- 1Department of Neuroscience, Washington University School of Medicine, St. Louis, MO, United States
- 2Department of Developmental Biology, Washington University School of Medicine, St. Louis, MO, United States
- 3Center of Regenerative Medicine, Washington University School of Medicine, St. Louis, MO, United States
- 4Department of Pathology and Immunology, Washington University School of Medicine, St. Louis, MO, United States
- 5Hope Center for Neurological Disorders, Washington University School of Medicine, St. Louis, MO, United States
Injured sensory neurons activate a transcriptional program necessary for robust axon regeneration and eventual target reinnervation. Understanding the transcriptional regulators that govern this axon regenerative response may guide therapeutic strategies to promote axon regeneration in the injured nervous system. Here, we used cultured dorsal root ganglia neurons to identify pro-regenerative transcription factors. Using RNA sequencing, we first characterized this neuronal culture and determined that embryonic day 13.5 DRG (eDRG) neurons cultured for 7 days are similar to e15.5 DRG neurons in vivo and that all neuronal subtypes are represented. This eDRG neuronal culture does not contain other non-neuronal cell types. Next, we performed RNA sequencing at different time points after in vitro axotomy. Analysis of differentially expressed genes revealed upregulation of known regeneration associated transcription factors, including Jun, Atf3 and Rest, paralleling the axon injury response in vivo. Analysis of transcription factor binding sites in differentially expressed genes revealed other known transcription factors promoting axon regeneration, such as Myc, Hif1α, Pparγ, Ascl1a, Srf, and Ctcf, as well as other transcription factors not yet characterized in axon regeneration. We next tested if overexpression of novel candidate transcription factors alone or in combination promotes axon regeneration in vitro. Our results demonstrate that expression of Ctcf with Yy1 or E2f2 enhances in vitro axon regeneration. Our analysis highlights that transcription factor interaction and chromatin architecture play important roles as a regulator of axon regeneration.
Introduction
Neurons within the central nervous system lack the intrinsic capacity to regenerate their axons after injury, leading to permanent functional deficits. In contrast, peripheral sensory neurons with cell soma in the dorsal root ganglia (DRG) can switch to a regenerative state after nerve injury to enable axon regeneration and functional recovery. Defining how injured sensory neurons transition to a pro-regenerative state may suggest future therapeutic approaches to improve neuronal recovery following axon injury.
Sensory neurons project both a peripheral axon branch into peripheral nerves and a central axon branch through the dorsal root into the spinal cord, providing a unique model system to study the mechanisms that control the axon regeneration program. Lesion of the peripheral axon is followed by robust and successful regeneration, whereas outgrowth of the centrally projecting axons is weak and does not lead to functional recovery. Multiple studies have utilized this differential response to gain insight into the early transcriptional events associated with successful regeneration (Smith and Skene, 1997; Stam et al., 2007; Blackmore, 2012; Mahar and Cavalli, 2018). A growing number of transcription factors (TFs) have been functionally linked to axon growth and regeneration (Venkatesh and Blackmore, 2017; Mahar and Cavalli, 2018). TFs work both independently of and in concert with epigenetic modifiers to increase the expression of pro-regenerative genes after injury (Weng et al., 2016; Fawcett and Verhaagen, 2018; Mahar and Cavalli, 2018) and represent ideal targets for therapeutic treatment of CNS injury (Fagoe et al., 2014; Venkatesh and Blackmore, 2017). One of the transcription factors that has emerged in many studies is ATF3, which was shown to drive the injured state and to be necessary for axonal regeneration and functional recovery (Chandran et al., 2016; Renthal et al., 2020). However, while ATF3 overexpression can promote peripheral axon regeneration (Seijffers et al., 2007), ATF3 fails to do so in several models of CNS injury (Seijffers et al., 2006; Fagoe et al., 2015; Venkatesh et al., 2016). This may be due to the fact that differentially expressed genes containing an ATF3 binding motif and the epigenomic signatures are largely distinct after spinal cord injury compared to nerve injury (Palmisano et al., 2019; Ewan et al., 2021). Another possibility is that a combination of transcription factors may provide more robust regeneration than a single TF alone (Chandran et al., 2016; Wang et al., 2018; Venkatesh et al., 2021). While genetic manipulation of TFs is a promising strategy, no single TF will likely be sufficient to fully restore neuron-intrinsic growth potential, and multiple, functionally interacting factors will be needed.
Selecting the optimal combination of TF has remained a difficult challenge (Venkatesh and Blackmore, 2017). One reason for this is that previous sequencing and bioinformatics studies have used whole DRG as input for sequencing (Michaelevski et al., 2010; Puttagunta et al., 2014; Chandran et al., 2016; Palmisano et al., 2019). The DRG is a highly heterogeneous cell population that consists of several cell types, including neurons, satellite glial cells, and macrophages (Kwon et al., 2013; Niemi et al., 2013; Avraham et al., 2020), limiting downstream bioinformatics analysis (Omura et al., 2015; Chandran et al., 2016; Tedeschi et al., 2016; Palmisano et al., 2019; Shin et al., 2019). Single-cell approaches have been used to unravel the transcriptional response of sensory neurons to nerve injury in vivo (Renthal et al., 2020), but the limited depth of sequencing also limits the analysis of the TFs implicated in the regeneration program.
To address this challenge, we characterized an in vitro model of embryonic DRG neurons that is widely used for studies of axon injury responses (Miller et al., 2009; Sasaki et al., 2009, 2020; Cho and Cavalli, 2012; Cho et al., 2013, 2015; Oh et al., 2018; Bloom et al., 2022). RNA-sequencing of eDRG cultures treated with a mitotic inhibitor revealed that a week after plating, these cultures are highly enriched in sensory neurons, and contain some neural progenitors but do not contain non-neuronal cells. Comparison to an in vivo single-cell data set of DRG cells traversing the primary sensory neuron lineage (Sharma et al., 2020) indicated that neurons in these eDRG cultures remained at an embryonic stage with all neuronal subtypes represented. Analysis of differentially expressed genes after injury revealed downregulation of pathways related to ion channels and upregulation of known regeneration-associated genes, including Jun, Atf3, and Rest, paralleling the axon injury response in vivo (Broude et al., 1997; Tedeschi et al., 2016; Lisi et al., 2017; Oh et al., 2018; Renthal et al., 2020). Analysis of TFs binding sites in differentially expressed genes revealed other known transcription factors promoting axon regeneration, such as Myc (Belin et al., 2015), Hif1a (Cho et al., 2015), Pparg (Lezana et al., 2016), Ascl1a (Williams et al., 2015), Srf (Stern et al., 2013), Ctcf (Palmisano et al., 2019), as well as other transcription factors not yet characterized in axon regeneration. We tested if overexpression of candidate pro-regenerative TFs alone or in combination promotes axon regeneration in vitro. Our results demonstrate that the expression of Ctcf, which is also known for its role in chromatin three-dimensional organization, does not enhance axon regeneration. However, a combination of Ctcf with Yy1 or E2f2 promotes in vitro axon regeneration. Our analysis reveals that pairs of TFs can functionally synergize to promote axon regeneration and highlights that TF interactions and chromatin architecture are essential regulators of axon regeneration.
Results
The eDRG spot culture model contains mostly neurons
The in vitro model of embryonic DRG neurons is widely used for studies of axon injury responses related to both axon degeneration and axon regeneration (Miller et al., 2009; Sasaki et al., 2009, 2020; Cho and Cavalli, 2012; Cho et al., 2013, 2015; Oh et al., 2018; Bloom et al., 2022). Yet, what type of neurons and what developmental stage is present in this model have not been examined in detail. In this culture model, embryonic day 13.5 DRG are spot cultured in the presence of the mitotic inhibitor 5-fluorodeoxyuridine (FDU). To determine the purity of these cultures, we stained for satellite glial cells, the most abundant cell type in DRG, with antibodies to FABP7 (Avraham et al., 2020). We observed that at days in vitro 1 (DIV1), FABP7 positive glial cells are present in high numbers (Figures 1A,B) but are nearly absent by DIV7 (Figures 1A,B), consistent with our previous results (Avraham et al., 2020). FDU treatment eliminates non-neuronal proliferating cells from the culture, without affecting neuronal properties and viability (Lesslich et al., 2022). Consistently, we did not observe pycnotic nuclei in our FDU-treated cultures compared to no FDU. We also did not detect any differences in neuronal morphology or axon elongation between the cultures that were treated with or without FDU (Figure 1C and Supplementary Figure 1). We next collected the mRNA from DRG spot cultures at DIV7 and submitted them for RNA sequencing. We examined the expression level of cell type-specific marker genes: satellite glial cells (Fabp7, Kir4.1, and Cdh19), myeloid/macrophages (Cd45, Cd68, and Iba1), endothelial (Pecam, Chh5, and Cldn5), Schwann cells (Egr2, Gfap, and Mpz), neural crest (Foxd3, Sox2, and Nes), neural progenitor markers (Rtn4, Plxna4, and Dpysl3) and sensory neurons (Avil, Calca, TrkA, and Tubb3). Compared to the average expression of sensory neuron marker genes and neural progenitor marker genes, the expression of non-neuronal cells marker genes and neural crest genes were all very low, indicating that these non-neuronal cell types are very low or absent in our preparation (Figure 1D). We conclude that the presence of FDU causes the progressive death of essentially all non-neuronal cells, resulting in a culture model that includes only sensory neurons and neural progenitors after 7 days in vitro.
