- 1Biogeochemistry Program, Research Institute for Marine Resources Utilization, Japan Agency for Marine-Earth Science and Technology, Yokosuka, Japan
- 2Geological Institute, Department of Earth Sciences, ETH Zürich, Zurich, Switzerland
Tracing of biogeochemical pathways using molecular approaches has advanced our basic understanding of the carbon cycle and life’s legacy in the sedimentary record. To this end, compound-specific radiocarbon analysis has been instrumental in shedding light on the turnover, age, and sources of a range of biomarkers embedded within complex environmental matrices. However, despite their foundational importance for life and their omnipresence throughout geologic space and time, the biogeochemical cycling of amino acids remains largely unexplored. Here, we discuss the potential of using amino acid-specific radiocarbon to deepen our knowledge of the biogeochemistry of food webs and sedimentary organic carbon.
Introduction
The “building blocks of life” constitute a common foundation which all life on Earth shares and depends on (Miller and Urey, 1959; Kitadai and Maruyama, 2018) and amino acids are one of the largest pools of characterizable organic matter found in sedimentary environments (Trask, 1936; Degens, 1970; Lee et al., 1983; Wakeham et al., 1997; Hedges et al., 2001). In contrast to other biomarkers where source specificity is a key attribute for understanding sedimentary and biogeochemical cycling pathways [e.g., lignin from the terrestrial biosphere (Hedges and Parker, 1976), lipids from the marine biosphere (Volkman et al., 1980)], amino acids are arguably among the least specific “biomarkers” as the molecules themselves are omnipresent in all life forms and occur across most of geologic space and time (Abelson, 1954; Erdman et al., 1956; Degens and Bajor, 1960; Hare, 1969). Embracing this property of environmental omnipresence, previous workers, as summarized by Kvenvolden (1975), have provided insights on the utility of amino acids (and their degradation products) as molecular clocks, (paleo)thermometers, and diagenesis proxies. Since then, appreciable progress on compound-specific isotope measurements has been made. Here, we share our perspectives on how amino acid-specific radiocarbon content can theoretically provide insights into a variety of biogeochemical studies.
Biogeochemistry of Amino Acids in Food Webs
Although amino acids are part of the living fabric of all organisms, not all amino acids can be synthesized by all organisms leading to facultative or mandatory dietary acquisition of certain amino acids. This circumstance leads to different pathways of carbon and nitrogen propagating through the food web up to higher trophic positions. Δ14C is successfully used to trace carbon pathways in bulk biomass (Williams et al., 1987; Ishikawa et al., 2013), a point we will get back to later. Analytical developments first opened stable carbon and nitrogen isotopic compositions of amino acids (δ13C and δ15N, respectively) to exploration and our current state of knowledge rests on what these isotopes have revealed. The isotopic composition of amino acids within organisms reflect the isotopic composition of carbon and nitrogen sources in the food web with superimposed effects of metabolic processes (Hare et al., 1991; Carstens et al., 2013). “Trophic” amino acids fractionate heavily against 15N due to preferential deamination or transamination during metabolism, enriching the 15N in the remaining trophic amino acids (Macko et al., 1986; Goto et al., 2018). In contrast, “source” amino acids show no or a lesser extent of 15N fractionation, because their amino group is preserved during metabolism (Chikaraishi et al., 2009). This allows amino acid nitrogen isotopic compositions to be used as excellent proxies for determination of trophic position of organisms in recent and paleo studies (Ohkouchi et al., 2017) and provides useful information for identifying ecological niches of organisms in food webs (Ishikawa, 2018). The “trophic” and “source” categories are controlled by catabolism of amino acids, such as deamination (Chikaraishi et al., 2015), whereas the “essential” and “non-essential” categories are controlled by anabolism such as biosynthesis (McMahon et al., 2010). Metabolic energy is produced in the former and consumed in the latter (Bender, 2012). In autotrophs, the δ13C profile of “essential” amino acids are influenced by biosynthetic processes specific to phylogeny because diverse precursors, intermediates, and reactions produce a different pattern of 13C fractionation among amino acids (Larsen et al., 2009). Using normalized δ13C of several essential amino acids, Larsen et al. (2009) found that plants, fungi, and bacteria can be discriminated from each other in multivariate space. Higher organisms such as vertebrates are unable to biosynthesize essential amino acids, which are exclusively derived from their diets. This is reflected in the findings of McMahon et al. (2010) which show that δ13C of essential amino acids in fish are consistent with those of their diet, whereas δ13C of non-essential amino acids deviate, suggesting that non-essential amino acids are at least partly biosynthesized by higher organisms using different compound and carbon skeleton precursors, which can be exploited to trace dietary sources at higher trophic positions such as for zooplankton and fish (Larsen et al., 2013; McMahon et al., 2016). Therefore, the stable carbon isotopic compositions of amino acids, especially of essential amino acids, are used as a fingerprint of primary producers. Stable carbon and nitrogen isotopic investigations of amino acids in organisms have currently led literature to two contrasting dichotomies, i.e., “trophic” vs. “source” and “essential” vs. “non-essential” (Figure 1). While there are wide overlaps between the source and essential categories and for trophic and non-essential categories, a few amino acids behave out of line. Additionally, the carbon pools from which the carbon atoms of the amino acids stem remain loosely constrained. Catalyzed by recent advances in accelerator mass spectrometry enabling the routine analysis of small-scale radiocarbon measurements (McIntyre et al., 2017), methods have been developed to investigate amino acid-specific radiocarbon in soft tissue of organisms opening a new frontier in research possibilities (Ishikawa et al., 2018).
