et al., 2015). The signaling pathway underlying activated D1R regulation of cyp19a1b expression involves cAMP, protein kinase A (PKA) and phosphorylated cAMP response element binding protein (p-CREB). Whether this regulation involves nuclear ERs has yet to be characterized. Since it is likely that both mechanisms are acting simultaneously to affect multifactorial control over neuroestrogen synthesis, and given the close association between RGCs and DA neurons we directly tested for interactive effects on RGCs in vitro.
In this report, we show that both E2 and D1R agonist SKF 38393 alone significantly increased the expression of cyp19a1b. Moreover, the expression of cyp19a1b in RGCs may be synergistically up-regulated or antagonistically down-regulated, respectively, under low-concentration or high-concentration exposure to E2 and SKF 38393.
Materials and Methods
Ethics Statement
All procedures used were approved by the University of Ottawa Protocol Review Committee and followed standard Canadian Council on Animal Care guidelines on the use of animals in research.
Cell Culture
Common adult female goldfish (Carassius auratus) were purchased in April from a commercial supplier (Mt. Parnell Fisheries Inc., Mercersburg, PA, USA) and acclimated for at least 3 weeks prior to experimentation. The fish were maintained at 18°C under a natural simulated photoperiod and fed standard goldfish flakes. Sexually mature, pre-spawning female goldfish (20–35 g) were anesthetized using 3-aminobenzoic acid ethyl ester (MS222) for all handling, injection, and dissection procedures.
Cell culture methods have been established and validated previously by Xing et al. (2015). Briefly, female goldfish hypothalamus and telencephalon were dissected and rinsed with Hanks Balanced Salt Solution (HBSS; 400 mg KCl, 600 mg KH2PO4, 350 mg NaHCO3, 8 g NaCl, 48 mg Na2HPO4, and 1 g D-Glucose in 1 L ddH2O) with Antibiotic-Antimycotic solution (Gibco) and minced into small explants. Radial glial cells were dissociated with trypsin (0.25%, Gibco) and cultured in Leibovitz's L-15 medium with 15% Fetal Bovine Serum (FBS) and Antibiotic-Antimycotic solution. Cell culture medium was changed 4–7 days after isolation to remove tissue explants and suspended cells. Following the initial media change, cells received fresh media once a week until they were ready for their next passage. RGCs were sub-cultured following trypsinization (0.125%) for three passages and then used for experiments.
Drugs and Exposures
Cells were exposed to E2 (Sigma-Aldrich) and selective DA D1R agonist SKF 38393 (Tocris) for 24 h to study their effects on cyp19a1b, esr1, esr2a, esr2b, and drd1a mRNA and p-CREB protein level. To investigate ERs involvement in cyp19a1b mRNA regulation by E2 and SKF 38393, cells were pre-exposed to estrogen receptor antagonist MPP (Tocris) for 1 h, and then exposed to E2 or SKF 38393 for 24 h.
RNA Extraction, Quality Control and cDNA Synthesis
RNA was isolated using the RNeasy Micro kit (Qiagen) as described in the manufacturer's protocol. Upon purification, concentration and quality of all samples were assessed using the NanoDrop ND-1000 spectrophotometer (Thermo Scientific) and the 2100 Bioanalyzer (Agilent). RNA integrity values of samples were above the recommended minimum value of 5 for quantitative real-time RT PCR applications and ranged from 8.5 to 9.8 (Fleige and Pfaffl, 2006). Total cDNA was prepared using Maxima First Strand cDNA Synthesis Kit for RT-qPCR (Thermo Scientific). Each 20 μl reaction was diluted 10-fold in nuclease-free water and used as the template for the real-time RT-PCR assays.
Quantitative Real-Time RT-PCR
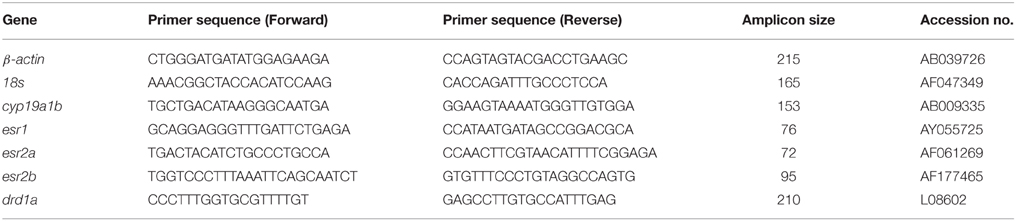
Quantitative real-time RT-PCR (qPCR) assays based on SYBR green detection were used to validate relative gene expression. Primers used in this study were designed using the Primer3 (http://primer3.sourceforge.net/) software and synthesized by Invitrogen (Table 1). The Maxima SYBR green qPCR Master Mix (Thermo Scientific) and CFX96 Real-Time PCR Detection System (Bio-Rad) were used to amplify and detect the transcripts of interest. Thermal cycling conditions included a denaturation step at 95°C for 3 min, and then 40 cycles of denaturation at 95°C for 10 s, annealing at 60°C for 30 s followed by melt curve analysis to confirm product specificity for all transcripts. Serial 1/2 dilution of the pool of all cDNA samples was analyzed to build the standard curves, from which the mRNA abundance in samples was calculated using the CFX Manager™ Software package (Bio-Rad). The efficiencies for all standard curves were between 92 and 106%. Data were normalized by using NORMA-GENE algorithm (Heckmann et al., 2011) and then presented as means + SEM of fold change to control group (n = 4; assayed in duplicate).
Droplet Digital PCR
The low expression of D1R (drd1a) in RGC cultures is not quantifiable using qPCR, so we used a more sensitive method to measure drd1a mRNA level. Droplet digital PCR (ddPCR) was performed using 25 ng of each cDNA sample and QX200™ ddPCR™ EvaGreen Supermix by employing QX200™ Droplet Digital™ PCR System (Bio-Rad). The ddPCR conditions were as follows: an initial denaturation for 5 min at 95°C followed by 40 cycles of denaturation for 30 s at 95°C and annealing and extension for 1 min at 56°C, then signal stabilization for 5 min at 4°C and 5 min at 90°C. Template DNA was omitted from the ddPCR reaction as a no template control (NTC) and the results of ddPCR were analyzed using the QuantaSoft Software (Bio-Rad). The absolute copy number of targeted gene was divided by the average of the absolute copy number of two reference genes β-actin and 18s, and presented as means + SEM of fold change to control group (n = 4; assayed in duplicate).
Western Blot
Total protein extract from RGCs was denatured and separated by electrophoresis on a 10% SDS–polyacrylamide gel and transferred to a PVDF membrane as described previously by Zhao et al. (2006). Membranes were incubated with anti-Phospho-CREB (Ser133, Cell signaling technology, 9198S) antibodies overnight at 4°C and then incubated with donkey anti-rabbit IgG antibody (GE Health Care, NA934VS). Blots were visualized using the Amersham ECL Prime Western Blotting Detection System (GE Health Care, RPN2232). Images were obtained using a ChemiDoc XRS+ system (Bio-Rad) and analyzed with image lab (Bio-Rad). Same blot incubated with anti-Phospho-CREB antibodies before was stripped and re-probed with primary anti-actin (Cedarlane, CLT9001) antibodies overnight at 4°C and then incubated with goat anti-mouse IgG-HRP (Santa Cruz, sc-2005), where actin served as an internal control.
