- 1Key Laboratory of Sustainable Development of Marine Fisheries, Ministry of Agriculture, P.R.China, Yellow Sea Fisheries Research Institute, Chinese Academy of Fishery Sciences, Qingdao, China
- 2Function Laboratory for Marine Fisheries Science and Food Production Processes, Qingdao National Laboratory for Marine Science and Technology, Qingdao, China
- 3College of marine technology and environment, Dalian Ocean University, Dalian, China
Salt-alkali tolerance is one of the important breeding traits of Portunus trituberculatus. Identification of molecular markers linked to salt-alkali tolerance is prerequisite to develop such molecular marker-assisted breeding. In this study, Bulked Segregant Analysis (BSA) was used to screen molecular markers associated with salt-alkali tolerance trait in P. trituberculatus. Two DNA mixing pools with significant difference in salt-alkali tolerance were prepared and 94.83G of high-quality sequencing data was obtained. 855 SNPs and 1051 Indels were firstly selected as candidate markers by BSA analysis, out of which, 20 markers were further selected via △index value (close to 0 or 1) and eight of those were successfully verified. In addition, based on the located information of the markers in genome, eight candidate genes related to salt-alkali tolerance were anchored including ubiquitin-conjugating enzyme, aspartate–tRNA ligase, vesicle-trafficking protein, and so on. qPCR results showed that the expression patterns of all these genes changed significantly after salt-alkali stress, suggesting that they play certain roles in salt-alkali adaptation. Our results will provide applicable markers for molecular marker-assisted breeding and help to clarify the mechanisms of salt-alkali adaptation of P. trituberculatus.
Introduction
Portunus trituberculatus (Juan and Gaoli, 2021; Ye and Fangmin, 2021), which belongs to Crustacea, Decapoda, Portunus family, commonly known as Portunus, is an important breeding species along the coast of China (Wu, 2014). Saline-alkali waters are a form of worldwide low-yield water resources, and there are about 690 million mu of low-lying saline-alkali waters in inland China alone. However, due to its high salinity, high alkalinity, high pH, and complex ion composition (Li, 2020), common aquatic animals cannot survive and reproduce in this environment suitably. The use of this type of water resource is thus, greatly hindered. High salt-alkali stimulation will reduce the feeding, metamorphosis, and survival rates of juvenile crabs (Hu et al., 2014). Therefore, the salt-alkali tolerance trait is one of the important breeding traits of P. trituberculatus.
At present, research on salt-alkali tolerance in aquatic animals mainly focus on fishes, especially on species with natural resistance to salt-alkali, such as Oncorhynchus clarkihenshawi (Wilkie and Wright, 1994), and Chalcalburnus tarichi (Arabacı and Sarı, 2004). Studies of molecular physiological mechanisms have been carried out on Oreochromis niloticus (Biao and Li, 2012; Zhao and Wu, 2016) and Oryzias latipes (Yao and Wang, 2012). Some important ion transport genes including Na+ -K+-ATPase, slc4a1, slc4a4, and slc26a6, were cloned and proved to play an important role in salt-alkali tolerance (Whittamore, 2012; Wood and Bergman, 2012). Compared with fish, less studies documented salt-alkali stress in aquatic crustaceans (Fu, 2002; Fu-yi et al., 2005; Li et al., 2021). Out of those, most studies are similar to the work in fish, focusing on expression of ion transport-related genes (Na+/K+-ATPase, HSP, proPO) (Shen et al., 2014; Xu et al., 2020) and the analysis of enzyme activity (ACP, AKP, SOD) (Liu, 2016; Xu et al., 2020) after salt-alkali stress. However, just a few cases were reported on whether crustaceans have special salt-alkali tolerance genes.
Molecular markers are prerequisite for marker-assisted breeding. Due to the important economic value of P. trituberculatus, some growth-, sex-, and immunity-related markers have been discovered and verified based on quantitative trait locus (QTL) and association analysis (Lv et al., 2017; Lv et al., 2018; Zhou and Zhao, 2019). Recently, in terms of environmental adaptation, some salinity-related molecular markers have also been reported (Lv et al., 2019). However, there were no reports on the development of salt-alkali tolerance molecular markers in crabs.
In this study, we explored the salt-alkali tolerance molecular markers in P. trituberculatus by the Bulked Segregant Analysis (BSA) strategy. In addition, based on the location information of the salt-alkali tolerance molecular markers on genome, we tried to anchor genes related to salt-alkali adaptation in crab. Our results will provide applicable markers for molecular marker-assisted breeding and help to clarify the mechanism of salt-alkali adaptation of P. trituberculatus.
