Abstract
Parkinson's disease (PD) patient care is limited by inadequate, sporadic symptom monitoring, infrequent access to care, and sparse encounters with healthcare professionals leading to poor medical decision making and sub-optimal patient health-related outcomes. Recent advances in digital health approaches have enabled objective and remote monitoring of impaired motor function with the promise of profoundly changing the diagnostic, monitoring, and therapeutic landscape in PD. We recently demonstrated that by using a variety of upper limb functional tests iMotor, an artificial intelligence powered, cloud-based digital platform differentiated PD subjects from healthy volunteers (HV). The objective of this paper is to provide preliminary evidence that artificial intelligence systems may allow one to discriminate PD patients from (HV) further and determine different features of the disease within a cohort of PD subjects. The recently introduced Neural Network Construction (NNC) technique was used here to classify data collected by a mobile application (iMotor, Apptomics Inc., Wellesley, MA) into two categories: PD for patients and HV. The method was tested on a series of data previously collected, and the results were compared against more traditional techniques for neural network training. The NNC algorithm discriminated individual PD patients from HVs with 93.11% accuracy and ON vs. OFF state with 76.5% accuracy. Future applications of artificial intelligence-powered digital platforms can enhance clinical care and research by generating rich, reliable, and sensitive datasets that can be used for medical decision-making during and between office visits. Additional artificial intelligence-based studies in larger cohorts of patients are warranted.
Introduction
Parkinson's disease (PD) is a prevalent neurodegenerative disease affecting about 1% of the world population over the age of 55 (Nussbaum and Ellis, 2003). About five million people worldwide are estimated to have PD. PD Prevalence is expected to double by the year 2030 (Dorsey et al., 2007). PD is diagnosed in the presence of two or more cardinal motor symptoms such as rest tremor, bradykinesia (slow movement), or rigidity (stiffness). Bradykinesia is universally present in PD and responds relatively well to existing treatments (Hughes et al., 1992).
For the last 50 years symptomatic treatment of PD has focused on replacing the declining levels of the neurotransmitter dopamine with the orally administered dopamine precursor, levodopa (L-Dopa). However, chronic administration of L-Dopa leads to side effects, such as fluctuations in motor performance and dyskinesias (Sweet and McDowell, 1975) as the disease progresses, after more than 3–5 years of therapy. These phenomena can be as troublesome as the disease itself (Olanow et al., 2006).
Parkinson's disease (PD) patient care is limited by inadequate, sporadic symptom monitoring, infrequent access to care, and sparse encounters with healthcare professionals leading to poor medical decision making and sub-optimal patient health-related outcomes. More frequent patient monitoring and treatment adjustments can lead to better symptomatic management and reduction in treatment-related complications such as motor fluctuations (Papapetropoulos et al., 2015).
Recent advances in digital health approaches have enabled objective and remote monitoring of impaired motor function with the promise of profoundly changing the diagnostic, monitoring, and therapeutic landscape in PD (Pasluosta et al., 2015). Sensing technologies, mobile networks, cloud computing, the Internet of Things and big data analytics innovations that have the potential to transform healthcare and our approach to patients with chronic, complex, and disorders like PD (Espay et al., 2016).
Tapping tests conducted through the interface of fingers and/or other areas of the hand and a touch sensing screen has been shown to measure motor function in PD (Djuric-Jovicic et al., 2016; Giancardo et al., 2016; Mitsi et al., 2017). PD patients frequently develop fluctuations in motor function as a side effect of commonly used anti-parkinsonian medications. These fluctuations, termed “ON” and “OFF” states, are difficult to manage in clinical care and often serve as endpoints in PD clinical trials (Papapetropoulos, 2012).
iMotor is a clinically validated digital platform (iMotor; Apptomics Inc., Wellesley Hills, MA, USA) that utilizes the smart-tablet's screen sensing capabilities during upper limb function tests (similar to ones conducted in the neurologists' office) to objectively collect data that allow detection and quantification of neuromotor function. It has been tested in various movement disorders including PD and more recently Essential tremor (Mitsi et al., 2017). A series of screenshots from the application are presented in Figure 1.