Statistics
In all cases and before any analysis, normality and homogeneity of variances was verified using Shapiro-Wilk's and Levene's test, respectively. Comparison of two groups was performed using Student's t-test in GraphPad Prism (Version 6.0). Two-way analysis of variance (ANOVA) followed by Tukey's post-hoc test in GraphPad Prism (Version 6.0) was used to determine the effects of single or combined drug treatments. P = 0.05 were considered statistically significant. Data were presented as mean + SEM. Note that the t- and F-statistics, degrees of freedom (df) of interaction (DFn), the total degrees of freedom (DFd), and P-values are reported in the Results Section. The P-values associated with main effects and the interactions are reported directly on the graphs.
Results
Regulation of cyp19a1b and ER Expression by SKF 38393 and E2
Here, the qPCR data show that 10 μM SKF 38393 was able to increase the expression of cyp19a1b (t = 2.454, df = 6, P = 0.0225), but not esr1 (t = 1.744, df = 6, P = 0.0952), esr2a (t = 1.338, df = 6, P = 0.1947), and esr2b (t = 1.638, df = 22, P = 0.1157) after 24 h incubation (Figure 1A). The 100 nM concentration of E2 did not affect cyp19a1b (t = 1.557, df = 6, P = 0.1705), esr1 (t = 0.5747, df = 6, P = 0.5864), esr2a (t = 0.5106, df = 6, P = 0.6993), and esr2b (t = 0.9012, df = 6, P = 0.4022) mRNAs after 24 h incubation (Figure 1B). However, when the concentration of E2 was increased to 1 μM, the expression of cyp19a1b (t = 4.791, df = 6, P = 0.003) and esr2b (t = 10.65, df = 6, P < 0.0001) were increased dramatically 6.7-fold and 2.9-fold, respectively, but not esr1 (t = 0.3295, df = 6, P = 0.7530) and esr2a (t = 0.2395, df = 6, P = 0.779; Figure 1C). These data suggest the up-regulation of esr2b may contribute to the stimulatory effect of E2 in cyp19a1b expression.
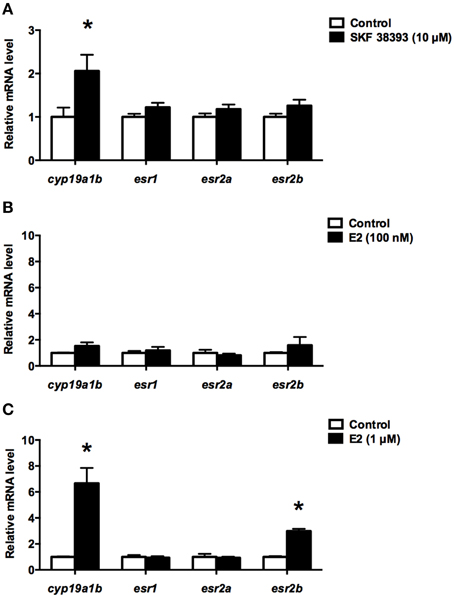
Figure 1. Effects of SKF 38393 and E2 on the expression of cyp19a1b, esr1, esr2a, and esr2b in primary RGC culture. Quantitative real-time PCR analysis showing variations in the relative amounts of the cyp19a1b, esr1, esr2a, and esr2b mRNAs in primary RGCs culture exposed to 10 μM SKF38393 (A) 100 nM E2 (B) and 1 μM E2 (C). Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM. Treatment groups marked by asterisks have significantly different mRNA levels compared to control (P < 0.05).
The Involvement of ERs in cyp19a1b Regulation by E2 and SKF 38393
When RGC cultures are exposed to SKF38393 and E2 for 24 h there is no change in the expression of esr1 and esr2a. This result led us to investigate the possible contribution of pre-existing ERs in RGCs to cyp19a1b up-regulation. The ERα and ERβ antagonist MPP was administered to RGC culture 1 h before SKF 38393 and E2. Data revealed that MPP alone decreases the expression of all three ERs (t = 2.462, df = 6, P = 0.049; t = 2.285, df = 6, P = 0.0624; t = 3.134, df = 6, P = 0.0202, respectively) by 29.9–57.6% (Figure 2A), but has no effect on cyp19a1b (F = 32.65, DFn = 1, DFd = 12, P = 0.8582; F = 18.15, DFn = 1, DFd = 12, P = 0.3065, respectively) expression (Figures 2B,C). Two-way ANOVAs were conducted that examined the effect of MPP and SKF 38393 as well as MPP and E2 on cyp19a1b expression. There was a statistically significant interaction between MPP and SKF 38393 (F = 32.65, DFn = 1, DFd = 12, P < 0.0001), and between MPP and E2 (F = 18.15, DFn = 1, DFd = 12, P = 0.0011). The stimulatory effect of SKF 38393 (F = 32.65, DFn = 1, DFd = 12, P = 0.0002) and E2 (F = 18.15, DFn = 1, DFd = 12, P < 0.0001) on cyp19a1b mRNA was completely blocked by MPP (Figures 2B,C).
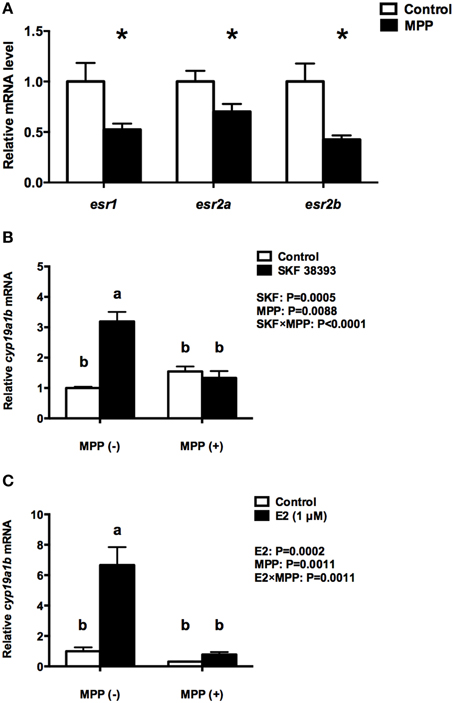
Figure 2. Effects of MPP on the expression of esr1, esr2a, and esr2b and the regulation of cyp19a1b expression by DA and E2 in primary RGC culture. Quantitative real-time PCR analysis showing variations in the relative amounts of the cyp19a1b, esr1, esr2a, and esr2b mRNAs in primary RGCs culture exposed to 100 nM MPP (A), 100 nM MPP and 10 μM SKF 38393 (B), 100 nM MPP and 1 μM E2 (C). Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM. Treatment groups marked by an asterisk (A) are significantly different from controls (P < 0.05). Treatment groups marked with different letters (B,C) are significantly different (P < 0.05).
