- Department of Cardiovascular Disease, Inova Schar Heart and Vascular, Falls Church, VA, United States
Despite significant advances in organ preservation, surgical techniques, and immunosuppressive regimens, rejection continues to pose a major challenge in the care of heart transplant patients. Endomyocardial biopsy (EMB) remains the gold standard test for surveillance and diagnosis of rejection, but is limited by its invasiveness, interobserver variability, procedural risk, and cost thus prompting the widespread use of non-invasive biomarkers such as donor-derived cell-free DNA (dd-cfDNA). Due to its high negative predictive value, dd-cfDNA is often routinely used for surveillance of asymptomatic patients. However, it is a non-specific marker of allograft injury and elevated levels in the presence of a reassuring EMB creates a diagnostic dilemma. This review explores the pathophysiological basis and clinical utility of dd-cfDNA in monitoring of heart transplant recipients with particular focus on evaluation and management of discordant findings.
Introduction
Heart transplantation is the definitive treatment for patients with heart disease refractory to conventional medical, surgical, and device therapies. Despite advances in organ preservation, surgical techniques, immunosuppressive therapies, and longitudinal care, the survival rates post heart transplant (HT) have only marginally increased in the last decade (1). Patient survival is often limited by the competing problems of rejection and infection, and complications of immunosuppression such as kidney disease and malignancy. The precise calibration of immunosuppressive therapies to mitigate these risks remains a complex challenge in the management of transplant recipients. Clinically, patients with acute rejection (AR) can present with a wide spectrum of findings, ranging from asymptomatic rejection detected on routine surveillance testing to cardiogenic shock or sudden death (2). Since its introduction in the 1970s, the EMB has been the cornerstone of AR detection, allowing for histopathological evaluation of myocardial tissue and grading of rejection based on the criteria established by the International Society for Heart and Lung Transplantation (3–5). However, it is an invasive procedure associated with inherent risks such as cardiac arrhythmias (0.25%) (6), perforation (0.1%–0.5%) (7–9), damage to the tricuspid valve apparatus (0.27%–6.3%) (10, 11), and in some cases the risk of sedation. There is also significant variability in its histological and immunopathological interpretation (12, 13). To minimize the risk of a false negative test, multiple EMB specimens (usually 3) are obtained. Despite this, false negatives do exist either through sampling error, sample preservation artifacts, confusion with quilty lesions, or biopsy negative rejection (BNR) (14). BNR is an uncommon clinical diagnosis defined by the presence of allograft dysfunction without evidence of AR on histopathology, either cellular (ACR) or antibody-mediated (AMR) (15).
To address these limitations, the field of HT has witnessed tremendous growth in the development of non-invasive strategies promising earlier, safer, and more consistent detection of rejection in the last decade (16). Most commonly used in clinical practice is donor-derived cell-free DNA (dd-cfDNA), which assesses the fragments of DNA released into the recipient's bloodstream from apoptotic and necrotic donor cells (17). Elevated levels of dd-cfDNA have been shown to correlate with episodes of ACR and AMR (18–20). Due to its high negative predictive value (>98%), its widespread use in recent years has led to a significant reduction in the frequency of routine surveillance EMBs (18, 20). Nevertheless, in either asymptomatic patients biopsied following an abnormal dd-cfDNA result, or for cause biopsies in response to patient symptoms or other evidence of graft dysfunction, discordant results namely abnormal dd-cfDNA without evidence of AR on biopsy represent a diagnostic dilemma. In broader context, is the dd-cfDNA a false positive, is the biopsy a false negative, or is there another process causing allograft injury?
This review aims to explore the evolving role of dd-cfDNA in the monitoring of heart transplant recipients, emphasizing its strengths, limitations, and clinical implications when used in conjunction with EMB. We also propose a multimodal diagnostic framework to enhance the understanding of this clinical dilemma and optimize care for patients with HT in this complex and rapidly evolving field.
