- 1Key Laboratory of Freshwater Fisheries and Germplasm Resources Utilization, Ministry of Agriculture, Freshwater Fisheries Research Center, Chinese Academy of Fishery Sciences, Wuxi, China
- 2Wuxi Fisheries College, Nanjing Agricultural University, Wuxi, China
Previous studies have revealed that SDHB has potential functions in the male sexual differentiation and development in M. nipponense through providing ATP. In this study, the functions of Mn-SDHB were further analyzed in depth using quantitative polymerase chain reaction (qPCR), in situ hybridization, western-blot, and RNA interference (RNAi), combined with the histological observations. The full-genome sequence of Mn-SDHB was 54,608 bp at Chromosome 34, including 7 introns and 6 exons. The full-length cDNA sequence of Mn-SDHB was 1,268 base pairs (bp) long with an open reading frame of 807 bp, encoding for 268 amino acids. The highest expression level of Mn-SDHB in different tissues was observed in the testis, and male prawns at post-larval developmental stage 25 during different developmental stages, indicating that SDHB was potentially involved in the male sexual development in M. nipponense. In situ hybridization and western-blot analysis indicated that SDHB plays essential roles in the testis development. The in situ hybridization analysis also implies the potential roles of Mn-SDHB in ovarian development. The expressions of Mn-IAG were decreased after Mn-SDHB dsRNA injection, indicating SDHB has the positive regulatory effects on IAG in M. nipponese. Thus, SDHB was involved in the mechanism of the male sexual development. The testis development was inhibited, and sperms were rarely observed after 10 days of Mn-SDHB dsRNA injection, indicating SDHB has positive effects on the male sexual development in M. nipponense. This study highlights the functions of SDHB in M. nipponense, which provide new insights for the future studies of the male sexual development in other crustacean species.
Introduction
The oriental river prawn, Macrobrachium nipponense, (Crustacea; Decapoda; Palaemonidae) is an important economic freshwater species in China (Fu et al., 2012). The annual aquaculture production was gradually increased to 205.010 tons in 2016 in China with an output value of over 20 billion. Sexual precocity is the main problem in M. nipponense aquaculture, restricting the sustainable development of the M. nipponense industry. Our previous study has been proven that testis was matured at 19 days after metamorphosis (Post-larval developmental stage 19, PL19) (Jin et al., 2016). Testis began to be distinguished at PL13, and sperms can be seen for the first time in the testis at PL19. Thus, inbreeding will be happened between the new born shrimps, leading to the short life span, small size, and low disease resistance. Thus, the long-term goal of M. nipponense aquaculture is to establish a technique to regulate the testis development in M. nipponense. Therefore, a full understanding of the mechanism of male sexual development of M. nipponense is urgently needed.
The androgenic gland is a special tissue in crustacean species, which plays essential roles in male sexual differentiation and development in crustacean species. The androgenic gland and its secreted hormones have been proven to promote the testes development, male sexual differentiation, and the male sexual characteristics development in crustacean species (Sagi et al., 1986, 1990). The IAG gene is an important hormone, secreted by the androgenic gland, playing essential roles in male sexual differentiation and development in crustacean species (Ventura et al., 2009, 2011; Rosen et al., 2010). Knockdown in the expression of IAG in M. rosenbergii resulted in the sex reversal (Ventura et al., 2012). The previous studies also confirmed the multiple and important functions of testis in male sexual differentiation, sexual maturity, and reproductive capability in M. nipponense (Qiu et al., 1995; Yang et al., 1999; Cao, 2006; Guo, 2007). The histological observation of gonad development during the post-larval development stages of M. nipponense indicated that the development of the androgenic gland had important regulatory roles in the testis development in M. nipponense (Jin et al., 2016).
Our previous study predicted that Succinate dehydrogenase complex iron sulfur subunit B (SDHB) may play essential roles in male sexual development in M. nipponense through promoting ATP generation because it was differentially expressed in the testis and the androgenic gland between the reproductive season and non-reproductive season, and was enriched in the metabolic pathways of oxidative phosphorylation and tricarboxylic acid cycle (Jin et al., 2020). Oxidative phosphorylation and tricarboxylic acid cycle were the most enriched metabolic pathways in the transcriptome profiling analysis of the testis and androgenic gland between the reproductive season and non-reproductive season (Jin et al., 2020). SDHB is one of four protein subunits that form succinate dehydrogenase (SDH), which catalyzes the oxidation of succinate, playing essential roles in pro-inflammatory response (Kita et al., 1990; Au et al., 1995). The specific structure of iron-sulfur protein was proven to be responsible for the oxidative stress in oxidative phosphorylation. It is also the only enzyme to be involved in the tricarboxylic acid cycle through association with the inner mitochondrial membrane. It is believed that succinate dehydrogenase complex is bound to two small integral membrane proteins of 13.5 and 15.5 kDa through iron protein subunit, which played vital roles in the electron transport chain. Spermatogenic cells of Lymnaea stagnalis possessed the succinate dehydrogenase, which prevent the metabolic acidosis in the testis resulted from the anaerobic production of lactate and succinate by Sertoli cells. It also played essential roles in developing rat testis (Gaździk et al., 1986). SDHB also had multiple functions, including the resistance to carboxin in Ustilago maydis (Broomfield and Hargreaves, 1992), the prevention of superoxide generation, and premature aging in C. elegans (Huang and Lemire, 2009), resistance to boscalid in Sclerotinia sclerotiorum (Wang et al., 2015).
In this study, we aimed to verify the important role of SDHB in male sexual development mechanism in M. nipponense, by using rapid amplification of cDNA ends (RACE) cloning, qPCR analysis, western-blot and in situ hybridization analysis. This study analyzes the male sexual development from energy metabolism for the first time, providing new insight for the analysis of male sexual development in M. nipponense and the whole crustacean species.
Materials and Methods
Ethics Statement
The Institutional Animal Care and Use Ethics Committee of the Freshwater Fisheries Research Center, Chinese Academy of Fishery Sciences (Wuxi, China) was used to approve all experiments involving M. nipponense in this study.
Sample Preparation
A total of 100 healthy adult M. nipponense were obtained from Tai Lake in Wuxi, China (120°13’44“E, 31°28’22”N), maintained in the lab condition for at least 72 h before tissue collection with the dissolved oxygen of 6 mg/L. Different tissues for qPCR analysis included hepatopancreas, testis, muscle, ovary, gill, eyestalk, heart, and brain. Specimens for the qPCR analysis of different developmental stages were collected from the full-sibs population, collected with their maturation process. The characteristics of various phases of the ovarian reproductive cycle have been well described in the previous report (Qiao et al., 2015). The reproductive season of testis and the androgenic gland were collected in July in summer for western-blot analysis with the water temperature of ≥28°C and illumination time of ≥15 h, while the non-reproductive season of tests and androgenic gland were collected in January in winter with the water temperature of ≤15°C and illumination time of ≤10 h. The samples were washed by phosphate buffer saline (PBS), and immediately frozen in liquid nitrogen until being used for RNA and protein extraction to prevent total RNA and protein degradation.
Rapid Amplification of cDNA Ends (RACE)
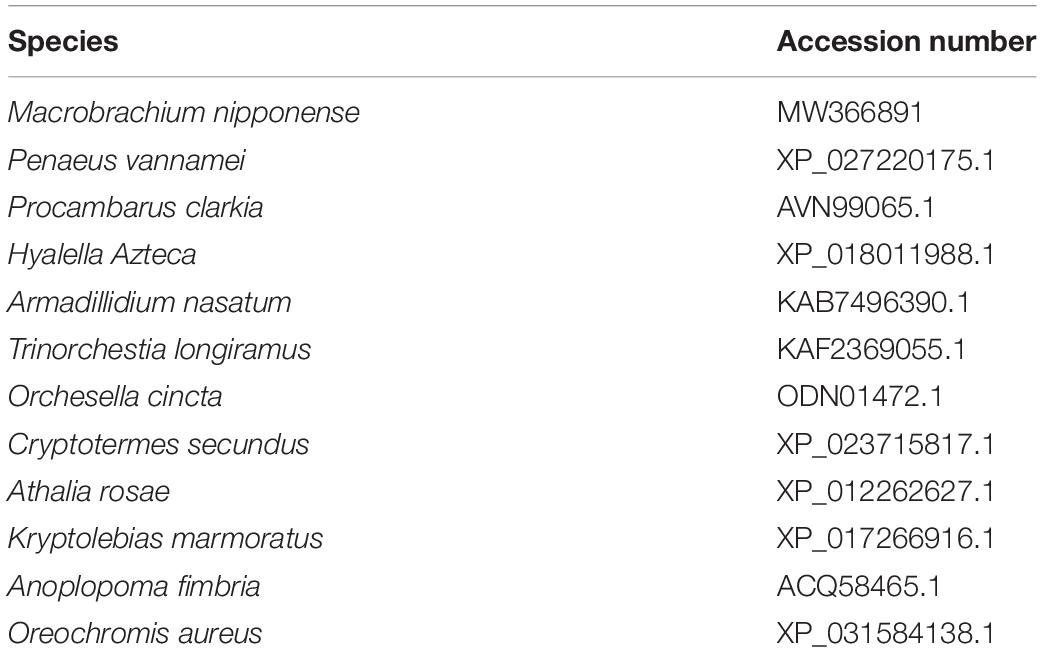
The procedures for RACE cloning have been well-described in previous studies (Jin et al., 2014, 2018). Briefly, RNAiso Plus Reagent (Takara Bio Inc.) was used to extract the total RNA from testis, and then the extracted total RNA was used to synthesize the templates for 3′cDNA and 5′cDNA cloning by using 3′-Full RACE Core Set Ver.2.0 kit and the 5′-Full RACE kit (Takara Bio Inc., Japan). Primer-BLAST tool in NCBI1 was used to design the specific primers used for Mn-SDHB cloning (Table 1). The BLASTX and BLASTN search program2 and the ORF Finder tool3 were employed to analyze the structural characteristics. The phylogenetic tree was constructed by MEGA X, followed by the maximum-likelihood method with Bootstrap method of 1000 replications. The species used for the construction of phylogenetic tree were listed in Table 2.
qPCR Analysis
The mRNA expressions of Mn-SDHB were measured by qPCR analysis. The Bio-Rad iCycler iQ5 Real-Time PCR System (Bio-Rad) was used to carry out the SYBR Green RT-qPCR assay. The previous studies have well described the detailed procedures of qPCR analysis (Jin et al., 2014, 2018). The primers used for qPCR analysis were listed in Table 1. EIF was used as a reference gene in this study (Hu et al., 2018). The amplification efficiency between the Mn-SDHB and EIF were measured, and the amplification efficiency is the same in this study. The relative mRNA expressions of Mn-SDHB were measured by using 2–ΔΔCT method.