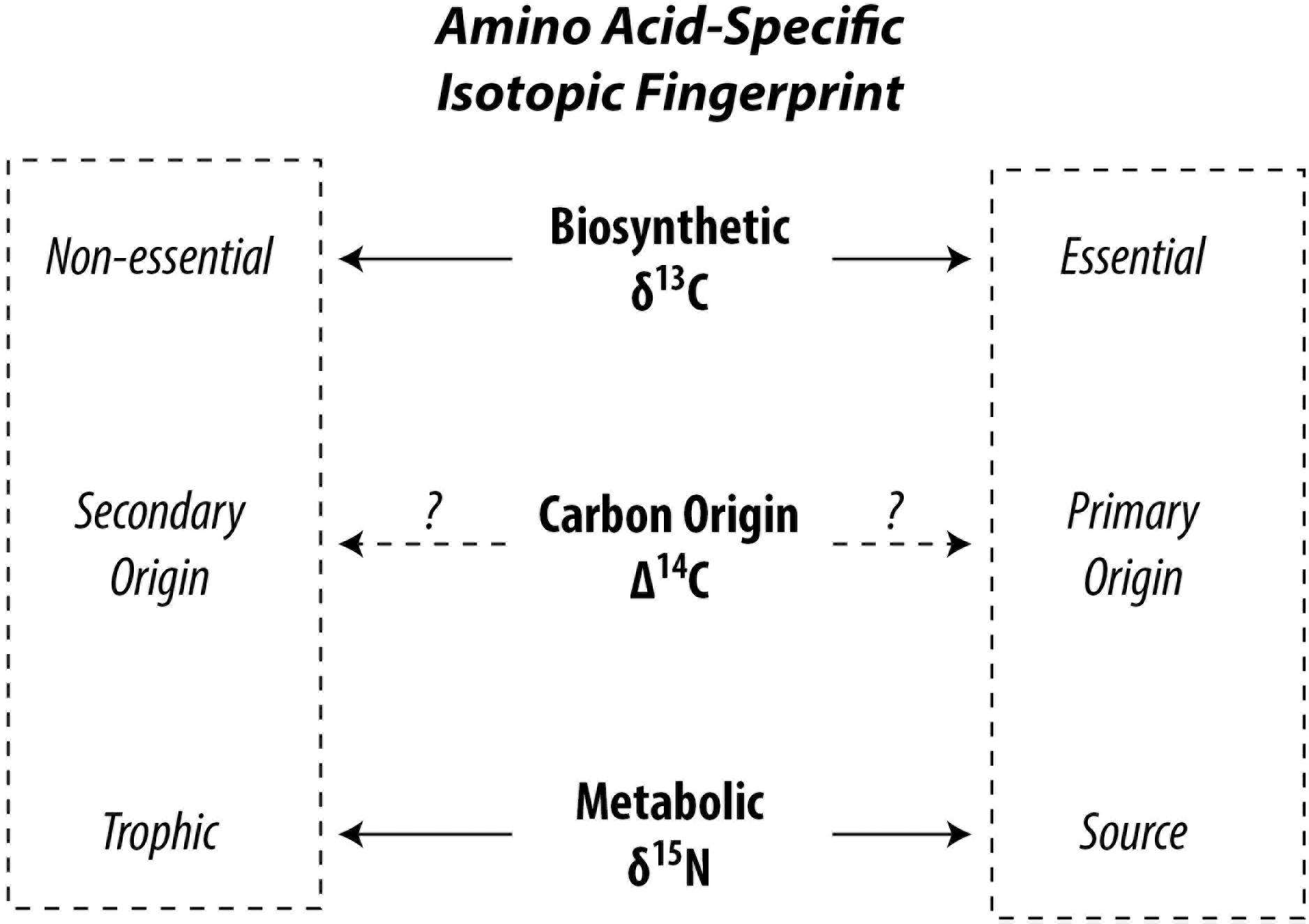
Figure 1. Conceptual division for categorizing amino acids in the context of food web studies. The Δ14C signature of amino acids has potential to integrate δ13C and δ15N.
In this context, natural abundance Δ14C appears a promising tool to help disentangle complex amino acid metabolic pathways and offer new leads in resolving the differences between these competing dichotomies (Figure 1). Unlike δ15N and δ13C, isotopic fractionation for Δ14C is internally canceled out by definition (Stuiver and Polach, 1977). Therefore, metabolic effects should be canceled out with Δ14C and its value should reflect that of their carbon origin(s), such as dissolved inorganic carbon (Broecker and Walton, 1959; Guillemette et al., 2017) or organic materials (Petsch et al., 2001). Most amino acids in the biosphere can be expected to bear a “modern” signature (Wang et al., 1996, 1998; Hwang et al., 2005). However, carbon originating from the atmosphere, dissolved inorganic and organic carbon, surface sediments, methane, and kerogen (often) show contrasting radiocarbon fingerprints. If an ecosystem intersects two or more of these domains, the radiocarbon gradient should be exploitable. To date, efforts have focused on radiocarbon dating of amino acids (particularly hydroxyproline) found in bone collagen for archeological applications (McCullagh et al., 2010). For the “building blocks of life,” there is nearly no data on amino acid radiocarbon in flora and fauna and there is clearly incentive to shed light on this area of basic research (c.f. Bour et al., 2016).
Biogeochemistry of Sedimentary Amino Acids
The importance of organic matter-mineral interactions in soils began to emerge in the mid-20th Century (Beutelspacher, 1955) and their importance for the stabilization of organic matter in marine sediments became recognized in the 1990s (Keil et al., 1994b; Mayer, 1994; Kennedy et al., 2002). These processes influence contrasting degrees of loss-and-replacement of terrestrial with marine organic matter along land-ocean transitions (Keil et al., 1997; Blattmann et al., 2018, 2019).
Nitrogenous compounds are classically considered biogeochemically labile and therefore are efficiently remineralized (Vernadsky, 1930). In the case of amino acids, their sustained existence is unlikely due to their intrinsic lability (Kirchman, 1990; Pantoja and Lee, 2003) leaving stabilization by mineral surfaces as the likeliest explanation (Keil et al., 1994a; Hedges et al., 2001). The modes of interactions between organic matter and minerals include ligand exchange, ion exchange, cation bridging, van der Waals forces, hydrogen bonding, hydrophobic interactions, and combinations thereof (Beutelspacher, 1955; Keil and Mayer, 2014). The stabilization of amino acids and their polymers have been hypothesized and identified as important in the formation of recalcitrant organic nitrogen (e.g. Müller, 1977; Henrichs and Sugai, 1993; Keil et al., 1994a; Hedges et al., 2001; Estes et al., 2019). Different amino acids and structurally related compounds exhibit very different sorption affinities for phyllosilicate mineral surfaces (Hedges, 1978; Hedges and Hare, 1987). Sorption experiments have demonstrated the strong participation of basic amino acids by way of electrostatic interactions in forming phyllosilicate-amino acid/protein complexes (Ensminger and Gieseking, 1941; Hedges, 1978; Hedges and Hare, 1987; Cowie and Hedges, 1992). This effect appears mirrored in nature with the high abundance of basic amino acids found in continental margin sediments (Keil et al., 1998). Amino acids and proteins associated with phyllosilicates and natural sediments have revealed sometimes erratic and irreversible desorption behavior (Henrichs and Sugai, 1993; Wang and Lee, 1993; Montluçon and Lee, 2001; Ding and Henrichs, 2002), which may ultimately influence their bioavailability (Pinck and Allison, 1951). Amino acids are likely more representative of bulk sedimentary organic matter than most other compounds, yet their ubiquity limits their value in quantitatively assessing contributions from sedimentary organic matter sources. On a molecular level, sedimentary amino acids are subject to differential microbial processing and loss-and-replacement of terrestrial with marine amino acids as revealed by stable carbon isotopes (Keil and Fogel, 2001) and therefore source apportionments recorded by individual amino acids are overlain by a variety of effects.