Materials and Methods
Experimental Material
The crab used in this study was cultured in Changyi Haifeng Aquaculture Co., Ltd., Weifang City, Shandong Province, China. Six hundred healthy crabs were selected and randomly divided into three groups (experimental, control, and sample), and the average weight was 55.28 g. During the temporary rearing (holding for 7 days), the water temperature was maintained at 22 ± 1 °C and pH 8.2 ± 0.5. Seawater of the experimental group and sample group was adjusted to 16.7 mmol/L by adding carbonate, and for the control group, normal seawater was used. Then, salt-alkali stress was observed every 2 h and dead individuals were collected in time. The experiment lasted for 72 h, until the survival rate of the experimental group was 5%, while no death was recorded in the control group. The first 20 individuals that died were from the saline-alkali-sensitive group (S) and the last 20 individuals that survived were from the saline-alkali-tolerant group (T). Muscle tissues from the crabs were collected and stored in liquid nitrogen. In the sampling group, gill tissues were sampled at 0, 6, 12, 24, 48, and 72 h after stress, and stored in liquid nitrogen for cDNA for qPCR detection analysis.
Library Construction and Sequencing
DNA degradation and contamination was monitored on 1% agarose gels. DNA purity was checked using NanoPhotometer® spectrophotometer (IMPLEN, CA, United States). DNA concentration was measured using Qubit® DNA Assay Kit in Qubit® 2.0 Fluorometer (LifeTechnologies, CA, United States). DNA sample was fragmented by sonication to a size of 350 bp, which were end polished, A-tailed, and ligated with the full-length adapter for Illumina sequencing with further PCR amplification. PCR products were purified (AMPure XP system) and libraries were analyzed for size distribution. These libraries were sequenced by using Illumina HiSeq platform and 150 bp paired-end reads were generated.
BSA Analysis
Burrows-Wheeler Aligner (Heng and Richard, 2009) was used to align the clean reads of each sample against the reference genome. Alignment files were converted to BAM files using SAMtools software (Li et al., 2009). Variant calling was performed for all samples using the Unified Genotyper function in GATK3.3 (McKenna and Hanna, 2010) software. SNP was selected by using the Variant Filtration parameter in GATK and InDel was filtered by using the Variant Filtration parameter.
The reads depth information for the SNPs/InDels above in the offspring pools was gained to calculate the SNP/InDel index (Takagi and Abe, 2013). We filtered out those points whose SNP/InDel index in both pools were less than 0.3 and depth less than 7. The difference of the SNP/InDel index of two pools was calculated as the △ SNP/InDel index. The markers with an absolute value of △index between 0.69 and 1 were screened out as candidate molecular markers.
Validation of Molecular Markers Related to Salt-Alkali Tolerance
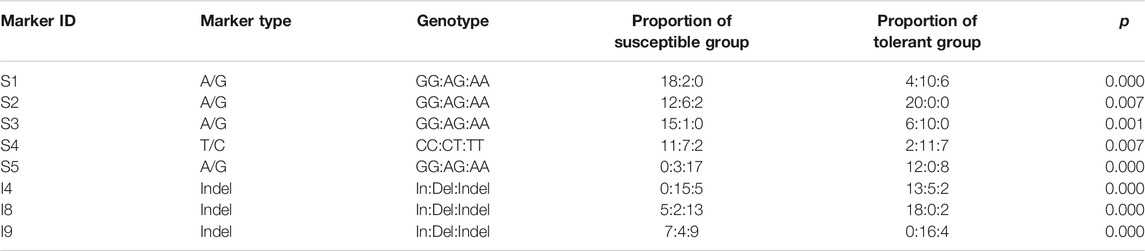
SNPs and Indels conducted two rounds of verification (Table 1). The first round used a mixed template made by mixing the extracted DNA individual templates for PCR amplification and preliminary screening. The second round used individual DNA templates for amplification and checked by sequencing, or 2% agarose electrophoresis. Statistics of the genotype of each individual were based on the results of sequencing and electrophoresis. SPSS 17.0 software was used for one-way analysis of variance (ANOVA), and p < 0.05 was considered to be significantly related to salt-alkali tolerance.
qPCR
The volume of the qPCR reaction system was 10 μL, containing 5 μL of 2 × SYBR Master Mix (RR420A, Takara Bio, Japan), 0.2 μL of each primer and ROX Reference Dye II, 1 μL of cDNA template, and 3.4 μL of rnase Free dH2O. qPCR was carried out in a FAST-7500 system (ABI-7500, ThermoFisher, Singapore) as follows: 95°C or 30 s, and 40 cycles of 95°C for 5 s and 60°C for 34 s. Results from qPCR were analyzed by SPSS software’s ANOVA for group differences, and Origin 9.1 was used for mapping.
Results
Screening of Candidate Molecular Markers
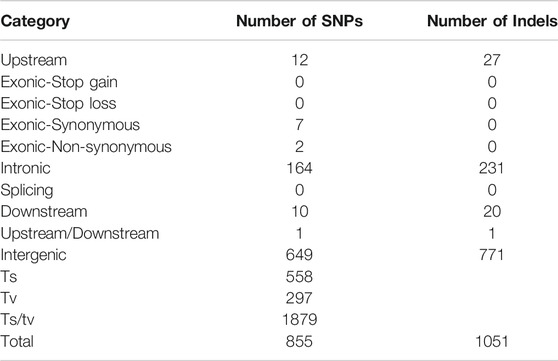
In this study, raw data of 45,854,625,300 and 49,159,475,100 were obtained from the populations with significant differences to salt-alkali tolerance (Table 2). The sequencing depth is 44.65X and 48.31X. A total of 855 SNPs (Supplementary Table S1) and 1051 Indels candidate polymorphic sex sites were selected with 95% confidence level (Supplementary Table S2). Among them, 395 were in the intron region and 1,420 in the intergenic region (Table 3).
