1
Smilow Neuroscience Program, Department of Physiology & Neuroscience, New York, USA
2
Medical Scientist Training Program, New York University School of Medicine, New York, USA
3
Department of Molecular and Cellular Physiology, Stanford University, Stanford, USA
The gene encoding the dipeptidyl peptidase-like protein DPP6 (also known as DPPX) has been associated with human neural disease. However, until recently no function had been found for this protein. It has been proposed that DPP6 is an auxiliary subunit of neuronal Kv4 K+ channels, the ion channels responsible for the somato-dendritic A-type K+ current, an ionic current with crucial roles in the regulation of firing frequency, dendritic integration and synaptic plasticity. This view has been supported mainly by studies showing that DPP6 is necessary to generate channels with biophysical properties resembling the native channels in some neurons. However, independent evidence that DPP6 is a component of neuronal Kv4 channels in the brain, and whether this protein has other functions in the CNS is still lacking. We generated antibodies to DPP6 proteins to compare their distribution in brain with that of the Kv4 pore-forming subunits. DPP6 proteins were prominently expressed in neuronal populations expressing Kv4.2 proteins and both types of protein were enriched in the dendrites of these cells, strongly supporting the hypothesis that DPP6 is an associated protein of Kv4 channels in brain neurons. The observed similarity in the cellular and subcellular patterns of expression of both proteins suggests that this is the main function of DPP6 in brain. However, we also found that DPP6 antibodies intensely labeled the hippocampal mossy fiber axons, which lack Kv4 proteins, suggesting that DPP6 proteins may have additional, Kv4-unrelated functions.
Mutations in the gene encoding DPP6 (also known as DPPX) have been recently associated with human disease, including amyotrophic lateral sclerosis (ALS) (Cronin et al., 2008
; Garber, 2008
; van Es et al., 2008
) and autism (Marshall et al., 2008
). The latter include de-novo copy number variations (CNVs), which are considered particularly significant since these events are extremely rare in the human genome (Sebat, 2007
; Sebat et al., 2007
). Moreover, mutations in the DPP6 gene have not been detected in a large control population (Marshall et al., 2008
). These discoveries underscore the importance of understanding the function of DPP6 proteins in the CNS.
DPP6 is a type II membrane glycoprotein expressed predominantly in the CNS. The protein consists of a large extracellular C-terminal domain and a single membrane-spanning domain followed by a short alternatively spliced intracellular N-terminal sequence (Kin et al., 2001
; Nadal et al., 2003
; Strop et al., 2004
). DPP6 proteins are related and share a similar structure as dipeptidyl peptidase IV (DPPIV, also known as CD26) a serine protease and cell adhesion protein with important functions in the endocrine and immune systems (Boonacker and Van Noorden, 2003
; De Meester et al., 1999
; Lambeir et al., 2003
). However, DPP6 proteins lack enzymatic activity due to mutations in the catalytic region, and until recently no function for these proteins had been described.
Nadal et al. (2003)
found that DPP6 proteins can be
co-purified with Kv4.2 K+ channel proteins from rat cerebellar membranes, suggesting that they might be associated proteins of Kv4 K+ channels. Consistent with this hypothesis, DPP6 proteins were found to regulate the membrane trafficking of Kv4 proteins in heterologous cells and to modify channel properties (Amarillo et al., 2008
; Jerng et al., 2004a
; Nadal et al., 2003
). Products of another DPP gene closely related to DPP6 known as DPP10 were later discovered, and found to have similar effects on Kv4 channels in heterologous cells (Jerng et al., 2004b
, 2005
; Qi et al., 2003
; Ren et al., 2005
; Zagha et al., 2005
).
These studies have led to the view that Kv4 channels in neurons are ternary molecular complexes that include, in addition to pore-forming subunits of the Kv4 subfamily, two types of auxiliary subunits, K+ channel interacting proteins (KChIPs) and the DPP-like (DPPL) proteins DPP6 or DPP10 (Amarillo et al., 2008
; Jerng et al., 2004b
, 2007
; Nadal et al., 2003
). Co expression of Kv4 proteins with KChIPs and DPPLs in heterologous cells results in channels that more closely resemble native channels in certain neurons (Amarillo et al., 2008
; Jerng et al., 2005
, 2007
; Nadal et al., 2003
).
KChIPs were identified as Kv4 channel associated proteins in yeast two hybrid screens using the amino terminal region of Kv4.3 as bait (An et al., 2000
). Rhodes et al. (2004)
studied the distribution of KChIP proteins in brain and showed that it resembles the distribution of Kv4 proteins. Given that Kv4 proteins have relatively restricted cellular and subcellular distributions in brain, this is a powerful argument supporting the view that KChIPs are integral components of native ISA channel complexes in brain tissue. Similar studies with DPPLs have not been performed due to the lack of good quality antibodies suitable for high-resolution immunolocalization in native tissue.