Figure 1
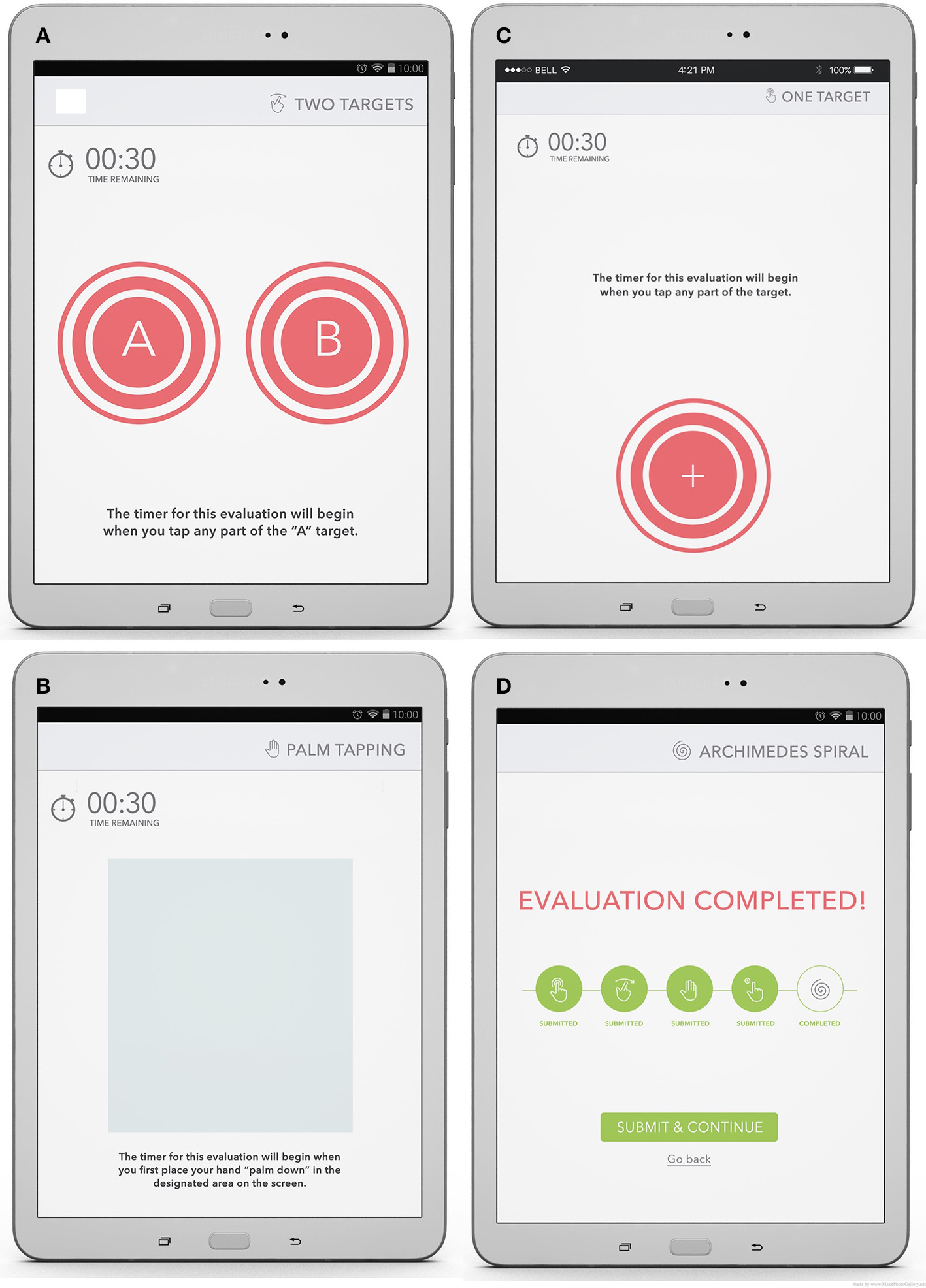
Screenshots of iMotor-based tapping tests. (A) Two-target test: the participants alternatingly tapped with the index finger, as fast and as accurately as possible, the centers of two concentric circles on the tablet screen. (B) Pronation-supination test: the participants alternatingly tapped the palmar and dorsal surfaces of their hand as fast as possible on the tablet screen. (C) Reaction time: the participants tap on the target with their index finger following a visual queue as fast as possible. (D) Task completion screen: sample patient score report, available immediately to the patient.
