- 1Department of Chemistry, University of California at Davis, Davis, CA, USA
- 2Basic Sciences, Pennsylvania College of Optometry, Salus University, Elkins Park, PA, USA
Neuronal calcium sensor (NCS) proteins, a sub-branch of the calmodulin superfamily, are expressed in the brain and retina where they transduce calcium signals and are genetically linked to degenerative diseases. The amino acid sequences of NCS proteins are highly conserved but their physiological functions are quite different. Retinal recoverin controls Ca2+-dependent inactivation of light-excited rhodopsin during phototransduction, guanylyl cyclase activating proteins 1 and 2 (GCAP1 and GCAP2) promote Ca2+-dependent activation of retinal guanylyl cyclases, and neuronal frequenin (NCS-1) modulates synaptic activity and neuronal secretion. Here we review the molecular structures of myristoylated forms of NCS-1, recoverin, and GCAP1 that all look very different, suggesting that the attached myristoyl group helps to refold these highly homologous proteins into different three-dimensional folds. Ca2+-binding to both recoverin and NCS-1 cause large protein conformational changes that ejects the covalently attached myristoyl group into the solvent exterior and promotes membrane targeting (Ca2+-myristoyl switch). The GCAP proteins undergo much smaller Ca2+-induced conformational changes and do not possess a Ca2+-myristoyl switch. Recent structures of GCAP1 in both its activator and Ca2+-bound inhibitory states will be discussed to understand structural determinants that control their Ca2+-dependent activation of retinal guanylyl cyclases.
Introduction
Intracellular calcium ions (Ca2+) regulate neuronal signaling in the central nervous system (Berridge et al., 2000; Augustine et al., 2003). Neuronal Ca2+ signals are detected by a family of neuronal calcium sensor (NCS) proteins (Ames et al., 1996, 2012; Braunewell and Gundelfinger, 1999; Burgoyne and Weiss, 2001; Burgoyne et al., 2004; Weiss et al., 2010) that contain EF-hand motifs (Moncrief et al., 1990; Ikura, 1996; Ikura and Ames, 2006) as well as by a family of C2-domain containing proteins (synaptotagmin and protein kinase C isoforms; Nalefski and Falke, 1996; Corbalan and Gomez, 2014). At least sixteen different NCS proteins are known (Weiss and Burgoyne, 2002; Burgoyne and Haynes, 2012) and are conserved from yeast to humans (Figure 1). Recoverin (Dizhoor et al., 1991) and guanylyl cyclase activating proteins 1 and 2 (GCAP1 and GCAP2; Dizhoor et al., 1994; Palczewski et al., 1994) are expressed in the retina, where they regulate phototransduction in photoreceptor cells (Palczewski et al., 2000; Ames and Ikura, 2002; Stephen et al., 2008; Ames et al., 2012). NCS proteins are also expressed in the brain such as neurocalcin (Hidaka and Okazaki, 1993), frequenin (NCS-1; Pongs et al., 1993; McFerran et al., 1998), visinin-like proteins (VILIPs; Bernstein et al., 1999; Braunewell and Klein-Szanto, 2009), K+ channel interacting proteins (KChIPs; An et al., 2000), calsenilin/DREAM (Buxbaum et al., 1998; Carrion et al., 1999), and hippocalcin (Kobayashi et al., 1992, 1993; Tzingounis et al., 2007).
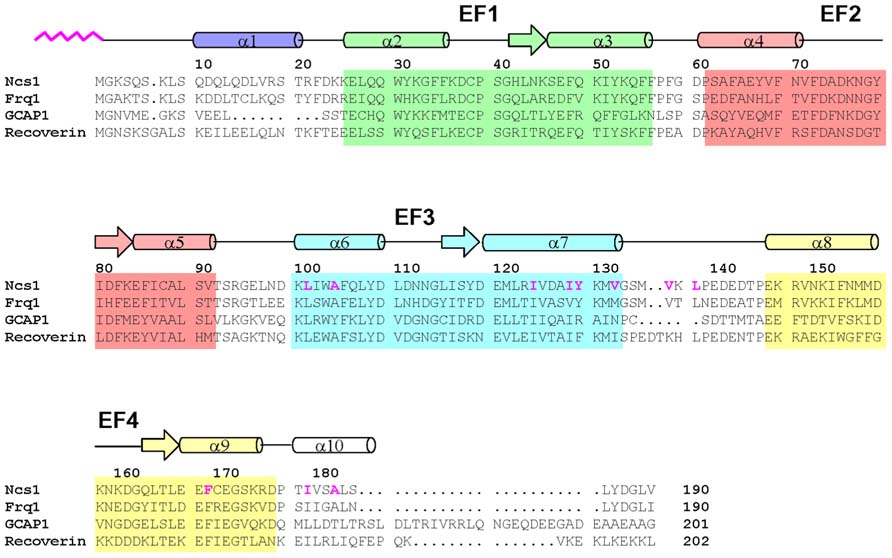
FIGURE 1. Amino acid sequence alignment of selected NCS proteins (sequence numbering is for S. pombe NCS-1). Secondary structure elements (helices and strands), EF-hand motifs (EF1 green, EF2 red, EF3 cyan, and EF4 yellow), and residues that interact with the myristoyl group (highlighted magenta) are indicated. Swiss Protein Database accession numbers are Q09711 (S. pombe NCS-1), Q06389 (S. cerevisiae Frq1), P21457 (bovine recoverin), and P43080 (human GCAP1).
Recoverin also called S-modulin (Dizhoor et al., 1991; Kawamura and Murakami, 1991), the first NCS protein to be discovered, controls the lifetime of photo-excited rhodopsin (Kawamura, 1993; Erickson et al., 1998; Makino et al., 2004) by regulating rhodopsin kinase (Calvert et al., 1995; Chen et al., 1995; Klenchin et al., 1995; Komolov et al., 2009). Recoverin decreases the lifetime of rhodopsin at low Ca2+ levels to control visual recovery and promote photoreceptor adaptation to background light. More recent evidence indicates that recoverin can also modulate the decay of the light-activated phsophodiesterase activity. Such modulation may help accelerate visual recovery in the presence of background light (Chen et al., 2012). Recoverin is also located in the rod inner segment (Strissel et al., 2005) and is associated with cancer-associated retinopathy (Polans et al., 1991; Subramanian and Polans, 2004).
Guanylyl cyclase activating proteins 1 and 2 are also expressed in photoreceptor cells where they activate retinal guanylyl cyclase at low cytosolic Ca2+ levels upon light activation (Dizhoor et al., 1994; Palczewski et al., 1994, 2004). The EF-hand motifs in GCAPs can bind both Mg2+ and Ca2+ (Peshenko and Dizhoor, 2004, 2006). Mg2+ binding stabilizes a structural form of GCAPs that activates cyclase activity (Peshenko and Dizhoor, 2006; Lim et al., 2009), whereas Ca2+-bound GCAPs inhibit the cyclase (Dizhoor and Hurley, 1996; Dizhoor et al., 1998). GCAPs are important for regulating the recovery phase of visual excitation and particular mutants are linked to various forms of retinal degeneration (Semple-Rowland et al., 1996; Sokal et al., 1998; Baehr and Palczewski, 2007; Bondarenko et al., 2010; Jiang and Baehr, 2010).
Neuronal calcium sensor proteins (frequenin or NCS-1) are expressed in other tissues beside the brain (Kapp et al., 2003) and in lower organisms including flies (Pongs et al., 1993), worms (Gomez et al., 2001), and yeast (Frq1; Hendricks et al., 1999; Huttner et al., 2003; Hamasaki et al., 2004). Yeast NCS homologs (called Frq1) activate a phosphatidyl inositol 4-OH kinase isoform (Pik1; Hendricks et al., 1999; Kapp et al., 2003; Strahl et al., 2003, 2007) required for vesicle trafficking and secretion (Hama et al., 1999; Walch-Solimena and Novick, 1999). Mammalian NCS-1 interacts with voltage-gated Ca2+ and K+ channels (Weiss et al., 2000; Nakamura et al., 2001) and activates inositol trisphosphate receptors (Boehmerle et al., 2006).
NCS proteins typically contain about 200 amino acid residues in chain length with four EF-hand motifs, a first EF-hand that does not bind Ca2+, and a myristoylation consensus sequence at the N-terminus. NCS proteins have similar sequences, ranging from 35 to 60% identity (Figure 1). EF-hand residues are the most highly conserved, particularly in the Ca2+ binding loops. The fourth EF-hand sequence is variable, and Ca2+ is able to bind to EF4 in frequenin (Cox et al., 1994; Ames et al., 2000) and GCAPs (Peshenko and Dizhoor, 2007; Stephen et al., 2007) but Ca2+ does not bind to EF4 in recoverin (Ames et al., 1995a) and VILIPs (Cox et al., 1994; Li et al., 2011). Ca2+-binding to EF4 in GCAP1 controls whether GCAP1 can activate or inhibit guanylyl cyclase (Peshenko and Dizhoor, 2007). The residues near the C-terminus and linker between EF3 and EF4 are non-conserved, suggesting that these regions may play a role in target specificity for recoverin but not for GCAPs.
Retinal recoverin and most other NCS proteins are myristoylated at the amino terminus (Dizhoor et al., 1992; Kobayashi et al., 1993; Ladant, 1995). Recoverin and GCAPs contain a saturated myristoyl (14:0) or related fatty acyl group (12:0, 14:1, 14:2), because N-myristoyl transferase (Gordon et al., 1991) can efficiently utilize C12 and/or C14 acetyl-CoA as fatty acyl donors in the retina. In tissues other than the retina, myristoylation is the predominant modification. Myristoylated recoverin binds to cell membranes only at high Ca2+ levels (Zozulya and Stryer, 1992; Dizhoor et al., 1993), whereas unmyristoylated recoverin does not bind to membranes. Likewise, bovine neurocalcin (Ladant, 1995) and hippocalcin (Kobayashi et al., 1993) both are myristoylated and exhibit Ca2+-induced localization at the plasma membrane in response to neuronal stimulation. Ca2+-induced membrane targeting by NCS proteins has been termed, Ca2+-myristoyl switch. The attached fatty acyl group is buried inside the protein structure of Ca2+-free recoverin (Tanaka et al., 1995). Ca2+ binding to recoverin causes extrusion of the fatty acid, enabling it to interact with lipid bilayer membranes. Recoverin’s Ca2+-myristoyl switch may control its light-induced movement into the rod inner segment (Strissel et al., 2005). GCAP proteins are also myristoylated (Palczewski et al., 1994; Frins et al., 1996; Olshevskaya et al., 1997). However, unlike recoverin, GCAPs do not possess a functional Ca2+-myristoyl switch (Olshevskaya et al., 1997; Hwang and Koch, 2002). Instead the N-terminal myristoyl group remains sequestered inside GCAP1 in both Ca2+-free and Ca2+-bound states (Hughes et al., 1998; Lim et al., 2009). Indeed, the crystal structure of Ca2+-bound GCAP1 shows the myristoyl group surrounded by the protein (Stephen et al., 2007), and a recent nuclear magnetic resonance (NMR) structural analysis reveals that the activator state of GCAP1 has an overall structure similar to that of Ca2+-bound inhibitory state in which the N-terminal myristoyl group is buried in both the Ca2+-free and Ca2+-bound states (Lim et al., 2013).