Donor-derived cell-free DNA: mechanism, diagnostic role, and limitations
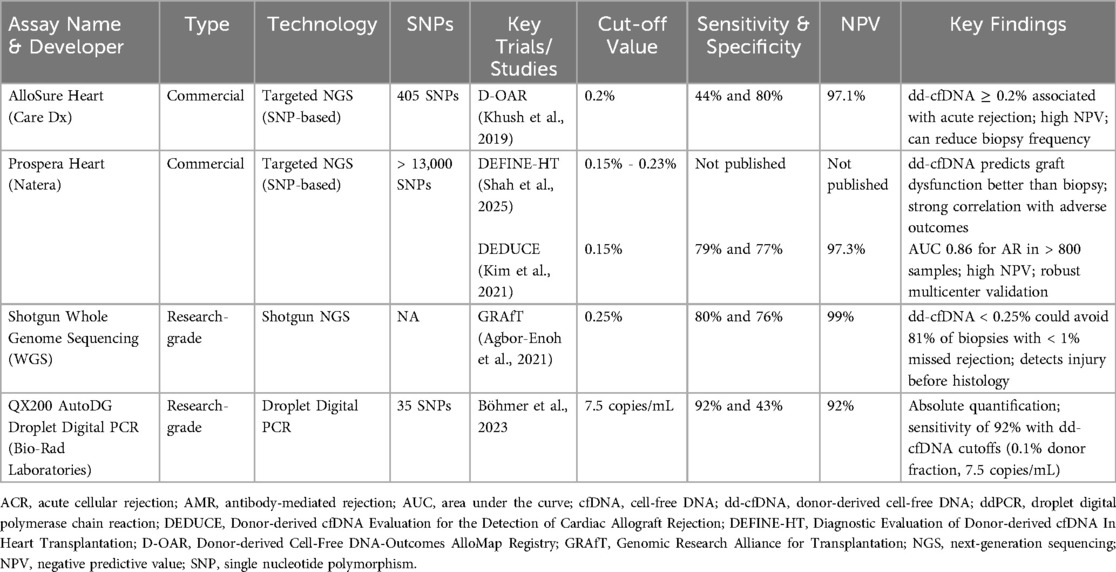
Cell-free DNA is released into the bloodstream with cell death due to normal cell turnover or disease processes (21). Normally, healthy individuals have small amounts of cfDNA, most of which is released by hematopoietic cells with a small amount released from the heart (22). In organ transplant recipients, it was hypothesized that acute rejection leads to cell death in the allograft resulting in elevated levels of dd-cfDNA in the blood of transplant recipients (17). The clinical use of earlier techniques such as shot-gun sequencing and targeted quantification using polymerase chain reaction were impractical due to cost, complexity, requirement of previous genotyping of both donor and recipient, and sex-mismatch between the donor and recipient (23). These limitations were overcome with the advent of targeted amplification. Allosure (CareDx, Inc., Brisbane, CA) is a targeted amplification sequencing assay which has been analytically and clinically validated to quantify the percentage of dd-cfDNA in the transplant recipients’ blood (18). This technique was first utilized in the D-OAR study to target highly polymorphic single nucleotide polymorphisms (SNPs) to distinguish dd-cfDNA from recipient-derived cfDNA (rd-cfDNA) (24). The Allosure assay currently includes a panel of 405 SNPs. Another commercially available assay is the Prospera test (Natera, Inc., Austin, TX), validated for clinical use in the DEDUCE trial in 2021, and includes more than 13,000 SNPs (20). Table 1 summarizes the commercially available and research grade dd-cfDNA assays.
The advantages of dd-cfDNA testing include its noninvasive nature, potential for early detection, and opportunity for longitudinal monitoring. Elevated percentage of dd-cfDNA levels have been correlated with ACR and AMR, as well as other forms of allograft injury (25). Prior studies have demonstrated that an elevated dd-cfDNA fraction can precede biopsy-proven rejection by up to 3 months, highlighting its important role in screening allografts in routine surveillance, especially in the first year post-transplant (17, 19, 26). Apart from earlier detection, researchers from Genomic Research Alliance for Transplantation (GRAfT) consortium demonstrated that dd-cfDNA from patients with AMR were rich in guanosine-cytosine bases, quantitatively 5–11 times higher, and often detected sub-clinical AMR (19). By utilizing dd-cfDNA, up to 80% of the routine surveillance biopsies could be eliminated, highlighting the high NPV (>98%) of the test (19). Elevated dd-cfDNA fraction also offers prognostic information as higher levels have been linked to unfavorable outcomes including allograft dysfunction and death at one year (13). However, it is important to note that dd-cfDNA is non-specific marker of graft injury, therefore it is used as a screening tool for AR while EMB is the test of choice in cases of suspected rejection (27). Figure 1 shows the surveillance protocol for HT recipients at our center.
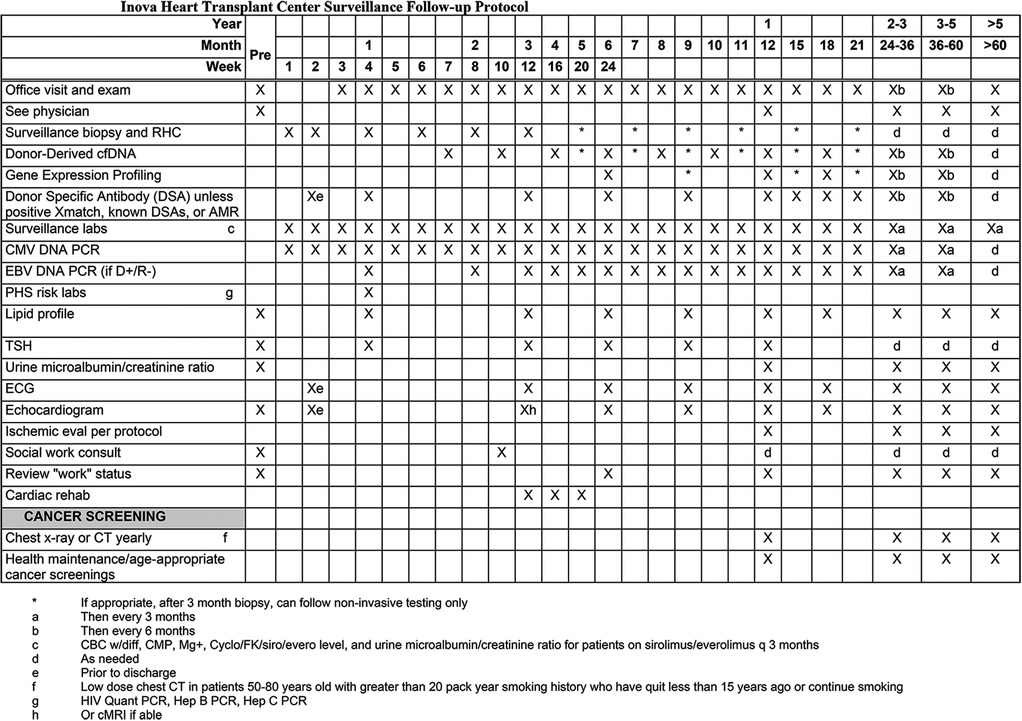
Figure 1. Routine surveillance protocol for patients after heart transplantation at Inova Schar Heart and Vascular (ISHV) Hospital. The figure shows the routine surveillance protocol for patients after heart transplantation. RHC with EMB at weeks 1, 2, 4, 6, 8, and 12. Dd-cfDNA at weeks 7 and 10, then monthly for months 4 through 12. Surveillance echocardiogram at weeks 2, then at months 3, 6, 9, and 12. DSAs at weeks 2 and 4, followed by at months 3, 6, 9, and 12. AMR, antibody mediated rejection; cfDNA, cell-free DNA; CMV, cytomegalovirus; EBV, Epstein Barr virus; PCR, polymerase chain reaction; PHS, public health service; TSH, thyroid stimulating hormone; ECG, electrocardiogram.
