- 1Division of Life Science, The Hong Kong University of Science and Technology, Hong Kong, China
- 2HKUST Shenzhen Research Institute, Shenzhen, China
Cyanobacteria exhibit biological rhythms as an adaptation to the daily light-dark (diel) cycle. Light is also crucial for bacteriophages (cyanophages) that infect cyanobacteria. As the first step of infection, the adsorption of some cyanophages to their host cells is light-dependent. Moreover, cyanophage replication is affected by light intensity and possibly the host cell cycle. Photosynthesis and carbon metabolism genes have been found in cyanophage genomes. With these genes, cyanophages may affect the host metabolic rhythm. Field studies suggest that cyanophage infection of cyanobacteria in aquatic environments is synchronized directly or indirectly to the light-dark cycle. These discoveries are beginning to reveal how the daily light-dark cycle shapes the interaction of cyanophages and cyanobacteria, which eventually influences matter and energy transformation in aquatic environments.
Introduction
The cyanobacteria are a large and diverse group of microorganisms that are typically oxygenic photoautotrophs. This group comprises the unicellular cyanobacteria Synechococcus and Prochlorococcus, which are responsible for a vast majority of primary production in freshwater and oligotrophic marine ecosystems (Scanlan and West, 2002). Bacteriophages (cyanophages) that infect cyanobacteria are abundant in aquatic environments (Bergh et al., 1989; Proctor and Fuhrman, 1990; Suttle et al., 1990), and 15% of marine cyanobacteria are estimated to be infected by cyanophages at any given time (Proctor and Fuhrman, 1990). Three morphotypes of lytic cyanophages have been found to infect Synechococcus (Suttle and Chan, 1993; Waterbury and Valois, 1993; Wilson et al., 1993) and Prochlorococcus (Sullivan et al., 2003): T4-like myoviruses, T7-like podoviruses, and siphoviruses. They are all double-stranded DNA viruses. Similar to bacteriophages infecting heterotrophic bacteria, the life cycle of cyanophages consists of adsorption to a host cell and replication, formation, and eventually release of mature progeny phages after lysis of the infected host cells. Different from other bacteriophage groups, the life cycle of cyanophages is influenced by light, which might be an adaptation to their photosynthetic host cells. The importance of light to cyanophages was reviewed by Clokie and Mann (2006) 10 years ago, and here we review the progress in the past 10 years with a focus on cyanophage infection of Synechococcus and Prochlorococcus under daily light-dark (diel) cycles.
Biological Rhythms of Cyanobacteria
Circadian rhythms are ~24 h cycles of physiological processes found in most living organisms as a fundamental adaptation to the Earth's daily fluctuation in light and temperature (Dunlap, 1999). The endogenous circadian clock systems generate these ~24 h rhythms in constant conditions (continuous light). The freshwater cyanobacterium Synechococcus elongatus PCC 7942 has been used as a model system to study the circadian clock of prokaryotes (Kondo et al., 1994). The cyanobacterial circadian clock regulates many important cellular processes, including cell cycle, amino acid uptake, nitrogen fixation, photosynthesis, carbohydrate synthesis, and respiration (Golden et al., 1997). The molecular mechanism of cyanobacterial circadian clock has been reviewed extensively (Johnson and Egli, 2014; Cohen and Golden, 2015), and here we introduce its major components for reference. The circadian clock of S. elongatus PCC 7942 is composed of three proteins KaiA, KaiB, and KaiC (Ishiura et al., 1998). KaiC is an autokinase and also an autophosphatase. KaiA can stimulate KaiC phosphorylation, while KaiB can promote KaiC dephosphorylation by inactivating KaiA. Interactions of these three Kai proteins generate circadian oscillations in the phosphorylation state of KaiC (reviewed by Dong and Golden, 2008; Johnson et al., 2008; Dong et al., 2010a). The KaiABC circadian clock is synchronized with the light-dark cycle through the intracellular adenosine triphosphate (ATP) (Rust et al., 2011) and oxidized plastoquinone levels (Kim et al., 2012; Figure 1A) that reflect photosynthetic activity. This circadian clock is responsible for genome-wide transcriptional oscillations in S. elongatus (Liu et al., 1995; Ito et al., 2009), with one group of genes peaking at subjective dawn and the other group peaking at subjective dusk (Ito et al., 2009). The response regulator RpaA serves as the master transcriptional regulator of these clock outputs (Markson et al., 2013).