Atomic-resolution structures are known for myristoylated forms of recoverin (Ames et al., 1997), GCAP1 (Stephen et al., 2007), and NCS-1 (Lim et al., 2011) that each fold differently around the attached myristoyl group (Figure 2). For NCS-1, the attached myristoyl group is located in a protein crevice formed by helices from EF3 and EF4 near the C-terminus (Figure 2A). The covalently attached fatty acyl group in NCS-1 protrudes in a parallel fashion between four helices from EF3 and EF4 (Figure 2B). The C-terminal location of the myristoyl binding site in NCS-1 is quite different from that of recoverin in which the myristate is positioned inside a cavity near the N-terminus (Figure 2C). The attached fatty acyl chain in recoverin is wedged between the helices of EF1 and EF2 in a perpendicular fashion (Figure 2D), which contrasts with the parallel arrangement of the fatty acyl chain in NCS-1 (Figure 2B). For GCAP1, the myristoyl group is sequestered in a cavity formed by the N-terminal domain with participation of a C-terminal helix (Figure 2E). The myristoyl group in GCAP1 bridges the N-terminal and C-terminal ends of the protein by contacting helices at each end (Figure 2F). In short, the protein structural environment around the myristoyl group is quite different in recoverin, GCAP1 and NCS-1 (Figure 2). This suggests that each NCS protein folds differently around the N-terminal myristoyl group by contacting non-conserved patches of hydrophobic residues that are unique to each NCS protein. However, myristoylation of GCAP2 is not essential for its ability to activate its target (retinal guanylyl cyclase), because unmyristoylated GCAP2 can activate cyclase activity nearly as well as myristoylated GCAP2 (Olshevskaya et al., 1997).
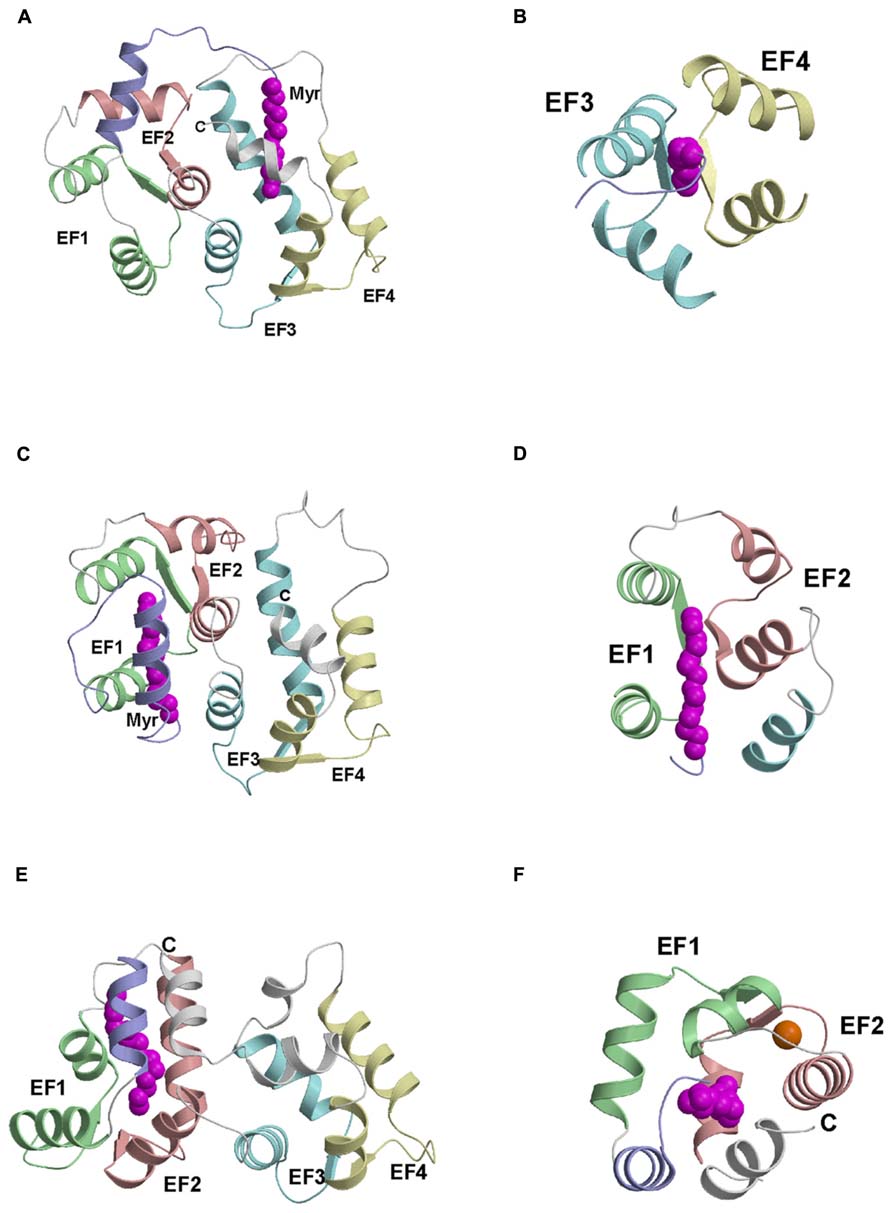
FIGURE 2. Main chain structures of Ca2+-free myrisoylated NCS-1 (PDB ID: 212e) (A), recoverin (PDB ID: 1iku) (C), and GCAP1 (PDB ID: 2r2i) (E). Close-up views of the myristate binding pocket in NCS-1 (B), recoverin (D) and GCAP1 (F). EF-hands and myristoyl group (magenta) are colored as defined in Figure 1. Adapted from and originally published by Lim et al. (2011).
In this review, we discuss recent structures of GCAP1 in both Ca2+-free activator and Ca2+-bound inhibitor states to understand structural determinants that control Ca2+-dependent activation of retinal guanylyl cyclases.
Ca2+-Dependent Activation of Retinal Guanylyl Cyclase
Guanylyl Cyclase Activation and Photoreceptor Recovery
In vertebrate rods and cones, photon absorption by rhodopsin or cone visual pigments triggers a phototransduction cascade that hydrolyzes cGMP, resulting in the closure of cGMP-gated cation channels on the plasma membrane that causes membrane hyperpolarization [see reviews (Pugh et al., 1997, 1999)]. To reset the resting dark-state of retinal rods (known as visual recovery), cGMP levels are replenished very quickly (Burns and Baylor, 2002) by retina-specific guanylyl cyclases (RetGCs), a membrane enzyme present in rod and cone outer segments (Dizhoor et al., 1994; Lowe et al., 1995). RetGC is a Ca2+-regulated enzyme (Koch and Stryer, 1988; Koutalos and Yau, 1996) whose activity is controlled by intracellular domains (Laura et al., 1996; Duda et al., 2005) that interact with soluble EF-hand Ca2+ sensor proteins, called guanylyl cyclase activating proteins (GCAPs: GCAP1 and GCAP2; Dizhoor et al., 1994, 1995; Gorczyca et al., 1994, 1995; Koutalos et al., 1995).
Light-induced channel closure in photoreceptor cells causes a decrease in the cytosolic free Ca2+ concentration (Gray-Keller and Detwiler, 1994), in mammals from 250 nM in the dark to 25 nM in the light (Woodruff et al., 2002). The catalytic activity of RetGC in the dark is negatively controlled by Ca2+-bound GCAPs (Dizhoor and Hurley, 1996; Dizhoor et al., 1998; Burns and Baylor, 2002), whereas the release of Ca2+ from GCAPs at low Ca2+ levels in light-activated photoreceptors causes activation of RetGC (Dizhoor et al., 1994, 1995; Gorczyca et al., 1994, 1995; Dizhoor and Hurley, 1996; Mendez et al., 2001). Light stimulation of the rod cell causes a more than 10-fold increase in cGMP production due to the activation of RetGC by GCAPs (Hodgkin and Nunn, 1988; Burns and Baylor, 2002) and is a critical step for controlling the recovery rate of a single-photon response (Pugh et al., 1999; Burns and Baylor, 2002) as well as the cone response to stronger light stimuli (Sakurai et al., 2011). In mouse rods, GCAPs have been demonstrated to have different Ca2+ sensitivities (Dizhoor et al., 1998; Hwang et al., 2003) and therefore contribute to the recovery by activating guanylyl cyclase at different steps of excitation and recovery, thus imparting proper recovery kinetics to the rod response (Mendez et al., 2001; Makino et al., 2008, 2012).
Mg2+ and Ca2+ Binding to GCAPs Reciprocally Control Cyclase Activation
Guanylyl cyclase activating proteins activate RetGC at low Ca2+ levels (less than 50 nM) and only in the presence of physiological Mg2+ levels (Peshenko and Dizhoor, 2004, 2006, 2007; Dizhoor et al., 2010). This Mg2+ requirement for RetGC activation by GCAPs initially suggested that Mg2+ binding to GCAPs might be important for their activation of RetGC. Indeed, Mg2+ was shown to bind directly to at least two of the EF-hands in GCAP1 (Lim et al., 2009), and NMR studies showed that Mg2+ binding to GCAP1 at EF2 and EF3 was needed to stabilize the overall tertiary fold of the protein (Lim et al., 2009). By contrast, the Ca2+-free/Mg2+-free GCAP1 (apo-state) forms a molten globule-like structure, that contains regular secondary structure (Dell’Orco et al., 2010) but does not form a stable tertiary fold (Peshenko and Dizhoor, 2004; Lim et al., 2009). The flexible and unstructured molten-globule apo-protein could explain in part why GCAPs do not activate RetGC in the absence of Mg2+ (Dizhoor et al., 2010). Thus, Mg2+ binding to GCAP1 stablizes its protein structure in order to bind and activate RetGC (Dizhoor et al., 1994; Peshenko and Dizhoor, 2004), whereas Ca2+ binding to GCAP1 stabilizes a distinct structure important for the inhibition of RetGC (Dizhoor et al., 1998).