To address this problem, we raised antibodies to DPP6 proteins to investigate their distribution in brain tissue and compared it to the distribution of Kv4 proteins. If DPP6 proteins are associated with Kv4 channels in neurons one would predict that the two types of protein would be expressed in the same neurons and in the same neuronal compartments. We found that the cellular and subcellular pattern of expression of DPP6 closely matches that of Kv4.2, supporting the hypothesis that DPP6 is associated with Kv4 channels in the brain. However, we found one structure, the hippocampal mossy fiber axons, which lack Kv4 proteins but were intensely labeled by DPP6 antibodies, suggesting that DPP6 proteins may have additional, Kv4-unrelated functions and provide the first example of an axonal localization for this protein.
Recombinant DPP6 Expression and Antibody Production
Recombinant DPP6 protein was prepared essentially as described in Strop et al. (2004)
. Briefly, the extracellular domain of human DPP6-L (residues 117–865) was cloned into the pAcGP67A vector (Pharmingen) in frame with the gp67 secretion sequence. A His10 tag was added at the C terminus for affinity purification. Spodoptera frugiperda (Sf9) cells (Invitrogen) were transfected with pAcGP67A carrying the DPP6 cDNA and linearized AcNPV DNA (Sapphire Baculovirus DNA; Orbigen). Baculovirus was amplified in Sf9 cells. Large-scale protein expression was performed by infection of Trichoplusia ni (Hi5) cells. The media containing the secreted, glycosylated, recombinant protein was concentrated, dialyzed and purified with Ni-NTA resin (Qiagen) and by size-exclusion chromatography using a Superdex 200 16/60 column (Amersham Pharmacia Biotech). The peak corresponding to the DPP6 dimer was then concentrated to 10 mg/ml in 100 kDa Centricon (Millipore). The purified recombinant DPP6 protein was injected into rabbits for the generation of polyclonal antibodies at the Pocono Rabbit Farm (Canadensis, PA). Antibodies to DPP6 were purified by affinity chromatography.
Generation of DPP6 (−/−) Mice
We generated a conditional DPP6 knockout (DPP6fl/fl) by inserting lox P sites in the introns flanking exon 2 (see Figure 1
; Rudy et al., 2008
; Zagha et al., 2008
), which encodes the entire transmembrane and juxtamembrane domain, as well as the beginning of the extracellular C-terminal domain of DPP6. Exon 2 is the most 5′ sequence common to all spliced isoforms and is essential to the generation of a membrane protein. To generate null alleles we crossed the DPP6fl/fl mice to a Cre deleter strain expressing Cre recombinase in the germline (see Figure 1
). Mice were genotyped by PCR using primers flanking the lox P sites as indicated in Figure 1
. DPP6−/− mice lacked DPP6 mRNA as demonstrated by RT-PCR of brain RNA using primers: 5′-GGACCTTAGGGCTTTGGGA-3′ and 5′-CTGGTATCTTCCGCTGGG-3′. Expression of GAPDH mRNA was used as a control of mRNA quality and efficiency of reverse transcription. GAPDH PCR used primers: 5′-TGTTCCTACCCCCAATGTGT-3′ and 5′-TGTGAGGGAGATGCTCAGTG-3′.
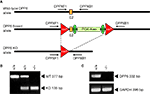
Figure 1. Strategy for the generation of conditional DPP6 knockout mice. (A) LoxP sites were introduced flanking exon 2 of the DPP6 gene. Floxed mice were bred to a Cre deleter strain expressing Cre recombinase in the germline to generate null alleles. (B) Genotyping for DPP6 alleles. Amplification products of wildtype, heterozygote and homozygote nulls following PCR with primers DPPXF1 and DPPXB1. (C) RT-PCR of rat brain mRNA demonstrates lack of DPP6 transcripts in DPP6−/− mice. PCR of GAPDH mRNA was used as control.
Transient Transfections and Immunocytochemistry
CHO-K1 (ATCC, Manassas, VA) cells were cultured, transfected with DPP cDNAs, and membrane extracts collected as described previously in Chang et al. (2007)
. Cultured cells were immunostained with purified polyclonal rabbit DPP6 antibodies (1:50). DPP6 immunoreactivity was visualized with CY3-congugated goat anti-rabbit secondary antibodies.
Mouse Brain Membrane Extracts and Immunoblot Analysis
Mouse brain membrane fractions from wild type and DPP6 null animals (Rudy et al., 2008
; Zagha et al. 2008
) and preparation of immunoblots were carried out as described in Chang et al. (2007)
. The immunoblots were incubated with DPP6 antibodies (1:1000 dilution), followed by horseradish peroxidase-linked anti-rabbit secondary antibodies prepared in donkey (Pierce). Bound antibodies were detected by chemiluminescence with an ECL detection kit (Pierce).