Amino acids stabilized by mineral surfaces are expected to exhibit very slow turnover rates and or would resist loss-and-replacement during land-ocean transit, giving rise to depleted radiocarbon fingerprints borne by mineral-stabilized amino acids. It is thus hypothesized, that radiocarbon isotopic compositions of individual amino acids can provide a biomarker tool to measure the effect of organic matter-mineral interactions in natural sedimentary environments (Figure 2; see also ideas by Keil et al., 1998). One of the greatest uncertainties in studying the stabilization effects of minerals on organic matter in nature is the longstanding and ongoing discussion whether organic matter is stabilized by minerals or if organic matter is intrinsically stable due to its structural characteristics or by other factors (compare Hedges et al., 2001; Eusterhues et al., 2005; Schmidt et al., 2011; Keil and Mayer, 2014; Kleber et al., 2015). To this end, amino acid-specific radiocarbon can offer new perspectives in this ongoing debate. Method developments have also opened up this frontier of amino acid-specific radiocarbon in sediments to exploration (Blattmann et al., 2020).
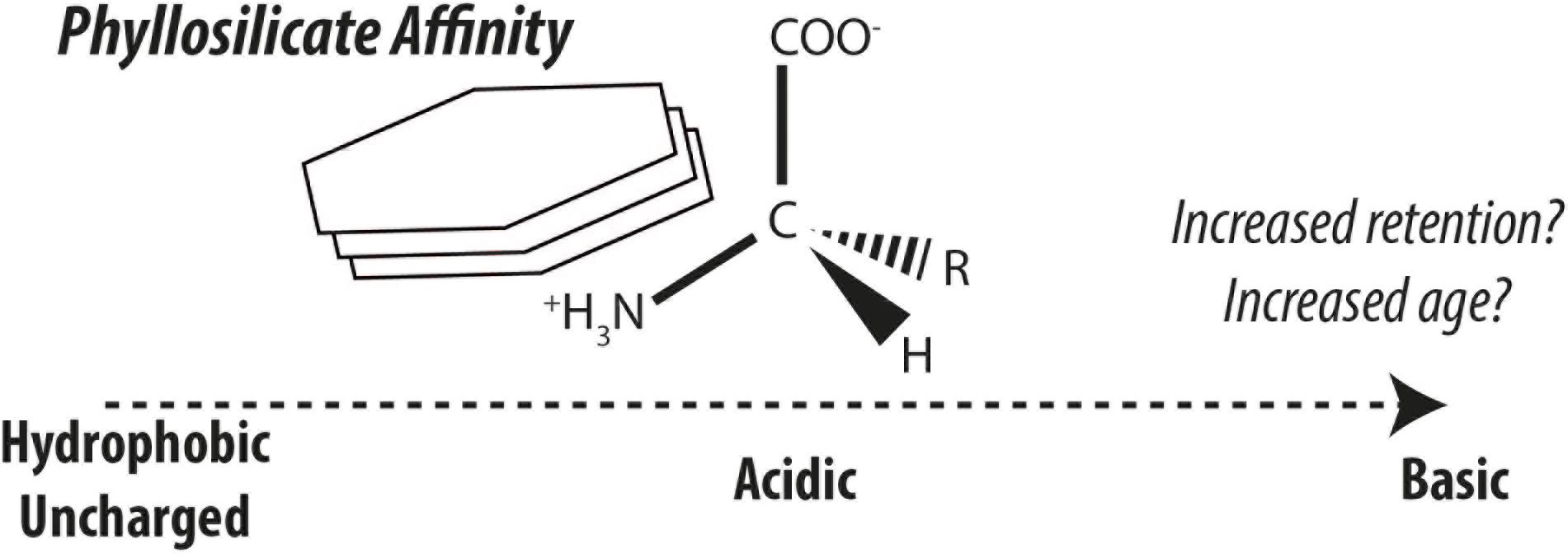
Figure 2. Generalized affinity of amino acids for kaolinite simplified after the findings of Hedges and Hare (1987). Increased affinity is hypothesized to enhance retention and protection of different amino acid classes, thereby leading to greater discrepancies in their radiocarbon ages.
Is Enantiomer-Specific Radiocarbon the Next Frontier?
With methods for amino acid-specific radiocarbon now developed for biological (Ishikawa et al., 2018) and earthen (Blattmann et al., 2020) samples, we share our thoughts on the merits of what we see as a promising next analytical frontier. Radiocarbon isotopic composition of individual compounds reflects a combination of influences stemming from aging, mixing, reservoir effects, etc. By isolating the effect of radioactive decay, constraints on the time since synthesis are attainable (Libby et al., 1949). In addition to this radioactive clock common to all organic compounds, amino acids distinguish themselves by having enantiomers (i.e., mirror image molecular structure like our left and right hands). In the biosphere, the L-form dominates, with certain organisms containing D-amino acids including bacteria (Corrigan, 1969; Lam et al., 2009), plants (Neuberger, 1948), freshwater and marine algae (Yokoyama et al., 2003), and higher organisms including insects and mammals (Neuberger, 1948; Corrigan, 1969). In a process called racemization, L-amino acids switch into D-configuration and vice versa. Racemization is a time and temperature dependent process and is also influenced by humidity, pH, and a host of matrix effects (see review by Bada, 1985). Over time, increasing amounts of D-amino acids are present in living tissue with slow turnover (e.g., dentin and eye lenses) such as for whales (Olsen and Sunde, 2002) and humans (Masters et al., 1977). This trend continues in organic remains such as in wood (Lee et al., 1976), bones (Bada et al., 1973, 1989), and in biospheric remains in the form of dissolved organic matter (Yamaguchi and McCarthy, 2018), subfossil foraminifera (Harada and Handa, 1995), and sedimentary organic matter (Bada et al., 1970; Kvenvolden et al., 1970; Bada and Schroeder, 1975; Bada and Man, 1980). Over time (within 15 million years), a steady state is approached where all amino acid enantiomers are present in a racemic mixture (i.e., D/L ratios equal to 1), which is mostly the case in kerogen (Kvenvolden, 1975) and extraterrestrial amino acids (Engel and Nagy, 1982). In addition to racemization producing D-amino acids, D-amino acids are actively removed from soils by way of enzymatic activity (racemase) by microbes (Zhang and Sun, 2014), further construing environmental amino acid enantiomeric ratios. However, despite these compications, with constraints on temperature history (e.g., organism and sediment temperature), amino acid enantiomers harbor rich information on age constraints (or vice versa), with less precision than radiocarbon, but extending back further in time (see also reviews by Kvenvolden, 1975 and Kaufman and Miller, 1992).
