- 1Division of Environmental Medicine, Department of Medicine, University of Louisville, KY, United States
- 2Division of Molecular and Cellular Pathology, Department of Pathology, University of Alabama at Birmingham, Birmingham, AL, United States
- 3Division of Anatomic Pathology, Department of Pathology, University of Alabama at Birmingham, Birmingham, AL, United States
- 4Division of Cardiovascular Disease, Department of Medicine, University of Alabama at Birmingham, Birmingham, AL, United States
Transmission electron microscopy (TEM) has long been an important technique, capable of high degree resolution and visualization of subcellular structures and organization. Over the last 20 years, TEM has gained popularity in the cardiovascular field to visualize changes at the nanometer scale in cardiac ultrastructure during cardiovascular development, aging, and a broad range of pathologies. Recently, the cardiovascular TEM enabled the studying of several signaling processes impacting mitochondrial function, such as mitochondrial fission/fusion, autophagy, mitophagy, lysosomal degradation, and lipophagy. The goals of this review are to provide an overview of the current usage of TEM to study cardiac ultrastructural changes; to understand how TEM aided the visualization of mitochondria, autophagy, and mitophagy under normal and cardiovascular disease conditions; and to discuss the overall advantages and disadvantages of TEM and potential future capabilities and advancements in the field.
Introduction
Since the development of the first electron microscope in the 1930's, transmission electron microscopy (TEM) has been an essential technique for the visualization of cellular ultrastructure. The timeline of the key events in the development and evolution of TEM has been described in an previous study (Lukasz Mielanczyk et al., 2015). TEM has steadily gained popularity in the cardiovascular research field over the last 20 years, being utilized in over 2,000 peer-reviewed cardiovascular research publications (Figure 1), to examine cardiac ultracellular organization at the nanometer scale. In the cardiovascular field, TEM is often used to assist in studies investigating: (1) cardiovascular development and aging; (2) cardiovascular disease processes, such as heart failure, myocardial infarction/ischemia (MI), and diabetic cardiomyopathy; (3) phenotyping of cardiovascular transgenic models; and (4) examination of complex signaling processes, many of which impact mitochondrial function and dynamics, such as apoptosis, autophagy, and mitophagy. In this review, we discuss the application of TEM as a tool for viewing cardiac ultrastructure and how this has been leveraged in cardiovascular studies to visualize complex signaling processes, with a special focus on the mitochondria. Although it is important to appreciate that, in addition to TEM, scanning electron microscopy (SEM) has also been used in several cardiovascular studies, its primary application is to provide information on cell surface and composition, rather than cellular ultrastructure; therefore, SEM will not be discussed in this review (Klang et al., 2012; Lukasz Mielanczyk et al., 2015).
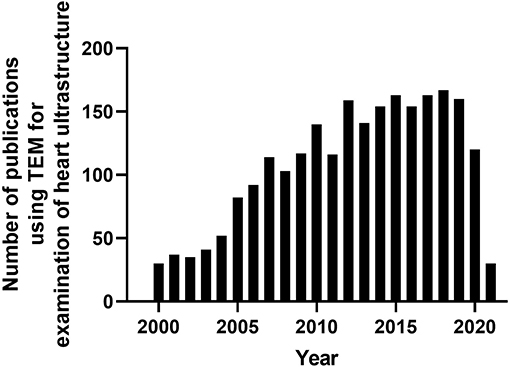
Figure 1. Timeline of peer-reviewed cardiovascular publications utilizing transmission electron microscopy to examine cardiac ultrastructure. Data taken from Pubmed analytics using a search of “Transmission electron microscopy” and “heart.” Data presented were taken from 2000-present.
Cardiac Structures Visualized With TEM
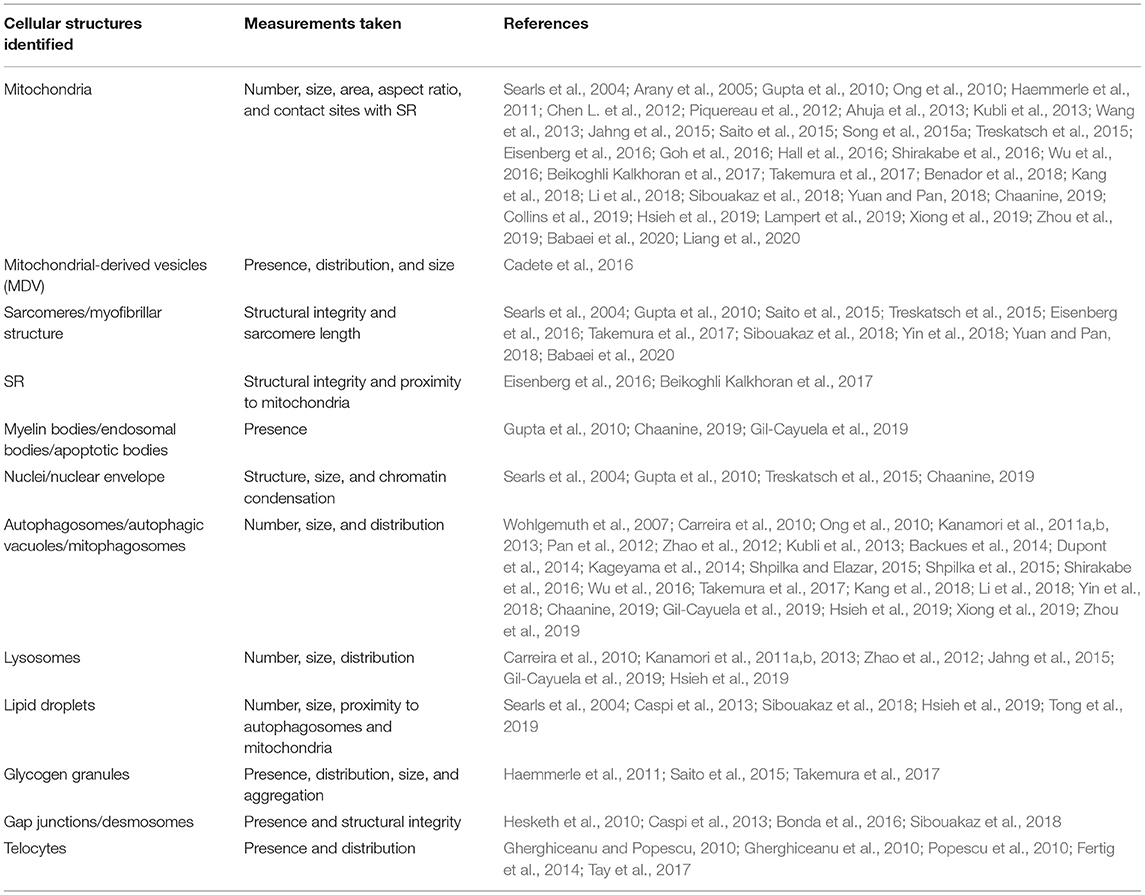
Transmission electron microscopy has been used to provide high magnification and high-resolution images of sarcomeric structures (i.e., H-zone, A-band, M-line, and Z-line of the sarcomere, myofibrils, etc.), and at the subcellular level, numerous structures have been characterized, such as endoplasmic/sarcoplasmic reticulum (ER/SR), nuclei and chromatin structure/organization, lipid droplets (LDs), glycogen deposits, lysosomes, autophagolysosomes, gap junctions, caveolae, and extracellular matrix (ECM), and mitochondrial-associated structures, which include cristae, intact and disrupted mitochondria, mitochondrial-derived vesicles (MDV) (Cadete et al., 2016), mitochondrial-associated membranes (MAMs), and mitophagosomes. TEM has also been used to reveal or confirm the presence of new cell populations in the heart, such as telocytes (Popescu et al., 2010; Fertig et al., 2014; Tay et al., 2017). Figure 2 shows examples of cardiac structures visualized using TEM. From the images, quantification can be performed (Glancy, 2020) for the assessment of number, length, aspect ratio, and areas of mitochondria, lysosomes, and autophagosomes, in addition to qualitatively scoring cellular integrity (e.g., cristae of mitochondria, cristae junctions, and matrix remodeling).
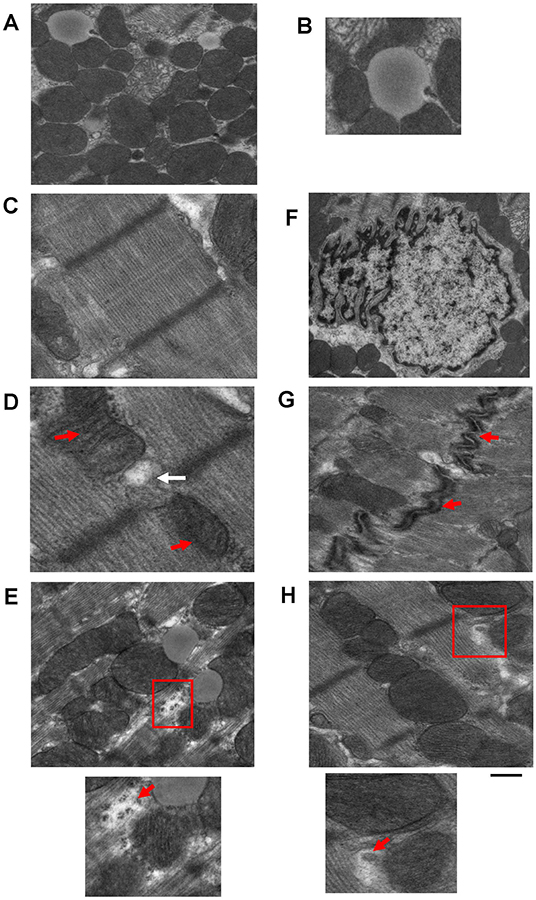

Figure 2. Example images of cardiac cellular structures examined using TEM studies from wild-type mouse hearts. (A) TEM image showing mitochondria surrounding a lipid droplet; (B) Magnification of Lipid droplet in (A); (C) TEM image of cardiac sarcomeric structure; (D) TEM image of SR-mitochondria connections. Red arrows depict mitochondria and white arrow depicts SR; (E) TEM image showing glycogen deposits (highlighted in red box); (F) TEM image of a nucleus; (G) TEM image of gap junctions (gap junctions depicted by red arrows); and (H) TEM image showing a mitochondrial derived vesicle budding from larger mitochondria (highlighted by red box). Images are unpublished images from Zhang lab studies. Scale bar depicts 500 nm.
TEM Sample Preparation
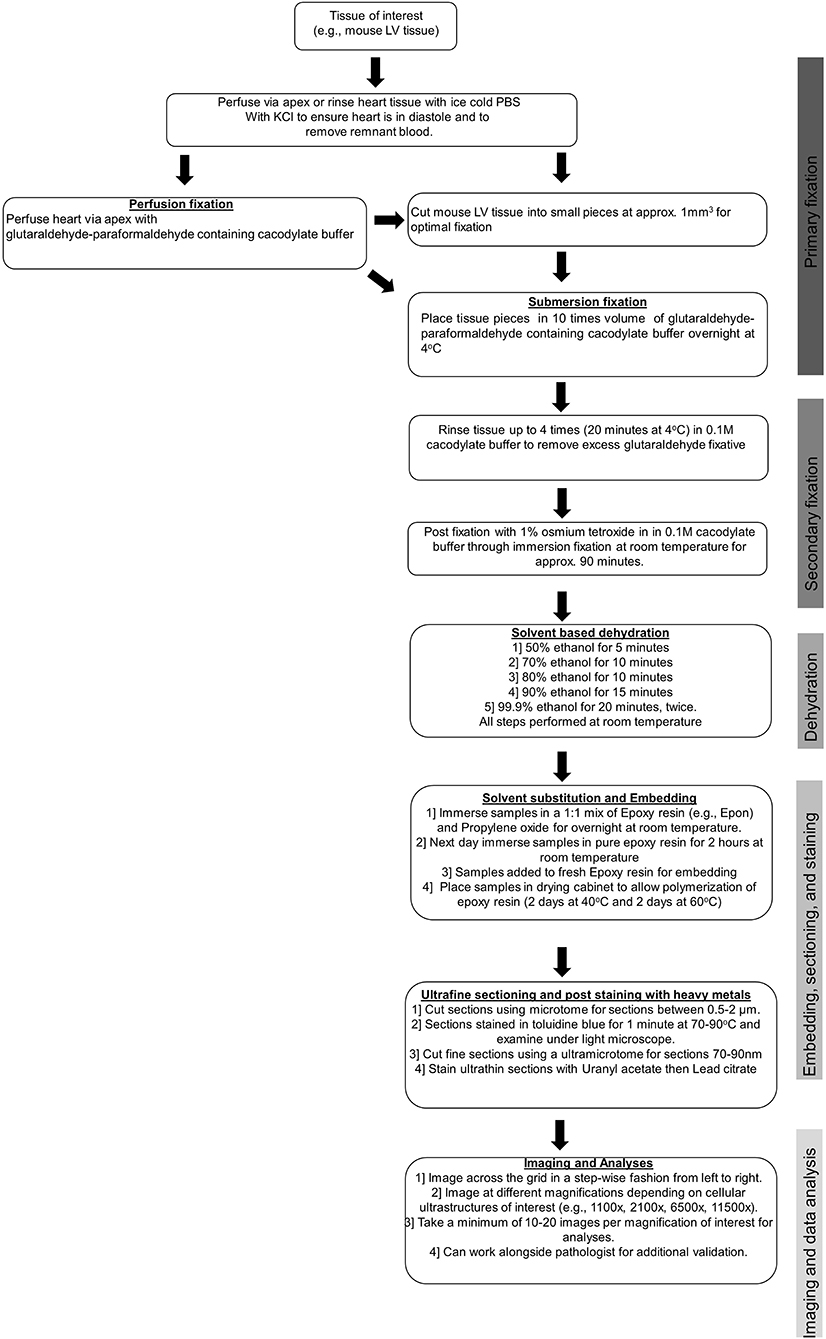
To obtain clear, reliable images suitable for accurate measurements, particular care must be taken at the sample preparation stage to avoid the production of artifacts. Typically, freshly isolated cardiac samples, usually left ventricular (LV) samples, are extracted and then either rinsed quickly in cold phosphate-buffered saline (PBS) or retrogradely perfused with cold PBS containing potassium chloride to arrest the heart in diastole and to remove any remnant red blood cells. Adequate removal of red blood cells prior to fixation is essential to improve fixation and subsequent image quality. Following the removal of red blood cells, tissues undergo either submersion fixation or perfusion fixation in a glutaraldehyde–paraformaldehyde containing cacodylate buffer. Most commonly, the submersion fixation is preferred for LV samples as only a small fraction of the heart is needed for TEM analysis, which allows the remaining LV tissues to be used for other downstream assays that are not compatible or optimal with the fixed tissues. For some tissues, such as those collected from humans during left ventricular assist device (LVAD) surgeries, perfusion is not possible. Prior to the submersion fixation, the heart must be cut into smaller pieces to allow for optimal fixation, and for the mouse heart, we immersed LV tissue pieces of ~1 mm3 in size in 1 ml of glutaraldehyde–paraformaldehyde containing cacodylate buffer. After this, a post-fixation step with osmium tetroxide and uranyl acetate containing cacodylate buffer was performed to preserve lipid structures. This post-fixation stage, although important for creating the appropriate contrast required for high-quality TEM, if prolonged (i.e., longer than 90 min), can result in protein and lipid damage and distortion of the cellular ultrastructure. Following the post-fixation, samples underwent a solvent-based dehydration step followed by embedding in epoxy resin (e.g., Epon), prior to ultra-fine sectioning and post-staining with heavy metal stains. The ultra-fine sectioning is performed using an ultramicrotome, where sections of 1 μm are cut and stained with toluidine blue. The samples are then cut further using a diamond knife so that sections of 50–70 nm are achieved, and these are placed onto a metal grid. The heavy metal staining is performed with lead citrate, which interacts with the proteins and glycogens in the tissue sample and is required to enhance image contrast.