We demonstrated that by using a variety of upper limb functional tests iMotor differentiated PD subjects from healthy volunteers (HV) (Mitsi et al., 2017; Wissel et al., 2017). The objective of this paper is to provide preliminary evidence that artificial intelligence systems may allow one to discriminate PD patients from HV further and determine different features of the disease within a cohort of PD subjects.
Materials and Methods
Data from a study (10) with the iMotor application were collected from 36 participants (19 with PD and 17 healthy controls) for the following tasks: (1) Two-target finger tapping test, (2) Pronation–supination test, and (3) Reaction time test: Details on the execution of these tests are provided in Figure 1 and described in details in previous papers on iMotor (Mitsi et al., 2017; Wissel et al., 2017).
In this paper, experiments were conducted on the data collected by iMotor to automatically classify records into two categories: HV and PD. The method used for the classification is the recently introduced classification model of constructed neural networks. Neural networks are well-established parametric tools (Cybenko, 1989; Bishop, 1995), used with success in many areas such as recognition of patterns (Artyomov and Yadid-Pecht, 2005), signal processing (Uncini, 2003), astronomy (Valdas and Bonham-Carter, 2006), and mathematics (Lagaris et al., 1998).
A neural network can be expressed as a function N where is the input vector and is the vector of the parameters to be estimated (weight vector). Modification of the vector of parameters (neural network training) is conducted by minimizing the equation below (also named error function):
In the above the set is the data used to train the neural network. The symbol yi stands for the output of the function estimated at the point . Some training methods used are Back Propagation (Rumelhart et al., 1986), RPROP (Riedmiller and Braun, 1993), Genetic Algorithms (Yao, 1999), and Particle Swarm Optimization (Zhang et al., 2000). The constructed neural networks with grammatical evolution have been described by Tsoulos et al. (2008) and are based on the Grammatical Evolution technique (O'Neill and Ryan, 2001) to evolve the neural network topology along with the network parameters. The method of constructing neural networks has been also used with success in other problems such as solving differential equations (Tsoulos et al., 2009) and in problem of locating Amide I bonds in chemistry (Papamokos et al., 2009). The results from this method are compared against the results from neural network trained with a local optimization procedure and against the results from neural network trained with a hybrid genetic algorithm. The Grammatical Evolution procedure, as well as the proposed method, are outlined in the following subsections.
Grammatical Evolution
Every chromosome in the genetic population is a vector of integers, which is mapped through a mapping procedure described in O'Neill and Ryan (2001) governed by the BNF grammar of Figure 2 into an artificial neural network with one hidden level. The output of the constructed neural network is a summation of different sigmoidal units, and it can be formulated as:
where x ϵ Rd, is the number of hidden nodes and the vector p denotes the parameters (weights) of the neural network. The function σ (x) is the sigmoid function:
The constant nodes represent the total number of input parameters. The sigmoid function is used as activation function because it is continuous and differentiable at every point. Also, the derivatives of this function can easily be estimated and have been used with success in many applications of neural networks.
Figure 2

Example of a one-point crossover.
Steps of the Algorithm
The main steps of the algorithm have as follows:
- Initialize
the following parameters:
The number of chromosomes S.
The maximum generations allowed K.
The crossover rate pc ϵ [0, 1].
The mutation rate pm ϵ [0, 1].
A positive number ε < 1, used for the termination rule.
