Abstract
Paleo-drainage connections and headwater stream-captures are two main historical processes shaping the distribution of strictly freshwater fishes. Recently, bathymetric-based methods of paleo-drainage reconstruction have opened new possibilities to investigate how these processes have shaped the genetic structure of freshwater organisms. In this context, the present study used paleo-drainage reconstructions and single-locus cluster delimitation analyses to examine genetic structure on the whole distribution of Pareiorhaphis garbei, a ‘near threatened’ armored catfish from the Fluminense freshwater ecoregion in Southeastern Brazil. Sequences of two mitochondrial genes (cytochrome b and cytochrome c oxidase subunit 1) were obtained from five sampling sites in four coastal drainages: Macaé (KAE), São João (SJO), Guapi-Macacu [sub-basins Guapiaçu (GAC) and Guapimirim (GMI)], and Santo Aleixo (SAL). Pronounced genetic structure was found, involving 10 haplotypes for cytB and 6 for coi, with no haplotypes shared between localities. Coalescent-based delineation methods as well as distance-based methods revealed genetic clusters corresponding to each sample site. Paleo-drainage reconstructions showed two putative paleo-rivers: an eastern one connecting KAE and SJO; and a western one merging in the Guanabara Bay (GAC, GMI, and SAL). A disagreement was uncovered between the inferred past riverine connections and current population genetic structure. Although KAE and SJO belong to the same paleo-river, the latter is more closely related to specimens from the Guanabara paleo-river. This discordance between paleo-drainage connections and phylogenetic structure may indicate an ancient stream-capture event in headwaters of this region. Furthermore, all analyses showed high divergence between KAE and the other lineages, suggesting at least one cryptic species in the latter, and that the nominal species should be restricted to the Macaé river basin, its type locality. In this drainage, impacts such as the invasive species and habitat loss can be especially threatening for such species with a narrow range. Our results also suggest that freshwater fishes from headwaters in the Serra do Mar mountains might have different biogeographical patterns than those from the lowlands, indicating a complex and dynamic climatic and geomorphological history.
Introduction
Two alternative hypotheses are typically invoked to explain disjoint distributions encompassing more than one drainage basin of strictly freshwater species: past anastomoses of coastal basins due to marine regression episodes that resulted in paleo-drainage connections or headwater stream-captures caused by tectonic adjustments (Albert and Reis, 2011). Stream capture occurs when a river changes its course and connects with another drainage system as a result of geomorphological changes (Bishop, 1995; Wilkinson et al., 2006), whereas paleo-drainage connections happen when sea levels are lowered by climatic changes, causing basins that previously were isolated to coalesce (Voris, 2000; Dias et al., 2014). To assess how these processes have influenced the genetic structure of freshwater fishes, recently developed GIS-based methods have been used to reconstruct paleo-river systems (Dias et al., 2014; Thomaz et al., 2015, 2017). These methods use topographical and bathymetrical data to reconstruct land exposure, fine-scale depth and steepness of exposed areas (which represent putative riverbeds), as well as putative flow and basin limits (Thomaz et al., 2015). Bathymetric-based reconstructions proved to be highly concordant with the available geological record for paleo-drainages (Dias et al., 2014), supporting GIS-based reconstruction as a powerful and reliable tool for paleo-drainage reconstructions, especially in regions with scarce geological data (Thomaz et al., 2015).
Headwater regions usually exhibit high levels of endemism and microhabitat specialization (Albert et al., 2011; Buckup, 2011). One such example of headwater taxa involves loricariid catfishes of the genus Pareiorhaphis. These rheophilic (fast-water) fishes usually show a narrow distribution, with 20 of 25 described species (80%) restricted to a single basin (Pereira et al., 2017). One of the exceptions within the genus is Pareiorhaphis garbei (Ihering, 1911), a species found in four coastal basins of the Rio de Janeiro State in southeastern Brazil: Macaé, São João, Guapi-Macacu, and Santo Aleixo drainages (Maia et al., 2013). This armored catfish is restricted to clear-water streams with fast-flowing high-oxygen waters and predominance of rocky (bouldered) substrate (Lazzarotto et al., 2007), and these habitats only occur in the headwaters or upper reaches of coastal basins (Maia et al., 2013). The geographic distribution of P. garbei lies entirely within the Atlantic Forest biodiversity hotspot (Myers et al., 2000), which in turn is situated within the Fluminense freshwater ecoregion, a small area comprising about 110 freshwater fish species and a high proportion (42%) of endemic species (Albert et al., 2011).
Due to the intense modification of natural areas by rural and urban expansion, many freshwater fish species in the Fluminense ecoregion are threatened. Previously, P. garbei was classified as an ‘endangered’ species [Ministério do Meio Ambiente [MMA], 2004], but it is currently considered ‘near threatened’ and is, thus, a priority species for research [Instituto Chico Mendes de Conservação da Biodiversidade [ICMBio], 2014]. Deforestation, chemical pesticides, and the introduction of rainbow trout Oncorhynchus mykiss (Walbaum, 1792) were shown to be threats for P. garbei (Pereira and Brito, 2008), even though most records for the species are in protected areas (Maia et al., 2013). However, molecular studies indicate that some taxa putatively distributed along coastal basins of the Fluminense ecoregion may be species complexes with individually narrow distributions (Villa-Verde et al., 2012; Cherobim et al., 2016; Roxo et al., 2017).