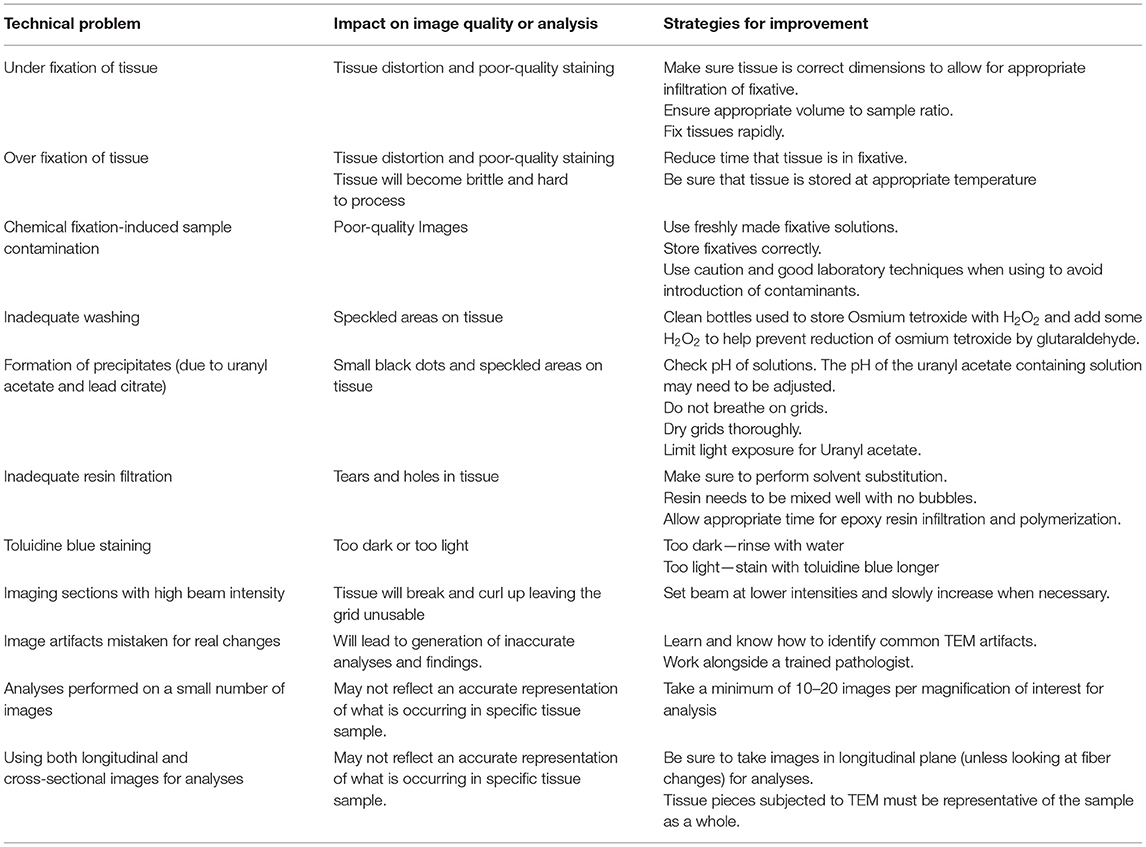
Sample preparation is the single most important stage in TEM. Sample artifacts have been reported to arise from each stage in the sample preparation stage (i.e., from fixation to post-staining), which reduce resolution and can produce spurious findings. These sample artifacts include chemical fixation-induced sample contamination; small darkened speckled areas from inadequate washing; formation of precipitates by the heavy metals, uranyl acetate, and lead citrate; shrinkage artifacts introduced through rapid dehydration; tears and holes formed due to inadequate resin infiltration; sectioning of tissue resulting in compression, scratches, and tears; and high TEM beam setting resulting in tissue breakage. Some common technical problems experienced during TEM and strategies for avoiding these issues are outlined in Table 1. Therefore, with TEM, there is a constant need and requirement to improve sample preparation and the staining methods to improve image quality. Furthermore, there is a great need to adapt TEM to create innovative new methods for sample examination and quantification, which includes the use and development of immunogold labeling techniques and cryo-TEM, which have proven to be essential tools in the visualization of physiological processes and various cardiovascular pathologies (see below). Key steps of sample preparation are illustrated in Figure 3.
Analysis of TEM Images
Analysis of TEM images is an important step in understanding and assessing the ultrastructural changes occurring in a given tissue sample. Identification of image artifacts is a key step in the analysis of TEM images, as, to the untrained eye, these may appear to be physiological changes. The recruitment of a trained pathologist to help assess ultrastructural changes is recommended. In addition, for the quantitative analysis, it is important that at least 10–20 images are taken from each sample at each magnification and in the same plane for an accurate representation. Similarly, the tissue pieces selected for TEM must be representative of the sample as a whole to ensure accurate analysis of findings. As mentioned in Table 2, many cardiac ultrastructures have been identified, examined, and measured using TEM. For these cellular structures, descriptive analyses have often been performed to describe visual differences between samples. In addition, across the field there does not appear to be consistent reporting of TEM findings and analyses. Often, the quantitative information on cell number and size is collected during TEM analyses using software with specific cell counting and cell size capabilities, such as Image J software (National Institutes of Health, Bethesda, MD, USA). For these analyses, it is important, in terms of quantification, that cell borders are intact and completely visible in order to be included in the analyses, and care must be taken to keep measurement criteria standard across all samples in a study. In addition, it is important to normalize data obtained back to the known dimensions/area of the field assessed. Although measurements of cell size and aspect ratio (length:width) can be obtained, one significant disadvantage of TEM in terms of measuring size changes is that it does not provide a 3D image of various structures, such as mitochondria, which are spherical/tubular in form, so only crude measurements of size are possible with traditional TEM, and additional electron tomography studies are required for the assessment of 3D structures with TEM. The next section discusses the current usage and application of TEM to visualize changes in cardiac ultrastructure.
Use of TEM to Study Cardiovascular Disease Processes
Cardiovascular diseases, such as hypertrophy, hypertension, myocardial infarction (MI), ischemia/reperfusion (I/R) injury, heart failure (HF), and metabolic-induced perturbations in cardiac function, such as those associated with diabetic cardiomyopathy, are all characteristically associated with significant alterations in the cardiac ultrastructure. Broadly, these ultrastructural changes, most often visualized using TEM, include myofibril and sarcomeric disorganization; damaged gap junctions; changes in nuclear organization, such as chromatin aggregation; changes in mitochondrial shape, content, and distribution; swollen, cristae-damaged mitochondria; LD deposition; glycogen aggregation; changes in the SR-mitochondrial association and SR dilatation; and the presence of autophagosomes and lysosomes. Cardiac ultrastructures identified, examined, and measured using TEM are summarized in Table 2. This section discusses the current usage and application of TEM to visualize changes in the cardiac ultrastructure during these respective disease processes.
Non-ischemic and Non-metabolic Heart Disease
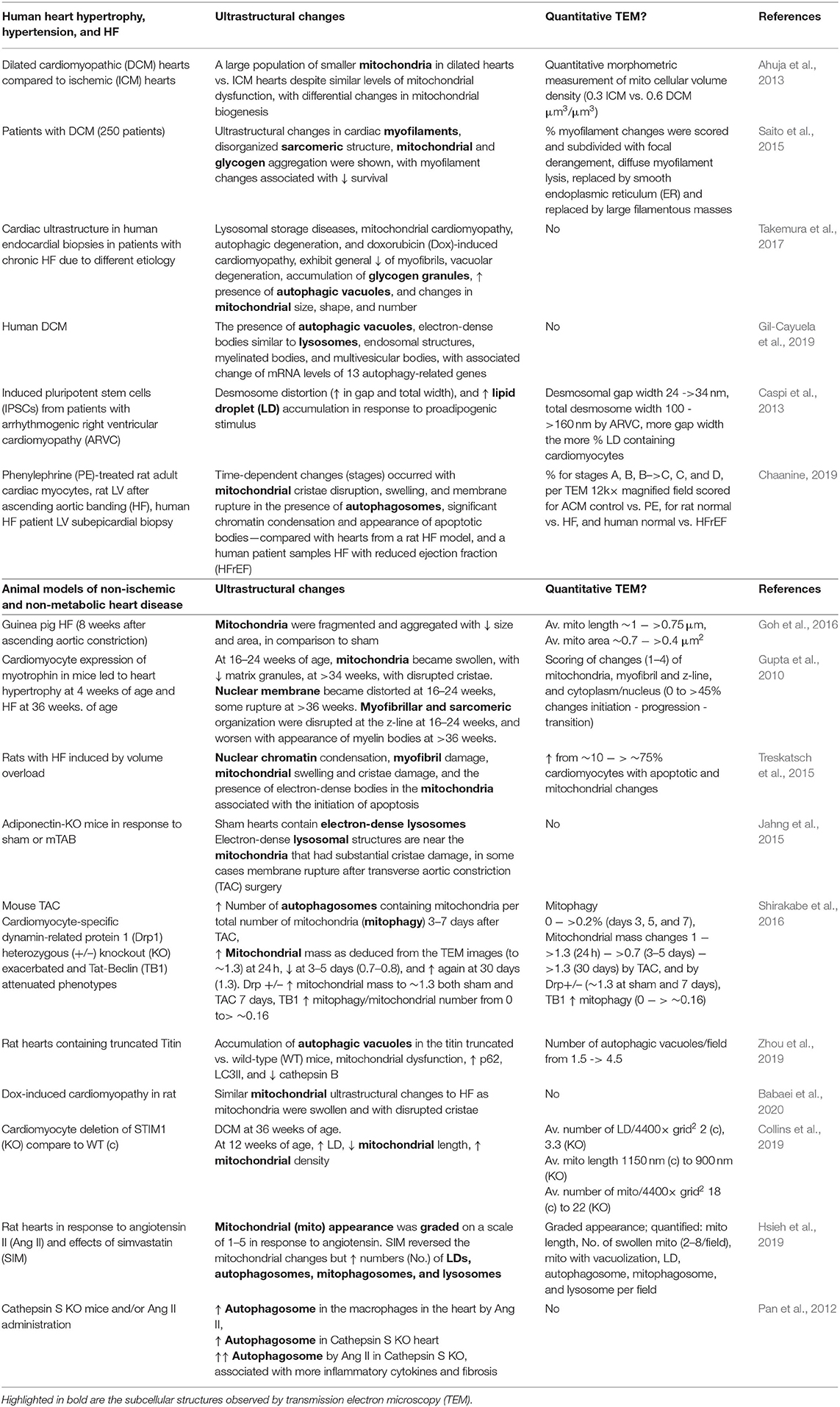
As mentioned above, TEM has been utilized to document ultrastructural changes in the heart in cases of pathological remodeling, such as hypertrophy, hypertension, cardiomyopathy, and ultimately progression to HF, in various models including human tissues, animal models, and in cultured cells (Table 3). Historically, many of the TEM studies performed in the failing heart focused significantly on sarcomeric and myofibrillar structure, density, and organization. For example, degeneration of cardiac sarcomeres has been observed in the hypertrophic cardiac tissue that was associated with significant thickening of z-bands and shortening of the sarcomere and with changes in the basement membrane and tubular structure (Maron et al., 1975a). In addition, the TEM analyses of cardiac tissues from patients with hypertrophy and valvular disease showed focal myofibril lysis, loss of thick myofilaments, and T-tubular changes, which were associated with expansion of the SR and mitochondria into these areas (Maron et al., 1975b). Furthermore, TEM of LV endomyocardial biopsies showed that myofibrillar density was lower in the hearts of patients with HFrEF than those with HFpEF and was associated with significant myofibrillar loss (van Heerebeek et al., 2006). End-stage dilated cardiomyopathic heart tissue obtained from patients undergoing heart transplant showed similar changes in myofilament and sarcomeric structures, which have been linked to changes in cytoskeletal proteins including desmin (Schaper et al., 1991; Mudhar et al., 2001).