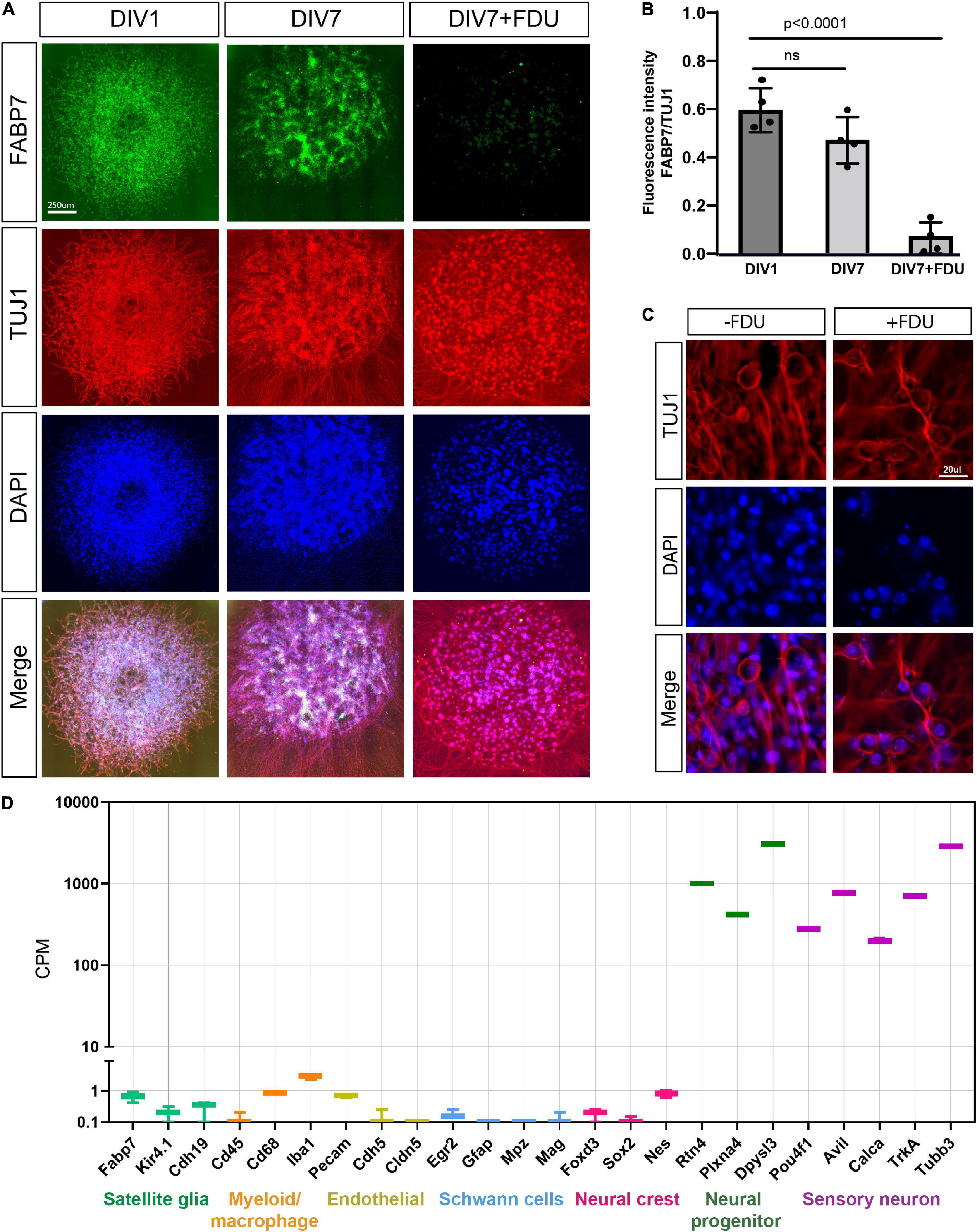
Figure 1. eDRG spot culture contains mostly sensory neurons after treatment with FDU. (A) Representative images of eDRG culture stained with the neuronal marker TUJ1 (red) and the Satellite glia marker FABP7 (green) at DIV1 and at DIV7 with and without treatment with FDU. Scale Bar: 250 μm. (B) Quantification of the fluorescence intensity of FABP7 normalized to TUJ1 at DIV1 and DIV7 ± FDU. n = 4 biologically independent animals. P values are determined by One way ANOVA. Data are presented as mean values ± SEM. (C) Representative images of eDRG culture in high magnification, with and without FDU, stained with TUJ1 (red) and DAPI (blue). Scale Bar: 20 μm. (D) Quantification of selected neuronal and non-neuronal markers (counts per million) from RNAseq analysis of eDRG spot culture at DIV7 treated with FDU.
The eDRG spot culture model contains most neuronal subtypes present at E15.5 in vivo
Using this model, we next sought to understand whether neurons mature in vitro or remain at an unspecified stage and whether neuronal subtypes are present. Indeed, sensory neurons represent a heterogenous neuronal population that is specified during development in part by target-derived specific cues, such as NGF. In the eDRG spot culture model (eDRG spot), neurons are collected at E13.5 and cultured for 7 days in the presence of NGF. We thus compared the transcriptional profile of eDRG 7 days after plating to the transcriptomic atlas of cells traversing the somatosensory neuron lineage in mice (Sharma et al., 2020). We obtained the data from Sharma et al. for DRG at four developmental stages: E15.5, when innervation of peripheral and central target fields occurs; P0 when the maturation of sensory nerve ending with skin and other organs occurs and P5, when most peripheral endings mature into morphological states and central projection terminals are properly organized within select spinal cord laminae, and adult (P28-P42) (Sharma et al., 2020). From the single cell data, 15 neuronal subtypes were defined across these four developmental stages (Figure 2A and Supplementary Table 1) and each subtype was defined by the 4 most expressed genes (Supplementary Table 2). Fifty subtype-specific genes were then identified using the FindMarker function in Seurat (Supplementary Table 3). All genes were spot-checked by overlaying the expression levels on the t-SNE plot to ensure the computational method was correctly identifying genes with the prescribed features. We then combined read counts per subtypes, including scRNAseq and our eDRG spot RNAseq. The data were normalized with respect to library size and count differences were minimized between samples. We then calculated counts per million (CPM) and reads per kilobase million (RPKM) values and generated heat maps for the 50 enriched genes in each subtype. eDRG spot clustered closely to E15.5 in 11 out of 15 subtypes (Figure 2B). eDRG spot clustered with CGRP-Zeta from both E15.5 and P0 and with AbetaRA-LTMR and CGRP_Eta at E15.5, P0, and P5 (Figure 2B). Interestingly, the eDRG spot did not cluster with the unknown subtype (Figure 2B). This unknown subtype does not contain neural crest markers in the top 50 DEGs but expresses some neural progenitor genes, such as Mllt11, Tuba1a, Stmn1, and Stmn2 (Supplementary Table 3). The neural progenitor genes expressed in the eDRG spot (Figure 1D) may reflect a different population of neural progenitors compared to the in vivo unknown cluster or represent a different developmental stage of this cluster. We also visualized the expression of genes in each neuronal subtype during development using RPKM with two different cut-offs (Figure 2C). These data further indicate that the eDRG spot model is most similar to E15.5 DRG in vivo and that most neuronal subtypes are present. These results suggest that while the eDRG spot is an embryonic culture, their stage at DIV7 is in between E17.5, in which axon growth in vitro is decreased and synapses have already formed in vivo, and E12.5, in which axon growth is active and synaptogenesis has not yet occurred (Prasad and Weiner, 2011; Tedeschi et al., 2016).
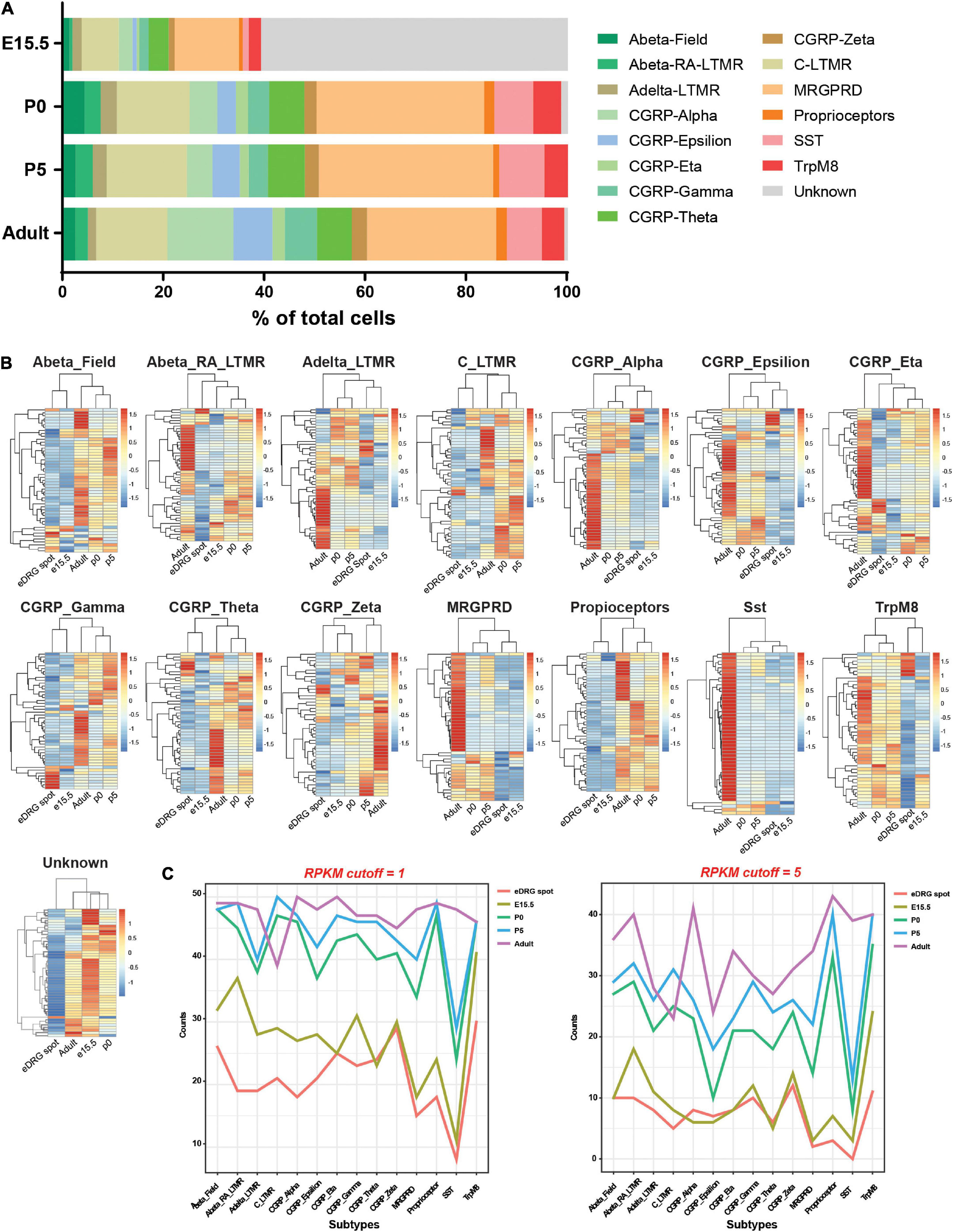
Figure 2. Neurons in the eDRG at DIV7 are similar to DRG neurons at the E15.5 developmental stage. (A) Fraction of 15 neuronal subtypes across four developmental stages of DRG neurons. (B) Heatmap shows the scaled z-score of the average CPM of 50 subtype-specific genes from eDRG controls and 4 developmental stages. Both row and column clustering were applied. High and low expressions are indicated in red and blue, respectively. (C) Plots of the counts of 50 subtype-specific genes expressed in each neuronal subtype during development using RPKM with cutoffs of 1 and 5.