The four EF-hands in the GCAPs have quite distinct divalent metal binding properties that control whether GCAPs can activate or inhibit RetGC. The first EF-hand (EF1) does not bind to either Ca2+ or Mg2+ because the residue at the 3-position in the EF-hand binding loop (Cys29 in GCAP1, see Figure 1) is not suitable for ligating either Ca2+ or Mg2+. Ca2+ binds to GCAP1 at the other three EF-hands (EF2, EF3, and EF4) in an independent fashion (Lim et al., 2009) in contrast to the cooperative binding of two Ca2+ to recoverin (Ames et al., 1995a). The apparent dissociation constant for Ca2+ binding to GCAPs is in the submicromolar range (Lim et al., 2009; Dizhoor et al., 2010), whereas Mg2+ binds with ~1000-fold lower affinity (Gifford et al., 2007) in the sub-millimolar range (Peshenko and Dizhoor, 2004, 2007; Lim et al., 2009). These binding affinities imply that three Ca2+ bind per mole of GCAP1 in dark-adapted rod cells, which have relatively high cytosolic Ca2+ levels [Ca2+]free = 250–500 nM [(Woodruff et al., 2002; Matthews and Fain, 2003) and [Mg2+] ~1 mM (Chen et al., 2003)]. Light-activation of the rod cell causes a dramatic lowering of the cytosolic Ca2+ level [Ca2+]free = 5–50 nM (Gray-Keller and Detwiler, 1994; Sampath et al., 1998; Woodruff et al., 2002) while the Mg2+ level remains fixed at [Mg2+]free ~1 mM (Chen et al., 2003). Therefore in light-adapted rods, GCAPs do not bind Ca2+ but instead bind to at least two Mg2+. Thus, Ca2+-free/Mg2+-bound GCAPs activate RetGC in light exposed rods (Dizhoor et al., 1994, 1995; Gorczyca et al., 1995), whereas Ca2+-bound GCAPs (with Ca2+ bound at EF2, EF3, and EF4) inhibit RetGC in dark-adapted rods (Dizhoor and Hurley, 1996; Dizhoor et al., 1998).
Constitutively Active Mutants of GCAP1 Cause Retinal Disease
Various point mutations in the EF-hand motifs of GCAP1 that weaken Ca2+ binding (but do not affect Mg2+ binding) cause GCAP1 to constitutively activate RetGC in rods and cones, which is genetically linked to various retinal diseases (Jiang and Baehr, 2010). These mutations in the EF-hand motifs [Y99C (Dizhoor et al., 1998; Payne et al., 1998) and E155G (Wilkie et al., 2000, 2001)] weaken the Ca2+ binding affinity beyond the photoreceptor Ca2+ concentration and cause the Ca2+-free/Mg2+-bound GCAP1 activator state to persist even at high Ca2+ levels in dark-adapted rods, which causes persistent activation of RetGC (Sokal et al., 1998; Olshevskaya et al., 2004). The GCAP mutants that constitutively activate RetGC then cause elevated cGMP levels in photoreceptor cells that promotes apoptosis and disease (Olshevskaya et al., 2004, 2012; Woodruff et al., 2007).
Mutagenesis studies of the individual EF-hands in GCAP1 have revealed that Ca2+-binding to EF4 is critical for controlling Ca2+-dependent activation of RetGC (Peshenko and Dizhoor, 2007). Mutants of GCAP1 that weaken or abolish Ca2+ binding to EF4 but retain Ca2+ binding at EF2 and EF3 [D144N/D148G (Peshenko and Dizhoor, 2007 and E155G Wilkie et al., 2000, 2001)] are constitutively active even in the presence of high Ca2+ levels in dark-adapted photoreceptors. Furthermore, these mutants that disable Ca2+ binding to EF4 (but not EF2 and EF3) are unable to inhibit RetGC at high Ca2+ levels in the dark (Peshenko and Dizhoor, 2007). In summary, GCAP1 can activate RetGC even if Ca2+ is bound to EF2 and EF3 (but not bound to EF4). Also, Ca2+ binding to EF4 is essential for having Ca2+-induced inhibition of RetGC. Therefore, Ca2+ binding to EF4 is critical for controlling whether GCAP1 can activate or inhibit RetGC.
GCAPs do not Possess a Ca2+-Myristoyl Switch
Ca2+ binding to GCAP1 and GCAP2 does not cause ejection of the covalently attached myristoyl group (Hughes et al., 1998; Lim et al., 2009, 2013) and Ca2+ binding to GCAPs do not promote their membrane targeting (Olshevskaya et al., 1997; Hwang and Koch, 2002). This is in stark contrast to the Ca2+-induced exposure of the N-terminal myristoyl group [termed Ca2+-myristoyl switch (Zozulya and Stryer, 1992; Dizhoor et al., 1993)] that promotes membrane targeting of recoverin (Zozulya and Stryer, 1992; Dizhoor et al., 1993; Valentine et al., 2003), neurocalcin (Ladant, 1995), hippocalcin (O’Callaghan et al., 2003), VILIPs (Spilker et al., 1997, 2002), and NCS-1 (Hamasaki et al., 2004). Instead, the covalently attached myristoyl group of GCAP1 remains sequestered inside the protein hydrophobic core in both Ca2+-free and Ca2+-bound forms of GCAP1 (Lim et al., 2009, 2013). NMR studies on the myristate attached to recoverin (Ames et al., 1995b, 1997; Hughes et al., 1995), VILIP1 (Li et al., 2011), and NCS-1 (Lim et al., 2011) reveal that the covalently attached myristoyl group is buried inside these proteins only in the Ca2+-free state. Ca2+-binding causes protein conformational changes that lead to exposure of the fatty acyl chain in recoverin (Ames et al., 1997), VILIP1 (Li et al., 2011), and NCS-1 (Lim et al., 2011). By stark contrast, NMR studies on GCAP1 indicate that the covalently attached myristoyl group is buried inside both Ca2+-free and Ca2+-bound GCAP1 (Lim et al., 2009, 2013). Further evidence that GCAP1 lacks a Ca2+-myristoyl switch comes from SPR studies that show myristoylation of GCAP1 has little effect on membrane binding (Hwang and Koch, 2002). Finally, the recent atomic level structures of GCAP1 directly demonstrated that the myristoyl group is buried inside the protein in a similar environment in both the Ca2+-free activator (Lim et al., 2013) and Ca2+-bound inhibitor states (Stephen et al., 2007).
Solid-state NMR and other spectroscopic studies on GCAP2 have suggested that Ca2+-free GCAP2 might be targeted to cell membranes by a reversed Ca2+-myristoyl switch (Theisgen et al., 2010, 2011). The covalently attached fatty acyl group has been suggested to become exposed in Ca2+-free GCAP2 in the presence of lipid bilayer membranes (Theisgen et al., 2011), in contrast to having the myristoyl group sequestered inside Ca2+-bound GCAP2 (Schroder et al., 2011). However, other studies on GCAP2 indicate that the myristoyl group remains sequestered inside the protein environment in both Ca2+-free and Ca2+-bound GCAP2 (Hughes et al., 1998). Also, unmyristoylated GCAP2 activates RetGC with nearly the same potency as that of myristoylated GCAP2, and myristoylation of GCAP2 is not essential for its targeting to the membrane-bound cyclase (Olshevskaya et al., 1997; Hwang and Koch, 2002).
Ca2+-Induced Protein Conformational Changes in GCAPs
Atomic level structures are known for Ca2+-bound forms of GCAP1 (Stephen et al., 2007) and GCAP2 (Ames et al., 1999). The four EF-hands in GCAP1 (Figures 1 and 3) are grouped into two globular domains: N-domain includes EF1 and EF2 (residues 18–83) and C-domain contains EF3 and EF4 (residues 88–161). Ca2+ is bound to GCAP1 at EF2, EF3 and EF4, and the structure of each Ca2+-bound EF-hand in GCAP1 (Figure 3B) adopts the familiar open conformation of EF-hands as seen in calmodulin and other Ca2+-bound EF-hand proteins (Ikura, 1996). Indeed, the interhelical angles for each Ca2+-bound EF-hand in GCAP1 are similar to those of recoverin (Ames et al., 1997) and NCS-1 (Bourne et al., 2001). Although the internal structure of each EF-hand in GCAP1 is similar to that of recoverin, the overall three-dimensional packing arrangement and spatial organization of the four EF-hands is very different for GCAP1 vs. recoverin. Indeed, the overall root-mean-squared deviation of main chain atoms is 3.4 Å when comparing the structures of GCAP1 and recoverin (Figure 2). A unique structural feature of GCAP1 is that the N-terminal α-helix (residues 5–15) upstream of EF1 and C-terminal helix (residues 175–183) downstream of EF4 are held closely together by their mutual interaction with the N-terminal myristoyl group (Figure 2F). Thus, the covalently attached myristoyl group in GCAP1 is sequestered within a unique environment inside the Ca2+-bound protein, which is quite different from that of recoverin as described above in Figure 2. The myristoyl group attached to GCAP1 makes contacts with N-terminal residues (V9, L12, and F42) and the C-terminal helix (L174, V178 and I181; Figure 2F). In essence, the myristoyl group serves to bridge both the N-terminal and C-terminal ends of the protein, which explains how Ca2+-induced conformational changes in the C-terminal domain (particularly in EF4) might be transmitted to a possible target binding site in EF1. A Ca2+-myristoyl tug mechanism (Peshenko et al., 2012) has been proposed to explain how Ca2+-induced conformational changes in EF4 serve to “tug” on the adjacent C-terminal helix that connects structurally to the myristoyl group and EF1. This tug mechanism helps explain how Ca2+-induced structural changes in EF4 might be relayed to the cyclase binding region in EF1 (Lim et al., 2013). The Ca2+-induced structural changes involving the C-terminal helix might also be related to Ca2+-dependent phosphorylation of S201 in GCAP2 (Peshenko et al., 2004).