Since mitochondrial dysfunction is a common feature of adverse remodeling and HF, it is not surprising that mitochondrial derangements are among the most common ultrastructural finding in TEM studies. For example, TEM revealed a large population of smaller mitochondria in dilated cardiomyopathic hearts compared with ischemic cardiomyopathic (ICM) hearts, and this was associated with differential changes in mitochondrial biogenesis (Ahuja et al., 2013). In this study, a morphometric measurement was performed to demonstrate that the mitochondrial cellular volume density is 0.3 μm3/μm3 in ICM vs. ~0.6 in the dilated cardiomyopathic heart (Ahuja et al., 2013). Furthermore, in a study involving 250 patients, ultrastructural changes in cardiac myofilaments, disorganized sarcomeric structure, and mitochondrial and glycogen aggregation were shown with TEM in patients with dilated cardiomyopathy (DCM), with myofilament changes (% changes scored) associated with reduced survival (Saito et al., 2015). A more recent study compared cardiac ultrastructure in endocardial biopsies in patients with chronic HF, lysosomal storage diseases, mitochondrial cardiomyopathy, and doxorubicin (Dox)-induced cardiomyopathy. Dependent on disease type and stage, loss of myofibrils, vacuolar degeneration, accumulation of glycogen granules, the increased presence of autophagic vacuoles and changes in mitochondrial size, shape, and number have been observed. The importance of TEM as a tool in differential cardiac disease diagnoses was highlighted by the recommendation that tissues from every endocardial biopsy should be saved for the cardiac TEM analysis (Takemura et al., 2017). The presence of autophagic structures in the heart tissue has also been observed using TEM in cases of human DCM (Te Rijdt et al., 2016; Gil-Cayuela et al., 2019). These included autophagic vacuoles, electron-dense bodies similar to lysosomes, endosomal structures, myelinated bodies, aggresomes, and multivesicular bodies (Gil-Cayuela et al., 2019). In this same study, RNA-seq analysis showed that, out of 175 autophagy-related genes detected, these ultrastructural changes were associated with five increased and eight decreased transcripts (Gil-Cayuela et al., 2019). Analysis of autopsied and explanted hearts from patients with DCM and with phospholamban mutations showed the presence of p62 and LC3 containing aggresomes at TEM, suggestive of changes in autophagy (Te Rijdt et al., 2016).
In addition, TEM has been used to examine changes in cell culture models of cardiomyopathy. For example, induced pluripotent stem cells (IPSCs) from patients with arrhythmogenic right ventricular cardiomyopathy (ARVC) were shown to have distortion of the cell-to-cell adhesion cell type, desmosome (desmosomal gap width and total desmosome width scored), and extensive LD accumulation (scored as % LD containing cardiomyocytes), which were further augmented in response to proadipogenic stimulus (Caspi et al., 2013). In a study comparing phenylephrine (PE)-treated adult cardiomyocytes, hearts from a rat transverse aortic constriction (TAC) model and from human patients with systolic HF, it was reported that similar time-dependent changes (scored as stages A–D) occurred with mitochondrial cristae disruption, swelling, and membrane rupture in the presence of autophagosomes, in the condensation of significant chromatins, and in the appearance of apoptotic bodies (Chaanine, 2019).
Dilated cardiomyopathy is commonly associated with truncations in the large structural protein, titin. TEM studies in rat hearts containing truncated titin showed the presence of damaged mitochondria and the accumulation of autophagic vacuoles (with numbers scored per field) vs. wild-type (WT) hearts, suggestive of changes in autophagy. These changes were associated with hyperacetylation of mitochondrial proteins, increased p62 and LC3II, and decreased cathepsin B (Zhou et al., 2019). Dox-induced cardiomyopathy was shown to have similar mitochondrial ultrastructural changes to HF as mitochondria were swollen and exhibited disruption of cristae (Babaei et al., 2020). However, WT mice subjected to TAC or sham were shown to have a normal mitochondrial morphology and sarcomeric structure and organization. Since adiponectin plays a key role in DCM, adiponectin-KO mice were examined in the same study. Compared to WT, cardiac tissues in sham adiponectin-KO mice contained electron-dense lysosomes, and following TAC, these electron-dense lysosomes were located near the mitochondria that have substantial cristae damage and in some cases membrane rupture (Jahng et al., 2015). Collectively, these studies have used TEM to highlight significant structural changes in mitochondria and autophagic processes in DCM.
Many of the cardiac ultrastructural changes documented in human cardiac samples have also been examined and visualized in animal cardiac samples through the use of TEM. For example, in a guinea pig model of HF induced by pressure overload, mitochondria were fragmented and aggregated with decreased size and area, in comparison to mitochondria in sham hearts. Quantitative changes were reported for mitochondrial diameters and areas with TEM images (Goh et al., 2016). In an extensive TEM-based ultrastructural study of the transition from hypertrophy to HF in a myotrophin overexpression (OE) mouse model, mitochondria were found to become swollen and cristae were found to be disrupted. In addition, the nuclear membranes were distorted, and the myofibrillar and sarcomeric organization was disrupted at the z-line and was associated with the presence of myelin bodies, immediately prior to and during the progression to HF phase (Gupta et al., 2010). In this study, mitochondrial changes were scored as matrix electron lucent appearance, swelling, and concentric cristae; myofibril structure and z-line changes were scored as wavy, blurring, and no defined structure; and cytoplasm and nucleus were scored according to the shape change, nuclear membrane distortion, and rupture of cell membrane (Gupta et al., 2010). Rats with HF induced by volume overload were shown to exhibit changes, such as nuclear chromatin condensation, myofibril damage, mitochondrial swelling, and cristae damage, and the presence of electron-dense bodies in the mitochondria associated with the initiation of apoptosis (Treskatsch et al., 2015). In this study, the percent of cardiomyocytes with apoptotic and mitochondrial changes were scored (Treskatsch et al., 2015). In a study in C57BL/6J mice subjected to TAC, mitophagy was found to be upregulated as early as 3 days after pressure overload, and mitochondrial mass was deduced from the TEM images to be up at 24 h and down at 3–5 days after pressure overload and up again at 30 days (Shirakabe et al., 2016). The number of autophagosomes containing mitochondria per total number of mitochondria (mitophagy) was scored with the mitochondrial mass normalized to control (Shirakabe et al., 2016). In the same study, cardiomyocyte-specific heterozygous knockout (KO) of dynamin-related protein 1 (Drp1) resulted in a greater mitochondrial mass at 7 days and exacerbated HF phenotypes. Injection of Tat-Beclin 1, an inducer of autophagy, partially rescued the phenotypes (Shirakabe et al., 2016). In addition, cardiomyocyte-specific KO of STIM1, which is a key calcium signaling protein involved in cardiac hypertrophy, developed DCM at 36 weeks of age, which was associated with decreased mitochondrial length, an increase in mitochondrial density, and an increase in the number of LDs from 12 weeks of age (Collins et al., 2019).
Extensive TEM analyses were performed in hypertrophic rat hearts to investigate the potential cardiac benefits of the 3-hydroxy-3-methyl-glutaryl-coenzyme A (HMG-CoA) reductase inhibitor, simvastatin (SIM), in response to 14 days of angiotensin II (Ang II) osmotic pump infusion (Hsieh et al., 2019). In this study, the mitochondrial appearance was subjectively graded in control, Ang II, and Ang II + SIM on a scale of 1–5 with 1 referring to normal mitochondria, 2 referring to occasional swollen, 3 referring to major distortions, 4 referring to membranes/cristae dissociated into particulates, and 5 referring to mitochondria with damaged cristae, mitochondrial membrane damage, and mitochondrial vacuolization. In addition, the mitochondrial length (decreased with Ang II vs. control), the number of swollen mitochondria (increased in Ang II vs. control), mitochondrial vacuolization (increased in Ang II vs. control), LDs, autophagosomes, mitophagosomes, and lysosomes (increased in Ang II vs. control) were quantified. Interestingly, SIM reversed the Ang II-induced mitochondrial changes but increased the numbers of LDs, autophagosomes, mitophagosomes, and lysosomes in response to Ang II (Hsieh et al., 2019). Cathepsin S deficiency was shown to result in more fibrosis and more inflammatory cytokines in the heart in response to Ang II. These pathologies were also associated with more autophagic vacuoles in macrophages in the heart (Pan et al., 2012).
Collectively, these studies show that many significant ultrastructural changes occur with pressure overload and HF, which are associated with changes in mitochondrial morphology and accumulation of autophagic vacuoles. More than half of these studies performed morphometric or graded quantifications of ultrastructural changes in diseases or disease models. These observations are useful in the investigation of pathologies and effects of potential therapeutic interventions in both human patient specimen and animal models.
Ischemic Heart Diseases
Many of the ultrastructural changes occurring with non-ischemic heart diseases also occur in I/R and MI despite the differences in the underlying pathology (Table 4). Compared to HF tissues, the ultrastructural studies in ICM are relatively fewer, and most studies lack quantification of the changes. However, TEM has been used to examine changes in cardiomyocyte apoptosis in hearts with ICM (Abbate et al., 2007). In addition, human LV tissues subjected to ischemia showed damaged contractile proteins and cytoskeletal structures, which were associated with the disruption of several cytoskeletal proteins, including desmin, tubulin, and myomesin (Hein et al., 1995).
In addition, there have been some studies that have used TEM to examine cardiac tissues extracted during LVAD placement and during transplantation. For example, TEM has been used to examine cardiac tissue in patients with end-stage HF before and after LVAD placement and to examine soft tissue reaction to LVAD placement. In these studies, it has been shown that there are significant ultrastructural changes that occur in the hearts of these patients, which include basal membrane changes (Bruggink et al., 2007), changes in collagen fibers and remodeling (Bruggink et al., 2007), changes in myofilament structure, and changes in mitochondrial size and number (Ikeda et al., 2015). A recent study compared pre-LVAD and post-LVAD myocardial tissue samples from patients with DCM and ischemic heart disease. It was found that, although mitochondrial morphology was similar between samples from patients with ischemic heart disease and DCM, differences were observed in some mitochondrial proteins (de Weger et al., 2011). TEM has been used to examine aortic valve ultrastructure in patients with LVAD and non-LVAD during transplant, where it has been shown that there was an increase in the fiber size in patients with LVAD, which was linked to reduced compliance and changes in proteins associated with the valve activation (Stephens et al., 2018). Of note, many of these ultrastructural changes are not impacted by LVAD placement. For example, it has been shown that after LVAD unloading, the LV structure was not impacted and was not indicative of atrophic changes (Diakos et al., 2014).
Similarly, the cardiac ultrastructural changes have been observed in animal models of ischemic heart disease. In a rat model of MI, TEM showed that the myocardial fiber network was disrupted, mitochondria were smaller and rounder, and mitochondrial and autophagosome numbers were also increased. These ultrastructural changes were associated with increased LC3II/I ratio and decreased Parkin protein level, with decreased autophagic flux as assessed by chloroquine treatment (Wu et al., 2016). In a mouse model of MI, there was an increase in large vacuoles that resembled autophagosomes at the border zone and smaller autophagosomes at the remote zone of the infarct at 1-week post-MI (Kanamori et al., 2011a). By 3-week post-MI, more autophagosomes and lysosomes were present in the remote zone, and autophagosome size in the border zone was smaller (Kanamori et al., 2011a). The time-dependent accumulation of autophagosomes and electron-dense bodies, at both the infarct and border zones following MI from 30 min to 24 h, was shown in GFP-LC3 transgenic mice (Kanamori et al., 2011b). Immune-TEM with a cathepsin D antibody and enzyme cytochemistry for acid phosphatase were used to confirm that the electron-dense bodies were, in fact, lysosomes. The immunoblot analysis reported the increase of Cathepsin D from 30 min to 24 h. This study highlighted the combined use of TEM and organelle-specific labeling method in examination of cardiac ultrastructure. The accumulation of autophagosomes was also shown with increased GFP-LC3 puncta in both border and infarct areas at 30 min and in border but not infarct areas at 4–24 h. LC3II is increased at 4–24 h, and p62 increased at 24 h. Furthermore, the impact of starvation (24 h before MI) appeared to increase autophagosomes that encircle partially degraded intracellular materials and/or Bafilomycin A treatment (30 min, 0.3 mg/kg before MI, blocking autophagy and qualitatively appeared to increase electron-dense lysosomes) on both the infarct sizes (Kanamori et al., 2011b). TEM was also used to qualitatively demonstrate increased accumulation of autophagic vacuoles and lysosomes in post-MI hearts, which were further increased with resveratrol and were associated with a reduction in adverse remodeling (Kanamori et al., 2013). The effect of resveratrol on autophagic vacuoles was recapitulated qualitatively and attenuated by the adenosine monophosphate-activated protein kinase (AMPK) inhibitor compound C in neonatal cardiomyocytes in vitro (Kanamori et al., 2013). Collectively, these studies have highlighted the use of TEM to examine ultrastructural changes during ICM, and these studies, like those in non-ICM, have shown extensive changes in mitochondria and autophagic signaling.