In situ Hybridization
The mRNA locations of Mn-SDHB were analyzed by using in situ hybridization. The different tissues included various reproductive cycles of the ovary, the testis, and androgenic gland in reproductive season. The anti-sense and sense probes of CISH (Chromogenic in situ hybridization) study with DIG signal were designed by using Primer5 software based on the cDNA sequence of Mn-SDHB (Table 1). Shanghai Sangon Biotech Company synthesized the probes. The previous studies have well described the detailed procedures of in situ hybridization (Jin et al., 2018). Slides were examined under light microscope for evaluation.
Western-Blot Analysis
Testis and androgenic gland samples (20 mg) were respectively obtained in the reproductive season and non-reproductive season. The Bradford method was used to quantify the total protein concentration (Bradford, 1976). The detailed procedure of western-blot was well described in a previous study (Sun et al., 2017). Briefly, a 10% SDS-polyacrylamide (SDS-PAGE) was used to separate 50 mg protein from each sample, and then transferred to a PVDF membrane (Millipore, Bedford, MA, United States).
RNA Interference (RNAi) Analysis
The potentially regulatory roles of Mn-SDHB on male sexual development in M. nipponense were analyzed by using RNAi. Snap Dragon tools,4 which were used to design the specific RNAi primer with T7 promoter site, shown in Table 2. The Transcript AidTM T7 High Yield Transcription kit (Fermentas, Inc, United States) was used to synthesize the Mn-SDHB dsRNA, following the procedures of the manufacturer. A total of 300 healthy mature male M. nipponense were collected with body weights of 3.12–4.87 g, and divided into two groups. As described in previous study (Jiang et al., 2014; Li et al., 2018), the prawns from experimental group were injected with 4 μg/g Mn-SDHB dsRNA. The concentration of the Mn-SDHB dsRNA was adjusted to 4 μg/μl. Thus, the injected volume of Mn-SDHB dsRNA into each prawn was the same as that of their body weight, while the prawns from the control group were injected with equal volume of vehicles based on the body weight. The SDHB mRNA expression was investigated in the testis by qPCR after the injection of 1, 7, and 14 days, in order to detect the interference efficiency (N ≥ 5).
Histological Observation
The morphological changes of the testis between different days after RNAi treatment were observed by Hematoxylin and eosin (HE) staining. Five testicular samples were respectively collected after 1, 4, 7, 10, and 14 days of RNAi treatment for HE staining. The procedures have been well described in previous studies (ShangGuan et al., 1991; Ma et al., 2006). Olympus SZX16 microscope was used to observe the slides (Olympus Corporation, Tokyo, Japan). The various cell types were labeled following the description of the previous report (Jin et al., 2016).
Statistical Analysis
SPSS Statistics 23.0 was used to measure the statistical differences, estimated by one-way ANOVA followed by LSD and Duncan’s multiple range test. p < 0.05 was used to indicate statistical significance. Quantitative data were expressed as mean ± SD.
Results
Genome and cDNA Sequence Analysis
The full-length cDNA sequence of Mn-SDHB was 1,268 base pairs (bp) long with an open reading frame of 807 bp, encoding for 268 amino acids. The 5′ untranslated region (UTRs) and the 3′ UTR were 74 bp and 387 bp long, respectively. The sequence has been submitted to NCBI with the accession number of MW366891. The molecular weight and theoretical isoelectric point of the protein were 30.684 kDa and 8.68, respectively. The full-genome sequence of Mn-SDHB was 54,608 bp at Chromosome 34 at position 47,658,691-47,713,299, including 7 introns and 6 exons. One PLN00129 superfamily was detected in the cDNA sequence of Mn-SDHB (Figure 1). In addition, Mn-SDHB contained a 2Fe–2S iron–sulfur cluster binding domain (44–152 aa) and a 4Fe–4S dicluster domain (185–261 aa) (Figure 2).
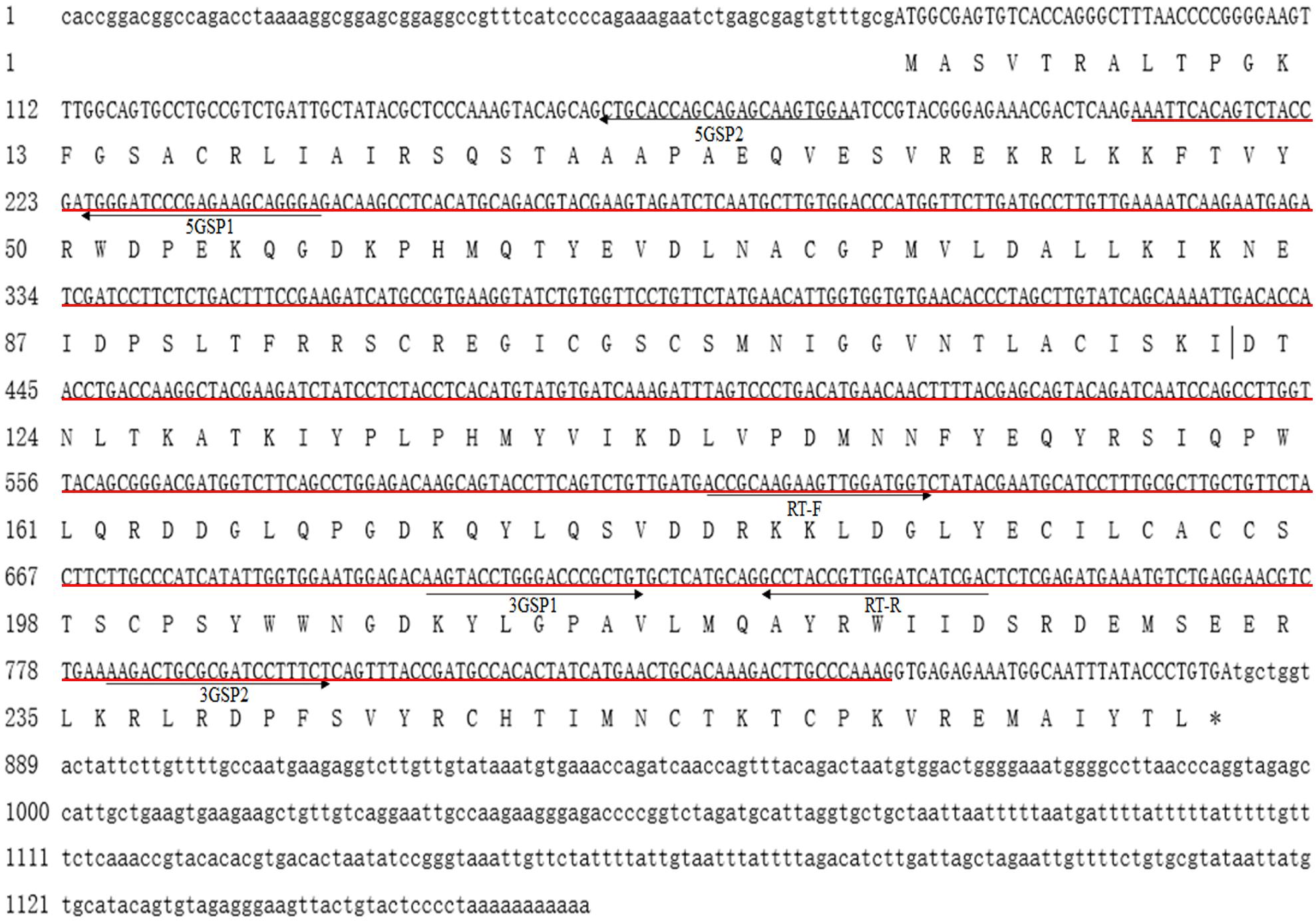
Figure 1. Nucleotide and deduced amino acid sequence of Mn-SDHB. The nucleotide sequence is displayed in the 5′–3′ directions and numbered at the left. The deduced amino acid sequence is shown in a single capital letter amino acid code. 3′ UTR and 5′ UTR are shown with lowercase letters. Codons are numbered at the left with the methionine (ATG) initiation codon, an asterisk denotes the termination codon (TGA). Red line indicated the PLN00129 superfamily.