Application of molecular phylogenetic analyses and paleo-drainage reconstruction of headwater taxa might reveal their population histories, including information relevant for taxonomy and conservation. Single-locus cluster delineation, often based on mitochondrial DNA, is drastically changing investigations of global biodiversity (Hajibabaei et al., 2007; Pereira L.H. et al., 2013). Clusters revealed by this analysis are considered operational taxonomic units (OTUs). Units that are supported by several clustering methods with different assumptions offer a starting point for taxonomic reevaluation (Kekkonen and Hebert, 2014). This study addresses the genetic structure of P. garbei to infer historical processes that may have influenced the species’ disjointed distribution along coastal basins of the Serra do Mar. In particular, we will address possible paleo-drainage connections and headwater captures. If the lineage patterns are consistent with reconstructed paleo-rivers, then anastomosis of coastal basins by regressive glacial episodes would likely explain the current distribution. However, if the observed phylogeographic pattern is incongruent with the paleo-river reconstruction, then stream capture would likely explain lineage patterns along the headwaters of distinct basins. Furthermore, the species delimitation analyses will provide complementary data for systematic studies, as well as useful information for evaluating the conservation status of P. garbei.
Materials and Methods
Sampling
Specimens of P. garbei were captured in five localities of four hydrographic basins covering the entire known species distribution (Maia et al., 2013): Macaé (KAE), São João (SJO), Guapi-Macacu [sub-basins Guapiaçu (GAC) and Guapimirim (GMI)], and Santo Aleixo (SAL) (Table 1). Whereas the rivers Santo Aleixo and Guapi-Macacu flow into the Guanabara Bay in the same mangrove area (and their river mouths are less than 2 km apart), the São João and Macaé rivers flow to the east coast of Rio de Janeiro State directly into the Atlantic Ocean, about 100 km from Guanabara Bay (Costa, 2014) (Figure 1). Muscle tissues or fin clips were obtained from specimens preserved in absolute ethanol. These individuals are stored at the ichthyological collections of the Universidade Federal do Rio Grande do Norte and of the Departamento de Ecologia of the Universidade Federal do Rio de Janeiro (Table 1).
Table 1
| Locality | Basin | Geographic Coordinates | Altitude (m) | CU | Vouchers (UFRN) | cytB | coi |
|---|---|---|---|---|---|---|---|
| Macaé (KAE) | Macaé | 22°25′50.1″ S 42°32′20.2″ W | 1090 | 1 | 098, 108 | 6 | 6 |
| São João (SJO) | São João | 22°29′54.5″ S 42°30′34.6″ W | 90 | 2 | 010, 1025 | 4 | 10 |
| Guapiaçu (GAC) | Guapi-Macacu | 22°24′53″ S 42°43′37″ W | 180 | 3 | DEPR J8483 | 3 | 5 |
| Guapimirim (GMI) | Guapi-Macacu | 22°29′32.1″S 43°00′03.3″ W | 400 | 3 | 092 | 4 | 4 |
| Santo Aleixo (SAL) | Santo Aleixo | 22°31′29.7″ S 43°01′54.1″ W | 270 | 3 | 103, 111 | 1 | – |
Sampling sites of Pareiorhaphis garbei in the Serra do Mar coastal basins, Rio de Janeiro State, southeastern Brazil.
Conservation Units (CU) are as follows: (1) Três Picos State Park, (2) Bacia do Rio São João Environmental Protection Area, and (3) Petrópolis Environmental Protection Area/Serra dos Órgãos National Park. Vouchers: (UFRN) Universidade Federal do Rio Grande do Norte fish collection, (DEPRJ) Departamento de Ecologia Universidade Federal do Rio de Janeiro fish collection. Columns cytB and coi show number of specimens studied at each gene.
FIGURE 1

(A) Paleo-drainages reconstruction of coastal basins from Serra do Mar mountains, Rio de Janeiro, southeastern Brazil. Dark gray represents the continent at current sea level and light gray a putative exposure of the continental shelf during a sea regression of 125 m. The Atlantic Forest distribution is represented in grey on the inset map. (B) Haplotypes networks of cytochrome b (cytB) and cytochrome oxidase I (coi) in Pareiorhaphis garbei.
Paleo-Drainages Reconstruction
To access paleo-drainages connections during the Last Glacial Maximum (LGM), topographical and bathymetrical data were retrieved from the digital elevation model at 30 arc-second resolution1. This layer was uploaded into ArcGIS10 software, and the Hydrological tools add-in was used to reconstruct the paleo-drainages, following the steps described in Thomaz et al. (2015). First, the area exposed during the LGM (-125 m) was identified using the Contour followed by Mask. Afterwards, the tools Fill, Flow Direction, and Basin were used, respectively, to fill depressions on the surface, identify the steepness within each cell, and define the basin borders. Finally, the Stream order function was used to estimate putative paleo-rivers.