Validation of Molecular Markers
A total of ten SNPs and ten Indels were screened out based on △index value (close to 0 or 1). Using the mixed template for PCR amplification, a total of 20 sites were successfully amplified by the target PCR product. Then, 40 individuals were typed for the successfully verified markers in the mixed template by PCR product sequencing and gel agarose electrophoresis. The association analysis test was performed by SPSS, and eight of the markers showed significant correlation with salt-alkali tolerance (p < 0.05) (Table 4), including five SNPs and three Indels (Table 5).
Enrichment Analysis of Candidate Genes
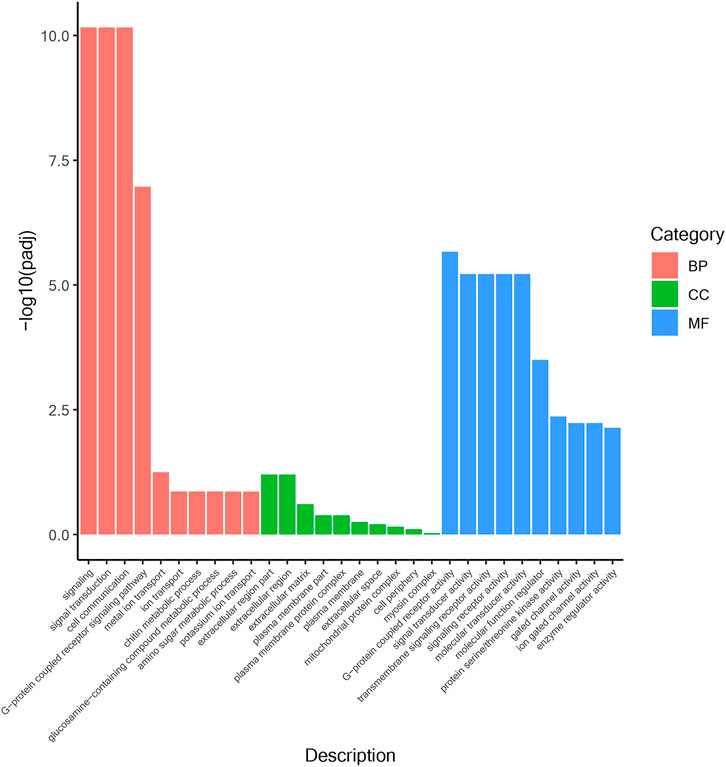
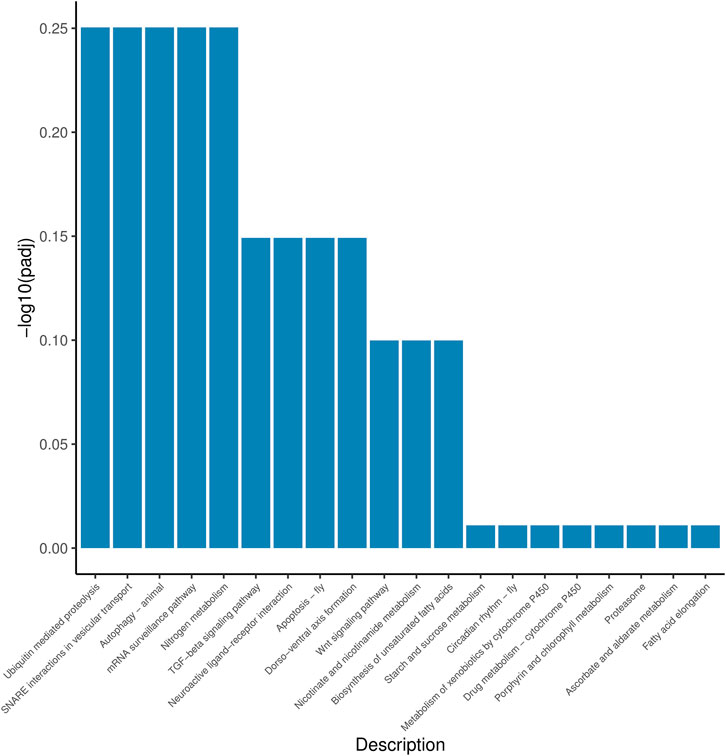
According to the position information of the candidate marker in the genome, a total of 2171 genes were located, which were used to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses. The GO results (Figure 1) showed that 30 processes were significantly enriched, of which “signaling”, “extracellular region part”, and “G-protein coupled receptor activity” were the main processes with the largest number of genes in “Biological Process”, “Cellular Component” and “Molecular Function”, respectively. Besides, a total of 20 KEGG pathways (Figure 2) were enriched, among which “Ubiquitin mediated proteolysis”, “SNARE interactions in vesicular transport”, “Autophagy”, “mRNA surveillance pathway” and “Nitrogen metabolism” were the most enriched pathways (p < 0.05).