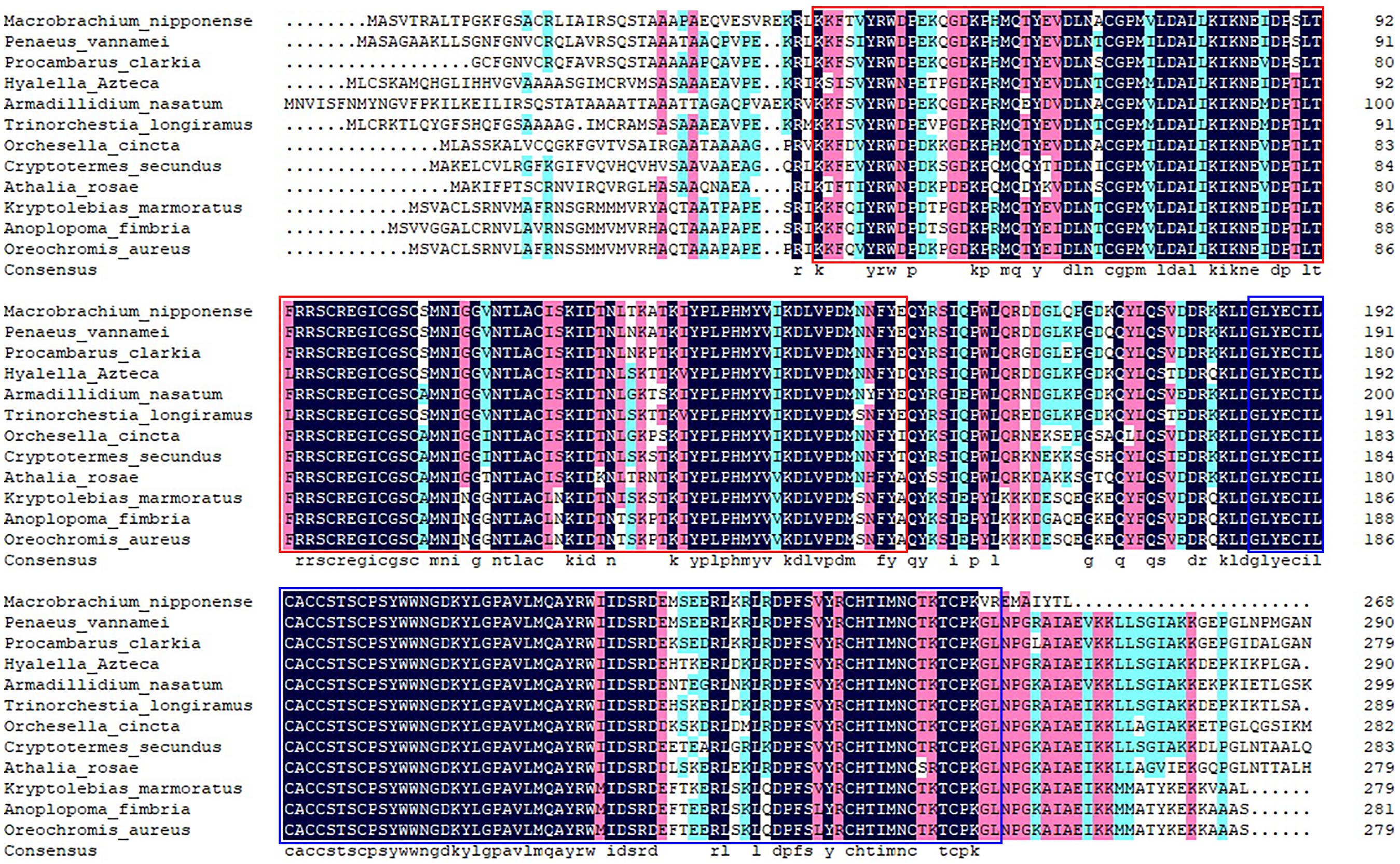

Figure 2. The similarity identity of amino acid sequences of SDHB between different species. Red box indicates the 2Fe–2S iron–sulfur cluster binding domain. Blue box indicates the 4Fe–4S dicluster domain.
According to the BLASTN similarity in NCBI, the similarities of Mn-SDHB and SDHB in other species were over 70%, including Salmo salar (74.74%), Salvelinus alpinus (74.55%), Oncorhynchus kisutch (74.20%), and Megachile rotundata (73.99%) (Figure 2). MEGA 5.1 was further used to analyze the evolutionary relationship between the amino acid sequences of Mn-SDHB and other well-defined SDHB sequences in NCBI, followed the maximum-likelihood method to construct a condensed phylogenetic tree. According to the phylogenetics tree analysis, the amino acids sequence of Mn-SDHB has the closest evolutionary relationship with that of crustacean species, including Penaeus vannamei and Procambarus clarkia. The amino acids sequence of Mn-SDHB was then clustered with the amino acid sequences of insect species as a group, while Mn-SDHB has a long evolutionary relationship with that of fish species (Figure 3).
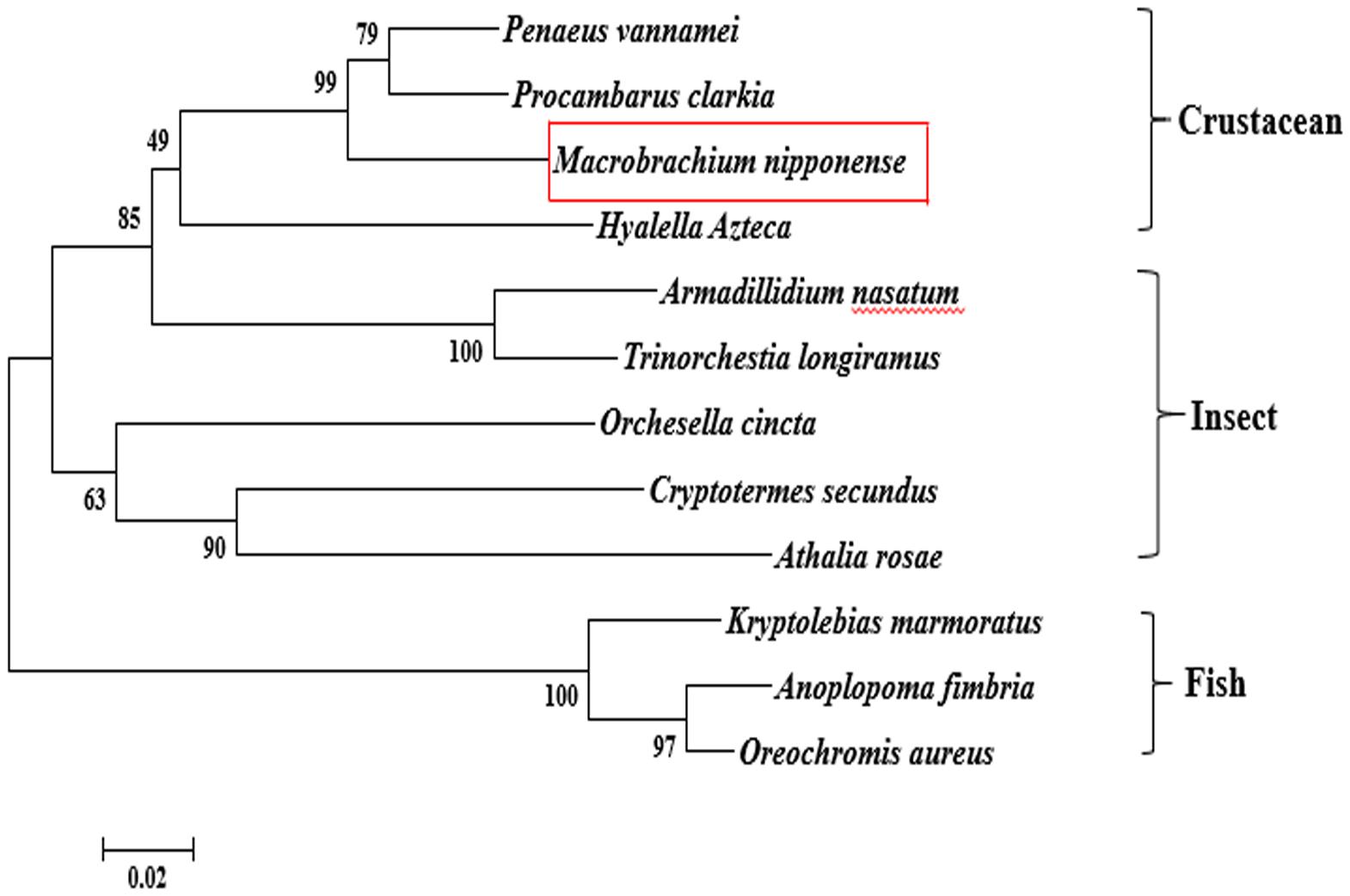
Figure 3. The phylogenetic tree of SDHB from different organisms based on amino acid sequence comparisons. Species names of SDHB are listed on the right of the tree. Red rectangles indicated M. nipponense.
Expression Analysis in Different Tissues and Developmental Stages
The physiological function of a gene may be reflected by the mRNA tissue distribution, evaluated by qPCR analysis. The qPCR analysis of Mn-SDHB in different tissues revealed that the highest RNA expression of Mn-SDHB was observed in the testis, and showed significant difference with other tested tissues (p < 0.05), followed by the heart, gill and ovary. The lowest mRNA expression of Mn-SDHB was observed in muscle. The expression in testis was 20.66-folder and 2.3-folder higher than that of muscle and ovary, respectively (Figure 4A).