The Combined Effects of E2 and SKF 38393 on cyp19a1b Regulation
Based on the previous dose-dependent studies (Xing et al., 2015 and unpublished data), we examined the combined effects of two concentrations of SKF 38393 and E2. Our results indicate that there is either a synergistic or antagonistic effect in cyp19a1b regulation by E2 and SKF 38393, which is dependent on the concentration of E2. Two-way ANOVAs were conducted that examined the effect of SKF 38393 and E2 on cyp19a1b expression. As shown in Figure 3A, the low concentration of E2 (100 nM) alone increased cyp19a1b expression 1.5 fold, but this was not statistically significant (F = 0.1071, DFn = 1, DFd = 12, P = 0.1498) regardless of the combination with SKF 38393 (1 μM). In contrast, the high concentration of E2 (1 μM) alone increased cyp19a1b 6.7-fold (F = 0.2028, DFn = 1, DFd = 12, P = 0.0002, Figure 3B) significantly regardless of the combination with SKF 38393 (1 μM). There was no statistically significant interaction between SKF 38393 (1 μM) and either the low (100 nM; F = 0.1071, DFn = 1, DFd = 12, P = 0.7491, Figure 3A) or higher (1 μM; F = 0.2028, DFn = 1, DFd = 12, P = 0.6601, Figure 3B) concentration of E2. When the concentration of SKF 38393 was increased to 10 μM, statistically significant interactions between SKF 38393 and both low concentration (F = 8.112, DFn = 1, DFd = 12, P = 0.0147, Figure 3C) and high concentration of E2 (F = 14.91, DFn = 1, DFd = 12, P = 0.0023, Figure 3D) were evident. The Two-way ANOVA indicates that both 10 μM SKF 38393 and 100 nM E2 increased cyp19a1b expression 2.8-fold and 1.5-fold, consistent with the earlier experiment shown in Figures 1A,B, respectively. Post-hoc tests showed that the combination of high concentration of SKF 38393 and low concentration of E2 synergistically up-regulated cyp19a1b (F = 8.112, DFn = 1, DFd = 12, P = 0.0003, Figure 3C) expression 8.0-fold. Contrasting this is the experiment where high concentration of SKF 38393 and high concentration of E2 were used. Exposure of RGCs to 1 μM E2 alone increased cyp19a1b expression 6.7-fold, and the high concentration of SKF 38393 inhibited this effect (F = 14.91, DFn = 1, DFd = 12, P = 0.0141, Figure 3D).
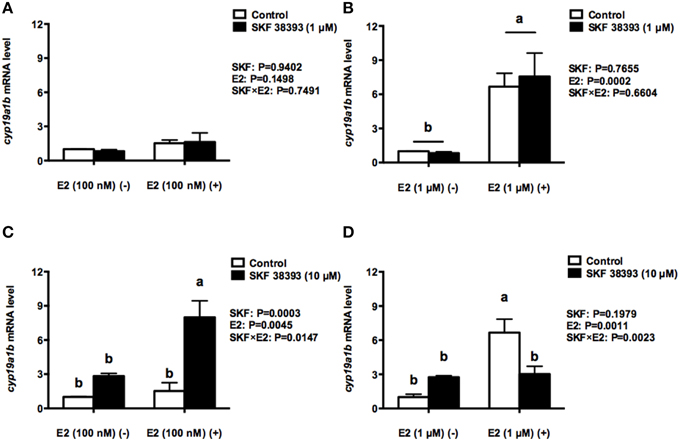
Figure 3. Quantitative real-time PCR analysis showing variations in the relative amounts of the cyp19a1b mRNA in primary RGCs culture exposed to 100 nM E2 and 1 μM SKF 38393 (A) 1 μM E2 and 1 μM SKF 38393 (B) 100 nM E2 and 10 μM SKF 38393 (C) 1 μM E2 and 10 μM SKF 38393 (D). Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM. Treatment groups marked by different letters are significantly different (P < 0.05).
Synergistic Effects of Low Levels of E2 on SKF 38393-Induced cyp19a1b Expression are Associated with Parallel Increases in p-CREB
To further understand the underlying mechanisms of synergistic up-regulation of cyp19a1b by low concentration of E2 and high concentration of SKF 38393, we studied two components of the D1R/cAMP/PKA/p-CREB pathway, drd1a in mRNA level and p-CREB in protein level as well as the expression of esr1, esr2a, and esr2b. Two-way ANOVAs were conducted that examined the effect of SKF 38393 and E2 on drd1a, esr1 esr2a, and esr2b mRNA level and p-CREB protein level in cultured RGCs. There was a statistically significant interaction (F = 7.393, DFn = 1, DFd = 12, P = 0.01786) between SKF 38393 and E2 for drd1a mRNA. Post-hoc tests showed that the combination of SKF 38393 and E2 up-regulated drd1a mRNA (F = 7.393, DFn = 1, DFd = 12, P = 0.0029; Figure 4A), although increases were not very large, in the range of 1.7 fold. There was also a statistically significant interaction (F = 10.35, DFn = 1, DFd = 12, P = 0.0123) between SKF 38393 and E2 for p-CREB protein level (Figures 4B–D). The D1R agonist SKF 38393 alone increased p-CREB 2.2-fold, and this was enhanced to 3.5-fold in the presence of E2, paralleling these results for cyp19a1b mRNA (Figure 3C). No effects on the expression of esr1, esr2a and esr2b (F = 0.2907, DFn = 1, DFd = 12, P = 0.5996; F = 0.0061, DFn = 1, DFd = 12, P = 0.9389; F = 0.0078, DFn = 1, DFd = 12, P = 0.9312, respectively) were observed for the E2 and SKF 38393 treatments (Figure 5).
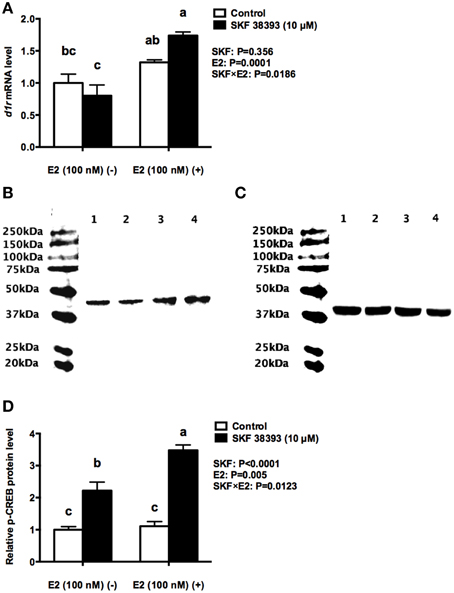
Figure 4. Synergistic up-regulation of D1R and p-CREB by E2 and SKF 38393. Droplet digital PCR analysis showing variations in the absolute amounts of the drd1a mRNA in primary RGCs culture exposed to 100 nM E2 and 10 μM SKF38393 (A). Western blot showing variations in the relative protein level of p-CREB in primary RGCs culture exposed to 100 nM E2 and 10 μM SKF 38393. Western blot image (B) showing the p-CREB immunoreactivity in control (lane 1), 100 nM E2 treated (lane 2), 10 μM SKF 38393 treated (lane3), and both 100 nM and 10 μM SKF 38393 treated (lane 4) groups, where actin served as internal control (C). The densitometric analysis of western blot is reported as an arbitrary value relative to the average of all bands on the same blot. Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM (n = 4, D). Treatment groups marked by different letters are significantly different (P < 0.05).