Immunoperoxidase and Immunofluorescent Histology
Rabbit anti-DPP6 antibodies and mouse monoclonal anti-Kv4.2; K57/1 and anti-Kv4.3 K75/41 Neuromab; www.neuromab.org were used for immunolocalization experiments in brain. Immunoperoxidase [DPP6 antibodies diluted (1:1000); Kv4.2 (1:2000)] and immunofluorescent histology [DPP6 antibodies diluted (1:50); Kv4.2 (1:1000); Kv4.3 (1:1000)] was carried out as described in Chang et al. (2007)
. Images were obtained as described in Chang et al. (2007)
.
Generation and Characterization of DPP6 Antibodies
Several attempts to raise site-specific antibodies to DPP6 proteins using synthetic peptides or fusion proteins, methods that have been used traditionally to raise antibodies to ion channel components, failed to produce good quality antibodies. These attempts included the use of peptide antigens corresponding to exposed sequences based on the crystallographic structure of the extracellular domain of DPP6 (Strop et al., 2004
). Some of these anti-peptide antibodies recognized DPP6 proteins in immunoblots, but did not produce good quality immunostaining in brain tissue e.g. (Kin et al., 2001
; Nadal et al., 2003
). Based on this experience, we used the Baculovirus system to produce the entire glycosylated extracellular domain of DPP6. Size-exclusion chromatography demonstrated that the protein formed dimers, indicating that it had folded properly and retained its native structure (Strop et al., 2004
). Recombinant DPP6 was used to immunize rabbits and antibodies to DPP6 were purified by affinity chromatography. These antibodies proved to be of good quality for immunoblotting and immunostaining in brain tissue.
The antibodies recognized a band of approximately 115 kDa in immunoblots of membrane extracts from CHO cells transfected with DPP6-S or DPP6-L (two alternative spliced DPP6 isoforms with different intracellular amino termini, but identical extracellular domains (Nadal et al., 2006
; Figure 2
A). This band was absent in immunoblots of membrane extracts from untransfected cells or extracts from cells transfected with cDNAs of products of the DPP6-related gene DPP10 (Figure 2
A). A band of similar size was observed in immunoblots of brain membrane fractions prepared from wildtype mice, whereas no bands were observed in immunoblots of extracts from DPP6 knockout mice (Figure 2
B).
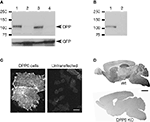
Figure 2. Specificity of antibodies against DPP6 proteins. (A) DPP6 antibodies detected a band of approximately 115 kD in immunoblots of extracts from CHO cells transfected with DPP6-S (lane 1) or DPP6-L (lane 3). The antibodies were specific for detection of DPP6 proteins, since no bands were detected in extracts from non-transfected cells (lane 2) or cells transfected with DPP10 (lane 4). All cells were also transfected with GFP to control for transfection efficiency. (B) DPP6 antibodies detected a band of similar size in immunoblots of brain membranes obtained from wild type mice (lane 1) but not from DPP6 knockout mice (lane 2). (C) Immunofluorescent detection of DPP6 protein in DPP6-transfected CHO cells (left panel). No labeling is observed in untransfected CHO cells (right panel). Note that, as previously reported, transfection with DPPs produces flattening of CHO cells (Nadal et al., 2003
; Zagha et al., 2005
). (D) DPP6 antibodies labeled several brain structures in wild type mice. Immunoperoxidase staining was reduced to background levels in DPP6 knockout mice. Scale bar: 10 μm (C); 2 mm (D).
The antibodies also performed well in immunohistochemistry of transfected cultured cells and mouse brain tissue. Cells transfected with DPP6-S, but not untransfected cells, had surface immunostaining (Figure 2
C). In brain tissue, the antibodies produced immunostaining patterns consistent with the patterns of distribution of DPP6 mRNA transcripts (Nadal et al., 2006
; Zagha et al., 2005
; Figure 2
D). Furthermore, the immunostaining produced by the DPP6 antibodies in mouse brain was entirely absent in brains from DPP6 knockout mice (Figure 2
D). Together these data demonstrate that the antibodies are specific and do not cross react with other brain components.
The Pattern of Expression of DPP6 in Brain Closely Resembles the Pattern of Kv4.2 Proteins
There are three known Kv4 genes, Kv4.1, Kv4.2 and Kv4.3, two of which, Kv4.2 and Kv4.3, are highly expressed in the brain (Serodio and Rudy, 1998
). Some neurons contain both Kv4.2 and Kv4.3 products, which might be forming heteromultimeric channels containing both Kv4 subunits. However, many neuronal populations express predominantly Kv4.2 or Kv4.3 (Rhodes et al., 2004
; Serodio and Rudy, 1998
; Strassle et al., 2005
). In heterologous cells, DPP6 and DPP10 can interact with all three Kv4 proteins, however, in-situ hybridization studies showed that the two DPPL genes tended to be expressed in different neurons, with neuronal populations expressing mainly Kv4.2 containing DPP6 mRNAs, while DPP10 was the predominant DPPL in cells expressing Kv4.3, although exceptions to this pattern were noted (Nadal et al., 2006
; Zagha et al., 2005
). We therefore first focused our analysis on a comparison of the pattern of expression of DPP6 and Kv4.2.