Metabolic-Induced Cardiac Perturbations
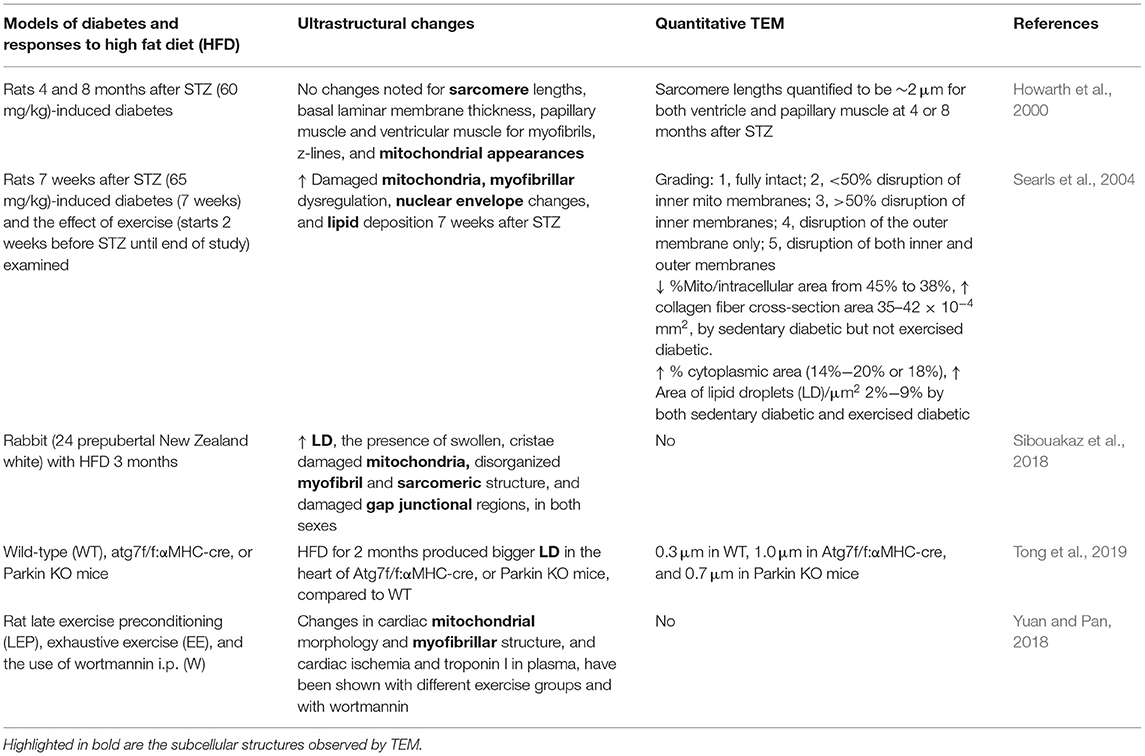
Diabetic cardiomyopathy, obesity, and utilization of high fat diet (HFD) have been shown to impact the structure and function of the myocardium, mitochondrial function, and mitochondrial driven processes (Bournat and Brown, 2010; Duncan, 2011; Miotto et al., 2018). Hearts from both diabetic and obese patients were the first to suggest changes in lipid content and lipotoxicity in hearts. The increased presence of LDs or cardiac steatosis has been observed through the use of TEM in diabetes (Sugawara et al., 2021). TEM analyses in hearts of type 2 diabetics exhibited the presence of fragmented mitochondria (Montaigne et al., 2014); however, the study also investigated parameters regarding mitochondrial function in the heart samples from patients with obesity, TEM analyses were not performed under these conditions, and the TEM analyses in this study were performed in right atrial tissue. Similar to human studies, the TEM analyses have documented the effect of diabetes and obesity on cardiac ultrastructure in animal models (Howarth et al., 2000; Searls et al., 2004; Boudina et al., 2007; Li et al., 2020). Earlier studies using a rat model of type 1 diabetes (STZ) documented no specific changes in sarcomere lengths, myofibrils, and mitochondrial appearance in hearts at 4 and 8 months after following diabetes induction (Howarth et al., 2000). A later study examined hearts from rats with STZ-induced diabetes and observed damaged mitochondria (i.e., disrupted cristae), myofibrillar disarray, nuclear envelope changes, and increased lipid deposition (Searls et al., 2004). The changes in mitochondrial area in the cell and collagen fiber cross-section area were attenuated by exercise, whereas the area of cytoplasm and LDs were not affected (Searls et al., 2004). In addition, several studies in diabetic mice and rats showed increased cardiac numbers of LDs, increased mitochondrial size, and changes in glycogen levels (Zhou et al., 2000; Christoffersen et al., 2003; Boudina et al., 2007; Li et al., 2020). In contrast, studies in obese Zucker rats showed that there were no changes in LDs or changes in mitochondrial mass and density; despite the lack of change in LDs, cardiac TAG levels were increased (Holloway et al., 2011). However, it is important to note that the ultrastructural changes found in diabetic hearts in animal studies are dependent on the animal model used, duration of diabetes, or the model used to generate diabetes, and some dynamic changes may not be effectively captured in these studies (Table 5).
High fat diet also induced significant changes in rabbit cardiac ultrastructure consisting of the accumulation of LDs, the presence of swollen, cristae damaged mitochondria, disorganized myofibril and sarcomeric structure, and damaged gap junctional regions (Sibouakaz et al., 2018). The diameters of LDs have been quantified using TEM after 2 months of HFD in the hearts of WT, Atg7, or Parkin-deficient mice (Tong et al., 2019), which indicated the presence of larger LD in hearts of both the Atg7 KO and Parkin KO mice vs. WT hearts. Furthermore, cardiac mitochondrial morphology and myofibrillar structure have been shown to be altered by different exercise protocols (Yuan and Pan, 2018). Together, these studies have shown that TEM is an important tool in assessing the contribution of diabetes and obesity with the development of lipotoxicity and similar to many of the previous cardiac studies also have shown changes in autophagy.
Aging
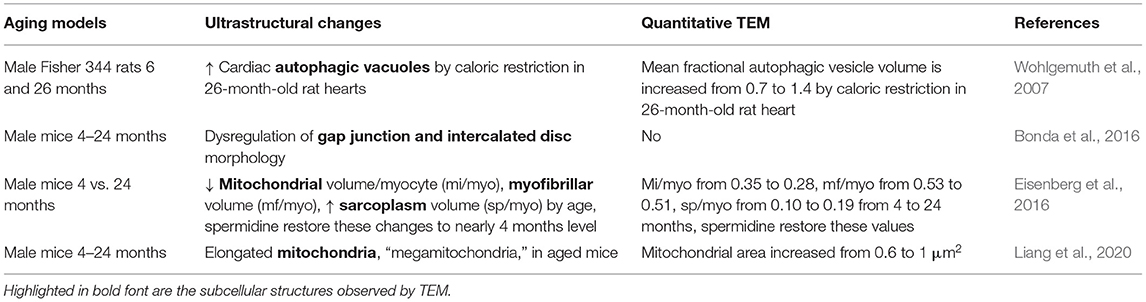
Aging increases the risk for the development of cardiovascular disease and is associated with changes in sarcomeric and mitochondrial structure. TEM analyses have been performed to investigate aging-related changes in cardiac ultrastructure (Table 6). Earlier studies have demonstrated that the presence of cardiac autophagic vacuoles is increased by caloric restriction in aging rat hearts (26 months of age) (Wohlgemuth et al., 2007). Dysregulation of gap junction and intercalated disc morphology have been observed via TEM in aged mouse hearts (24-month-old) in comparison to hearts from younger mice (4-month-old). These changes in gap junction morphology were associated with decreased Cx43, increased β-catenin, and increased collagen (Bonda et al., 2016). In addition, a study by Eisenberg et al. showed a significant decrease in mitochondrial volume, a decrease in myofibrillar volume, and an increase in sarcoplasmic volume in aging hearts (Eisenberg et al., 2016). They also showed that spermidine feeding from 18 to 24 months of age improved cardiac diastolic function and restored these structural changes to close to those in the young (4-month-old) mice (Eisenberg et al., 2016). Notably, the spermidine cardiac protective effect is dependent on the autophagy gene Atg5 (Eisenberg et al., 2016). However, a more recent study demonstrated the presence of elongated mitochondria of aging hearts, termed as “megamitochondria,” which was associated with the increase of mitochondrial-associated p62 and Parkin and decreased autophagy-related 9B (Atg9b), nuclear respiratory factor 1 (Nrf1), and mtDNA/nDNA (Liang et al., 2020). The differences between these two studies may be due to fact that the volumes of mitochondria were normalized to the volumes of the myocytes in the first study, and the absolute area visualized in TEM was used for the latter study. Collectively, these TEM studies show ultrastructural changes in the aging hearts that are associated with significant changes in mitochondrial morphology and autophagy.
Overall, these studies all suggest that the most common TEM observations in models of cardiac physiological and pathological remodeling are those associated with changes in mitochondrial morphology and distribution, and several studies show a link to alterations in autophagy through the visualization of autophagosomes and lysosomes. The next section of the review specifically examines the usage of TEM to visualize autophagy and mitophagy in cardiac tissue.
Use of TEM to Visualize Changes in Autophagic Mechanisms and Pathways
A common theme in many of the studies discussed in the previous section is the contribution of substantial changes in mitochondrial form and function, and mitochondrial quality control processes, such as autophagy and mitophagy, to the dysregulation of the heart during disease. TEM has been heavily utilized for the examination of autophagy due to the ability to visualize autophagosomes and vacuolar structures (Yla-Anttila et al., 2009; Backues et al., 2014) and has been a recommended methodology (Klionsky et al., 2016) for visualizing autophagosomes. This section discusses the application of TEM to visualize changes in cardiac autophagy and mitophagy in transgenic and pharmacological animal models.
Autophagy
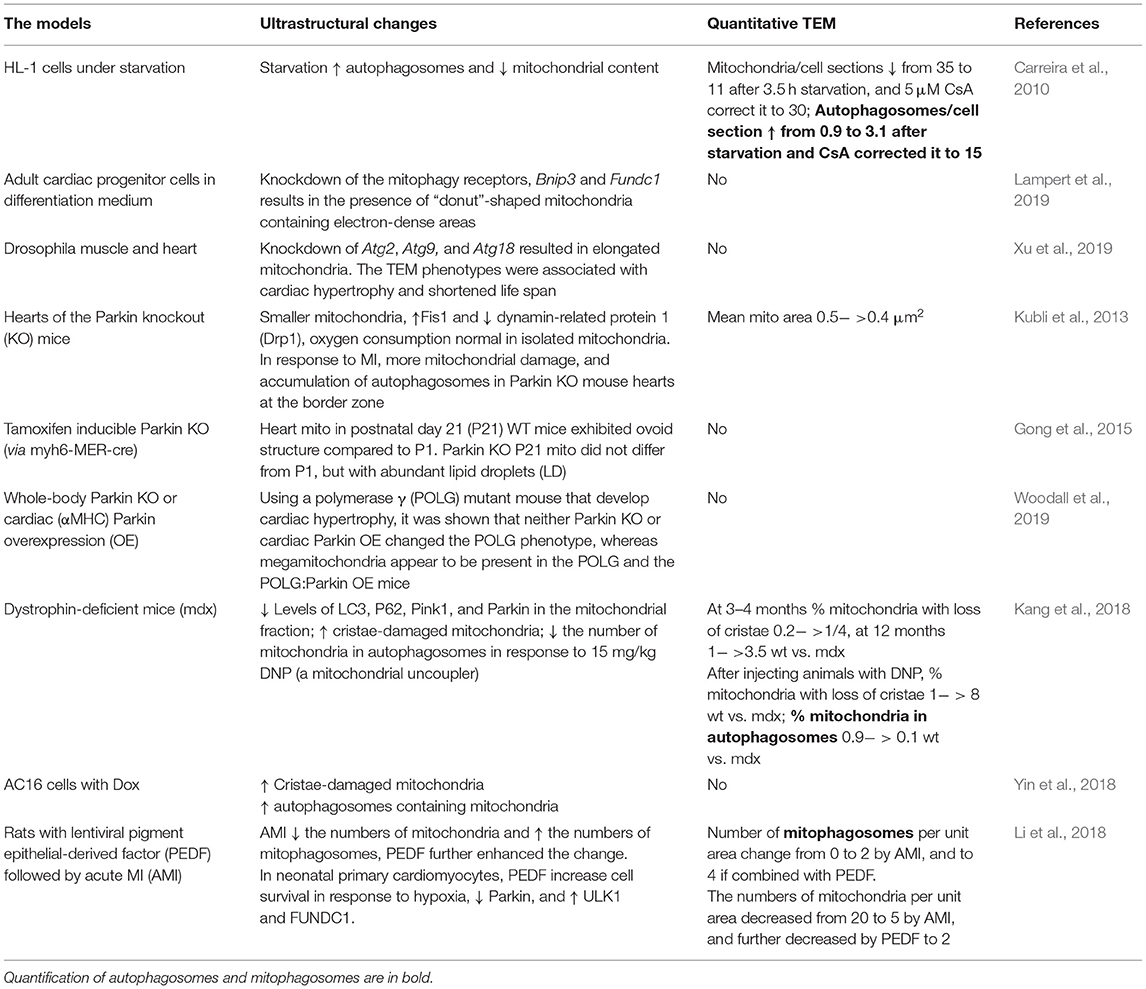
Transmission electron microscopy is an essential tool for visualizing autophagy in the heart and has been described as key to the visualization and quantification of autophagosomes and lysosomes. TEM has proved useful in the assessment of ultrastructural changes in response to starvation-induced autophagy in HL-1 cells, which have been shown to be associated with a decrease of mitochondrial content and an increase in the number of autophagolysosomes (Carreira et al., 2010). As described above, several TEM studies in biopsies from human patients, rat HF, and PE-treated rat adult cardiac myocytes have revealed the presence of autophagic vacuoles (Takemura et al., 2017; Chaanine, 2019; Gil-Cayuela et al., 2019) (Table 3). The numbers of autophagic vacuoles were quantified in rat HF and hypertension models with truncated titin and exposure to Ang II and SIM (Hsieh et al., 2019; Zhou et al., 2019) and qualitatively evaluated in Ang II model in combination with cathepsin S KO (Pan et al., 2012) (Table 3). Qualitative analyses were performed in rat and mouse MI models (Kanamori et al., 2011a,b, 2013; Wu et al., 2016) (Table 4). Several studies have used TEM in combination with manipulations that promote or inhibit autophagy to examine changes in autophagy. An aging study measured autophagic vesicle volume in hearts from the 26-month-old rat with ad libitum vs. caloric restriction feeding (Wohlgemuth et al., 2007) (Table 6). In the mouse MI models (Table 4), 1 week of daily rapamycin injections increased the presence of autophagosomes and lysosomes, whereas bafilomycin-injected animals remained free of these organelles in the heart (Kanamori et al., 2011a). Starvation increased autophagosomes and bafilomycin given prior to MI increased electron-dense bodies in the ischemic border zone in mouse heart (Kanamori et al., 2011b). The autophagy inhibitor, chloroquine, increased the presence of lysosomes without changing autophagosomes in a neonatal cardiomyocyte MI model; in contrast, resveratrol increased autophagosomes (Kanamori et al., 2013).