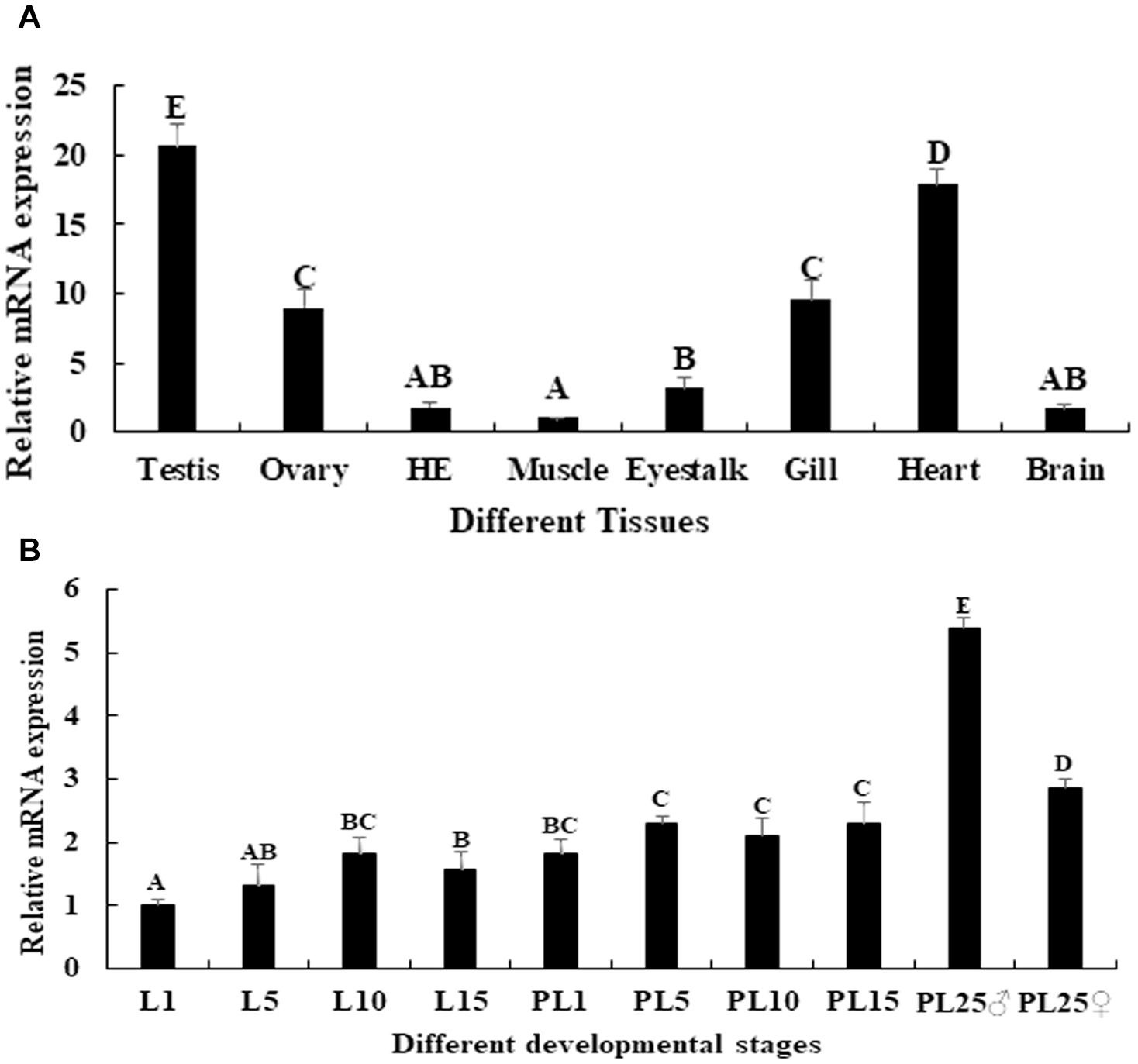
Figure 4. Expression characterization of Mn-SDHB in different mature tissues and developmental stages. The amount of Mn-SDHB mRNA was normalized to the EIF transcript level. Data are shown as mean + SD (standard deviation) of tissues from three separate individuals. Capital letters indicate expression difference between different samples. (A) Expression characterization in different mature tissues. (B) Expression characterization in different developmental stages. HE, hepatopancreas.
qPCR was further used to evaluate the mRNA expression of Mn-SDHB in different larval and post-larval developmental stages. The mRNA expression of Mn-SDHB was gradually increased from larval developmental stage day 1 (L1) to L10, and reached the peak at L10 during the larval developmental stages. The expression at L10 was 1.83-fold higher than that of L1, and showed significant difference (p < 0.05) (Figure 4B). The expression of Mn-SDHB was significantly increased during the post-larval developmental stages, compared with that of larval developmental stages (p < 0.05). The expression remained stable from post-larval developmental stage 1 (PL1) to PL15, and showed no significant difference (p > 0.05). The new born prawns can be distinguished between male and female for the first time in PL25. The expressions were dramatically increased in both PL25♂ (male) and PL25♀ (female), and showed significant difference with other tested developmental stages. However, the expression in PL25♂ was almost 2-fold higher than that of PL25♀ (Figure 4B).
Western-Blot Analysis
Western-blot analysis was performed in the testis and androgenic gland of reproductive season and non-reproductive season. The molecular mass of Mn-SDHB was approximately 30 kDa, according to the western-blot analysis, which was similar to the predicted molecular weight by software. The previous study has indicated that the mRNA expressions of Mn-SDHB in the testis and androgenic gland were higher than that of non-reproductive season (Jin et al., 2020). As shown in Figure 5, clear bands were observed at approximately 30 kDa in the testis and androgenic gland from both of the reproductive season and non-reproductive season, indicating the Mn-SDHB protein was transcribed in these tissues. It is observed that the protein expression levels of Mn-SDHB were up-regulated in the testis and androgenic gland from reproductive season than that in non-reproductive season, which were consistent with that of qPCR analysis. Clear bands were also observed in control group, using β-actin as a reference gene.
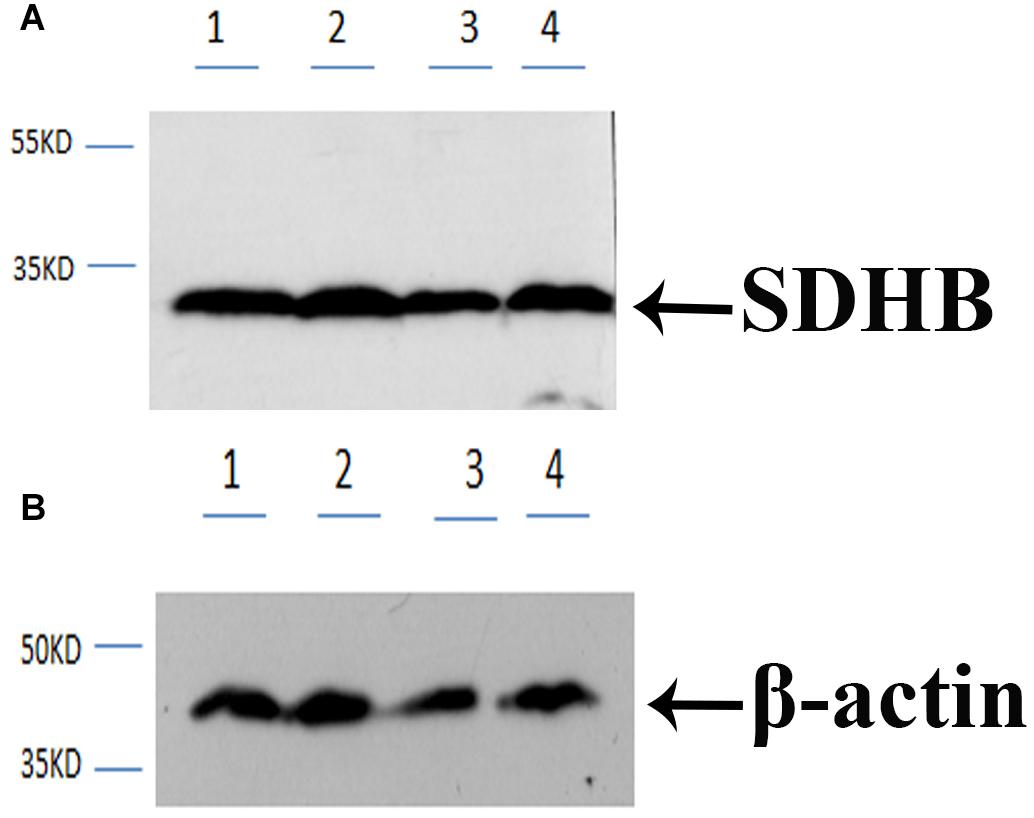

Figure 5. Western-blot analysis in testis and androgenic gland from reproductive season and non-reproductive season. (A) Western-blot for SDHB. (B) Western-blot for B -action. Line 1: Testis from the non-reproductive season; Line 2: Testis from the reproductive season; Line 3: Androgenic gland from the non-reproductive season; Line 4: Androgenic gland from the reproductive season. B -action was used as reference gene.
Expression Analysis in Different Reproductive Cycle of Ovarian Development
Mn-SDHB was expressed during the whole ovarian reproductive cycle. The lowest expression level of Mn-SDHB was observed in O I, while the expression of Mn-SDHB reached the peak at O II. The expression in O II is more than 2-folder higher than that of O I, which showed significant difference with other ovarian developmental stage (p < 0.05). The expression level of Mn-SDHB was then gradually decreased from O III to O V, while the expressions showed no significant difference (p > 0.05) (Figure 6).