DPP6 immunostaining was distributed throughout the mouse brain. The three areas most prominently labeled—the striatum, the hippocampus and the cerebellar cortex—were the same as those most strongly labeled with antibodies to the Kv4.2 pore-forming subunits (Figure 3
). Strong to moderate staining for both proteins was also observed in the neocortex, globus pallidus (Figure 3
), pontine nucleus (Figure 4
) and the medial habenula (data not shown).
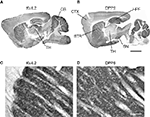
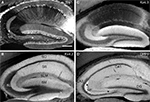
Figure 3. DPP6 and Kv4.2 are expressed in similar structures in mouse brain. (A,B) Both Kv4.2 and DPP6 proteins are widely distributed throughout the mouse brain. The three structures most prominently labeled with Kv4.2 antibodies (A) the striatum (STR), hippocampus (HPF) and cerebellar (CB) cortex, are prominently labeled by antibodies to DPP6 (B). (C,D) Magnification of the Kv4.2 (C) and DPP6 (D) immunostaining of the striatum. As previously reported for antibodies to Kv4.2 and KChIPs (Rhodes et al., 2004
) the labeling is concentrated in the neuropil. Neuronal somata are weakly labeled and are not apparent. All images are immunoperoxidase staining. Scale bar: 2 mm (A,B); 100 μm (C,D).
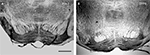
Figure 4. Immunolocalization of Kv4.2 and DPP6 proteins in pontine nuclei. Kv4.2 antibodies (A) and DPP6 antibodies (B) both prominently labeled the neuropil of the pontine nuclei. Immunoperoxidase staining. Scale bar: 250 μm.
However, the regional expression patterns of both proteins were not identical. For example, there is moderate to strong Kv4.2 staining of the thalamus (TH), however this area was weakly labeled with the DPP6 antibodies (Figure 3
). Thalamic nuclei contain both Kv4.2 and Kv4.3 (Serodio and Rudy, 1998
) and strongly express DPP10 mRNAs (Jerng et al., 2007
; Zagha et al., 2005
). It is thus possible that DPP10 is the preferred DPPL subunit of Kv4 channels in this brain area. Alternatively, it is possible that Kv4 channels in some neuronal populations may lack DPPLs as appears to be the case in rodent heart (Radicke et al., 2005
). In addition, as discussed later, some structures apparently lacking Kv4.2 expressed DPP6 protein. Next, we analyzed in more detail the three brain areas where Kv4 protein localization and function have been best characterized.
Kv4.2 and DPP6 Proteins are Expressed in the Same Neuronal Populations and are Enriched in the Same Subcellular Compartments
In the CA1 field of the hippocampus, ISA channels have been shown to be concentrated in pyramidal cell dendrites (Hoffman et al., 1997
; Johnston et al., 2000
). Consistent with this observation, Kv4.2 proteins are most prominently expressed in the dendritic fields of CA1–CA3 pyramidal cells and dentate granule cells (Jinno et al., 2005
; Kim et al., 2005
; Rhodes et al., 2004
; Sheng et al., 1992
). A predominant dendritic localization of Kv4.2 proteins has also been observed in other neuronal populations such as in cerebellar granule cells (Rhodes et al., 2004
; Sheng et al., 1992
; Strassle et al., 2005
). DPP6 was also predominantly dendritic in these neurons (Figures 5–
8
).
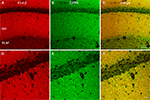
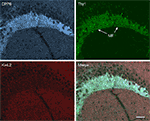
Figure 5. Immunolocalization of Kv4 and DPP6 in the mouse hippocampus. (B–D) Immunofluorescent staining for Kv4.2, Kv4.3 and DPP6 is concentrated in dendritic fields, including the apical and basal dendrites of CA pyramidal cells and the apical dendritic fields of the granule cells in the dentate gyrus (DG). The somata and dendrites of GABAergic interneurons are stained with Kv4.3, but not with Kv4.2 or DPP6 antibodies. The mossy fibers (MF) are labeled with antibodies to DPP6, but not with antibodies to Kv4.2 or Kv4.3. (A) As reference, the hippocampus of a Thy1-H YFP mouse (Feng et al., 2000
) expressing YFP in a subset of pyramidal cells in the CA1–CA3 fields and granule cells of the DG is shown. Note the labeled cell bodies and dendrites of pyramidal and granule cells. The axons of the granule cells (MF) are also YFP labeled. Scale bar: 200 μm. SO, stratum oriens; SR, stratum radiatum; SM, stratum moleculare; SL, stratum lucidum; SLM, stratum lacunosum moleculare.