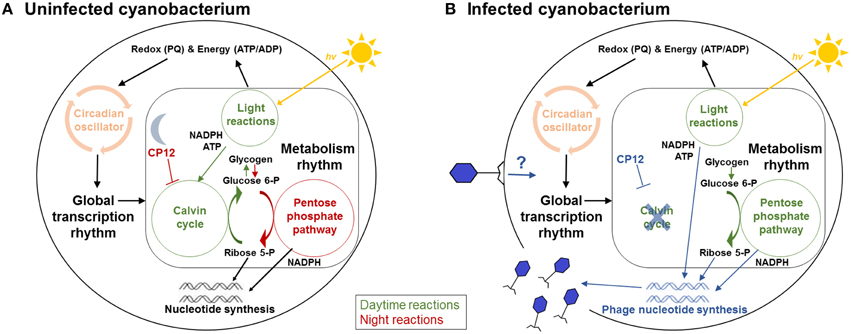
Figure 1. Cyanobacterial metabolism in the uninfected (A) and infected cells (B). (A) In the uninfected host cells, the light reactions of photosynthesis harness energy from the sunlight (hv) to produce NADPH and ATP, which are used for carbon fixation in the Calvin cycle. The Calvin cycle is active during the daytime and is inhibited by CP12 at night. The pentose phosphate pathway is mainly active at night. This metabolism rhythm is under the control of the circadian oscillator, which is affected by the intracellular redox (PQ, plastoquinone) and energy (ATP/ADP) state. The reactions with higher activity during the daytime (daytime reactions) are indicated by green color, and ones with higher activity at night (night reactions) are indicated by red color. (B) In the infected host cells, the pentose phosphate pathway is proposed to be active during the daytime, and the Calvin cycle is proposed to be inhibited by phage-encoded CP12 (Thompson et al., 2011). The energy (ATP) and reducing power (NADPH) from the light reactions flow to phage nucleotide synthesis. Reactions that are potentially affected by cyanophages are indicated by blue color. The effects of phage infection on the host's biological rhythms remain to be explored.
As the most abundant photosynthetic organism in the ocean, the unicellular cyanobacterium Prochlorococcus is responsible for almost 50% of the total primary production in oligotrophic regions (Partensky et al., 1999). Sequenced Prochlorococcus strains all encode kaiB and kaiC, but lack kaiA (Holtzendorff et al., 2008; Axmann et al., 2014), suggesting that their biological timing system is different from that of S. elongatus (Holtzendorff et al., 2008; Johnson and Egli, 2014). Indeed, synchronized S. elongatus cells are able to maintain periodic oscillations of cell cycle (Mori et al., 1996) and gene expression under continuous light (Liu et al., 1995), while Prochlorococcus lacks this ability (Holtzendorff et al., 2008). Therefore, Prochlorococcus maintains a diurnal rhythm rather than a circadian rhythm. Recombinant KaiC protein from Prochlorococcus MED4 is constitutively phosphorylated, and dephosphorylation of KaiC cannot be promoted by KaiB proteins from Prochlorococcus or S. elongatus (Axmann et al., 2009). These data suggest that Prochlorococcus may have lost the autonomous circadian clock, but retain an hourglass-like timing system that needs a daily resetting (Holtzendorff et al., 2008; Axmann et al., 2009).
When Prochlorococcus strain MED4 was grown under light-dark cycles, the expression of around 80% genes showed diel oscillations, with the majority of transcripts peaking at dawn or dusk (Zinser et al., 2009; Waldbauer et al., 2012). Photosystem I and II genes showed maximum expression at dawn, just in time to harvest energy from sunlight (Zinser et al., 2009). The Calvin cycle genes had maximum transcript abundances at dawn, and this was thought to be an anticipatory regulation for the immediate increase of biomass when the light energy was captured (Zinser et al., 2009). On the other hand, the pentose phosphate pathway genes had maximum expression at dusk, which may prepare for glycogen catabolism at night (Zinser et al., 2009). Diel oscillations were also observed for nearly half of all Prochlorococcus population transcripts in the North Pacific Subtropical Gyre (Ottesen et al., 2014), which were similar to those of lab cultures (Zinser et al., 2009; Waldbauer et al., 2012). Moreover, the co-occurring heterotrophic bacterioplankton groups displayed diel periodicity in many transcripts as well, which is probably caused by the tight metabolic coupling between primary producers like Prochlorococcus and consumers living in the open ocean (Ottesen et al., 2014). Since microorganisms constitute more than 90% of the living biomass in the sea (Azam and Malfatti, 2007), biological rhythms of marine cyanobacteria are likely to influence matter and energy transformation in the world's oceans.
Light-dependent and Light-independent Adsorption of Cyanophages onto Their Host Cells
Phage adsorption to a host cell is the first step of infection. During adsorption, phage tails recognize specific receptors on the host cell, such as lipopolysaccharide, membrane proteins, pili, or flagella (Labrie et al., 2010). Cyanophages infecting freshwater (Cséke and Farkas, 1979; Kao et al., 2005) and marine Synechococcus (Jia et al., 2010) have been used to study the effect of light on cyanophage adsorption onto their host cells. Cyanophage AS-1 is a myovirus that infects closely related freshwater S. elongatus strains PCC 6301 and PCC 7942. When S. elongatus PCC 6301 was grown under continuous light, 70% of cyanophage AS-1 were adsorbed within 15 min, and a maximum adsorption of 80% was reached after 20 min (Cséke and Farkas, 1979). When the PCC 6301 culture was moved to the dark after infection, only 40% of cyanophage AS-1 were adsorbed after 20 min (Cséke and Farkas, 1979). The adsorption of cyanophage AS-1 to S. elongatus PCC 7942 was also lower in the dark than that in the light (Kao et al., 2005). Jia et al. studied the adsorption of nine different cyanophages to their host marine Synechococcus WH7803 (Jia et al., 2010). Using unsynchronized Synechococcus WH7803 cultures, they found that the adsorption of myoviruses S-BnM1, S-PWM3, S-PM2, and podovirus S-BP3 was lower in the dark than that in the light (light-dependent adsorption), while the adsorption of myoviruses S-PWM1, S-BM3, S-MM4, S-MM1, and S-MM5 was not affected by darkness (light-independent adsorption) (Jia et al., 2010). Light-dependent adsorption of cyanophage S-PM2 was also seen with synchronized Synechococcus WH7803 cells, and the percentage of adsorbed phage did not change significantly at different times of the day, indicating that the adsorption of cyanophage S-PM2 was not influenced by the circadian rhythm of the host cells (Jia et al., 2010). The authors did not check whether the adsorption of light-independent cyanophages was affected by the circadian rhythm of the host cells. This light-dependent adsorption could be explained by light-induced conformational changes in either the phage tails or the host receptors (Jia et al., 2010), but the exact mechanism is still unexplored.