Expression Analysis of Candidate Genes
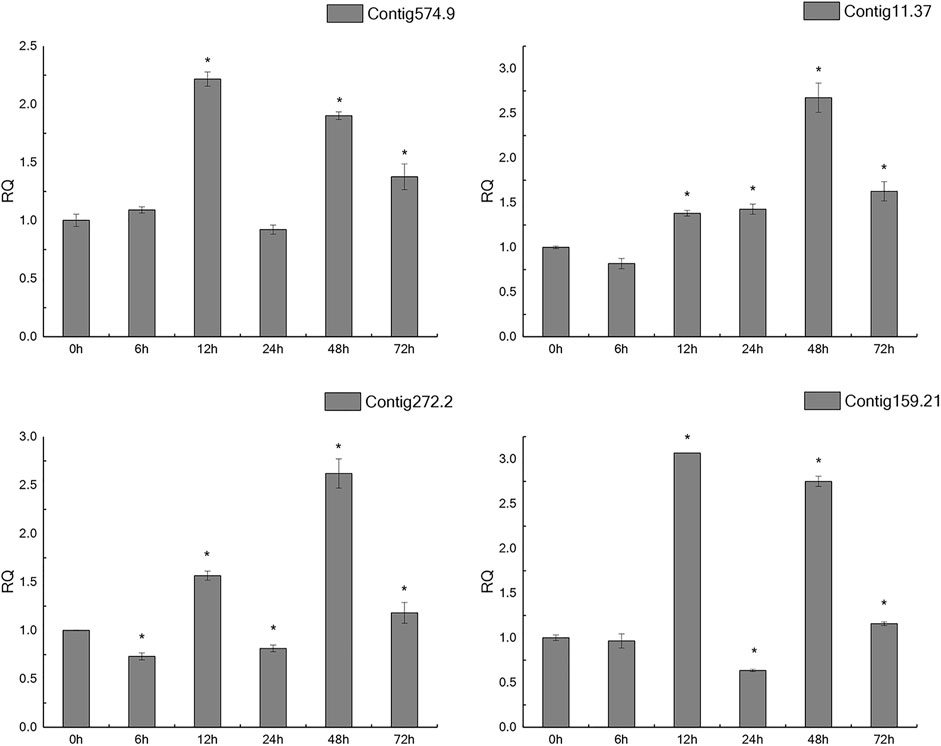
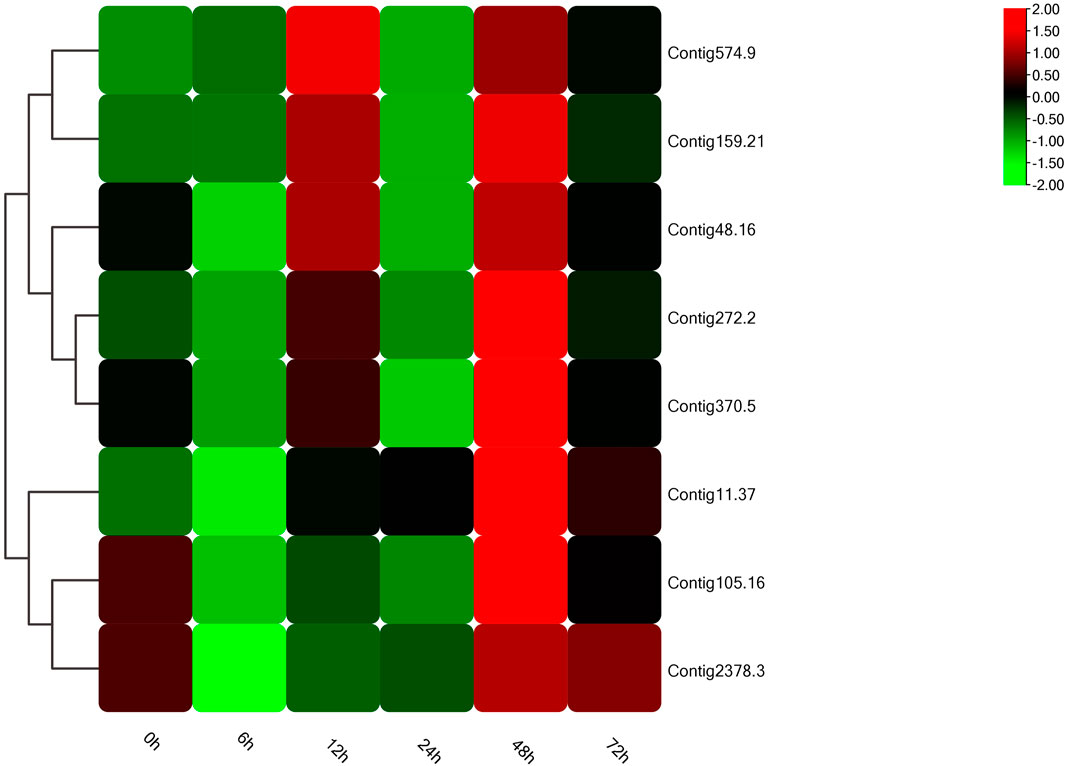
In the light of gene annotation and enrichment analysis results, eight genes related to proteolysis, autophagy, nitrogen metabolism, and vesicle transport were screened. qPCR results showed that the expression patterns of these genes changed significantly after salt-alkali stress (Figure 3), suggesting that they play a role in the adaptation to salt-alkali. The results show that these genes can be divided into two types (Figure 4). One group was significantly up-regulated at 48 h and the other group of genes was significantly up-regulated at 12 and 48 h with a trend of fluctuating expression.