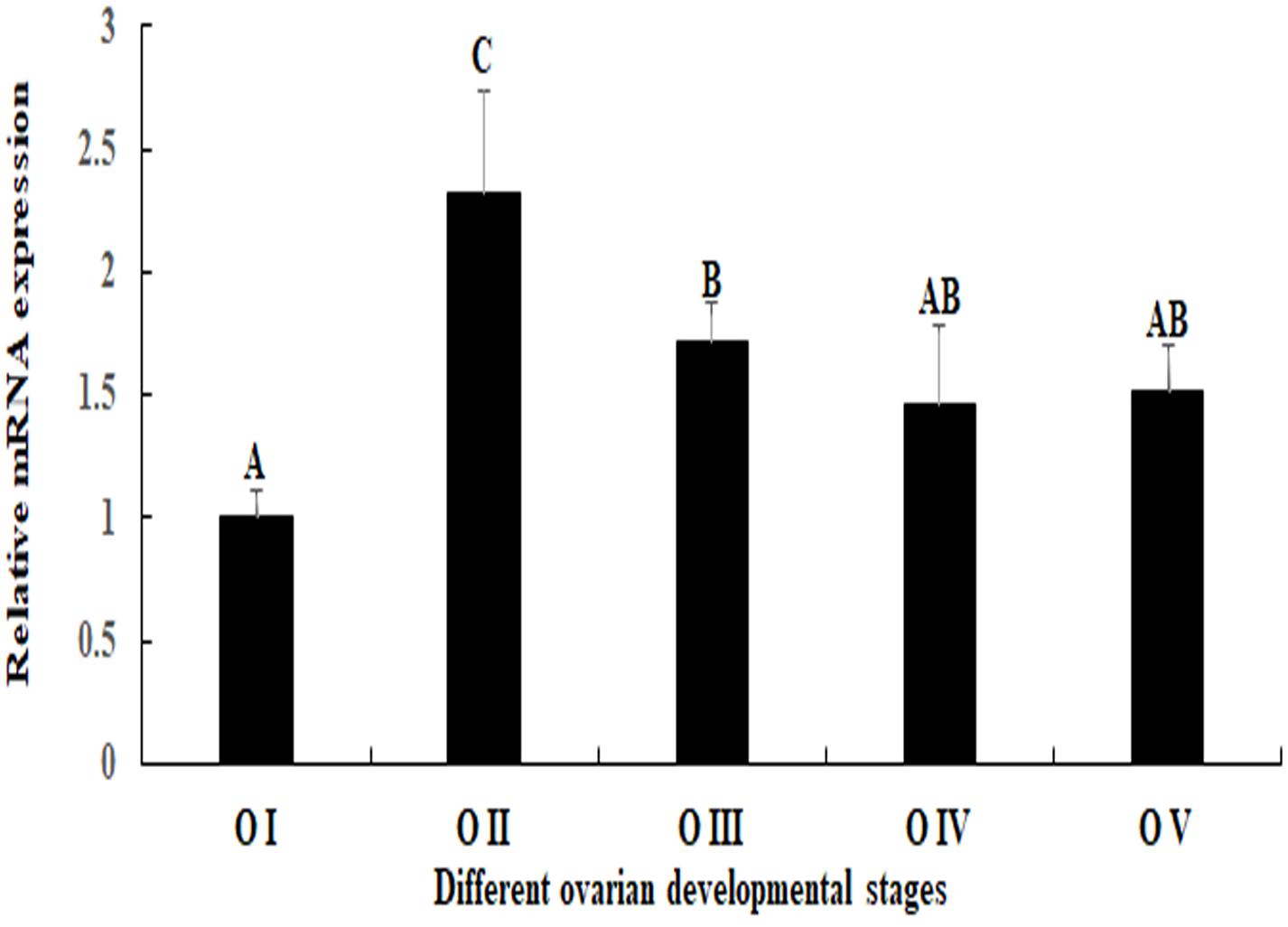
Figure 6. Expression characterization of Mn-SDHB in different reproductive cycles ovary. The amount of Mn-SDHB mRNA was normalized to the EIF transcript level. Data are shown as mean + SD (standard deviation) of tissues from three separate individuals. Capital letters indicate expression difference between different samples.
In situ Hybridization Observation
In situ hybridization analysis was performed in order to detect the locations of Mn-SDHB mRNA in the testis, androgenic gland, and different reproductive cycles of ovarian development. Hematoxylin and eosin (HE) staining showed that the cells in the testis included spermatogonia, spermatocyte, sperms, and collecting tissues. mRNA signals of Mn-SDHB were observed in spermatogonium (Figure 7). Androgenic gland consisted of androgenic gland cells and the ejaculatory bulb. Strong mRNA signals Mn-SDHB in androgenic gland were only observed in the ejaculatory bulb, while no signals were found in androgenic gland cells (Figure 7).
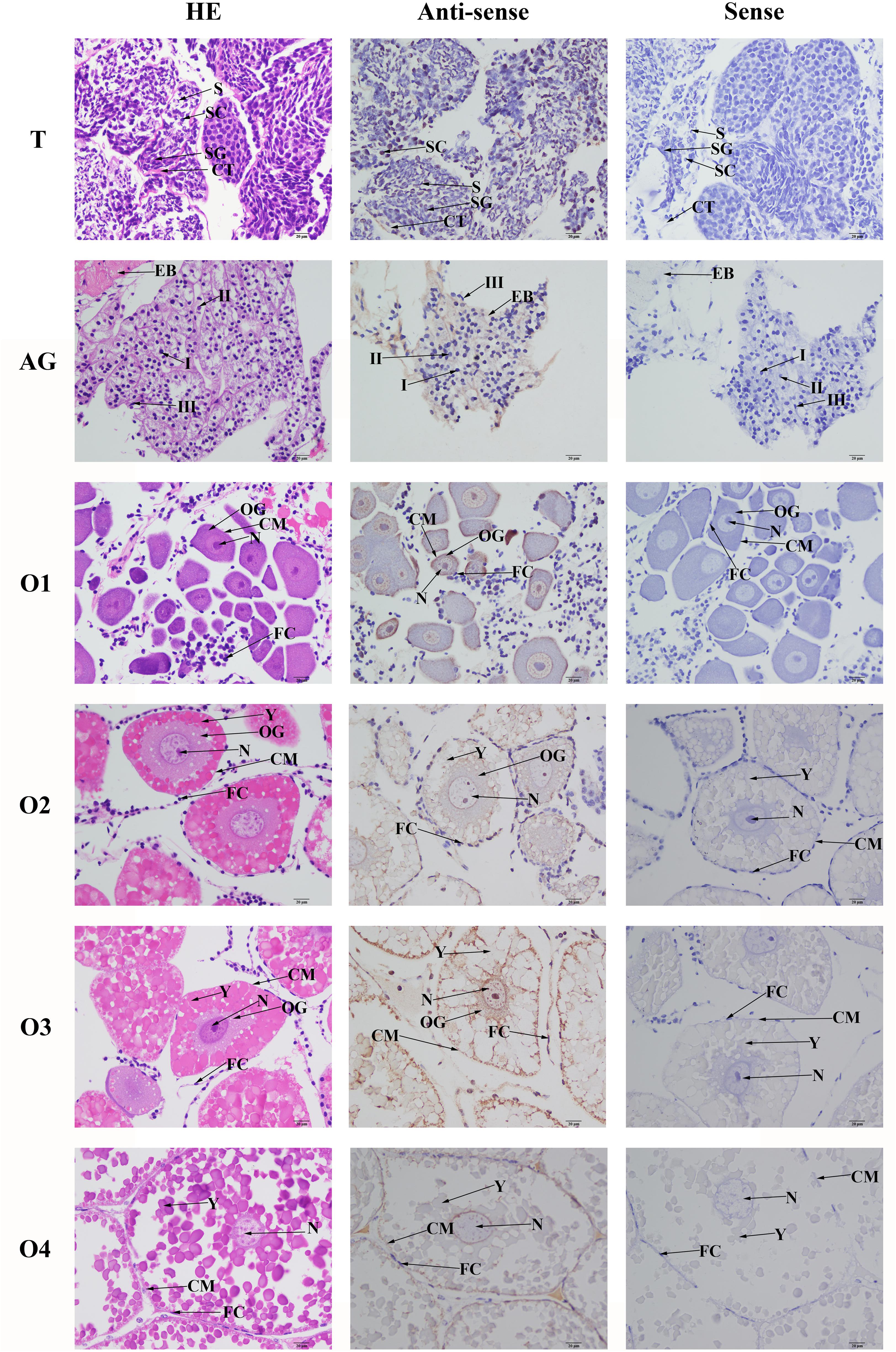
Figure 7. In situ hybridization analysis of Mn-SDHB gene in testis and androgenic gland from reproductive season, and different ovarian reproductive cycle of M. nipponense. ST, seminiferous tubule; SG, Spermatogonia; SC, spermatocyte; S, sperm; CT, collecting tissue; E, wall epithelium; EM, eosinophilic matrix; VD, vas deferens; EB, ejaculatory bulb; OG, oogonium; OC, oocyte; CM, cytoplasmic membrane; N, nucleus; Y, yolk granule; FC, follicle membrane. Scale bars = 20 μm.
Oogonia and follicle cells were observed in ovary stage I, which are derived via differentiation from ovarian epithelial cells, the follicular cavity formed in stage II, which were derived from the follicle cells. Oocyte volume gradually increased in stage III, and yolk granules accumulated in the oocyte (stage IV) during oogenesis and vitellogenesis. According to the Figure 6, the mRNA signals of Mn-SDHB were observed in all of the cell types from O I to O IV, including nucleus, oogoniums, oocytes, cytoplasmic membrane, yolk granule, follicle cells, and follicle membrane (Figure 7).
RNAi Analysis
The mediated functions of Mn-SDHB in male sexual development was evaluated by using RNAi analysis in male prawns. The Mn-SDHB dsRNA was injected twice and the RNAi experiment lasted for 2 weeks. The Mn-SDHB mRNA expression remained at a stable level during the 2 weeks in control group. However, the Mn-SDHB mRNA expressions in the RNAi group were significantly decreased at day 7 and day 14, compared with that in day 1. The mRNA expressions of Mn-SDHB were respectively decreased over 95 and 85% at day 7 and day 14 in the RNAi group, compared to those in the control group (P < 0.01) (Figure 8A). The mRNA expression of Mn-IAG was also measured in the androgenic gland from the same prawn, treated by Mn-SDHB dsRNA. The expressions of Mn-IAG were decreased with the decrease of Mn-SDHB. The expressions were respectively decreased by about 57 and 48% at day 7 and day 14 in the RNAi group, compared with those in the control group at the same day (P < 0.01) (Figure 8B).