Effect of Light on Cyanophage Replication in the Host Cells
A strong positive correlation between light intensity and burst size has been shown on cyanophages infecting cyanobacteria Plectonema boryanum (Padan et al., 1970; Sherman and Haselkorn, 1971), Nostoc muscorum (Adolph and Haselkorn, 1972), Synechococcus (Mackenzie and Haselkorn, 1972; Sherman, 1976; Kao et al., 2005), and Prochlorococcus (Lindell et al., 2005; Thompson et al., 2011). Similar to darkness, photosynthesis inhibitors can also significantly reduce the burst sizes of cyanophages (Adolph and Haselkorn, 1972; Mackenzie and Haselkorn, 1972; Sherman, 1976; Lindell et al., 2005). When Prochlorococcus MED4 was infected by the T4-like myovirus P-HM2 in the dark, phage genomic DNA replication inside the host cells stopped, and phage transcript abundances decreased significantly (Thompson et al., 2011). These studies indicate that cyanophage replication derives most of its energy and certain resources from photosynthetic metabolism of the host cell (Sherman, 1976).
The importance of light to cyanophage replication is also revealed by phage-encoded genes for the light reactions of photosynthesis (Mann et al., 2003; Lindell et al., 2004; Millard et al., 2004; Sullivan et al., 2006). Cyanophages carry psbA and psbD genes, encoding the D1 and D2 proteins of the core of photosystem II, respectively (Mann et al., 2003). Phage D1 protein was detected when Prochlorococcus MED4 was infected by cyanophage P-SSP7 (Lindell et al., 2005). Phage psbA was highly transcribed when Synechococcus WH7803 was infected by cyanophage S-PM2 (Clokie et al., 2006a; Millard et al., 2010). The expression of cyanophage-encoded D1 and D2 proteins is proposed to maintain the photosynthetic activity of the infected host cells and therefore provide energy for cyanophage replication (Mann et al., 2003). Many genes related to photosynthesis have also been found in cyanophage genomes (reviewed by Clokie and Mann, 2006; Puxty et al., 2015).
Cyanophage Infection and the Host's Cell Cycle
Studies on phages infecting Escherichia coli showed that the largest burst size was achieved after the completion of host DNA replication and immediately prior to cell division (Starka, 1962; Storms et al., 2014). This is likely due to the increased availability of intracellular resources for the formation of phage progeny (Storms et al., 2014). Under light-dark cycles, S. elongatus PCC 7942 cells divide mainly during the daytime and stop dividing early at night (Mori et al., 1996). The cell division pattern of synchronized S. elongatus PCC 7942 cells persists under continuous light for several days (Mori et al., 1996), and it is controlled by the circadian clock (Mori et al., 1996; Dong et al., 2010b). Using S. elongatus PCC 7942 and cyanophage AS-1, Kao et al. found that the infections started between dawn and noon resulted in larger burst sizes than the ones started in the afternoon or at night (Kao et al., 2005). It is not clear whether the diel infection pattern of cyanophage AS-1 is determined by the light-dark cycle or the cell cycle, or both. Future infection experiments using light-dark synchronized S. elongatus PCC 7942 cells under continuous light will help distinguish the effects of the light-dark cycle and the host cell cycle on burst sizes. Different from S. elongatus PCC 7942, Prochlorococcus cells replicate their DNA in the afternoon and divide at night (Vaulot et al., 1995; Liu et al., 1997; Shalapyonok et al., 1998; Holtzendorff et al., 2008; Zinser et al., 2009). It remains to be studied how the cell cycle of Prochlorococcus affects cyanophage infection.
Potential Interaction between Cyanophage and the Host's Timing System
For mammalian viruses, it has been shown that viral proteins can interact with the host's circadian clock proteins (Kalamvoki and Roizman, 2010), and these interactions are important for viral replication and gene expression (Kalamvoki and Roizman, 2010; Benegiamo et al., 2013). Kao et al. examined whether mutations in the circadian clock of S. elongatus PCC 7942 affected cyanophage AS-1 infection (Kao et al., 2005). They infected the wild-type strain and two mutants (ΔkaiA and ΔkaiB) with cyanophage AS-1 under continuous light and did not see any significant difference in burst sizes (Kao et al., 2005). They did not examine whether the burst size of AS-1 was affected when infecting the ΔkaiA or ΔkaiB mutants under light-dark cycles.