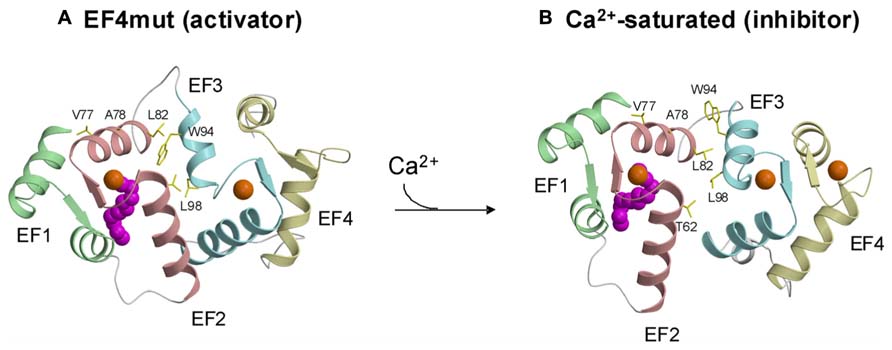
FIGURE 3. Structures of activator (A) vs. inhibitor forms (B) of GCAP1 adapted from and originally published by Lim et al. (2013).
The atomic level structure of Ca2+-free/Mg2+-bound activator form of GCAP1 or GCAP2 is currently not known. The main difficulty is that Ca2+-free/Mg2+-bound GCAP proteins form dimers and higher order protein oligomers that causes considerable sample heterogeneity at high protein concentrations needed for NMR or to make crystals for X-ray crystallography (Ames et al., 1999; Lim et al., 2009, 2013). Ca2+-dependent dimerization of GCAP2 has been suggested to be important for activating the cyclase (Olshevskaya et al., 1999b; Ermilov et al., 2001). Protein dimerization was also reported for GCAP1, and a GCAP1 mutant (V77E) that prevents protein dimerization also abolishes its ability to activate RetGC, suggesting that dimerization of Ca2+-free/Mg2+-bound activator state might be important for activating RetGC (Lim et al., 2013). However, protein dimerization of GCAP1 is not Ca2+ dependent and only occurs at relatively high protein concentrations above 100 μ M (Lim et al., 2013). Future studies are needed to investigate whether GCAPs might form a functional dimer upon binding to the dimeric RetGC to form a 2:2 complex (Ramamurthy et al., 2001; Peshenko et al., 2010), and whether the dimeric quaternary structure may play a regulatory role.
A GCAP1 mutant, D144N/D148G called EF4mut, (Lim et al., 2013) that binds Ca2+ at EF2 and EF3 (and does not bind Ca2+ at EF4) can activate RetGC at high Ca2+ concentrations (Peshenko and Dizhoor, 2007). Therefore, EF4mut (with Ca2+ bound at EF2 and EF3) serves as a model of the Ca2+-free/Mg2+-bound activator state. The EF4mut protein is more soluble and stable than Ca2+-free/Mg2+-bound wild type, and EF4mut exhibits NMR spectra with much higher resolution and sensitivity compared to that of Ca2+-free/Mg2+-bound wild type (Lim et al., 2009, 2013). NMR structural studies on EF4mut provide some insights for a structural model of the GCAP1 activator state (Figure 3A). The overall structure of EF4mut is similar to the crystal structure of Ca2+-bound GCAP1 (root mean squared deviation of main chain atoms is 1.3 Å when comparing the two structures). However, residues at the domain interface (between EF2 and EF3) are structured somewhat differently in EF4mut compared to the crystal structure of Ca2+-bound GCAP1. Many of the GCAP1 residues at the domain interface have quite broad NMR resonances, suggesting that these residues are conformationally dynamic (Lim et al., 2013). The corresponding residues in recoverin (Figure 1) also exhibited broad NMR resonances and 15N NMR relaxation dispersion studies reveal that these domain interface residues exhibit millisecond exchange kinetics (Xu et al., 2011). Ca2+-induced rearrangement of residues at the domain interface in recoverin gives rise to a 45° swiveling of the two domains (Ames et al., 1997). A structural comparison between EF4mut and Ca2+-bound GCAP1 suggests a related but much smaller Ca2+-induced structural change at the domain interface in GCAP1 (Figure 3). The most noteworthy Ca2+-induced structural difference in GCAP1 can be seen in the entering helix of EF3 that unravels a half turn in EF4mut, which causes a repositioning of the W94 side-chain at the domain interface. A Ca2+-induced change in the structural environment around W94 is consistent with previous tryptophan fluorescence and electron paramagnetic resonance studies of GCAP1 (Sokal et al., 2001; Peshenko and Dizhoor, 2006, 2007). We suggest that Ca2+-induced rearrangement of residues at the domain interface (V77, A78, L82, K85, and W94) plays a role in modulating Ca2+-dependent contacts with RetGC1.
Activation Mechanism for RetGC by GCAPs
The structural information for the GCAPs above provides insights into the activation mechanism of RetGC (Figure 4). GCAP1 residues in EF1 (Krylov et al., 1999; Olshevskaya et al., 1999a,b; Ermilov et al., 2001) and at the domain interface (Krylov et al., 1999; Lim et al., 2013; Peshenko et al., 2014) are suggested to make direct contact with RetGC (see labeled residues in Figure 4). In our model (Figure 4), Ca2+-induced conformational changes in EF4 (Lim et al., 2013) are transmitted to the cyclase binding site (see labeled residues in EF1, EF2, and EF3, Figure 4) by a Ca2+-myristoyl tug mechanism as described by (Peshenko et al., 2012). In the GCAP1 activator state under physiological conditions (Figure 4, left panel), EF2 and EF3 are bound to Mg2+ (blue circles in Figure 4) with EF1 and EF4 unoccupied. The Ca2+-free state of EF4 forms a loose and dynamic structure (Lim et al., 2013), which allows the adjacent C-terminal helix to reach all the way to the N-terminal myristoyl group (magenta in Figure 4) and thus indirectly form hydrophobic contacts with residues in EF1 and EF2. In essence, the myristoyl group forms a bridge between Ca2+-induced conformational changes in the C-terminal domain and the cyclase binding site in the N-terminal domain. In the GCAP1 activator state, residues in the cyclase binding site [see Figure 4, labeled residues in EF1, EF2 and EF3 (Peshenko et al., 2014)] are spatially close together and form particular contacts with RetGC that require close proximity between W94 and V77 (see arrow in Figure 4). In the Ca2+-bound GCAP1 inhibitor state (Figure 4, right panel), Ca2+ is bound to EF2, EF3, and EF4 (orange circles in Figure 4). Ca2+-binding to EF4 causes local conformational changes that in turn “tug” on the C-terminal helix which causes a slight reorientation of the N-terminal domain (EF1 and EF2) with respect to the C-terminal domain (EF3 and EF4). This domain swiveling causes key residues in EF3 at the domain interface (K85, W94, and K97) to move farther away from cyclase binding site residues in EF2 [F73, V77, and A78 (Peshenko et al., 2014)], which disrupts key contacts to RetGC that we suggest may be important for regulating cyclase activation.
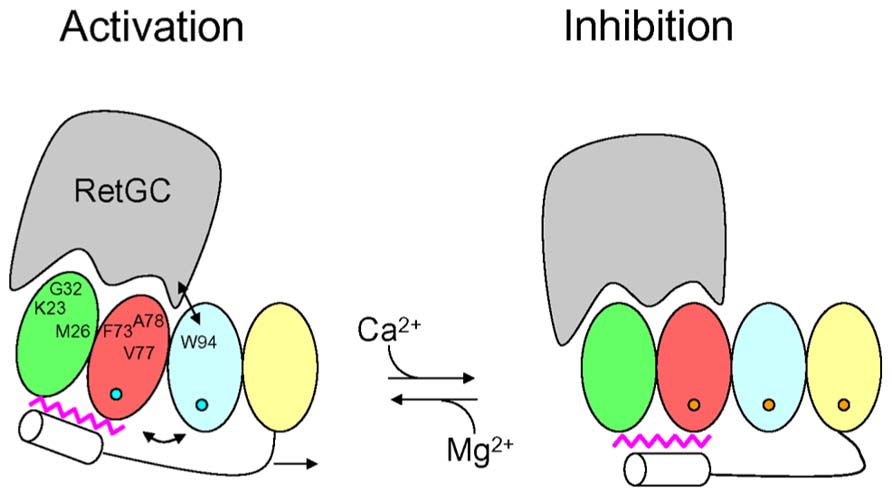
FIGURE 4. Schematic diagram of retinal guanylyl cyclase activation by GCAP1. RetGC is shown in gray. The four EF-hands in GCAP1 are colored coded as defined in Figure 1, bound Mg2+ are depicted by blue circles, bound Ca2+ are orange circles, the C-terminal helix is white, and N-terminal myristoyl group is magenta. The labeled GCAP1 residues (K23, M26, G32, F73, V77, A78, and W94) have been implicated in the RetGC binding site (Lim et al., 2013; Peshenko et al., 2014). The binding of Ca2+ at EF4 causes conformational changes that are transmitted to the RetGC binding site by a Ca2+-myristoyl tug mechanism (see arrows) modified from Peshenko et al. (2012).
Conclusion
N-terminal myristoylation serves to remodel the structure of NCS proteins as seen for recoverin, GCAP1 and NCS-1. Each NCS protein folds differently around the attached myristoyl group, which causes each NCS protein to adopt a unique structure. Most NCS proteins possess a functional Ca2+-myristoyl switch that promotes their Ca2+-induced membrane targeting. By contrast, GCAP1 contains a sequestered myristoyl group in both its Ca2+-free and Ca2+-bound states and undergoes rather small Ca2+-induced protein conformational changes. Ca2+ binding to the fourth EF-hand in GCAP1 triggers conformational changes in the N-terminal domain via a Ca2+-myristoyl tug mechanism that controls the exposure of residues in EF1 and EF2 that we propose may serve as a target binding site for guanylyl cyclase.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by grants to James B. Ames (EY012347) and Alexander M. Dizhoor (EY11522) from the National Institutes of Health.
