- 1R&D Division, Kyowa Kirin Co., Ltd., Tokyo, Japan
- 2Department of Endocrinology, Metabolism and Hematology, Tokushima University Graduate School of Medical Sciences, Tokushima, Japan
- 3Department of Molecular Endocrinology, Fujii Memorial Institute of Medical Sciences, Institute of Advanced Medical Sciences, Tokushima University, Tokushima, Japan
Fibroblast growth factor 23 (FGF23), a hormone, mainly produced by osteocytes, regulates phosphate and vitamin D metabolism. By contrast, 1,25-dihydroxyvitamin D3, the active form of vitamin D, has been shown to enhance FGF23 production. While it is likely that osteocytes are heterogenous in terms of gene expression profiles, specific subpopulations of Fgf23-expressing osteocytes have not been identified. Single-cell RNA sequencing (scRNA-seq) technology can characterize the transcriptome of an individual cell. Recently, scRNA-seq has been used for bone tissue analysis. However, owing to technical difficulties associated with isolation of osteocytes, studies using scRNA-seq analysis to characterize FGF23-producing osteocytes are lacking. In this study, we characterized osteocytes secreting FGF23 from murine femurs in response to calcitriol (1,25-dihydroxyvitamin D3) using scRNA-seq. We first detected Dmp1, Mepe, and Phex expression in murine osteocytes by in situ hybridization and used these as marker genes of osteocytes. After decalcification, enzyme digestion, and removal of CD45+ cells, femoral bone cells were subjected to scRNA-seq. We identified cell clusters containing osteocytes using marker gene expression. While Fgf23 expression was observed in some osteocytes isolated from femurs of calcitriol-injected mice, no Fgf23 expression was detected in untreated mice. In addition, the expression of several genes which are known to be changed after 1,25-dihydroxyvitamin D3 treatment such as Ccnd2, Fn1, Igfbp7, Pdgfa, and Timp1 was also affected by calcitriol treatment in Fgf23-expressing osteocytes, but not in those lacking Fgf23 expression, even after calcitriol administration. Furthermore, box-and-whisker plots indicated that Fgf23 expression was observed in osteocytes with higher expression levels of the Fam20c, Dmp1, and Phex genes, whose inactivating mutations have been shown to cause FGF23-related hypophosphatemic diseases. These results indicate that osteocytes are heterogeneous with respect to their responsiveness to 1,25-dihydroxyvitamin D3, and sensitivity to 1,25-dihydroxyvitamin D3 is one of the characteristics of osteocytes with Fgf23 expression. It is likely that there is a subpopulation of osteocytes expressing several genes, including Fgf23, involved in phosphate metabolism.
Introduction
FGF23 has been identified as a gene responsible for autosomal dominant hypophosphatemic rickets (ADHR Consortium, 2000). Fibroblast growth factor 23 (FGF23) has also been identified as a factor responsible for tumor-induced osteomalacia, a rare paraneoplastic syndrome characterized by hypophosphatemia (Shimada et al., 2001). FGF23 has been shown to be physiologically produced mainly by osteocytes (Liu et al., 2003) and to act as a hormone regulating phosphate and vitamin D metabolism (Shimada et al., 2004a; Shimada et al., 2004b). In addition to autosomal dominant hypophosphatemic rickets and tumor-induced osteomalacia, several types of hypophosphatemic rickets/osteomalacia have been shown to be caused by overexpression of FGF23 (Fukumoto and Martin, 2009). For example, X-linked hypophosphatemic rickets is caused by inactivating mutations in the PHEX gene located on the X chromosome, which has homology to a family of endopeptidases (Francis et al., 1995). It is known that the Phex/PHEX is expressed in mouse bones and in human bones, lungs, and ovaries (Du et al., 1996; Beck et al., 1997; Grieff et al., 1997; Guo and Quarles., 1997; Lipman et al., 1998). Autosomal recessive hypophosphatemic rickets 1 is caused by mutations in the DMP1 gene (Feng et al., 2006; Lorenz-Depiereux et al., 2006). DMP1 is a glycoprotein that is highly expressed in the bones and teeth and controls mineralization (George et al., 1993). Raine syndrome is caused by mutations in the FAM20C gene (Simpson et al., 2007). FAM20C mutations were identified during whole-exome sequencing in patients with FGF23-related hypophosphatemia, dental anomalies, and ectopic calcification (Rafaelsen et al., 2013). In the absence of functional PHEX, DMP1, or FAM20C, FGF23 expression is considered to be elevated in osteocytes and in circulation, leading to phosphate excretion from the kidneys and reduction in circulating phosphate (Martin et al., 2011; Rafaelsen et al., 2013; Miyagawa et al., 2014).
Recently, single-cell RNA sequencing (scRNA-seq) technology has been developed to characterize the transcriptomes of individual cells (Macosko et al., 2015; Zheng et al., 2017; Zilionis et al., 2017). This method has been used to map the complex cell diversity (Baryawno et al., 2019; Tikhonova et al., 2019) and identify gene expression signatures related to cell differentiation (Kanton et al., 2019; Wolock et al., 2019). Bone tissue is composed of various cells, and recently several scRNA-seq analyses data of bone cells have been reported (Ayturk et al., 2020; Wang et al., 2021). However, technical difficulties associated with isolation of osteocytes limit scRNA-seq analysis. Additionally, since Fgf23 is normally expressed at considerably low levels in osteocytes (Liu et al., 2006), difficulties in detection of Fgf23 gene expression are expected. Recently, Wang et al. performed scRNA-seq using murine femoral cells and demonstrated defects in osteocyte maturation in the absence of Sp7. They were able to isolate osteocytes using expression of Dmp1 or Mepe, which encodes a bone matrix protein. However, in this study, Fgf23-expressing osteocytes were not detected (Wang et al., 2021).
Several studies have elucidated various signals that regulate FGF23 secretion. A high phosphate diet has been shown to increase FGF23 levels in both mice and humans (Ferrari et al., 2005; Perwad et al., 2005; Antoniucci et al., 2006). In addition to phosphate, 1,25-dihydroxyvitamin D3 has been reported to increase serum FGF23 levels (Saito et al., 2005). Moreover, vitro analysis using MC3T3-E1 osteocyte progenitor cells, 1,25-dihydroxyvitamin D3 induced FGF23 production, and Fgf23 expression in osteocyte-like cells transfected with a knockdown sequence against vitamin D receptor was significantly decreased compared with that in control cells (Yashiro et al., 2020). In this study, we validated the utility of scRNA-seq to quantify the abundance and describe the characteristics of subpopulations of osteocytes from murine femurs. In particular, we focused on the characterization of Fgf23-expressing osteocytes responded to 1,25-dihydroxyvitamin D3.
Materials and methods
Animals and calcitriol injection
Twelve 7-week-old male C57BL/6J mice (The Jackson Laboratory Japan, Inc.) were divided into two groups: untreated and treated with calcitriol (Kyowa Kirin Co., Ltd.). The administration protocol was modified from previous reports (Kolek et al., 2005; Saito et al., 2005). Mice in treated group were intravenously injected with 2.5 μg/kg calcitriol every day for 3 days. Twenty-four hours after the final injection, the mice were euthanized under anesthesia, and femurs were collected for scRNA-seq analysis.
Ethics approval statement
All animal studies were performed in accordance with the Standards for Proper Conduct of Animal Experiments at Kyowa Kirin Co., Ltd., under the approval of the company’s Institutional Animal Care and Use Committee. Tokyo Research Park of Kyowa Kirin Co., Ltd. is fully accredited by AAALAC International.
In situ hybridization
DNA fragments corresponding to murine Dmp1 cDNA (nucleotides 1129–1744), Mepe cDNA (nucleotides 810–1468), and Phex cDNA (nucleotides 239–1161) were individually cloned into pGEM-T vector (Promega K.K.). Sense and antisense probes were labeled with digoxigenin (DIG) using in vitro transcription. Paraffin-embedded blocks of mouse femurs were purchased from Genostaff Co., Ltd. Blocks were cut into 6 μm-thick sections and fixed on glass slides. The sections were acetylated with 0.2% hydrochloric acid and permeabilized with 4 μg/mL proteinase K (FUJIFILM Wako Pure Chemical Corporation) at 37°C for 10 min, with each step followed by two 5 min washes in PBS. Hybridization with DIG-labeled riboprobes was performed at 60°C for 16 h. For mRNA detection, serial sections were used for hybridization with sense and antisense riboprobes. The sections were then washed with 50% formamide in 0.5x G-Wash (Genostaff Co., Ltd.) and blocked with G-Block (Genostaff Co., Ltd.). The hybridized DIG-labeled RNA probes were detected by alkaline phosphatase–conjugated anti-DIG antibody (Roche Diagnostics) in G-Block-TBST and nitro blue tetrazolium chloride/5-bromo-4-chloro-3-indolyl phosphate (NBT/BCIP) substrates (Sigma-Aldrich). The sections were counterstained with nuclear fast red dye (Muto Pure Chemicals Co., Ltd.), fixed with G-Mount (Genostaff Co., Ltd.), cleared in xylene, and mounted using Marinol (Muto Pure Chemicals Co., Ltd.). Samples were imaged using a virtual slide scanner (NanoZoomer S210; Hamamatsu Photonics K.K.) at ×40 magnification. The collected images were adjusted for brightness.