Discussions
In this study, we successfully screened the salt-alkali tolerance markers in P. trituberculatus, using the BSA strategy for the first time, and then anchored several genes related to salt-alkali tolerance. The BSA method that was proposed by Michelmore and Paran, (1991) can lock the target trait genes or markers by comparing the degree of difference in the allele frequency at the polymorphic site between the two groups with markedly differentiated traits (June 2021; Rujia and Yiming, 2021; Zaiqing and Anmin, 2021). At present, research on trait-related markers of aquatic animals is mainly based on QTL and association analysis (Wang and Liu, 2017; Cao and Wang, 2021). QTL analysis relies on pedigree materials, and only excavated markers are applicable (Zhixue and Khorshed, 2021). Association analysis is often based on re-sequencing and the cost is relatively high (Hussain and Rivandi, 2007; Vermerris, 2011; Ali, 2013). Compared with the above two methods, BSA has the advantages of being able to use population materials and being cost effective and time efficient (Takagi Abe, 2013). Using the BSA strategy, trait-related markers have been successfully developed in plants (Cheng and Abass, 2020; Marc, 2020; Farhan and Qiutao, 2021; Hossein and Azin, 2021), proving it to be a feasible and economical method. However, its applicability in aquatic animals had not been reported in the past. In this study, 20 markers were selected based on BSA analysis, and eight salt-alkali tolerance related markers were finally developed efficiently. The results of the study show that BSA strategy is suitable for the mining of trait-related markers in aquatic crustaceans.
We noticed that most of the successfully verified SNPs and Indels are located on the intergenic region (81.82%). The intergenic region was once considered unimportant. However, in recent years, it has been found that these regions contain functionally important elements (Cai, 2016) and independent transcription of miRNAs with their own promoters (Hess et al., 2006). Intergenic regions may also contain unidentified functional elements, such as non-coding RNA (Zhong et al., 2014; Fariha and Changrui, 2019). It is currently believed that intergenic regions can control gene expression and inhibition, and help RNA to identify genes with low conservation and diversity (Ingvarsson et al., 2016; Bakhtiarizadeh and Salehi, 2018). The fact that most of the saline-alkali markers are located in the intergenic region, indicates that some potential functional regions of the intergenic region may have important functions in the regulation of saline-alkali adaptation. However, the specific mechanisms are still unclear and necessitate further research.
Based on the location information of markers in the genome, we initially anchored 2171 potential saline-alkali adaptation genes, which were mainly enriched in the pathway including ubiquitin-mediated proteolysis, SNARE interactions in vesicle transport and autophagy. These results indicate that the mechanism of saline-alkali adaptation is complex and co-regulated by an equally complex regulatory network. It is worth noting that the salt-alkali-related ion transport genes such as Na+-K+-ATPase are not included here. This suggests that, although ion transport plays an important role in the process of saline-alkali adaptation, it does not play a prominent role in the differentiation of salt-alkali tolerance.
In addition, according to annotation information, 8 genes were screened from the five most enriched KEGG pathways for qPCR verification. The expression patterns of these eight genes changed significantly after salt-alkali stress, indicating that they may have certain functions in salt-alkali adaptation, or salt-alkali stress has affected their normal functions. We also found that most genes were up-regulated at 12 and 48 h, suggesting the critical time for salt-alkali adaptation.
In this study, we screened the salt-tolerant markers to prove the feasibility of the BSA strategy in aquatic research. The 8 successfully verified markers provide usable markers for subsequent molecular marker-assisted breeding. Furthermore, this experiment anchored some candidate genes related to salt-alkali tolerance. These results will help to systematically clarify the molecular mechanism of salt-alkali adaptation of P. trituberculatus.
Data Availability Statement
The original contributions presented in the study are publicly available in NCBI using accession number PRJNA755748.
Author Contributions
WZ: conducted experiments, analyzed data and results, paper writing. XZ: conducted experiments. JW: collect the samples. LJ: idea for the project. JL: analyzed data and results, paper revising. BG: idea for the project. PL: paper revising.
Funding
This research was supported by the National Key RD Program of China (2018YFD0901304), Ministry of Agriculture of China and China Agriculture Research System (CARS-48), National Natural Science Foundation of China (42076116) and Projects of International Exchange and Cooperation in Agriculture, Ministry of Agriculture and Rural Affairs of China-Science, Technology and Innovation Cooperation in Aquaculture with Tropical Countries along the Belt and Road.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fgene.2022.755004/full#supplementary-material
References
Ali, F. (2013). Exploring the Genetic Basis of Northern Leaf Blight via Genome Wide Association Mapping in Maize. Wuhan, China: Huazhong Agricultural University.