Going a step beyond amino acid-specific radiocarbon, development of a method for enantiomer-specific radiocarbon could be pursued. The technical feasibility of such an approach was recently made plausible, where underivatized individual amino acids, which were previously isolated from sediments, were chromatographically separated into their L and D forms (Blattmann, 2018). If collected, such isolates can provide insight into the radiocarbon population distributions of a single compound on a coarsest of levels. Other efforts to constrain the complex age distributions of organic matter using compound-specific radiocarbon (using fatty acids) have recently come to fruition using time series inversion approaches deconvolving pools of organic matter with different turnovers (French et al., 2018; Vonk et al., 2019). However, even without performing inversions, enantiomer-specific radiocarbon can provide direct insight into the radiocarbon population distributions of individual amino acids. In a simplest end member of cases involving conservative mixing, contributions of biospheric L-amino acids can be teased apart from petrogenic sources, which exhibit D/L ratios equal to 1. In such a case, the age of biospheric L-amino acids can be assessed by mass balance as petrogenic amino acids would equally contribute to the radiocarbon fingerprint of L- and D-amino acids, while biospheric sources would exclusively contribute to the L-amino acid pool (c.f. Silfer et al., 1994). In open systems involving dynamic changes such as contributions of D-amino acids stemming from peptidoglycan from bacteria (c.f. Pelz et al., 1998; Glaser and Amelung, 2002; Veuger et al., 2005), an enantiomer-specific approach can provide insight into microbial pathways of carbon flow embedded in the tapestry of radiocarbon ages contained within dissolved and sedimentary organic matter (e.g., incorporation of carbon derived from young, labile organic matter into bacterial biomass). Additionally, for calibrations of archeological and geochronological time using amino acid enantiomer ratios (e.g., Bada et al., 1984; Kosnik et al., 2013; Simonson et al., 2013), the integrity of purportedly closed systems such as within biominerals can be tested with enantiomer-specific radiocarbon, which can provide insight into radiocarbon disequilibria between enantiomers thereby unveiling open system behavior. The possibility of performing enantiomer-specific radiocarbon for amino acids is unique among most other compounds and the hypothesized benefits for understanding biogeochemical cycles for ecological as well as environmental applications look promising.
Synthesis and Outlook
Following the biospheric pathways along which amino acids are synthesized, propagate through food webs, and decay with a very small fraction finding its way into the geosphere, many questions arise regarding the origin and fate of carbon in these biosphere internal and biosphere-geosphere exchanges (Figure 3). Following their purposes in the biosphere, amino acids face extensive degradation and diagenetic changes, yet remain among the largest characterizable fractions of organic matter preserved in the geologic record (Trask, 1936; Erdman et al., 1956; Wakeham et al., 1997). Amino acid-specific radiocarbon appears a promising tool to confront a range of ecological, sedimentological, and biogeochemical challenges. The authors will report on their progress in future contributions.
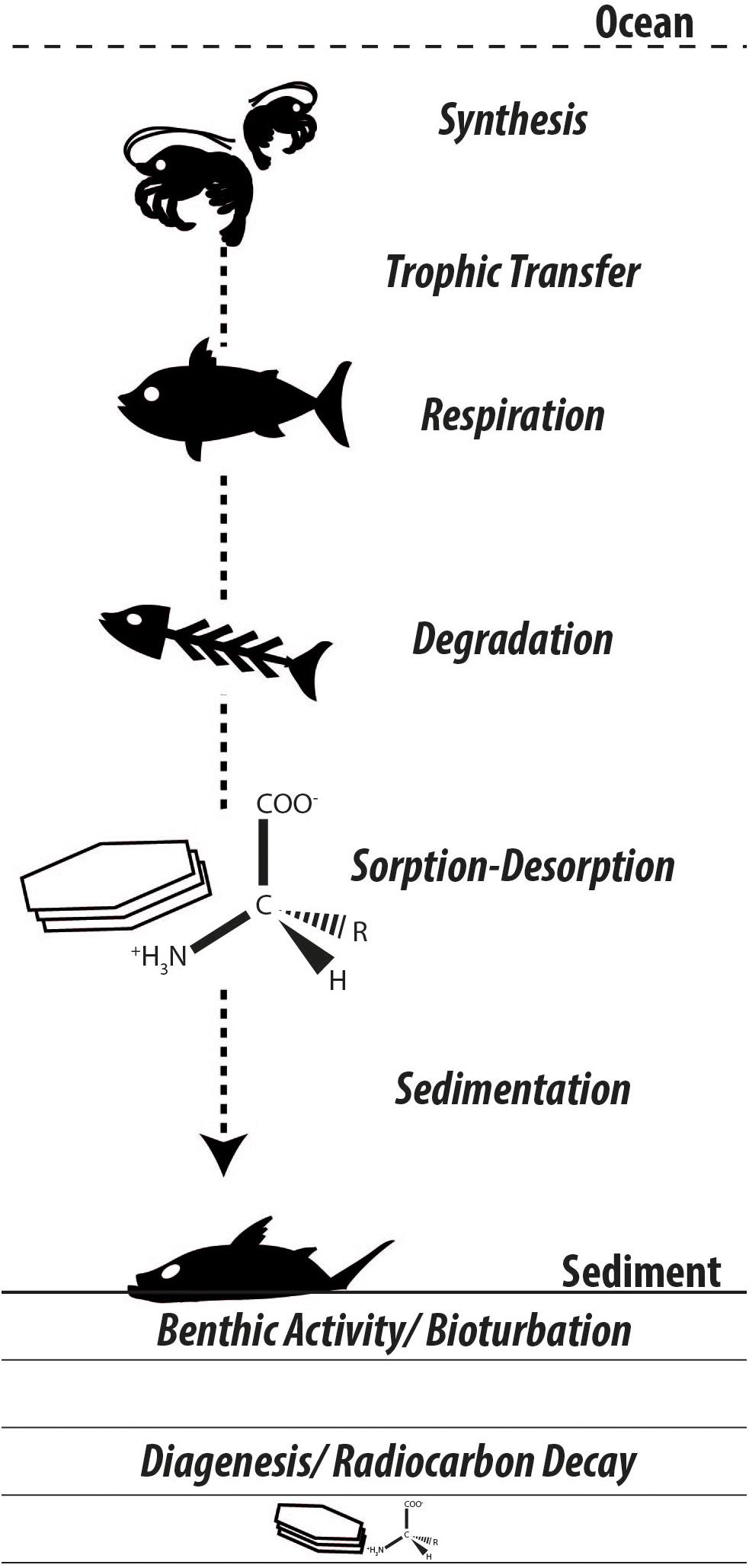
Figure 3. Synthesizing the many unknowns in amino acid biogeochemistry. As carbon is incorporated in biomass and propagates through the food chain, radiocarbon isotopic signatures in individual amino acids should reflect primary (e.g., dissolved inorganic carbon) and secondary (e.g., benthic-derived organic matter) carbon sources. Biomass subject to decay releases amino acids that undergo differential degradation and interact with available mineral substrates, thereby influencing their bioavailability and sealing their fate in the geosphere.