Figure 6. Enrichment of Kv4.2 and DPP6 in CA1 pyramidal cell dendrites. Confocal image of the CA1 field of the hippocampus in mouse double immunostained for Kv4.2 (A, pseudocolored red) and DPP6 (B, pseudocolored green) and the overlay of the two channels (C). Strong staining of the stratum radiatum (SR) was evident with both antibodies, however DPP6 antibodies produced stronger staining in the SLM than Kv4.2. (D–F) Higher magnification confocal image of CA1. Strong staining with both antibodies is observed in pyramidal cells’ apical and basal dendrites, while pyramidal cell somata are only weakly stained. Scale bar: 50 μm (A–C); 20 μm (D–F).
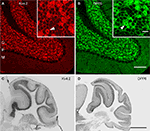
Figure 7. Immunolocalization of Kv4.2 and DPP6 proteins in the cerebellar cortex. Confocal fluorescent images of the cerebellar cortex single labeled with antibodies to Kv4.2 (A, pseudocolored red) or DPP6 (B, pseudocolored green). Immunostaining for both Kv4.2 and DPP6 is prominent in the granule cell layer (G), while the molecular layer (M) is weakly stained in both instances. Also, neither antibody labeled Purkinje cells (P). The insets show high magnifications of the granule cell layer demonstrating high density labeling of the cerebellar glomeruli (arrowheads). Granule cell somata are more weakly stained, and the label is restricted to the periphery of the cell. Scale bars: 100 μm for main panels in (A) and (B); 10 μm for the insets in (A) and (B). (C,D) Immunoperoxidase staining showing anterior-posterior gradient of expression of Kv4.2 and DPP6 proteins in the granule cell layer in lateral cerebellar lobules. Shown are sagittal sections of the mouse cerebellum immunostained with antibodies to Kv4.2 (C) or DPP6 (D). Immunolabeling for both, Kv4.2 and DPP6, is more prominent in anterior than posterior lobules. Scale bar: 1 mm for (C) and (D).
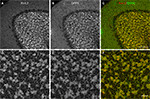
Figure 8. Double-labeling of Cerebellum with Kv4.2 and DPP6 antibodies. Low magnification of cerebellum showing confocal immumofluorescence staining of Kv4.2 (A), DPP6 (B), and the overlay of the two images (C). (D–F) High magnification confocal images of granule cell layer double-labeled with Kv4.2 (D) and DPP6 (E) antibodies showing colocalization in cerebellar granule cells. Scale bar: 100 μm in upper panels; 20 μm in lower panels.
Hippocampus
There was dense immunoreactivity for both Kv4.2 and DPP6 in the hippocampal formation, and overall, both proteins produced similar staining patterns (Figure 5
). As previously reported (Rhodes et al., 2004
), in the CA1–CA3 subfields immunoreactivity for Kv4.2 was concentrated in the stratum oriens and stratum radiatum (Figure 5
B), which contain the basal and apical dendritic arbors of pyramidal cells, respectively (Figure 5
A). The dendrites of CA1–CA3 pyramidal cells were also prominently labeled with DPP6 antibodies (Figure 5
D). By comparison, the somata of the pyramidal cells were only faintly labeled for both Kv4.2 and DPP6 (Figures 5
B,D; see also Figure 6
). There are high levels of expression of Kv4.2 and DPP6 mRNAs in these cells (Nadal et al., 2006
; Serodio and Rudy, 1998
; Zagha et al., 2005
), indicating that most of the protein is transported to the dendrites in these neurons.
Rhodes et al. (2004)
reported that the staining for Kv4.2 in CA1 apical dendrites is not uniform, but drops significantly in the distal portion of the dendrite producing weak staining of stratum lacunosum moleculare (SLM). DPP6 staining also drops in the CA1 distal apical dendritic field, however, SLM staining is stronger for DPP6 than for Kv4.2 (Figures 5 and 6
A–C).
In the dentate gyrus, immunoreactivity for both Kv4.2 and DPP6 was concentrated in the molecular layer (Figures 5
B,D), where the dendrites of dentate granule cells are located (Figure 5
A). As in the CA subfields there was weak staining for these proteins in the somata of the dentate granule cells (Figures 5
B,D).
However, there was one major difference between the Kv4.2 and the DPP6 immunolabeling of the hippocampal formation. There was little or no staining for Kv4.2 proteins in the stratum lucidum in the CA3 subfield (Figure 5
B), but this region was strongly stained with antibodies to DPP6 (Figure 5
D). This difference is significant and will be discussed further below.