References
Ames, J. B., and Ikura, M. (2002). Structure and membrane-targeting mechanism of retinal Ca2+-binding proteins, recoverin and GCAP-2. Adv. Exp. Med. Biol. 514, 333–348. doi: 10.1007/978-1-4615-0121-3_20
Ames, J. B., Dizhoor, A. M., Ikura, M., Palczewski, K., and Stryer, L. (1999). Three-dimensional structure of guanylyl cyclase activating protein-2, a calcium-sensitive modulator of photoreceptor guanylyl cyclases. J. Biol. Chem. 274, 19329–19337. doi: 10.1074/jbc.274.27.19329
Ames, J. B., Hendricks, K. B., Strahl, T., Huttner, I. G., Hamasaki, N., and Thorner, J. (2000). Structure and calcium-binding properties of Frq1, a novel calcium sensor in the yeast Saccharomyces cerevisiae. Biochemistry 39, 12149–12161. doi: 10.1021/bi0012890
Ames, J. B., Ishima, R., Tanaka, T., Gordon, J. I., Stryer, L., and Ikura, M. (1997). Molecular mechanics of calcium-myristoyl switches. Nature 389, 198–202. doi: 10.1038/38310
Ames, J. B., Lim, S., and Ikura, M. (2012). Molecular structure and target recognition of neuronal calcium sensor proteins. Front. Mol. Neurosci. 5:10. doi: 10.3389/fnmol.2012.00010
Ames, J. B., Porumb, T., Tanaka, T., Ikura, M., and Stryer, L. (1995a). Amino-terminal myristoylation induces cooperative calcium binding to recoverin. J. Biol. Chem. 270, 4526–4533. doi: 10.1074/jbc.270.9.4526
Ames, J. B., Tanaka, T., Ikura, M., and Stryer, L. (1995b). Nuclear magnetic resonance evidence for Ca(2+)-induced extrusion of the myristoyl group of recoverin. J. Biol. Chem. 270, 30909–30913. doi: 10.1074/jbc.270.52.30909
Ames, J. B., Tanaka, T., Stryer, L., and Ikura, M. (1996). Portrait of a myristoyl switch protein. Curr. Opin. Struct. Biol. 6, 432–438. doi: 10.1016/S0959-440X(96)80106-0
An, W. F., Bowlby, M. R., Betty, M., Cao, J., Ling, H. P., Mendoza, G., et al. (2000). Modulation of A-type potassium channels by a family of calcium sensors. Nature 403, 553–556. doi: 10.1038/35000592
Augustine, G. J., Santamaria, F., and Tanaka, K. (2003). Local calcium signaling in neurons. Neuron 40, 331–346. doi: 10.1016/S0896-6273(03)00639-1
Baehr, W., and Palczewski, K. (2007). Guanylate cyclase-activating proteins and retina disease. Subcell. Biochem. 45, 71–91. doi: 10.1007/978-1-4020-6191-2_4
Bernstein, H. G., Baumann, B., Danos, P., Diekmann, S., Bogerts, B., Gundelfinger, E. D., et al. (1999). Regional and cellular distribution of neural visinin-like protein immunoreactivities (VILIP-1 and VILIP-3) in human brain. J. Neurocytol. 28, 655–662. doi: 10.1023/A:1007056731551
Berridge, M. J., Lipp, P., and Bootman, M. D. (2000). The versatility and universality of calcium signalling. Nat. Rev. Mol. Cell Biol. 1, 11–21. doi: 10.1038/35036035
Boehmerle, W., Splittgerber, U., Lazarus, M. B., McKenzie, K. M., Johnston, D. G., Austin, D. J., et al. (2006). Paclitaxel induces calcium oscillations via an inositol 1,4,5-trisphosphate receptor and neuronal calcium sensor 1-dependent mechanism. Proc. Natl. Acad. Sci. U.S.A. 103, 18356–18361. doi: 10.1073/pnas.0607240103
Bondarenko, V. A., Hayashi, F., Usukura, J., and Yamazaki, A. (2010). Involvement of rhodopsin and ATP in the activation of membranous guanylate cyclase in retinal photoreceptor outer segments (ROS-GC) by GC-activating proteins (GCAPs): a new model for ROS-GC activation and its link to retinal diseases. Mol. Cell. Biochem. 334, 125–139. doi: 10.1007/s11010-009-0323-y
Bourne, Y., Dannenberg, J., Pollmann, V. V., Marchot, P., and Pongs, O. (2001). Immunocytochemical localization and crystal structure of human frequenin (neuronal calcium sensor1). J. Biol. Chem. 276, 11949–11955. doi: 10.1074/jbc.M009373200
Braunewell, K. H., and Gundelfinger, E. D. (1999). Intracellular neuronal calcium sensor proteins: a family of EF-hand calcium-binding proteins in search of a function. Cell Tissue Res. 295, 1–12. doi: 10.1007/s004410051207
Braunewell, K. H., and Klein-Szanto, A. J. (2009). Visinin-like proteins (VSNLs): interaction partners and emerging functions in signal transduction of a subfamily of neuronal Ca2+ -sensor proteins. Cell Tissue Res. 335, 301–316. doi: 10.1007/s00441-008-0716-3
Burgoyne, R. D., and Haynes, L. P. (2012). Understanding the physiological roles of the neuronal calcium sensor proteins. Mol. Brain 5, 2. doi: 10.1186/1756-6606-5-2
Burgoyne, R. D., O’Callaghan, D. W., Hasdemir, B., Haynes, L. P., and Tepikin, A. V. (2004). Neuronal Ca2+-sensor proteins: multitalented regulators of neuronal function. Trends Neurosci. 27, 203–209. doi: 10.1016/j.tins.2004.01.010
Burgoyne, R. D., and Weiss, J. L. (2001). The neuronal calcium sensor family of Ca2+-binding proteins. Biochem. J. 353, 1–12. doi: 10.1042/0264-6021:3530001
Burns, M. E., and Baylor, D. A. (2002). Activation, deactivation, and adaptation in vertebrate photoreceptor cells. Annu. Rev. Neurosci. 24, 779–805. doi: 10.1146/annurev.neuro.24.1.779
Buxbaum, J. D., Choi, E. K., Luo, Y., Lilliehook, C., Crowley, A. C., Merriam, D. E., et al. (1998). Calsenilin: a calcium-binding protein that interacts with the presenilins and regulates the levels of a presenilin fragment. Nat. Med. 4, 1177–1181. doi: 10.1038/2673
Calvert, P. D., Klenchin, V. A., and Bownds, M. D. (1995). Rhodopsin kinase inhibition by recoverin. Function of recoverin myristoylation. J. Biol. Chem. 270, 24127–24129. doi: 10.1074/jbc.270.27.16147
Carrion, A. M., Link, W. A., Ledo, F., Mellstrom, B., and Naranjo, J. R. (1999). DREAM is a Ca2+-regulated transcriptional repressor. Nature 398, 80–84. doi: 10.1038/18044
Chen, C. K., Inglese, J., Lefkowitz, R. J., and Hurley, J. B. (1995). Ca(2+)-dependent interaction of recoverin with rhodopsin kinase. J. Biol. Chem. 270, 18060–18066. doi: 10.1074/jbc.270.30.18060
Chen, C. K., Woodruff, M. L., Chen, F. S., Chen, Y., Cilluffo, M. C., Tranchina, D., et al. (2012). Modulation of mouse rod response decay by rhodopsin kinase and recoverin. J. Neurosci. 32, 15998–16006. doi: 10.1523/JNEUROSCI.1639-12.2012
Chen, C., Nakatani, K., and Koutalos, Y. (2003). Free magnesium concentration in salamander photoreceptor outer segments. J. Physiol. 553, 125–135. doi: 10.1113/jphysiol.2003.053280
Corbalan, S., and Gomez, J. C. (2014). Signaling through C2 domains: more than one lipid target. Biochim. Biophys. Acta doi: 10.1016/j.bbamem.2014.01.008 [Epub ahead of print].
Cox, J. A., Durussel, I., Comte, M., Nef, S., Nef, P., Lenz, S. E., et al. (1994). Cation binding and conformational changes in VILIP and NCS-1, two neuron-specific calcium-binding proteins. J. Biol. Chem. 269, 32807–32813.
Dell’Orco, D., Behnen, P., Linse, S., and Koch, K. W. (2010). Calcium binding, structural stability and guanylate cyclase activation in GCAP1 variants associated with human cone dystrophy. Cell. Mol. Life Sci. 67, 973–984. doi: 10.1007/s00018-009-0243-8
Dizhoor, A. M., Boikov, S. G., and Olshevskaya, E. V. (1998). Constitutive activation of photoreceptor guanylate cyclase by Y99C mutant of GCAP-1. Possible role in causing human autosomal dominant cone degeneration. J. Biol. Chem. 273, 17311–17314. doi: 10.1074/jbc.273.28.17311
Dizhoor, A. M., Chen, C. K., Olshevskaya, E., Sinelnikova, V. V., Phillipov, P., and Hurley, J. B. (1993). Role of the acylated amino terminus of recoverin in Ca(2+)-dependent membrane interaction. Science 259, 829–832. doi: 10.1126/science.8430337
Dizhoor, A. M., Ericsson, L. H., Johnson, R. S., Kumar, S., Olshevskaya, E., Zozulya, S., et al. (1992). The NH2 terminus of retinal recoverin is acylated by a small family of fatty acids. J. Biol. Chem. 267, 16033–16036.
