In the original article, there was a mistake in Table 1 as published. The references indicated in row B are wrong. The corrected Table 1 appears in the attached below.
Table 1
| Integration strategies | Advantages | Disadvantages | References | ||
|---|---|---|---|---|---|
| A | Endogenous locus | 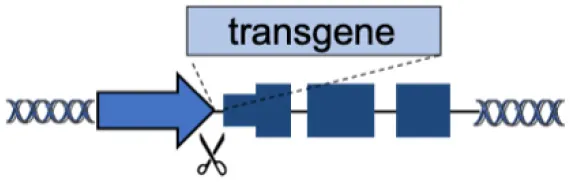 | Physiological transgene expression Corrects multiple mutations | Gene-specific strategy Limited to gene body mutations | Urnov et al., 2005; Lombardo et al., 2007; Li et al., 2011; Genovese et al., 2014; Voit et al., 2014; Dever et al., 2016; Hubbard et al., 2016; Schiroli et al., 2017; Sweeney et al., 2017; Kuo et al., 2018; Wang et al., 2019; Rai et al., 2020; Wang L. et al., 2020 |
| B | Superactive promoters (ALB, HBA) | 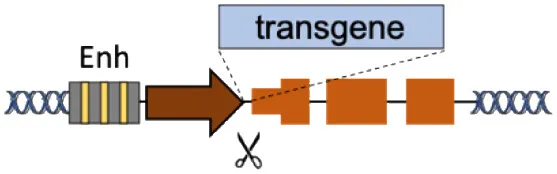 | Accommodates different transgenes Supraphysiological expression Few integrations required | Partial gene disruption Limited to non-cell autonomous disorders Extensive validation required | Barzel et al., 2015; Sharma et al., 2015; Davidoff and Nathwani, 2016; Laoharawee et al., 2018; Chen et al., 2019; Conway et al., 2019; De Caneva et al., 2019; Ou et al., 2019, 2020; Zhang et al., 2019; Wang Q. et al., 2020 |
| C | Tolerant to integration (AAVS1, CCR5, Rosa26) | 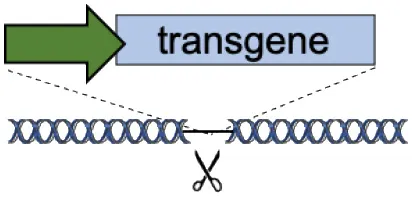 | Accommodates different transgenes | Artificial promoters required Variable expression | De Ravin et al., 2016; Diez et al., 2017; Stephens et al., 2018, 2019; Gomez-Ospina et al., 2019; Scharenberg et al., 2020 |
| D | Chromatin domains (NAD) | 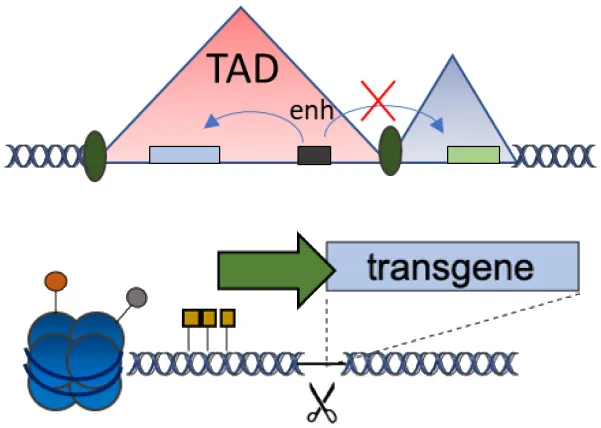 | Fine gene regulation Far from oncogenic genes | No proof-of-principle in clinically relevant models | Schenkwein et al., 2020 |
| E | Disease-modifier genes (CCR5, HBA) | 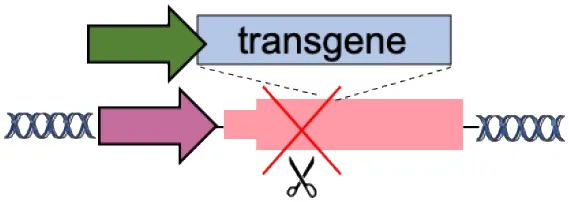 | Improve therapeutic effect Lower therapeutic threshold | Extensive validation required Limited to well-known diseases | Voit et al., 2013; Wiebking et al., 2018 |
| F | Specificity Exchange (TCR, BCR) | 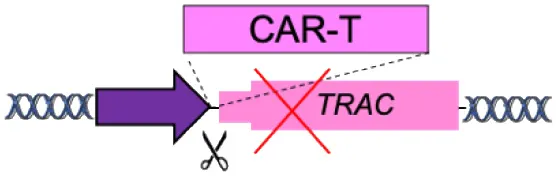 | Improved CAR expression and potency | Off-targets Translocations risk (for multiple edits) | Eyquem et al., 2017; MacLeod et al., 2017; Greiner et al., 2019; Hartweger et al., 2019; Moffett et al., 2019; Voss et al., 2019 |
(A–F) The advantages and disadvantages of different integration strategies.