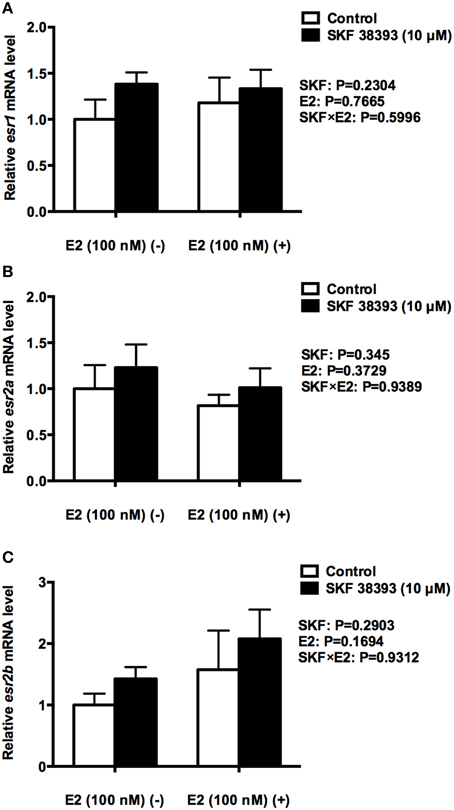
Figure 5. Quantitative real-time PCR analysis showing variations in the relative amounts of the esr1 (A), esr2a (B) and esr2b mRNA (C) in primary RGC cultures exposed to 100 nM E2 and 10 μM SKF 38393. Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM. There were no statistically signficant treatment effects.
Antagonistic Effects of High Levels of E2 on SKF 38393-Induced cyp19a1b Expression are Associated with Parallel Decreases in p-CREB Protein and esr2b mRNA
Further analysis was essential in understanding the potential mechanisms of antagonistic regulation of cyp19a1b by the combined high concentration of E2 and high concentration of SKF 38393. As with the previous experiment, two-way ANOVAs were conducted that examined the potential interactive effects of SKF 38393 and E2 on drd1a, esr1 esr2a, and esr2b mRNA level and p-CREB protein level. Exposure to 10 μM SKF 38393 and 1 μM E2 alone did not affect drd1a (F = 0.9571, DFn = 1, DFd = 12, P = 0.3472) mRNA levels in cultured RGCs (Figure 6A). In contrast, there was a statistically significant interaction between SKF 38393 and E2 on p-CREB protein (F = 6.252, DFn = 1, DFd = 12, P = 0.0369) and esr2b mRNA (F = 86.01, DFn = 1, DFd = 12, P < 0.0001) levels. Post-hoc tests showed that higher concentration of E2 inhibited the up-regulation of p-CREB protein induced by SKF 38393 (F = 6.252, DFn = 1, DFd = 12, P = 0.0484; Figures 6B–D) and that SKF 38393 inhibited the up-regulation of esr2b mRNA induced by higher concentration of E2 (F = 86.01, DFn = 1, DFd = 12, P < 0.0001; Figure 7C) in RGCs, both following a similar pattern to that of cyp19a1b expression (Figure 3D). No effects on esr1 and esr2a were evident (Figures 7A,B).
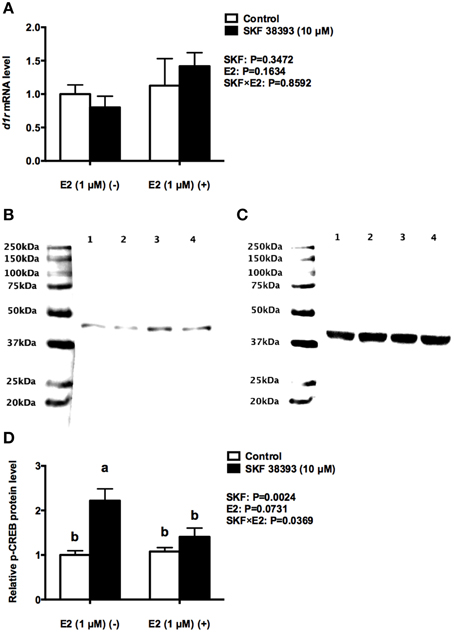
Figure 6. Antagonistic regulation of D1R and p-CREB by E2 and SKF 38393. Droplet digital PCR analysis showing variations in the absolute amounts of the d1r mRNA in primary RGCs culture exposed to 1 μM E2 and 10 μM SKF 38393 (A). Western blot image (B) showing the p-CREB immunoreactivity in control (lane 1), 1 μM E2 treated (lane 2), 10 μM SKF 38393 treated (lane3), and both 1 μM and 10 μM SKF 38393 treated (lane 4) groups, where actin served as internal control (C). The densitometric analysis of western blot is reported as an arbitrary value relative to the average of all bands on the same blot. Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM (n = 4, D). Treatment groups marked by different letters are significantly different (P < 0.05).
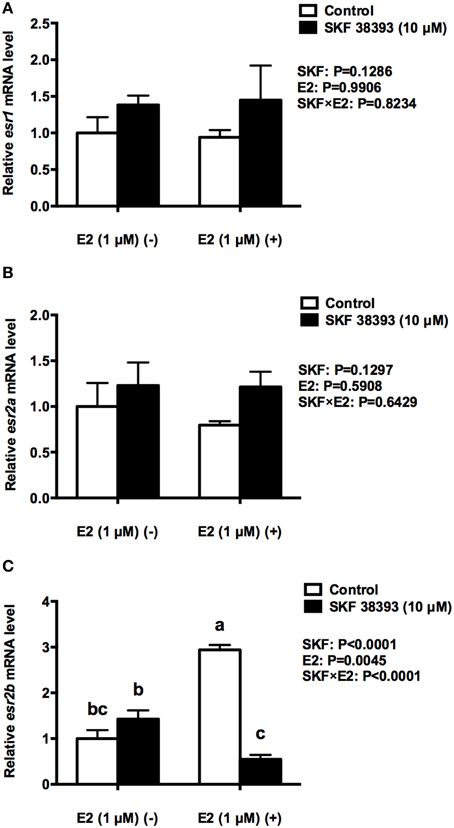
Figure 7. Quantitative real-time PCR analysis showing variations in the relative amounts of the esr1 (A), esr2a (B) and esr2b mRNA (C) in primary RGCs culture exposed to 1 μM E2 and 10 μM SKF 38393. Data were normalized and defined as fold change relative to control, and bars represent the mean + SEM. Treatment groups marked by different letters are significantly different (P < 0.05).