Author Contributions
Both authors equally contributed and wrote this manuscript.
Funding
The authors acknowledge financial support from ETH Research Grant ETH-41 14-1 and JAMSTEC.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank colleagues at JAMSTEC and ETH for stimulating discussions and are grateful for the reviews provided by A. McNichol and S. Kusch. We acknowledge the Hanse-Wissenschaftskolleg Delmenhorst, Germany, for sponsoring the “Marine Organic Biogeochemistry” workshop in April 2019. The workshop was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – project number: 422798570.
References
Abelson, H. (1954). Organic Constituents of Fossils: Carnegie Institute of Washington Year Book, Vol. 53. Washington, DC: Carnegie Institution of Washington, 97–101.
Bada, J. L., Luyendyk, B. P., and Maynard, J. B. (1970). Marine sediments: dating by the racemization of amino acids. Science 170, 730–732. doi: 10.1126/science.170.3959.730
Bada, J. L., Kvenvolden, K. A., and Peterson, E. (1973). Racemization of amino acids in bones. Nature 245, 308–310. doi: 10.1038/245308a0
Bada, J. L., and Schroeder, R. A. (1975). Amino acid racemization reactions and their geochemical implications. Naturwissenschaften 62, 71–79. doi: 10.1007/BF00592179
Bada, J. L., and Man, E. H. (1980). Amino acid diagenesis in deep sea drilling project cores: kinetics and mechanisms of some reactions and their applications in geochronology and in paleotemperature and heat flow determinations. Earth Sci. Rev. 16, 21–55. doi: 10.1016/0012-8252(80)90003-3
Bada, J. L., Gillespie, R., Gowlett, J. A. J., and Hedges, R. E. M. (1984). Accelerator mass spectrometry radiocarbon ages of amino acid extracts from Californian palaeoindian skeletons. Nature 312, 442–444. doi: 10.1038/312442a0
Bada, J. L. (1985). “Racemization of amino acids,” in Chemistry and Biochemistry of the Amino Acids, ed. G. C. Barrett (Dordrecht: Springer), 376–398. doi: 10.1007/978-94-009-4832-7_12
Bada, J. L., Herrmann, B., Payan, I. L., and Man, E. H. (1989). Amino acid racemization in bone and the boiling of the German Emperor Lothar I. Appl. Geochem. 4, 325–327. doi: 10.1016/0883-2927(89)90036-X
Bender, D. A. (2012). Amino Acid Metabolism, 3rd Edn. Oxford: Wiley-Blackwell. doi: 10.1002/9781118357514
Beutelspacher, H. (1955). Wechselwirkung zwischen anorganischen und organischen Kolloiden des Bodens. Zeitschrift Pflanzenernährung Düngung Bodenkunde 69, 108–115. doi: 10.1002/jpln.19550690118
Blattmann, T. M. (2018). Topics on Radiocarbon Geochemistry and Organic Matter-Mineral Interactions. Ph.D. thesis, ETH Zürich, Zurich. doi: 10.3929/ethz-b-000313397
Blattmann, T. M., Zhang, Y., Zhao, Y., Wen, K., Lin, S., Li, J., et al. (2018). Contrasting fates of petrogenic and biospheric carbon in the South China Sea. Geophys. Res. Lett. 45, 9077–9086. doi: 10.1029/2018GL079222
Blattmann, T. M., Liu, Z., Zhang, Y., Zhao, Y., Haghipour, N., Montluçon, D. B., et al. (2019). Mineralogical control on the fate of continentally derived organic matter in the ocean. Science 366:eaax5345. doi: 10.1126/science.aax5345
Blattmann, T. M., Montlucon, D. B., Haghipour, N., Ishikawa, N. F., and Eglinton, T. I. (2020). Liquid chromatographic isolation of individual amino acids extracted from sediments for radiocarbon analysis. Front. Mar. Sci. 7:174. doi: 10.3389/fmars.2020.00174
Bour, A. L., Walker, B. D., Broek, T. A. B., and McCarthy, M. D. (2016). Radiocarbon analysis of individual amino acids: carbon blank quantification for a small-sample high-pressure liquid chromatography purification method. Anal. Chem. 88, 3521–3528. doi: 10.1021/acs.analchem.5b03619
Broecker, W. S., and Walton, A. (1959). The geochemistry of C14 in fresh-water systems. Geochim. Cosmochim. Acta 16, 15–38. doi: 10.1016/0016-7037(59)90044-4
Carstens, D., Lehmann, M. F., Hofstetter, T. B., and Schubert, C. J. (2013). Amino acid nitrogen isotopic composition patterns in lacustrine sedimenting matter. Geochim. Cosmochim. Acta 121, 328–338. doi: 10.1016/j.gca.2013.07.020
Chikaraishi, Y., Ogawa, N. O., Kashiyama, Y., Takano, Y., Suga, H., Tomitani, A., et al. (2009). Determination of aquatic food-web structure based on compound-specific nitrogen isotopic composition of amino acids. Limnol. Oceanogr. Methods 7, 740–750. doi: 10.1016/j.jchromb.2016.09.004
Chikaraishi, Y., Steffan, S. A., Takano, Y., and Ohkouchi, N. (2015). Diet quality influences isotopic discrimination among amino acids in an aquatic vertebrate. Ecol. Evol. 5, 2048–2059. doi: 10.1002/ece3.1491
Corrigan, J. J. (1969). D-Amino acids in animals. Science 164, 142–149. doi: 10.1126/science.164.3876.142
Cowie, G. L., and Hedges, J. I. (1992). Improved amino acid quantification in environmental samples: charge-matched recovery standards and reduced analysis time. Mar. Chem. 37, 223–238. doi: 10.1016/0304-4203(92)90079-P
Degens, E. T., and Bajor, M. (1960). Die Verteilung von Aminosäuren in bitumin sen Sedimenten und ihre Bedeutung für die Kohlen- und Erdölgeologie. Glückauf 96, 1525–1534.
Degens, E. T. (1970). “Molecular nature of nitrogenous compounds in sea water and recent marine sediments,” in Proceedings of the Symposium on Organic Matter in Natural Waters, ed. D. H. Hood (Alaska: Institute of Marine Science University), 77–106.
Ding, X., and Henrichs, S. M. (2002). Adsorption and desorption of proteins and polyamino acids by clay minerals and marine sediments. Mar. Chem. 77, 225–237. doi: 10.1016/S0304-4203(01)00085-8
Engel, M. H., and Nagy, B. (1982). Distribution and enantiomeric composition of amino acids in the Murchison meteorite. Nature 296, 837–840. doi: 10.1038/296837a0
Ensminger, L. E., and Gieseking, J. E. (1941). The adsorption of proteins by montmorillonitic clays and its effect on base-exchange capacity. Soil Sci. 51, 125–132.