Femoral cell isolation
The femurs were pooled for each group and treated as one sample. Cells were isolated from mouse femurs utilizing a modified protocol derived from previous reports (Stern et al., 2012; Miyagawa et al., 2014). Briefly, epiphyses were cut off, and the marrow was flushed out by centrifugation. Mouse femurs were disrupted by Cryo-Press (Microtec Co., Ltd.) and digested with 1.6 U/mL collagenase D (Roche Diagnostics) in HBSS; pH = 7.8; FUJIFILM Wako Pure Chemical Corporation) for 30 min. The supernatant was discarded. Enzyme treatment and removal of supernatant steps were repeated twice. Residual bone pieces were treated with 20 mM ethylene glycol bis(β-aminoethylether)-N,N,N′,N′-tetraacetic acid disodium salt solution (neutral) (EGTA; Nacalai Tesque, Inc.) in HBSS for 15 min and then with 1.6 U/mL collagenase D for 20 min to collect the cells. Treatment with collagenase D was repeated twice. All digestion steps were carried out in a 3.5 mL solution in a six-well Petri dish, on a rotating shaker at 120 rpm in a 37°C incubator. After centrifugation and removal of supernatant, cell pellets were suspended in 1% (w/v) BSA-PBS(−); pH = 7.0 (Nacalai Tesque, Inc.). To remove CD45-positive cells, cell suspensions were mixed with CD45 microbeads (Miltenyi Biotec) and incubated at 4°C for 15 min in PEB buffer and rinsed. PEB buffer contains MACS® BSA stock solution (Miltenyi Biotec) and autoMACS rinsing solution (Miltenyi Biotec) in a 1:20 ratio. Cell suspensions were passed through the LD column (Miltenyi Biotec). The flow-through was collected, centrifuged, and resuspended in 1% (w/v) BSA-PBS(−). Cells were passed through a 40 μm nylon strainer and used for scRNA-seq.
scRNA-seq library preparation and sequencing
For each sample, the quantity and viability of the cells were evaluated. Absence of aggregated cells or cell debris was confirmed microscopically. Single cells were encapsulated into emulsion droplets using Chromium Connect (10x Genomics). scRNA-seq libraries were constructed using Chromium Next GEM Automated Single Cell 3′ Library and Gel Bead Kit v. 3.1 (10x Genomics), Chromium Next GEM Automated Chip G Single Cell Kit, and Single Index Kit T Set A (10x Genomics) according to the manufacturer’s protocol. The libraries were sequenced using Illumina NextSeq 2000.
Data processing for scRNA-seq datasets
Sequencing reads (FASTQ files) were mapped to the mouse reference genome (mm10) using Cell Ranger (v. 4.0.0; 10x Genomics; https://support.10xgenomics.com/single-cell-gene-expression/software/overview/welcome). The downstream analysis was performed using the Seurat R package (version 3.2.2). The output matrices from Cell Ranger were converted into Seurat objects and barcodes with <200 genes were excluded to eliminate low-quality cells or cell-free droplets as a primary quality control. Following the primary quality control, each object was normalized and scaled according to the Seurat standard workflow, followed by principal component analysis based on the top 2000 highly variable genes, Uniform Manifold Approximation and Projection (UMAP) dimension reduction, and unsupervised clustering. After the clusters were obtained, the clusters with a high mitochondrial gene fraction (median of percent. mt >10%), low gene counts (median of nFeature <1000), and low Unique Molecular Identifier (UMI) counts (median of nCount <2000) were excluded to eliminate dead cells as the secondary quality control. Following the secondary quality control, the objects were reprocessed following the Seurat standard workflow and subsequently integrated by the Seurat anchor-based integration workflow followed by principal component analysis, dimensionality reduction, and unsupervised clustering. The UMAP coordinate and the unsupervised cluster information of each cell was incorporated in the cloupe file for downstream analysis.
Real-time quantitative PCR
Total RNA was isolated with a phenol-chloroform extraction method using ISOGEN kit (Nippon Gene Co., Ltd.) according to the manufacturer’s protocol, and reverse transcribed using QuantiTect® Reverse Transcription Kit (QIAGEN). cDNA from 1 µg of total RNA was subjected to real-time PCR. TaqMan Gene Expression Master Mix (Life Technologies Co., Ltd.) was used for real-time quantitative PCR. The primer sets were obtained from Applied Biosystems (Fgf23, no. Mm00445621_m1; Actb, no. Mm02619580_g1). Reactions were run on and analyzed with a QuantStudio™ 5 real-time PCR system (Thermo Fisher Scientific K.K.). Fgf23 mRNA expression levels were normalized with respect to those of Actb mRNA.
Serum FGF23 measurements
Serum FGF23 levels were measured in mice using a sandwich enzyme-linked immunosorbent assay kit (Kainos Laboratories) (Yamazaki et al., 2002).
Results
Detection of Dmp1, Mepe, and Phex gene expression by in situ hybridization
We first confirmed the expression of Dmp1, Mepe, and Phex, genes which are already known to be expressed in osteocytes, by in situ hybridization of murine femur cortical bone tissue (Petersen et al., 2000; Gluhak-Heinrich et al., 2003; Kalajzic et al., 2004; Lu et al., 2004; Dallas et al., 2013). Positive signals were detected only with antisense probes for each gene (Figures 1A, C, E), and no non-specific signals were detected with sense probes (Figures 1B, D, F). Thus, in this scRNA-seq analysis, we defined a cell cluster with higher expression levels of Dmp1, Mepe, and Phex as osteocytes.
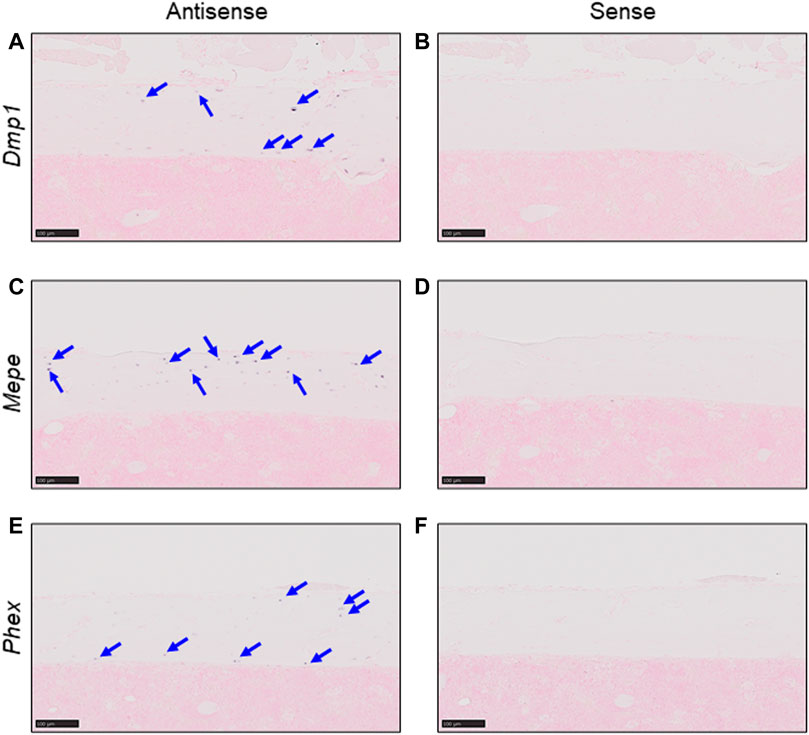
FIGURE 1. Detection of osteocyte marker mRNA using in situ hybridization. Expression of osteocyte markers was detected in serial sections from untreated C57BL/6J murine femurs by in situ hybridization. (A,B) Expression of Dmp1 in cortical bone. (C,D) Expression of Mepe in cortical bone. (E,F) Expression of Phex in cortical bone. Markers detected using antisense probes [panels (A, C, E)] and sense probes [panels (B, D, F)] (Scale bar: 100 μm).
scRNA-seq identifies diverse murine femoral cell populations and their transcriptomes
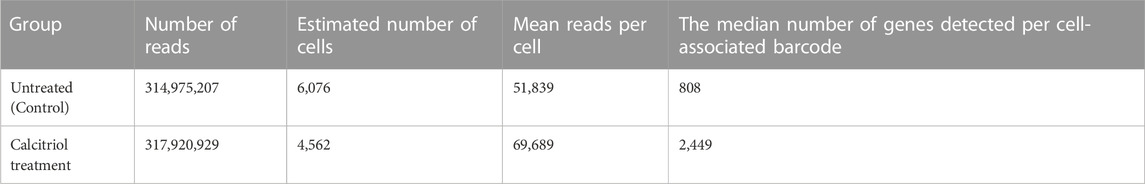
C57BL/6J mice were injected with calcitriol every day for 3 days. Untreated C57BL/6J mice were used as controls (Figure 2A). The femurs from six mice per group were collected, and used for isolation of femoral cells. Subsequently, scRNA-seq was performed on isolated cells (Figure 2B). More than 60,000 cells could be collected with a cell viability rate of over 80% from each group sample (Table 1), which were used to generate scRNA-seq libraries. We obtained over 300 million reads from the femoral single-cell libraries in each group. In the control group, 51,839 reads per cell and 6,076 cells per library were obtained. On the other hand, 69,689 reads per cell and 4,562 cells per library were obtained in the calcitriol-injected group. These libraries were used for preprocessing and filtering (Table 2).