Cerebellum
In the cerebellar cortex both Kv4.2 and DPP6 were expressed predominantly in the granule cell layer. There was no staining of Purkinje cells or the interneurons in the molecular layer (Figure 7
). In the granule cell layer there is a gradient of expression of Kv4.2 and Kv4.3 mRNAs. Kv4.2 is much less prominent in posterio-lateral lobules than in the rest of the cerebellar cortex. These lobules express instead mainly Kv4.3 transcripts (Amarillo et al., 2008
; Serodio and Rudy, 1998
; Strassle et al., 2005
). Interestingly, DPP6 labeling showed a similar gradient of expression as Kv4.2 (Figures 7
C,D).
In the granule cells both Kv4.2 and DPP6 proteins were enriched in granule cell dendrites, producing strong staining of the cerebellar glomeruli, where the dendrites of the granule cells contact the unstained cerebellar mossy fiber axons (Figures 7
A,B and 8
). There was weaker but clear staining of the granule cell somata, which was limited to the periphery of the cell (Figures 7
A,B and 8
).
Striatum
In the caudate/putamen, immunoreactivity for Kv4.2 and DPP6 was concentrated in the neuropil, as previously reported for Kv4.2 in rat (Rhodes et al., 2004
; Figure 3
), where the staining is likely to be associated with the somata and dendrites of striatal projection neurons (Rhodes et al., 2004
). Although the density of the staining precludes association to a specific cell type or subcellular domain, the pattern of fiber labeling is clearly the same for Kv4.2 as for DPP6.
Coexpression of DPP6 and Kv4.3 Proteins
Kv4.3 proteins are co-expressed with Kv4.2 in several of the neuronal populations examined above. In the hippocampus for example, Kv4.3 mRNA is expressed in CA2 and CA3 pyramidal cells, but not in CA1 (Serodio and Rudy, 1998
). Consistent with this, Kv4.3 immunoreactivity in rat hippocampus is observed in the basal and apical dendrites of CA2 and CA3 pyramidal neurons but not in the CA1 subfield (Rhodes et al., 2004
), as we were able to confirm in our studies (Figure 5
C). In CA1, immunoreactivity for Kv4.3 is concentrated exclusively in interneurons (Rhodes et al., 2004
; Figure 5
C). DPP6 was prominently expressed in the basal and apical dendrites of all CA subfields (Figure 5
D). Thus the presence of Kv4.3 proteins in Kv4.2-expressing neurons does not preclude DPP6 expression.
On the other hand, in other neurons such as Purkinje cells in the cerebellar cortex and hippocampal interneurons, which express Kv4.3 but not Kv4.2 (Menegola and Trimmer, 2006
; Rhodes et al., 2004
; Serodio and Rudy, 1998
; Strassle et al., 2005
), there was no immunolabeling for DPP6 proteins (Figures 5 and 7
), consistent with the results of DPP6 in-situ hybridization (Nadal et al., 2006
; Zagha et al., 2005
). As discussed earlier, the analysis of mRNA distribution suggests that instead, the Kv4 channels in these neurons are mainly associated with DPP10 (Zagha et al., 2005
).
However, the in situ hybridization studies also showed that DPP6 mRNAs were present in some neurons expressing predominantly Kv4.3 rather than Kv4.2, most notably in neurons in the substancia nigra (Nadal et al., 2006
; Serodio and Rudy, 1998
). Consistent with the mRNA distribution studies, DPP6, but not Kv4.2, was observed in the substantia nigra (Figure 3
).
Kv4-Unrelated DPP6 Expression
So far the analysis shows that Kv4 and DPP6 proteins are expressed in the same brain regions, cell types and neuronal structures, consistent with the view that they are components of the same channel complex and suggesting that this is the main function of DPP6 proteins in brain. However, in one brain structure lacking Kv4.2 or Kv4.3 proteins there was prominent expression of DPP6, the hippocampal mossy fiber (Figures 5
B–D), the axons of the granule cells of the dentate gyrus (Figure 5
A). The entire mossy fiber was immunolabeled with DPP6 antibodies, however the staining was particularly strong in the terminal fields in the stratum lucidum of the CA3 subfield (Figure 5
D). We confirmed that the strong DPP6 immunoreactivity in stratum lucidum was associated with the mossy fiber by double staining with antibodies to DPP6 and Kv4.2 in mouse tissue in which the mossy fiber axons are YFP-labeled (Figure 9
). The immunolabeling of the mossy fiber with DPP6 antibodies appears to be specific, since it was completely absent in knockout mouse brains (Figure 2
). Hence, our observations raise the intriguing possibility that DPP6 may have Kv4-unrelated functions in brain (see Discussion).