Erdman, J. G., Marlett, E. M., and Hanson, W. E. (1956). Survival of amino acids in marine sediments. Science 124, 1026–1026. doi: 10.1126/science.124.3230.1026
Estes, E. R., Pockalny, R., D’Hondt, S., Inagaki, F., Morono, Y., Murray, R. W., et al. (2019). Persistent organic matter in oxic subseafloor sediment. Nat. Geosci. 12, 126–131. doi: 10.1038/s41561-018-0291-5
Eusterhues, K., Rumpel, C., and Kögel-Knabner, I. (2005). Stabilization of soil organic matter isolated via oxidative degradation. Organ. Geochem. 36, 1567–1575. doi: 10.1016/j.orggeochem.2005.06.010
French, K. L., Hein, C. J., Haghipour, N., Wacker, L., Kudrass, H. R., Eglinton, T. I., et al. (2018). Millennial soil retention of terrestrial organic matter deposited in the Bengal Fan. Sci. Rep. 8:11997. doi: 10.1038/s41598-018-30091-8
Glaser, B., and Amelung, W. (2002). Determination of 13C natural abundance of amino acid enantiomers in soil: methodological considerations and first results. Rapid Commun. Mass Spectrom. 16, 891–898. doi: 10.1002/rcm.650
Goto, A. S., Miura, K., Korenaga, T., Hasegawa, T., Ohkouchi, N., and Chikaraishi, Y. (2018). Fractionation of stable nitrogen isotopes (15N/14N) during enzymatic deamination of glutamic acid: implications for mass and energy transfers in the biosphere. Geochem. J. 52, 273–280. doi: 10.2343/geochemj.2.0513
Guillemette, F., Bianchi, T. S., and Spencer, R. G. M. (2017). Old before your time: ancient carbon incorporation in contemporary aquatic foodwebs. Limnol. Oceanogr. 62, 1682–1700. doi: 10.1002/lno.10525
Harada, N., and Handa, N. (1995). Amino acid chronology in the fossil planktonic foraminifera, Pulleniatina obliquiloculata from Pacific Ocean. Geophys. Res. Lett. 22, 2353–2356. doi: 10.1029/95GL02112
Hare, E. (1969). “Geochemistry of proteins, peptides, and amino acids,” in Organic Geochemistry: Methods and Results, eds G. Eglinton and M. T. J. Murphy (Berlin: Springer), 438–462. doi: 10.1007/978-3-642-87734-6_22
Hare, E., Fogel, M. L., Stafford, T. W. Jr., Mitchell, A. D., and Hoering, T. C. (1991). The isotopic composition of carbon and nitrogen in individual amino acids isolated from modern and fossil proteins. J. Arch. Sci. 18, 277–292. doi: 10.1016/0305-4403(91)90066-X
Hedges, J. I, and Parker, L. (1976). Land-derived organic matter in surface sediments from the Gulf of Mexico. Geochim. Cosmochim. Acta 40, 1019–1029. doi: 10.1016/j.marpolbul.2017.04.042
Hedges, J. I. (1978). The formation and clay mineral reactions of melanoidins. Geochim. Cosmochim. Acta 42, 69–76. doi: 10.1016/0016-7037(78)90218-1
Hedges, I., and Hare, E. (1987). Amino acid adsorption by clay minerals in distilled water. Geochim. Cosmochim. Acta 51, 255–259. doi: 10.1016/0016-7037(87)90237-7
Hedges, I., Baldock, J. A., Gélinas, Y., Lee, C., Peterson, M., and Wakeham, S. G. (2001). Evidence for non-selective preservation of organic matter in sinking marine particles. Nature 409, 801–804. doi: 10.1038/35057247
Henrichs, S. M., and Sugai, S. F. (1993). Adsorption of amino acids and glucose by sediments of Resurrection Bay, Alaska, USA: functional group effects. Geochim. Cosmochim. Acta 57, 823–835. doi: 10.1016/0016-7037(93)90171-R
Hwang, J., Druffel, E. R. M., and Komada, T. (2005). Transport of organic carbon from the California coast to the slope region: a study of Δ14C and δ13C signatures of organic compound classes. Glob. Biogeochem. Cycles 19:GB2018. doi: 10.1029/2004GB002422
Ishikawa, N. F., Hyodo, F., and Tayasu, I. (2013). Use of carbon-13 and carbon-14 natural abundances for stream food web studies. Ecol. Res. 28, 759–769. doi: 10.1007/s11284-012-1003-z
Ishikawa, N. F., Itahashi, Y., Blattmann, T. M., Takano, Y., Ogawa, N. O., Yamane, M., et al. (2018). Improved method for isolation and purification of underivatized amino acids for radiocarbon analysis. Anal. Chem. 90, 12035–12041. doi: 10.1021/acs.analchem.8b02693
Ishikawa, N. F. (2018). Use of compound-specific nitrogen isotope analysis of amino acids in trophic ecology: assumptions, applications, and implications. Ecol. Res. 33, 825–837. doi: 10.1007/s11284-018-1616-y
Kaufman, D. S., and Miller, G. H. (1992). Overview of amino acid geochronology: comparative biochemistry and physiology Part B. Comp. Biochem. 102, 199–204. doi: 10.1016/0305-0491(92)90110-D
Keil, R. G., Montluçon, D. B., Prahl, F. G., and Hedges, J. I. (1994a). Sorptive preservation of labile organic matter in marine sediments. Nature 370, 549–552. doi: 10.1038/370549a0
Keil, R. G., Tsamakis, E., Fuh, C. B., Giddings, J. C., and Hedges, J. I. (1994b). Mineralogical and textural controls on the organic composition of coastal marine sediments: hydrodynamic separation using SPLITT-fractionation. Geochim. Cosmochim. Acta 58, 879–893. doi: 10.1016/0016-7037(94)90512-6