Scissors: nuclease; Solid arrows: promoters; Enh, enhancers; TAD, topologically associating; d, domain; Solid ovals: histone modifications; Solid squares: DNA modifications.
The authors apologize for this error and state that this does not change the scientific conclusions of the article in any way. The original article has been updated.
References
1
BarzelA.PaulkN. K.ShiY.HuangY.ChuK.ZhangF.et al. (2015). Promoterless gene targeting without nucleases ameliorates haemophilia B in mice. Nature517, 360–364. 10.1038/nature13864
2
ChenH.ShiM.GilamA.ZhengQ.ZhangY.AfrikanovaI.et al. (2019). Hemophilia A ameliorated in mice by CRISPR-based in vivo genome editing of human factor. Sci. Rep. 9:16838. 10.1038/s41598-019-53198-y
3
ConwayA.MendelM.KimK.McGovernK.BoykoA.ZhangL.et al. (2019). Non-viral delivery of zinc finger nuclease mRNA enables highly efficient in vivo genome editing of multiple therapeutic gene targets. Mol. Ther. 27, 866–877. 10.1016/j.ymthe.2019.03.003
4
DavidoffA. M.NathwaniA. C. (2016). Genetic targeting of the albumin locus to treat Hemophilia. N. Engl. J. Med. 374, 1288–1290. 10.1056/NEJMcibr1600347
5
De CanevaA.PorroF.BortolussiG.SolaR.LisjakM.BarzelA.et al. (2019). Coupling AAV-mediated promoterless gene targeting to SaCas9 nuclease to efficiently correct liver metabolic diseases. JCI Insight. 5:128863. 10.1172/jci.insight.128863
6
De RavinS. S.ReikA.LiuP. Q.LiL.WuX.SuL.et al. (2016). Targeted gene addition in human CD34(+) hematopoietic cells for correction of X-linked chronic granulomatous disease. Nat. Biotechnol. 34, 424–429. 10.1038/nbt.3513
7
DeverD. P.BakR. O.ReinischA.CamarenaJ.WashingtonG.NicolasC. E.et al. (2016). CRISPR/Cas9 beta-globin gene targeting in human haematopoietic stem cells. Nature539, 384–389. 10.1038/nature20134
8
DiezB.GenoveseP.Roman-RodriguezF. J.AlvarezL.SchiroliG.UgaldeL.et al. (2017). Therapeutic gene editing in CD34(+) hematopoietic progenitors from Fanconi anemia patients. EMBO Mol. Med. 9, 1574–1588. 10.15252/emmm.201707540
9
EyquemJ.Mansilla-SotoJ.GiavridisT.van der StegenS. J.HamiehM.CunananK. M.et al. (2017). Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection. Nature543, 113–117. 10.1038/nature21405
10
GenoveseP.SchiroliG.EscobarG.TomasoT. D.FirritoC.CalabriaA.et al. (2014). Targeted genome editing in human repopulating haematopoietic stem cells. Nature510, 235–240. 10.1038/nature13420
11
Gomez-OspinaN.ScharenbergS. G.MostrelN.BakR. O.MantriS.QuadrosR. M.et al. (2019). Human genome-edited hematopoietic stem cells phenotypically correct Mucopolysaccharidosis type. Nat. Commun. 10:4045. 10.1038/s41467-019-11962-8
12
GreinerV.Bou PuertoR.LiuS.HerbelC.CarmonaE. M.GoldbergM. S. (2019). CRISPR-mediated editing of the B cell receptor in primary human B cells. iScience12, 369–378. 10.1016/j.isci.2019.01.032
13
HartwegerH.McGuireA. T.HorningM.TaylorJ. J.DosenovicP.YostD.et al. (2019). HIV-specific humoral immune responses by CRISPR/Cas9-edited B cells. J. Exp. Med. 216, 1301–1310. 10.1084/jem.20190287
14
HubbardN.HaginD.SommerK.SongY.KhanI.CloughC.et al. (2016). Targeted gene editing restores regulated CD40L function in X-linked hyper-IgM syndrome. Blood127, 2513–2522. 10.1182/blood-2015-11-683235
15