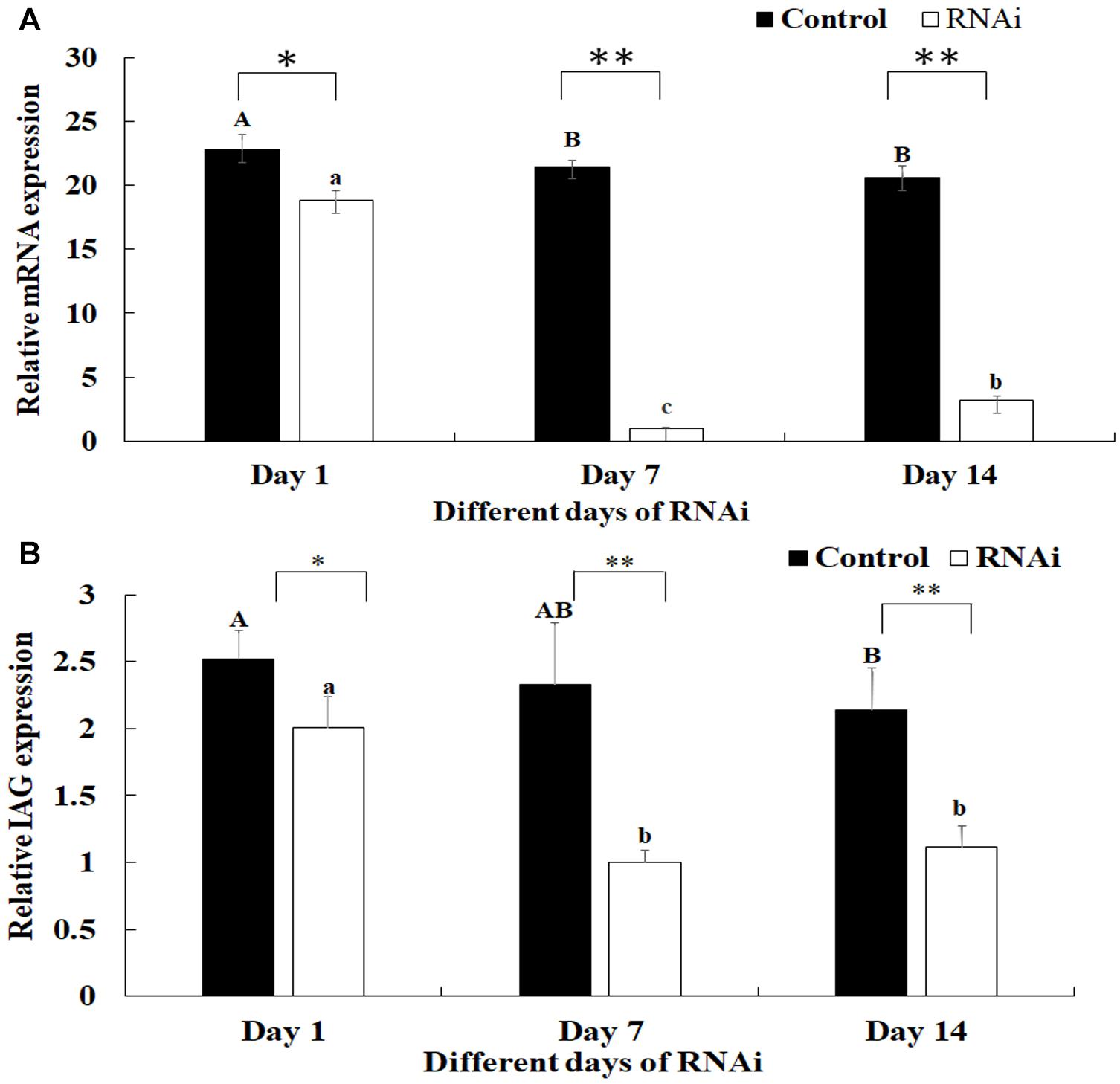
Figure 8. Expression characterization of Mn-SDHB and Mn-IAG at different days after Mn-SDHB dsRNA injection. The amount of Mn-SDHB and Mn-IAG mRNA was normalized to the EIF transcript level. Data are shown as mean + SD (standard deviation) of tissues from three separate individuals. Capital letters indicate expression difference between different days after vehicle injection in control group. Lowercase indicate expression difference between different days after Mn-SDHB dsRNA injection in RNAi group. * (P < 0.05) and ** (P < 0.01) indicates significant expression difference between the RNAi group and control group at the sample day. (A) Expression characterization of Mn-SDHB at different days after Mn-SDHB dsRNA injection. (B) Expression characterization of Mn-IAG at different days after Mn-SDHB dsRNA injection.
The histological observations of testis revealed that the majority of cells in the testis from control group were sperms, while only a few of spermatogonias and spermatocytes were observed. In RNAi group, the number of sperms were deceased with the time of Mn-SDHB dsRNA treatment, while the number of spermatogonias were increased. Sperms were rarely found at day 14 after Mn-SDHB dsRNA treatment (Figure 9).
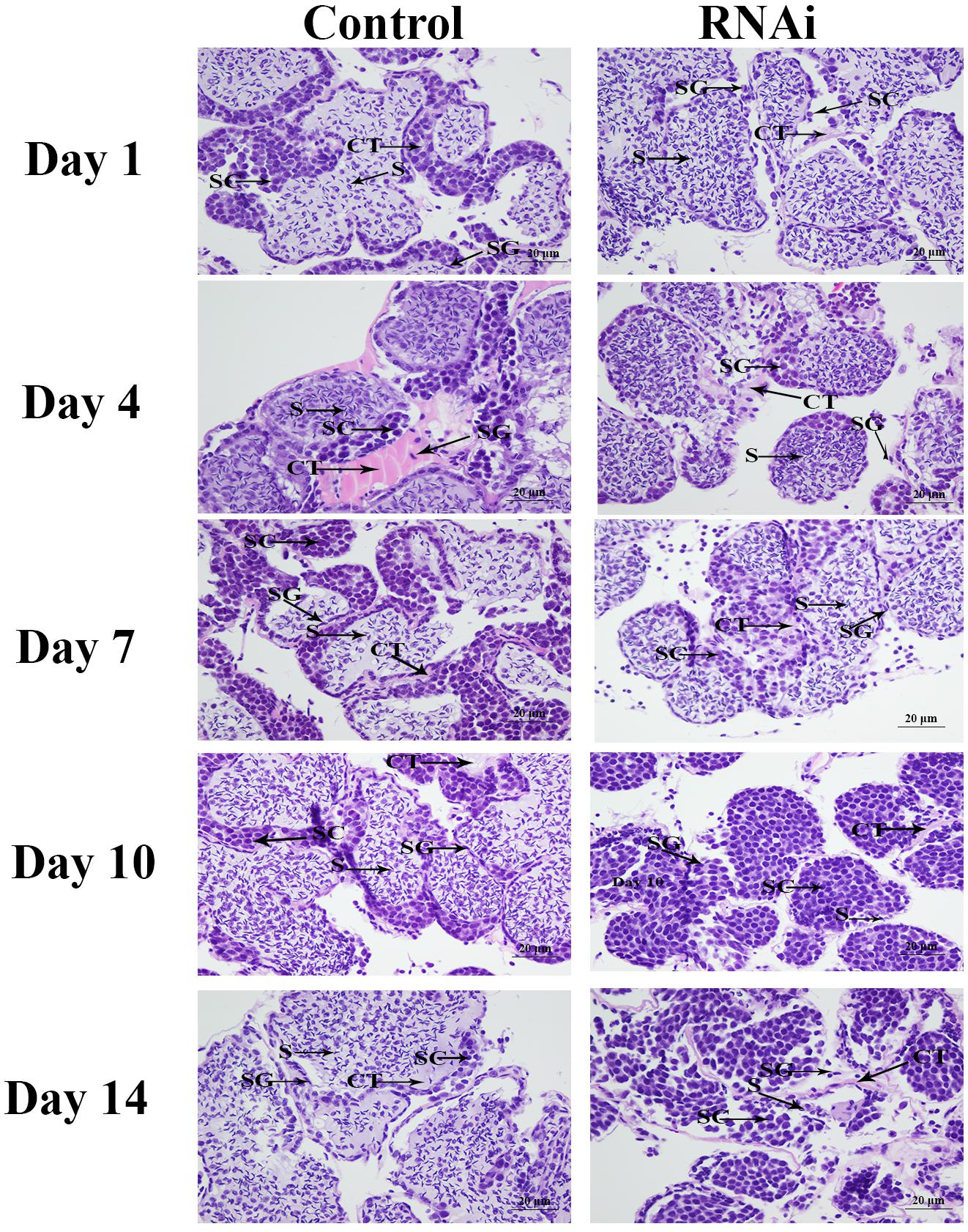
Figure 9. The histological observations of testis between RNAi and control group. SG, Spermatogonia; SC, spermatocyte; S, sperm; CT, collecting tissue. Scale bars = 20 μm.
Discussion
The previous studies have been proven that SDH plays vital roles in the immune system and immune response to infection (Mills et al., 2016; Pinegin et al., 2018). SDHB is a component of SDH, and has been proven to be an important gene in the ATP-releasing process. It also proved to have a function in testis development of Lymnaea stagnalis and rat (Gaździk et al., 1986). SDHB was predicted to be involved in the male sexual differentiation and development in M. nipponense (Jin et al., 2020). In this study, we aimed to focus on the potential functions of SDHB in male sexual differentiation and development through performing the qPCR, in situ hybridization and RNAi analysis in male prawns, combined with the histological observations.