Keil, R. G., Mayer, L. M., Quay, D., Richey, J. E., and Hedges, J. I. (1997). Loss of organic matter from riverine particles in deltas. Geochim. Cosmochim. Acta 61, 1507–1511. doi: 10.1016/S0016-7037(97)00044-6
Keil, R. G., Tsamakis, E., Giddings, J. C., and Hedges, J. I. (1998). Biochemical distributions (amino acids, neutral sugars, and lignin phenols) among size-classes of modern marine sediments from the Washington coast. Geochim. Cosmochim. Acta 62, 1347–1364. doi: 10.1016/S0016-7037(98)00080-5
Keil, R. G., and Fogel, M. L. (2001). Reworking of amino acid in marine sediments: stable carbon isotopic composition of amino acids in sediments along the Washington coast. Limnol. Oceanogr. 46, 14–23. doi: 10.4319/lo.2001.46.1.0014
Keil, R. G., and Mayer, L. M. (2014). “Mineral matrices and organic matter,” in Treatise on Geochemistry, eds H. D. Holland and K. K. Turekian (Oxford: Elsevier), 337–359. doi: 10.1016/B978-0-08-095975-7.01024-X
Kennedy, M. J., Pevear, D. R., and Hill, R. J. (2002). Mineral surface control of organic carbon in black shale. Science 295, 657–660. doi: 10.1126/science.1066611
Kirchman, D. L. (1990). Limitation of bacterial growth by dissolved organic matter in the subarctic Pacific. Mar. Ecol. Prog. Ser. 62, 47–54. doi: 10.3354/meps062047
Kitadai, N., and Maruyama, S. (2018). Origins of building blocks of life: a review. Geosci. Front. 9, 1117–1153. doi: 10.1016/j.gsf.2017.07.007
Kleber, M., Eusterhues, K., Keiluweit, M., Mikutta, C., Mikutta, R., and Nico, S. (2015). “Mineral–organic associations: formation, properties, and relevance in soil environments,” in Advances in Agronomy, Vol. 130, ed. D. L. Sparks (Cambridge, MA: Academic Press), 1–140. doi: 10.1016/bs.agron.2014.10.005
Kosnik, M. A., Kaufman, D. S., and Hua, Q. (2013). Radiocarbon-calibrated multiple amino acid geochronology of Holocene molluscs from Bramble and Rib Reefs (Great Barrier Reef, Australia). Quat. Geochronol. 16, 73–86. doi: 10.1016/j.quageo.2012.04.024
Kvenvolden, K. A., Peterson, E., and Brown, F. S. (1970). Racemization of amino acids in sediments from Saanich Inlet, British Columbia. Science 169, 1079–1082. doi: 10.1126/science.169.3950.1079
Kvenvolden, K. A. (1975). Advances in the geochemistry of amino acids. Annu. Rev. Earth Planet. Sci. 3, 183–212. doi: 10.1146/annurev.ea.03.050175.001151
Lam, H., Oh, D.-C., Cava, F., Takacs, C. N., Clardy, J., de Pedro, M. A., et al. (2009). D-Amino acids govern stationary phase cell wall remodeling in bacteria. Science 325, 1552–1555. doi: 10.1126/science.1178123
Larsen, T., Taylor, D. L., Leigh, M. B., and O’Brien, D. M. (2009). Stable isotope fingerprinting: a novel method for identifying plant, fungal, or bacterial origins of amino acids. Ecology 90, 3526–3535. doi: 10.1890/08-1695.1
Larsen, T., Ventura, M., Andersen, N., O’Brien, D. M., Piatkowski, U., and McCarthy, M. D. (2013). Tracing carbon sources through aquatic and terrestrial food webs using amino acid stable isotope fingerprinting. PLoS One 8:e73441. doi: 10.1371/journal.pone.0073441
Lee, C., Bada, J. L., and Peterson, E. (1976). Amino acids in modern and fossil woods. Nature 259, 183–186. doi: 10.1038/259183a0
Lee, C., Wakeham, S. G., and Farrington, J. W. (1983). Variations in the composition of particulate organic matter in a time-series sediment trap. Mar. Chem. 13, 181–194. doi: 10.1016/0304-4203(83)90013-0
Libby, W. F., Anderson, E. C., and Arnold, J. R. (1949). Age determination by radiocarbon content – World-wide assay of natural radiocarbon. Science 109, 227–228. doi: 10.1126/science.109.2827.227
Macko, S. A., Estep, M. L. F., Engel, M. H., and Hare, E. (1986). Kinetic fractionation of stable nitrogen isotopes during amino acid transamination. Geochim. Cosmochim. Acta 50, 2143–2146. doi: 10.1016/0016-7037(86)90068-2
Masters, M., Bada, J. L., and Samuel Zigler, J. (1977). Aspartic acid racemisation in the human lens during ageing and in cataract formation. Nature 268, 71–73. doi: 10.1038/268071a0
Mayer, L. M. (1994). Surface area control of organic carbon accumulation in continental shelf sediments. Geochim. Cosmochim. Acta 58, 1271–1284. doi: 10.1016/0016-7037(94)90381-6
McCullagh, J. S. O., Marom, A., and Hedges, R. E. M. (2010). Radiocarbon dating of individual amino acids from archaeological bone collagen. Radiocarbon 52, 620–634. doi: 10.1017/S0033822200045653
McIntyre, C. P., Wacker, L., Haghipour, N., Blattmann, T. M., Fahrni, S., Usman, M., et al. (2017). Online 13C and 14C gas measurements by EA-IRMS–AMS at ETH Zürich. Radiocarbon 59, 893–903.