Figure 9. DPP6, but not Kv4.2, is expressed in the hippocampal mossy fibers. Confocal images of the hippocampus of a Thy1-H mouse expressing YFP (green) in the mossy fibers (MF) double labeled for Kv4.2 (red) and DPP6 (light blue). The three images are shown superimposed in the lower right panel (merge). Note the strong immunolabeling of the mossy fibers in the stratum lucidum with DPP6 antibodies and lack of staining with antibodies to Kv4.2. Scale bar: 50 μm.
Kv4 channels mediate most of the somato-dendritic, subthreshold-operating transient, or A-type K+ currents (ISA) in neurons. These currents have fundamental roles in neuronal function. They are important in the control of the frequency of repetitive firing, subthreshold excitability, signal processing in dendrites and spike timing-dependent plasticity (Chen and Johnston, 2006
; Connor and Stevens, 1971
; Hoffman et al., 1997
; Johnston et al., 2003
; Kim et al., 2007
; Liss et al., 2001
; Ramakers and Storm, 2002
; Schoppa and Westbrook, 1999
) Kv4 channels also have medical importance, as they have been associated with epilepsy and pain regulation (Bernard et al., 2004
; Hu et al., 2006
; Singh et al., 2006
). These functions rely on the precise subcellular distribution, voltage dependence and kinetic properties of the underlying K+ channels, all of which depend on the molecular composition of the channels. Hence, knowledge of the molecular components of Kv4 channels is critical to understanding their function, their modulation and their participation in disease.
Immunoprecipitation of native Kv4 channels from brain led to the suggestion that these channels are molecular complexes that include in addition to Kv4 pore-forming subunits, the associated proteins KChIPs and the dipeptidyl peptidase-like protein DPP6 (Nadal et al., 2003
). The observation that DPP6, as well as the products of a closely related gene, DPP10, modify the properties of Kv4 channels expressed in heterologous cells producing channels that more closely resemble native channels in neurons, has provided additional support to the hypothesis that these DPPs are molecular components of Kv4 channels in neurons (Amarillo et al., 2008
; Jerng et al., 2005
, 2007
;Nadal et al., 2003
; Takimoto et al., 2006
; Zagha et al., 2005
). Furthermore, we recently showed that treatment of hippocampal neurons with an siRNA to DPP6 modified somatic currents as expected from the results of heterologous expression (Kim et al., 2008
).
However, there are several issues that compel the need to obtain independent validation that DPPs are components of Kv4 channels in brain. First, the finding that DPP6 and DPP10 associate and modulate Kv4 channels is surprising, since all other known DPPs are enzymes and there is no precedent for these proteins functioning as modulators of other membrane proteins. Second, the interaction between DPP6 and Kv4 subunits appears to be relatively weak, and in the experiments of Nadal et al. (2003)
cross-linking of channel complexes was necessary to stabilize their association during solubilization.
Moreover, several other proteins have been observed to modify Kv4 channels in heterologous cells, including minK, various members of the MiRP family, NCS-1 (frequenin), Kvβs and the Na+ channel β1 subunit (Birnbaum et al., 2004
; Deschenes and Tomaselli, 2002
). However, none of these proteins has been validated as an associated protein of native Kv4 channels in neurons, suggesting that caution is required when extrapolating results from heterologous expression experiments with over-expressed proteins.
In this study we used recombinant DPP6 to raise antibodies against this protein suitable for immunolocalization in brain tissue, and asked whether DPP6 proteins are found in the same neuronal populations and subcellular compartments as the pore-forming subunits. Distribution studies provide a global view of protein localization and can be critical to infer or predict function. This is especially true in the brain given its complex cellular makeup and regional specializations.
We found remarkable similarities in the regional and cellular patterns of expression of Kv4.2 and DPP6 proteins. In those neurons where the subcellular pattern of expression of Kv4.2 proteins has been analyzed and shown to have a predominantly dendritic localization, DPP6 was also found predominantly in the dendritic compartment. In one neuronal population, the cerebellar granule cells, where there is an anterior–posterior gradient of expression of Kv4.2, there was also a corresponding gradient of expression of DPP6.
Given the large number of possible combinations, it is unlikely that these similarities in expression patterns of Kv4.2 and DPP6 could arise by chance. Therefore the findings from immunolocalization provide strong additional support to the notion that DPP6 is a component of Kv4 channels in neurons, and suggest that this is the main function of this DPP in brain.
However, we found one structure, the hippocampal mossy fibers, which contain the axons of the granule cells of the dendate gyrus, that strongly expressed DPP6 proteins (Figures 5 and 9
). Neither Kv4.2 nor Kv4.3, the two prominent Kv4 subunits in brain, are found in these axons (Figure 5
), and for that matter have not been described in axons in any other CNS neurons. The function of DPP6 proteins in the mossy fibers remains to be elucidated, but our results strongly suggest that in addition to being a Kv4-associated protein, DPP6 may have Kv4-unrelated functions in some neuronal populations. The results suggest that in addition to postsynaptic roles as a Kv4 channel auxiliary subunit, DPP6 proteins have still undetermined presynaptic functions.