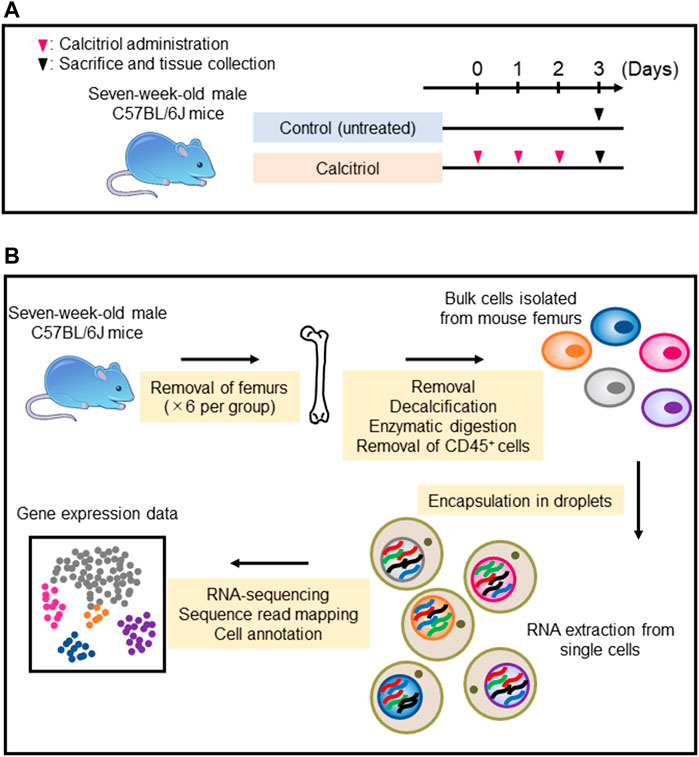
FIGURE 2. Single-cell RNA sequencing sample preparation and RNA sequencing protocol. (A) Seven-week-old male C57BL/6J mice were injected with 2.5 μg per kg of calcitriol (n = 6) every day for 3 days. As a control, six untreated C57BL/6J mice were sacrificed on day 3. (B) Each cell isolated from six femurs by collagenase digestion and EGTA treatment was encapsulated in a droplet for mRNA extraction and RNA sequencing.
Murine femoral cells were divided into 18 clusters using Seurat’s unbiased cluster detection algorithms (Figure 3A). Cluster #17 was identified as an osteocyte cluster based on expression of Dmp1, Mepe, and Phex, which had been defined as osteocyte markers (Figure 3B). In other cell types, clusters were identified based on the expression profiles of osteoblast, chondrocyte, hematopoietic stem cell, endothelial cell, smooth muscle cell, and red blood cell marker genes that have been reported previously (Spangrude and Brooks., 1993; Ducy et al., 1996; Komori et al., 1997; Letamendía et al., 1998; Bi et al., 1999; Horiuchi et al., 1999; Ikegawa et al., 2000; Wang et al., 2006; Runck et al., 2009; Toffalini and Demoulin, 2010). Clusters #8 and #12 exhibited osteoblast markers (e.g., Bglap, Postn, and Runx2). Clusters #4, #10, and #14 exhibited chondrocyte markers (e.g., Col2a1, Prg4, and Sox9). The remaining femoral cell clusters expressed transcripts found in hematopoietic stem cells (Ly6a and Pdgfra; clusters #1, #3, #6, and #13), endothelial cells (Eng; clusters #7 and #16), smooth muscle cells (Acta2, cluster #5), and red blood cells (Hbb-bs; clusters #0, #2, #11, and #15) (Figure 3B). Vdr expression that dictates calcitriol responsiveness was detected in clusters #4, #8, #12, and #17, that contain chondrocytes (#4), osteoblasts (#8, #12), and osteocytes (#17). (Figure 3B).
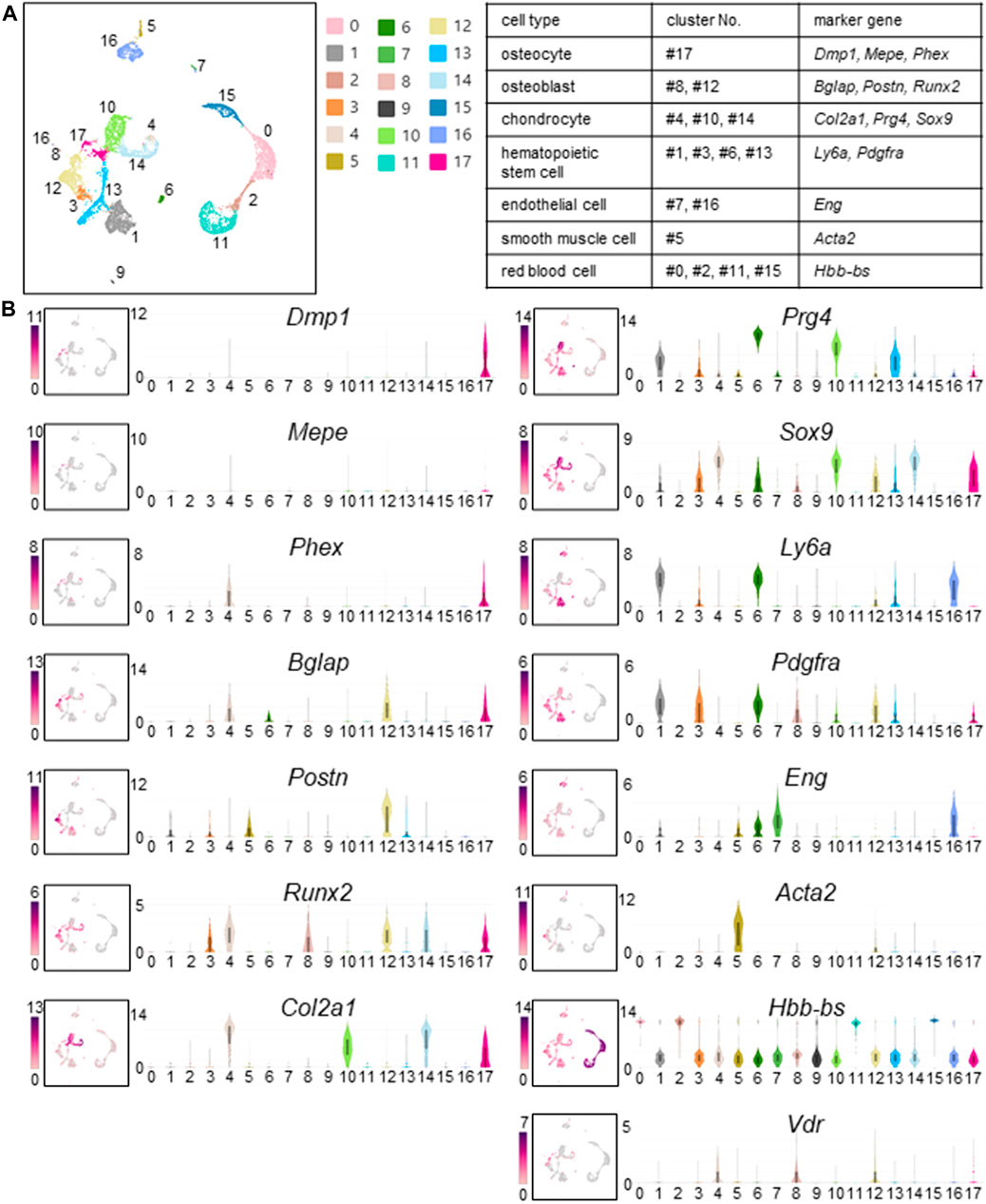
FIGURE 3. Cell clusters identified in fresh cells isolated from murine femurs. (A) Cell clustering was performed on cells isolated from femurs of calcitriol-injected and untreated C57BL/6J mice. Uniform Manifold Approximation and Projection (UMAP) plot wherein each dot represents a single cell, and cells sharing the same color code indicate discrete populations of transcriptionally similar cells. Table shows the correspondence between cell type, cluster number, and marker gene when identified. (B) UMAP plots of gene expression (left) and associated violin plots (right) indicate cluster-specific expression of representative genes. Osteocytes are marked by expression of Dmp1, Mepe, and Phex (#17). Similarly, osteoblasts express Bglap, Postn, and Runx2 (#8 and #12). Chondrocytes express Col2a1, Prg4, and Sox9 (#4, #10, and #14). Hematopoietic stem cells express Ly6a and Pdgfra (#1, #3, #6, and #13). Endothelial cells express Eng (#7 and #16). Smooth muscle cells are distinctly marked by the expression of Acta2 (#5). Red blood cells express Hbb-Bs (#0, #2, #11, and #15). Vdr expression that dictates calcitriol responsiveness is shown by UMAP plot and violin plot. Vdr was expressed in osteoblasts, chondrocytes and osteocytes (#4, #8, #12, and #17).