McMahon, K. W., Fogel, M. L., Elsdon, T. S., and Thorrold, S. R. (2010). Carbon isotope fractionation of amino acids in fish muscle reflects biosynthesis and isotopic routing from dietary protein. J. Anim. Ecol. 79, 1132–1141. doi: 10.1111/j.1365-2656.2010.01722.x
McMahon, K. W., Thorrold, S. R., Houghton, L. A., and Berumen, M. L. (2016). Tracing carbon flow through coral reef food webs using a compound-specific stable isotope approach. Oecologia 180, 809–821. doi: 10.1007/s00442-015-3475-3
Miller, S. L., and Urey, H. C. (1959). Organic compound synthesis on the primitive Earth. Science 130, 245–251. doi: 10.1126/science.130.3370.245
Montluçon, D. B., and Lee, C. (2001). Factors affecting lysine sorption in a coastal sediment. Organ. Geochem. 32, 933–942. doi: 10.1016/S0146-6380(01)00050-X
Müller, J. (1977). C/N ratios in Pacific deep-sea sediments: effect of inorganic ammonium and organic nitrogen compounds sorbed by clays. Geochim. Cosmochim. Acta 41, 765–776. doi: 10.1016/0016-7037(77)90047-3
Neuberger, A. (1948). “Stereochemistry of amino acids,” in Advances in Protein Chemistry, eds M. L. Anson and J. T. Edsall (Cambridge, MA: Academic Press), 297–383. doi: 10.1016/S0065-3233(08)60009-1
Ohkouchi, N., Chikaraishi, Y., Close, H. G., Fry, B., Larsen, T., Madigan, D. J., et al. (2017). Advances in the application of amino acid nitrogen isotopic analysis in ecological and biogeochemical studies. Organ. Geochem. 113, 150–174. doi: 10.1016/j.orggeochem.2017.07.009
Olsen, E., and Sunde, J. (2002). Age determination of minke whales (Balaenoptera acutorostrata) using the aspartic acid racemization technique. Sarsia 87, 1–8. doi: 10.1080/003648202753631686
Pantoja, S., and Lee, C. (2003). Amino acid remineralization and organic matter lability in Chilean coastal sediments. Organ. Geochem. 34, 1047–1056. doi: 10.1016/S0146-6380(03)00085-8
Pelz, O., Cifuentes, L. A., Hammer, B. T., Kelley, C. A., and Coffin, R. B. (1998). Tracing the assimilation of organic compounds using δ13C analysis of unique amino acids in the bacterial peptidoglycan cell wall. FEMS Microbiol. Ecol. 25, 229–240. doi: 10.1111/j.1574-6941.1998.tb00475.x
Petsch, S. T., Eglinton, T. I., and Edwards, K. J. (2001). 14C-dead living biomass: evidence for microbial assimilation of ancient organic carbon during shale weathering. Science 292, 1127–1131. doi: 10.1126/science.1058332
Pinck, L. A., and Allison, F. E. (1951). Resistance of a protein-montmorillonite complex to decomposition by soil microorganisms. Science 114, 130–131. doi: 10.1126/science.114.2953.130
Schmidt, M. W. I., Torn, M. S., Abiven, S., Dittmar, T., Guggenberger, G., Janssens, I. A., et al. (2011). Persistence of soil organic matter as an ecosystem property. Nature 478, 49–56. doi: 10.1038/nature10386
Silfer, J. A., Qian, Y., Macko, S. A., and Engel, M. H. (1994). Stable carbon isotope compositions of individual amino acid enantiomers in mollusc shell by GC/C/IRMS. Organ. Geochem. 21, 603–609. doi: 10.1016/0146-6380(94)90006-X
Simonson, A. E., Lockwood, R., and Wehmiller, J. F. (2013). Three approaches to radiocarbon calibration of amino acid racemization in Mulinia lateralis from the Holocene of the Chesapeake Bay, USA. Quat. Geochronol. 16, 62–72. doi: 10.1016/j.quageo.2012.06.005
Stuiver, M., and Polach, H. A. (1977). Discussion: reporting of 14C data. Radiocarbon 19, 355–363. doi: 10.1017/S0033822200003672
Trask, D. (1936). Inferences about the origin of oil as indicated by the composition of the organic constituents of sediments. Paper Presented at the Washington: United States Government Printing Office, Professional Paper / United States Department of the Interior, Geological Survey, Washington, DC.
Vernadsky, W. J. (1930). Geochemie in Ausgewählten Kapiteln. Leipzig: Akademische Verlagsgesellschaft, 370.
Veuger, B., Middelburg, J. J., Boschker, H. T. S., and Houtekamer, M. (2005). Analysis of 15N incorporation into D-alanine: a new method for tracing nitrogen uptake by bacteria. Limnol. Oceanogr. Methods 3, 230–240. doi: 10.4319/lom.2005.3.230
Volkman, J. K., Eglinton, G., Corner, E. D. S., and Forsberg, T. E. (1980). Long-chain alkenes and alkenones in the marine coccolithophorid Emiliania huxleyi. Phytochemistry 19, 2619–2622. doi: 10.1016/s0031-9422(00)00120-5
Vonk, J. E., Drenzek, N. J., Hughen, K. A., Stanley, R. H. R., McIntyre, C., Montluçon, D. B., et al. (2019). Temporal deconvolution of vascular plant-derived fatty acids exported from terrestrial watersheds. Geochim. Cosmochim. Acta 244, 502–521. doi: 10.1016/j.gca.2018.09.034
Wakeham, S. G., Lee, C., Hedges, J. I., Hernes, J., and Peterson, M. J. (1997). Molecular indicators of diagenetic status in marine organic matter. Geochim. Cosmochim. Acta 61, 5363–5369. doi: 10.1016/S0016-7037(97)00312-8
Wang, X.-C., and Lee, C. (1993). Adsorption and desorption of aliphatic amines, amino acids and acetate by clay minerals and marine sediments. Mar. Chem. 44, 1–23. doi: 10.1016/0304-4203(93)90002-6
Wang, X.-C., Druffel, E. R. M., and Lee, C. (1996). Radiocarbon in organic compound classes in particulate organic matter and sediment in the deep northeast Pacific ocean. Geophys. Res. Lett. 23, 3583–3586. doi: 10.1029/96GL03423
Wang, X.-C., Druffel, E. R. M., Griffin, S., Lee, C., and Kashgarian, M. (1998). Radiocarbon studies of organic compound classes in plankton and sediment of the northeastern Pacific ocean. Geochim. Cosmochim. Acta 62, 1365–1378. doi: 10.1016/S0016-7037(98)00074-X
Williams, M., Druffel, E. R. M., and Smith, K. L. (1987). Dietary carbon sources for deep-sea organisms as inferred from their organic radiocarbon activities: deep sea research part A. Oceanogr. Res. Pap. 34, 253–266. doi: 10.1016/0198-0149(87)90085-9
Yamaguchi, Y. T., and McCarthy, M. D. (2018). Sources and transformation of dissolved and particulate organic nitrogen in the North Pacific Subtropical Gyre indicated by compound-specific δ15N analysis of amino acids. Geochim. Cosmochim. Acta 220, 329–347. doi: 10.1016/j.gca.2017.07.036
Yokoyama, T., Kan-No, N., Ogata, T., Kotaki, Y., Sato, M., and Nagahisa, E. (2003). Presence of free D-amino acids in microalgae: bioscience. Biotechnol. Biochem. 67, 388–392. doi: 10.1271/bbb.67.388
Keywords: amino acid, radiocarbon, ecology, food web, organic matter-mineral interactions, peptide, compound specific isotope analysis, protein
Citation: Blattmann TM and Ishikawa NF (2020) Theoretical Amino Acid-Specific Radiocarbon Content in the Environment: Hypotheses to Be Tested and Opportunities to Be Taken. Front. Mar. Sci. 7:302. doi: 10.3389/fmars.2020.00302
Received: 29 October 2019; Accepted: 15 April 2020;
Published: 05 May 2020.
Edited by:
Stuart Wakeham, University of Georgia, United StatesReviewed by:
Ann P. McNichol, Woods Hole Oceanographic Institution, United StatesStephanie Kusch, University of Cologne, Germany
Copyright © 2020 Blattmann and Ishikawa. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Thomas M. Blattmann, YmxhdHRtYW5udEBqYW1zdGVjLmdvLmpw; Naoto F. Ishikawa, aXNoaWthd2FuQGphbXN0ZWMuZ28uanA=
 Thomas M. Blattmann
Thomas M. Blattmann Naoto F. Ishikawa
Naoto F. Ishikawa