Transcriptional changes are induced by in vitro axotomy over time
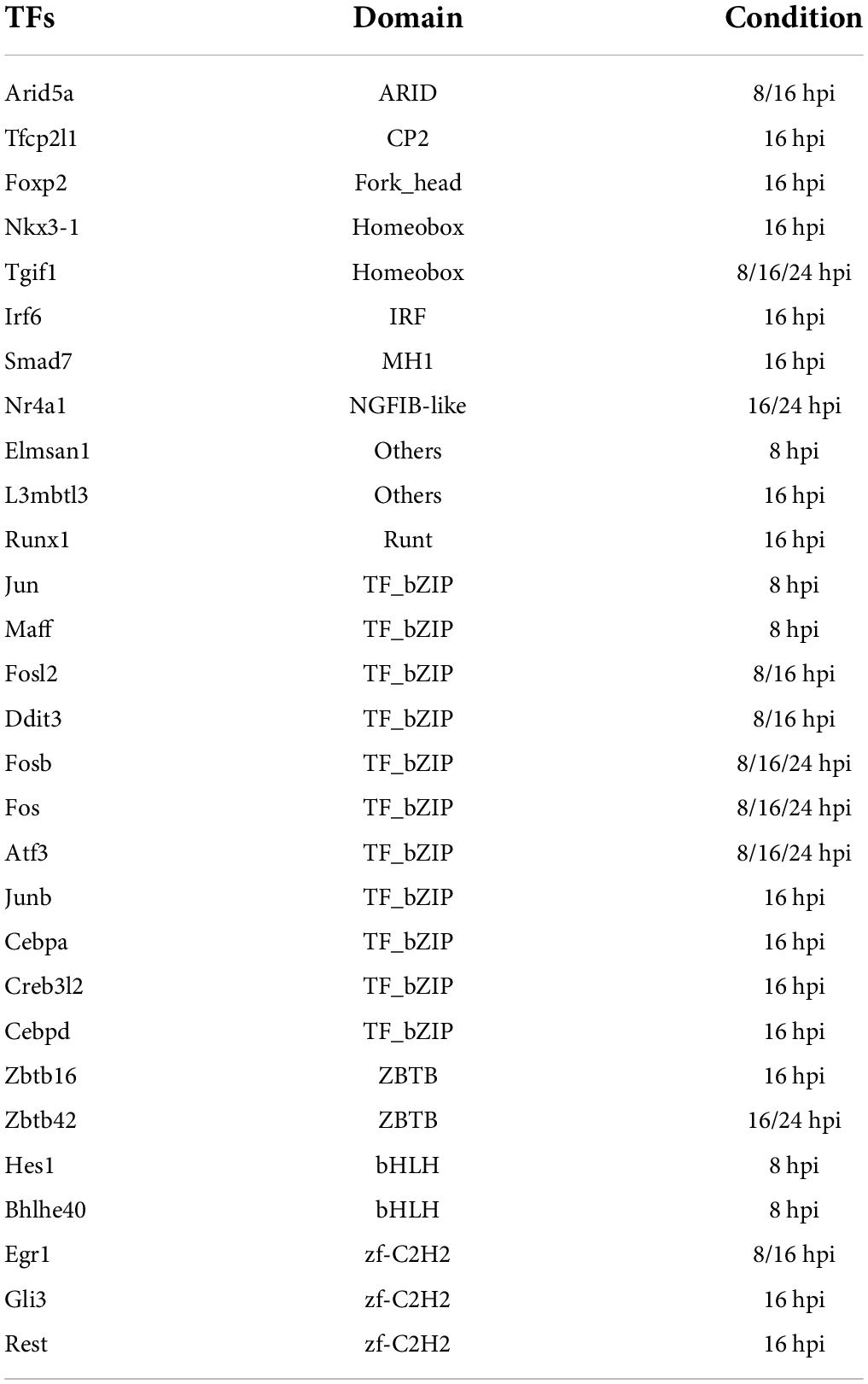
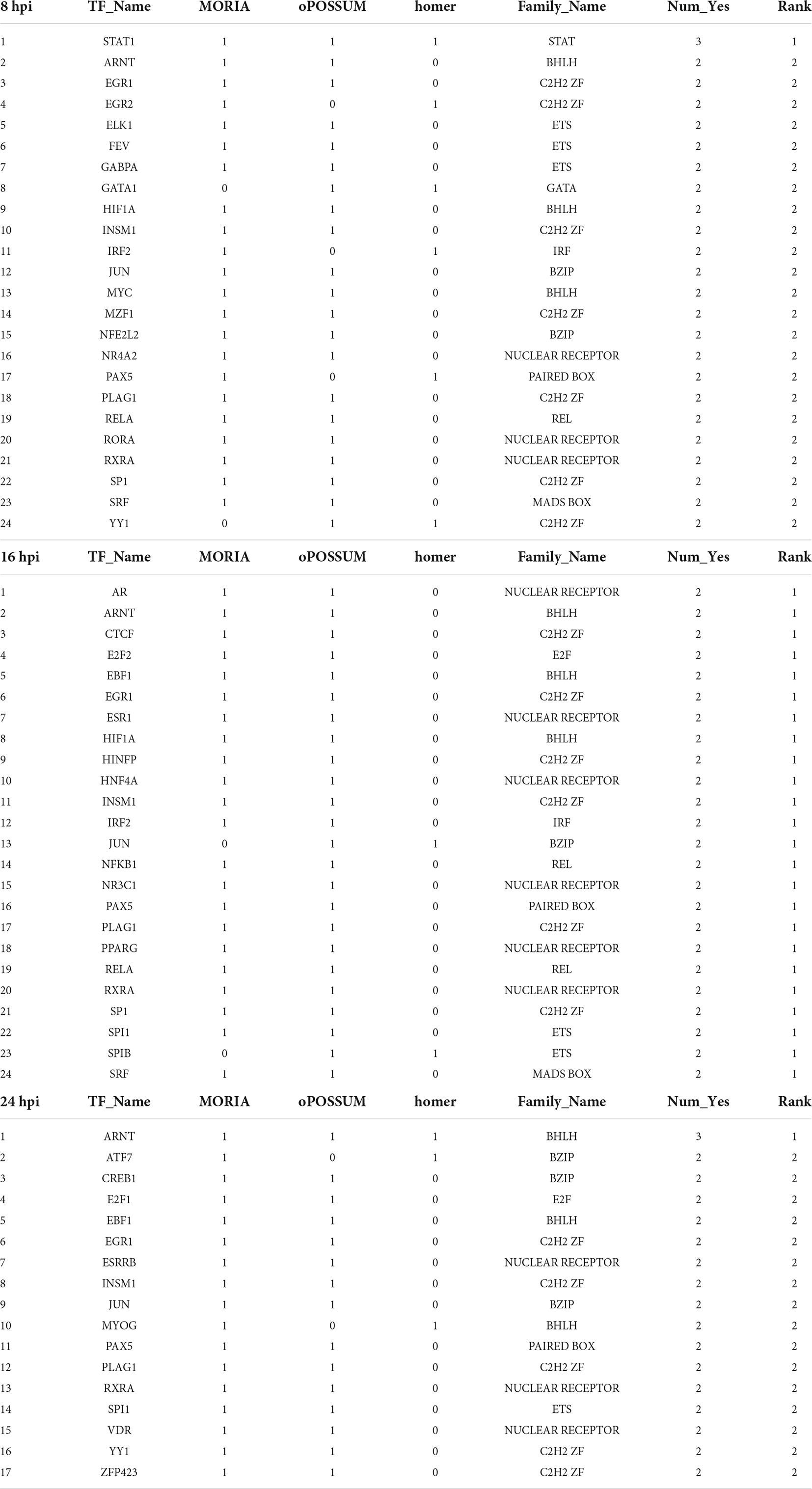
To unravel the neuron-intrinsic transcriptional changes in DRG neurons that arise after axon injury, we cut the axons of DRG neurons at DIV7 with a blade to induce an injury response and collected mRNA at 1, 3, 8, 16, and 24 h post injury (HPI). Hierarchical clustering and PCA analysis with batch correction revealed that 16 and 24 HPI displayed the most significant changes (Figures 3A,B). We identified a total of 467 differentially expressed genes (DEG), with most changes occurring 8–24 HPI (> 2-fold change, Padj < 0.05) and most DEG genes being upregulated (Figure 3C and Supplementary Table 4). Among the DEG, we identified 5 unique gene profile clusters with similar expression dynamics over time post-injury (Figures 3D,E and Supplementary Table 5). Gene ontology (GO) analysis of each cluster revealed unique biological pathways (Supplementary Figure 2). Clusters 4 and 5, in which genes are downregulated after injury, revealed enrichment for pathways related to ion channel activity, neurotransmitter, and synapse. These results parallel those of previous reports of the axon injury response in vivo, which demonstrated that ion channels are downregulated after injury, a process required for PNS regeneration (Tedeschi et al., 2016; Lisi et al., 2017; Oh et al., 2018). Pro-regenerative TFs known for their role in vivo, such as Jun, Atf3, and Rest, were found in clusters 2 and 3, in which expression peaked at 8 and 16 HPI, respectively. Cluster 2 and cluster 3 were enriched for pathways related to transcription factor, DNA binding, MAPK signaling, p53 signaling, and Wnt signaling, which are also known to regulate axon regeneration in vivo (Smith and Skene, 1997; Di Giovanni et al., 2006; Shin et al., 2012; Mahar and Cavalli, 2018). To further determine if the DEG elicited by in vitro axotomy is related to the in vivo situation, we compared our results to two recent studies that identified DEG in adult mice sensory neurons following sciatic nerve crush injury (Renthal et al., 2020; Ewan et al., 2021). We found 67 genes that were shared between the eDRG spot and at least one of the in vivo injury datasets (Figure 3F and Supplementary Table 4). To further determine if known pro-regenerative TFs are expressed in the eDRG spot model, we examined the up-regulated DEG and found that 29 were TFs, with 10 of them