While mutations in the circadian clock proteins of cyanobacteria do not seem to affect cyanophage infection, it is not clear whether cyanophage infection affects the host's circadian clock. In cyanobacteria, the Calvin cycle is linked to the pentose phosphate pathway, with several reactions shared by both pathways (Zinser et al., 2009; Figure 1A). During the daytime, the light reactions of photosynthesis harness energy from the sunlight and split water to produce NADPH (an electron carrier) and ATP (an energy carrier). The Calvin cycle consumes NADPH and ATP to fix CO2 and to produce glucose 6-phosphate, which is stored as the branched glucose polymer glycogen (Figure 1A). At night, the Calvin cycle is inhibited by the Calvin cycle inhibitor CP12 (Wedel and Soll, 1998; Tamoi et al., 2005), and glycogen is mostly oxidized in the pentose phosphate pathway (Pelroy et al., 1972) to produce NADPH and ribose 5-phosphate (Figure 1A), which can be used for carbon skeletons. The rhythm of glycogen metabolism, which is controlled by the circadian clock, can affect the cellular ATP/ADP ratio (Pattanayak et al., 2014). Thus, the circadian clock outputs (glycogen) can feed back through metabolism to control the circadian clock inputs (ATP) (Figure 1A; Pattanayak et al., 2014). Cyanophage genomes contain the pentose phosphate pathway genes (Mann et al., 2005; Sullivan et al., 2005, 2010; Weigele et al., 2007; Clokie et al., 2010) and the Calvin cycle inhibitor gene cp12 (Thompson et al., 2011). Experiments with Prochlorococcus and Synechococcus phages suggested (Thompson et al., 2011) that cyanophages can use these auxiliary metabolic genes (Breitbart et al., 2007) to enhance the host's photosynthesis and pentose phosphate pathway for the production of NADPH, ATP, and ribose 5-phosphate, which can power nucleotide biosynthesis that is critical for phage reproduction (Figure 1B). During the daytime, cyanophage infection may inhibit the Calvin cycle by phage-encoded CP12 protein (Figure 1B). By reversing the host's metabolic rhythm, cyanophage infection might affect the cellular ATP/ADP ratio, which can control the circadian clock of the infected host cells.
Diel Oscillations in Cyanophage Abundance in Aquatic Environments
Several studies suggest that cyanophage infection of cyanobacteria in aquatic environments is synchronized directly or indirectly to the light-dark cycle. In natural aquatic environments, virus decay has been attributed to adsorption to heat-labile particles, inactivation by solar radiation, and grazing by flagellates (Suttle and Chen, 1992), which may not exist in lab cultures. Therefore, cyanophage abundances in aquatic environments are affected by both production and decay. Bettarel et al. (2002) measured the abundances of Synechococcus, heterotrophic bacteria, heterotrophic nanoflagellates, and viruses in September 2001 every 3 h for 3 days at a depth of 1 m in the N.W. Mediterranean Sea. They only found apparent diel patterns for Synechococcus and viruses: Synechococcus was maximal at midnight, and viruses were maximal in the late afternoon and were minimal at midnight (Bettarel et al., 2002). Although they did not measure Synechococcus-specific viruses, they suggested that the diel pattern of viruses was mainly due to cyanophage production through Synechococcus mortality since viral infection of heterotrophic bacteria was rare (Bettarel et al., 2002). Clokie et al. used marine Synechococcus WH7803 to measure cyanophages from seawater samples at different depths (10, 25, and 50 m) in the Indian Ocean (Clokie et al., 2006b). With sampling every 6 h for a 24-h period, cyanophage abundances were similar for most sampling time-points at 10 m, but showed a distinct peak shortly after midnight (Clokie et al., 2006b). Different from the 10 m sample, cyanophage abundances at 25 and 50 m did not show strong diel variations (Clokie et al., 2006b). Using real-time PCR and strain-specific primers, Kimura et al. measured Microcystis and its cyanophage abundances from surface waters of a freshwater pond in Japan (Kimura et al., 2012). They sampled every 3 h over a period of 24 h in two different days (Kimura et al., 2012). For both days, Microcystis abundances were highest after midnight, which may be caused by diel vertical migration of Microcystis (Kimura et al., 2012). The highest Microcystis cyanophage abundances were found at dusk in one day and about 3 h after dusk in the other day (Kimura et al., 2012). Recently, the growth and mortality of Prochlorococcus in the subtropical Pacific gyre was found to be synchronized to the light-dark cycle, with the highest mortality rate at night (Ribalet et al., 2015). This could be explained by light-dark synchronized infection of Prochlorococcus by cyanophages, although synchronized grazing by protists is also possible (Ribalet et al., 2015).