Non-rejection causes of elevated donor-derived cell-free DNA
The lack of histological evidence of rejection on EMB has typically been reassuring to clinicians, but the interpretation of elevated dd-cfDNA fraction in the setting of a “negative” EMB is unclear. Plausible explanations for the discordance include subclinical rejection, non-rejection allograft injury, false positive dd-cfDNA, and false negative EMB. In instances where percentages of dd-cfDNA are elevated despite a negative biopsy, it is imperative to systematically consider these possibilities. This should include an initial assessment of technical reasons for dd-cfDNA fraction to be falsely elevated, such as improper handling of the sample, artifactual increase due to recent cardiac procedure such as biopsy or pacemaker implantation, or even after vigorous exercise (19, 28, 29). The absolute quantity of cfDNA levels is subject to large variations and have been reported in leukocytosis, systemic lupus erythematosus, myocardial infarction, and psychiatric disorders (30–33). Significant reduction in total rd-cfDNA can increase the dd-cfDNA fraction without cardiac injury. Pre-transplant considerations such as an autonomic storm in brain-dead donors or donor coronary artery disease can also lead to baseline cellular injury in the donor heart contributing to elevations in dd-cfDNA fraction in the recipient immediately post HT (34). Additionally, factors such as ischemic time, surgical complexity, pulmonary vascular resistance, and crossmatch status are speculated to lead to elevations in dd-cfDNA immediately post HT. To mitigate or reduce the likelihood of false positives, centers have adopted protocols where the timing of initial dd-cfDNA testing occurs 28 days following transplant (see Figure 1) (19).
If there are no such technical reasons, then a closer look at the immunosuppression trough levels and medication adherence is critical. Viral infections, most commonly cytomegalovirus (CMV), are associated with increased dd-cfDNA levels (27, 35, 36). CMV frequently causes leukopenia which may lead to reduction in rd-cfDNA and resulting increase in percentage of dd-cfDNA. Afzal et al. reported 4 clusters in their single center study of patients with elevated dd-cfDNA without biopsy evidence of AR: CMV viremia, non-CMV infections (which included SARS-CoV-2, Ebstein-Barr virus, and hepatitis B infection), females (with 40% having de novo DSA), and lastly patients with right ventricular dysfunction (27). Of note, ACR 1R was present in all patients with RV dysfunction, 80% of females, and none in the non-CMV infection group. It is unclear whether any patients with ACR 1R progressed to higher-grade rejection, but it will be important to study in future larger multicenter studies as currently most episodes of ACR 1R are not treated. However, in the D-OAR study by Kanwar et al., CMV infections were not associated with significantly higher percentages of dd-cfDNA as compared to those without CMV infection (36). Our group has observed transient elevations in dd-cfDNA immediately following vaccinations.
Elevated dd-cfDNA fraction may reflect ischemic injury secondary to cardiac allograft vasculopathy (CAV) (34). Therefore, further ischemic evaluation with coronary angiography or noninvasive techniques should be considered. Finally, myocarditis is a possible etiology of non-rejection myocardial injury. Elevated percentage of dd-cfDNA due to toxoplasmosis has been described (37). Recurrent sarcoidosis and giant cell myocarditis following transplant are well-established, presumably they could also cause an elevated dd-cfDNA fraction. A high index of suspicion is required to make these diagnoses along with close collaboration with pathologists. To evaluate these possibilities, the incorporation of supplemental diagnostic techniques is warranted, as outlined in the central illustration (Figure 2). We, briefly discuss the role of multimodal diagnostics in evaluation of such cases when dd-cfDNA fraction is elevated in the absence of biopsy-proven rejection.