showing up-regulation at 8, 16, and 24 HPI (Table 1). These included TF known for their pro-regenerative function, such as Rest, Jun, and Atf3, paralleling the axon injury response in vivo (Broude et al., 1997; Tsujino et al., 2000; Mahar and Cavalli, 2018; Oh et al., 2018; Carlin et al., 2019; Renthal et al., 2020). We found that 11 of the 29 TFs were also differentially expressed in adult sensory neurons following nerve injury (Figure 3G and Table 1). To identify TFs that may regulate the expression of the upregulated DEG at 8, 16, and 24 HPI, we used an in-house algorithm that combines different analysis methods to predict TF binding sites (MORA, oPOSSUM3, and HOMER) (Zhao et al., 2007). We found a total of 43 TFs that were predicted in at least 2 different analysis methods (Table 2). These included previously characterized TFs, such as Myc (Belin et al., 2015), Hif1a (Cho et al., 2015), Pparg (Lezana et al., 2016), Ascl1a (Williams et al., 2015), Srf (Stern et al., 2013) and Ctcf (Palmisano et al., 2019), as well as other transcription factors not yet characterized in axon regeneration.
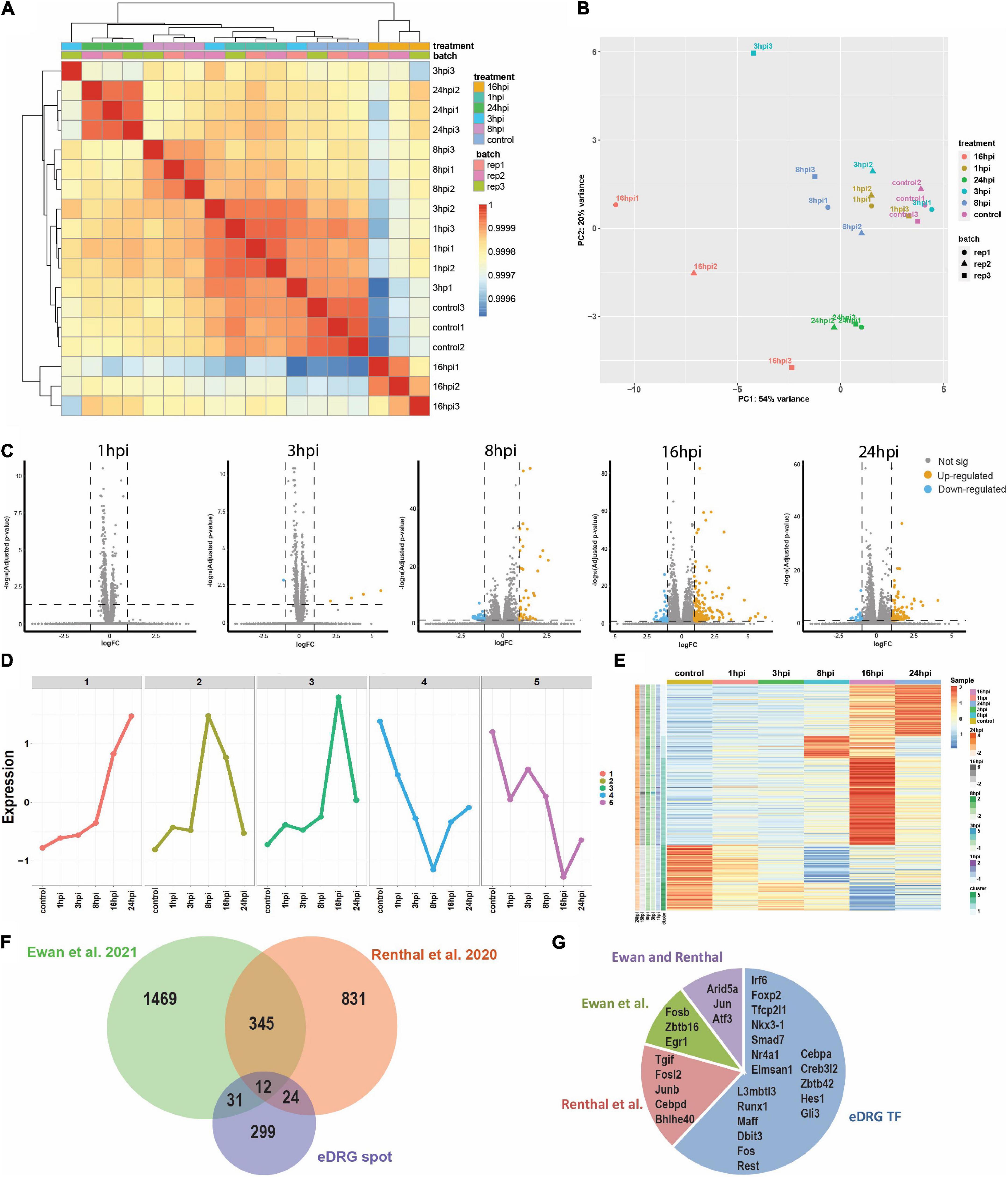
Figure 3. Time course analysis (1, 3, 8, 16, and 24 h) of the transcriptional response to injury in eDRG spot culture. (A) Heatmap of correlation of samples from time-course RNAseq analysis after axotomy. (B) PCA analysis of time-course RNAseq analysis after axotomy illustrates the relative similarity between sample groups at control, 1, 3, 8, 16, and 24 hpi (3 replicates per time point). (C) Volcano plots of differentially regulated genes after injury (p adj < 0.05, FC > 2). (D) K-means clustering identified five unique gene profile clusters with similar gene expression dynamics according to the time-course gene expression data. (E) Heatmap shows the scale z-score of the average CPM of 365 differentially expressed genes from time-course data. Red and blue cells indicate relative gene upregulation and downregulation, respectively. Green cells indicate cluster types identified in D. the cells next to cluster type are genes that are significantly expressed at different time points between treated and control groups. (F) Venn diagram comparing the differentially expressed genes after injury in the eDRG spot culture and 2 datasets of in vivo adult injury using enriched neuronal populations. (G) Out of the 29 up-regulated TFs in the eDRG spot, 11 were also identified in adult DRG neurons following sciatic nerve crush.