Concluding Remarks
Cyanobacteria exhibit biological rhythms as an adaptation to the natural daily light-dark cycle, with many biological processes connected by the circadian clock into a cellular network (Shultzaberger et al., 2015). Light is also crucial for cyanophage infection of cyanobacteria. It remains to be explored how cyanophage infection affects the biological rhythms of cyanobacteria. However, most cyanophage infection studies have been conducted under continuous light, and the host cells were not synchronized. Future infection experiments with light-dark synchronized cyanobacteria are needed to reveal the interaction between cyanophage infection and the host biological rhythms. Additionally, more field studies should be carried out to understand the diel infection patterns of cyanobacteria by cyanophages in different aquatic environments. Having coevolved with cyanobacteria under light-dark cycles for millions of years, cyanophages will give us new insights into the biological rhythms of their host cells. With these data, we can begin to understand how the daily light-dark cycle shapes the interaction of cyanophages and cyanobacteria, which eventually influences matter and energy transformation in aquatic environments.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Juliene Co, Xiaoting Fang, and Xingqin Lin for valuable discussions. This publication is supported by grants to QZ from the Research Grants Council of the Hong Kong Special Administrative Region, China (Project No. 16103414), the National Basic Research Program of China (973 Program) (Project No. 2013CB955700), and the National Natural Science Foundation of China (Project No. 41476147).
References
Adolph, K. W., and Haselkorn, R. (1972). Photosynthesis and the development of blue-green algal virus N-1. Virology 47, 370–374. doi: 10.1016/0042-6822(72)90272-3
Axmann, I. M., Dühring, U., Seeliger, L., Arnold, A., Vanselow, J. T., Kramer, A., et al. (2009). Biochemical evidence for a timing mechanism in Prochlorococcus. J. Bacteriol. 191, 5342–5347. doi: 10.1128/JB.00419-09
Axmann, I. M., Hertel, S., Wiegard, A., Dörrich, A. K., and Wilde, A. (2014). Diversity of KaiC-based timing systems in marine cyanobacteria. Mar. Genomics 14, 3–16. doi: 10.1016/j.margen.2013.12.006
Azam, F., and Malfatti, F. (2007). Microbial structuring of marine ecosystems. Nat. Rev. Microbiol. 5, 782–791. doi: 10.1038/nrmicro1747
Benegiamo, G., Mazzoccoli, G., Cappello, F., Rappa, F., Scibetta, N., Oben, J., et al. (2013). Mutual antagonism between circadian protein period 2 and hepatitis C virus replication in hepatocytes. PLoS ONE 8:e60527. doi: 10.1371/journal.pone.0060527
Bergh, O., Børsheim, K. Y., Bratbak, G., and Heldal, M. (1989). High abundance of viruses found in aquatic environments. Nature 340, 467–468. doi: 10.1038/340467a0
Bettarel, Y., Dolan, J. R., Hornak, K., Lemee, R., Masin, M., Pedrotti, M. L., et al. (2002). Strong, weak, and missing links in a microbial community of the N.W. Mediterranean Sea. FEMS Microbiol. Ecol. 42, 451–462. doi: 10.1111/j.1574-6941.2002.tb01034.x
Breitbart, M., Thompson, L. R., Suttle, C. A., and Sullivan, M. B. (2007). Exploring the vast diversity of marine viruses. Oceanography 20, 135–139. doi: 10.5670/oceanog.2007.58
Clokie, M. R. J., Millard, A. D., Mehta, J. Y., and Mann, N. H. (2006b). Virus isolation studies suggest short-term variations in abundance in natural cyanophage populations of the Indian Ocean. J. Mar. Biol. Assoc. U.K. 86, 499–505. doi: 10.1017/S0025315406013403
Clokie, M. R., and Mann, N. H. (2006). Marine cyanophages and light. Environ. Microbiol. 8, 2074–2082. doi: 10.1111/j.1462-2920.2006.01171.x
Clokie, M. R., Millard, A. D., and Mann, N. H. (2010). T4 genes in the marine ecosystem: studies of the T4-like cyanophages and their role in marine ecology. Virol. J. 7:291. doi: 10.1186/1743-422x-7-291
Clokie, M. R., Shan, J., Bailey, S., Jia, Y., Krisch, H. M., West, S., et al. (2006a). Transcription of a ‘photosynthetic’ T4-type phage during infection of a marine cyanobacterium. Environ. Microbiol. 8, 827–835. doi: 10.1111/j.1462-2920.2005.00969.x
Cohen, S. E., and Golden, S. S. (2015). Circadian rhythms in cyanobacteria. Microbiol. Mol. Biol. Rev. 79, 373–385. doi: 10.1128/MMBR.00036-15
Cséke, C. S., and Farkas, G. L. (1979). Effect of light on the attachment of cyanophage AS-1 to Anacystis nidulans. J. Bacteriol. 137, 667–669.