Detection of Fgf23-expressing osteocytes in femurs of calcitriol-injected mice
We compared the number of Fgf23-expressing cells between untreated and calcitriol-injected murine femoral cells (Figure 4A). Fgf23-expressing cells were not detected in the untreated mice. On the other hand, Fgf23-expressing cells were observed in clusters #1, #3, #12, and #17 of calcitriol-injected murine femoral cells. Eight Fgf23-expressing cells were observed in cluster #17 (osteocytes) of calcitriol-injected murine femoral cells (Figures 4A, B). In the calcitriol-injected group, Fgf23 mRNA levels in cells isolated from femurs were higher than in the control group as measured by real-time quantitative PCR (Supplementary Figure S1). We compared another osteocyte marker gene, Sost, between untreated and calcitriol-injected murine femoral cells. Sost-expressing cells were detected at the same existence in both of the two groups (Figures 4D, E). Matching the distribution of Fgf23 expression with the cell type of the cluster revealed that some of the osteocytes classified in cluster #17 expressed Fgf23 (Figure 4C). Serum FGF23 levels were also higher in the calcitriol-injected group compared with the control group (Supplementary Figure S2).
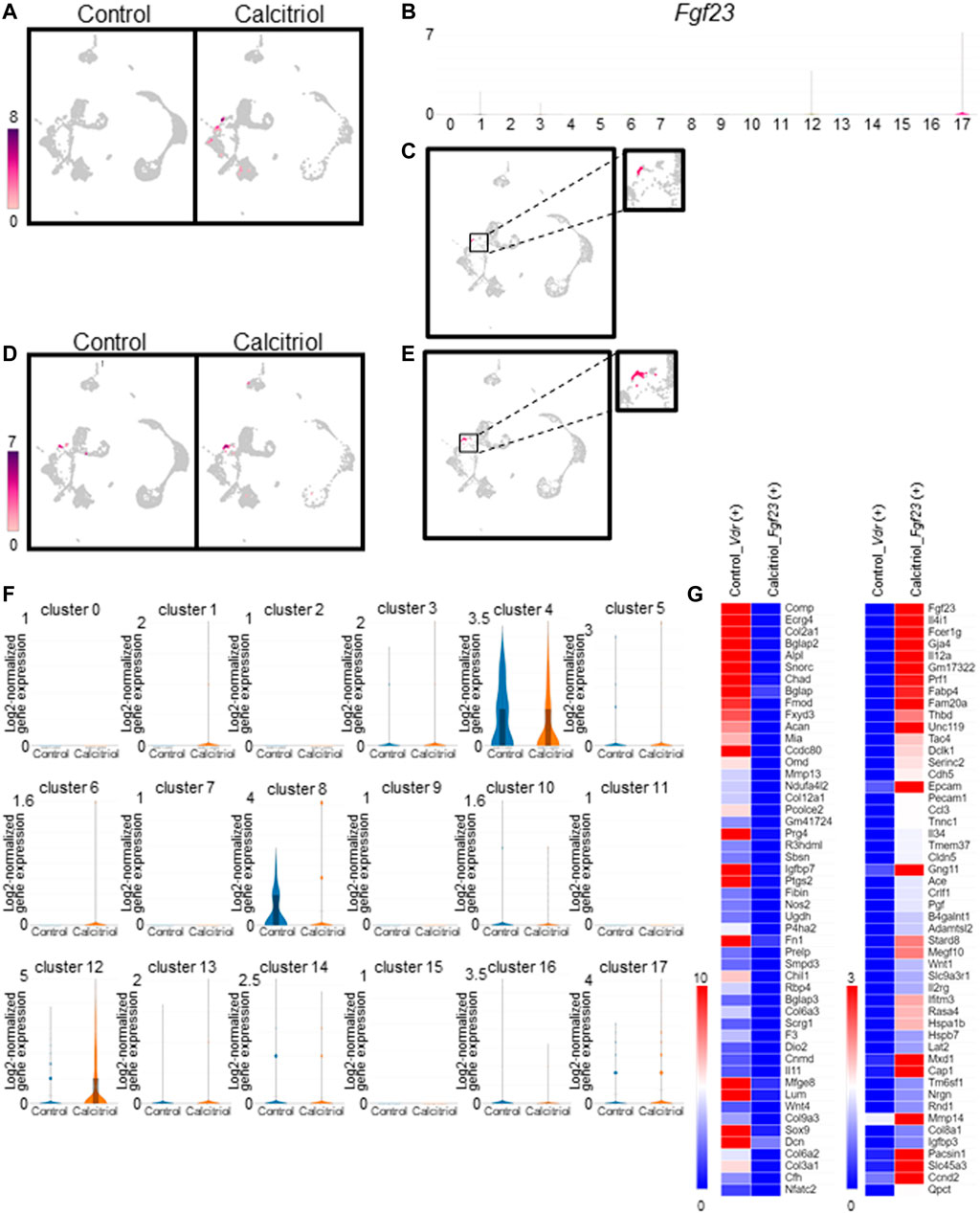
FIGURE 4. Identification and characterization of Fgf23-expressing cells in fresh cells isolated from femurs of calcitriol-injected mice. (A) Uniform Manifold Approximation and Projection (UMAP) plot showing Fgf23 gene expression in all cells from calcitriol-injected mice or control mice. Color scale bar shows log2-normalized gene expression values. (B) Violin plots indicate cluster-specific expression of Fgf23 gene. (C) UMAP plot showing Fgf23 gene expression in osteocytes (cluster #17) of calcitriol-injected mice. (D) UMAP plot showing Sost gene expression in all cells from calcitriol-injected mice or control mice. Color scale bar shows log2-normalized gene expression values. (E) UMAP plot showing Sost gene expression in osteocytes (cluster #17) of calcitriol-injected mice. (F) Violin plots indicate Vdr gene expression in the untreated and calcitriol-injected mice in each cluster. (G) The gene expression was compared between Fgf23-expressing osteocytes in calcitriol-injected group and Vdr-expressing osteocytes in control group. Heat maps indicate gene expression profiles upregulated (right) or downregulated (left) in Fgf23-expressing osteocytes in calcitriol-injected group.
To estimate the origin of Fgf23-expressing cells, we compared gene expression profiles in Vdr-expressing cells in the control group and Fgf23-expressing cells in the calcitriol-injected group. Vdr-expressing cells were observed in clusters #4, #8, #12, and #17 (Figure 3B). In clusters #4 and #8, Vdr gene expression was higher in the control group than in calcitriol-injected group (Figure 4F). This suggested that the percentage of Vdr-high expressing cells in cluster #4 and #8, i.e., chondrocytes and osteoblasts, was reduced by calcitriol treatment. We compared the gene expression profiles in Vdr-expressing osteocytes (cluster #17) in the control group and Fgf23-expressing osteocytes in the calcitriol-injected group. As a result, osteoblast and chondrocyte markers such as Bglap, Col2a1, Prg4, and Sox9 (Ducy et al., 1996; Bi et al., 1999; Ikegawa et al., 2000) were detected as downregulated genes, especially in Fgf23-expressing cells (Figure 4G).
Fgf23-expressing osteocytes are defined by specific gene expression profiles
To characterize Fgf23-expressing osteocytes, we compared the expression levels of Dmp1, Fam20c, and Phex genes, whose inactivating mutations have been shown to cause FGF23-related hypophosphatemic diseases, between osteocytes isolated from the control mice, as well as non–Fgf23-expressing osteocytes and Fgf23-expressing osteocytes isolated from the calcitriol-treated mice. Fgf23-expressing osteocytes showed higher expression levels of Dmp1, Fam20c, and Phex genes (Figure 5A). In the calcitriol-injected group, Dmp1 mRNA levels in cells isolated from femurs were higher than in the control group as measured by real-time quantitative PCR (Supplementary Figure S3). We further compared gene expression profiles in these osteocytes. The expression levels of several genes that are known to be enhanced by 1,25-dihydroxyvitamin D3, such as Cap1, Ccnd2, Dnmt3a, Dyrk3, Il12a, Il4i1, Mxd1, Pdgfa, Serinc2, Sulf2, Timp1, Tmem37, and Vdr, were increased by calcitriol in Fgf23-expressing osteocytes compared with untreated osteocytes (Zhang et al., 1995; Timms et al., 2002; Satoh et al., 2005; Wang et al., 2005; An et al., 2010; Ramagopalan et al., 2010; Heikkinen et al., 2011; Salehi-Tabar et al., 2012; Ding et al., 2013; Goeman et al., 2014; Ryynänen et al., 2014; Seuter et al., 2014; Salehi-Tabar et al., 2019; Fernández-Barral et al., 2020). The expression of Igfbp7 and Fn1 genes, which are also known to be suppressed by 1,25-dihydroxyvitamin D3, was decreased by calcitriol in Fgf23-expressing osteocytes (Shang et al., 2000; Zhang et al., 2008; Satoh and Tabunoki., 2013) (Figure 5B). The expression levels of these genes were not influenced in osteocytes lacking Fgf23 expression even after calcitriol injection (Figure 5B).