Discussion
The findings of this study indicate that cyp19a1b expression is up-regulated in goldfish RGC cultures in response to individual treatments with E2 or D1R agonist. However, co-exposure to these substances leads to either a further up-regulation or a down-regulation of cyp19a1b mRNA levels that are dependent on the concentration of E2. This is the first evidence for direct multifactorial control of aromatase in RGCs from the adult teleost brain. In mammals, aromatase is expressed in both neurons and glia (Tsuruo et al., 1995; Azcoitia et al., 2003). In contrast, RGCs are the exclusive cells to express aromatase in teleost brains (Forlano et al., 2001; Tong et al., 2009). This makes teleosts amenable models to delineate direct control of glial aromatase expression. Our data provides the foundation for further analysis of neurotransmitter and estrogenic regulation of RGCs, and contributes to our understanding of neuronal-glial interactions. This is important because estrogens in the CNS control fundamental processes such as sexual behavior and neurogenesis (Okada et al., 2008; Ubuka and Tsutsui, 2014; Pellegrini et al., 2015).
The ability of E2 to regulate brain aromatase mRNA has been reported for a variety of species, including fish, birds, and rats (Hutchison and Steimer, 1986; Pellegrini et al., 2005; Zhao et al., 2007; Strobl-Mazzulla et al., 2008). Here, we confirm and extend these observations because we show a direct effects of E2 on isolated goldfish RGCs. Our pharmacological experiments using goldfish RGCs in culture indicate the involvement of ERs since the up-regulation of cyp19a1b caused by exogenous E2 was completely blocked by the ER antagonist MPP (Pinto et al., 2014), consistent with previous findings in zebrafish in vivo (Diotel et al., 2010). Here, we report that E2 in vitro can increase esr2b mRNA only at a relatively high concentration. Contrary to in vivo reports in teleost, we did not find effects of exogenous E2 on esr1 and esr2a expression in RGC cultures (Menuet et al., 2005; Marlatt et al., 2008). To further understand the involvement of ERs, we administrated MPP together with SKF 38393 and E2. Exposure to MPP decreased the expression of all three nuclear ERs, suggesting that endogenous estrogen may be involved. Moreover, MPP blocked the stimulatory effect of added E2 on cyp19a1b expression. This finding indicates that the up-regulation of cyp19a1b mRNA by E2 is potentially due to the recruitment of the existing esr1 and esr2a and/or increasing expression of esr2b to contribute to the positive feedback loop between aromatase, estrogens, and ERs (Diotel et al., 2010). Further studies on the involvement of esr2b are currently limited by the lack of a selective esr2b antagonist, although we do hypothesize that esr2b may be contributing to the up-regulation of cyp19a1b in RGCs, given the potency of esr2b to drive reporter gene expression for a consensus ERE construct or for the zebrafish cyp19a1b promoter construct in transfected cell assays (Menuet et al., 2002). It is unlikely that the G-protein coupled membrane estrogen receptor (GPER) is involved because we have been unable to identify GPER in RGCs using targeted PCR and RNA sequencing (data not shown).
We confirm here the results from our recent study (Xing et al., 2015) of the direct stimulatory effect of D1R activation of the p-CREB signaling pathway to increase the expression of cyp19a1b in adult female goldfish RGCs. Interestingly, in the current study, pretreatment with the estrogen antagonist MPP completely blocks SKF 38393-stimulated cyp19a1b expression. This suggests that DA does not increase the expression of ERs but somehow activates or recruits the pre-existing ERs that contribute to the increase in expression of cyp19a1b.
While it is clear from our data that treatments with E2 or SKF 38393 alone can increase expression of cyp19a1b, this is unlikely to represent situations in vivo where RGCs would be responding to neurotransmitters and endogenous estrogens simultaneously. Therefore, we treated RGCs with combinations of low and high concentrations of the D1R agonists SKF 38393 and low and high concentrations of E2. It is evident that the RGC cyp19a1b response to D1R activation is dependent on the level of E2. The maximal response in cyp19a1b expression (i.e., 8.0-fold) was obtained from RGCs exposed to the lower E2 level of 100 nM plus the concentration of 10 μM SKF 38393, which we showed previously to be the highest effective concentration in the culture system (Xing et al., 2015). These experiments revealed a synergistic interaction between E2 and SKF 38393 that was supported statistically. Additionally, the levels of cellular p-CREB most closely followed the pattern of cyp19a1b expression, however, a more complex effect on D1R and ER expression was evident. The low level E2 treatment appears to amplify signaling via D1R/cAMP/PKA/p-CREB. Our absolute quantification of drd1a mRNA abundance showed that the low concentration of E2 increased D1R gene expression, especially in the presence of the D1R agonist SKF 38393. With the increased expression level of D1R, we hypothesized the activity of the downstream signal transduction pathway will be higher than the control level. Our data showed that the protein level of p-CREB also increased upon exposure to E2 and SKF 38393, similar to the drd1a expression pattern. It is well-known that nuclear ERs bind to EREs thereby enhancing cyp19a1b transcription (Callard et al., 2001; Menuet et al., 2002, 2005) as transcription factors themselves or potentially through interactions with other transcription factors, such as activator protein 1 (Marino et al., 2006; Zhang and Trudeau, 2006). While our experiments with the ER antagonist MPP indicate that endogenous pre-existing ERs are involved, our findings do not support that an increase in ERs is required for the synergistic up-regulation of cyp19a1b by E2 and SKF 38393 in RGCs since no increase in expression was observed for any of the three nuclear ERs. Together these data indicate that the up-regulation of drd1a expression and p-CREB protein may contribute to the overall synergistic effects of the low concentration of E2 and SKF 38393 on cyp19a1b expression.
While we do not have an estimate of the intracellular or extracellular levels of estrogens at or near regions of high RGC density in the brain, we consider the high 1 μM concentration of E2 to be supraphysiological and the low 100 nM concentration to be physiological relative to circulating E2 (Kobayashi et al., 1988) based on previous in vivo responses to exogenous estrogens in goldfish (Martyniuk et al., 2006). Nevertheless, we originally hypothesized that the higher combined concentrations of E2 and SKF 38393 would cause up-regulation of cyp19a1b expression. However, co-treatment with 10 μM SKF 38393 significantly reduced the high cyp19a1b expression induced by 1 μM E2. This was evident at the level of intracellular signaling because p-CREB protein levels were also reduced by the co-treatment of high concentration of E2 and high concentration of SKF 38393. The effects of E2 on CREB are variable since both inhibition and stimulation have been reported, perhaps reflecting tissue-specific regulation (Duan et al., 1999; Choi et al., 2004). Our findings did not show any change in p-CREB protein levels after exposure to E2 alone regardless of the concentration. Moreover, 10 μM SKF 38393 reduced the increase in esr2b expression caused by the high 1 μM concentration of E2 in our RGCs. Even though SKF 38393 did not cause any down-regulation after 24 h exposure, time-course studies have shown that esr2b decreased after 8 h exposure to SKF 38393 alone in RGC culture (unpublished data). Although speculative, together these data indicate that the attenuation of p-CREB protein and esr2b expression may contribute to the overall antagonistic effects of the high concentrations of E2 and SKF 38393 on cyp19a1b expression.