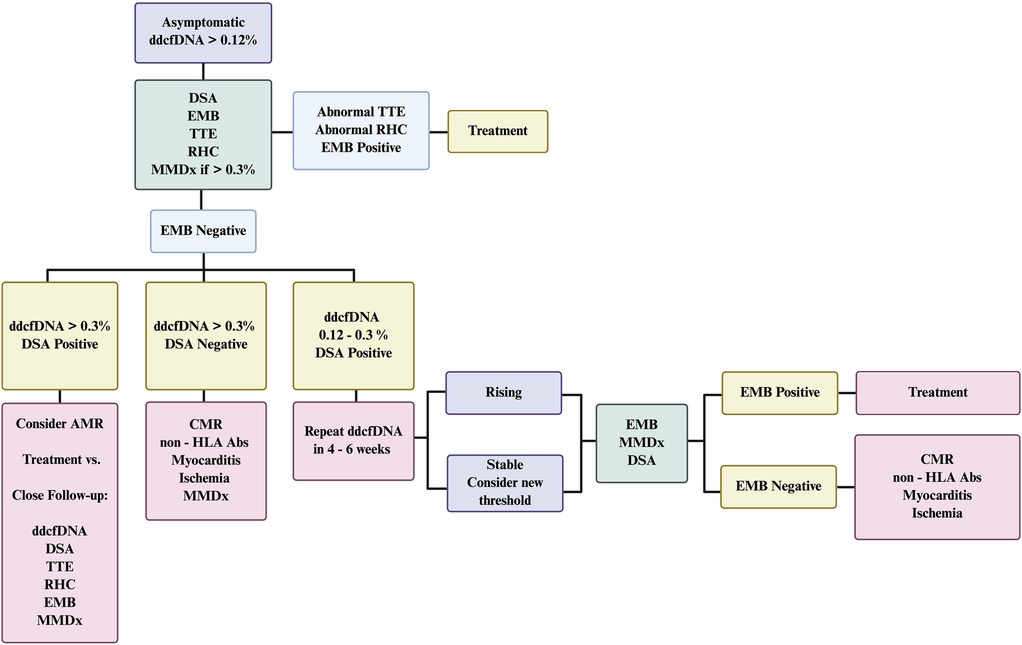
Figure 2. Suggested algorithm for management of asymptomatic patients with elevated dd-cfDNA without biopsy-proven rejection. The figure outlines a stepwise approach to the evaluation and management of an asymptomatic heart transplant recipient with elevated donor-derived cell-free DNA. In cases where dd-cfDNA is elevated but endomyocardial biopsy (EMB) results are negative, the algorithm incorporates a multimodal diagnostic strategy to guide further assessment and clinical decision-making. Close follow-up of possible AMR would include subsequent testing with dd-cfDNA, DSAs, TTE, RHC with EMB and MMDx. AMR, antibody-mediated rejection; CMR, cardiac magnetic resonance; dd-cfDNA, donor-derived cell-free DNA; DSA, donor-specific antibodies; EMB, endomyocardial biopsy; HLA, human leukocyte antigen; MMDx, molecular microscope diagnostic system; RHC, right heart catheterization; TTE, transthoracic echocardiogram.
Additional biomarkers have been investigated to predict AR. Early work using troponin T and I to diagnose AR was not encouraging and recent efforts with high sensitivity assays have been shown to be more promising (38). Similarly, B type natriuretic peptide and its fragments correlated poorly with AR, though patient level trending was noted to be more informative (38). Current ISHLT guidelines state these biomarkers may have a role in a comprehensive strategy for rejection surveillance (38). The combination of dd-cfDNA fraction and gene expression profiling (GEP) has been extensively studied with incremental improvement in predicting AR on biopsy (39). A recent publication combined dd-cfDNA with a readily available biomarker NT-proBNP to improve accuracy in predicting rejection (40). Both these studies relied on EMB as the gold standard for diagnosing rejection, despite its known limitations, and neither study addressed the clinical dilemma of elevated dd-cfDNA levels in the context of reassuring biopsy findings.
The Molecular Microscope Diagnostic System (MMDx, Thermo Fisher Scientific, Waltham, MA) analyzes EMBs using microarrays to quantify mRNA transcripts associated with rejection and injury (41). It is unique in that is uses machine-based learning algorithms to compare each biopsy against a reference database of samples and follows the principle that depending on the type and severity of transplanted organ injury, specific genes are activated and produce unique mRNA patterns (42). While MMDx does not replace histology, it can provide additional information to reconcile discordant results. A contemporary study included more than 3,200 EMBs to identify 7 rejection archetypes as well as non-rejection injury and atrophy-fibrosis (43). The ongoing Trifecta-Heart study seeks to incorporate all three tools (dd-cfDNA, MMDx, and EMB histopathology) to improve accuracy in diagnosing AR (NCT04707872) (44).
Donor specific antibodies (DSAs), primarily targeting donor human leukocyte antigen molecules, are associated with AMR and poor graft survival (45). DSAs may be present at the time of transplant, or may develop following transplant termed de novo. The development of de novo DSA(s) or significant increase in the strength of known DSA(s) coupled with allograft dysfunction even in the presence of a negative EMB may warrant treatment. Moreover, antibodies against non-HLA antigens such as angiotensin II type-1 receptor and others have been linked to AMR and BNR (46, 47); however, monitoring is highly variable across centers, and standardized treatment protocols remain poorly defined.
Cardiac MRI is another promising non-invasive alternative surveillance tool for AR in HT. Besides assessing graft function and valve lesions, parametric mapping with T1 and T2 sequences can potentially enable detection of myocardial edema, fibrosis, and interstitial expansion. In a small study of 60 HT patients, T2 relaxation has been shown to be an independent predictor of AR (48). Another small randomized trial comparing EMB and CMR for AR surveillance demonstrated high diagnostic accuracy of CMR achieving 92% sensitivity and 92% NPV for AR (49). However, CMR is not able to distinguish between different types of AR. CMR can also aid in the diagnosis of myocarditis, including toxoplasmosis, giant cell, and sarcoidosis.