Many studies have also examined cardiac ultrastructural changes in response to the genetic manipulation of autophagy genes. As mentioned above, cathepsin S deficiency was shown to result in more autophagosomal vacuoles and exacerbates the response to Ang II (Pan et al., 2012) (Table 3). Mice with cardiac-specific knockdown of Atg7 or whole-body KO of Parkin showed the presence of large diameter LDs associated with mitochondria in the heart, which was confirmed using standard biochemical staining of LDs (i.e., Oil Red O staining) (Table 5) (Tong et al., 2019). Knockdown of the mitophagy receptors, Bcl2 interacting protein (Bnip3), and FUN14 domain containing 1 (Fundc1) were associated with the presence of “donut”-shaped mitochondria containing electron-dense areas upon differentiation of adult cardiac progenitor cells (Lampert et al., 2019) (Table 7). Knockdown of Atg2, Atg9, and Atg18 genes was known to play a role in autophagosome formation, in drosophila muscle, and in heart and was found to elongate mitochondria (Xu et al., 2019). The TEM phenotypes were associated with cardiac hypertrophy and shortened life span (Xu et al., 2019). Taken together, these studies indicate that TEM has been a useful tool to visually document autophagy in the heart.
Pink1 and Parkin and Mitophagy
Mitophagy is the specific autophagic degradation of mitochondrial cargo and is mediated by Pink1–Parkin-dependent and -independent pathways. TEM was used to examine cardiac ultrastructure in hearts of the Parkin KO mice under basal conditions and in the response to MI (Kubli et al., 2013). Parkin KO mice have smaller body weight (BW) and heart weight (HW), but the HW/BW is normal, so are the heart functions measured by echocardiography at 3–12 months of age. Disorganized and smaller mitochondria were present at baseline, with decreased Drp1 and increased Fis1 fission proteins, and normal oxygen consumption was measured in isolated mitochondria. In response to MI, WT mice subjected to MI had greater levels of Parkin protein, and Parkin KO mouse hearts had worsened functional outcome, a greater level of mitochondrial damage, and accumulation of autophagosomes at the border zone of the Parkin KO mice (Kubli et al., 2013) (Table 7). In mice with inducible Parkin deletion, heart mitochondria in KO mice at postnatal day 21 (P21) did not change compared to that at P1 (when tamoxifen was injected), whereas in WT mice they become ovoid (Gong et al., 2015). In this study, P21 Parkin KO mouse hearts also exhibited abundant LD (Gong et al., 2015). As mentioned above, studies in the whole-body Parkin KO mouse have shown larger diameter LD in the adult heart (Table 5) (Tong et al., 2019). The proof-reading defective mtDNA polymerase γ (POLG) mutant mouse strains that develop age-dependent cardiac hypertrophy are used as a model of aging, and it was shown using TEM that megamitochondria appear to be present in the POLG and the POLG:Parkin OE mice, whereas neither whole-body Parkin KO or cardiac Parkin OE rescued these POLG-dependent changes in mitochondrial morphology (Woodall et al., 2019).
In hearts from mdx mice [dystrophin-deficient mice as an animal model of Duchenne muscular dystrophy (DMD)], the percent of mitochondria with loss of cristae was increased both at 3–4 months of age and at 12 months of age compared to WT. With the exposure to mitochondrial uncoupler dinitrophenol, the percent of mitochondria in autophagosomes were lower in mdx mice compared to control mice shown using TEM. The levels of Pink1, Parkin, LC3II, and p62 in the mitochondrial fraction were also lower in mdx mice, consistent with decreased mitophagy (Kang et al., 2018). In AC16 cells, the Dox exposure resulted in the increased mitochondrial vacuolization with loss of cristae and the increase of autophagosomes containing mitochondria (Yin et al., 2018). In a study with rats, TEM was used to show that acute MI decreased the numbers of mitochondria and increased the numbers of mitophagosomes, and pigment epithelial-derived factor (PEDF) treatment further elevated the differences (Li et al., 2018). In neonatal primary cardiomyocytes, PEDF increased cell survival in response to hypoxia, decreased Parkin, and increased ULK1 and FUNDC1 (Li et al., 2018) (Table 7). Altogether, TEM studies on autophagy and mitophagy are generally supported with additional biochemical and cell biology studies, which delve into molecular cellular mechanisms and provide assessment of autophagy and mitophagy flux. Nonetheless, more and more studies are adopting TEM as one of the key elements to provide ultrastructural insights into these processes. Figure 4 illustrates the percentage of studies using TEM as a tool to examine autophagy and mitophagy in the heart.
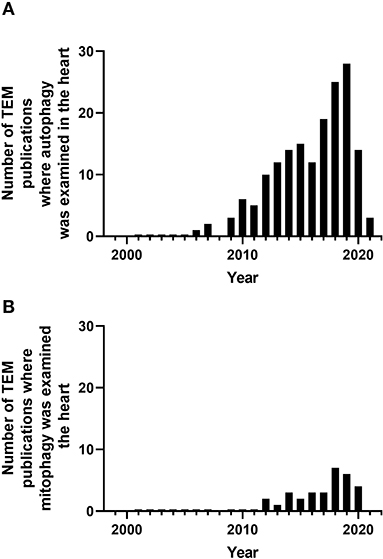
Figure 4. Number of peer-reviewed cardiovascular publications utilizing transmission electron microscopy to examine cardiovascular autophagy (A) and mitophagy (B). Data taken from Pubmed analytics using a search of “Transmission electron microscopy,” “heart,” and “autophagy” or “mitophagy” for year 2000 to present.
Mitochondrial Biogenesis, Fission, and Fusion
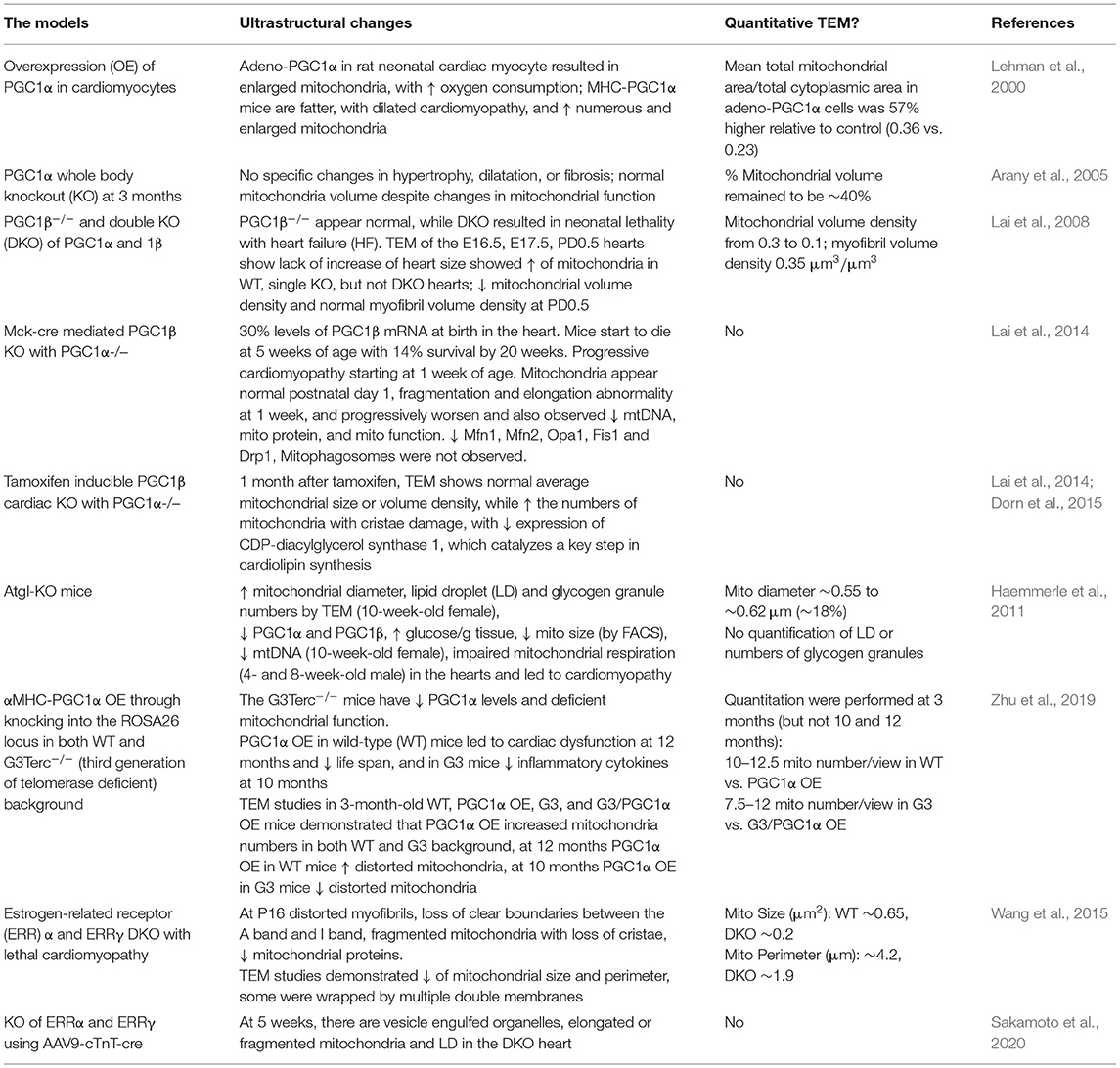
Many TEM studies that have investigated mitochondria often describe changes in the size and number of the mitochondria pool, which are regulated through the balance between biogenesis, mitochondrial degradation, and the dynamic nature of mitochondrial fission and fusion. Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC1) has been shown to play a key role in mitochondrial biogenesis. Adenoviral OE of PGC1? in rat neonatal cardiac myocytes resulted in enlarged mitochondria as viewed by TEM, which was associated with increased oxygen consumption (Lehman et al., 2000). Mice overexpressing cardiomyocyte PGC1α are Larger and develop DCM. One TEM study demonstrated numerous and enlarged mitochondria (Lehman et al., 2000). Despite changes in the mitochondrial function in PGC1α KO mice, there were no changes in mitochondrial morphology in the heart despite changes in the mitochondrial function (Arany et al., 2005). PGC1β KO mice showed normal heart function, whereas double KO (DKO) of PGC1α and PGC1β resulted in neonatal lethality with HF. TEM of the hearts from DKO mice showed decreased mitochondrial volume density and myofibril volume density at PD0.5, likely due to deficient mitochondrial biogenesis starting E17.5 (Lai et al., 2008). The heart and skeletal muscle PGC1α/β−/− KO resulted in a decrease of the PGC1β mRNA level to 30% at birth in the heart (Martin et al., 2014). Mice start to die at 5 weeks of age with 14% survival rate at 20 weeks with progressive cardiomyopathy starting at 1 week of age. Mitochondria appear normal at postnatal day 1, but with fragmentation and elongation abnormality at 1 week, which progressively became worse and decreased mtDNA, mitochondrial proteins, and mitochondrial function were observed. Mitophagosomes were not observed, while genes involved in mitochondrial dynamics, including Mfn1, Mfn2, Opa1, Fis1, and Drp1 were decreased. Mfn1 exhibited the most dramatic decrease (Martin et al., 2014). TEM studies of mice with inducible PGC1β cardiac KO in a PGC1α−/− background showed normal average mitochondrial size or volume density, while the numbers of mitochondria with cristae damage were increased and were associated with a decreased expression of CDP-diacylglycerol synthase 1, which catalyzes a key step in cardiolipin synthesis (Lai et al., 2014; Dorn et al., 2015). In Atgl-KO mice, the decrease in PGC1α and PGC1β has been proposed to mediate the observed mitochondrial defects, with increased mitochondrial diameter shown by TEM in the hearts of 10-week-old female mice (Haemmerle et al., 2011). More recently, a study generated moderate cardiac-specific PGC1α OE through knocking into ROSA26 locus in WT mice and in mice with a G3Terc−/− (third generation of telomerase deficient; G3) background (Zhu et al., 2019). The G3 mice have decreased PGC1α levels and deficient basal and maximal mitochondrial function. TEM studies in 3-month-old WT, PGC1α OE, G3, and G3/ PGC1α OE mice demonstrated that PGC1α OE increased mitochondria numbers in both WT and G3 background (Zhu et al., 2019). Furthermore, at 12 months, PGC1α OE in WT background mice showed increased distorted mitochondria, and at 10 months, PGC1α OE in G3 mice showed decreased distorted mitochondria (Zhu et al., 2019). In addition to PGC1α, estrogen-related receptor α (ERRα) and ERRγ also have a combined role in mitochondrial biogenesis, as DKO of these genes resulted in lethal cardiomyopathy (Wang et al., 2015). At postnatal day 16, there was a significant decrease in the levels of mitochondrial electron transport chain proteins. TEM studies demonstrated distorted myofibrils, loss of clear boundaries between the A band and I band, fragmented mitochondria with loss of cristae, and decrease of mitochondrial size and perimeter (Wang et al., 2015). Postnatal KO of ERRα and ERRγ was generated using AAV9-cTnT-cre i.p. injection at postnatal day 0. At 5 weeks, there was an organelle containing vesicles suggestive of autophagic vacuoles, elongated or fragmented mitochondria, and LD in the DKO heart (Sakamoto et al., 2020). Taken together, these studies indicate that TEM has been a useful tool to demonstrate perturbation of mitochondrial quality control in the heart (Table 8).
Mitochondrial fission proteins are comprised of Drp1 and mitochondrial fission 1 protein (Fis1), and the mitochondrial fusion proteins consist of the mitofusins 1/2 (Mfn1/2) and Optic atrophy protein 1 (Opa1), and alterations in the expression and activity of these proteins impact the available mitochondrial pool. Inducible cardiac-specific Drp1-KO resulted in DCM and HF at 6–8 weeks after KO. TEM demonstrated that mitochondria have preserved cristae; however, they were larger and more elongated (Song et al., 2015a). However, changes in mitochondrial autophagy and mitochondrial stress were confirmed using p62, LC3, LONP2, AFG3L2, and Hsp60 immunoblotting in this study (Song et al., 2015a). Furthermore, early cardiac-specific Drp1 KO mice were found to have compromised LV function at postnatal day 7 and decreased mitochondrial respiratory activity. TEM showed that myofibrils appeared unaffected; however, cardiomyocytes contained heterogeneous mitochondria and mitophagosome-like structures and mitochondrial vacuoles. Additional measurements of mitochondrial volume were made using 3D reconstruction with electron tomography (Kageyama et al., 2014), highlighting the need for additional technologies for the enhancement of traditional TEM. Similar findings were documented using the Drp1 inhibitor mdivi-1 (0.24–1.2 mg/kg; 15 min; i.v); however, the effects of mdivi were more pronounced in MI heart (Ong et al., 2010). Cardiac Parkin KO mice exhibited normal cardiac size, HW, and cardiac function for over 20 weeks. No cardiac OE in Parkin mice led to cardiac damage. TEM studies at 6 weeks after Parkin KO look normal, and Parkin transgenic mice at 30 weeks did not show abnormalities in mitochondrial content, mitochondrial area, and aspect ratio. Inducible Drp1 KO hearts exhibited loss of mitochondria 6 weeks after Drp1 deletion, whereas Parkin/Drp1 DKO delays cardiomyopathy of the Drp1 KO and partially restores mitochondrial content (Song et al., 2015b).