Arabacı, M., and Sarı, M. (2004). Induction of Ovulation in Endemic Pearl Mullet (Chalcalburnus Tarichi), Living in the Highly Alkaline Lake Van, Using GnRHa ([D-Ser(T Bu)6, Pro9-Net]-GnRH) Combined with Haloperidol. Aquaculture. 238 (1), 529–535. doi:10.1016/j.aquaculture.2004.05.004
Bahadur, K., and Tripathi, P. (1978). Investigation if Inorganic and Organic Nitrogenous Compounds on Microbial Nitrogen Fixation with Respect to the Fixation Cycle. Zentralbl Bakteriol Naturwiss. 133 (2), 154. doi:10.1016/s0323-6056(78)80013-5
Bakhtiarizadeh, M. R., Salehi, A., and Rivera, R. M. (2018). Genome-wide Identification and Analysis of A-To-I RNA Editing Events in Bovine by Transcriptome Sequencing. PLOS ONE. 13 (2), e0193316. doi:10.1371/journal.pone.0193316
Biao, Y., and Li, Z. (2012). miR-429 Regulation of Osmotic Stress Transcription Factor 1 (OSTF1) in tilapia during Osmotic Stress. Biochem. biophysical Res. Commun. 426 (3), 294–298. doi:10.1016/j.bbrc.2012.08.029
Cai, L. (2016). Comprehensive Characterization of LincRNAs' Functions in Transcription Regulation through Long-Range Chromatin Interactions. Wuhan, China: Huazhong Agricultural University.
Cao, j., and Wang, w. (2021). Construction of a High-Density Genetic Linkage Map and QTL Mapping for Gender of Megalobrama amblycephala. J. Huazhong Agric. Univ. 40 (02), 188–196. doi:10.13300/j.cnki.hnlkxb.2021.02.020
Cheng, Q., and Abass, A. M. (2020). Comparative Transcriptome Analysis Reveals the Regulatory Effects of Acetylcholine on Salt Tolerance of Nicotiana Benthamiana. Phytochemistry. 181, 112582. doi:10.1016/j.phytochem.2020.112582
De Carvalho, I. P. C., and Detmann, E. (2012). In Vitro and In Situ Activity of Carboxymethyl Cellulase and Glutamate Dehydrogenase According to Supplementation with Different Nitrogenous Compounds. Revista Brasileira de Zootecnia. 41 (3), 683. doi:10.1590/s1516-35982012000300031
Evans, D. H. (2010). A Brief History of the Study of Fish Osmoregulation: The Central Role of the Mt. Desert Island Biological Laboratory. Front. Physiol. 1, 13. doi:10.3389/fphys.2010.00013
Farhan, U., and Qiutao, X. (2021). Histone Deacetylase HDA710 Controls Salt Tolerance by Regulating ABA Signaling in rice. J. Integr. Plant Biol. 63 (3), 451–467. doi:10.1111/jipb.13042
Fariha, K., and Changrui, L. (2019). A Review on Native and Denaturing Purification Methods for Non-coding RNA (ncRNA). J. Chromatogr. B, Anal. Tech. Biomed. Life Sci. 1120, 71–79. doi:10.1016/j.jchromb.2019.04.034
Fu, L. (2002). Effccts of Environmental Factors on the Osmoregulation of Eriocheir Sinensis. Qingdao, China: Ocean University of China.
Fu-yi, Y., Xiu-jun, L., Wang, Z-c., Zhao, C-s., and Sun, L-m. (2005). Possibility Test of Transplanting Prawn in Carbonate Saline-Alkali Waters. J. ofJilin Agric. Univ. 27 (5), 559–564. doi:10.1360/biodiv.050121
Hassan, N. M., and Fadwa, S. (2021). Dysregulation of CCAAT/enhancer Binding Protein-Alpha (CEBPA) Expression in the Bone Marrow of Acute Myeloid Leukemia Patients. Egypt. J. Med. Hum. Genet. 22 (1), 30. doi:10.1186/s43042-021-00154-z
Hess, N. K., Singer, P. A., Trinh, K., Nikkhoy, M., and Bernstein, S. I. (2006). Transcriptional Regulation of the Drosophila melanogaster Muscle Myosin Heavy-Chain Gene. Gene Expr. Patterns. 7 (4), 413–422. doi:10.1016/j.modgep.2006.11.007
Hossein, S., and Azin, T. (2021). Crushed maize Seeds Enhance Soil Biological Activity and Salt Tolerance in Caper (Capparis Spinosa L.). Ind. Crops Prod. 160, 113103. doi:10.1016/j.indcrop.2020.113103
Hu, Z., Li, Z., Yu, J., Tong, C., Lin, Y., Guo, X., et al. (2014). Association Analysis Identifies New Risk Loci for Non-obstructive Azoospermia in Chinese Menfluences of Different Salinities on Growth and Survival Rate of Portunus Trituberculatus. Nat. Commun. 5 (10), 3857–3866. doi:10.1038/ncomms4857
Hussain, S. S., and Rivandi, A. (2007). Molecular Breeding for Drought Tolerance in Plants:Wheat Perspective. Proc. Pak. Acad. Sci., 31–58.