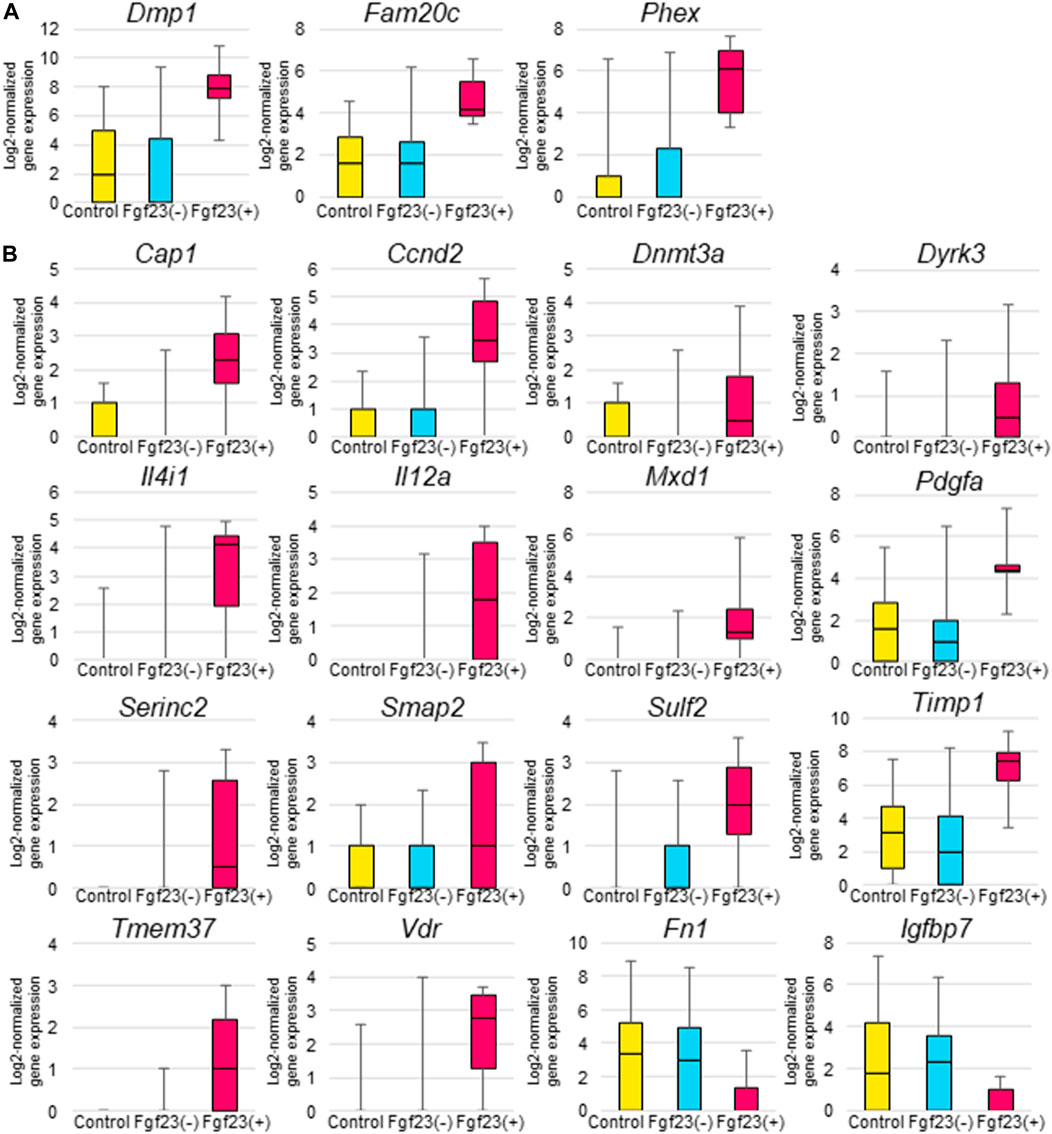
FIGURE 5. Comparison of gene expression levels among Fgf23-expressing osteocytes and non-Fgf23-expressing osteocytes from femurs of untreated or calcitriol-injected mice. (A) Box-and-whisker plots indicate Dmp1, Fam20c, and Phex gene expression in osteocytes from untreated mice, non-Fgf23-expressing osteocytes from calcitriol-injected mice, and Fgf23-expressing osteocytes from calcitriol-injected mice. The Y-axes of the plots show log2-normalized gene expression values. (B) Box-and-whisker plots indicate 1,25-dihydroxyvitamin D3–stimulated gene expression in untreated osteocytes from untreated mice, non-Fgf23-expressing osteocytes from calcitriol-injected mice, and Fgf23-expressing osteocytes from calcitriol-injected mice. The vertical axes of the plots show log2-normalized gene expression values.
Discussion
Osteocytes are embedded in the mineralized bone matrix, connecting and interacting with each other via gap junctions and through a bone fluid flow (Parfitt, 1977; Kamioka et al., 2001). One of the distinguishing characteristics of osteocytes is their lifespan, which can extend up to several decades in the bone matrix (Lanyon, 1993). However, recent studies have demonstrated the role of osteocytes as secretory cells as well (Bonewald, 2011). Osteocytes can organize bone remodeling, control calcium and phosphate homeostasis, and transmit signals to distant tissues. Recent studies also show that they have high heterogeneity in gene expression. For example, matured and deeply embedded osteocytes express high level of sclerostin, which is the product of the SOST gene (Dallas et al., 2013).
In this study, we focused on the genetic definition of Fgf23-expressing osteocytes. FGF23 is already known to be expressed in bone cells, especially in osteocytes (Feng et al., 2006; Liu et al., 2006). In addition to FGF23, we know that some proteins specifically expressed in osteocytes play critical roles in phosphate homeostasis, for example, PHEX and DMP1 (Bonewald, 2007; Bonewald and Johnson., 2008). Dmp1, Mepe, and Phex are highly expressed in osteocytes compared with osteoblasts or other cell types (Petersen et al., 2000; Gluhak-Heinrich et al., 2003; Kalajzic et al., 2004; Lu et al., 2004; Dallas et al., 2013). We were able to detect Dmp1, Mepe, and Phex expression specifically in osteocytes using in situ hybridization (Figure 1). By using these genes as marker genes, we could define osteocytes in scRNA-seq analysis (Figure 3).
To characterize femoral cells including osteocytes, we used the scRNA-seq technology. Ayturk et al. performed scRNA-seq using neonatal mouse calvarial cells and compared the relative cell type abundance and the transcriptomes of freshly isolated cells. However, in this study, osteocytes were not detected from neonatal murine calvarial cells (Ayturk et al., 2020). This is probably because osteocytes are present in the bone matrix and it is difficult to isolate them. In our current study, we focused on osteocyte detection from murine femurs. We optimized the protocols for tissue disruption, enzyme digestion, and depletion of CD45+ cells by MACS protocol (Figure 2), and we were able to detect osteocytes as a cell cluster that was defined by osteocyte-specific marker gene expression (Figure 3A). In addition, we defined other cell types using marker genes for osteoblasts, chondrocytes, hematopoietic stem cells, endothelial cells, smooth muscle cells, and red blood cells (Figure 3B). We obtained transcriptome profiles of murine femurs and analyzed characteristics of each cell type (Figure 3).
FGF23 is normally expressed at considerably low levels in osteocytes but its expression is considered to be increased in patients with hypophosphatemic rickets (Liu et al., 2006; Endo et al., 2008) and in patients with chronic kidney disease (Pereira et al., 2009). We could not detect Fgf23-expressing osteocytes by scRNA-seq from untreated murine femurs (Figure 4A). Our results are consistent with the previous reports.
To isolate Fgf23-expressing osteocytes, mice were injected with calcitriol. 1,25-dihydroxyvitamin D3 is known to induce expression of Fgf23 in the osteocytes (Liu et al., 2006; Woo et al., 2011), suggesting a negative feedback system. The induction of Fgf23 by 1,25-dihydroxyvitamin D3 is mediated by vitamin D receptor (Saji et al., 2010; Yashiro et al., 2020). In this study, Vdr gene expression was detected in osteoblasts, chondrocytes, and osteocytes (Figure 3A). It is known that Vdr is expressed in immature osteoblasts and chondrocytes (Wang et al., 2014). After calcitriol treatment, we could detect Fgf23 expression in some cells that we had categorized as osteocytes (Figure 4A). Osteoblast and chondrocyte markers, Bglap, Col2a1, Prg4, and Sox9 (Ducy et al., 1996; Bi et al., 1999; Ikegawa et al., 2000) were downregulated in Fgf23-expressing cells compared with Vdr-expressing cells in control group (Figure 4G). From these results, it was presumed that calcitriol treatment enhanced differentiation of Vdr-expressing osteoblasts into osteocytes, and increased the number of Fgf23-expressing cells. VDR is important in the late stage of osteogenic differentiation (Yang et al., 2018). Our results are consistent with the previous reports.
To genetically characterize Fgf23-expressing osteocytes, we analyzed the gene expressions regulating Fgf23 expression. DMP1, PHEX, and FAM20C proteins are highly expressed in osteocytes and regulate FGF23 production and bone mineralization (Francis et al., 1995; Thompson et al., 2002; Bonewald, 2007; Kinoshita et al., 2014). In Fgf23-expressing osteocytes responding to calcitriol treatment, the expression of Dmp1, Phex, and Fam20c was higher than in non-Fgf23-expressing osteocytes (Figure 5A). However, the expression levels of these genes were variable in osteocytes from the untreated mice. It is probable that some osteocytes in untreated mice had expression levels of Dmp1, Phex, and Fam20c that were similar to those in Fgf23-expressing osteocytes. Since there have been no previous reports indicating that 1,25-dihydroxyvitamin D3 upregulates the expression of these genes, these results suggest that there is a population of osteocytes with higher expression levels of Dmp1, Phex, and Fam20c that can produce FGF23 in response to 1,25-dihydroxyvitamin D3.