Lentiviral-based expression of pro-regenerative TF identifies TF combination that improves axon regeneration in vitro
Because it is likely that a single TF is not sufficient to fully restore neuron-intrinsic axon growth potential, and that a combination of TFs is needed (Venkatesh and Blackmore, 2017), we decided to overexpress combinations of TFs identified in our analysis, and test if a given TF combination can promote regeneration. We used the eDRG spot culture model to screen the effect of lentiviral mediated expression of TFs combinations on regenerative axon growth 24 h after in vitro axotomy, as described previously (Cho et al., 2013, 2015; Avraham et al., 2020). We selected 8 TFs that were predicted to regulate DEG after injury in our analysis and were also identified in different in vivo regeneration models but have not been directly tested for their pro-regenerative potential. These are Atf3 (Tsujino et al., 2000; Chandran et al., 2016; Renthal et al., 2020), Fos (Stam et al., 2007; Michaelevski et al., 2010), Egr1 (Zou et al., 2009; Michaelevski et al., 2010), Nfya (Smith et al., 2011), Ebf1 (Smith et al., 2011), E2f2 (Pita-Thomas et al., 2021), Ctcf (Palmisano et al., 2019; Pita-Thomas et al., 2021), Yy1 (Stam et al., 2007). Interestingly, CTCF is also known as a chromatin remodeler and conditional deletion of CTCF in vivo impairs nerve regeneration, implicating chromatin organization in the regenerative competence (Palmisano et al., 2019). Whether CTCF expression is sufficient to enhance axon regeneration had not been tested.
eDRG spot cultured neurons were infected with lentiviruses expressing the selected TFs in pairwise combination 4 days before axotomy (Figure 4A). Axons were injured at DIV7, immunostained for the regeneration marker SCG10 24 h later and, analysis for axon growth past the injury site was performed as described previously (Figure 4A; Cho et al., 2015; Avraham et al., 2020). Axonal length after treatment with an expression of TF combinations was normalized to lentivirus expressing mCherry as a control. We verified that in these conditions, transduction efficiency was high by detection of mCherry in > 90% of neurons (Supplementary Figure 3). We observed that a combination of the TFs Ctcf with Yy1 or E2f2 had a significant effect on regenerative axon growth (Figures 4B,C), but there was no significant increase in axon growth when Ctcf, E2f2, or Yy1 were expressed alone (Figures 4C,D). These results identify Ctcf, Yy1, and E2f2 as new pro-regenerative TFs and highlight that TF interaction and chromatin remodeling play an important role as a regulator of axon regeneration.
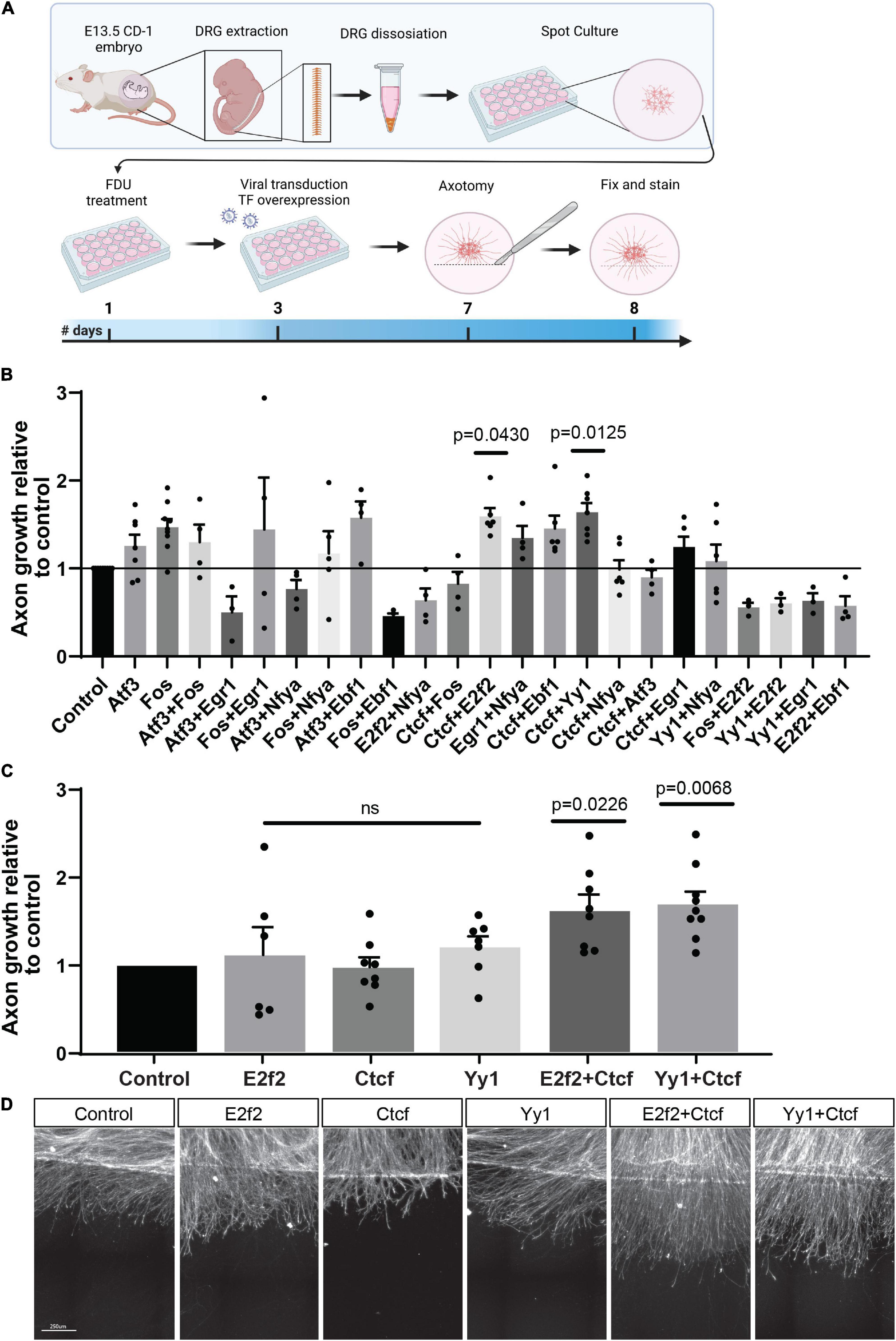
Figure 4. Overexpression of TF combinations in eDRG axotomy model. (A) Schematic timeline of the experimental procedure for TF overexpression in injured cultured eDRG neurons. (B) Axon growth of infected cells with TF combinations relative to control. (C) Significant TF were tested alone and repeated in combinations. (D) Represented images of axotomized cultures infected with the pro-regenerative TFs alone and in combinations. Scale Bar: 250 μm. n = 3–9 biologically independent animals examined over 2 independent experiments P-values determined by one-way ANOVA followed by Dunnett’s multiple comparisons test. Data are presented as mean values ± SEM.
Discussion
In the present study, we used RNA-sequencing and bioinformatic analyses to identify pairs of TFs that synergize to promote axon regeneration. Our analysis provides a framework for the development of combinatorial gene over-expression approaches which may provide strategies to induce axon regeneration in vivo. We also characterized the eDRG spot culture model and determined that this model partially mimics the in vivo injury responses.
Our data suggest that the eDRG spot culture is a useful model to identify genes and transcription factors that regulate the neuron’s intrinsic capacity for axon regeneration. Our analysis reveals that this model contains sensory neurons that are in a stage between E12.5, in which axon growth is active and synaptogenesis has not occurred in vivo, and E17.5, in which axon growth in vitro is decreased and synapses have already formed in vivo. The DEG elicited by in vitro injury parallels in part those observed in vivo. However, this model does present limitations, given that non-neuronal cells are not present and thus contributions from satellite glial cells (Avraham et al., 2020, 2021), immune cells (Niemi et al., 2013) or other extrinsic factors, such as the microbiome (Serger et al., 2022), are not contributing to the neuronal response to injury. This model also does not allow to study the age-dependent neuronal regenerative decline (Zhou et al., 2022). The strengths of this model lie in the ability to study intrinsic neuronal mechanisms that regulate axon growth capacity in a defined and characterized neuronal population that are accessible to genetic and pharmacological manipulations.
Previous studies have taken different approaches to identify regulators of axon regeneration, including high throughput phenotypic screening (Simpson et al., 2015; Huebner et al., 2018; Karney-Grobe et al., 2018; Sekine et al., 2018) and transcriptional-based approaches (Michaelevski et al., 2010; Puttagunta et al., 2014; Cho et al., 2015; Omura et al., 2015; Chandran et al., 2016). Each method has provided key insight into the molecular mechanisms of regeneration and identified targets that can promote some axon regeneration. ATF3 is a key TF inducing the pro-regenerative state in sensory neurons and loss of ATF3 impairs axon regeneration (Tsujino et al., 2000; Seijffers et al., 2007; Renthal et al., 2020). Overexpression of ATF3 alone or in combination with c-jun was shown to improve DRG axon growth in vitro (Chandran et al., 2016) and had only a mild effect on cortical neuron neurite outgrowth in vitro (Simpson et al., 2015). However, ATF3 expression fails to promote robust axon regrowth in several models of CNS injury (Seijffers et al., 2006; Fagoe et al., 2015; Venkatesh et al., 2016). This may be due to the fact that differentially expressed genes containing an ATF3 binding motif and the epigenomic signatures are largely distinct after spinal cord injury compared to nerve injury (Palmisano et al., 2019; Ewan et al., 2021). Indeed, several epigenetic mechanisms implicating histone deacetylase (HDAC3 and HDAC5) and histone acyltransferase (PCAF) operate after peripheral but not central axon injury (Cho and Cavalli, 2012; Finelli et al., 2013; Puttagunta et al., 2014; Hervera et al., 2019). The combination of TFs expression with epigenetic modifiers might synergize to stimulate long-range axon regeneration.