DNA Extraction and Amplification
Genomic DNA samples were extracted using phenol/chloroform/isopropanol with ethanol precipitation protocol (Milligan, 1998), and mitochondrial DNA fragments of genes encoding cytochrome oxidase subunit I (coi) and cytochrome b (cytB) were amplified and sequenced using primers and conditions proposed by Villa-Verde et al. (2012). Overall, 18 and 25 specimens were amplified and sequenced for cytB (1056 bp) and coi (878 bp), respectively (GenBank MG251217-251259) (Table 1).
Phylogenetic Analysis and Species Delimitation
Sequences were edited in MEGA6 (Tamura et al., 2013), then aligned using the MUSCLE algorithm (Edgar, 2004). P-distance (cytB) and Kimura-2-parameter (K2P) (coi) distances for markers were also calculated in MEGA6. A Bayesian phylogenetic reconstruction for both cytB and coi data was done in BEAST v. 1.7 (Drummond et al., 2012) using HKY+G (cytB) and K2P+G (coi) as nucleotide substitution models, as suggested by jModeltest 2 (Darriba et al., 2012). An uncorrelated relaxed lognormal model with estimated rate was used, with ucld.mean parameter set and uniform distribution (0 and 10 as lower and upper boundaries). Remaining parameters were set as default. The length of the MCMC chain was 10,000,000 runs with sampling every 1000 runs. ESS (> 200) values were checked using Tracer v. 1.5 (Rambaut, 2009). The initial 2000 trees were discarded as burn-in period, and a final tree was reconstructed using TreeAnnotator v.1.50. Each haplotype network was inferred using the TCS method on PopART (Leigh and Bryant, 2015). Pareiorhaphis cf. bahianus from Contas river basin in Bahia State was used as out-group in an additional Bayesian phylogeny, using the same parameters described above, for each gene to corroborate the relationships among P. garbei lineages (Supplementary Figure S1).
For increased robustness in delimiting OTUs, five single-locus species-delimitation analyses were performed using both markers separately: single-threshold of Generalized Mixed Yule-Coalescent (sGMYC) (Fujisawa and Barraclough, 2013); multiple-threshold GMYC (mGMYC) (Fujisawa and Barraclough, 2013); Bayesian implementation of Poisson Tree Process (bPTP) (Zhang et al., 2013); multiple rate PTP (mPTP) (Kapli et al., 2017); and Automatic Barcode Gap Discovery (ABGD) (Puillandre et al., 2012). Ultrametric trees generated from cytB and coi data of the Bayesian phylogeny were used as an input file for sGMYC and mGMYC. Previous studies have shown that the clock model and tree prior have low impact on the results of both sGMYC and mGMYC (Tavalera et al., 2013). These two analyses were conducted in R (R Core Team, 2017), using the package SPLITS (Ezard et al., 2009). We re-ran the phylogenetic reconstructions under the same parameters used in BEAST using MrBayes v. 3.2.6 (Huelsenbeck and Ronquist, 2001) to generate a bifurcating phylogram (with branch length representing the number of substitutions) to be the input file for PTPs analyses. bPTP analyses were performed in the online server2. The analysis length was 500,000 generations with sampling of 500 and burn-in of 0.1. Convergence was visualized on the MCMC interactions plots vs. log-likelihood. mPTP was run using the online server3, under the same parameters as for bPTP. ABGD distance-based analyses were run through the software command line, with a gap width value of 1.0 for all the distances available (p-distance, K2P, and Jukes–Cantor). The ABGD delineation taken into account was the one with the P-value of ∼0.01, as advocated by previous studies (Puillandre et al., 2012; Blair and Bryson, 2017). To have a species-delimitation partition based on a genetic distance threshold value, we used cut-off values of 5% divergence for cytB and 2% divergence for coi as indicators of distinct species. These values were based on previous reviews on genetic distances on fish species (Ward, 2009; Kartavtsev, 2011).
Results
A total of 10 cytB and six coi haplotypes were found, none of which were shared among localities. For both mtDNA markers, the haplotype networks showed multiple mutational steps among all haplotypes (Figure 1). Mutations were especially abundant along the branch connecting KAE and the other populations (Figure 1). Genetic p-distances and K2P distances are presented in Table 2. Between KAE and the other localities, divergence values ranged from 7.5 to 8.0% for cytB and 4.9 to 5.0% for coi; between SJO and rivers that flow to the Guanabara Bay (GAC, GMI, and SAL) from 3.6 to 4.4% (cytB) and 2.2% (coi); and, among the watersheds that run into Guanabara Bay, from 2.6 to 3.2% (cytB) and 1.4% (coi) (Table 2).