Mutations in PHEX, DMP1, and FAM20C have been reported to be responsible for FGF23-related hypophosphatemia. Inactivating mutations in these genes cause the hypophosphatemic diseases X-linked hypophosphatemic rickets, autosomal recessive hypophosphatemic rickets 1, and Raine syndrome, respectively (George et al., 1993; Francis et al., 1995; Feng et al., 2006; Simpson et al., 2007). Therefore, it seems rather paradoxical that FGF23-producing osteocytes also express these genes. However, serial analysis of gene expression identified DMP1 and PHEX as overexpressed genes in tumors responsible for tumor-induced osteomalacia and secreting FGF23 (De Beur et al., 2002). Additionally, Dmp1 and Fam20c expression levels were higher in osteocytes from Hyp mice compared with those from wild-type mice (Miyagawa et al., 2014). While the detailed mechanism is unclear, our results suggest that PHEX, DMP1, and FAM20C are involved in the regulation of FGF23 production in a cell-autonomous manner.
There are several limitations in our study. Osteocytes make up 90%–95% of cells in the bone tissue (Bonewald, 2007), However, the number of isolated osteocytes and Fgf23-expressing osteocytes were small. Since isolation of single cells from bones is a time-consuming procedure, it is possible that the gene expression profiles of isolated cells are different from their profiles in vivo. Additionally, we could not detect Fgf23 expression in bone cells in untreated mice. Further refinement of the methods for preparing bone cells would be necessary to address these issues.
In this study, we detail the first report of Fgf23-expressing osteocytes isolated using scRNA-seq. We also show that osteocytes are heterogeneous with respect to their responsiveness to 1,25-dihydroxyvitamin D3, and sensitivity to active vitamin D is one of the characteristics of osteocytes with Fgf23 expression. It is likely that there is a subpopulation of osteocytes which expresses several genes, including Fgf23, that can affect phosphate metabolism.
Data availability statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/search/all/?term=GSE220836.
Ethics statement
The animal study was reviewed and approved by the Institutional Animal Care and Use committee Kyowa Kirin Co., Ltd.
Author contributions
AH, KN, and SF contributed to conception and design of the study. AH and KN performed in vivo experiments. AK and IU performed the data processing, database construction and data analysis. AH wrote the first draft of the manuscript. AH, AK, KN, and IU wrote sections of the manuscript. All authors contributed to manuscript revision, read, and approved the submitted version.
Funding
This work was supported in part by Grant-in-Aid for Scientific Research no. 22H03089 (to SF).
Conflict of interest
Authors AH, AK, KN, KM, IU, and YY were employed by Kyowa Kirin Co., Ltd.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fphys.2023.1102751/full#supplementary-material
Abbreviations
cDNA, complementary DNA; DIG, digoxigenin; mRNA, messenger RNA; PCR, polymerase chain reaction; scRNA-seq, single-cell RNA sequencing.
References
ADHR Consortium (2000). Autosomal dominant hypophosphataemic rickets is associated with mutations in FGF23. Nat. Genet. 26, 345–348. doi:10.1038/81664
An B. S., Tavera-Mendoza L. E., Dimitrov V., Wang X., Calderon M. R., Wang H. J., et al. (2010). Stimulation of Sirt1-regulated FoxO protein function by the ligand-bound vitamin D receptor. Mol. Cell Biol. 30, 4890–4900. doi:10.1128/MCB.00180-10
Antoniucci D. M., Yamashita T., Portale A. A. (2006). Dietary phosphorus regulates serum fibroblast growth factor-23 concentrations in healthy men. J. Clin. Endocrinol. Metab. 91, 3144–3149. doi:10.1210/jc.2006-0021
Ayturk U. M., Scollan J. P., Goz Ayturk. D., Suh E. S., Vesprey A., Jacobsen C. M., et al. (2020). Single-cell RNA sequencing of calvarial and long-bone endocortical cells. J. Bone Min. Res. 35, 1981–1991. doi:10.1002/jbmr.4052
Baryawno N., Przybylski D., Kowalczyk M. S., Kfoury Y., Severe N., Gustafsson K., et al. (2019). A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell 177, 1915–1932. doi:10.1016/j.cell.2019.04.040
Beck L., Soumounou Y., Martel J., Krishnamurthy G., Gauthier C., Goodyer C. G., et al. (1997). Pex/PEX tissue distribution and evidence for a deletion in the 3' region of the Pex gene in X-linked hypophosphatemic mice. J. Clin. Invest. 99, 1200–1209. doi:10.1172/JCI119276
Bi W., Deng J. M., Zhang Z., Behringer R. R., de Crombrugghe B. (1999). Sox9 is required for cartilage formation. Nat. Genet. 22, 85–89. doi:10.1038/8792
Bonewald L. F., Johnson M. L. (2008). Osteocytes, mechanosensing and Wnt signaling. Bone 42, 606–615. doi:10.1016/j.bone.2007.12.224
Bonewald L. F. (2007). Osteocytes as dynamic multifunctional cells. Ann. N. Y. Acad. Sci. 1116, 281–290. doi:10.1196/annals.1402.018
Dallas S. L., Prideaux M., Bonewald L. F. (2013). The osteocyte: An endocrine cell. and more. Endocr. Rev. 34, 658–690. doi:10.1210/er.2012-1026
De Beur S. M., Finnegan R. B., Vassiliadis J., Cook B., Barberio D., Estes S., et al. (2002). Tumors associated with oncogenic osteomalacia express genes important in bone and mineral metabolism. J. Bone Min. Res. 17, 1102–1110. doi:10.1359/jbmr.2002.17.6.1102
Ding N., Yu R. T., Subramaniam N., Sherman M. H., Wilson C., Rao R., et al. (2013). A vitamin D receptor/SMAD genomic circuit gates hepatic fibrotic response. Cell 153, 601–613. doi:10.1016/j.cell.2013.03.028
Du L., Desbarats M., Viel J., Glorieux F. H., Cawthorn C., Ecarot B. (1996). cDNA cloning of the murine Pex gene implicated in X-linked hypophosphatemia and evidence for expression in bone. Genomics 36, 22–28. doi:10.1006/geno.1996.0421
Ducy P., Desbois C., Boyce B., Pinero G., Story B., Dunstan C., et al. (1996). Increased bone formation in osteocalcin-deficient mice. Nature 382, 448–452. doi:10.1038/382448a0
Endo I., Fukumoto S., Ozono K., Namba N., Tanaka H., Inoue D., et al. (2008). Clinical usefulness of measurement of fibroblast growth factor 23 (FGF23) in hypophosphatemic patients: Proposal of diagnostic criteria using FGF23 measurement. Bone 42, 1235–1239. doi:10.1016/j.bone.2008.02.014
Feng J. Q., Ward L. M., Liu S., Lu Y., Xie Y., Yuan B., et al. (2006). Loss of DMP1 causes rickets and osteomalacia and identifies a role for osteocytes in mineral metabolism. Nat. Genet. 38, 1310–1315. doi:10.1038/ng1905
Fernández-Barral A., Costales-Carrera A., Buira S. P., Jung P., Ferrer-Mayorga G., Larriba M. J., et al. (2020). Vitamin D differentially regulates colon stem cells in patient-derived normal and tumor organoids. FEBS J. 287, 53–72. doi:10.1111/febs.14998
Ferrari S. L., Bonjour J. P., Rizzoli R. (2005). Fibroblast growth factor-23 relationship to dietary phosphate and renal phosphate handling in healthy young men. J. Clin. Endocrinol. Metab. 90, 1519–1524. doi:10.1210/jc.2004-1039
Francis F., Hennig S., Korn B., Reinhardt R., Jong P., Poustka A., et al. (1995). A gene (PEX) with homologies to endopeptidases is mutated in patients with X-linked hypophosphatemic rickets. The HYP Consortium. Nat. Genet. 11, 130–136. doi:10.1038/ng1095-130
Fukumoto S., Martin T. J. (2009). Bone as an endocrine organ. Trends Endocrinol. Metab. 20, 230–236. doi:10.1016/j.tem.2009.02.001
George A., Sabsay B., Simonian P. A., Veis A. (1993). Characterization of a novel dentin matrix acidic phosphoprotein. Implications for induction of biomineralization. J. Biol. Chem. 268, 12624–12630. doi:10.1016/S0021-9258(18)31434-0
Gluhak-Heinrich J., Ye L., Bonewald L. F., Feng J. Q., MacDougall M., Harris S. E., et al. (2003). Mechanical loading stimulates dentin matrix protein 1 (DMP1) expression in osteocytes in vivo. J. Bone Min. Res. 18, 807–817. doi:10.1359/jbmr.2003.18.5.807