Recent epigenomics studies revealed that neuronal conditional deletion of CTCF impaired nerve regeneration, implicating chromatin organization in the axon regenerative competence (Palmisano et al., 2019). Whether CTCF expression could promote axon regeneration had not been tested. CTCF is known as a chromatin remodeler with a critical role in connecting higher-order chromatin folding in pluripotent stem cells (Beagan et al., 2017). Our results suggest that CTCF expression in combination with E2F2 or YY1, but not alone, stimulates axon growth in vitro. This co-factor requirement appears specific since CTCF did not synergize with NFYA, FOS, ATF3, EGR1, or EBF1 in our assay or with KLF6 in a cortical growth assay (Venkatesh et al., 2021). In the context of genome imprinting, YY1 is a required cofactor for CTCF function in the X chromosome binary switch (Donohoe et al., 2007). Many clustered YY1 and CTCF binding sites are conserved among humans, mice, and cows (Kang et al., 2009). YY1 contributes to enhancer-promoter structural interactions in a manner analogous to DNA interactions mediated by CTCF (Weintraub et al., 2017), suggesting that enhancer-promoter looping facilitates gene expression required for axon regeneration. This may also underlie why combined TF expression without chromatin remodeler can increase collateral sprouting but not long-range regenerative growth through sites of the spinal lesion (Venkatesh et al., 2021). Indeed, prior work shows that the epigenetic landscape of pro-growth genes acquires marks of heterochromatin and transcriptional repression with age (Venkatesh et al., 2016). Reactivating the pro-growth program will require changes in the epigenomic landscape. Our algorithm that combines different analysis methods to predict TF binding sites provides useful insights into the selection of TF combinations and suggests that CTCF represents a potent target for those combinations. In combination with previous bioinformatics analyses of axon regeneration (Chandran et al., 2016; Venkatesh and Blackmore, 2017), our results will help guide future experiments to select optimal TFs combinations for promoting axon regeneration.
Materials and methods
Animals and procedures
All animal procedures were performed in accordance with the WashU animal care committee’s regulations. Time pregnant e13.5 CD-1 mice were used for all experiments.
Embryonic DRG cultures and regeneration assay
Dorsal root ganglia were isolated from time pregnant e13.5 CD-1 mice into dissection media consisting of DMEM and Pen/Strep. After a short centrifugation, dissection media was aspirated and cells were digested in 0.05% Trypsin-EDTA for 25 min. Next, cells were pelleted by centrifuging for 2 min at 500 × g, the supernatant was aspirated, and Neurobasal was added. Cells were then triturated 25x and added to the growth medium containing Neurobasal, B27 Plus, 5 μM FDU, 1 ng/ml NGF, Glutamax, and Pen/Strep. Approximately 10,000 cells were added to each well in a 2.5 μl spot. Spotted cells were allowed to adhere for 10 min before the addition of the growth medium. Plates were pre-coated with 100 μg/ml poly-D-lysine overnight and washed with sterile water prior to plating. For the regeneration assay, lentivirus was added on DIV3. Cells were then injured using an 8 mm microtome blade on DIV 7 and fixed 24 h later with 4% PFA. For immunostaining, wells were incubated in PBS-0.1% Triton (PBST) for 1 h at room temperature containing SCG10 primary antibody (Novus Bio NBP1-49461; RRID:AB_10011569). The wells were then washed 3x with PBST and then incubated in PBST solution containing fluorescent-labeled goat anti-rabbit secondary antibody (AlexaFluor-555; Invitrogen A21428; RRID:AB_141784) for 1 h at room temperature. Finally, wells were washed 3x with PBS. For immunostaining of spot culture at DIV1 and DIV7, wells were stained with TUJ1 primary antibody (Biolegend 802001; RRID:AB_2564645), Fabp7 (Thermo Fisher Scientific Cat# PA5-24949, RRID:AB_2542449) and DAPI (1:1,000).
Viral production
To produce lentivirus, HEK293T cells were grown on 15 cm plates to a confluency of 70–90%. On DIV0, 3 μg PMD2.G, 9 μg PsPax2, and 12 μg of target plasmid were incubated for 15 min with 96 μg of PEI Max (Polysciences 24765-1) per 15 cm dish in Optimem at room temperature. Following incubation, the transfection solution was added dropwise to the plate, gently mixed, and incubated for 96 h. Following incubation, viral-containing supernatant was collected and centrifuged at 500 × g for 10 min to remove cellular debris. Cleared supernatants were further filtered with a 0.45 μm PES filter to remove the remaining debris. A total of 30 μl of viral-containing supernatant was added into 500 μl of media in each well in a 24-well plate. Virus with mCherry was used as a control for infection efficiency and for axon regeneration. Overexpression clones in the lentiviral backbone vector Plx304 were obtained from the human lentiviral ORF library (transOMICs).

RNA sequencing and bioinformatics analysis
For RNA sequencing experiments, embryonic dorsal root ganglia were injured with a blade, isolated at 1, 3, 8, 16, and 24 h post-injury. Cells were lysed and total RNA was extracted using the PureLink RNA Mini Kit (Thermo Fisher Scientific 12183018A) with on-column DNase treatment (Thermo Fisher Scientific 12185010). Next, RNA concentration was determined using a NanoDrop 2000 (Thermo Fisher Scientific). First-strand synthesis was then performed using the High Capacity cDNA Reverse Transcription kit (Applied Biosystems).
Samples were submitted to the Genome Access Technology Center at Washington University in St. Louis for library preparation and sequencing. Libraries were sequenced on an Illumina HiSeq2500 using 2 × 101 bp runs.
Reads were processed using an in-house pipeline and open-source R packages. Briefly, raw reads were first trimmed using cutadapt to remove low-quality bases and reads. Trimmed reads were then aligned to the mouse genome 10 mm with GENCODE annotation vM20 using STAR (v2.5.4) with default parameters. Transcript quantification was performed using featureCounts from the subread package (v1.6.3). further quality control assessments were made using RSeQC and RSEM, and the batch correction was performed using edgeR, EDASeq, and RUVSeq. Gene type and transcription factor (TF) annotation were performed using mouse GENCODE vM20 and AnimalTFDB, respectively.
Principle component analysis and differential expression analysis for neurons collected at 1, 3, 8, 16, and 24 h post injury (HPI) and control groups were determined using DESeq2 in negative binomial mode using batch-corrected transcripts from featureCounts (> 2-fold expression change, > 1 count per million (CPM), Benjamini corrected P < 0.05). Pairwise comparisons were made between time points vs control to determine differentially expressed genes (DEGs) within each group. To determine transcription factor binding site enrichment, the significantly upregulated genes were used as input for oPOSSUM 3.0 for TF enrichment analysis with a default cutoff for statistical significance1. TF enrichment was also confirmed using the prediction of TF binding sites (MORIA and HOMER). The HOMER (Heinz et al., 2010) and motif over-representation analysis (MORA) (Zhao et al., 2007) algorithms were used to identify transcription factor binding sites enriched in the upstream regions compared with background sequences in the genome. An in-house script was used to generate a combined ranking of predicted TFs.
Image analysis
All images were acquired at 10x using a Nikon TE2000 microscope and image analysis was completed using Nikon Elements. For embryonic dorsal root ganglia experiments, regenerative length was measured from the visible blade mark to the end of the regenerating axons. Each technical replicate was measured 4–6 times and three technical replicates were measured per biological replicate.
Statistical analysis
All experimenters were blinded to treatment conditions while performing image analysis. All statistical analysis was completed using GraphPad Prism. Data are presented as ± SEM. All statistical values are reported in the text as appropriate.
Data availability statement
The datasets presented in this study can be found in online repositories. The name of the repository and accession number can be found below: National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO), https://www.ncbi.nlm.nih.gov/geo/, GSE138480.
Ethics statement
The animal study was reviewed and approved by the Washington University School of Medicine Institutional Animal Care and Use Committee (IACUC) under protocol A-3381-01. All experiments were performed in accordance with the relevant guidelines and regulations. All experimental protocols involving mice were approved by the Washington University School of Medicine (protocol #21-0104 and #20-0173). Mice were housed and cared for in the Washington University School of Medicine animal care facility. This facility is accredited by the Association for Assessment and Accreditation of Laboratory Animal Care (AALAC) and conforms to the PHS guidelines for Animal Care. Accreditation - 7/18/97, USDA Accreditation: Registration #43- R-008.
Author contributions
OA, JL, GZ, and VC conceived and designed the experiments. OA, JL, and KL performed the experiments. TL and GZ performed the bioinformatic analyses. OA and JL performed the data analysis. VC secured the funding. OA, GZ, and VC wrote the manuscript. All authors approved the submitted version.
Funding
This work was funded in part by a bioinformatic grant from the Center of Regenerative Medicine and NIH grants R35 NS122260 and R01 NS096034 to VC.
Acknowledgments
We would like to thank members of the Cavalli lab for valuable comments and discussions. We would also like to thank Marcus Mahar for his technical help with eDRG culture. We also thank Bo Zhang from the bioinformatic core for his assistance and help with bioinformatic analyses.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fnmol.2022.967472/full#supplementary-material
Supplementary Figure 1 | Representative images of eDRG spot culture at DIV7 at low magnification, with and without FDU, stained with TUJ1. Scale bar: 1,000 μm.
Supplementary Figure 2 | Gene Ontology and KEGG pathway analysis (mouse 2019) for the different gene profile clusters with similar gene expression dynamics.
Supplementary Figure 3 | eDRG spot culture transduced with a lentivirus expressing mCherry 3 days after infection.
Supplementary Table 1 | Quantification of neuronal subtypes in the DRG at different developmental stages.
Supplementary Table 2 | Four most expressed genes in each neuronal subtype.
Supplementary Table 3 | 50 neuronal subtype-specific genes (FindMarker, Seurat).
Supplementary Table 4 | Differentially regulated genes at different time points after axotomy. (p adj < 0.05, FC > 2).
Supplementary Table 5 | 5 unique gene profile clusters with similar expression dynamics over time post-injury.
Footnotes
References
Avraham, O., Deng, P. Y., Jones, S., Kuruvilla, R., Semenkovich, C. F., Klyachko, A. K., et al. (2020). Satellite glial cells promote regenerative growth in sensory neurons. Nat. Commun. 11:4891.