Dizhoor, A. M., and Hurley, J. B. (1996). Inactivation of EF-hands makes GCAP-2 (p24) a constitutive activator of photoreceptor guanylyl cyclase by preventing a Ca2+-induced “activator-to-inhibitor” transition. J. Biol. Chem. 271, 19346–19350. doi: 10.1074/jbc.271.32.19346
Dizhoor, A. M., Lowe, D. G., Olsevskaya, E. V., Laura, R. P., and Hurley, J. B. (1994). The human photoreceptor membrane guanylyl cyclase, RetGC, is present in outer segments and is regulated by calcium and a soluble activator. Neuron 12, 1345–1352. doi: 10.1016/0896-6273(94)90449-9
Dizhoor, A. M., Olshevskaya, E. V., and Peshenko, I. V. (2010). Mg2+/Ca2+ cation binding cycle of guanylyl cyclase activating proteins (GCAPs): role in regulation of photoreceptor guanylyl cyclase. Mol. Cell. Biochem. 334, 117–124. doi: 10.1007/s11010-009-0328-6
Dizhoor, A. M., Olshevskaya, E. V., Henzel, W. J., Wong, S. C., Stults, J. T., Ankoudinova, I., et al. (1995). Cloning, sequencing and expression of a 24-kDa Ca2+-binding protein activating photoreceptor guanylyl cyclase. J. Biol. Chem. 270, 25200–25206. doi: 10.1074/jbc.270.42.25200
Dizhoor, A. M., Ray, S., Kumar, S., Niemi, G., Spencer, M., Rrolley, D., et al. (1991). Recoverin: a calcium sensitive activator of retinal rod guanylate cyclase. Science 251, 915–918. doi: 10.1126/science.1672047
Duda, T., Fik, E., Venkataraman, V., Krishnan, R., Koch, K. W., and Sharma, R. K. (2005). The calcium-sensor guanylate cyclase activating protein type 2 specific site in rod outer segment membrane guanylate cyclase type 1. Biochemistry 44, 7336–7345. doi: 10.1021/bi050068x
Erickson, M. A., Lagnado, L., Zozulya, S., Neubert, T. A., Stryer, L., and Baylor, D. A. (1998). The effect of recombinant recoverin on the photoresponse of truncated rod photoreceptors. Proc. Natl. Acad. Sci. U.S.A. 95, 6474–6479. doi: 10.1073/pnas.95.11.6474
Ermilov, A. N., Olshevskaya, E. V., and Dizhoor, A. M. (2001). Instead of binding calcium, one of the EF-hand structures in guanylyl cyclase activating protein-2 is required for targeting photoreceptor guanylyl cyclase. J. Biol. Chem. 276, 48143–48148. doi: 10.1074/jbc.M107539200
Frins, S., Bonigk, W., Muller, F., Kellner, R., and Koch, K. W. (1996). Functional characterization of a guanylyl cyclase activating protein from vertebrate rods. Cloning, heterologous expression, and localization. J. Biol. Chem. 271, 8022–8027. doi: 10.1074/jbc.271.14.8022
Gifford, J. L., Walsh, M. P., and Vogel, H. J. (2007). Structures and metal-ion-binding properties of the Ca2+-binding helix-loop-helix EF-hand motifs. Biochem. J. 405, 199–221. doi: 10.1042/BJ20070255
Gomez, M., De Castro, E., Guarin, E., Sasakura, H., Kuhara, A., Mori, I., et al. (2001). Ca2+ signaling via the neuronal calcium sensor-1 regulates associative learning and memory in C. elegans. Neuron 30, 241–248. doi: 10.1016/S0896-6273(01)00276-8
Gorczyca, W. A., Polans, A. S., Surgucheva, I. G., Subbaraya, I., Baehr, W., and Palczewski, K. (1995). Guanylyl cyclase activating protein. A calcium-sensitive regulator of phototransduction. J. Biol. Chem. 270, 22029–22036. doi: 10.1074/jbc.270.37.22029
Gorczyca, W. A., Van Hooser, J. P., and Palczewski, K. (1994). Nucleotide inhibitors and activators of retinal guanylyl cyclase. Biochemistry 33, 3217–3222. doi: 10.1021/bi00177a011
Gordon, J. I., Duronio, R. J., Rudnick, D. A., Adams, S. P., and Gokel, G. W. (1991). Protein myristoylation. J. Biol. Chem. 266, 8647–8650.
Gray-Keller, M. P., and Detwiler, P. B. (1994). The calcium feedback signal in the phototransduction cascade of vertebrate rods. Neuron 13, 849–861. doi: 10.1016/0896-6273(94)90251-8
Hama, H., Schnieders, E. A., Thorner, J., Takemoto, J. Y., and DeWald, D. B. (1999). Direct involvement of phosphatidylinositol 4-phosphate in secretion in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 274, 34294–34300. doi: 10.1074/jbc.274.48.34294
Hamasaki, N., Molchanova, T., Takeda, K., and Ames, J. B. (2004). Fission yeast homolog of neuronal calcium sensor-1 (Ncs1p) regulates sporulation and confers calcium tolerance. J. Biol. Chem. 279, 12744–12754. doi: 10.1074/jbc.M311895200
Hendricks, K. B., Wang, B. Q., Schnieders, E. A., and Thorner, J. (1999). Yeast homologue of neuronal frequenin is a regulator of phosphatidylinositol-4-OH kinase. Nat. Cell Biol. 1, 234–241. doi: 10.1038/12058
Hidaka, H., and Okazaki, K. (1993). Neurocalcin family: a novel calcium-binding protein abundant in bovine central nervous system. Neurosci. Res. 16, 73–77. doi: 10.1016/0168-0102(93)90074-Z
Hodgkin, A. L., and Nunn, B. J. (1988). Control of light-sensitive current in salamander rods. J. Physiol. 403, 439–471.
Hughes, R. E., Brzovic, P. S., Dizhoor, A. M., Klevit, R. E., and Hurley, J. B. (1998). Ca2+-dependent conformational changes in bovine GCAP-2. Protein Sci. 7, 2675–2680. doi: 10.1002/pro.5560071222
Hughes, R. E., Brzovic, P. S., Klevit, R. E., and Hurley, J. B. (1995). Calcium-dependent solvation of the myristoyl group of recoverin. Biochemistry 34, 11410–11416. doi: 10.1021/bi00036a013
Huttner, I. G., Strahl, T., Osawa, M., King, D. S., Ames, J. B., and Thorner, J. (2003). Molecular interactions of yeast frequenin with Pik1. J. Biol. Chem. 278, 4862–4874. doi: 10.1074/jbc.M207920200
Hwang, J. Y., and Koch, K. W. (2002). Calcium- and myristoyl-dependent properties of guanylate cyclase-activating protein-1 and protein-2. Biochemistry 41, 13021–13028. doi: 10.1021/bi026618y
Hwang, J. Y., Lange, C., Helten, A., Hoppner-Heitmann, D., Duda, T., Sharma, R. K., et al. (2003). Regulatory modes of rod outer segment membrane guanylate cyclase differ in catalytic efficiency and Ca(2+)-sensitivity. Eur. J. Biochem. 270, 3814–3821. doi: 10.1046/j.1432-1033.2003.03770.x
Ikura, M. (1996). Calcium binding and conformational response in EF-hand proteins. Trends Biochem. Sci. 21, 14–17. doi: 10.1016/0968-0004(96)80879-6
Ikura, M., and Ames, J. B. (2006). Genetic polymorphism and protein conformational plasticity in the calmodulin superfamily: two ways to promote multifunctionality. Proc. Natl. Acad. Sci. U.S.A. 103, 1159–1164. doi: 10.1073/pnas.0508640103
Jiang, L., and Baehr, W. (2010). GCAP1 mutations associated with autosomal dominant cone dystrophy. Adv. Exp. Med. Biol. 664, 273–282. doi: 10.1007/978-1-4419-1399-9_31
Kapp, Y., Melnikov, S., Shefler, A., Jeromin, A., and Sagi, R. (2003). NCS-1 and phosphatidylinositol 4-kinase regulate IgE receptor-triggered exocytosis in cultured mast cells. J. Immunol. 171, 5320–5327.
Kawamura, S. (1993). Rhodopsin phosphorylation as a mechanism of cyclic GMP phosphodiesterase regulation by S-modulin. Nature 362, 855–857. doi: 10.1038/362855a0
Kawamura, S., and Murakami, M. (1991). Calcium-dependent regulation of cyclic GMP phosphodiesterase by a protein from frog retinal rods. Nature 349, 420–423. doi: 10.1038/349420a0
Klenchin, V. A., Calvert, P. D., and Bownds, M. D. (1995). Inhibition of rhodopsin kinase by recoverin. Further evidence for a negative feedback system in phototransduction. J. Biol. Chem. 270, 16147–16152. doi: 10.1074/jbc.270.41.24127
Kobayashi, M., Takamatsu, K., Saitoh, S., Miura, M., and Noguchi, T. (1992). Molecular cloning of hippocalcin, a novel calcium-binding protein of the recoverin family exclusively expressed in hippocampus [published erratum appears in Biochem. Biophys. Res. Commun. (1993). 196, 1017]. Biochem. Biophys. Res. Commun. 189, 511–517. doi: 10.1016/0006-291X(92)91587-G
Kobayashi, M., Takamatsu, K., Saitoh, S., and Noguchi, T. (1993). Myristoylation of hippocalcin is linked to its calcium-dependent membrane association properties. J. Biol. Chem. 268, 18898–18904.
Koch, K. W., and Stryer, L. (1988). Highly cooperative feedback control of retinal rod guanylate cyclase by calcium ions. Nature 334, 64–66. doi: 10.1038/334064a0
Komolov, K. E., Senin, I. I., Kovaleva, N. A., Christoph, M. P., Churumova, V. A., Grigoriev, I. I., et al. (2009). Mechanism of rhodopsin kinase regulation by recoverin. J. Neurochem. 110, 72–79. doi: 10.1111/j.1471-4159.2009.06118.x
Koutalos, Y., Nakatani, K., Tamura, T., and Yau, K. W. (1995). Characterization of guanylate cyclase activity in single retinal rod outer segments. J. Gen. Physiol. 106, 863–890. doi: 10.1085/jgp.106.5.863
Koutalos, Y., and Yau, K. W. (1996). Regulation of sensitivity in vertebrate rod photoreceptors by calcium. Trends Neurosci. 19, 73–81. doi: 10.1016/0166-2236(96)89624-X
Krylov, D. M., Niemi, G. A., Dizhoor, A. M., and Hurley, J. B. (1999). Mapping sites in guanylyl cyclase activating protein-1 required for regulation of photoreceptor membrane guanylyl cyclases. J. Biol. Chem. 274, 10833–10839. doi: 10.1074/jbc.274.16.10833
Ladant, D. (1995). Calcium and membrane binding properties of bovine neurocalcin expressed in Escherichia coli. J. Biol. Chem. 270, 3179–3185.