Goeman F., Nicola F. D., Onorio D. P. D., Meo D., Pallocca M., Elmi B., et al. (2014). VDR primary targets by genome-wide transcriptional profiling. J. Steroid Biochem. Mol. Biol. 143, 348–356. doi:10.1016/j.jsbmb.2014.03.007
Grieff M., Mumm S., Waeltz P., Mazzarella R., Whyte M. P., Thakker R. V., et al. (1997). Expression and cloning of the human X-linked hypophosphatemia gene cDNA. Biochem. Biophys. Res. Commun. 231, 635–639. doi:10.1006/bbrc.1997.6153
Guo R., Quarles L. D. (1997). Cloning and sequencing of human PEX from a bone cDNA library: Evidence for its developmental stage-specific regulation in osteoblasts. J. Bone Min. Res. 12, 1009–1017. doi:10.1359/jbmr.1997.12.7.1009
Heikkinen S., Väisänen S., Pehkonen P., Seuter S., Benes V., Carlberg C. (2011). Nuclear hormone 1α,25-dihydroxyvitamin D3 elicits a genome-wide shift in the locations of VDR chromatin occupancy. Nucleic Acids Res. 39, 9181–9193. doi:10.1093/nar/gkr654
Horiuchi K., Amizuka N., Takeshita S., Takamatsu H., Katsuura M., Ozawa H., et al. (1999). Identification and characterization of a novel protein, periostin, with restricted expression to periosteum and periodontal ligament and increased expression by transforming growth factor beta. J. Bone Min. Res. 14, 1239–1249. doi:10.1359/jbmr.1999.14.7.1239
Ikegawa S., Sano M., Koshizuka Y., Nakamura Y. (2000). Isolation, characterization and mapping of the mouse and human PRG4 (proteoglycan 4) genes. Cytogenet. Cell Genet. 90, 291–297. doi:10.1159/000056791
Kalajzic I., Braut A., Guo D., Jiang X., Kronenberg M. S., Mina M., et al. (2004). Dentin matrix protein 1 expression during osteoblastic differentiation, generation of an osteocyte GFP-transgene. Bone 35, 74–82. doi:10.1016/j.bone.2004.03.006
Kamioka H., Honjo T., Takano-Yamamoto T. (2001). A three-dimensional distribution of osteocyte processes revealed by the combination of confocal laser scanning microscopy and differential interference contrast microscopy. Bone 28, 145–149. doi:10.1016/s8756-3282(00)00421-x
Kanton S., Boyle M. J., He Z., Santel M., Weigert A., Sanchís-Calleja F., et al. (2019). Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature 574, 418–422. doi:10.1038/s41586-019-1654-9
Kinoshita Y., Hori M., Taguchi M., Fukumoto S. (2014). Functional analysis of mutant FAM20C in Raine syndrome with FGF23-related hypophosphatemia. Bone 67, 145–151. doi:10.1016/j.bone.2014.07.009
Kolek O. I., Hines E. R., Jones M. D., LeSueur L. K., Lipko M. A., Kiela P. R., et al. (2005). 1alpha,25-Dihydroxyvitamin D3 upregulates FGF23 gene expression in bone: The final link in a renal-gastrointestinal-skeletal axis that controls phosphate transport. Am. J. Physiol. Gastrointest. Liver Physiol. 289, G1036–G1042. doi:10.1152/ajpgi.00243.2005
Komori T., Yagi H., Nomura S., Yamaguchi A., Sasaki K., Deguchi K., et al. (1997). Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89, 755–764. doi:10.1016/s0092-8674(00)80258-5
Lanyon L. E. (1993). Osteocytes, strain detection, bone modeling and remodeling. Calcif. Tissue Int. 53, S102–S106. doi:10.1007/BF01673415
Letamendía A., Lastres P., Botella L. M., Raab U., Langa C., Velasco B., et al. (1998). Role of endoglin in cellular responses to transforming growth factor-beta. A comparative study with betaglycan. J. Biol. Chem. 273, 33011–33019. doi:10.1074/jbc.273.49.33011
Lipman M. L., Panda D., Bennett H. P., Henderson J. E., Shane E., Shen Y., et al. (1998). Cloning of human PEX cDNA. Expression, subcellular localization, and endopeptidase activity. J. Biol. Chem. 273, 13729–13737. doi:10.1074/jbc.273.22.13729
Liu S., Guo R., Simpson L. G., Xiao Z. S., Burnham C. E., Quarles L. D. (2003). Regulation of fibroblastic growth factor 23 expression but not degradation by PHEX. J. Biol. Chem. 278, 37419–37426. doi:10.1074/jbc.M304544200
Liu S., Zhou J., Tang W., Jiang X., Rowe D. W., Quarles L. D. (2006). Pathogenic role of Fgf23 in Hyp mice. Am. J. Physiol. Endocrinol. Metab. 291, E38–E49. doi:10.1152/ajpendo.00008.2006
Lorenz-Depiereux B., Bastepe M., Benet-Pagès A., Amyere M., Wagenstaller J., Müller-Barth U., et al. (2006). DMP1 mutations in autosomal recessive hypophosphatemia implicate a bone matrix protein in the regulation of phosphate homeostasis. Nat. Genet. 38, 1248–1250. doi:10.1038/ng1868
Lu C., Huang S., Miclau T., Helms J. A., Colnot C. (2004). Mepe is expressed during skeletal development and regeneration. Histochem. Cell Biol. 121, 493–499. doi:10.1007/s00418-004-0653-5
Macosko E. Z., Basu A., Satija R., Nemesh J., Shekhar K., Goldman M., et al. (2015). Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214. doi:10.1016/j.cell.2015.05.002
Martin A., Liu S., David V., Li H., Karydis A., Feng J. Q., et al. (2011). Bone proteins PHEX and DMP1 regulate fibroblastic growth factor Fgf23 expression in osteocytes through a common pathway involving FGF receptor (FGFR) signaling. FASEB J. 25, 2551–2562. doi:10.1096/fj.10-177816
Miyagawa K., Yamazaki M., Kawai M., Nishino J., Koshimizu T., Ohata Y., et al. (2014). Dysregulated gene expression in the primary osteoblasts and osteocytes isolated from hypophosphatemic Hyp mice. PLoS One 9, e93840. doi:10.1371/journal.pone.0093840
Parfitt A. M. (1977). The cellular basis of bone turnover and bone loss: A rebuttal of the osteocytic resorption--bone flow theory. Clin. Orthop. Relat. Res. 127, 236–247. doi:10.1097/00003086-197709000-00036
Pereira R. C., Juppner H., Azucena-Serrano C. E., Yadin O., Salusky I. B., Wesseling-Perry K. (2009). Patterns of FGF-23, DMP1, and MEPE expression in patients with chronic kidney disease. Bone 45, 1161–1168. doi:10.1016/j.bone.2009.08.008
Perwad F., Azam N., Zhang M. Y., Yamashita T., Tenenhouse H. S., Portale A. A. (2005). Dietary and serum phosphorus regulate fibroblast growth factor 23 expression and 1,25-dihydroxyvitamin D metabolism in mice. Endocrinology 146, 5358–5364. doi:10.1210/en.2005-0777
Petersen D. N., Tkalcevic G. T., Mansolf A. L., Rivera-Gonzalez R., Brown T. A. (2000). Identification of osteoblast/osteocyte factor 45 (OF45), a bone-specific cDNA encoding an RGD-containing protein that is highly expressed in osteoblasts and osteocytes. J. Biol. Chem. 275, 36172–36180. doi:10.1074/jbc.M003622200
Rafaelsen S. H., Raeder H., Fagerheim A. K., Knappskog P., Carpenter T. O., Johansson S., et al. (2013). Exome sequencing reveals FAM20c mutations associated with fibroblast growth factor 23-related hypophosphatemia, dental anomalies, and ectopic calcification. J. Bone Min. Res. 28, 1378–1385. doi:10.1002/jbmr.1850
Ramagopalan S. V., Heger A., Berlanga A. J., Maugeri N. J., Lincoln M. R., Burrell A., et al. (2010). A ChIP-seq defined genome-wide map of vitamin D receptor binding: Associations with disease and evolution. Genome Res. 20, 1352–1360. doi:10.1101/gr.107920.110
Runck A. M., Moriyama H., Storz J. F. (2009). Evolution of duplicated beta-globin genes and the structural basis of hemoglobin isoform differentiation in Mus. Mol. Biol. Evol. 26, 2521–2532. doi:10.1093/molbev/msp165
Ryynänen J., Neme A., Tuomainen T. P., Virtanen J. K., Voutilainen S., Nurmi T., et al. (2014). Changes in vitamin D target gene expression in adipose tissue monitor the vitamin D response of human individuals. Mol. Nutr. Food Res. 58, 2036–2045. doi:10.1002/mnfr.201400291
Saito H., Maeda A., Ohtomo S., Hirata M., Kusano K., Kato S., et al. (2005). Circulating FGF-23 is regulated by 1alpha,25-dihydroxyvitamin D3 and phosphorus in vivo. J. Biol. Chem. 280, 2543–2549. doi:10.1074/jbc.M408903200