Future directions
As the clinical utility of dd-cfDNA continues to evolve in HT surveillance, several avenues hold promise for advancing the field and addressing current diagnostic limitations. Transitioning from relative quantification or donor fraction to absolute quantification of dd-cfDNA, may enhance diagnostic specificity and reduce variability across diverse clinical scenarios. However, absolute level of dd-cfDNA has also been correlated to AR providing a stable, reproducible measure of donor-specific injury (18, 50–52). This approach may be particularly beneficial in patients with high baseline cfDNA due to concurrent illness or those with multi-organ transplants, where interpreting percentage of dd-cfDNA can be challenging. The primary results of the DEFINE-HT (Development of Non-Invasive Cell-Free DNA to Supplant Invasive Biopsy in Heart Transplantation) study recently reported that Prospera (Natera, Inc, Austin, TX) with donor quantity score (DQS) incorporating both absolute dd-cfDNA and donor fraction, predicted allograft dysfunction 3 times more than EMB and demonstrated strong correlation with clinical outcomes at one year (53).
The use of dd-cfDNA in multiorgan transplant recipients is not well-described. Dd-cfDNA fractions of healthy allografts vary by the transplanted organ, the heart is typically the lowest at < 0.20%, kidney < 0.5% (54), lung < 1% (55), and liver the highest ranging from 5 to 30% (56), which makes selecting a threshold for multiorgan recipients challenging. Intriguingly, dd-cfDNA can be fractionated if recipients have organs from multiple donors, permitting one to follow each organ independently (57). If the transplanted organs are from a single donor however, setting an appropriate threshold must be done on a case-by-case basis, which has been reported in the literature (58) and we have done at our center.
Integration of dd-cfDNA with other emerging diagnostics, such as MMDx, CMR, and DSA profiling, represents an important step toward a precision medicine approach in HT. By combining molecular, immunological, and imaging data, clinicians may be able to generate a more comprehensive and nuanced risk profile for each patient, enabling more timely and targeted interventions. Lastly, longitudinal studies are needed to understand the kinetics of dd-cfDNA over time, establish optimal surveillance intervals, and determine the clinical significance of transient elevations in the absence of rejection.
Conclusion
Elevated dd-cfDNA in the context of a negative EMB poses a significant diagnostic challenge, highlighting the shift from traditional reliance on EMB towards an integrated, multimodal diagnostic approach for AR surveillance in the contemporary era. While dd-cfDNA offers the advantage of non-invasive testing with high NPV, its interpretation must consider the broader clinical context, including other potential causes of graft injury and limitations of the “gold standard” EMB. Future research should focus on refining multimodal diagnostic algorithms to improve early detection and management of AR, ultimately personalizing the diagnostic testing for individual patients based on their clinical characteristics and, thereby improving overall outcomes for patients with HT.
Author contributions
RB: Writing – review & editing, Writing – original draft. AM: Writing – review & editing, Writing – original draft. JK: Writing – original draft, Writing – review & editing.
Funding
The author(s) declare that no financial support was received for the research and/or publication of this article.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Abbreviations
ACR, acute cellular rejection; AMR, antibody-mediated rejection; AR, acute rejection; BNR, biopsy negative rejection; CMR, cardiac magnetic resonance; CMV, cytomegalovirus; dd-cfDNA, donor-derived, cell-free DNA; DF, donor fraction; DQS, donor quantity score; DSA, donor-specific antigen antibody; EMB, endomyocardial biopsy; HT, heart transplant; MMDx, molecular microscope diagnostic system; Rd-cfDNA, recipient-derived cfDNA; SNP, single nucleotide polymorphism.
References
1. Chambers DC, Cherikh WS, Harhay MO, Hayes D Jr, Hsich E, Khush KK, et al. The international thoracic organ transplant registry of the international society for heart and lung transplantation: thirty-sixth adult lung and heart-lung transplantation report-2019; focus theme: donor and recipient size match. J Heart Lung Transplant. (2019) 38(10):1042–55. doi: 10.1016/j.healun.2019.08.001
2. Colvin MM, Cook JL, Chang P, Francis G, Hsu DT, Kiernan MS, et al. Antibody-mediated rejection in cardiac transplantation: emerging knowledge in diagnosis and management: a scientific statement from the American Heart Association. Circulation. (2015) 131(18):1608–39. doi: 10.1161/CIR.0000000000000093
3. Caves PK, Stinson EB, Billingham M, Shumway NE. Percutaneous transvenous endomyocardial biopsy in human heart recipients. Experience with a new technique. Ann Thorac Surg. (1973) 16(4):325–36. doi: 10.1016/S0003-4975(10)65002-3