Table 2
| KAE | SJO | GAC | GMI | SAL | |
|---|---|---|---|---|---|
| Macaé (KAE) | 0.049 | 0.050 | 0.050 | – | |
| São João (SJO) | 0.080 | 0.022 | 0.022 | – | |
| Guapiaçu (GAC) | 0.080 | 0.037 | 0.014 | – | |
| Guapimirim (GMI) | 0.080 | 0.044 | 0.032 | – | |
| Santo Aleixo (SAL) | 0.075 | 0.036 | 0.028 | 0.026 |
P-distances of cytochrome b gene (below diagonal) and K2P distances of cytochrome oxidase subunit I gene (above diagonal) for Pareiorhaphis garbei in the Serra do Mar coastal basins, Rio de Janeiro State, southeastern Brazil.
Our phylogenetic reconstructions showed high genetic structuring within P. garbei, with all sampling sites representing monophyletic clusters in both the cytB and the coi data. According to this result, SJO is more closely related to the Guanabara Bay populations (GAC, GMI, and SAL) than to the KAE. Within the Guanabara Bay watersheds, GMI is more closely related to SAL (from an adjacent basin) than it is to GAC (that belongs to the same drainage, Guapi-Macacu), although supported by lower posterior probabilities (Figure 2).
FIGURE 2
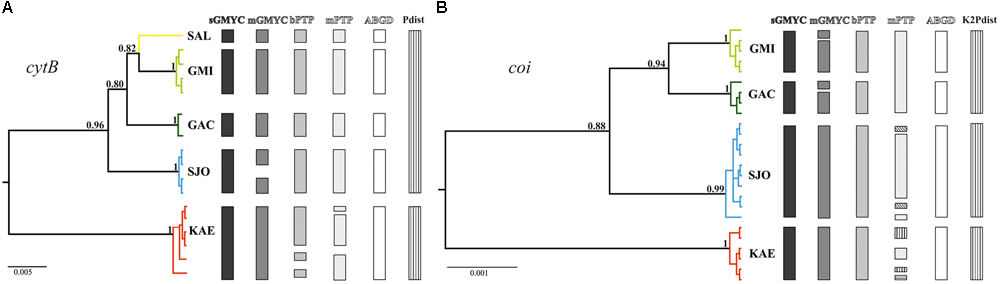
Ultrametric trees generated by the Bayesian phylogenetic reconstructions for (A) cytochrome b (cytB) and (B) cytochrome oxidase I (coi) in Pareiorhaphis garbei. The bars in different colors and patterns represent the operational taxonomic units (OTUs) partitions obtained by the different species delimitation methods.
The sGMYC and ABGD delimitation analyses indicated five distinct OTUs, corresponding to each sampling site for cytB. mGMYC split the SJO clade into two different OTUs, resulting in six genetic clusters. Both PTP methods (bPTP and mPTP) indicated potential distinct lineages within the KAE clade. The strict species threshold (p-distance > 5%) separates KAE from the other sampling sites (Figure 2A). Regarding to the coi reconstruction, four OTUs corresponding to sampling sites were shown as the best genetic partition in three of the five species-delineation methods (sGMYC, bPTP, and ABGD). However, some incongruences were found. Similarly to the cytB reconstruction, mGMYC tended to split lineages, dividing the GAC clade into two different clusters. Furthermore, mPTP found putative distinct genetic lineages within the SJO and KAE clades, resulting in a total of seven OTUs. Using the K2P distance species threshold value (>2%), the Guanabara Bay drainages represent one species while SJO and KAE each represent different putative species (Figure 2B).
Our paleo-drainage reconstruction places Macaé (KAE) and São João (SJO) in the same paleo-drainage (the Macaé-São João paleo-river), whereas Guapiaçu (GAC), Guapimirim (GMI), and Santo Aleixo (SAL) all reside in the Guanabara paleo-river (Figure 1A). This paleo-reconstruction does not corroborate the phylogenetic result, according to which SJO is somewhat more closely related to Guanabara Bay rivers populations than to KAE.
Discussion
Our results revealed a discordance between past paleo-river connections and current genetic structure, which might indicate an ancient stream capture event in the headwaters of the Serra do Mar mountains. According to the paleo-drainage scenario, it was expected that SJO would be closely related to KAE; however, it appears to be a sister group to lineages from the Guanabara paleo-river. This result corroborates the river-capture event suggested by Maia et al. (2013) based on the occurrence of P. garbei in a single tributary adjacent to the headwaters of Macaé river basin. These drainage rearrangements are the result of tectonic reactivation that started in the Paleogene and continues to the present (Ribeiro et al., 2006; Lima and Ribeiro, 2011; Lima et al., 2016).
Although located within the Guapi-Macacu river basin, the water path connecting GAC and GMI is a lowland area where micro- and meso-habitats suitable for P. garbei are absent. The fact that GAC and GMI share no haplotypes, and belong to distinct lineages, reveals that any dispersal of P. garbei through the lower portions of rivers is very limited (and might explain in part why the current genetic structure of P. garbei does not perfectly reflect the paleo-drainage reconstruction).