Avraham, O., Feng, R., Ewan, E. E., Rustenhoven, J., Zhao, G., and Cavalli, V. (2021). Profiling sensory neuron microenvironment after peripheral and central axon injury reveals key pathways for neural repair. Elife 10:e68457. doi: 10.7554/eLife.68457
Beagan, J. A., Duong, M. T., Titus, K. R., Zhou, L., Cao, Z., Ma, J., et al. (2017). YY1 and CTCF orchestrate a 3D chromatin looping switch during early neural lineage commitment. Genome Res. 27, 1139–1152. doi: 10.1101/gr.215160.116
Belin, S., Nawabi, H., Wang, C., Tang, S., Latremoliere, A., Warren, P., et al. (2015). Injury-induced decline of intrinsic regenerative ability revealed by quantitative proteomics. Neuron 86, 1000–1014. doi: 10.1016/j.neuron.2015.03.060
Blackmore, M. G. (2012). Molecular control of axon growth: Insights from comparative gene profiling and high-throughput screening. Int. Rev. Neurobiol. 105, 39–70. doi: 10.1016/B978-0-12-398309-1.00004-4
Bloom, A. J., Mao, X., Strickland, A., Sasaki, Y., Milbrandt, J., and DiAntonio, A. (2022). Constitutively active SARM1 variants that induce neuropathy are enriched in ALS patients. Mol. Neurodegener. 17:1. doi: 10.1186/s13024-021-00511-x
Broude, E., McAtee, M., Kelley, M. S., and Bregman, B. S. (1997). c-Jun expression in adult rat dorsal root ganglion neurons: Differential response after central or peripheral axotomy. Exp. Neurol. 148, 367–377.
Carlin, D., Halevi, A. E., Ewan, E. E., Moore, A. M., and Cavalli, V. (2019). Nociceptor deletion of Tsc2 enhances axon regeneration by inducing a conditioning injury response in dorsal root ganglia. eNeuro 6, ENEURO.168–ENEURO.219. doi: 10.1523/ENEURO.0168-19.2019
Chandran, V., Coppola, G., Nawabi, H., Omura, T., Versano, R., Huebner, E. A., et al. (2016). A systems-level analysis of the peripheral nerve intrinsic axonal growth program. Neuron 89, 956–970. doi: 10.1016/j.neuron.2016.01.034
Cho, Y., and Cavalli, V. (2012). HDAC5 is a novel injury-regulated tubulin deacetylase controlling axon regeneration. EMBO J. 31, 3063–3078. doi: 10.1038/emboj.2012.160
Cho, Y., Shin, J. E., Ewan, E. E., Oh, Y. M., Pita-Thomas, W., and Cavalli, V. (2015). Activating injury-responsive genes with hypoxia enhances axon regeneration through neuronal HIF-1alpha. Neuron 88, 720–734. doi: 10.1016/j.neuron.2015.09.050
Cho, Y., Sloutsky, R., Naegle, K. M., and Cavalli, V. (2013). Injury-induced HDAC5 nuclear export is essential for axon regeneration. Cell 155, 894–908.
Di Giovanni, S., Knights, C. D., Rao, M., Yakovlev, A., Beers, J., Catania, J., et al. (2006). The tumor suppressor protein p53 is required for neurite outgrowth and axon regeneration. EMBO J. 25, 4084–4096.
Donohoe, M. E., Zhang, L. F., Xu, N., Shi, Y., and Lee, J. T. (2007). Identification of a Ctcf cofactor, Yy1, for the X chromosome binary switch. Mol. Cell. 25, 43–56. doi: 10.1016/j.molcel.2006.11.017
Ewan, E. E., Avraham, O., Carlin, D., Goncalves, T. M., Zhao, G., and Cavalli, V. (2021). Ascending dorsal column sensory neurons respond to spinal cord injury and downregulate genes related to lipid metabolism. Sci. Rep. 11:374. doi: 10.1038/s41598-020-79624-0
Fagoe, N. D., Attwell, C. L., Kouwenhoven, D., Verhaagen, J., and Mason, M. R. (2015). Overexpression of ATF3 or the combination of ATF3, c-Jun, STAT3 and Smad1 promotes regeneration of the central axon branch of sensory neurons but without synergistic effects. Hum. Mol. Genet. 24, 6788–6800.
Fagoe, N. D., van Heest, J., and Verhaagen, J. (2014). Spinal cord injury and the neuron-intrinsic regeneration-associated gene program. Neuromol. Med. 16, 799–813.
Fawcett, J. W., and Verhaagen, J. (2018). Intrinsic determinants of axon regeneration. Dev. Neurobiol. 78, 890–897.
Finelli, M. J., Wong, J. K., and Zou, H. (2013). Epigenetic regulation of sensory axon regeneration after spinal cord injury. J. Neurosci. 33, 19664–19676. doi: 10.1523/JNEUROSCI.0589-13.2013
Heinz, S., Benner, C., Spann, N., Bertolino, E., Lin, Y. C., Laslo, P., et al. (2010). Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell. 38, 576–589. doi: 10.1016/j.molcel.2010.05.004
Hervera, A., Zhou, L., Palmisano, I., McLachlan, E., Kong, G., Hutson, T. H., et al. (2019). PP4-dependent HDAC3 dephosphorylation discriminates between axonal regeneration and regenerative failure. EMBO J. 38, e101032. doi: 10.15252/embj.2018101032
Huebner, E. A., Budel, S., Jiang, Z., Omura, T., Ho, T. S., Barrett, L., et al. (2018). Diltiazem promotes regenerative axon growth. Mol. Neurobiol. 56, 3948–3957. doi: 10.1007/s12035-018-1349-5
Kang, K., Chung, J. H., and Kim, J. (2009). Evolutionary conserved motif finder (ECMFinder) for genome-wide identification of clustered YY1- and CTCF-binding sites. Nucleic Acids Res. 37, 2003–2013. doi: 10.1093/nar/gkp077
Karney-Grobe, S., Russo, A., Frey, E., Milbrandt, J., and DiAntonio, A. (2018). HSP90 is a chaperone for DLK and is required for axon injury signaling. Proc. Natl. Acad. Sci. U.S.A. 115, E9899–E9908. doi: 10.1073/pnas.1805351115
Kwon, M. J., Kim, J., Shin, H., Jeong, S. R., Kang, Y. M., Choi, J. Y., et al. (2013). Contribution of macrophages to enhanced regenerative capacity of dorsal root ganglia sensory neurons by conditioning injury. J. Neurosci. 33, 15095–15108.
Lesslich, H. M., Klapal, L., Wilke, J., Haak, A., and Dietzel, I. D. (2022). Adjusting the neuron to astrocyte ratio with cytostatics in hippocampal cell cultures from postnatal rats: A comparison of cytarabino furanoside (AraC) and 5-fluoro-2’-deoxyuridine (FUdR). PLoS One 17:e0265084. doi: 10.1371/journal.pone.0265084
Lezana, J. P., Dagan, S. Y., Robinson, A., Goldstein, R. S., Fainzilber, M., Bronfman, F. C., et al. (2016). Axonal PPARgamma promotes neuronal regeneration after injury. Dev. Neurobiol. 76, 688–701.
Lisi, V., Singh, B., Giroux, M., Guzman, E., Painter, M. W., Cheng, Y. C., et al. (2017). Enhanced neuronal regeneration in the CAST/Ei mouse strain is linked to expression of differentiation markers after injury. Cell Rep. 20, 1136–1147. doi: 10.1016/j.celrep.2017.07.010
Mahar, M., and Cavalli, V. (2018). Intrinsic mechanisms of neuronal axon regeneration. Nat. Rev. Neurosci. 19, 323–337.
Michaelevski, I., Segal-Ruder, Y., Rozenbaum, M., Medzihradszky, K. F., Shalem, O., Coppola, G., et al. (2010). Signaling to transcription networks in the neuronal retrograde injury response. Sci. Signal. 3:ra53.
Miller, B. R., Press, C., Daniels, R. W., Sasaki, Y., Milbrandt, J., and DiAntonio, A. (2009). A dual leucine kinase-dependent axon self-destruction program promotes Wallerian degeneration. Nat. Neurosci. 12, 387–389. doi: 10.1038/nn.2290
Niemi, J. P., Defrancesco-Lisowitz, A., Roldan-Hernandez, L., Lindborg, J. A., Mandell, D., and Zigmond, R. E. (2013). A critical role for macrophages near axotomized neuronal cell bodies in stimulating nerve regeneration. J. Neurosci. 33, 16236–16248. doi: 10.1523/JNEUROSCI.3319-12.2013
Oh, Y. M., Mahar, M., Ewan, E. E., Leahy, K. M., Zhao, G., and Cavalli, V. (2018). Epigenetic regulator UHRF1 inactivates REST and growth suppressor gene expression via DNA methylation to promote axon regeneration. Proc. Natl. Acad. Sci. U.S.A. 115, E12417–E12426. doi: 10.1073/pnas.1812518115
Omura, T., Omura, K., Tedeschi, A., Riva, P., Painter, M. W., Rojas, L., et al. (2015). Robust axonal regeneration occurs in the injured CAST/Ei Mouse CNS. Neuron 86, 1215–1227.