A positive integer number G which determines the frequency of the application of the local search procedure.
A positive integer parameter M which provides the number of chromosomes that participate in the local search procedure.
Set iters = 0, as the current number of generation.
Initialize the S chromosomes. Each chromosome is mapped to a neural network as noted in the subsection of Grammatical Evolution.
Calculate the fitness for every chromosome, using Equation 1 and
Applied the genetic operations of selection and mutation to the population.
Selection procedure: The chromosomes are sorted in descending order according to their fitness value. The first (1 − ps)∗S chromosomes are transferred to the next generation. The rest of the chromosomes are substituted by offsprings created through crossover procedure: For every offspring, two chromosomes (parents) are selected from the old population using tournament selection. The procedure of tournament selection is as follows: A set of N > 1 randomly selected chromosomes is produced, the chromosome with the best fitness value in this set is selected, and the others are discarded. Each offspring is created by the parents using the one-point crossover. During one point crossover, the parent chromosomes are cut at a randomly selected point, and their right-hand side sub-chromosomes are exchanged as shown in Figure 2.
Mutation procedure: For every element of each chromosome a random number r in a range [0,1] is produced. If r ≤ pm then the corresponding element is randomly altered. For the case of grammatical evolution simply a new integer in the range [0,255] is produced.
Replace the ps∗S worst chromosomes in the population with the offsprings created by the genetic operators.
Set iters = iters+1.
If iters mod G = 0 then
For i = 1..M do
Select a chromosome Ri randomly from the genetic population.
Construct with the Grammatical Evolution procedure the corresponding neural network (Ri).
Train the neural network N(Ri) with a local optimization procedure. This process enhances the weights created by the Grammatical Evolution procedure. The local search method used is a BFGS variant of Powell.
End for
Endif
If iters ≥ K or the best chromosomes has fitness value below the predefined threshold & epsi; terminate, else go to step 3.
Data Description
The method was evaluated on three different datasets collected using the iMotor digital platform using methodologies previously described (Mitsi et al., 2017; Wissel et al., 2017). In summary:
The first dataset named TWO_TARGET is produced from the two-target finger tapping test: (tapping of the index finger on the tablet screen). The dataset contains the following variables:
Total number of taps during a test period of 30 s.
Speed the index finger travels between two targets during the test period.
Average interval between two consecutive finger screen taps.
Average accuracy as measured by the average number of pixels from the center of the target center.
The second dataset is called PALM (tapping of the hand surface on the tablet screen). This dataset is produced from the pronation-supination test and contains the following variables:
Total number of taps during a test period of 30 s.
Average interval between two consecutive finger screen taps.
Average accuracy as measured by the average number of pixels from the center of the target center.
The third test is called REACTION (tapping of index finger on a target on following a color shift on the tablet screen). This dataset is produced by the reaction time test and has the following features:
Average interval between two consecutive finger screen taps.
Average accuracy as measured by the average number of pixels from the center of the target center.
A cluster of Intel machines running Ubuntu Linux 14.04 were utilized to conduct the analysis. Experiments were run 30 times with random seeds for the embedded generator of the C programming language and averages were used each time. Values for parameters of the Neural Network Construction method are shown in Table 1. The 10-folding validation was used.
Table 1
| Parameter | Value |
|---|---|
| S | 200 |
| K | 50 |
| G | 200 |
| M | 2 |
| pc | 0.9 |
| pm | 0.05 |
Experimental parameters.
The following classification methods were tested:
BAYES. The Naive Bayes classifier methodology (Webb et al., 2005).
SIMPLE. A neural network with 10 hidden nodes was used. The network was trained using the optimization method of Powell.
GENETIC. A neural network with 10 hidden nodes was used. The network was trained using a hybrid genetic algorithm using the same parameters shown in Table 1. After the termination of the genetic algorithm, the BFGS variant of Powell was used to minimize further the training error.