Optic atrophy 1 is involved in the fusion of the mitochondrial inner membrane, and its mutation causes autosomal dominant optic atrophy and the Behr syndrome in humans. Whole-body Opa1 heterozygous KO (+/–) mice exhibit cardiac dysfunction at 12 months but not at 3 months. However, TEM studies demonstrated disruption of orderly arrays of mitochondria between myofilaments and decreased mitochondrial density even at 3 months of age. Loss of cristae has been visualized at 12 months. Decreased complex IV activity and decreased mtDNA was seen at 3 months, and decreased complex I activity was seen at 12 months (Chen L. et al., 2012). A parallel study showed normal cardiac function of Opa+/− mice at 6 months, whereas there was a decreased percentage of small mitochondria (<1 μm3 from 46 to 41%) and an increase in large mitochondria (>1.8 m3 from 21 to 27%) by MitoTracker image analyses. In these studies, mitochondria exhibited variability in size and shape at 3 months, with cristae deformation and the presence of dark material consistent with deficient fusion. There was also an increase of mean surface of individual mitochondrion, and there was no change in SR mitochondrial contacts (Chen L. et al., 2012; Piquereau et al., 2012).
The TEM imaging of cardiac-specific Mfn2 KO hearts has shown the increased presence of autophagosomes at 4 months, associated with increased LC3II and p62 protein levels, cardiac dysfunction at 17 months, and increased sensitivity to I/R injury at 6 months (Zhao et al., 2012). There was an increase of mitochondrial area; however, this neither correlated with changes in mitochondrial volume density nor in mtDNA (Zhao et al., 2012). Mfn2 deletion at early postnatal day 2 exhibited increased mitochondrial mean perimeter, decreased contact length with jSR, decreased Ca2+ uptake, and Ca2+-induced change of NAD(P)H/FAD+, whereas hearts from Mfn1 KO in the same study were normal (Chen Y. et al., 2012). However, in an independent study, where Mfn1 was deleted as early as embryonic day 9.5, mitochondria from Mfn1-KO mice were shown to have an increased presence of small, spherical, fragmented mitochondria; cardiac function and respiration in isolated mitochondria were maintained, but adult cardiac myocytes had better survival in response to H2O2 (Papanicolaou et al., 2012). Not surprisingly, as Drp1 deficiency resulted in cardiomyopathy and mitochondrial morphology changes in cardiac-specific Drp1-KO (Song et al., 2015a), significant changes in mitochondrial morphology and abnormal cardiac remodeling were also observed in the hearts of Mfn1/2 DKO mice (Song et al., 2015a). The cardiac expression of Mfn2 mutation (AA, T111A/S442A, Mfn2 that cannot be phosphorylated by PINK1) resulted in the presence of smaller and elongated mitochondria, associated with decreased succinate dehydrogenase (SDHB) complex II subunit compared to WT Mfn2 transgenics at 3 weeks of age (Gong et al., 2015). At 5 weeks of age, the Mfn2-AA mutant mice developed DCM (Gong et al., 2015). While Drp1 KO hearts have higher LV/wall thickness, Mfn DKO mice have enlarged hearts, but the LV/wall thickness remains unchanged. Mitochondrial area is decreased while the total content is increased in Mfn DKO (Song et al., 2015a). Using Mfn DKO, it was further shown that Mfn1/2 DKO mice are more resistant than WT mice to I/R (Hall et al., 2016). It was again shown that mitochondrial area decreased, while the presence of fragmented interfibrillar and cristae-disrupted mitochondria was reported, along with a decrease of maximal respiration in isolated mitochondria from isolated cardiomyocytes (Hall et al., 2016).
The analyses of TEM images using 3D electron tomography of the cardiomyocyte-specific Mfn2 KO mouse hearts suggest significant increases in mitochondrial volume and aspect ratio, which were associated with increased ER/SR–mitochondrial junctional distances and a reduction in the number of ER/SR–mitochondrial connections (Beikoghli Kalkhoran et al., 2017). As with the study using Mfn2 mutant mice in vivo (Gong et al., 2015), OE of Mfn2 (ad-Mfn2) in neonatal rat cardiomyocytes in vitro did not change mitochondrial number, length and size, aspect ratio, or number of mitophagic vacuoles (Xiong et al., 2019). However, addition of Ang II resulted in an accumulation of mitophagic vacuoles and increased mitochondrial size, indicative of increased mitochondrial damage (Xiong et al., 2019). Collectively, these studies clearly demonstrated that mitochondrial size and shape changes in response to genetic and pharmacological perturbations of fission and fusion proteins and in the context of heart diseases (Table 9).
Lipid Droplets
Transmission electron microscopy is capable of imaging LD at high resolution and has been used to describe conditions impacting the size and distribution of LD, such as cardiac lipotoxicity. In a study on Atgl, which is a lipase associated with LD homeostasis, TEM was used to examine cardiomyocyte LD size, glycogen granules, and mitochondrial diameters in Atgl-KO mice (Haemmerle et al., 2011). As discussed above in Table 8, Atgl-KO mice have lower PGC1α and 1β and reduced mitochondrial diameter and size, glycogen deposition, and LD deposition in the hearts of the KO mice, resulting in cardiomyopathy (Haemmerle et al., 2011). Additionally, cardiac OE of an Atgl-interacting protein, perilipin 5 (Plin5), resulted in the accumulation of LD in the vicinity of enlarged mitochondria in the heart, as shown by TEM (Wang et al., 2013). There was an increase in the thickness of LV wall while the cardiac function was normal and a significant alteration of gene expression occurred (Wang et al., 2013). Although the accumulation of LD may be related to, or affect, autophagy, these earlier studies did not investigate autophagy per se. In addition, there is a lack of studies on the use of TEM to study the role of lipophagy in heart diseases, and this warrants further investigation.
Conclusions
When utilized appropriately, TEM is a powerful tool at not only visualizing the cardiac ultrastructure at a higher degree of resolution and magnification in comparison to traditional microscopy and imaging techniques but also at shedding significant light on the contribution of mitochondrial-derived signaling processes to dysregulation of the myocardium during health and disease. Despite the fact that TEM can be costly, sample preparation (and analyses) can be laborious and can introduce artifacts, these issues can be overcome with appropriate training. Importantly, TEM is a powerful imaging technique in the cardiovascular field and provides the powerful information regarding cardiac ultrastructure and signaling processes, which readily complements other imaging techniques. For example, for dynamic events, such as mitochondrial fission/fusion or engulfment of cargo by autophagosomes, live-cell imaging is preferred as TEM will only provide a snapshot of what is occurring with regard to mitophagy (and the number of mitophagosomes) but does not provide a real-time estimate of mitophagic flux. Furthermore, there is an additional level of complexity for assessing mitophagic flux, given the need for lysosomal inhibitors, which is currently not possible with traditional TEM imaging. Future advances in TEM sample preparation and staining and the development of highly innovative TEM imaging methodologies, such as 3D TEM, cryo-EM tomography, electron tomography, correlative light and electron microscopy (CLEM), and imaging systems, will likely advance the usage and capabilities of TEM in the cardiovascular field. The introduction of these new advances should help with the examination and quantification of measurements of 3D cardiac structures, such as mitochondria, so that accurate measurements may be obtained. In addition, newer labeling techniques, such as immunogold labeling, are promising new developments in the field of cardiac TEM. This allows specific proteins in tissues to be labeled during TEM, although this labeling method needs to be further developed since it is also associated with the production of image artifacts. Taken together, TEM provides essential information for visualization and quantification of cellular ultrastructure in both animal and human hearts. Furthermore, TEM has shed light on the presence of new cell types in the heart, and as technology improves, the capabilities of TEM will likely shift the field further.
Author Contributions
HEC and JZ wrote the manuscript. MSK, SHL, VDU, MEY, and JCC critically read and edited the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work has been partially supported by an American Diabetes Association Postdoctoral Fellowship 1-16-PDF-024 (HEC), NIH T32 HL007457 (MSK), HL152354 (JCC), HL149354 (JCC, VDU, and MEY), HL142216 (VDU, MEY, JCC, and JZ), Nathan Shock Center NIH AG050886 (VDU and JZ), and a UAB AMC21 reload multi-investigator grant (VDU, MEY, JCC, and JZ).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to thank all members of the JZ, MEY, and JCC laboratories for valuable discussions.
Abbreviations
Atg9b, autophagy-related 9B; BW, body weight; CLEM, correlative light and electron microscopy; CVS, cardiovascular system; DCM, dilated cardiomyopathy; DKO, double knockout; Dox, doxorubicin; Drp1, dynamin-related protein 1; ECM, extracellular matrix; ER, endoplasmic reticulum; ERR, estrogen-related receptor α; HF, heart failure; HFD, high fat diet; HW, heart weight; I/R, ischemia/reperfusion; ICM, ischemic cardiomyopathy; IPSCs, induced pluripotent stem cells; jSR, junctional SR; KO, knockout; LD, lipid droplets; LV, left ventricular; LVAD, left ventricular assist device; MAM, mitochondrial-associated membrane; MDV, mitochondrial-derived vesicles; Mfn1, mitofusin 1; Mfn2, mitofusin 2; MI, myocardial infarction; Microtubule-associated protein 1A/1B-light chain 3, LC3; mito, mitochondria; mTOR, mechanistic target of rapamycin; Nrf1, nuclear respiratory factor 1; OE, overexpression; Opa1, optic atrophy 1; PBS, phosphate-buffered saline; PEDF, pigment epithelial-derived factor; PGC-1α, peroxisome proliferator-activated receptor gamma coactivator 1-alpha; Plin5, perilipin 5; SDHB, succinate dehydrogenase [ubiquinone] iron sulfur subunit, mitochondrial; SEM, scanning electron microscopy; sequestosome 1, p62; SR, sarcoplasmic reticulum; TAC, transverse aortic constriction; TEM, transmission electron microscopy.