KuoC. Y.LongJ. D.Campo-FernandezB.de OliveiraS.CooperA. R.RomeroZ.et al. (2018). Site-specific gene editing of human hematopoietic stem cells for X-linked hyper-IgM syndrome. Cell Rep. 23, 2606–2616. 10.1016/j.celrep.2018.04.103
16
LaoharaweeK.DeKelverR. C.Podetz-PedersenK. M.RohdeM.SproulS.NguyenH. O.et al. (2018). Dose-dependent prevention of metabolic and neurologic disease in murine MPS II by ZFN-mediated in vivo genome editing. Mol. Ther. 26, 1127–1136. 10.1016/j.ymthe.2018.03.002
17
LiH.HaurigotV.DoyonY.LiT.WongS. Y.BhagwatA. S.et al. (2011). In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature475, 217–221. 10.1038/nature10177
18
LombardoA.GenoveseP.BeausejourC. M.ColleoniS.LeeY. L.KimK. A.et al. (2007). Gene editing in human stem cells using zinc finger nucleases and integrase-defective lentiviral vector delivery. Nat. Biotechnol. 25, 1298–1306. 10.1038/nbt1353
19
MacLeodD. T.AntonyJ.MartinA. J.MoserR. J.HekeleA.WetzelK. J.et al. (2017). Integration of a CD19 CAR into the TCR alpha chain locus streamlines production of allogeneic gene-edited CAR T cells. Mol. Ther. 25, 949–961. 10.1016/j.ymthe.2017.02.005
20
MoffettH. F.HarmsC. K.FitzpatrickK. S.TooleyM. R.BoonyaratanakornkitJ.TaylorJ. J. (2019). B cells engineered to express pathogen-specific antibodies protect against infection. Sci. Immunol. 4:aax0644. 10.1126/sciimmunol.aax0644
21
OuL.DeKelverR. C.RohdeM.TomS.RadekeR.St MartinS. J.et al. (2019). ZFN-mediated in vivo genome editing corrects murine hurler syndrome. Mol. Ther. 27, 178–187. 10.1016/j.ymthe.2018.10.018
22
OuL.PrzybillaM. J.AhlatO.KimS.OvernP.JarnesJ.et al. (2020). A highly efficacious PS gene editing system corrects metabolic and neurological complications of Mucopolysaccharidosis type I. Mol. Ther. 28, 1442–1454. 10.1016/j.ymthe.2020.03.018
23
RaiR.RomitoM.RiversE.TurchianoG.BlattnerG.VetharoyW.et al. (2020). Targeted gene correction of human hematopoietic stem cells for the treatment of Wiskott - Aldrich Syndrome. Nat. Commun. 11:4034. 10.1038/s41467-020-17626-2
24
ScharenbergS. G.PolettoE.LucotK. L.ColellaP.SheikaliA.MontineT. J.et al. (2020). Engineering monocyte/macrophage-specific glucocerebrosidase expression in human hematopoietic stem cells using genome editing. Nat. Commun. 11:3327. 10.1038/s41467-020-17148-x
25
SchenkweinD.AfzalS.NousiainenA.SchmidtM.Yla-HerttualaS. (2020). Efficient nuclease-directed integration of lentivirus vectors into the human ribosomal DNA locus. Mol. Ther. 28, 1858–1875. 10.1016/j.ymthe.2020.05.019
26
SchiroliG.FerrariS.ConwayA.JacobA.CapoV.AlbanoL.et al. (2017). Preclinical modeling highlights the therapeutic potential of hematopoietic stem cell gene editing for correction of SCID-X1. Sci. Transl. Med. 9:aan0820. 10.1126/scitranslmed.aan0820
27
SharmaR.AnguelaX. M.DoyonY.WechslerT.DeKelverR. C.SproulS.et al. (2015). In vivo genome editing of the albumin locus as a platform for protein replacement therapy. Blood126, 1777–1784. 10.1182/blood-2014-12-615492
28
StephensC. J.KashentsevaE.EverettW.KaliberovaL.CurielD. T. (2018). Targeted in vivo knock-in of human alpha-1-antitrypsin cDNA using adenoviral delivery of CRISPR/Cas9. Gene Ther. 25, 139–156. 10.1038/s41434-018-0003-1
29