4. Stewart S, Winters GL, Fishbein MC, Tazelaar HD, Kobashigawa J, Abrams J, et al. Revision of the 1990 working formulation for the standardization of nomenclature in the diagnosis of heart rejection. J Heart Lung Transplant. (2005) 24(11):1710–20. doi: 10.1016/j.healun.2005.03.019
5. Berry GJ, Burke MM, Andersen C, Bruneval P, Fedrigo M, Fishbein MC, et al. The 2013 international society for heart and lung transplantation working formulation for the standardization of nomenclature in the pathologic diagnosis of antibody-mediated rejection in heart transplantation. J Heart Lung Transplant. (2013) 32(12):1147–62. doi: 10.1016/j.healun.2013.08.011
6. Baraldi-Junkins C, Levin HR, Kasper EK, Rayburn BK, Herskowitz A, Baughman KL. Complications of endomyocardial biopsy in heart transplant patients. J Heart Lung Transplant. (1993) 12(1 Pt 1):63–7.8443204
7. Cooper LT, Baughman KL, Feldman AM, Frustaci A, Jessup M, Kuhl U, et al. The role of endomyocardial biopsy in the management of cardiovascular disease. Circulation. (2007) 116(19):2216–33. doi: 10.1161/CIRCULATIONAHA.107.186093
8. Galeone A, Bernabei A, Pesarini G, Lucchetti R, Onorati M, Luciani F, et al. Ten-Year experience with endomyocardial biopsy after orthotopic heart transplantation: comparison between trans-jugular and trans-femoral approach. J Cardiovasc Dev Dis. (2024) 11(4):115. doi: 10.3390/jcdd11040115
9. From AM, Maleszewski JJ, Rihal CS. Current status of endomyocardial biopsy. Mayo Clin Proc. (2011) 86(11):1095–102. doi: 10.4065/mcp.2011.0296
10. Beck-da-Silva L, Bridi LH, Matte BS, Valle FH. Endomyocardial biopsy using rigid bioptome technique and the risk of tricuspid regurgitation after heart transplantation. Arq Bras Cardiol. (2024) 121(10):e20240223. doi: 10.36660/abc.20240223
11. Wiklund L, Caidahl K, Kjellström C, Nilsson B, Svensson G, Berglin E. Tricuspid valve insufficiency as a complication of endomyocardial biopsy. Transpl Int. (1992) 5(Suppl 1):S255–8. doi: 10.1111/tri.1992.5.s1.255
12. Bermpeis K, Esposito G, Gallinoro E, Paolisso P, Bertolone DT, Fabbricatore D, et al. Safety of right and left ventricular endomyocardial biopsy in heart transplantation and cardiomyopathy patients. JACC Heart Fail. (2022) 10(12):963–73. doi: 10.1016/j.jchf.2022.08.005
13. Mehta A, Goldberg J, Bagchi P, Marboe C, Shah KB, Najjar SS, et al. Pathologist interrater reliability and clinical implications of elevated donor-derived cell-free DNA beyond heart transplant rejection, on behalf of the GRAfT investigators. J Heart Lung Transplant. (2025) 44(5):803–12. doi: 10.1016/j.healun.2024.10.006
14. Marboe CC, Billingham M, Eisen H, Deng MC, Baron H, Mehra M, et al. Nodular endocardial infiltrates (quilty lesions) cause significant variability in diagnosis of ISHLT grade 2 and 3A rejection in cardiac allograft recipients. J Heart Lung Transplant. (2005) 24(7, Supplement):S219–S26. doi: 10.1016/j.healun.2005.04.001
15. Tang Z, Kobashigawa J, Rafiei M, Stern LK, Hamilton M. The natural history of biopsy-negative rejection after heart transplantation. J Transplant. (2013) 2013:236720. doi: 10.1155/2013/236720
16. Holzhauser L, DeFilippis EM, Nikolova A, Byku M, Contreras JP, De Marco T, et al. The End of endomyocardial biopsy?: a practical guide for noninvasive heart transplant rejection surveillance. JACC Heart Fail. (2023) 11(3):263–76. doi: 10.1016/j.jchf.2022.11.002
17. De Vlaminck I, Valantine HA, Snyder TM, Strehl C, Cohen G, Luikart H, et al. Circulating cell-free DNA enables noninvasive diagnosis of heart transplant rejection. Sci Transl Med. (2014) 6(241):241ra77. doi: 10.1126/scitranslmed.3007803
18. Khush KK, Patel J, Pinney S, Kao A, Alharethi R, DePasquale E, et al. Noninvasive detection of graft injury after heart transplant using donor-derived cell-free DNA: a prospective multicenter study. Am J Transplant. (2019) 19(10):2889–99. doi: 10.1111/ajt.15339
19. Agbor-Enoh S, Shah P, Tunc I, Hsu S, Russell S, Feller E, et al. Cell-Free DNA to detect heart allograft acute rejection. Circulation. (2021) 143(12):1184–97. doi: 10.1161/CIRCULATIONAHA.120.049098
20. Kim PJ, Olymbios M, Siu A, Wever Pinzon O, Adler E, Liang N, et al. A novel donor-derived cell-free DNA assay for the detection of acute rejection in heart transplantation. J Heart Lung Transplant. (2022) 41(7):919–27. doi: 10.1016/j.healun.2022.04.002
21. Burnham P, Kim MS, Agbor-Enoh S, Luikart H, Valantine HA, Khush KK, et al. Single-stranded DNA library preparation uncovers the origin and diversity of ultrashort cell-free DNA in plasma. Sci Rep. (2016) 6:27859. doi: 10.1038/srep27859
22. Kustanovich A, Schwartz R, Peretz T, Grinshpun A. Life and death of circulating cell-free DNA. Cancer Biol Ther. (2019) 20(8):1057–67. doi: 10.1080/15384047.2019.1598759
23. Lo YM, Tein MS, Pang CC, Yeung CK, Tong KL, Hjelm NM. Presence of donor-specific DNA in plasma of kidney and liver-transplant recipients. Lancet. (1998) 351(9112):1329–30. doi: 10.1016/S0140-6736(05)79055-3