NNC. A hybrid method utilizing constructed neural networks by grammatical evolution and a local optimization procedure is used to train a neural network. The local optimization procedure was a combination of the BFGS variant of Powell and Differential Evolution.
Results
PD subjects included in our analysis had an average age of 67.9 (SD 8.8) years, a mean disease duration of 6.5 (SD 4.6) years, and a mean UPDRS score was 24.6 (SD 6.7). All PD subjects were Hoehn & Yahr stage 2 indicating moderate disease severity.
The NNC algorithm discriminated individual PD patients from HVs with 93.11% accuracy (Table 2) and ON vs. OFF state with 76.5% accuracy (Table 3). Results from alternative classification methodologies are presented in Tables 2, 3. The discriminatory precision of Naive Bayes appears lower compared to the NNC algorithm when all variables were combined.
Table 2
| Dataset | Bayes (%) | Simple (%) | Genetic (%) | NNC (%) |
|---|---|---|---|---|
| 2-target tapping | 73.33 | 64.11 | 67.10 | 69.98 |
| Pronation/Supination | 70.00 | 57.67 | 63.89 | 69.00 |
| Reaction time | 76.67 | 71.33 | 74.67 | 83.90 |
| ALL variable combined | 80.00 | 74.33 | 79.89 | 93.11 |
Accuracy of the NNC algorithm in healthy volunteers vs. PD patients (% correct).
Dataset (sensitivity/specificity) for NNC, 2 Target tapping (93/53%), Pronation/supination (92.50%/50%), Reaction time (89.50/78.50%), all variable (97.50%/85%).
Table 3
| Dataset | Simple (%) | Genetic (%) | NNC (%) |
|---|---|---|---|
| MOST vs. LESS affected side during ON | 59.50 | 61.50 | 75.83 |
| MOST vs. LESS affected side during OFF | 61.67 | 65.00 | 71.50 |
| ON vs. OFF MOST affected side | 68.16 | 68.50 | 76.50 |
| ON vs. OFF less affected side | 74.33 | 75.00 | 75.83 |
Accuracy of the NNC algorithm in detecting motor fluctuations in PD patients (% correct).
Discussion
This study investigated the ability of artificial intelligence systems to discriminate PD patients from HV and determine the patient motor status (ON vs. OFF) in a clinic-based cohort. To our knowledge, this is the first attempt to compare different artificial intelligence classification methodologies in finger tapping data. Despite the small sample size, the discriminatory precision of the NNC machine learning algorithm for both diagnostic purposes and motor status detection is encouraging. A recent study with similar objectives, attempted automatic discrimination of a movement pattern related to the risk of fall in PD termed as “freezing of gait (FoG) using machine learning techniques in a wearable accelerometry dataset derived from 36 patients (Aich et al., 2018). The classification accuracy was ~88% generating evidence for the real-life application of wearable accelerometer-based system to assess and monitor the FoG. Additional studies report similar results in FoG classification (Palmerini et al., 2017).
Early detection of Parkinson's disease and systematic early and accurate detection of motor fluctuations (and other motor symptoms) using artificial intelligence may have significant therapeutic implications leading to improvements research methodologies and most importantly in PD patient care.
Enrichment of machine learning algorithms with additional sensor data (i.e., accelerometry, gyrometry, spirography) may enhance the precision of the algorithm. More recent versions of iMotor have incorporated some of these capabilities, and additional studies are currently ongoing.
Future applications of artificial intelligence-powered digital platforms can supplement standard clinical care and research methodologies by providing rich, reliable, and sensitive datasets during and between office visits. Additional artificial intelligence-based studies in larger cohorts of patients are warranted.
Statements
Author contributions
IT: project conception, design, statistical analysis, execution, manuscript authoring, review, and final approval. GM: project conception and execution, and manuscript authoring, review and final approval. AS: research execution, manuscript review, and final approval. SP: project conception, design, analysis interpretation, execution, manuscript authoring, review, and final approval.