In summary, we have described an example of complex control of aromatase B expression in adult teleost RGCs. Anatomical and physiological data (Xing et al., 2015) indicate a close relationship between catecholaminergic cell bodies and fibers and D1R-expressing RGCs in female goldfish brain. Pharmacological activation of D1R up-regulates cyp19a1b expression in RGCs, which ultimately would increase estrogen synthesis. It has been previously shown and confirmed here, that teleost RGCs respond positively to E2 with an increase in cyp19a1b expression. We extend these observations to demonstrate a potentiating or synergistic interaction between dopaminergic and estrogenic stimulation of cyp19a1b expression. On the other hand, it appears that overstimulation of RGCs with the combined high concentration of E2 and high concentration of SKF 38393 may limit cyp19a1b expression, and thus reduce estrogen synthesis in a manner similar to classical negative feedback control (Figure 8). In the teleost brain, the two main roles of estrogens in the adult brain that have been documented thus far are regulation of expression of neuroendocrine-related genes (Martyniuk et al., 2006), and the inhibition of neurogenesis (Diotel et al., 2013; Makantasi and Dermon, 2014). Interactive regulation of aromatase B by dopamine and estrogens therefore, would finely regulate local estrogen levels that in turn could regulate adjacent RGCs that express nuclear ERs, or neurons in the vicinity that express these receptors or GPER. However, further studies on the conversion of androgen precursors by aromatase B are needed to further understand the control of estrogen synthesis in the brain. Given the array of neurotransmitter and neuropeptide-expressing neurons and fibers in the neurogenic regions where cyp19a1b expressing RGCs are found, it is likely that many other neurohormones regulate the steroidogenic function of this critical cell. The culture system we have developed provides an amenable approach to the understanding of multifactorial control of RGCs.
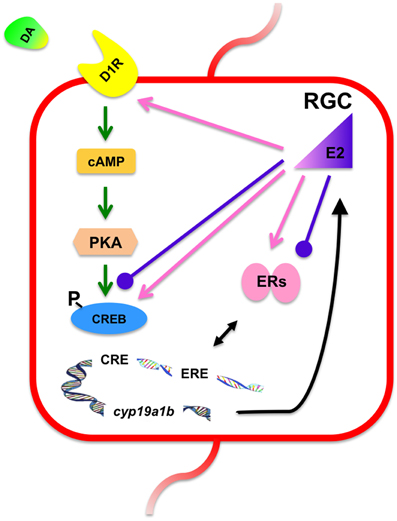
Figure 8. Proposed mechanisms underlying the synergistic and antagonistic regulation of cyp19a1b expression in cultured RGCs. Activation of the D1R by SKF 38393 (10 μM) up-regulates cyp19a1b expression, most likely through cAMP, PKA, and the phosphorylation of CREB. This up-regulation can be enhanced by low concentration of E2 (100 nM) through increasing the expression of D1R and the level of p-CREB protein (pink arrows). Also, up-regulation of cyp19a1b expression by E2 and DA both requires the involvement of esr1 and esr2a. However, a high concentration of E2 (1 μM) and SKF38393 (10 μM) together failed to up-regulate cyp19a1b expression, potentially due to attenuation of esr2b expression and p-CREB levels (purple bulbous arrows). RGC, radial glial cells; DA, dopamine; D1R, dopamine D1 receptor; cAMP, cyclic adenosine monophosphate; PKA, protein kinase A; CREB, cAMP response element binding protein; p-CREB, phosphorylated cAMP response element binding protein; CRE, cAMP response element; E2, 17β-estradiol; ERs, estrogen receptors; ERE, estrogen response element.
Author Contributions
LX, designed the study, developed the methodology, conducted the majority of experiment, and analysis and wrote the manuscript. CE, maintained cell culture and helped with qPCR analysis. VT, helped with the design of the study and editing the article.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Funding from The Ontario Trillium Scholarship (LX), and NSERC Discovery Grants and University of Ottawa Research Chair in Neuroendocrinology (VT) are acknowledged with appreciation. The authors of the manuscript have no conflicts of interest to declare.
References
Alvarez-Buylla, A., Garcia-Verdugo, J. M., and Tramontin, A. D. (2001). A unified hypothesis on the lineage of neural stem cells. Nat. Rev. Neurosci. 2, 287–293. doi: 10.1038/35067582
Arevalo, M. A., Azcoitia, I., and Garcia-Segura, L. M. (2015). The neuroprotective actions of oestradiol and oestrogen receptors. Nat. Rev. Neurosci. 16, 17–29. doi: 10.1038/nrn3856
Azcoitia, I., Sierra, A., Veiga, S., and Garcia-Segura, L. M. (2003). Aromatase expression by reactive astroglia is neuroprotective. Ann. N.Y. Acad. Sci. 1007, 298–305. doi: 10.1196/annals.1286.028
Azcoitia, I., Yague, J. G., and Garcia-Segura, L. M. (2011). Estradiol synthesis within the human brain. Neuroscience 191, 139–147. doi: 10.1016/j.neuroscience.2011.02.012
Balthazart, J., and Ball, G. F. (1998). New insights into the regulation and function of brain estrogen synthase (aromatase). Trends Neurosci. 21, 243–249. doi: 10.1016/S0166-2236(97)01221-6
Belcher, S. M., and Zsarnovszky, A. (2001). Estrogenic actions in the brain: estrogen, phytoestrogens, and rapid intracellular signaling mechanisms. J. Pharmacol. Exp. Ther. 299, 408–414.