Dong, G., and Golden, S. S. (2008). How a cyanobacterium tells time. Curr. Opin. Microbiol. 11, 541–546. doi: 10.1016/j.mib.2008.10.003
Dong, G., Kim, Y. I., and Golden, S. S. (2010a). Simplicity and complexity in the cyanobacterial circadian clock mechanism. Curr. Opin. Genet. Dev. 20, 619–625. doi: 10.1016/j.gde.2010.09.002
Dong, G., Yang, Q., Wang, Q., Kim, Y. I., Wood, T. L., Osteryoung, K. W., et al. (2010b). Elevated ATPase activity of KaiC applies a circadian checkpoint on cell division in Synechococcus elongatus. Cell 140, 529–539. doi: 10.1016/j.cell.2009.12.042
Dunlap, J. C. (1999). Molecular bases for circadian clocks. Cell 96, 271–290. doi: 10.1016/S0092-8674(00)80566-8
Golden, S. S., Ishiura, M., Johnson, C. H., and Kondo, T. (1997). Cyanobacterial circadian rhythms. Annu. Rev. Plant Physiol. Plant Mol. Biol. 48, 327–354. doi: 10.1146/annurev.arplant.48.1.327
Holtzendorff, J., Partensky, F., Mella, D., Lennon, J. F., Hess, W. R., and Garczarek, L. (2008). Genome streamlining results in loss of robustness of the circadian clock in the marine cyanobacterium Prochlorococcus marinus PCC 9511. J. Biol. Rhythms 23, 187–199. doi: 10.1177/0748730408316040
Ishiura, M., Kutsuna, S., Aoki, S., Iwasaki, H., Andersson, C. R., Tanabe, A., et al. (1998). Expression of a gene cluster kaiABC as a circadian feedback process in cyanobacteria. Science 281, 1519–1523. doi: 10.1126/science.281.5382.1519
Ito, H., Mutsuda, M., Murayama, Y., Tomita, J., Hosokawa, N., Terauchi, K., et al. (2009). Cyanobacterial daily life with Kai-based circadian and diurnal genome-wide transcriptional control in Synechococcus elongatus. Proc. Natl. Acad. Sci. U.S.A. 106, 14168–14173. doi: 10.1073/pnas.0902587106
Jia, Y., Shan, J., Millard, A., Clokie, M. R., and Mann, N. H. (2010). Light-dependent adsorption of photosynthetic cyanophages to Synechococcus sp. WH7803. FEMS Microbiol. Lett. 310, 120–126. doi: 10.1111/j.1574-6968.2010.02054.x
Johnson, C. H., and Egli, M. (2014). Metabolic compensation and circadian resilience in prokaryotic cyanobacteria. Annu. Rev. Biochem. 83, 221–247. doi: 10.1146/annurev-biochem-060713-035632
Johnson, C. H., Egli, M., and Stewart, P. L. (2008). Structural insights into a circadian oscillator. Science 322, 697–701. doi: 10.1126/science.1150451
Kalamvoki, M., and Roizman, B. (2010). Circadian CLOCK histone acetyl transferase localizes at ND10 nuclear bodies and enables herpes simplex virus gene expression. Proc. Natl. Acad. Sci. U.S.A. 107, 17721–17726. doi: 10.1073/pnas.1012991107
Kao, C. C., Green, S., Stein, B., and Golden, S. S. (2005). Diel infection of a cyanobacterium by a contractile bacteriophage. Appl. Environ. Microbiol. 71, 4276–4279. doi: 10.1128/AEM.71.8.4276-4279.2005
Kim, Y. I., Vinyard, D. J., Ananyev, G. M., Dismukes, G. C., and Golden, S. S. (2012). Oxidized quinones signal onset of darkness directly to the cyanobacterial circadian oscillator. Proc. Natl. Acad. Sci. U.S.A. 109, 17765–17769. doi: 10.1073/pnas.1216401109
Kimura, S., Yoshida, T., Hosoda, N., Honda, T., Kuno, S., Kamiji, R., et al. (2012). Diurnal infection patterns and impact of Microcystis cyanophages in a Japanese pond. Appl. Environ. Microbiol. 78, 5805–5811. doi: 10.1128/AEM.00571-12
Kondo, T., Tsinoremas, N. F., Golden, S. S., Johnson, C. H., Kutsuna, S., and Ishiura, M. (1994). Circadian clock mutants of cyanobacteria. Science 266, 1233–1236. doi: 10.1126/science.7973706
Labrie, S. J., Samson, J. E., and Moineau, S. (2010). Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 8, 317–327. doi: 10.1038/nrmicro2315
Lindell, D., Jaffe, J. D., Johnson, Z. I., Church, G. M., and Chisholm, S. W. (2005). Photosynthesis genes in marine viruses yield proteins during host infection. Nature 438, 86–89. doi: 10.1038/nature04111
Lindell, D., Sullivan, M. B., Johnson, Z. I., Tolonen, A. C., Rohwer, F., and Chisholm, S. W. (2004). Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc. Natl. Acad. Sci. U.S.A. 101, 11013–11018. doi: 10.1073/pnas.0401526101