Along the geographic range of P. garbei, a geographic barrier known as the Cabo Frio Magmatic Lineament (CFML) (Riccomini et al., 2005) may have influenced the evolution and diversification of many freshwater fishes. Based on molecular data, Pereira T.L. et al. (2013) suggested that this barrier had a vicariant effect isolating an eastern lineage (including a clade in the São João and Macaé basins) from a southeastern lineage (from Guanabara Bay to Paranaguá Bay) in the trahira, Hoplias malabaricus (Bloch, 1794). This pattern is also corroborated by the geographic distribution of Atlantirivulus rivulids in lowland areas (Costa, 2014). Although the CFLM seems to be an effective barrier to P. garbei (because different lineages are on its alternate sides), specimens from the São João basin are more closely related to Guanabara Bay drainages than to Macaé drainage, suggesting that different biogeographic forces may apply to headwater versus lowland fishes. Although not exclusively a headwater species, a biogeographic pattern similar to that found in P. garbei is also shown by Hisonotus loricariids, with H. notatus Eigenmann and Eigenmann, 1889 occurring from the São João river basin to the south (including Guanabara Bay’s drainages), whereas H. thayeri Martins and Langeani, 2016 occurs from the Macaé river northward (Martins and Langeani, 2016). Altogether, these various patterns indicate a complex and dynamic history of the coastal basins of the Fluminense ecoregion. Curiously, the CFLM is also the boundary between two marine provinces: the Tropical Southwestern Atlantic and the Warm Temperate Southwestern Atlantic (Spalding et al., 2007).
Pareiorhaphis garbei was the subject of a taxonomic review (Pereira and Reis, 2002; Pereira, 2005) that did not include molecular data, preventing the discovery of deep genetic structure. Several other demersal Neotropical freshwater fishes that display only subtle morphological differences have proved to show substantial genetic divergences that support the description of new and often endemic species (Benine et al., 2009; Melo et al., 2011; Cherobim et al., 2016), and some of these support an ancient split between rivers flowing to the Guanabara Bay versus the São João and Macaé drainages (Villa-Verde et al., 2012; Costa, 2014; Roxo et al., 2017). All molecular analyses herein performed indicate high divergence of the KAE population of P. garbei from all others, strongly suggesting that individuals from these two groups represent different species. If this outcome is formally recognized, P. garbei (sensu stricto) would be confined to the Macaé river basin (its type locality), with another species distributed along the headwaters of São João, Guapi-Macacu, and Santo Aleixo river basins. Moreover, according to high divergence at coi, the São João lineage could also be a distinct species from the one inhabiting drainages entering Guanabara Bay. In either case, P. garbei itself would be limited to Macaé river basin, in which the invasive rainbow trout poses a risk factor (Lazzarotto et al., 2007; Pereira and Brito, 2008). Currently, P. garbei is classified as ‘near threatened’ [Instituto Chico Mendes de Conservação da Biodiversidade [ICMBio], 2014], a category that probably stems from the recent record of this species in the São João river and the confirmation of its occurrence in protected areas (Maia et al., 2013). However, if the deep phylogenetic lineages found here corroborate a species complex in P. garbei, the conservation status of the entire assemblage may need reexamination, mainly due to a reduction in the range of each species due to the taxonomic splitting.
In summary, our molecular analysis together with paleo-drainage reconstructions revealed that the current phylogeographic patterns of the rheophilic catfish P. garbei were importantly impacted by headwater stream-captures. This taxon proved to encompass at least two highly divergent lineages, and furthermore, each headwater seems to represent a genetically diagnosable OTU. The integrative approach employed here helps to introduce a useful way to test hypotheses regarding to the distribution and conservation of Neotropical freshwater fishes.
Statements
Author contributions
SL had substantial contribution to the acquisition of samples, analyses, and interpretation of the results, writing the manuscript. WB-F and TA also contributed in the analyses, writing, and figures. HL helped in the fish collection, design, and writing the manuscript. AT and JA provided logistic support for molecular data during SL’s postdoctoral activities at UCI. All authors contributed in conception and elaboration of the manuscript, and read and approved its final version.
Funding
SL and HL received postdoctoral scholarships from Science without Borders Program/CNPq (203476/2014-0 and 202253/2015-5, respectively). WB-F receives a Ph.D. scholarship from Science without Borders Program/CNPq (233161/2014-7).
Acknowledgments
We are grateful to Edson H. L. Pereira for receiving and analyzing specimens for morphological differences. We are further indebted to CNPq (Conselho Nacional de Desenvolvimento Cientifico e Tecnologico) for supporting the research and to ICMBio (Instituto Chico Mendes de Conservação da Biodiversidade) and INEA/RJ (Instituto Estadual do Ambiente do Rio de Janeiro) for providing collecting permits to SL (28332-1, 30532-1, and 049/2011). Thiago Barros and Érica Caramaschi were responsible for providing specimens from Guapiaçu sub-basin, and therefore, we are also thankful to them. We are thankful to Andrea Thomaz for helpful advices on paleo-drainages reconstruction.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fgene.2017.00199/full#supplementary-material
FIGURE S1Bayesian phylogenetic reconstructions for cytochrome b (cytB) and cytochrome oxidase I (coi) in Pareiorhaphis garbei, using HKY+I (cytB) and TrN+G (coi) as nucleotide substitution models. Pareiorhaphis cf. bahianus (MG496258 and MG496259) from Contas river basin (northeastern Brazil) was used as out-group. Posterior probabilities are shown above the nodes.