Bowman, C. J., Kroll, K. J., Gross, T. G., and Denslow, N. D. (2002). Estradiol-induced gene expression in largemouth bass (Micropterus salmoides). Mol. Cell. Endocrinol. 196, 67–77. doi: 10.1016/S0303-7207(02)00224-1
Callard, G. V., Petro, Z., and Ryan, K. J. (1981). Estrogen synthesis in vitro and in vivo in the brain of a marine teleost (Myoxocephalus). Gen. Comp. Endocrinol. 43, 243–255. doi: 10.1016/0016-6480(81)90318-X
Callard, G. V., Tchoudakova, A. V., Kishida, M., and Wood, E. (2001). Differential tissue distribution, developmental programming, estrogen regulation and promoter characteristics of cyp19 genes in teleost fish. J. Steroid Biochem. Mol. Biol. 79, 305–314. doi: 10.1016/S0960-0760(01)00147-9
Choi, Y. C., Lee, J. H., Hong, K. W., and Lee, K. S. (2004). 17 Beta-estradiol prevents focal cerebral ischemic damages via activation of Akt and CREB in association with reduced PTEN phosphorylation in rats. Fundam. Clin. Pharmacol. 18, 547–557. doi: 10.1111/j.1472-8206.2004.00284.x
Denslow, N. D., Lee, H. S., Bowman, C. J., Hemmer, M. J., and Folmar, L. C. (2001). Multiple responses in gene expression in fish treated with estrogen. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 129, 277–282. doi: 10.1016/S1096-4959(01)00322-0
Diotel, N., Do Rego, J. L., Anglade, I., Vaillant, C., Pellegrini, E., Gueguen, M.-M., et al. (2011). Activity and expression of steroidogenic enzymes in the brain of adult zebrafish. Eur. J. Neurosci. 34, 45–56. doi: 10.1111/j.1460-9568.2011.07731.x
Diotel, N., Le Page, Y., Mouriec, K., Tong, S.-K., Pellegrini, E., Vaillant, C., et al. (2010). Aromatase in the brain of teleost fish: expression, regulation and putative functions. Front. Neuroendocrinol. 31:3. doi: 10.1016/j.yfrne.2010.01.003
Diotel, N., Vaillant, C., Gabbero, C., Mironov, S., Fostier, A., Gueguen, M.-M., et al. (2013). Effects of estradiol in adult neurogenesis and brain repair in zebrafish. Horm. Behav. 63, 193–207. doi: 10.1016/j.yhbeh.2012.04.003
Duan, W. R., Shin, J. L., and Jameson, J. L. (1999). Estradiol suppresses phosphorylation of cyclic adenosine 3',5'-monophosphate response element binding protein (CREB) in the pituitary: evidence for indirect action via gonadotropin-releasing hormone. Mol. Endocrinol. 13, 1338–1352.
Fergus, D. J., and Bass, A. H. (2013). Localization and divergent profiles of estrogen receptors and aromatase in the vocal and auditory networks of a fish with alternative mating tactics. J. Comp. Neurol. 521, 2850–2869. doi: 10.1002/cne.23320
Fleige, S., and Pfaffl, M. W. (2006). RNA integrity and the effect on the real-time qRT-PCR performance. Mol. Aspects Med. 27, 126–139. doi: 10.1016/j.mam.2005.12.003
Forlano, P. M., Deitcher, D. L., and Bass, A. H. (2005). Distribution of estrogen receptor alpha mRNA in the brain and inner ear of a vocal fish with comparisons to sites of aromatase expression. J. Comp. Neurol. 483, 91–113. doi: 10.1002/cne.20397
Forlano, P. M., Deitcher, D. L., Myers, D. A., and Bass, A. H. (2001). Anatomical distribution and cellular basis for high levels of aromatase activity in the brain of teleost fish: aromatase enzyme and mRNA expression identify glia as source. J. Neurosci. 21, 8943–8955.
Forlano, P. M., Schlinger, B. A., and Bass, A. H. (2006). Brain aromatase: new lessons from non-mammalian model systems. Front. Neuroendocrinol. 27:2. doi: 10.1016/j.yfrne.2006.05.002
Garcia-Segura, L. M. (2008). Aromatase in the brain: not just for reproduction anymore. J. Neuroendocrinol. 20, 705–712. doi: 10.1111/j.1365-2826.2008.01713.x
Garcia-Segura, L. M., Veiga, S., Sierra, A., Melcangi, R. C., and Azcoitia, I. (2003). Aromatase: a neuroprotective enzyme. Prog. Neurobiol. 71, 31–41. doi: 10.1016/j.pneurobio.2003.09.005
Garcia-Segura, L. M., Wozniak, A., Azcoitia, I., Rodriguez, J. R., Hutchison, R. E., and Hutchison, J. B. (1999). Aromatase expression by astrocytes after brain injury: implications for local estrogen formation in brain repair. NSC 89, 567–578. doi: 10.1016/s0306-4522(98)00340-6
Heckmann, L. H., Sørensen, P. B., Krogh, P. H., and Sørensen, J. G. (2011). NORMA-Gene: a simple and robust method for qPCR normalization based on target gene data. BMC Bioinformatics 12:250. doi: 10.1186/1471-2105-12-250
Hutchison, J. B., and Steimer, T. (1986). Formation of behaviorally effective 17 beta-estradiol in the dove brain: steroid control of preoptic aromatase. Endocrinology 118, 2180–2187. doi: 10.1210/endo-118-6-2180
Kato, J., Yamada-Mouri, N., and Hirata, S. (1997). Structure of aromatase mRNA in the rat brain. J. Steroid Biochem. Mol. Biol. 61, 381–385. doi: 10.1016/S0960-0760(97)80036-2
Kobayashi, M., Aida, K., and Hanyu, I. (1988). Hormone changes during the ovulatory cycle in goldfish. Gen. Comp. Endocrinol. 69, 301–307. doi: 10.1016/0016-6480(88)90018-4
Lephart, E. D. (1996). A review of brain aromatase cytochrome P450. Brain Res. Brain Res. Rev. 22, 1–26. doi: 10.1016/0165-0173(96)00002-1
Makantasi, P., and Dermon, C. R. (2014). Estradiol treatment decreases cell proliferation in the neurogenic zones of adult female zebrafish (Danio rerio) brain. Neuroscience 277, 306–320. doi: 10.1016/j.neuroscience.2014.06.071
Marino, M., Galluzzo, P., and Ascenzi, P. (2006). Estrogen signaling multiple pathways to impact gene transcription. Curr. Genomics 7, 497–508. doi: 10.2174/138920206779315737
Marlatt, V. L., Martyniuk, C. J., Zhang, D., Xiong, H., Watt, J., Xia, X., et al. (2008). Auto-regulation of estrogen receptor subtypes and gene expression profiling of 17β-estradiol action in the neuroendocrine axis of male goldfish. Mol. Cell. Endocrinol. 283, 38–48. doi: 10.1016/j.mce.2007.10.013
Martínez-Cerdeno, V., Noctor, S. C., and Kriegstein, A. R. (2006). Estradiol stimulates progenitor cell division in the ventricular and subventricular zones of the embryonic neocortex. Eur. J. Neurosci. 24, 3475–3488. doi: 10.1111/j.1460-9568.2006.05239.x
Martyniuk, C. J., Xiong, H., Crump, K., Chiu, S., Sardana, R., Nadler, A., et al. (2006). Gene expression profiling in the neuroendocrine brain of male goldfish (Carassius auratus) exposed to 17 -ethinylestradiol. Physiol. Genomics 27, 328–336. doi: 10.1152/physiolgenomics.00090.2006
McEwen, B. (2002). Estrogen actions throughout the brain. Recent Prog. Horm. Res. 57, 357–384. doi: 10.1210/rp.57.1.357
Menuet, A., Pellegrini, E., Anglade, I., Blaise, O., Laudet, V., Kah, O., et al. (2002). Molecular characterization of three estrogen receptor forms in zebrafish: binding characteristics, transactivation properties, and tissue distributions. Biol. Reprod. 66, 1881–1892. doi: 10.1095/biolreprod66.6.1881
Menuet, A., Pellegrini, E., Brion, F., Gueguen, M. M., Anglade, I., Pakdel, F., et al. (2005). Expression and estrogen-dependent regulation of the zebrafish brain aromatase gene. J. Comp. Neurol. 485, 304–320. doi: 10.1002/cne.20497
Okada, M., Murase, K., Makino, A., Nakajima, M., Kaku, T., Furukawa, S., et al. (2008). Effects of estrogens on proliferation and differentiation of neural stem/progenitor cells. Biomed. Res. 29, 163–170. doi: 10.2220/biomedres.29.163
Pellegrini, E., Diotel, N., Vaillant-Capitaine, C., Pérez Maria, R., Gueguen, M. M., Nasri, A., et al. (2015). Steroid modulation of neurogenesis: focus on radial glial cells in zebrafish. J. Steroid Biochem. Mol. Biol. doi: 10.1016/j.jsbmb.2015.06.011. [Epub ahead of print].