Liu, H., Nolla, H. A., and Campbell, L. (1997). Prochlorococcus growth rate and contribution to primary production in the equatorial and subtropical North Pacific Ocean. Aquat. Microbial Ecol. 12, 39–47. doi: 10.3354/ame012039
Liu, Y., Tsinoremas, N. F., Johnson, C. H., Lebedeva, N. V., Golden, S. S., Ishiura, M., et al. (1995). Circadian orchestration of gene expression in cyanobacteria. Genes Dev. 9, 1469–1478. doi: 10.1101/gad.9.12.1469
Mackenzie, J. J., and Haselkorn, R. (1972). Photosynthesis and the development of blue-green algal virus SM-1. Virology 49, 517–521. doi: 10.1016/0042-6822(72)90503-X
Mann, N. H., Clokie, M. R., Millard, A., Cook, A., Wilson, W. H., Wheatley, P. J., et al. (2005). The genome of S-PM2, a “photosynthetic” T4-type bacteriophage that infects marine Synechococcus strains. J. Bacteriol. 187, 3188–3200. doi: 10.1128/JB.187.9.3188-3200.2005
Mann, N. H., Cook, A., Millard, A., Bailey, S., and Clokie, M. (2003). Bacterial photosynthesis genes in a virus. Nature 424, 741. doi: 10.1038/424741a
Markson, J. S., Piechura, J. R., Puszynska, A. M., and O'shea, E. K. (2013). Circadian control of global gene expression by the cyanobacterial master regulator RpaA. Cell 155, 1396–1408. doi: 10.1016/j.cell.2013.11.005
Millard, A., Clokie, M. R., Shub, D. A., and Mann, N. H. (2004). Genetic organization of the psbAD region in phages infecting marine Synechococcus strains. Proc. Natl. Acad. Sci. U.S.A. 101, 11007–11012. doi: 10.1073/pnas.0401478101
Millard, A. D., Gierga, G., Clokie, M. R., Evans, D. J., Hess, W. R., and Scanlan, D. J. (2010). An antisense RNA in a lytic cyanophage links psbA to a gene encoding a homing endonuclease. ISME J. 4, 1121–1135. doi: 10.1038/ismej.2010.43
Mori, T., Binder, B., and Johnson, C. H. (1996). Circadian gating of cell division in cyanobacteria growing with average doubling times of less than 24 hours. Proc. Natl. Acad. Sci. U.S.A. 93, 10183–10188. doi: 10.1073/pnas.93.19.10183
Ottesen, E. A., Young, C. R., Gifford, S. M., Eppley, J. M., Marin, R. III, Schuster, S. C., et al. (2014). Multispecies diel transcriptional oscillations in open ocean heterotrophic bacterial assemblages. Science 345, 207–212. doi: 10.1126/science.1252476
Padan, E., Ginzburg, D., and Shilo, M. (1970). The reproductive cycle of cyanophage LPP1-G in Plectonema boryanum and its dependence on photosynthetic and respiratory systems. Virology 40, 514–521. doi: 10.1016/0042-6822(70)90194-7
Partensky, F., Hess, W. R., and Vaulot, D. (1999). Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol. Mol. Biol. Rev. 63, 106–127.
Pattanayak, G. K., Phong, C., and Rust, M. J. (2014). Rhythms in energy storage control the ability of the cyanobacterial circadian clock to reset. Curr. Biol. 24, 1934–1938. doi: 10.1016/j.cub.2014.07.022
Pelroy, R. A., Rippka, R., and Stanier, R. Y. (1972). Metabolism of glucose by unicellular blue-green algae. Arch. Microbiol. 87, 303–322. doi: 10.1007/bf00409131
Proctor, L. M., and Fuhrman, J. A. (1990). Viral mortality of marine bacteria and cyanobacteria. Nature 343, 60–62. doi: 10.1038/343060a0
Puxty, R. J., Millard, A. D., Evans, D. J., and Scanlan, D. J. (2015). Shedding new light on viral photosynthesis. Photosyn. Res. 126, 71–97. doi: 10.1007/s11120-014-0057-x
Ribalet, F., Swalwell, J., Clayton, S., Jiménez, V., Sudek, S., Lin, Y., et al. (2015). Light-driven synchrony of Prochlorococcus growth and mortality in the subtropical Pacific gyre. Proc. Natl. Acad. Sci. U.S.A. 112, 8008–8012. doi: 10.1073/pnas.1424279112
Rust, M. J., Golden, S. S., and O'shea, E. K. (2011). Light-driven changes in energy metabolism directly entrain the cyanobacterial circadian oscillator. Science 331, 220–223. doi: 10.1126/science.1197243
Scanlan, D. J., and West, N. J. (2002). Molecular ecology of the marine cyanobacterial genera Prochlorococcus and Synechococcus. FEMS Microbiol. Ecol. 40, 1–12. doi: 10.1111/j.1574-6941.2002.tb00930.x
Shalapyonok, A., Olson, R. J., and Shalapyonok, L. S. (1998). Ultradian growth in Prochlorococcus spp. Appl. Environ. Microbiol. 64, 1066–1069.