Laura, R. P., Dizhoor, A. M., and Hurley, J. B. (1996). The membrane guanylyl cyclase, retinal guanylyl cyclase-1, is activated through its intracellular domain. J. Biol. Chem. 271, 11646–11651. doi: 10.1074/jbc.271.20.11646
Li, C., Pan, W., Braunewell, K. H., and Ames, J. B. (2011). Structural analysis of Mg2+ and Ca2+ binding, myristoylation, and dimerization of the neuronal calcium sensor and visinin-like protein 1 (VILIP-1). J. Biol. Chem. 286, 6354–6366. doi: 10.1074/jbc.M110.173724
Lim, S., Peshenko, I. V., Dizhoor, A. M., and Ames, J. B. (2009). Effects of Ca2+, Mg2+, and myristoylation on guanylyl cyclase activating protein 1 structure and stability. Biochemistry 48, 850–862. doi: 10.1021/bi801897p
Lim, S., Peshenko, I. V., Dizhoor, A. M., and Ames, J. B. (2013). Structural insights for activation of retinal guanylate cyclase by GCAP1. PLoS ONE 8:e81822. doi: 10.1371/journal.pone.0081822
Lim, S., Strahl, T., Thorner, J., and Ames, J. B. (2011). Structure of a Ca2+- myristoyl switch protein that controls activation of a phosphatidylinositol 4-kinase in fission yeast. J. Biol. Chem. 286, 12565–12577. doi: 10.1074/jbc.M110.208868
Lowe, D. G., Dizhoor, A. M., Liu, K., Gu, Q., Spencer, M., Laura, R., et al. (1995). Cloning and expression of a second photoreceptor-specific membrane retina guanylyl cyclase (RetGC), RetGC-2. Proc. Natl. Acad. Sci. U.S.A. 6, 5535–5539. doi: 10.1073/pnas.92.12.5535
Makino, C. L., Dodd, R. L., Chen, J., Burns, M. E., Roca, A., Simon, M. I., et al. (2004). Recoverin regulates light-dependent phosphodiesterase activity in retinal rods. J. Gen. Physiol. 123, 729–741. doi: 10.1085/jgp.200308994
Makino, C. L., Peshenko, I. V., Wen, X. H., Olshevskaya, E. V., Barrett, R., and Dizhoor, A. M. (2008). A role for GCAP2 in regulating the photoresponse. Guanylyl cyclase activation and rod electrophysiology in GUCA1B knock-out mice. J. Biol. Chem. 283, 29135–29143. doi: 10.1074/jbc.M804445200
Makino, C. L., Wen, X. H., Olshevskaya, E. V., Peshenko, I. V., Savchenko, A. B., and Dizhoor, A. M. (2012). A role for GCAP2 in regulating the photoresponse. Guanylyl cyclase activation and rod electrophysiology in GUCA1B knock-out mice. PLoS ONE 7:e47637. doi: 10.1371/journal.pone.0047637
Matthews, H. R., and Fain, G. L. (2003). The effect of light on outer segment calcium in salamander rods. J. Physiol. 552, 763–776. doi: 10.1113/jphysiol.2003.050724
McFerran, B. W., Graham, M. E., and Burgoyne, R. D. (1998). Neuronal Ca2+ sensor 1, the mammalian homologue of frequenin, is expressed in chromaffin and PC12 cells and regulates neurosecretion from dense-core granules. J. Biol. Chem. 273, 22768–22772. doi: 10.1074/jbc.273.35.22768
Mendez, A., Burns, M. E., Izabela, S., Dizhoor, A. M., Baehr, W., Palczewski, K., et al. (2001). Role of guanylate cyclase-activating proteins (GCAPs) in setting the flash sensitivity of rod photoreceptors. Proc. Natl. Acad. Sci. U.S.A. 98, 9948–9953. doi: 10.1073/pnas.171308998
Moncrief, N. D., Kretsinger, R. H., and Goodman, M. (1990). Evolution of EF-hand calcium-modulated proteins. J. Mol. Evol. 30, 522–562. doi: 10.1007/BF02101108
Nakamura, T. Y., Pountney, D. J., Ozaita, A., Nandi, S., Ueda, S., Rudy, B., et al. (2001). A role for frequenin, a Ca2+-binding protein, as a regulator of Kv4 K+-currents. Proc. Natl. Acad. Sci. U.S.A. 98, 12808–12813. doi: 10.1073/pnas.221168498
Nalefski, E. A., and Falke, J. J. (1996). The C2 domain calcium-binding motif: structural and functional diversity. Protein Sci. 5, 2375–2390. doi: 10.1002/pro.5560051201
O’Callaghan, D. W., Tepikin, A. V., and Burgoyne, R. D. (2003). Dynamics and calcium sensitivity of the Ca2+/myristoyl switch protein hippocalcin in living cells. J. Cell Biol. 163, 715–721. doi: 10.1083/jcb.200306042
Olshevskaya, E. V., Boikov, S., Ermilov, A., Krylov, D., Hurley, J. B., and Dizhoor, A. M. (1999a). Mapping functional domains of the guanylate cyclase regulator protein, GCAP-2. J. Biol. Chem. 274, 10823–10832. doi: 10.1074/jbc.274.16.10823
Olshevskaya, E. V., Ermilov, A. N., and Dizhoor, A. M. (1999b). Dimerization of guanylyl cyclase-activating protein. J. Biol. Chem. 274, 25583–25587. doi: 10.1074/jbc.274.36.25583
Olshevskaya, E. V., Calvert, P. D., Woodruff, M. L., Peshenko, I. V., Savchenko, A. B., Makino, C. L., et al. (2004). The Y99C mutation in guanylyl cyclase-activating protein 1 increases intracellular Ca2+ and causes photoreceptor degeneration in transgenic mice. J. Neurosci. 24, 6078–6085. doi: 10.1523/JNEUROSCI.0963-04.2004
Olshevskaya, E. V., Hughes, R. E., Hurley, J. B., and Dizhoor, A. M. (1997). Calcium binding, but not a calcium-myristoyl switch, controls the ability of guanylyl cyclase-activating protein GCAP-2 to regulate photoreceptor guanylyl cyclase. J. Biol. Chem. 272, 14327–14333. doi: 10.1074/jbc.272.22.14327
Olshevskaya, E. V., Peshenko, I. V., Savchenko, A. B., and Dizhoor, A. M. (2012). Retinal guanylyl cyclase isozyme 1 is the preferential in vivo target for constitutively active GCAP1 mutants causing congenital degeneration of photoreceptors. J. Neurosci. 32, 7208–7217. doi: 10.1523/JNEUROSCI.0976-12.2012
Palczewski, K., Polans, A. S., Baehr, W., and Ames, J. B. (2000). Ca(2+)-binding proteins in the retina: structure, function, and the etiology of human visual diseases. Bioessays 22, 337–350. doi: 10.1002/(SICI)1521-1878(200004)22:4<337::AID-BIES4>3.0.CO;2-Z
Palczewski, K., Sokal, I., and Baehr, W. (2004). Guanylate cyclase-activating proteins: structure, function, and diversity. Biochem. Biophys. Res. Commun. 322, 1123–1130. doi: 10.1016/j.bbrc.2004.07.122
Palczewski, K., Subbaraya, I., Gorczyca, W. A., Helekar, B. S., Ruiz, C. C., Ohguro, H., et al. (1994). Molecular cloning and characterization of retinal photoreceptor guanylyl cyclase-activating protein. Neuron 13, 395–404. doi: 10.1016/0896-6273(94)90355-7
Payne, A. M., Downes, S. M., Bessant, D. A., Taylor, R., Holder, G. E., Warren, M. J., et al. (1998). A mutation in guanylate cyclase activator 1A (GUCA1A) in an autosomal dominant cone dystrophy pedigree mapping to a new locus on chromosome 6p21.1. Hum. Mol. Genet. 7, 273–277. doi: 10.1093/hmg/7.2.273
Peshenko, I. V., and Dizhoor, A. M. (2004). Guanylyl cyclase-activating proteins (GCAPs) are Ca2+/Mg2+ sensors: implications for photoreceptor guanylyl cyclase (RetGC) regulation in mammalian photoreceptors. J. Biol. Chem. 279, 16903–16906. doi: 10.1074/jbc.C400065200
Peshenko, I. V., and Dizhoor, A. M. (2006). Ca2+ and Mg2+ binding properties of GCAP-1. Evidence that Mg2+-bound form is the physiological activator of photoreceptor guanylyl cyclase. J. Biol. Chem. 281, 23830–23841. doi: 10.1074/jbc.M600257200
Peshenko, I. V., and Dizhoor, A. M. (2007). Activation and inhibition of photoreceptor guanylyl cyclase by guanylyl cyclase activating protein 1 (GCAP-1): the functional role of Mg2+/Ca2+ exchange in EF-hand domains. J. Biol. Chem. 282, 21645–21652. doi: 10.1074/jbc.M702368200
Peshenko, I. V., Olshevskaya, E. V., and Dizhoor, A. M. (2004). Ca(2+)-dependent conformational changes in guanylyl cyclase-activating protein 2 (GCAP-2) revealed by site-specific phosphorylation and partial proteolysis. J. Biol. Chem. 279, 50342–50349. doi: 10.1074/jbc.M408683200
Peshenko, I. V., Olshevskaya, E. V., Lim, S., Ames, J. B., and Dizhoor, A. M. (2012). Calcium-myristoyl Tug is a new mechanism for intramolecular tuning of calcium sensitivity and target enzyme interaction for guanylyl cyclase-activating protein 1: dynamic connection between N-fatty acyl group and EF-hand controls calcium sensitivity. J. Biol. Chem. 287, 13972–13984. doi: 10.1074/jbc.M112.341883
Peshenko, I. V., Olshevskaya, E. V., Lim, S., Ames, J. B., and Dizhoor, A. M. (2014). Identification of target binding site in photoreceptor guanylyl cyclase activating protein 1 (GCAP1). J. Biol. Chem. 289. doi: 10.1074/jbc.M113.540716 [Epub ahead of print].