Ingvarsson, P. K., Hvidsten, T. R., and Street, N. R. (2016). Towards Integration of Population and Comparative Genomics in forest Trees. New Phytol. 212 (2), 338–344. doi:10.1111/nph.14153
Juan, D., and Gaoli, Z. (2021). High-intensity Light of Full-Spectrum LED Promotes Survival Rate but Not Development of the Larval Swimming Crab Portunus Trituberculatus. Aquacultural Eng. 93, 102158. doi:10.1016/j.aquaeng.2021.102158
Jun, L. J. (2021). Comparative Transcriptomics and RNA-Seq-Based Bulked Segregant Analysis Reveals Genomic Basis Underlying Cronartium Ribicola Vcr2 Virulence& 13. Front. Microbiol. 12, 602812. doi:10.3389/fmicb.2021.602812
Kai, W., and Mingyao, L. (2010). ANNOVAR: Functional Annotation of Genetic Variants from High-Throughput Sequencing Data. Narnia. 38 (16).
Li, H., and Durbin, R. (2009). Fast and Accurate Short Read Alignment with Burrows-Wheeler Transform. Bioinformatics. 25 (14), 1754–1760. doi:10.1093/bioinformatics/btp324
Li, H., Wysoker, B. A., Fennell, T., Ruan, J., Homer, N., Marth, G., et al. (2009). The Sequence Alignment/Map Format and SAMtools. Bioinformatics. 25 (16), 2078–2079. doi:10.1093/bioinformatics/btp352
Li, M., Li, J., Shi, K., He, Y., Gao, B., Liu, P., et al. (2021). Estimation of Heritability and Genetic Correlation of saline-alkali Tolerance in Exopalaemon Carinicauda. Prog. Fish. Sci. 42 (1), 117–123. doi:10.19663/j.issn2095-9869.20200221001
Liu, F. (2016). Effects of Carbonate Alkalinity Stresses on the Growth and Reproduction of Exopalaemon Carinicauda. Shanghai, China: Shanghai Ocean University.
Lv, J., Sun, D., Huan, P., Song, L., Liu, P., and Li, J. (2018). QTL Mapping and Marker Identification for Sex-Determining: Indicating XY Sex Determination System in the Swimming Crab (Portunus Trituberculatus). Front. Genet. 9, 337. doi:10.3389/fgene.2018.00337
Lv, J., Sun, D., Yan, D., Ti, X., Liu, P., and Li, J. (2019). Quantitative Trait Loci Mapping and Marker Identification for Low Salinity Tolerance Trait in the Swimming Crab (Portunus Trituberculatus). Front. Genet. 10, 1193. doi:10.3389/fgene.2019.01193
Lv, J., Gao, B., Liu, P., Li, J., and Meng, X. (2017). Linkage Mapping Aided by De Novo Genome and Transcriptome Assembly in Portunus Trituberculatus: Applications in Growth-Related QTL and Gene Identification. Sci. Rep. 7, 7874. doi:10.1038/s41598-017-08256-8
Marc, S. (2020). How a Mangrove Tree Can Help to Improve the Salt Tolerance of Arabidopsis and Rice. Plant Physiol. 184 (4), 1630–1632. doi:10.1104/pp.20.01370
McKenna, A., Hanna, M., Banks, E., Sivachenko, A., Cibulskis, K., Kernytsky, A., et al. (2010). The Genome Analysis Toolkit: A MapReduce Framework for Analyzing Next-Generation DNA Sequencing Data. Genome Res. 20 (9), 1297–1303. doi:10.1101/gr.107524.110
Michelmore, R. W., Paran, I., and Kesseli, R. V. (1991). Identification of Markers Linked to Disease-Resistance Genes by Bulked Segregant Analysis: A Rapid Method to Detect Markers in Specific Genomic Regions by Using Segregating Populations. Proc. Natl. Acad. Sci. U S A. 88 (21), 9828–9832. doi:10.1073/pnas.88.21.9828
Nuo, S., and Zeya, Z. (2021). Generation of an Induced Pluripotent Stem Cell Line from a Congenital Microtia Patient with 4p16.1 Microduplication Involving the Long-Range Enhancer of HMX1. Stem Cel Res.
Rujia, Z., and Yiming, R. (2021). Mapping of Genetic Locus for Leaf Trichome Formation in Chinese Cabbage Based on Bulked Segregant Analysis. Plants. 10 (4), 771. doi:10.3390/plants10040771
Ryoji, K., and Yohko, K. (2021). Distinct Foxp3 Enhancer Elements Coordinate Development, Maintenance, and Function of Regulatory T Cells. Immunity.
Shen, A., Jiang, K., and Shen, X. (2014). Effects of Salinity on the mRNA Expression of Na+/K+-ATPase and HSP in the Ridgetail white Prawn, Exopalaemon Carinicauda. Mar. Environ. Sci. 33 (05), 693–698.