Palmisano, I., Danzi, M. C., Hutson, T. H., Zhou, L., McLachlan, E., Serger, E., et al. (2019). Epigenomic signatures underpin the axonal regenerative ability of dorsal root ganglia sensory neurons. Nat. Neurosci. 22, 1913–1924. doi: 10.1038/s41593-019-0490-4
Pita-Thomas, W., Goncalves, T. M., Kumar, A., Zhao, G., and Cavalli, V. (2021). Genome-wide chromatin accessibility analyses provide a map for enhancing optic nerve regeneration. Sci. Rep. 11:14924. doi: 10.1038/s41598-021-94341-y
Prasad, T., and Weiner, J. A. (2011). Direct and indirect regulation of spinal cord ia afferent terminal formation by the gamma-protocadherins. Front. Mol. Neurosci. 4:54. doi: 10.3389/fnmol.2011.00054
Puttagunta, R., Tedeschi, A., Soria, M. G., Hervera, A., Lindner, R., Rathore, K. I., et al. (2014). PCAF-dependent epigenetic changes promote axonal regeneration in the central nervous system. Nat. Commun. 5:3527. doi: 10.1038/ncomms4527
Renthal, W., Tochitsky, I., Yang, L., Cheng, Y. C., Li, E., Kawaguchi, R., et al. (2020). Transcriptional reprogramming of distinct peripheral sensory neuron subtypes after axonal injury. Neuron 108, 128.e–144.e. doi: 10.1016/j.neuron.2020.07.026
Sasaki, Y., Engber, T. M., Hughes, R. O., Figley, M. D., Wu, T., Bosanac, T., et al. (2020). cADPR is a gene dosage-sensitive biomarker of SARM1 activity in healthy, compromised, and degenerating axons. Exp. Neurol. 329:113252. doi: 10.1016/j.expneurol.2020.113252
Sasaki, Y., Vohra, B. P., Lund, F. E., and Milbrandt, J. (2009). Nicotinamide mononucleotide adenylyl transferase-mediated axonal protection requires enzymatic activity but not increased levels of neuronal nicotinamide adenine dinucleotide. J. Neurosci. 29, 5525–5535. doi: 10.1523/JNEUROSCI.5469-08.2009
Seijffers, R., Allchorne, A. J., and Woolf, C. J. (2006). The transcription factor ATF-3 promotes neurite outgrowth. Mol. Cell Neurosci. 32, 143–154. doi: 10.1016/j.mcn.2006.03.005
Seijffers, R., Mills, C. D., and Woolf, C. J. (2007). ATF3 increases the intrinsic growth state of DRG neurons to enhance peripheral nerve regeneration. J. Neurosci. 27, 7911–7920.
Sekine, Y., Lin-Moore, A., Chenette, D. M., Wang, X., Jiang, Z., Cafferty, W. B., et al. (2018). Functional genome-wide screen identifies pathways restricting central nervous system axonal regeneration. Cell Rep. 23, 415–428.
Serger, E., Luengo-Gutierrez, L., Chadwick, J. S., Kong, G., Zhou, L., Crawford, G., et al. (2022). The gut metabolite indole-3 propionate promotes nerve regeneration and repair. Nature 607, 585–592. doi: 10.1038/s41586-022-04884-x
Sharma, N., Flaherty, K., Lezgiyeva, K., Wagner, D. E., Klein, A. M., and Ginty, D. D. (2020). The emergence of transcriptional identity in somatosensory neurons. Nature 577, 392–398.
Shin, J. E., Cho, Y., Beirowski, B., Milbrandt, J., Cavalli, V., and DiAntonio, A. (2012). Dual leucine zipper kinase is required for retrograde injury signaling and axonal regeneration. Neuron. 74, 1015–1022.
Shin, J. E., Ha, H., Kim, Y. K., Cho, Y., and DiAntonio, A. (2019). DLK regulates a distinctive transcriptional regeneration program after peripheral nerve injury. Neurobiol. Dis. 127, 178–192. doi: 10.1016/j.nbd.2019.02.001
Simpson, M. T., Venkatesh, I., Callif, B. L., Thiel, L. K., Coley, D. M., Winsor, K. N., et al. (2015). The tumor suppressor HHEX inhibits axon growth when prematurely expressed in developing central nervous system neurons. Mol. Cell. Neurosci. 68, 272–283. doi: 10.1016/j.mcn.2015.08.008
Smith, D. S., and Skene, J. H. (1997). A transcription-dependent switch controls competence of adult neurons for distinct modes of axon growth. J. Neurosci. 17, 646–658. doi: 10.1523/JNEUROSCI.17-02-00646.1997
Smith, R. P., Lerch-Haner, J. K., Pardinas, J. R., Buchser, W. J., Bixby, J. L., and Lemmon, V. P. (2011). Transcriptional profiling of intrinsic PNS factors in the postnatal mouse. Mol. Cell Neurosci. 46, 32–44. doi: 10.1016/j.mcn.2010.07.015
Stam, F. J., MacGillavry, H. D., Armstrong, N. J., de Gunst, M. C., Zhang, Y., van Kesteren, R. E., et al. (2007). Identification of candidate transcriptional modulators involved in successful regeneration after nerve injury. Eur. J. Neurosci. 25, 3629–3637. doi: 10.1111/j.1460-9568.2007.05597.x
Stern, S., Haverkamp, S., Sinske, D., Tedeschi, A., Naumann, U., Di Giovanni, S., et al. (2013). The transcription factor serum response factor stimulates axon regeneration through cytoplasmic localization and cofilin interaction. J. Neurosci. 33, 18836–18848. doi: 10.1523/JNEUROSCI.3029-13.2013
Tedeschi, A., Dupraz, S., Laskowski, C. J., Xue, J., Ulas, T., Beyer, M., et al. (2016). The calcium channel subunit alpha2delta2 suppresses axon regeneration in the adult CNS. Neuron 92, 419–434. doi: 10.1016/j.neuron.2016.09.026
Tsujino, H., Kondo, E., Fukuoka, T., Dai, Y., Tokunaga, A., Miki, K., et al. (2000). Activating transcription factor 3 (ATF3) induction by axotomy in sensory and motoneurons: A novel neuronal marker of nerve injury. Mol. Cell Neurosci. 15, 170–182. doi: 10.1006/mcne.1999.0814
Venkatesh, I., and Blackmore, M. G. (2017). Selecting optimal combinations of transcription factors to promote axon regeneration: Why mechanisms matter. Neurosci. Lett. 652, 64–73. doi: 10.1016/j.neulet.2016.12.032
Venkatesh, I., Mehra, V., Wang, Z., Simpson, M. T., Eastwood, E., Chakraborty, A., et al. (2021). Co-occupancy identifies transcription factor co-operation for axon growth. Nat Commun. 12:2555. doi: 10.1038/s41467-021-22828-3
Venkatesh, I., Simpson, M. T., Coley, D. M., and Blackmore, M. G. (2016). Epigenetic profiling reveals a developmental decrease in promoter accessibility during cortical maturation in vivo. Neuroepigenetics 8, 19–26. doi: 10.1016/j.nepig.2016.10.002
Wang, Z., Mehra, V., Simpson, M. T., Maunze, B., Chakraborty, A., Holan, L., et al. (2018). KLF6 and STAT3 co-occupy regulatory DNA and functionally synergize to promote axon growth in CNS neurons. Sci. Rep. 8:12565. doi: 10.1038/s41598-018-31101-5
Weintraub, A. S., Li, C. H., Zamudio, A. V., Sigova, A. A., Hannett, N. M., Day, D. S., et al. (2017). YY1 Is a structural regulator of enhancer-promoter loops. Cell 171:e1528.
Weng, Y. L., Joseph, J., An, R., Song, H., and Ming, G. L. (2016). Epigenetic regulation of axonal regenerative capacity. Epigenomics 8, 1429–1442.
Williams, R. R., Venkatesh, I., Pearse, D. D., Udvadia, A. J., and Bunge, M. B. (2015). MASH1/Ascl1a leads to GAP43 expression and axon regeneration in the adult CNS. PLoS One. 10:e0118918. doi: 10.1371/journal.pone.0118918
Zhao, G., Schriefer, L. A., and Stormo, G. D. (2007). Identification of muscle-specific regulatory modules in Caenorhabditis elegans. Genome Res. 17, 348–357. doi: 10.1101/gr.5989907
Zhou, L., Kong, G., Palmisano, I., Cencioni, M. T., Danzi, M., De Virgiliis, F., et al. (2022). Reversible CD8 T cell-neuron cross-talk causes aging-dependent neuronal regenerative decline. Science 376:eabd5926. doi: 10.1126/science.abd5926
Keywords: axon regeneration, sensory neurons, transcription factors, bioinformatics analyses, CTCF, YY1, E2F2, dorsal root ganglia
Citation: Avraham O, Le J, Leahy K, Li T, Zhao G and Cavalli V (2022) Analysis of neuronal injury transcriptional response identifies CTCF and YY1 as co-operating factors regulating axon regeneration. Front. Mol. Neurosci. 15:967472. doi: 10.3389/fnmol.2022.967472
Received: 12 June 2022; Accepted: 28 July 2022;
Published: 23 August 2022.
Edited by:
Stephane Belin, Institut National de la Santé et de la Recherche Médicale (INSERM), FranceReviewed by:
Yongcheol Cho, Daegu Gyeongbuk Institute of Science and Technology (DGIST), South KoreaMelissa R. Andrews, University of Southampton, United Kingdom
Simone Di Giovanni, Imperial College London, United Kingdom
Copyright © 2022 Avraham, Le, Leahy, Li, Zhao and Cavalli. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Valeria Cavalli, Y2F2YWxsaUB3dXN0bC5lZHU=
 Oshri Avraham
Oshri Avraham Jimmy Le
Jimmy Le Kathleen Leahy
Kathleen Leahy Tiandao Li2,3
Tiandao Li2,3 Valeria Cavalli
Valeria Cavalli