BLASTN similarity comparisons revealed over 70% between Mn-SDHB and SDHB in other crustacean, insect, and fish species, indicating SDHB is a conserved gene. In addition, Mn-SDHB also contained a 2Fe–2S iron–sulfur cluster binding domain and a 4Fe–4S dicluster domain, which was consistent with a previous study (Sun et al., 2020). Mn-SDHB has the closest evolutionary relationship with amino acid sequences of SDHB from crustacean species, based on the maximum-likelihood analysis. These crustacean species include Penaeus vannamei and Procambarus clarkia. Mn-SDHB was then clustered with SDHB from insect species as a group, while the evolutionary relationship of Mn-SDHB with that of fish species was dramatically long. This is consistent with that of taxonomy.
qPCR analysis was used to detect the expression level of Mn-SDHB in various tissues and developmental stages. The previous studies showed that SDH performs multiple biological functions and exists in diverse vertebrate tissues and species (Hochachka et al., 1975; Hochachka and Dressendorfer, 1976; Chouchani et al., 2014). qPCR analysis was performed in Apostichopus japonicus, and the different tissues included coelomocytes, muscle, tentacle, respirating trees, and intestine. The highest expression level was observed in muscle, implying its high mitochondrial content and high oxidative potential in muscles (Sun et al., 2020). In this study, Mn-SDHB was detected in all tested tissues, while the highest expression levels were detected in the testis, and the expression level in the ovary was relatively high. However, the expression in the testis was 2.3-folder higher than that of the ovary. These results implied that the gonad may have high mitochondrial content and high oxidative potential, especially for that in the testis. The qPCR analysis in different developmental stages revealed that the mRNA expression levels of Mn-SDHB in post-larval developmental stages were generally higher than that of larval-developmental stages. The post-larval developmental stages have been proven to be the sensitive period for the gonad differentiation and development in M. nipponense (Jin et al., 2016). The new-born male and female prawn can be distinguished for the first-time at PL25 by morphological observation. The Mn-SDHB expressions were significantly increased at both of PL25♂ and PL25♀, while the expression in PL25♂ was almost 2-folder higher than that of PL25♀. Thus, SDHB was considered to have important roles in gonad development in M. nipponense, especially in the testis through combination of the qPCR results of different mature tissues and different developmental stages. According to the western-blot analysis, clear bands can be observed in the testis and androgenic gland from both the reproductive season and non-reproductive season, indicating that the Mn-SDHB protein can be transcribed in these tissues. The bands of testis and androgenic gland from the reproductive season were much clearer than those of the non-reproductive season, indicating that the protein expressions of Mn-SDHB were up-regulated in the reproductive season, compared with those of the non-reproductive season, which is consistent with those of mRNA expression (Jin et al., 2020).
The expression levels of Mn-SDHB were also detected during the different reproductive cycle of ovarian development. The Mn-SDHB mRNA expressions reached the peak at O II, and then gradually decreased from O III to O V, and the lowest expression level was observed in O I. According to the histological observation, the follicular cavities, derived from the follicle cells, were formed in stage II (Li et al., 2018). A reasonable explanation for the highest expression in O II was that Mn-SDHB plays essential roles in activating the ovarian development, especially for the formation of the follicular cavity.
In situ hybridization was also used to detect the mRNA locations of Mn-SDHB in the testis, androgenic gland, and different reproductive cycles of ovarian development. To the best of our knowledge, the in situ hybridization analysis of SDHB was not reported in any species. According to the in situ hybridization analysis, strong signals were observed in the spermatogonia and collecting tissues of testis, while no signals were observed in sperms, suggesting that Mn-SDHB plays essential roles in promoting the activation of testis development (Jin et al., 2018). No Mn-SDHB mRNA signals were observed in the androgenic gland cell, indicating that the androgenic gland did not directly secrete the SDHB. However, the strong signals in the ejaculatory bulb surrounding the androgenic gland cells indicated that SDHB plays essential roles in maintaining the normal structure and function of the androgenic gland, promoting and supporting the formation of androgenic gland cells (Jin et al., 2018). Strong mRNA signals of Mn-SDHB were observed in all cell types of the ovary, implying that SDHB was involved in the whole reproductive cycle of ovarian development.
RNAi can inhibit the gene expression or translation through the short double-stranded (ds) RNA molecules in the cell’s cytoplasm (Kusaba, 2004; Jones, 2005; Smith et al., 2009). RNAi has been widely used in gene function analysis in M. nipponense (Li et al., 2015, 2018; Jin et al., 2018). Previous studies have proven that SDH driven-mROS plays an important role in the development of antimicrobial innate immune responses (Pinegin et al., 2018), enhancing the host defense during infection (Mills et al., 2016). siRNA was used to knockdown the SDHB expression in A. japonicus, leading to the decrease of the mitochondrial membrane potential and further the decrease of mitochondrial ROS production in A. japonicus coelomocytes (Sun et al., 2020). However, a previous study predicted that SDHB may be involved in the male sexual development in M. nipponense (Jin et al., 2020). RNAi was used to analyze the potential functions of SDHB in male sexual development in M. nipponense. The mRNA expressions of Mn-SDHB were significantly decreased at day 7 and day 14 after the treatment of Mn-SDHB dsRNA, indicating that the Mn-SDHB dsRNA in this study was efficient to knockdown the Mn-SDHB expression. The expression levels of Mn-IAG were also measured in the androgenic gland from the same prawn, treated by Mn-SDHB dsRNA. The expressions of Mn-IAG were decreased with the decrease of Mn-SDHB, indicating that SDHB has a positive regulatory relationship with that of IAG in M. nipponense. The IAG gene has been proven to play essential roles in male sexual differentiation and development in crustacean species (Ventura et al., 2009, 2011, 2012; Rosen et al., 2010). The positive regulatory relationship between SDHB and IAG indicated that SDHB was involved in the male sexual development in M. nipponense. The morphological differences of testis between the RNAi group and control group were measured by histological observations. According to the morphological difference analysis of testis between the RNAi and control group, the sperms were decreased with the time of Mn-SDHB dsRNA treatment, and sperms were hardly observed at day 14 of Mn-SDHB dsRNA treatment, while spermatogonias were dramatically increased, indicating that Mn-SDHB has positive effects on testis development in M. nipponense.
In conclusion, we further analyzed the functions of SDHB in M. nipponense in this study, based on the previous study. Our data strongly suggest that Mn-SDHB played essential roles in gonad differentiation and development, especially for that in male sexual development. Mn-SDHB was proven to have positive regulatory effects on the testis development, based on the Mn-SDHB dsRNA treatment, combined with the histological observations. This study analyzes the male sexual development from energy metabolism for the first time in M. nipponense, promoting the future studies of male sexual development in other crustacean species as well.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/supplementary material.
Ethics Statement
The animal study was reviewed and approved by Macrobrachium nipponense Freshwater Fisheries Research Center. Written informed consent was obtained from the owners for the participation of their animals in this study.
Author Contributions
ShJ designed the experiment. YH performed the RNAi analysis. HF supervised the experiment. SuJ provided the experimental prawns. YX performed the western-blot analysis. HQ performed the qPCR analysis. WZ performed the in situ hybridization analysis. YG performed the histological observations. YW cloned the full cDNA sequence. All authors contributed to the article and approved the submitted version.
Funding
This research was supported by grants from the National Key R&D Program of China (2018YFD0900201); Central Public-interest Scientific Institution Basal Research Fund CAFS (2021JBFM02 and 2020TD36); Jiangsu Agricultural Industry Technology System (JATS[2020]461); the China Agriculture Research System-48 (CARS-48); and the New cultivar breeding Major Project of Jiangsu province (PZCZ201745).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
- ^ http://www.ncbi.nlm.nih.gov/tools/primer-blast/
- ^ http://www.ncbi.nlm.nih.gov/BLAST/
- ^ http://www.ncbi.nlm.nih.gov/gorf/gorf.html
- ^ http://www.flyrnai.org/cgibin/RNAifind_primers.pl
References
Au, H. C., Ream-Robinson, D., Bellew, L. A., Broomfield, P. L., Saghbini, M., and Scheffler, I. E. (1995). Structural organization of the gene encoding the human iron-sulfur subunit of succinate dehydrogenase. Gene 159, 249–253. doi: 10.1016/0378-1119(95)00162-y
Bradford, M. M. (1976). A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254. doi: 10.1016/0003-2697(76)90527-3
Broomfield, P. L., and Hargreaves, J. A. (1992). A single amino-acid change in the iron-sulphur protein subunit of succinate dehydrogenase confers resistance to carboxin in Ustilago maydis. Curr. Genet. 22, 117–121. doi: 10.1007/bf00351470
Cao, J. X. (2006). The Molecular Mechanism In Male Reproductive Tract Of The Prawn, Macrobrachium rosenbergii. Hangzhou: Zhejiang University.
Chouchani, E. T., Pell, V. R., Gaude, E., Aksentijević, D., Sundier, S. Y., Robb, E. L., et al. (2014). Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature 515, 431–435. doi: 10.1038/nature13909
Fu, H. T., Jiang, S. F., and Xiong, Y. W. (2012). Current status and prospects of farming the giant river prawn (Macrobrachium rosenbergii) and the oriental river prawn (Macrobrachium nipponense) in china. Aquac. Res. 43, 993–998. doi: 10.1111/j.1365-2109.2011.03085.x
Gaździk, T., Kamiński, M., Plewka, D., and Plewka, A. (1986). Evaluation of localization and intensity of the reactions of succinic dehydrogenase, NADH tetrazole reductase, and lactate dehydrogenase in developing rat testis. II: after cadmium chloride treatment. Acta Histochem. 78, 179–184. doi: 10.1016/s0065-1281(86)80051-4
Guo, Z. H. (2007). Study On Proliferation And Differentiation Of Spermatogenic Cells From Macrobrachium nipponense in vitro. Baoding: Hebei University.