Takagi, H., Abe, A., Yoshida, K., Kosugi, S., Natsume, S., Mitsuoka, C., et al. (2013). QTL-seq: Rapid Mapping of Quantitative Trait Loci in rice by Whole Genome Resequencing of DNA from Two Bulked Populations. Plant J. 74 (1), 174–183. doi:10.1111/tpj.12105
Vermerris, W. (2011). Survey of Genomics Approaches to Improve Bioenergy Traits in maize, Sorghum and Sugarcane. J. Integr. Plant Biol. 53 (02), 105–119. doi:10.1111/j.1744-7909.2010.01020.x
Wang, X., Liu, S., Jiang, C., Geng, X., Zhou, T., Li, N., et al. (2017). Multiple Across-Strain and Within-Strain QTLs Suggest Highly Complex Genetic Architecture for Hypoxia Tolerance in Channel Catfish. Mol. Genet. Genomics. 292 (1), 63–76. doi:10.1007/s00438-016-1256-2
Whittamore, J. M. (2012). Osmoregulation and Epithelial Water Transport: Lessons from the Intestine of marine Teleost Fish. J. Comp. Physiol. B. 182 (1), 1–39. doi:10.1007/s00360-011-0601-3
Wilkie, M. P., and Wright, P. A. (1994). The Physiological Adaptations of the Lahontan Cutthroat Trout (Oncorhynchus clarki Henshawi) Following Transfer from Well Water to the Highly Alkaline Waters of Pyramid Lake, Nevada (pH 9.4). Physiol. Zoolog. 67 (2), 355–380. doi:10.1086/physzool.67.2.30163853
Wood, C. M., Bergman, H. L., Bianchini, A., Laurent, P., Maina, J., Johannsson, O. E., et al. (2012). Transepithelial Potential in the Magadi tilapia, a Fish Living in Extreme Alkalinity. J. Comp. Physiol. B. 182 (2), 247–258. doi:10.1007/s00360-011-0614-y
Wu, X., Wang, Q., Lou, B., Liu, Z., and Cheng, Y. (2014). Effects of Fattening Period on Ovarian Development and Nutritional Quality of Female Swimming Crab ( Portunus Trituberculatus). Journal Offisheries China. 38 (02), 170–182. doi:10.3724/SP.J.1231.2014.48934
Xu, Z., Zhang, J., Zhang, W., Liang, J., Jian, L., and Li, J. (2020). Enzyme Activity and Gene Expressions of Immune-Related in Exopalaemon Carinicauda Following Long-Term Different Salinity Exposure. Henan Fish. 4 (01), 17–20.
Yao, Z. L., and Wang, H. (2012). Transcriptomic Profiles of Japanese Medaka (Oryzias latipes) in Response to Alkalinity Stress. Genet. Mol. Res. : GMR 11 (3), 2200. doi:10.4238/2012.June.15.2
Ye, Y., and Fangmin, X. (2021). Untargeted Lipidomics Reveals Metabolic Responses to Different Dietary N-3 PUFA in Juvenile Swimming Crab (Portunus Trituberculatus). Food Chem. 354, 129570. doi:10.1016/j.foodchem.2021.129570
Zaiqing, W., and Anmin, Y. (2021). Bulked Segregant Analysis Reveals Candidate Genes Responsible for dwarf Formation in Woody Oilseed Crop castor Bean. Scientific Rep. 11 (1), 6277. doi:10.1038/s41598-021-85644-1
Zhao, Y., Wu, J. W., Wang, Y., and Zhao, J. L. (2016). Role of miR-21 in Alkalinity Stress Tolerance in tilapia. Biochem. Biophys. Res. Commun. 471 (1), 26–33. doi:10.1016/j.bbrc.2016.02.007
Zhixue, D., and Khorshed, A. M. (2021). Mapping of a Major QTL Controlling Plant Height Using a High-Density Genetic Map and QTL-Seq Methods Based on Whole-Genome Resequencing in Brassica Napus. G3: Genes, Genomes, Genet. 11 (7), jkab118. doi:10.1093/g3journal/jkab118
Zhong, C., Andrews, J., and Zhang, S. (2014). Discovering Non-coding RNA Elements in Drosophila 3' Untranslated Regions. Int. J. Bioinform Res. Appl. 10 (4/5), 479–497. doi:10.1504/IJBRA.2014.062996
Keywords: BSA, INDEL, Portunus trituberculatus, salt-alkali, SNP
Citation: Zhang W, Zhao XY, Wu J, Jin L, Lv J, Gao B and Liu P (2022) Screening and Verification of Molecular Markers and Genes Related to Salt-Alkali Tolerance in Portunus trituberculatus. Front. Genet. 13:755004. doi: 10.3389/fgene.2022.755004
Received: 07 August 2021; Accepted: 12 January 2022;
Published: 08 February 2022.
Edited by:
Li Zhou, Institute of Hydrobiology (CAS), ChinaCopyright © 2022 Zhang, Zhao, Wu, Jin, Lv, Gao and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ping Liu, bGl1cGluZ0B5c2ZyaS5hYy5jbg==
 Wen Zhang
Wen Zhang Xiao Yan Zhao1
Xiao Yan Zhao1 Ping Liu
Ping Liu