StephensC. J.LauronE. J.KashentsevaE.LuZ. H.YokoyamaW. M.CurielD. T. (2019). Long-term correction of hemophilia B using adenoviral delivery of CRISPR/Cas9. J. Contr. Release. 298, 128–141. 10.1016/j.jconrel.2019.02.009
30
SweeneyC. L.ZouJ.ChoiU.MerlingR. K.LiuA.BodanskyA.et al. (2017). Targeted repair of CYBB in X-CGD iPSCs requires retention of intronic sequences for expression and functional correction. Mol. Ther. 25, 321–330. 10.1016/j.ymthe.2016.11.012
31
UrnovF. D.MillerJ. C.LeeY. L.BeausejourC. M.RockJ. M.AugustusS.et al. (2005). Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature435, 646–651. 10.1038/nature03556
32
VoitR. A.HendelA.Pruett-MillerS. M.PorteusM. H. (2014). Nuclease-mediated gene editing by homologous recombination of the human globin locus. Nucl. Acids Res. 42, 1365–1378. 10.1093/nar/gkt947
33
VoitR. A.McMahonM. A.SawyerS. L.PorteusM. H. (2013). Generation of an HIV resistant T-cell line by targeted “stacking” of restriction factors. Mol. Ther. 21, 786–795. 10.1038/mt.2012.284
34
VossJ. E.Gonzalez-MartinA.AndrabiR.FullerR. P.MurrellB.McCoyL. E.et al. (2019). Reprogramming the antigen specificity of B cells using genome-editing technologies. Elife8:42995. 10.7554/eLife.42995
35
WangL.YangY.BretonC.BellP.LiM.ZhangJ.et al. (2020). A mutation-independent CRISPR-Cas9-mediated gene targeting approach to treat a murine model of ornithine transcarbamylase deficiency. Sci. Adv. 6:eaax5701. 10.1126/sciadv.aax5701
36
WangL.YangY.BretonC. A.WhiteJ.ZhangJ.CheY.et al. (2019). CRISPR/Cas9-mediated in vivo gene targeting corrects hemostasis in newborn and adult factor IX-knockout mice. Blood133, 2745–2752. 10.1182/blood.2019000790
37
WangQ.ZhongX.LiQ.SuJ.LiuY.MoL.et al. (2020). CRISPR-Cas9-mediated in vivo gene integration at the albumin locus recovers hemostasis in neonatal and adult hemophilia B mice. Mol. Ther. Methods Clin. Dev. 18, 520–531. 10.1016/j.omtm.2020.06.025
38
WiebkingV.PattersonJ. O.MartinR.ChandaM. K.LeeC. M.SrifaW.et al. (2018). Metabolic engineering generates a transgene-free safety switch for cell therapy. Nat. Biotechnol2020:6. 10.1038/s41587-020-0580-6
39
ZhangJ. P.ChengX. X.ZhaoM.LiG. H.XuJ.ZhangF.et al. (2019). Curing hemophilia A by NHEJ-mediated ectopic F8 insertion in the mouse. Genome Biol. 20:276. 10.1186/s13059-019-1907-9
Summary
Keywords
genome editing, gene therapy, nuclease, CRISPR, targeted integration (TI), knock-in, safe harbor, homologous recombination (HR)
Citation
Pavani G and Amendola M (2021) Corrigendum: Targeted Gene Delivery: Where to Land. Front. Genome Ed. 3:682171. doi: 10.3389/fgeed.2021.682171
Received
17 March 2021
Accepted
06 April 2021
Published
17 May 2021
Volume
3 - 2021
Edited and reviewed by
Pietro Genovese, Boston Children's Hospital and Harvard Medical School, United States
Updates
Copyright
© 2021 Pavani and Amendola.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mario Amendola mamendola@genethon.fr
†Present address: Giulia Pavani, The Children's Hospital of Philadelphia, Raymond G. Perelman Center for Cellular and Molecular Therapeutics, Philadelphia, PA, United States
This article was submitted to Genome Editing in Blood Disorders, a section of the journal Frontiers in Genome Editing
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.