References
1
AlbertJ. S.PetryP.ReisR. E. (2011). “Major biogeographic and phylogenetic patterns,” inHistorical Biogeography of Neotropical Freshwater FishesedsAlbertJ. S.ReisR. (Berkeley, CA: University of California Press) 21–58. 10.1525/california/9780520268685.003.0002
2
AlbertJ. S.ReisR. E. (2011). “Introduction to neotropical freshwaters,” inHistorical Biogeography of Neotropical Freshwater FishesedsAlbertJ. S.ReisR. (Berkeley, CA: University of California Press) 3–19. 10.1525/california/9780520268685.003.0001
3
BenineR. C.MariguelaT. C.OliveiraC. (2009). New species of Moenkhausia Eigenmann, 1903 (Characiformes: Characidae) with comments on the Moenkhausia oligolepis species complex.Neotrop. Ichthyol.7161–168. 10.1590/S1679-62252009000200005
4
BishopP. (1995). Drainage rearrangement by river capture, beheading and diversion.Prog. Phys. Geogr.19449–473. 10.1177/030913339501900402
5
BlairC.BrysonR. W.Jr. (2017). Cryptic diversity and discordance in single-locus species delimitation methods within horned lizards (Phrynosomatidae: Phrynosoma).Mol. Ecol. Resour.171168–1182. 10.1111/1755-0998.12658
6
BuckupP. A. (2011). “The Eastern Brazilian shield,” inHistorical Biogeography of Neotropical Freshwater FishesedsAlbertJ. S.ReisR. (Berkeley, CA: University of California Press) 203–210. 10.1525/california/9780520268685.003.0012
7
CherobimA. M.LazzarottoH.LangeaniF. (2016). A new species of the catfish Neoplecostomus (Loricariidae: Neoplecostominae) from a coastal drainage in southeastern Brazil.Neotrop. Ichthyol.14:e160015. 10.1590/1982-0224-20160015
8
CostaW. J. E. M. (2014). Four new species of the genus Atlantirivulus (Cyprinodontiformes: Rivulidae) from the Brazilian Atlantic Forest.Vertebr. Zool.649–21.
9
DarribaD.TaboadaG. L.DoalloR.PosadaD. (2012). jModelTest 2: more models, new heuristics and parallel computing.Nat. Methods9772–772. 10.1038/nmeth.2109
10
DiasM. S.OberdorffT.HuguenyB.LeprieurF.JézéquelC.CornuJ. F.et al (2014). Global imprint of historical connectivity on freshwater fish biodiversity.Ecol. Lett.171130–1140. 10.1111/ele.12319
11
DrummondA. J.SuchardM. A.XieD.RambautA. (2012). Bayesian phylogenetics with BEAUti and the BEAST 1.7.Mol. Biol. Evol.291969–1973. 10.1093/molbev/mss075
12
EdgarR. C. (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput.Nucleic Acids Res.321792–1797. 10.1093/nar/gkh340
13
EzardT.FujisawaT.BarracloughT. G. (2009). Splits: Species’ Limits by Threshold Statistics. R Package Version 1.Available at: http://r-forge.r-project.org/projects/splits
14
FujisawaT.BarracloughT. G. (2013). Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: a revised method and evaluation on simulated data sets.Syst. Biol.62707–724. 10.1093/sysbio/syt033
15
HajibabaeiM.SingerG. A. C.HebertP. D. N.HickeyD. A. (2007). DNA barcoding: how it complements taxonomy, molecular phylogenetics and population genetics.Trends Genet.23167–172. 10.1016/j.tig.2007.02.001
16
HuelsenbeckJ. P.RonquistF. (2001). MRBAYES: Bayesian inference of phylogenetic trees.Bioinformatics17754–755. 10.1093/bioinformatics/17.8.754
17
Instituto Chico Mendes de Conservação da Biodiversidade [ICMBio] (2014). Lista de Espécies Quase Ameaçadas.Available at: http://www.icmbio.gov.br/portal/images/stories/biodiversidade/fauna-brasileira/quase-ameacadas/Lista_NT_site.xls
18
KapliP.LutteroppS.ZhangJ.KobertK.PavlidisP.StamatakisA.et al (2017). Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo.Bioinformatics331630–1638. 10.1101/063875
19
KartavtsevY. P. (2011). Divergence at Cyt-b and Co-1 mtDNA genes on different taxonomic levels and genetics of speciation in animals.Mitochondrial DNA2255–65. 10.3109/19401736.2011.588215
20
KekkonenM.HebertP. D. N. (2014). DNA barcode-based delineation of putative species: efficient start for taxonomic workflows.Mol. Ecol. Resour.14706–715. 10.1111/1755-0998.12233
21
LazzarottoH.BritoM. F.CaramaschiE. P. (2007). Threatened fishes of the world: Pareiorhaphis garbei (Ihering, 1911) (Ostariophysii: Loricariidae).Environ. Biol. Fishes7891–92. 10.1007/s10641-006-9036-1