Saji F., Shigematsu T., Sakaguchi T., Ohya M., Orita H., Maeda Y., et al. (2010). Fibroblast growth factor 23 production in bone is directly regulated by 1{alpha},25-dihydroxyvitamin D, but not PTH. Am. J. Physiol. Ren. Physiol. 299, F1212–F1217. doi:10.1152/ajprenal.00169.2010
Salehi-Tabar R., Memari B., Wong H., Dimitrov V., Rochel N., White J. H. (2019). The tumor suppressor FBW7 and the vitamin D receptor are mutual cofactors in protein turnover and transcriptional regulation. Mol. Cancer Res. 17, 709–719. doi:10.1158/1541-7786.MCR-18-0991
Salehi-Tabar R., Nguyen-Yamamoto L., Tavera-Mendoza L. E., Quail T., Dimitrov V., An B. S., et al. (2012). Vitamin D receptor as a master regulator of the c-MYC/MXD1 network. Proc. Natl. Acad. Sci. U. S. A. 109, 18827–18832. doi:10.1073/pnas.1210037109
Satoh J., Tabunoki H. (2013). Molecular network of chromatin immunoprecipitation followed by deep sequencing-based vitamin D receptor target genes. Mult. Scler. 19, 1035–1045. doi:10.1177/1352458512471873
Seuter S., Neme A., Carlberg C. (2014). Characterization of genomic vitamin D receptor binding sites through chromatin looping and opening. PLoS One 9, e96184. doi:10.1371/journal.pone.0096184
Shang Y., Hu X., DiRenzo J., Lazar M. A., Brown M. (2000). Cofactor dynamics and sufficiency in estrogen receptor-regulated transcription. Cell 103, 843–852. doi:10.1016/s0092-8674(00)00188-4
Shimada T., Hasegawa H., Yamazaki Y., Muto T., Hino R., Takeuchi Y., et al. (2004a). FGF-23 is a potent regulator of vitamin D metabolism and phosphate homeostasis. J. Bone Min. Res. 19, 429–435. doi:10.1359/JBMR.0301264
Shimada T., Kakitani M., Yamazaki Y., Hasegawa H., Takeuchi Y., Fujita T., et al. (2004b). Targeted ablation of Fgf23 demonstrates an essential physiological role of FGF23 in phosphate and vitamin D metabolism. J. Clin. Invest. 113, 561–568. doi:10.1172/JCI19081
Shimada T., Mizutani S., Muto T., Yoneya T., Hino R., Takeda S., et al. (2001). Cloning and characterization of FGF23 as a causative factor of tumor-induced osteomalacia. Proc. Natl. Acad. Sci. U. S. A. 98, 6500–6505. doi:10.1073/pnas.101545198
Simpson M. A., Hsu R., Keir L. S., Hao J., Sivapalan G., Ernst L. M., et al. (2007). Mutations in FAM20C are associated with lethal osteosclerotic bone dysplasia (Raine syndrome), highlighting a crucial molecule in bone development. Am. J. Hum. Genet. 81, 906–912. doi:10.1086/522240
Spangrude G. J., Brooks D. M. (1993). Mouse strain variability in the expression of the hematopoietic stem cell antigen Ly-6A/E by bone marrow cells. Blood 82, 3327–3332. doi:10.1182/blood.V82.11.3327.3327
Stern A. R., Stern M. M., Van Dyke M. E., Jähn K., Prideaux M., Bonewald L. F. (2012). Isolation and culture of primary osteocytes from the long bones of skeletally mature and aged mice. Biotechniques 52, 361–373. doi:10.2144/0000113876
Thompson D. L., Sabbagh Y., Tenenhouse H. S., Roche P. C., Drezner M. K., Salisbury J. L., et al. (2002). Ontogeny of Phex/PHEX protein expression in mouse embryo and subcellular localization in osteoblasts. J. Bone Min. Res. 17, 311–320. doi:10.1359/jbmr.2002.17.2.311
Tikhonova A. N., Dolgalev I., Hu H., Sivaraj K. K., Hoxha E., Cuesta-Domínguez Á., et al. (2019). The bone marrow microenvironment at single-cell resolution. Nature 569, 222–228. doi:10.1038/s41586-019-1104-8
Timms P. M., Mannan N., Hitman G. A., Noonan K., Mills P. G., Syndercombe-Court D., et al. (2002). Circulating MMP9, vitamin D and variation in the TIMP-1 response with VDR genotype: Mechanisms for inflammatory damage in chronic disorders? QJM 95, 787–796. doi:10.1093/qjmed/95.12.787
Toffalini F., Demoulin J. B. (2010). New insights into the mechanisms of hematopoietic cell transformation by activated receptor tyrosine kinases. Blood 116, 2429–2437. doi:10.1182/blood-2010-04-279752
Wang J. S., Kamath T., Mazur C. M., Mirzamohammadi F., Rotter D., Hojo H., et al. (2021). Control of osteocyte dendrite formation by Sp7 and its target gene osteocrin. Nat. Commun. 12, 6271. doi:10.1038/s41467-021-26571-7
Wang J., Zohar R., McCulloch C. A. (2006). Multiple roles of alpha-smooth muscle actin in mechanotransduction. Exp. Cell Res. 312, 205–214. doi:10.1016/j.yexcr.2005.11.004
Wang T. T., Tavera-Mendoza L. E., Laperriere D., Libby E., MacLeod N. B., Nagai Y., et al. (2005). Large-scale in silico and microarray-based identification of direct 1,25-dihydroxyvitamin D3 target genes. Mol. Endocrinol. 19, 2685–2695. doi:10.1210/me.2005-0106
Wang Y., Zhu J., DeLuca H. F. (2014). Identification of the vitamin D receptor in osteoblasts and chondrocytes but not osteoclasts in mouse bone. J. Bone Min. Res. 29, 685–692. doi:10.1002/jbmr.2081
Wolock S. L., Krishnan I., Tenen D. E., Matkins V., Camacho V., Patel S., et al. (2019). Mapping distinct bone marrow niche populations and their differentiation paths. Cell Rep. 28, 302–311. doi:10.1016/j.celrep.2019.06.031
Woo S. M., Rosser J., Dusevich V., Kalajzic I., Bonewald L. F. (2011). Cell line IDG-SW3 replicates osteoblast-to-late-osteocyte differentiation in vitro and accelerates bone formation in vivo. J. Bone Min. Res. 26, 2634–2646. doi:10.1002/jbmr.465
Yamazaki Y., Okazaki R., Shibata M., Hasegawa Y., Satoh K., Tajima T., et al. (2002). Increased circulatory level of biologically active full-length FGF-23 in patients with hypophosphatemic rickets/osteomalacia. J. Clin. Endocrinol. Metab. 87, 4957–4960. doi:10.1210/jc.2002-021105
Yang D., Anderson P. H., Wijenayaka A. R., Barratt K. R., Triliana R., Stapledon C. J. M., et al. (2018). Both ligand and VDR expression levels critically determine the effect of 1α,25-dihydroxyvitamin-D3 on osteoblast differentiation. J. Steroid Biochem. Mol. Biol. 177, 83–90. doi:10.1016/j.jsbmb.2017.09.005
Yashiro M., Ohya M., Mima T., Nakashima Y., Kawakami K., Yamamoto S., et al. (2020). Active vitamin D and vitamin D analogs stimulate fibroblast growth factor 23 production in osteocyte-like cells via the vitamin D receptor. J. Pharm. Biomed. Anal. 182, 113139. doi:10.1016/j.jpba.2020.113139
Zhang J. Z., Maruyama K., Ono I., Kaneko F. (1995). Production and secretion of platelet-derived growth factor AB by cultured human keratinocytes: Regulatory effects of phorbol 12-myristate 13-acetate, etretinate, 1,25-dihydroxyvitamin D3, and several cytokines. J. Dermatol 22, 305–309. doi:10.1111/j.1346-8138.1995.tb03393.x
Zhang Z., Sun L., Wang Y., Ning G., Minto A. W., Kong J., et al. (2008). Renoprotective role of the vitamin D receptor in diabetic nephropathy. Kidney Int. 73, 163–171. doi:10.1038/sj.ki.5002572
Zheng G. X., Terry J. M., Belgrader P., Ryvkin P., Bent Z. W., Wilson R., et al. (2017). Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049. doi:10.1038/ncomms14049
Keywords: FGF23, osteocyte, single-cell RNA sequencing, femur, transcriptome, gene expression, 1,25-dihydroxyvitamin D3 (calcitriol)
Citation: Hanai A, Kawabata A, Nakajima K, Masuda K, Urakawa I, Abe M, Yamazaki Y and Fukumoto S (2023) Single-cell RNA sequencing identifies Fgf23-expressing osteocytes in response to 1,25-dihydroxyvitamin D3 treatment. Front. Physiol. 14:1102751. doi: 10.3389/fphys.2023.1102751
Received: 19 November 2022; Accepted: 16 January 2023;
Published: 27 January 2023.
Edited by:
Ce Shi, Jilin University, ChinaReviewed by:
Marc Wein, Massachusetts General Hospital and Harvard Medical School, United StatesSylvain Provot, Institut National de la Santé et de la Recherche Médicale (INSERM), France
Copyright © 2023 Hanai, Kawabata, Nakajima, Masuda, Urakawa, Abe, Yamazaki and Fukumoto. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ayako Hanai, YXlha28uaGFuYWkuamFAa3lvd2FraXJpbi5jb20=
 Ayako Hanai
Ayako Hanai Ayako Kawabata
Ayako Kawabata Kenta Nakajima1
Kenta Nakajima1 Yuji Yamazaki
Yuji Yamazaki Seiji Fukumoto
Seiji Fukumoto