24. Kobashigawa JA, Khush K, Teuteberg J, Song M, Grskovic M, Hiller D, et al. Initial analysis of the donor-derived cell-free DNA - outcomes AlloMap registry (D-OAR) study in heart transplant recipients undergoing surveillance for rejection. J Heart Lung Transplant. (2016) 35(4):S33. doi: 10.1016/j.healun.2016.01.089
25. Keller M, Agbor-Enoh S. Donor-Derived cell-free DNA for acute rejection monitoring in heart and lung transplantation. Curr Transplant Rep. (2021) 8(4):351–8. doi: 10.1007/s40472-021-00349-8
26. Ortiz-Bautista C, Fernández-Avilés F, Delgado Jiménez JF. Serum biomarkers of acute rejection: towards precision medicine in heart transplant. J Heart Lung Transplant. (2021) 40(10):1090–7. doi: 10.1016/j.healun.2021.07.002
27. Afzal A, Alam A, van Zyl JS, Zafar H, Felius J, Hall SA, et al. Observed elevated donor-derived cell free DNA in orthotopic heart transplant recipients without clinical evidence of rejection. Clin Transplant. (2022) 36(3):e14549. doi: 10.1111/ctr.14549
28. Fridlich O, Peretz A, Fox-Fisher I, Pyanzin S, Dadon Z, Shcolnik E, et al. Elevated cfDNA after exercise is derived primarily from mature polymorphonuclear neutrophils, with a minor contribution of cardiomyocytes. Cell Rep Med. (2023) 4(6):101074. doi: 10.1016/j.xcrm.2023.101074
29. Atamaniuk J, Vidotto C, Kinzlbauer M, Bachl N, Tiran B, Tschan H. Cell-free plasma DNA and purine nucleotide degradation markers following weightlifting exercise. Eur J Appl Physiol. (2010) 110(4):695–701. doi: 10.1007/s00421-010-1532-5
30. Edwards RL, Menteer J, Lestz RM, Baxter-Lowe LA. Cell-free DNA as a solid-organ transplant biomarker: technologies and approaches. Biomark Med. (2022) 16(5):401–15. doi: 10.2217/bmm-2021-0968
31. Atamaniuk J, Hsiao YY, Mustak M, Bernhard D, Erlacher L, Fodinger M, et al. Analysing cell-free plasma DNA and SLE disease activity. Eur J Clin Invest. (2011) 41(6):579–83. doi: 10.1111/j.1365-2362.2010.02435.x
32. Lippi G, Sanchis-Gomar F, Cervellin G. Cell-free DNA for diagnosing myocardial infarction: not ready for prime time. Clin Chem Lab Med. (2015) 53(12):1895–901. doi: 10.1515/cclm-2015-0252
33. Melamud MM, Buneva VN, Ermakov EA. Circulating cell-free DNA levels in psychiatric diseases: a systematic review and meta-analysis. Int J Mol Sci. (2023) 24(4):3402. doi: 10.3390/ijms24043402
34. Hayward C. Cardiac allograft injuries: a review of approaches to a common dilemma, with emphasis on emerging techniques. Int J Heart Fail. (2022) 4(3):123–35. doi: 10.36628/ijhf.2021.0042
35. Alam AH, Van Zyl J, Shakoor HI, Farsakh D, Abdelrehim AB, Maliakkal N, et al. The impact of active cytomegalovirus infection on donor-derived cell-free DNA testing in heart transplant recipients. Clin Transplant. (2024) 38(3):e15287. doi: 10.1111/ctr.15287
36. Kanwar MK, Khush KK, Pinney S, Sherman C, Hall S, Teuteberg J, et al. Impact of cytomegalovirus infection on gene expression profile in heart transplant recipients. J Heart Lung Transplant. (2021) 40(2):101–7. doi: 10.1016/j.healun.2020.11.008
37. Backus B, Tanna S, Kennedy JL, Cochrane A, McCulloch M, Topor M, et al. Straight from the heart: late onset cardiac toxoplasmosis in a heart transplant recipient. J Heart Lung Transplant. (2025) 44(4):S301–S2. doi: 10.1016/j.healun.2025.02.636
38. Velleca A, Shullo MA, Dhital K, Azeka E, Colvin M, DePasquale E, et al. The international society for heart and lung transplantation (ISHLT) guidelines for the care of heart transplant recipients. J Heart Lung Transplant. (2023) 42(5):e1–e141. doi: 10.1016/j.healun.2022.10.015
39. Khush K, Hall S, Kao A, Raval N, Dhingra R, Shah P, et al. Surveillance with dual noninvasive testing for acute cellular rejection after heart transplantation: outcomes from the surveillance HeartCare outcomes registry. J Heart Lung Transplant. (2024) 43(9):1409–21. doi: 10.1016/j.healun.2024.05.003
40. Jiménez-Blanco M, Crespo-Leiro MG, García-Cosío Carmena MD, Gómez Bueno M, López-Vilella R, Ortiz-Bautista C, et al. Donor-derived cell-free DNA as a new biomarker for cardiac allograft rejection: a prospective study (FreeDNA-CAR). J Heart Lung Transplant. (2025) 44(4):560–9. doi: 10.1016/j.healun.2024.11.009
41. Halloran PF, Potena L, Van Huyen JD, Bruneval P, Leone O, Kim DH, et al. Building a tissue-based molecular diagnostic system in heart transplant rejection: the heart molecular microscope diagnostic (MMDx) system. J Heart Lung Transplant. (2017) 36(11):1192–200. doi: 10.1016/j.healun.2017.05.029
42. Benck L, Sato T, Kobashigawa J. Molecular diagnosis of rejection in heart transplantation. Circ J. (2022) 86(7):1061–7. doi: 10.1253/circj.CJ-21-0591
43. Halloran PF, Madill-Thomsen K, Aliabadi-Zuckermann AZ, Cadeiras M, Crespo-Leiro MG, Depasquale EC, et al. Redefining the molecular rejection states in 3230 heart transplant biopsies: relationships to parenchymal injury and graft survival. Am J Transplant. (2024) 24(8):1414–26. doi: 10.1016/j.ajt.2024.03.031
44. Uo A. Trifecta-Heart cfDNA-MMDx Study 2021. Available online at: https://clinicaltrials.gov/ct2/show/NCT04707872 (Accessed April 04, 2025).