22
LeighJ. W.BryantD. (2015). Popart: full-feature software for haplotype network construction.Methods Ecol. Evol.61110–1116. 10.1111/2041-210x.12410
23
LimaF. C. T.RibeiroA. C. (2011). “Continental-scale tectonic controls of biogeography and ecology,” inHistorical Biogeography of Neotropical Freshwater FishesedsAlbertJ. S.ReisR. (Berkeley, CA: University of California Press) 145–164. 10.1525/california/9780520268685.003.0009
24
LimaS. M. Q.VasconcellosA. V.Berbel-FilhoW. M.LazoskiC.RussoC. A.SazimaI.et al (2016). Effects of Pleistocene climatic and geomorphological changes on the population structure of the restricted-range catfish Trichogenes longipinnis (Siluriformes: Trichomycteridae).Syst. Biodivers.14155–170. 10.1080/14772000.2015.1104398
25
MaiaG. F.de PaivaR. E. C.dos SantosA. B.LazzarottoH.LimaS. M. Q. (2013). New distribution record for the threatened armoured suckermouth catfish Pareiorhaphis garbei (Ihering, 1911) (Siluriformes: Loricariidae) in São João river basin, Rio de Janeiro, southeastern Brazil.Check List91113–1116. 10.15560/9.5.1113
26
MartinsF. O.LangeaniF. (2016). Redescription of Hisonotus notatus Eigenmann & Eigenmann, 1889 (Loricariidae: Hypoptopomatinae), the type species of the genus, and description of a new species from coastal drainages of southeastern Brazil.Neotrop. Ichthyol.14:e150100. 10.1590/1982-0224-20150100
27
MeloB. F.BenineR. C.MariguelaT. C.OliveiraC. (2011). A new species of Tetragonopterus Cuvier, 1816 (Characiformes: Characidae: Tetragonopterinae) from the rio Jari, Amapá, northern Brazil.Neotrop. Ichthyol.949–56. 10.1590/s1679-62252011000100002
28
MilliganB. G. (1998). “Total DNA isolation,” inMolecular Genetic Analysis of Populationsed.HoelzelA. R. (Oxford: Oxford University Press) 28–64.
29
Ministério do Meio Ambiente [MMA] (2004). Lista Nacional das Espécies de Invertebrados Aquáticos e Peixes Ameaçados de Extinção com Categorias da IUCN. Instrução Normativa no. 5 de 21 de Maio de 2004. Diário Oficial da União 102: 102-142.Available at: http://www.mma.gov.br/estruturas/179/_arquivos/in_mma_005_04_179.pdf
30
MyersN.MittermeierR. A.MittermeierC. G.Da FonsecaG. A.KentJ. (2000). Biodiversity hotspots for conservation priorities.Nature403853–858. 10.1038/35002501
31
PereiraE. H.PessaliT. C.AndradeF. D.ReisR. E. (2017). Description of a new species of Pareiorhaphis (Loricariidae: Neoplecostominae) from the rio Jequitinhonha basin, Minas Gerais, eastern Brazil.Neotrop. Ichthyol.15:e170007. 10.1590/1982-0224-20170007
32
PereiraE. H.ReisR. E. (2002). Revision of the loricariid genera Hemipsilichthys and Isbrueckerichthys (Teleostei: Siluriformes), with descriptions of five new species of Hemipsilichthys.Ichthyol. Explor. Freshw.1397–146.
33
PereiraE. H. L. (2005). Resurrection of Pareiorhaphis Miranda Ribeiro, 1918 (Teleostei: Siluriformes: Loricariidae), and description of a new species from the rio Iguaçu basin, Brazil.Neotrop. Ichthyol.3271–276. 10.1590/s1679-62252005000200004
34
PereiraE. H. L.BritoM. F. J. (2008). “Hemipsilichthys garbei Ihering, 1911,” inLivro Vermelho da Fauna Brasileira Ameaçada de Extinção [Brazil Red Book of Threatened Species of Fauna]Vol. 2edsMachadoA. B. M.DrummondG. M. M.PagliaA. P. (Brasília: Ministério do Meio Ambiente) 217–218.
35
PereiraL. H.HannerR.ForestiF.OliveiraC. (2013). Can DNA barcoding accurately discriminate megadiverse Neotropical freshwater fish fauna?BMC Genet.14:20. 10.1186/1471-2156-14-20
36
PereiraT. L.SantosU.SchaeferC. E.SouzaG. O.PaivaS. R.MalabarbaL. R.et al (2013). Dispersal and vicariance of Hoplias malabaricus (Bloch, 1794) (Teleostei, Erythrinidae) populations of the Brazilian continental margin.J. Biogeogr.40905–914. 10.1111/jbi.12044
37
PuillandreN.LambertA.BrouilletS.AchazG. (2012). ABGD, automatic barcode gap discovery for primary species delimitation.Mol. Ecol.211864–1877. 10.1111/j.1365-294x.2011.05239.x
38
R Core Team (2017). R: A Language and Environment for Statistical Computing.Vienna: R Foundation for Statistical Computing.