Sherman, L. A. (1976). Infection of Synechococcus cedrorum by the cyanophage AS-1M. III. Cellular metabolism and phage development. Virology 71, 199–206. doi: 10.1016/0042-6822(76)90105-7
Sherman, L. A., and Haselkorn, R. (1971). Growth of the blue-green algae virus LPP-1 under conditions which impair photosynthesis. Virology 45, 739–746. doi: 10.1016/0042-6822(71)90188-7
Shultzaberger, R. K., Boyd, J. S., Diamond, S., Greenspan, R. J., and Golden, S. S. (2015). Giving time purpose: the Synechococcus elongatus clock in a broader network context. Annu. Rev. Genet. 49, 485–505. doi: 10.1146/annurev-genet-111212-133227
Starka, J. (1962). Cellular division and reproduction of bacteriophage in synchronized cultures of Escherichia coli. J. Gen. Microbiol. 29, 83–90. doi: 10.1099/00221287-29-1-83
Storms, Z. J., Brown, T., Cooper, D. G., Sauvageau, D., and Leask, R. L. (2014). Impact of the cell life-cycle on bacteriophage T4 infection. FEMS Microbiol. Lett. 353, 63–68. doi: 10.1111/1574-6968.12402
Sullivan, M. B., Coleman, M. L., Weigele, P., Rohwer, F., and Chisholm, S. W. (2005). Three Prochlorococcus cyanophage genomes: signature features and ecological interpretations. PLoS Biol. 3:e144. doi: 10.1371/journal.pbio.0030144
Sullivan, M. B., Huang, K. H., Ignacio-Espinoza, J. C., Berlin, A. M., Kelly, L., Weigele, P. R., et al. (2010). Genomic analysis of oceanic cyanobacterial myoviruses compared with T4-like myoviruses from diverse hosts and environments. Environ. Microbiol. 12, 3035–3056. doi: 10.1111/j.1462-2920.2010.02280.x
Sullivan, M. B., Lindell, D., Lee, J. A., Thompson, L. R., Bielawski, J. P., and Chisholm, S. W. (2006). Prevalence and evolution of core photosystem II genes in marine cyanobacterial viruses and their hosts. PLoS Biol. 4:e234. doi: 10.1371/journal.pbio.0040234
Sullivan, M. B., Waterbury, J. B., and Chisholm, S. W. (2003). Cyanophages infecting the oceanic cyanobacterium Prochlorococcus. Nature 424, 1047–1051. doi: 10.1038/nature01929
Suttle, C. A., and Chan, A. M. (1993). Marine cyanophages infecting oceanic and coastal strains of Synechococcus: abundance, morphology, cross-infectivity and growth characteristics. Mar. Ecol. Prog. Ser. 92, 99–109. doi: 10.3354/meps092099
Suttle, C. A., Chan, A. M., and Cottrell, M. T. (1990). Infection of phytoplankton by viruses and reduction of primary productivity. Nature 347, 467–469. doi: 10.1038/347467a0
Suttle, C. A., and Chen, F. (1992). Mechanisms and rates of decay of marine viruses in seawater. Appl. Environ. Microbiol. 58, 3721–3729.
Tamoi, M., Miyazaki, T., Fukamizo, T., and Shigeoka, S. (2005). The Calvin cycle in cyanobacteria is regulated by CP12 via the NAD(H)/NADP(H) ratio under light/dark conditions. Plant J. 42, 504–513. doi: 10.1111/j.1365-313X.2005.02391.x
Thompson, L. R., Zeng, Q., Kelly, L., Huang, K. H., Singer, A. U., Stubbe, J., et al. (2011). Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc. Natl. Acad. Sci. U.S.A. 108, E757–E764. doi: 10.1073/pnas.1102164108
Vaulot, D., Marie, D., Olson, R. J., and Chisholm, S. W. (1995). Growth of Prochlorococcus, a photosynthetic prokaryote, in the equatorial pacific ocean. Science 268, 1480–1482. doi: 10.1126/science.268.5216.1480
Waldbauer, J. R., Rodrigue, S., Coleman, M. L., and Chisholm, S. W. (2012). Transcriptome and proteome dynamics of a light-dark synchronized bacterial cell cycle. PLoS ONE 7:e43432. doi: 10.1371/journal.pone.0043432
Waterbury, J. B., and Valois, F. W. (1993). Resistance to co-occurring phages enables marine Synechococcus communities to coexist with cyanophages abundant in seawater. Appl. Environ. Microbiol. 59, 3393–3399.
Wedel, N., and Soll, J. (1998). Evolutionary conserved light regulation of Calvin cycle activity by NADPH-mediated reversible phosphoribulokinase/CP12/ glyceraldehyde-3-phosphate dehydrogenase complex dissociation. Proc. Natl. Acad. Sci. U.S.A. 95, 9699–9704. doi: 10.1073/pnas.95.16.9699
Weigele, P. R., Pope, W. H., Pedulla, M. L., Houtz, J. M., Smith, A. L., Conway, J. F., et al. (2007). Genomic and structural analysis of Syn9, a cyanophage infecting marine Prochlorococcus and Synechococcus. Environ. Microbiol. 9, 1675–1695. doi: 10.1111/j.1462-2920.2007.01285.x
Wilson, W. H., Joint, I. R., Carr, N. G., and Mann, N. H. (1993). Isolation and molecular characterization of five marine cyanophages propagated on Synechococcus sp. strain WH7803. Appl. Environ. Microbiol. 59, 3736–3743.
Keywords: cyanobacterium, Prochlorococcus, cyanophage, circadian rhythm, circadian clock
Citation: Ni T and Zeng Q (2016) Diel Infection of Cyanobacteria by Cyanophages. Front. Mar. Sci. 2:123. doi: 10.3389/fmars.2015.00123
Received: 14 June 2015; Accepted: 22 December 2015;
Published: 11 January 2016.
Edited by:
Sandie M. Degnan, The University of Queensland, AustraliaCopyright © 2016 Ni and Zeng. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Qinglu Zeng, emVuZ0B1c3QuaGs=
 Tianchi Ni
Tianchi Ni Qinglu Zeng
Qinglu Zeng