We also observed differences in the intensity of the SLM in the hippocampal formation. This partly reflects the fact that background staining is higher with DPP6 than it is with Kv4.2 antibodies. It is also possible that there is DPP6 protein that is not associated with Kv4.2 in the very distal apical dendrites of CA1 neurons, or that the two antibodies have different access to the subcellular compartments where channel complexes are located in distal dendrites. However, given the expression of DPP6 in the mossy fiber, it is also possible that there is DPP6 staining of perforant path axons, although DPP6 mRNA is only moderately to weakly expressed in enthorrinal cortex (Nadal et al., 2006
).
The possibility that DPP6 may have Kv4-unrelated functions in some neurons is reminiscent of observations with one member of the KChIP family, KChIP3, which has been suggested to have Kv4-unrelated functions as a Ca2+-dependent transcriptional regulator (also known as DREAM) (Buxbaum, 2004
; Carrion et al., 1999
; Cheng et al., 2002
; Vogt, 2002
; Zhang and Wang, 2005
) and as a presenilin-interacting protein also known as calsenilin (Buxbaum, 2004
; Buxbaum et al., 1998
; Lilliehook et al., 2003
). Additional studies are required to elucidate the role of DPP6 in the mossy fibers as this is clearly important to fully understand the function of DPP6 in the normal and pathological brain.
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supported by National Institutes of Health Grant NS045217 to BR and NINDS NRSA grant F30NS63761 to BDC. We thank Axel Brunger and his laboratory for providing purified DPP6 protein.
Cheng, H. Y., Pitcher, G. M., Laviolette, S. R., Whishaw, I. Q., Tong, K. I., Kockeritz, L. K., Wada, T., Joza, N. A., Crackower, M., Goncalves, J., Sarosi, I., Woodgett, J. R., Oliveira-dos-Santos, A. J., Ikura, M., van der Kooy, D., Salter, M. W., and Penninger, J. M. (2002). DREAM is a critical transcriptional repressor for pain modulation. Cell 108, 31–43.
Marshall, C. R., Noor, A., Vincent, J. B., Lionel, A. C., Feuk, L., Skaug, J., Shago, M., Moessner, R., Pinto, D., Ren, Y., Thiruvahindrapduram, B., Fiebig, A., Schreiber, S., Friedman, J., Ketelaars, C. E., Vos, Y. J., Ficicioglu, C., Kirkpatrick, S., Nicolson, R., Sloman, L., Summers, A., Gibbons, C. A., Teebi, A., Chitayat, D., Weksberg, R., Thompson, A., Vardy, C., Crosbie, V., Luscombe, S., Baatjes, R., Zwaigenbaum, L., Roberts, W., Fernandez, B., Szatmari, P., and Scherer, S. W. (2008). Structural variation of chromosomes in autism spectrum disorder. Am. J. Hum. Genet. 82, 477–488.
Rhodes, K. J., Carroll, K. I., Sung, M. A., Doliveira, L. C., Monaghan, M. M., Burke, S. L., Strassle, B. W., Buchwalder, L., Menegola, M., Cao, J., An, W. F., and Trimmer, J. S. (2004). KChIPs and Kv4 alpha subunits as integral components of A-type potassium channels in mammalian brain. J. Neurosci. 24, 7903–7915.
Sebat, J., Lakshmi, B., Malhotra, D., Troge, J., Lese-Martin, C., Walsh, T., Yamrom, B., Yoon, S., Krasnitz, A., Kendall, J., Leotta, A., Pai, D., Zhang, R., Lee, Y. H., Hicks, J., Spence, S. J., Lee, A. T., Puura, K., Lehtimaki, T., Ledbetter, D., Gregersen, P. K., Bregman, J., Sutcliffe, J. S., Jobanputra, V., Chung, W., Warburton, D., King, M. C., Skuse, D., Geschwind, D. H., Gilliam, T. C., Ye, K., and Wigler, M. (2007). Strong association of de novo copy number mutations with autism. Science 316, 445–449.
van Es, M. A., van Vught, P. W., Blauw, H. M., Franke, L., Saris, C. G., Van den Bosch, L., de Jong, S. W., de Jong, V., Baas, F., van’t Slot, R., Lemmens, R., Schelhaas, H. J., Birve, A., Sleegers, K., Van Broeckhoven, C., Schymick, J. C., Traynor, B. J., Wokke, J. H., Wijmenga, C., Robberecht, W., Andersen, P. M., Veldink, J. H., Ophoff, R. A., and van den Berg, L. H. (2008). Genetic variation in DPP6 is associated with susceptibility to amyotrophic lateral sclerosis. Nat. Genet. 40, 29–31.