Pellegrini, E., Menuet, A., Lethimonier, C., Adrio, F., Gueguen, M.-M., Tascon, C., et al. (2005). Relationships between aromatase and estrogen receptors in the brain of teleost fish. Gen. Comp. Endocrinol. 142, 60–66. doi: 10.1016/j.ygcen.2004.12.003
Pellegrini, E., Mouriec, K., Anglade, I., Menuet, A., Le Page, Y., Gueguen, M.-M., et al. (2007). Identification of aromatase-positive radial glial cells as progenitor cells in the ventricular layer of the forebrain in zebrafish. J. Comp. Neurol. 501, 150–167. doi: 10.1002/cne.21222
Pinto, C., Grimaldi, M., Boulahtouf, A., Pakdel, F., Brion, F., Aït-Aïssa, S., et al. (2014). Selectivity of natural, synthetic and environmental estrogens for zebrafish estrogen receptors. Toxicol. Appl. Pharmacol. 280, 60–69. doi: 10.1016/j.taap.2014.07.020
Strobl-Mazzulla, P. H., Lethimonier, C., Gueguen, M.-M., Karube, M., Fernandino, J. I., Yoshizaki, G., et al. (2008). Brain aromatase (Cyp19A2) and estrogen receptors, in larvae and adult pejerrey fish Odontesthes bonariensis: neuroanatomical and functional relations. Gen. Comp. Endocrinol. 158, 191–201. doi: 10.1016/j.ygcen.2008.07.006
Tchoudakova, A., Kishida, M., Wood, E., and Callard, G. V. (2001). Promoter characteristics of two cyp19 genes differentially expressed in the brain and ovary of teleost fish. J. Steroid Biochem. Mol. Biol. 78, 427–439. doi: 10.1016/S0960-0760(01)00120-0
Tong, S.-K., Mouriec, K., Kuo, M.-W., Pellegrini, E., Gueguen, M.-M., Brion, F., et al. (2009). A cyp19a1b-gfp(aromatase B) transgenic zebrafish line that expresses GFP in radial glial cells. Genesis 47, 67–73. doi: 10.1002/dvg.20459
Tsuruo, Y., Ishimura, K., and Osawa, Y. (1995). Presence of estrogen receptors in aromatase-immunoreactive neurons in the mouse brain. Neurosci. Lett. 195, 49–52. doi: 10.1016/0304-3940(95)11779-V
Ubuka, T., Haraguchi, S., Tobari, Y., Narihiro, M., Ishikawa, K., Hayashi, T., et al. (2014). Hypothalamic inhibition of socio-sexual behaviour by increasing neuroestrogen synthesis. Nat. Commun. 5, 3061. doi: 10.1038/ncomms4061
Ubuka, T., and Tsutsui, K. (2014). Review: neuroestrogen regulation of socio-sexual behavior of males. Front. Neurosci. 8:323. doi: 10.3389/fnins.2014.00323
Xing, L., Goswami, M., and Trudeau, V. L. (2014). Radial glial cell: critical functions and new perspective as a steroid synthetic cell. Gen. Comp. Endocrinol. 203, 181–185. doi: 10.1016/j.ygcen.2014.03.010
Xing, L., McDonald, H., Da Fonte, D. F., Gutierrez-Villagomez, J. M., and Trudeau, V. L. (2015). Dopamine D1 receptor activation regulates the expression of the estrogen synthesis gene aromatase B in radial glial cells. Front. Neurosci. 9:310. doi: 10.3389/fnins.2015.00310
Yague, J. G., Muñoz, A., de Monasterio-Schrader, P., Defelipe, J., Garcia-Segura, L. M., and Azcoitia, I. (2006). Aromatase expression in the human temporal cortex. Neuroscience 138, 389–401. doi: 10.1016/j.neuroscience.2005.11.054
Zhang, D., and Trudeau, V. L. (2006). Integration of membrane and nuclear estrogen receptor signaling. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 144, 306–315. doi: 10.1016/j.cbpa.2006.01.025
Zhao, C., Fujinaga, R., Tanaka, M., Yanai, A., Nakahama, K., and Shinoda, K. (2007). Region-specific expression and sex-steroidal regulation on aromatase and its mRNA in the male rat brain: immunohistochemical and in situ hybridization analyses. J. Comp. Neurol. 500, 557–573. doi: 10.1002/cne.21193
Zhao, E., Basak, A., Crump, K., and Trudeau, V. L. (2006). Proteolytic processing and differential distribution of secretogranin-II in goldfish. Gen. Comp. Endocrinol. 146, 100–107. doi: 10.1016/j.ygcen.2005.10.007
Keywords: 17β-estradiol, dopamine, aromatase, radial glial cell, teleost fish
Citation: Xing L, Esau C and Trudeau VL (2016) Direct Regulation of Aromatase B Expression by 17β-Estradiol and Dopamine D1 Receptor Agonist in Adult Radial Glial Cells. Front. Neurosci. 9:504. doi: 10.3389/fnins.2015.00504
Received: 26 September 2015; Accepted: 21 December 2015;
Published: 12 January 2016.
Edited by:
Sebastien G. Bouret, University of Southern California, USAReviewed by:
Ishwar Parhar, Monash University, MalaysiaPhilippe Ciofi, Institut National de la Santé et de la Recheche Médicale, France
Copyright © 2016 Xing, Esau and Trudeau. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Vance L. Trudeau, dHJ1ZGVhdXZAdW90dGF3YS5jYQ==
 Lei Xing
Lei Xing Crystal Esau
Crystal Esau Vance L. Trudeau
Vance L. Trudeau