References
Abbate, A., De Falco, M., Morales, C., Gelpi, R. J., Prisco, M., De Luca, A., et al. (2007). Electron microscopy characterization of cardiomyocyte apoptosis in ischemic heart disease. Int. J. Cardiol. 114, 118–120. doi: 10.1016/j.ijcard.2005.11.025
Ahuja, P., Wanagat, J., Wang, Z., Wang, Y., Liem, D. A., Ping, P., et al. (2013). Divergent mitochondrial biogenesis responses in human cardiomyopathy. Circulation 127, 1957–1967. doi: 10.1161/CIRCULATIONAHA.112.001219
Arany, Z., He, H., Lin, J., Hoyer, K., Handschin, C., Toka, O., et al. (2005). Transcriptional coactivator PGC-1 alpha controls the energy state and contractile function of cardiac muscle. Cell Metab. 1, 259–271. doi: 10.1016/j.cmet.2005.03.002
Babaei, H., Razmaraii, N., Assadnassab, G., Mohajjel Nayebi, A., Azarmi, Y., Mohammadnejad, D., et al. (2020). Ultrastructural and echocardiographic assessment of chronic doxorubicin-induced cardiotoxicity in rats. Arch. Razi Inst. 75, 55–62. doi: 10.22092/ari.2019.116862.1177
Backues, S. K., Chen, D., Ruan, J., Xie, Z., and Klionsky, D. J. (2014). Estimating the size and number of autophagic bodies by electron microscopy. Autophagy 10, 155–164. doi: 10.4161/auto.26856
Beikoghli Kalkhoran, S., Hall, A. R., White, I. J., Cooper, J., Fan, Q., Ong, S. B., et al. (2017). Assessing the effects of mitofusin 2 deficiency in the adult heart using 3D electron tomography. Physiol. Rep. 5:13437. doi: 10.14814/phy2.13437
Benador, I. Y., Veliova, M., Mahdaviani, K., Petcherski, A., Wikstrom, J. D., Assali, E. A., et al. (2018). Mitochondria bound to lipid droplets have unique bioenergetics, composition, and dynamics that support lipid droplet expansion. Cell Metab. 27, 869–85 e6. doi: 10.1016/j.cmet.2018.03.003
Bonda, T. A., Szynaka, B., Sokolowska, M., Dziemidowicz, M., Winnicka, M. M., Chyczewski, L., et al. (2016). Remodeling of the intercalated disc related to aging in the mouse heart. J. Cardiol. 68, 261–268. doi: 10.1016/j.jjcc.2015.10.001
Boudina, S., Sena, S., Theobald, H., Sheng, X., Wright, J. J., Hu, X. X., et al. (2007). Mitochondrial energetics in the heart in obesity-related diabetes: direct evidence for increased uncoupled respiration and activation of uncoupling proteins. Diabetes 56, 2457–2466. doi: 10.2337/db07-0481
Bournat, J. C., and Brown, C. W. (2010). Mitochondrial dysfunction in obesity. Curr. Opin. Endocrinol. Diabetes Obes. 17, 446–452. doi: 10.1097/MED.0b013e32833c3026
Bruggink, A. H., van Oosterhout, M. F., de Jonge, N., Cleutjens, J. P., van Wichen, D. F., van Kuik, J., et al. (2007). Type IV collagen degradation in the myocardial basement membrane after unloading of the failing heart by a left ventricular assist device. Lab. Invest. 87, 1125–1137. doi: 10.1038/labinvest.3700670
Cadete, V. J., Deschenes, S., Cuillerier, A., Brisebois, F., Sugiura, A., Vincent, A., et al. (2016). Formation of mitochondrial-derived vesicles is an active and physiologically relevant mitochondrial quality control process in the cardiac system. J. Physiol. 594, 5343–5362. doi: 10.1113/JP272703
Carreira, R. S., Lee, Y., Ghochani, M., Gustafsson, A. B., and Gottlieb, R. A. (2010). Cyclophilin D is required for mitochondrial removal by autophagy in cardiac cells. Autophagy 6, 462–472. doi: 10.4161/auto.6.4.11553
Caspi, O., Huber, I., Gepstein, A., Arbel, G., Maizels, L., Boulos, M., et al. (2013). Modeling of arrhythmogenic right ventricular cardiomyopathy with human induced pluripotent stem cells. Circ. Cardiovasc. Genet. 6, 557–568. doi: 10.1161/CIRCGENETICS.113.000188
Chaanine, A. H. (2019). Morphological stages of mitochondrial vacuolar degeneration in phenylephrine-stressed cardiac myocytes and in animal models and human heart failure. Medicina 55:60239. doi: 10.3390/medicina55060239
Chen, L., Liu, T., Tran, A., Lu, X., Tomilov, A. A., Davies, V., et al. (2012). OPA1 mutation and late-onset cardiomyopathy: mitochondrial dysfunction and mtDNA instability. J. Am. Heart Assoc. 1:e003012. doi: 10.1161/JAHA.112.003012
Chen, Y., Csordas, G., Jowdy, C., Schneider, T. G., Csordas, N., Wang, W., et al. (2012). Mitofusin 2-containing mitochondrial-reticular microdomains direct rapid cardiomyocyte bioenergetic responses via interorganelle Ca(2+) crosstalk. Circ. Res. 111, 863–875. doi: 10.1161/CIRCRESAHA.112.266585
Christoffersen, C., Bollano, E., Lindegaard, M. L., Bartels, E. D., Goetze, J. P., Andersen, C. B., et al. (2003). Cardiac lipid accumulation associated with diastolic dysfunction in obese mice. Endocrinology 144, 3483–3490. doi: 10.1210/en.2003-0242
Collins, H. E., Pat, B. M., Zou, L., Litovsky, S. H., Wende, A. R., Young, M. E., et al. (2019). Novel role of the ER/SR Ca(2+) sensor STIM1 in the regulation of cardiac metabolism. Am. J. Physiol. Heart Circ. Physiol. 316, H1014–H26. doi: 10.1152/ajpheart.00544.2018
de Weger, R. A., Schipper, M. E., Siera-de Koning, E., van der Weide, P., van Oosterhout, M. F., Quadir, R., et al. (2011). Proteomic profiling of the human failing heart after left ventricular assist device support. J. Heart Lung Transpl. 30, 497–506. doi: 10.1016/j.healun.2010.11.011
Diakos, N. A., Selzman, C. H., Sachse, F. B., Stehlik, J., Kfoury, A. G., Wever-Pinzon, O., et al. (2014). Myocardial atrophy and chronic mechanical unloading of the failing human heart: implications for cardiac assist device-induced myocardial recovery. J. Am. Coll. Cardiol. 64, 1602–1612. doi: 10.1016/j.jacc.2014.05.073
Dorn, G. W. II., Vega, R. B., and Kelly, D. P. (2015). Mitochondrial biogenesis and dynamics in the developing and diseased heart. Genes Dev. 29, 1981–1991. doi: 10.1101/gad.269894.115
Duncan, J. G. (2011). Mitochondrial dysfunction in diabetic cardiomyopathy. Biochim. Biophys. Acta 1813, 1351–1359. doi: 10.1016/j.bbamcr.2011.01.014
Dupont, N., Chauhan, S., Arko-Mensah, J., Castillo, E. F., Masedunskas, A., Weigert, R., et al. (2014). Neutral lipid stores and lipase PNPLA5 contribute to autophagosome biogenesis. Curr. Biol. 24, 609–620. doi: 10.1016/j.cub.2014.02.008
Eisenberg, T., Abdellatif, M., Schroeder, S., Primessnig, U., Stekovic, S., Pendl, T., et al. (2016). Cardioprotection and lifespan extension by the natural polyamine spermidine. Nat. Med. 22, 1428–1438. doi: 10.1038/nm.4222
Fertig, E. T., Gherghiceanu, M., and Popescu, L. M. (2014). Extracellular vesicles release by cardiac telocytes: electron microscopy and electron tomography. J. Cell Mol. Med. 18, 1938–1943. doi: 10.1111/jcmm.12436
Gherghiceanu, M., Manole, C. G., and Popescu, L. M. (2010). Telocytes in endocardium: electron microscope evidence. J. Cell Mol. Med. 14, 2330–2334. doi: 10.1111/j.1582-4934.2010.01133.x
Gherghiceanu, M., and Popescu, L. M. (2010). Cardiomyocyte precursors and telocytes in epicardial stem cell niche: electron microscope images. J. Cell Mol. Med. 14, 871–877. doi: 10.1111/j.1582-4934.2010.01060.x
Gil-Cayuela, C., Lopez, A., Martinez-Dolz, L., Gonzalez-Juanatey, J. R., Lago, F., Rosello-Lleti, E., et al. (2019). The altered expression of autophagy-related genes participates in heart failure: NRBP2 and CALCOCO2 are associated with left ventricular dysfunction parameters in human dilated cardiomyopathy. PLoS ONE 14:e0215818. doi: 10.1371/journal.pone.0215818
Glancy, B. (2020). Visualizing mitochondrial form and function within the cell. Trends Mol. Med. 26, 58–70. doi: 10.1016/j.molmed.2019.09.009
Goh, K. Y., Qu, J., Hong, H., Liu, T., Dell'Italia, L. J., Wu, Y., et al. (2016). Impaired mitochondrial network excitability in failing guinea-pig cardiomyocytes. Cardiovasc. Res. 109, 79–89. doi: 10.1093/cvr/cvv230
Gong, G., Song, M., Csordas, G., Kelly, D. P., Matkovich, S. J., and Dorn, G. W. II. (2015). Parkin-mediated mitophagy directs perinatal cardiac metabolic maturation in mice. Science 350:aad2459. doi: 10.1126/science.aad2459
Gupta, A., Gupta, S., Young, D., Das, B., McMahon, J., and Sen, S. (2010). Impairment of ultrastructure and cytoskeleton during progression of cardiac hypertrophy to heart failure. Lab. Invest. 90, 520–530. doi: 10.1038/labinvest.2010.43
Haemmerle, G., Moustafa, T., Woelkart, G., Buttner, S., Schmidt, A., van de Weijer, T., et al. (2011). ATGL-mediated fat catabolism regulates cardiac mitochondrial function via PPAR-alpha and PGC-1. Nat. Med. 17, 1076–1085. doi: 10.1038/nm.2439
Hall, A. R., Burke, N., Dongworth, R. K., Kalkhoran, S. B., Dyson, A., Vicencio, J. M., et al. (2016). Hearts deficient in both Mfn1 and Mfn2 are protected against acute myocardial infarction. Cell Death Dis. 7:e2238. doi: 10.1038/cddis.2016.139
Hein, S., Scheffold, T., and Schaper, J. (1995). Ischemia induces early changes to cytoskeletal and contractile proteins in diseased human myocardium. J. Thorac. Cardiovasc. Surg. 110, 89–98. doi: 10.1016/S0022-5223(05)80013-3
Hesketh, G. G., Shah, M. H., Halperin, V. L., Cooke, C. A., Akar, F. G., Yen, T. E., et al. (2010). Ultrastructure and regulation of lateralized connexin43 in the failing heart. Circ. Res. 106, 1153–1163. doi: 10.1161/CIRCRESAHA.108.182147
Holloway, G. P., Snook, L. A., Harris, R. J., Glatz, J. F., Luiken, J. J., and Bonen, A. (2011). In obese Zucker rats, lipids accumulate in the heart despite normal mitochondrial content, morphology and long-chain fatty acid oxidation. J. Physiol. 589, 169–180. doi: 10.1113/jphysiol.2010.198663
Howarth, F. C., Qureshi, M. A., Lawrence, P., and Adeghate, E. (2000). Chronic effects of streptozotocin-induced diabetes on the ultrastructure of rat ventricular and papillary muscle. Acta Diabetol. 37, 119–124. doi: 10.1007/s005920070013
Hsieh, C. C., Li, C. Y., Hsu, C. H., Chen, H. L., Chen, Y. H., Liu, Y. P., et al. (2019). Mitochondrial protection by simvastatin against angiotensin II-mediated heart failure. Br. J. Pharmacol. 176, 3791–3804. doi: 10.1111/bph.14781
Ikeda, Y., Inomata, T., Fujita, T., Iida, Y., Nabeta, T., Naruke, T., et al. (2015). Morphological changes in mitochondria during mechanical unloading observed on electron microscopy: a case report of a bridge to complete recovery in a patient with idiopathic dilated cardiomyopathy. Cardiovasc. Pathol. 24, 128–131. doi: 10.1016/j.carpath.2014.10.003
Jahng, J. W., Turdi, S., Kovacevic, V., Dadson, K., Li, R. K., and Sweeney, G. (2015). Pressure overload-induced cardiac dysfunction in aged male adiponectin knockout mice is associated with autophagy deficiency. Endocrinology 156, 2667–2677. doi: 10.1210/en.2015-1162
Kageyama, Y., Hoshijima, M., Seo, K., Bedja, D., Sysa-Shah, P., Andrabi, S. A., et al. (2014). Parkin-independent mitophagy requires Drp1 and maintains the integrity of mammalian heart and brain. EMBO J. 33, 2798–2813. doi: 10.15252/embj.201488658
Kanamori, H., Takemura, G., Goto, K., Maruyama, R., Ono, K., Nagao, K., et al. (2011b). Autophagy limits acute myocardial infarction induced by permanent coronary artery occlusion. Am. J. Physiol. Heart Circ. Physiol. 300, H2261–H2271. doi: 10.1152/ajpheart.01056.2010
Kanamori, H., Takemura, G., Goto, K., Maruyama, R., Tsujimoto, A., Ogino, A., et al. (2011a). The role of autophagy emerging in postinfarction cardiac remodelling. Cardiovasc. Res. 91, 330–339. doi: 10.1093/cvr/cvr073
Kanamori, H., Takemura, G., Goto, K., Tsujimoto, A., Ogino, A., Takeyama, T., et al. (2013). Resveratrol reverses remodeling in hearts with large, old myocardial infarctions through enhanced autophagy-activating AMP kinase pathway. Am. J. Pathol. 182, 701–713. doi: 10.1016/j.ajpath.2012.11.009
Kang, C., Badr, M. A., Kyrychenko, V., Eskelinen, E. L., and Shirokova, N. (2018). Deficit in PINK1/PARKIN-mediated mitochondrial autophagy at late stages of dystrophic cardiomyopathy. Cardiovasc. Res. 114, 90–102. doi: 10.1093/cvr/cvx201
Klang, V., Matsko, N. B., Valenta, C., and Hofer, F. (2012). Electron microscopy of nanoemulsions: an essential tool for characterisation and stability assessment. Micron 43, 85–103. doi: 10.1016/j.micron.2011.07.014
Klionsky, D. J., Abdelmohsen, K., Abe, A., Abedin, M. J., Abeliovich, H., Acevedo Arozena, A., et al. (2016). Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222. doi: 10.1080/15548627.2015.1100356
Kubli, D. A., Zhang, X., Lee, Y., Hanna, R. A., Quinsay, M. N., Nguyen, C. K., et al. (2013). Parkin protein deficiency exacerbates cardiac injury and reduces survival following myocardial infarction. J. Biol. Chem. 288, 915–926. doi: 10.1074/jbc.M112.411363
Lai, L., Leone, T. C., Zechner, C., Schaeffer, P. J., Kelly, S. M., Flanagan, D. P., et al. (2008). Transcriptional coactivators PGC-1alpha and PGC-lbeta control overlapping programs required for perinatal maturation of the heart. Genes Dev. 22, 1948–1961. doi: 10.1101/gad.1661708
Lai, L., Wang, M., Martin, O. J., Leone, T. C., Vega, R. B., Han, X., et al. (2014). A role for peroxisome proliferator-activated receptor gamma coactivator 1 (PGC-1) in the regulation of cardiac mitochondrial phospholipid biosynthesis. J. Biol. Chem. 289, 2250–2259. doi: 10.1074/jbc.M113.523654
Lampert, M. A., Orogo, A. M., Najor, R. H., Hammerling, B. C., Leon, L. J., Wang, B. J., et al. (2019). BNIP3L/NIX and FUNDC1-mediated mitophagy is required for mitochondrial network remodeling during cardiac progenitor cell differentiation. Autophagy 15, 1182–1198. doi: 10.1080/15548627.2019.1580095
Lehman, J. J., Barger, P. M., Kovacs, A., Saffitz, J. E., Medeiros, D. M., and Kelly, D. P. (2000). Peroxisome proliferator-activated receptor gamma coactivator-1 promotes cardiac mitochondrial biogenesis. J. Clin. Invest. 106, 847–856. doi: 10.1172/JCI10268
Li, X., Wu, Y., Zhao, J., Wang, H., Tan, J., Yang, M., et al. (2020). Distinct cardiac energy metabolism and oxidative stress adaptations between obese and non-obese type 2 diabetes mellitus. Theranostics 10, 2675–2695. doi: 10.7150/thno.40735
Li, Y., Liu, Z., Zhang, Y., Zhao, Q., Wang, X., Lu, P., et al. (2018). PEDF protects cardiomyocytes by promoting FUNDC1 mediated mitophagy via PEDF-R under hypoxic condition. Int. J. Mol. Med. 41, 3394–3404. doi: 10.3892/ijmm.2018.3536
Liang, W., Moyzis, A. G., Lampert, M. A., Diao, R. Y., Najor, R. H., and Gustafsson, A. B. (2020). Aging is associated with a decline in Atg9b-mediated autophagosome formation and appearance of enlarged mitochondria in the heart. Aging Cell 2020:e13187. doi: 10.1111/acel.13187
Lukasz Mielanczyk, N. M., Klymenko, O., and Wojnicz, R. (2015). The Transmission Electron Microscope- Theory and Applications (Intech), 193–239. doi: 10.5772/60680
Maron, B. J., Ferrans, V. J., and Jones, M. (1975b). The spectrum of degenerative changes in hypertrophied human cardiac muscle cells: an ultrastructural study. Recent Adv. Stud. Cardiac. Struct. Metab. 8, 447–466.