Conflict of interest
SP and GM have a financial interest in Apptomics Inc. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1
AichS.PradhanP. M.ParkJ.SethiN.VathsaV. S. S.KimH. C. (2018). A validation study of freezing of gait (FoG) detection and machine-learning-based FoG prediction using estimated gait characteristics with a wearable accelerometer. Sensors.18:E3287. 10.3390/s18103287
2
ArtyomovE.Yadid-PechtE. O. (2005). Modified high-order neural network for invariant pattern recognition. Pattern Recogn. Lett.26, 843–851. 10.1016/j.patrec.2004.09.029
3
BishopC. (1995). Neural Networks for Pattern Recognition. Oxford: Oxford University Press.
4
CybenkoG. (1989). Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst.2, 303–314. 10.1007/BF02551274
5
Djuric-JovicicM.PetrovicI.Jecmenica-LukicM.RadovanovicS.Dragasevic-MiskovicN.BelicM.et al. (2016). Finger tapping analysis in patients with Parkinson's disease and atypical parkinsonism. J. Clin. Neurosci.30, 49–55. 10.1016/j.jocn.2015.10.053
6
DorseyE. R.ConstantinescuR.ThompsonJ. P.BiglanK. M.HollowayR. G.KieburtzK.et al. (2007). Projected number of people with Parkinson disease in the most populous nations, 2005 through 2030. Neurology68, 384–386. 10.1212/01.wnl.0000247740.47667.03
7
EspayA. J.BonatoP.NahabF. B.MaetzlerW.DeanJ. M.KluckenJ.et al. (2016). Technology in Parkinson's disease: challenges and opportunities. Mov. Disord.31, 1272–1282. 10.1002/mds.26642
8
GiancardoL.Sanchez-FerroA.Arroyo-GallegoT.ButterworthI.MendozaC. S.MonteroP.et al. (2016). Computer keyboard interaction as an indicator of early Parkinson's disease. Sci. Rep.6:34468. 10.1038/srep34468
9
HughesA. J.DanielS. E.KilfordL.LeesA. J. (1992). Accuracy of clinical diagnosis of idiopathic Parkinson's disease: a clinico-pathological study of 100 cases. J. Neurol. Neurosurg. Psychiatry55, 181–4. 10.1136/jnnp.55.3.181
10
LagarisI. E.LikasA.FotiadisD. I. (1998). Artificial neural networks for solving ordinary and partial differential equations. IEEE Trans. Neural Netw.9, 987–1000. 10.1109/72.712178
11
MitsiG.MendozaE. U.WisselB. D.BarbopoulouE.DwivediA. K.TsoulosI.et al. (2017). Biometric digital health technology for measuring motor function in Parkinson's disease: results from a feasibility and patient satisfaction study. Front. Neurol.8:273. 10.3389/fneur.2017.00273
12
NussbaumR. L.EllisC. E. (2003). Alzheimer's disease and Parkinson's disease. N. Engl. J. Med.348, 1356–64. 10.1056/NEJM2003ra020003
13
OlanowC. W.ObesoJ. A.StocchiF. (2006). Continuous dopamine-receptor treatment of Parkinson's disease: scientific rationale and clinical implications. Lancet Neurol.5, 677–687. 10.1016/S1474-4422(06)70521-X
14
O'NeillM.RyanC. (2001). Grammatical Evolution. Evolu. Comput.5, 349–358. 10.1109/4235.942529
15
PalmeriniL.RocchiL.MaziluS.GazitE.HausdorffJ. M.ChiariL. (2017). Identification of characteristic motor patterns preceding freezing of gait in parkinson's disease using wearable sensors. Front. Neurol.8:394. 10.3389/fneur.2017.00394
16
PapamokosG. V.TsoulosI. G.DemetropoulosI. N.GlavasE. (2009). Location of amide I mode of vibration in computed data utilizing constructed neural networks. Expert Syst. Appli.36, 12210–12213. 10.1016/j.eswa.2009.04.065
17
PapapetropoulosS.MitsiG.EspayA. J. (2015). Digital health revolution: is it time for affordable remote monitoring for Parkinson's disease?Front. Neurol.6:34. 10.3389/fneur.2015.00034
18
PapapetropoulosS. S. (2012). Patient diaries as a clinical endpoint in Parkinson's disease clinical trials. CNS Neurosci Ther.18, 380–387. 10.1111/j.1755-5949.2011.00253.x
19
PasluostaC. F.GassnerH.WinklerJ.KluckenJ.EskofierB. M. (2015). An emerging era in the management of Parkinson's disease: wearable technologies and the internet of things. IEEE J. Biomed. Health Inform.19, 1873–1881. 10.1109/JBHI.2015.2461555
20
RiedmillerM.BraunH. (1993). A direct adaptive method for faster backpropagation learning: the RPROP algorithm, in IEEE International Conference on Neural Networks (San Francisco, CA), 586–591.