Hochachka, P. W., and Dressendorfer, R. H. (1976). Succinate accumulation in man during exercise. Eur. J. Appl. Physiol. Occup. Physiol. 35, 235–242. doi: 10.1007/bf00423282
Hochachka, P. W., Owen, T. G., Allen, J. F., and Whittow, G. C. (1975). Multiple end products of anaerobiosis in diving vertebrates. Comp. Biochem. Physiol. B 50, 17–22. doi: 10.1016/0305-0491(75)90292-8
Hu, Y. N., Fu, H. T., Qiao, H., Sun, S. M., Zhang, W. Y., Jin, S. B., et al. (2018). Validation and evaluation of reference genes for Quantitative real-time PCR in Macrobrachium nipponense. Int. J. Mol. Sci. 19:2258. doi: 10.3390/ijms19082258
Huang, J., and Lemire, B. D. (2009). Mutations in the C. elegans succinate dehydrogenase iron-sulfur subunit promote superoxide generation and premature aging. J. Mol. Biol. 387, 559–569. doi: 10.1016/j.jmb.2009.02.028
Jiang, F. W., Fu, H. T., Qiao, H., Zhang, W. Y., Jiang, S. F., Xiong, Y. W., et al. (2014). The RNA Interference Regularity of Transformer-2 Gene of Oriental River Prawn Macrobrachium nipponense. Chin. Agric. Sci. Bull. 30, 32–37.
Jin, S. B., Fu, H. T., Jiang, S. F., Xiong, Y. W., Sun, S. M., Qiao, H., et al. (2018). Molecular Cloning, Expression, and In Situ Hybridization Analysis of Forkhead Box Protein L2 during Development in Macrobrachium nipponense. J. World Aquac. Soc. 49, 429–440. doi: 10.1111/jwas.12510
Jin, S. B., Hu, Y. N., Fu, H. T., Sun, S. M., Jiang, S. F., Xiong, Y. W., et al. (2020). Analysis of testis metabolome and transcriptome from the oriental river prawn (Macrobrachium nipponense) in response to different temperatures and illumination times. Comp. Biochem. Physiol. D Genomics Proteomics 34:100662. doi: 10.1016/j.cbd.2020.100662
Jin, S. B., Jiang, S. F., Xiong, Y. W., Qiao, H., Sun, S. M., Zhang, W. Y., et al. (2014). Molecular cloning of two tropomyosin family genes and expression analysis during development in oriental river prawn, Macrobrachium nipponense. Gene 546, 390–397. doi: 10.1016/j.gene.2014.05.014
Jin, S. B., Zhang, Y., Guan, H. H., Fu, H. T., Jiang, S. F., Xiong, Y. W., et al. (2016). Histological observation of gonadal development during post-larva in oriental river prawn, Macrobrachium nipponense. Chin. J. Fish. 29, 11–16.
Jones, H. D. (2005). Wheat transformation: current technology and applications to grain development and composition. J. Cereal Sci. 41, 137–147. doi: 10.1016/j.jcs.2004.08.009
Kita, K., Oya, H., Gennis, R. B., Ackrell, B. A., and Kasahara, M. (1990). Human complex II (succinate-ubiquinone oxidoreductase): cDNA cloning of iron sulfur (Ip) subunit of liver mitochondria. Biochem. Biophys. Res. Commun. 166, 101–108. doi: 10.1016/0006-291x(90)91916-g
Kusaba, M. (2004). RNA interference in crop plants. Curr. Opin. Biol. 15, 139–143. doi: 10.1016/j.copbio.2004.02.004
Li, F., Qiao, H., Fu, H. T., Sun, S. M., Zhang, W. Y., Jin, S. B., et al. (2018). Identification and characterization of opsin gene and its role in ovarian maturation in the oriental river prawn Macrobrachium nipponense. Comp. Biochem. Physiol. B. 218, 1–12. doi: 10.1016/j.cbpb.2017.12.016
Li, F. J., Bai, H. K., Xiong, Y. W., Fu, H. T., Jiang, S. F., Jin, S. B., et al. (2015). Molecular characterization of insulin-like androgenic gland hormone-binding protein gene from the oriental river prawn Macrobrachium nipponense and investigation of its transcriptional relationship with the insulin-like androgenic gland hormone gene. Gen. Comp. Endocrinol. 216, 152–160. doi: 10.1016/j.ygcen.2014.12.007
Ma, X. K., Liu, X. Z., Wen, H. S., Xu, Y. J., and Zhang, L. J. (2006). Histological observation on gonadal sex differentiation in Cynoglossus semilaevis Günther. Mar. Fish. Res. 27, 55–61.
Mills, E. L., Kelly, B., Logan, A., Costa, A. S. H., Varma, M., Bryant, C. E., et al. (2016). Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell 167, 457–470. doi: 10.1016/j.cell.2016.08.064
Pinegin, B., Vorobjeva, N., Pashenkov, M., and Chernyak, B. (2018). The role of mitochondrial ROS in antibacterial immunity. J. Cell. Physiol. 233, 3745–3754. doi: 10.1002/jcp.26117
Qiao, H., Xiong, Y. W., Zhang, W. Y., Fu, H. T., Jiang, S. F., Sun, S. M., et al. (2015). Characterization, expression, and function analysis of gonad-inhibiting hormone in Oriental River prawn, Macrobrachium nipponense and its induced expression by temperature. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 185, 1–8. doi: 10.1016/j.cbpa.2015.03.005
Qiu, G. F., Du, N. S., and Lai, W. (1995). Studies on the male reproductive system of the freshwater prawn, Macrobrachium nipponense. J. Shanghai Fish. Univ. 4, 107–111.
Rosen, O., Manor, R., Weil, S., Gafni, O., Linial, A., Aflalo, E. D., et al. (2010). A sexual shift induced by silencing of a single insulin-like gene in crayfish: ovarian upregulation and testicular degeneration. PLoS One 5:e15281. doi: 10.1371/journal.pone.0015281
Sagi, A., Cohen, D., and Milner, Y. (1990). Effect of androgenic gland ablation on morphotypic differentiation and sexual characteristics of male freshwater prawns, Macrobrachium rosenbergii. Gen. Comp. Endocrinol. 77, 15–22. doi: 10.1016/0016-6480(90)90201-v
Sagi, A., Cohen, D., and Wax, Y. (1986). Production of Macrobrachium rosenbetgii in momosex population: yield characteristes under intensive monoculture conditions in cages. Aquaculture 51, 265–275. doi: 10.1016/0044-8486(86)90318-2
ShangGuan, B. M., Liu, Z. Z., and Li, S. Q. (1991). Histological Studies on Ovarian Development in Scylla serrata. J. Fish. China 15, 96–103.
Smith, C. A., Roeszler, K. N., Ohnesorg, T., Cummins, D. M., Farlie, P. G., Doran, T. M., et al. (2009). The avian Z-linked gene DMRT1 is required for male sex determination in the chicken. Nature 461, 267–271. doi: 10.1038/nature08298
Sun, L. L., Zhou, F. Y., Shao, Y. N., Lv, Z. M., and Li, C. H. (2020). The iron–sulfur protein subunit of succinate dehydrogenase is critical in driving mitochondrial reactive oxygen species generation in Apostichopus japonicus. Fish Shellfish Immunol. 102, 350–360. doi: 10.1016/j.fsi.2020.04.060
Sun, S. M., Xuan, F. J., Fu, H. Y., Zhu, J., Ge, X. P., and Wu, X. G. (2017). Molecular cloning, mRNA expression and characterization of membrane bound hemoglobin in oriental river prawn Macrobrachium nipponense. Comp. Biochem. Physiol. A 207, 36–42. doi: 10.1016/j.cbpa.2017.02.010
Ventura, T., Manor, R., Aflalo, E. D., Weil, S., Khalaila, I., Rosen, O., et al. (2011). Expression of an androgenic gland-specific insulin-like peptide during the course of prawn sexual and morphotypic differentiation. ISRN Endocrinol. 2011:476283.
Ventura, T., Manor, R., Aflalo, E. D., Weil, S., Raviv, S., Glazer, L., et al. (2009). Temporal silencing of an androgenic gland-specific insulin-like gene affecting phenotypical gender differences and spermatogenesis. Endocrinology 150, 1278–1286. doi: 10.1210/en.2008-0906
Ventura, T., Manor, R., Aflalo, E. D., Weil, S., Rosen, O., and Sagi, A. (2012). Timing sexual differentiation: full functional sex reversal achieved through silencing of a single insulin-like gene in the prawn, Macrobrachium rosenbergii. Biol. Reprod. 86:90.
Wang, Y., Duan, Y., Wang, J., and Zhou, M. (2015). A new point mutation in the iron-sulfur subunit of succinate dehydrogenase confers resistance to boscalid in Sclerotinia sclerotiorum. Mol. Plant Pathol. 16, 653–661. doi: 10.1111/mpp.12222
Keywords: Macrobrachium nipponense, SDHB, qPCR analysis, RNAi, male sexual development
Citation: Jin S, Hu Y, Fu H, Jiang S, Xiong Y, Qiao H, Zhang W, Gong Y and Wu Y (2021) Identification and Characterization of the Succinate Dehydrogenase Complex Iron Sulfur Subunit B Gene in the Oriental River Prawn, Macrobrachium nipponense. Front. Genet. 12:698318. doi: 10.3389/fgene.2021.698318
Received: 21 April 2021; Accepted: 24 May 2021;
Published: 15 July 2021.
Edited by:
Beide Fu, Institute of Hydrobiology, Chinese Academy of Sciences, ChinaCopyright © 2021 Jin, Hu, Fu, Jiang, Xiong, Qiao, Zhang, Gong and Wu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hongtuo Fu, ZnVodEBmZnJjLmNu
 Shubo Jin
Shubo Jin Yuning Hu2
Yuning Hu2 Sufei Jiang
Sufei Jiang