39
RambautA. (2009). Tracer Version v1. 5.0.Available at: http://tree.bio.ed.ac.uk/software/tracer/
40
RibeiroA. C.LimaF. C. T.RiccominiC.MenezesN. A. (2006). Fishes of the Atlantic Rainforest of Boracéia: testimonies of the Quaternary fault reactivation within a Neoproterozoic tectonic province in Southeastern Brazil.Ichthyol. Explor. Freshw.17157–164.
41
RiccominiC.VelázquezV. F.GomesC. B. (2005). “Tectonic controls of the Mesozoic and Cenozoic alkaline magmatism in central-southeastern Brazilian platform,” inMesozoic to Cenozoic Alkaline Magmatism in the Brazilian PlatformedsComin-ChiaramontiP.GomesC. B. (São Paulo: Edusp) 31–56. 10.2138/am.2007.485
42
RoxoF. F.MeloB. F.SilvaG. S.OliveiraC. (2017). New species of Parotocinclus (Siluriformes: Loricariidae) from coastal drainages of Rio de Janeiro, southeastern Brazil.Zootaxa4232260–270. 10.11646/zootaxa.4232.2.9
43
SpaldingM. D.FoxH. E.AllenG. R.DavidsonN.FerdañaZ. A.FinlaysonM. A. X.et al (2007). Marine ecoregions of the world: a bioregionalization of coastal and shelf areas.Bioscience57573–583. 10.1641/b570707
44
TamuraK.StecherG.PetersonD.FilipskiA.KumarS. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0.Mol. Biol. Evol.302725–2729. 10.1093/molbev/mst197
45
TavaleraG.DimcaV.VilaR. (2013). Factors affecting species delimitations with the GMYC model: insights from a butterfly survey.Methods Ecol. Evol.41101–1110. 10.1111/2041-210x.12107
46
ThomazA. T.MalabarbaL. R.BonattoS. L.KnowlesL. L. (2015). Testing the effect of palaeodrainages versus habitat stability on genetic divergence in riverine systems: study of a Neotropical fish of the Brazilian coastal Atlantic Forest.J. Biogeogr.422389–2401. 10.1111/jbi.12597
47
ThomazA. T.MalabarbaL. R.KnowlesL. L. (2017). Genomic signatures of paleodrainages in a freshwater fish along the southeastern coast of Brazil: genetic structure reflects past riverine properties.Heredity119287–294. 10.1038/hdy.2017.46
48
Villa-VerdeL.LazzarottoH.LimaS. M. Q. (2012). A new glanapterygine catfish of the genus Listrura (Siluriformes: Trichomycteridae) from southeastern Brazil, corroborated by morphological and molecular data.Neotrop. Ichthyol.10527–538. 10.1590/s1679-62252012000300005
49
VorisH. K. (2000). Maps of Pleistocene sea levels in Southeast Asia: shorelines, river systems and time durations.J. Biogeogr.271153–1167. 10.1046/j.1365-2699.2000.00489.x
50
WardR. D. (2009). DNA barcode divergence among species and genera of birds and fishes.Mol. Ecol. Resour.91077–1085. 10.1111/j.1755-0998.2009.02541.x
51
WilkinsonM. J.MarshallL. G.LundbergJ. G. (2006). River behavior on megafans and potential influences on diversification and distribution of aquatic organisms.J. South Am. Earth Sci.21151–172. 10.1016/j.jsames.2005.08.002
52
ZhangJ.KapliP.PavlidisP.StamatakisA. (2013). A general species delimitation method with applications to phylogenetic placements.Bioinformatics292869–2876. 10.1093/bioinformatics/btt499
Summary
Keywords
biogeography, Atlantic Forest, phylogeography, conservation genetics, Loricariidae
Citation
Lima SMQ, Berbel-Filho WM, Araújo TFP, Lazzarotto H, Tatarenkov A and Avise JC (2017) Headwater Capture Evidenced by Paleo-Rivers Reconstruction and Population Genetic Structure of the Armored Catfish (Pareiorhaphis garbei) in the Serra do Mar Mountains of Southeastern Brazil. Front. Genet. 8:199. doi: 10.3389/fgene.2017.00199
Received
01 September 2017
Accepted
21 November 2017
Published
05 December 2017
Volume
8 - 2017
Edited by
Roberto Ferreira Artoni, Ponta Grossa State University, Brazil
Reviewed by
Tomas Hrbek, Federal University of Amazonas, Brazil; Peter A. Ritchie, Victoria University of Wellington, New Zealand
Updates
Copyright
© 2017 Lima, Berbel-Filho, Araújo, Lazzarotto, Tatarenkov and Avise.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sergio M. Q. Lima, smaialima@gmail.com
This article was submitted to Evolutionary and Population Genetics, a section of the journal Frontiers in Genetics
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.