21
RumelhartD. E.HintonG. EWilliamsR. J. (1986). Learning representations by back-propagating errors. Nature323, 533–536. 10.1038/323533a0
22
SweetR. D.McDowellF. H. (1975). Five years' treatment of Parkinson's disease with levodopa. Therapeutic results and survival of 100 patients. Ann. Intern. Med.83, 456–463. 10.7326/0003-4819-83-4-456
23
TsoulosI. G.GavrilisD.GlavasE. (2009). Solving differential equations with constructed neural networks. Neurocomputing72, 2385–2391. 10.1016/j.neucom.2008.12.004
24
TsoulosI. G.GavrilisD. D.GlavasE. (2008). Neural network construction and training using grammatical evolution. Neurocomputing72, 269–277. 10.1016/j.neucom.2008.01.017
25
UnciniA. (2003). Audio signal processing by neural networks. Neurocomputing55, 593–625. 10.1016/S0925-2312(03)00395-3
26
ValdasJ. J.Bonham-CarterG. (2006). Time dependent neural network models for detecting changes of state in complex processes: applications in earth sciences and astronomy. Neural Netw.19, 196–207. 10.1016/j.neunet.2006.01.006
27
WebbG. I.BoughtonJ.WangZ. (2005). Not so naive bayes: aggregating one-dependence estimators. Mach. Learn.58, 5–24. 10.1007/s10994-005-4258-6
28
WisselB. D.MitsiG.DwivediA. K.PapapetropoulosS.LarkinS.López CastellanosJ. R.et al. (2017). Tablet-based application for objective measurement of motor fluctuations in Parkinson disease. Digital Biomark.1, 126–135. 10.1159/000485468
29
YaoX. (1999). Evolving artificial neural networks. Proc. IEEE87, 1423–1447. 10.1109/5.784219
30
ZhangC.ShaoH.LiY. (2000). Particle swarm optimisation for evolving artificial neural network, in IEEE International Conference on Systems, Man, and Cybernetics (Los Angeles, CA), 2487–2490.
Summary
Keywords
Parkinson disease, digital health, digital biomarker, artificial intelligence, objective monitoring, motor function, neural networks, grammatical evolution
Citation
Tsoulos IG, Mitsi G, Stavrakoudis A and Papapetropoulos S (2019) Application of Machine Learning in a Parkinson's Disease Digital Biomarker Dataset Using Neural Network Construction (NNC) Methodology Discriminates Patient Motor Status. Front. ICT 6:10. doi: 10.3389/fict.2019.00010
Received
17 January 2019
Accepted
25 April 2019
Published
14 May 2019
Volume
6 - 2019
Edited by
Ioanna Chouvarda, Aristotle University of Thessaloniki, Greece
Reviewed by
Mitsunori Ogihara, University of Miami, United States; Steven Demets, Quant ICT, Belgium
Updates
Copyright
© 2019 Tsoulos, Mitsi, Stavrakoudis and Papapetropoulos.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Spyros Papapetropoulos spapapetropoulos@mgh.harvard.edu
This article was submitted to Digital Health, a section of the journal Frontiers in ICT
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.