45. Zhang Q, Hickey M, Drogalis-Kim D, Zheng Y, Gjertson D, Cadeiras M, et al. Understanding the correlation between DSA, complement activation, and antibody-mediated rejection in heart transplant recipients. Transplantation. (2018) 102(10):e431–e8. doi: 10.1097/TP.0000000000002333
46. Fedrigo M, Berry GJ, Coutance G, Reed EF, Lin C-Y, Giarraputo A, et al. Report of the 2022 banff heart concurrent: focus on non-human leukocyte antigen antibodies in rejection and the pathology of rejection. Am J Transplant. (2024) 24(4):533–41. doi: 10.1016/j.ajt.2023.10.004
47. Kobashigawa JA, Patel J, Kittleson M, Kransdorf E, Dimbil S, Levine R, et al. Non-HLA antibodies as a cause for biopsy negative rejection: is it worth testing for them? J Heart Lung Transplant. (2019) 38(4):S87. doi: 10.1016/j.healun.2019.01.200
48. Butler CR, Savu A, Bakal JA, Toma M, Thompson R, Chow K, et al. Correlation of cardiovascular magnetic resonance imaging findings and endomyocardial biopsy results in patients undergoing screening for heart transplant rejection. J Heart Lung Transplant. (2015) 34(5):643–50. doi: 10.1016/j.healun.2014.12.020
49. Anthony C, Imran M, Pouliopoulos J, Emmanuel S, Iliff J, Liu Z, et al. Cardiovascular magnetic resonance for rejection surveillance after cardiac transplantation. Circulation. (2022) 145(25):1811–24. doi: 10.1161/CIRCULATIONAHA.121.057006
50. Richmond ME, Zangwill SD, Kindel SJ, Deshpande SR, Schroder JN, Bichell DP, et al. Donor fraction cell-free DNA and rejection in adult and pediatric heart transplantation. J Heart Lung Transplant. (2020) 39(5):454–63. doi: 10.1016/j.healun.2019.11.015
51. Böhmer J, Wåhlander H, Karason K, Sunnegårdh J, Wasslavik C, Jonsson M, et al. Clinical examples of the additive value of absolute quantification of cell-free DNA after heart transplantation. Clin Transplant. (2024) 38(10):e15477. doi: 10.1111/ctr.15477
52. Böhmer J, Wasslavik C, Andersson D, Ståhlberg A, Jonsson M, Wåhlander H, et al. Absolute quantification of donor-derived cell-free DNA in pediatric and adult patients after heart transplantation: a prospective study. Transpl Int. (2023) 36:11260. doi: 10.3389/ti.2023.11260
53. Shah P, Schlendorf KH, Kim PJ, Kennedy JL, Wever-Pinzon O, Nativi Nicolau JN, et al. Rationale, design and patient characteristics from the (DEFINE-HT) study. J Heart Lung Transplant. (2024) 43(4):S204. doi: 10.1016/j.healun.2024.02.1049
54. Bu L, Gupta G, Pai A, Anand S, Stites E, Moinuddin I, et al. Clinical outcomes from the assessing donor-derived cell-free DNA monitoring insights of kidney allografts with longitudinal surveillance (ADMIRAL) study. Kidney Int. (2022) 101(4):793–803. doi: 10.1016/j.kint.2021.11.034
55. Kattih Z, Aryal S. Using a combination of biomarkers to monitor allograft dysfunction in lung transplant recipients. Front Transplant. (2025) 4–2025:1574898. doi: 10.3389/frtra.2025.1574898
56. Zhong Y, Hu X, Li X, Qiao Y, Li H, Zhou S, et al. Advances and challenges in the application of donor-derived cell-free DNA for diagnosis and treatment in liver transplantation: a narrative review. BMC Surg. (2025) 25(1):203. doi: 10.1186/s12893-025-02911-y
57. Pettersson L, Frischknecht L, Westerling S, Ramezanali H, Weidmann L, Lopez KC, et al. Detection of donor-derived cell-free DNA in the setting of multiple kidney transplantations. Front Immunol. (2024) 15–2024:1282521. doi: 10.3389/fimmu.2024.1282521
Keywords: biomarkers, transplantation, rejection, monitoring, donor-specific antibodies, cell-free DNA, gene expression
Citation: Bahniwal RK, Mehta A and Kennedy JLW (2025) Diagnostic approach to elevated dd-cfDNA with reassuring EMB in heart transplantation. Front. Transplant. 4:1623514. doi: 10.3389/frtra.2025.1623514
Received: 6 May 2025; Accepted: 7 July 2025;
Published: 14 August 2025.
Edited by:
Reginald Gohh, Rhode Island Hospital, United StatesReviewed by:
Garrie Haas, The Ohio State University, United StatesCopyright: © 2025 Bahniwal, Mehta and Kennedy. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jamie L. W. Kennedy, amFtaWUua2VubmVkeUBpbm92YS5vcmc=
†These authors have contributed equally to this work and share first authorship
 Rupinder K. Bahniwal
Rupinder K. Bahniwal Aditya Mehta
Aditya Mehta Jamie L. W. Kennedy
Jamie L. W. Kennedy