Peshenko, I. V., Olshevskaya, E. V., Yao, S., Ezzeldin, H. H., Pittler, S. J., and Dizhoor, A. M. (2010). Activation of retinal guanylyl cyclase RetGC1 by GCAP1: stoichiometry of binding and effect of new LCA-related mutations. Biochemistry 49, 709–717. doi: 10.1021/bi901495y
Polans, A. S., Buczylko, J., Crabb, J., and Palczewski, K. (1991). A photoreceptor calcium binding protein is recognized by autoantibodies obtained from patients with cancer-associated retinopathy. J. Cell Biol. 112, 981–989. doi: 10.1083/jcb.112.5.981
Pongs, O., Lindemeier, J., Zhu, X. R., Theil, T., Engelkamp, D., Krah-Jentgens, I., et al. (1993). Frequenin-a novel calcium-binding protein that modulates synaptic efficacy. Neuron 11, 15–28. doi: 10.1016/0896-6273(93)90267-U
Pugh, E. N., Duda, T., Sitaramayya, A., and Sharma, R. K. (1997). Photoreceptor guanylate cyclases: a review. Biosci. Rep. 17, 429–473. doi: 10.1023/A:1027365520442
Pugh, E. N., Nikonov, S., and Lamb, T. D. (1999). Molecular mechanisms of vertebrate photoreceptor light adaptation. Curr. Opin. Neurobiol. 9, 410–418. doi: 10.1016/S0959-4388(99)80062-2
Ramamurthy, V., Tucker, C., Wilkie, S. E., Daggett, V., Hunt, D. M., and Hurley, J. B. (2001). Interactions within the coiled-coil domain of RetGC-1 guanylyl cyclase are optimized for regulation rather than for high affinity. J. Biol. Chem. 276, 26218–26229. doi: 10.1074/jbc.M010495200
Sakurai, K., Chen, J., and Kefalov, V. J. (2011). Role of guanylyl cyclase modulation in mouse cone phototransduction. J. Neurosci. 31, 7991–8000. doi: 10.1523/JNEUROSCI.6650-10.2011
Sampath, A. P., Matthews, H. R., Cornwall, M. C., and Fain, G. L. (1998). Bleached pigment produces a maintained decrease in outer segment Ca2+ in salamander rods. J. Gen. Physiol. 111, 53–64. doi: 10.1085/jgp.111.1.53
Schroder, T., Lilie, H., and Lange, C. (2011). The myristoylation of guanylate cyclase-activating protein-2 causes an increase in thermodynamic stability in the presence but not in the absence of Ca2+. Protein Sci. 20, 1155–1165. doi: 10.1002/pro.643
Semple-Rowland, S. L., Gorczyca, W. A., Buczylko, J., Helekar, B. S., Ruiz, C. C., Subbaraya, I., et al. (1996). Expression of GCAP1 and GCAP2 in the retinal degeneration (rd) mutant chicken retina. FEBS Lett. 385, 47–52. doi: 10.1016/0014-5793(96)00345-6
Sokal, I., Li, N., Klug, C. S., Filipek, S., Hubbell, W. L., Baehr, W., et al. (2001). Calcium-sensitive regions of GCAP1 as observed by chemical modifications, fluorescence, and EPR spectroscopies. J. Biol. Chem., 276, 43361–43373. doi: 10.1074/jbc.M103614200
Sokal, I., Li, N., Surgucheva, I., Warren, M. J., Payne, A. M., Bhattacharya, S. S., et al. (1998). GCAP1 (Y99C) mutant is constitutively active in autosomal dominant cone dystrophy. Mol. Cell 2, 129–133. doi: 10.1016/S1097-2765(00)80121-5
Spilker, C., Dresbach, T., and Braunewell, K. H. (2002). Reversible translocation and activity-dependent localization of the calcium-myristoyl switch protein VILIP-1 to different membrane compartments in living hippocampal neurons. J. Neurosci. 22, 7331–7339.
Spilker, C., Gundelfinger, E. D., and Braunewell, K. H. (1997). Calcium- and myristoyl-dependent subcellular localization of the neuronal calcium-binding protein VILIP in transfected PC12 cells. Neurosci. Lett. 225, 126–128. doi: 10.1016/S0304-3940(97)00201-2
Stephen, R., Bereta, G., Golczak, M., Palczewski, K., and Sousa, M. C. (2007). Stabilizing function for myristoyl group revealed by the crystal structure of a neuronal calcium sensor, guanylate cyclase-activating protein 1. Structure 15, 1392–1402. doi: 10.1016/j.str.2007.09.013
Stephen, R., Filipek, S., Palczewski, K., and Sousa, M. C. (2008). Ca2+ -dependent regulation of phototransduction. Photochem. Photobiol. 84, 903–910. doi: 10.1111/j.1751-1097.2008.00323.x
Strahl, T., Grafelmann, B., Dannenberg, J., Thorner, J., and Pongs, O. (2003). Conservation of regulatory function in calcium-binding proteins: human frequenin (neuronal calcium sensor-1) associates productively with yeast phosphatidylinositol 4-kinase isoform, Pik1. J. Biol. Chem. 278, 49589–49599. doi: 10.1074/jbc.M309017200
Strahl, T., Huttner, I. G., Lusin, J. D., Osawa, M., King, D., Thorner, J., et al. (2007). Structural insights into activation of phosphatidylinositol 4-kinase (Pik1) by yeast frequenin (Frq1). J. Biol. Chem. 282, 30949–30959. doi: 10.1074/jbc.M705499200
Strissel, K. J., Lishko, P. V., Trieu, L. H., Kennedy, M. J., Hurley, J. B., and Arshavsky, V. Y. (2005). Recoverin undergoes light-dependent intracellular translocation in rod photoreceptors. J. Biol. Chem. 280, 29250–29255. doi: 10.1074/jbc.M501789200
Subramanian, L., and Polans, A. S. (2004). Cancer-related diseases of the eye: the role of calcium and calcium-binding proteins. Biochem. Biophys. Res. Commun. 322, 1153–1165. doi: 10.1016/j.bbrc.2004.07.109
Tanaka, T., Ames, J. B., Harvey, T. S., Stryer, L., and Ikura, M. (1995). Sequestration of the membrane-targeting myristoyl group of recoverin in the calcium-free state. Nature 376, 444–447. doi: 10.1038/376444a0
Theisgen, S., Scheidt, H. A., Magalhaes, A., Bonagamba, T. J., and Huster, D. (2010). A solid-state NMR study of the structure and dynamics of the myristoylated N-terminus of the guanylate cyclase-activating protein-2. Biochim. Biophys. Acta 1798, 266–274. doi: 10.1016/j.bbamem.2009.06.028
Theisgen, S., Thomas, L., Schroder, T., Lange, C., Kovermann, M., Balbach, J., et al. (2011). The presence of membranes or micelles induces structural changes of the myristoylated guanylate-cyclase activating protein-2. Eur. Biophys. J. 40, 565–576. doi: 10.1007/s00249-011-0680-9
Tzingounis, A. V., Kobayashi, M., Takamatsu, K., and Nicoll, R. A. (2007). Hippocalcin gates the calcium activation of the slow after hyperpolarization in hippocampal pyramidal cells. Neuron 53, 487–493. doi: 10.1016/j.neuron.2007.01.011
Valentine, K. G., Mesleh, M. F., Opella, S. J., Ikura, M., and Ames, J. B. (2003). Structure, topology, and dynamics of myristoylated recoverin bound to phospholipid bilayers. Biochemistry 42, 6333–6340. doi: 10.1021/bi0206816
Walch-Solimena, C., and Novick, P. (1999). The yeast phosphatidylinositol-4-OH kinase Pik1 regulates secretion at the Golgi. Nat. Cell Biol. 1, 523–525. doi: 10.1038/70319
Weiss, J. L., Archer, D. A., and Burgoyne, R. D. (2000). Neuronal Ca2+ sensor-1/frequenin functions in an autocrine pathway regulating Ca2+ channels in bovine adrenal chromaffin cells. J. Biol. Chem. 275, 40082–40087. doi: 10.1074/jbc.M008603200
Weiss, J. L., and Burgoyne, R. D. (2002). “Neuronal calcium sensor proteins,” in Handbook of Cell Signaling, Vol. 2, ed. R. Bradshaw (San Diego: Academic Press), 79–82.
Weiss, J. L., Hui, H., and Burgoyne, R. D. (2010). Neuronal calcium sensor-1 regulation of calcium channels, secretion, and neuronal outgrowth. Cell. Mol. Neurobiol. 30, 1283–1292. doi: 10.1007/s10571-010-9588-7
Wilkie, S. E., Li, Y., Deery, E. C., Newbold, R. J., Garibaldi, D., Bateman, J. B., et al. (2001). Identification and functional consequences of a new mutation (E155G) in the gene for GCAP1 that causes autosomal dominant cone dystrophy. Am. J. Hum. Genet. 69, 471–480. doi: 10.1086/323265
Wilkie, S. E., Newbold, R. J., Deery, E., Walker, C. E., Stinton, I., Ramamurthy, V., et al. (2000). Functional characterization of missense mutations at codon 838 in retinal guanylate cyclase correlates with disease severity in patients with autosomal dominant cone-rod dystrophy. Hum. Mol. Genet. 9, 3065–3073. doi: 10.1093/hmg/9.20.3065
Woodruff, M. L., Olshevskaya, E. V., Savchenko, A. B., Peshenko, I. V., Barrett, R., Bush, R. A., et al. (2007). Constitutive excitation by Gly90Asp rhodopsin rescues rods from degeneration caused by elevated production of cGMP in the dark. J. Neurosci. 27, 8805–8815. doi: 10.1523/JNEUROSCI.2751-07.2007
Woodruff, M. L., Sampath, A. P., Mathews, H. R., Krasnoperova, N. V., Lem, J., and Fain, G. L. (2002). Measurement of cytoplasmic calcium concentration in the rods of wild-type and transducin knock-out mice. J. Physiol. 542, 843–854. doi: 10.1113/jphysiol.2001.013987
Xu, X., Ishima, R., and Ames, J. B. (2011). Conformational dynamics of recoverin’s Ca2+-myristoyl switch probed by 15N NMR relaxation dispersion and chemical shift analysis. Proteins 79, 1910–1922. doi: 10.1002/prot.23014
Keywords: calcium, EF-hand, Ca2+-myristoyl switch, NCS-1, recoverin, GCAP1, NCS protein, NMR
Citation: Lim S, Dizhoor AM and Ames JB (2014) Structural diversity of neuronal calcium sensor proteins and insights for activation of retinal guanylyl cyclase by GCAP1. Front. Mol. Neurosci. 7:19. doi: 10.3389/fnmol.2014.00019
Received: 27 January 2014; Paper pending published: 17 February 2014;
Accepted: 27 February 2014; Published online: 17 March 2014.
Edited by:
Rameshwar K. Sharma, Salus University, USAReviewed by:
Baojin Ding, University of Massachusetts Medical School, USAFrank Schmitz, Saarland University, Germany
Copyright © 2014 Lim, Dizhoor and Ames. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: James B. Ames, Department of Chemistry, University of California at Davis, One Shields Avenue, Davis, CA 95616, USA e-mail:amJhbWVzQHVjZGF2aXMuZWR1
 Sunghyuk Lim
Sunghyuk Lim Alexander M. Dizhoor
Alexander M. Dizhoor James B. Ames
James B. Ames