Maron, B. J., Ferrans, V. J., and Roberts, W. C. (1975a). Ultrastructural features of degenerated cardiac muscle cells in patients with cardiac hypertrophy. Am. J. Pathol. 79, 387–434.
Martin, O. J., Lai, L., Soundarapandian, M. M., Leone, T. C., Zorzano, A., Keller, M. P., et al. (2014). A role for peroxisome proliferator-activated receptor gamma coactivator-1 in the control of mitochondrial dynamics during postnatal cardiac growth. Circ. Res. 114, 626–636. doi: 10.1161/CIRCRESAHA.114.302562
Miotto, P. M., LeBlanc, P. J., and Holloway, G. P. (2018). High-fat diet causes mitochondrial dysfunction as a result of impaired ADP sensitivity. Diabetes 67, 2199–2205. doi: 10.2337/db18-0417
Montaigne, D., Marechal, X., Coisne, A., Debry, N., Modine, T., Fayad, G., et al. (2014). Myocardial contractile dysfunction is associated with impaired mitochondrial function and dynamics in type 2 diabetic but not in obese patients. Circulation 130, 554–564. doi: 10.1161/CIRCULATIONAHA.113.008476
Mudhar, H. S., Wagner, B. E., and Suvarna, S. K. (2001). Electron microscopy of myocardial tissue. A nine year review. J. Clin. Pathol. 54, 321–325. doi: 10.1136/jcp.54.4.321
Ong, S. B., Subrayan, S., Lim, S. Y., Yellon, D. M., Davidson, S. M., and Hausenloy, D. J. (2010). Inhibiting mitochondrial fission protects the heart against ischemia/reperfusion injury. Circulation 121, 2012–2022. doi: 10.1161/CIRCULATIONAHA.109.906610
Pan, L., Li, Y., Jia, L., Qin, Y., Qi, G., Cheng, J., et al. (2012). Cathepsin S deficiency results in abnormal accumulation of autophagosomes in macrophages and enhances Ang II-induced cardiac inflammation. PLoS ONE 7:e35315. doi: 10.1371/journal.pone.0035315
Papanicolaou, K. N., Ngoh, G. A., Dabkowski, E. R., O'Connell, K. A., Ribeiro, R. F. Jr., Stanley, W. C., et al. (2012). Cardiomyocyte deletion of mitofusin-1 leads to mitochondrial fragmentation and improves tolerance to ROS-induced mitochondrial dysfunction and cell death. Am. J. Physiol. Heart Circ. Physiol. 302, H167–H179. doi: 10.1152/ajpheart.00833.2011
Piquereau, J., Caffin, F., Novotova, M., Prola, A., Garnier, A., Mateo, P., et al. (2012). Down-regulation of OPA1 alters mouse mitochondrial morphology, PTP function, and cardiac adaptation to pressure overload. Cardiovasc. Res. 94, 408–417. doi: 10.1093/cvr/cvs117
Popescu, L. M., Manole, C. G., Gherghiceanu, M., Ardelean, A., Nicolescu, M. I., Hinescu, M. E., et al. (2010). Telocytes in human epicardium. J. Cell Mol. Med. 14, 2085–2093. doi: 10.1111/j.1582-4934.2010.01129.x
Saito, T., Asai, K., Sato, S., Takano, H., Mizuno, K., and Shimizu, W. (2015). Ultrastructural features of cardiomyocytes in dilated cardiomyopathy with initially decompensated heart failure as a predictor of prognosis. Eur. Heart J. 36, 724–732. doi: 10.1093/eurheartj/ehu404
Sakamoto, T., Matsuura, T. R., Wan, S., Ryba, D. M., Kim, J. U., Won, K. J., et al. (2020). A critical role for estrogen-related receptor signaling in cardiac maturation. Circ. Res. 126, 1685–1702. doi: 10.1161/CIRCRESAHA.119.316100
Schaper, J., Froede, R., Hein, S., Buck, A., Hashizume, H., Speiser, B., et al. (1991). Impairment of the myocardial ultrastructure and changes of the cytoskeleton in dilated cardiomyopathy. Circulation 83, 504–514. doi: 10.1161/01.CIR.83.2.504
Searls, Y. M., Smirnova, I. V., Fegley, B. R., and Stehno-Bittel, L. (2004). Exercise attenuates diabetes-induced ultrastructural changes in rat cardiac tissue. Med. Sci. Sports Exerc. 36, 1863–1870. doi: 10.1249/01.MSS.0000145461.38224.EC
Shirakabe, A., Zhai, P., Ikeda, Y., Saito, T., Maejima, Y., Hsu, C. P., et al. (2016). Drp1-dependent mitochondrial autophagy plays a protective role against pressure overload-induced mitochondrial dysfunction and heart failure. Circulation 133, 1249–1263. doi: 10.1161/CIRCULATIONAHA.115.020502
Shpilka, T., and Elazar, Z. (2015). Lipid droplets regulate autophagosome biogenesis. Autophagy 11, 2130–2131. doi: 10.1080/15548627.2015.1093719
Shpilka, T., Welter, E., Borovsky, N., Amar, N., Mari, M., Reggiori, F., et al. (2015). Lipid droplets and their component triglycerides and steryl esters regulate autophagosome biogenesis. EMBO J. 34, 2117–2131. doi: 10.15252/embj.201490315
Sibouakaz, D., Othmani-Mecif, K., Fernane, A., Taghlit, A., and Benazzoug, Y. (2018). Biochemical and ultrastructural cardiac changes induced by high-fat diet in female and male prepubertal rabbits. Anal. Cell Pathol. 2018:6430696. doi: 10.1155/2018/6430696
Song, M., Gong, G., Burelle, Y., Gustafsson, A. B., Kitsis, R. N., Matkovich, S. J., et al. (2015b). Interdependence of parkin-mediated mitophagy and mitochondrial fission in adult mouse hearts. Circ. Res. 117, 346–351. doi: 10.1161/CIRCRESAHA.117.306859
Song, M., Mihara, K., Chen, Y., Scorrano, L., and Dorn, G. W. 2nd. (2015a). Mitochondrial fission and fusion factors reciprocally orchestrate mitophagic culling in mouse hearts and cultured fibroblasts. Cell Metab. 21, 273–286. doi: 10.1016/j.cmet.2014.12.011
Stephens, E. H., Han, J., Trawick, E. A., Di Martino, E. S., Akkiraju, H., Brown, L. M., et al. (2018). Left-ventricular assist device impact on aortic valve mechanics, proteomics and ultrastructure. Ann. Thorac. Surg. 105, 572–580. doi: 10.1016/j.athoracsur.2017.08.030
Sugawara, R., Sugiyama, H., Nakamura, K., Tohgi, K., Hongo, T., Tsuchiya, M., et al. (2021). Electron microscopy revealed massive lipid droplets in cardiomyocytes in a patient with cardiogenic shock following a fulminant type 1 diabetes mellitus. Int Heart J. 62, 197–200. doi: 10.1536/ihj.20-537
Takemura, G., Kanamori, H., Okada, H., Tsujimoto, A., Miyazaki, N., Takada, C., et al. (2017). Ultrastructural aspects of vacuolar degeneration of cardiomyocytes in human endomyocardial biopsies. Cardiovasc. Pathol. 30, 64–71. doi: 10.1016/j.carpath.2017.06.012
Tay, H., Vandecasteele, T., and Van den Broeck, W. (2017). Identification of telocytes in the porcine heart. Anat. Histol. Embryol. 46, 519–527. doi: 10.1111/ahe.12296
Te Rijdt, W. P., van Tintelen, J. P., Vink, A., van der Wal, A. C., de Boer, R. A., van den Berg, M. P., et al. (2016). Phospholamban p.Arg14del cardiomyopathy is characterized by phospholamban aggregates, aggresomes, and autophagic degradation. Histopathology 69, 542–550. doi: 10.1111/his.12963
Tong, M., Saito, T., Zhai, P., Oka, S. I., Mizushima, W., Nakamura, M., et al. (2019). Mitophagy is essential for maintaining cardiac function during high fat diet-induced diabetic cardiomyopathy. Circ. Res. 124, 1360–1371. doi: 10.1161/CIRCRESAHA.118.314607
Treskatsch, S., Shakibaei, M., Feldheiser, A., Shaqura, M., Dehe, L., Roepke, T. K., et al. (2015). Ultrastructural changes associated with myocardial apoptosis, in failing rat hearts induced by volume overload. Int. J. Cardiol. 197, 327–332. doi: 10.1016/j.ijcard.2015.06.067
van Heerebeek, L., Borbely, A., Niessen, H. W., Bronzwaer, J. G., van der Velden, J., Stienen, G. J., et al. (2006). Myocardial structure and function differ in systolic and diastolic heart failure. Circulation 113, 1966–1973. doi: 10.1161/CIRCULATIONAHA.105.587519
Wang, H., Sreenivasan, U., Gong, D. W., O'Connell, K. A., Dabkowski, E. R., Hecker, P. A., et al. (2013). Cardiomyocyte-specific perilipin 5 overexpression leads to myocardial steatosis and modest cardiac dysfunction. J. Lipid Res. 54, 953–965. doi: 10.1194/jlr.M032466
Wang, T., McDonald, C., Petrenko, N. B., Leblanc, M., Wang, T., Giguere, V., et al. (2015). Estrogen-related receptor alpha (ERRalpha) and ERRgamma are essential coordinators of cardiac metabolism and function. Mol. Cell Biol. 35, 1281–1298. doi: 10.1128/MCB.01156-14
Wohlgemuth, S. E., Julian, D., Akin, D. E., Fried, J., Toscano, K., Leeuwenburgh, C., et al. (2007). Autophagy in the heart and liver during normal aging and calorie restriction. Rejuvenation Res. 10, 281–292. doi: 10.1089/rej.2006.0535
Woodall, B. P., Orogo, A. M., Najor, R. H., Cortez, M. Q., Moreno, E. R., Wang, H., et al. (2019). Parkin does not prevent accelerated cardiac aging in mitochondrial DNA mutator mice. JCI Insight 5:127713. doi: 10.1172/jci.insight.127713
Wu, L., Maimaitirexiati, X., Jiang, Y., and Liu, L. (2016). Parkin regulates mitochondrial autophagy after myocardial infarction in rats. Med. Sci. Monit. 22, 1553–1559. doi: 10.12659/MSM.898722
Xiong, W., Ma, Z., An, D., Liu, Z., Cai, W., Bai, Y., et al. (2019). Mitofusin 2 participates in mitophagy and mitochondrial fusion against angiotensin II-induced cardiomyocyte injury. Front. Physiol. 10:411. doi: 10.3389/fphys.2019.00411
Xu, P., Damschroder, D., Zhang, M., Ryall, K. A., Adler, P. N., Saucerman, J. J., et al. (2019). Atg2, Atg9 and Atg18 in mitochondrial integrity, cardiac function and healthspan in Drosophila. J. Mol. Cell Cardiol. 127, 116–124. doi: 10.1016/j.yjmcc.2018.12.006
Yin, J., Guo, J., Zhang, Q., Cui, L., Zhang, L., Zhang, T., et al. (2018). Doxorubicin-induced mitophagy and mitochondrial damage is associated with dysregulation of the PINK1/parkin pathway. Toxicol. in vitro 51, 1–10. doi: 10.1016/j.tiv.2018.05.001
Yla-Anttila, P., Vihinen, H., Jokitalo, E., and Eskelinen, E. L. (2009). Monitoring autophagy by electron microscopy in Mammalian cells. Methods Enzymol. 452, 143–164. doi: 10.1016/S0076-6879(08)03610-0
Yuan, Y., and Pan, S. S. (2018). Parkin mediates mitophagy to participate in cardioprotection induced by late exercise preconditioning but Bnip3 does not. J. Cardiovasc. Pharmacol. 71, 303–316. doi: 10.1097/FJC.0000000000000572
Zhao, T., Huang, X., Han, L., Wang, X., Cheng, H., Zhao, Y., et al. (2012). Central role of mitofusin 2 in autophagosome-lysosome fusion in cardiomyocytes. J. Biol. Chem. 287, 23615–23625. doi: 10.1074/jbc.M112.379164
Zhou, J., Ng, B., Ko, N. S. J., Fiedler, L. R., Khin, E., Lim, A., et al. (2019). Titin truncations lead to impaired cardiomyocyte autophagy and mitochondrial function in vivo. Hum. Mol. Genet. 28, 1971–1981. doi: 10.1093/hmg/ddz033
Zhou, Y. T., Grayburn, P., Karim, A., Shimabukuro, M., Higa, M., Baetens, D., et al. (2000). Lipotoxic heart disease in obese rats: implications for human obesity. Proc. Natl. Acad. Sci. U. S. A. 97, 1784–1789. doi: 10.1073/pnas.97.4.1784
Keywords: transmission electron microscopy, heart, heart failure, myocardial infarction, mitochondria, autophagy, mitophagy, aging
Citation: Collins HE, Kane MS, Litovsky SH, Darley-Usmar VM, Young ME, Chatham JC and Zhang J (2021) Mitochondrial Morphology and Mitophagy in Heart Diseases: Qualitative and Quantitative Analyses Using Transmission Electron Microscopy. Front. Aging 2:670267. doi: 10.3389/fragi.2021.670267
Received: 20 February 2021; Accepted: 26 March 2021;
Published: 06 May 2021.
Edited by:
Sruti Shiva, University of Pittsburgh, United StatesReviewed by:
Zhao Wang, University of Texas Southwestern Medical Center, United StatesQiangrong Liang, New York Institute of Technology, United States
Copyright © 2021 Collins, Kane, Litovsky, Darley-Usmar, Young, Chatham and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jianhua Zhang, Smlhbmh1YXpoYW5nQHVhYm1jLmVkdQ==
†Present address: Helen E. Collins, Division of Environmental Medicine, Department of Medicine, University of Louisville, Louisville, KY, United States
 Helen E. Collins
Helen E. Collins Mariame Selma Kane2
Mariame Selma Kane2 Martin E. Young
Martin E. Young John C. Chatham
John C. Chatham Jianhua Zhang
Jianhua Zhang