- 1Department of Rehabilitation Medicine, Center for Rehabilitation Medicine, Rehabilitation and Sports Medicine Research Institute of Zhejiang Province, Zhejiang Provincial People’s Hospital (Affiliated People’s Hospital, Hangzhou Medical College), Hangzhou, China
- 2Department of Rheumatology, Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, Hangzhou, China
- 3Zhejiang University-University of Edinburgh Institute, Zhejiang University, Hangzhou, China
Objective: The association of fat mass and obesity-related (FTO) gene with osteoarthritis (OA) risk has been investigated in multiple genome-wide association studies but showed inconsistent results. Our study aimed to assess FTO expression in different OA sequencing datasets and to meta-analyze whether FTO polymorphism was associated with the risk of osteoarthritis.
Method: Gene expression profiles were obtained from ArrayExpress, Gene Expression Omnibus (GEO), and BioProject databases. Three electronic databases including PubMed and EMBASE were systematically retrieved to identify articles exploring the association between FTO polymorphisms and OA risk published before September 2022. Summary odds ratios (ORs) and corresponding 95% confidence intervals (95% CIs) were calculated to perform the result. Stata software was utilized to conduct analyses on predetermined ethnicity and gender subgroups and sensitivity.
Results: FTO gene was differentially expressed in the datasets from the UK. This systematic review and meta-analysis encompasses eight studies that revealed a significant association between FTO polymorphisms and OA risk [OR 1.07, 95% CI (1.03, 1.11), P < 0.001] in the overall population. In subgroup analysis, a marked association was observed in European Caucasian [OR 1.08, 95% CI (1.04–1.12), P < 0.001] and North American Caucasian with the Asian subgroups [OR 0.98, 95% CI (0.83–1. 6), P = 0.83] as an exception. Among the studies, four of them demonstrated attenuation in their OA risk after body mass index (BMI) adjustment in Caucasian populations.
Conclusion: FTO significant differential expression was associated with the increased risk of OA in Caucasian populations. Nevertheless, the causality between FTO polymorphisms and OA risk remains largely elusive. Hence, further studies with larger sample size are necessary to validate whether FTO gene polymorphism contributes to OA susceptibility.
Introduction
Osteoarthritis (OA) is the most prevailing form of whole joint degenerative disease characterized by the degeneration of articular cartilage, bone remodeling, synovial inflammation, osteophyte formation, subchondral sclerosis, infrapatellar fat pad, and meniscus injuries, etc. (1). Its prevalence does not cease to escalate due to population aging, prolonged life expectancy and obesity, making the disease a major healthcare problem and socioeconomic burden affecting millions of people worldwide (1). OA is a multifactorial disease as its pathogenesis is an amalgamative effect of environmental factors such as traumatic joint injury and chronic mechanical overloading alongside genetic risk factors such as aging, gender, genetic predisposition, obesity, and inflammation (2). Previous studies frequently associate obesity with an augmented risk of OA, but how it contributes to the onset and progression of OA has not been well-established (3). On the other hand, it has been demonstrated the presence of OA in non-weight-bearing joints of obese subjects and obesity determines a low-grade inflammatory systemic inflammatory status. Thus, it is suggested that other factors other than mechanical loading contribute to the disease (4, 5).
The fat mass and obesity-associated (FTO) gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase which is the first well-established obesity-susceptibility gene (6). Recently, several genome-wide association studies (GWAS) explored the relationship between FTO gene variation and OA risk (7–9). However, these studies presented incongruent and inconclusive results attributed to the clinical heterogeneity of patients and various single nucleotide polymorphisms (SNP), different ethnic populations, and small sample sizes. In addition, the microarray and RNA-sequencing data provide us the possibility to investigate whether FTO is a candidate gene for OA susceptibility. To precisely elucidate the role of the FTO gene in the development of OA, we firstly detected the FTO expression between OA and normally followed by a comprehensive meta-analysis to determine the association between FTO polymorphisms and OA risk.
Method
Search strategy
Microarray and RNA-sequencing data from cartilage samples in OA patients were obtained from ArrayExpress, Gene Expression Omnibus (GEO), and BioProject databases using the search terms “osteoarthritis,” and “cartilage.” We conducted literature searches of databases which include PubMed, EMBASE to retrieve relevant articles that underlined the associations between FTO polymorphisms and OA up to September 1, 2022 with “FTO” AND (“OA” OR “osteoarthritis” OR “arthrosis”) as keywords. The search strategy in detail that we performed is illustrated in Supplementary Tables 1, 2. Additionally, the references of related studies were also screened to identify potentially relevant studies. This systematic review and meta-analysis was conducted by adhering to the Preferred Reporting Items for Systematic Reviews and Meta-analyses (PRISMA) reporting guideline (10).
Inclusion and exclusion criteria
Two investigators assessed the retrieved studies independently according to the pre-specified inclusion criteria as follows: studies that (1) case-control or cohort design; (2) evaluated the association between FTO gene polymorphism and knee or hip OA, no limitation in single-nucleotide polymorphisms (SNPs) sites; (3) contained genotype data for the calculation of odds ratios (ORs) and 95% confidence intervals (CIs); (4) were written in English. If several articles reported findings for repeated study populations, we only selected the most recent study or the one with the largest sample size. Any disagreements will be solved by discussion to decide for inclusion or exclusion of the study for the meta-analysis.
Data extraction and quality assessment
Two investigators extracted the following information from each eligible study independently: first author, year of publication, country, ethnic origin of the study population, names of SNPs, type of OA and sample size, age, female proportion of cases and controls.
Two investigators analyzed the methodological quality of each study by applying the Newcastle–Ottawa Scale (NOS), in terms of the selection of study participants, comparability of outcome groups and outcome measures.
Any disagreements will be resolved by discussion until consensus is reached.
Statistical analysis
Microarray datasets were obtained using the “GEOquery” R package (11), and after probe id conversion, the “edgR” R package was used to normalize the data with the CPM (computes counts per million) function (12). RNA sequencing datasets were normalized by applying the variance stabilizing transformation (VST) function from the “DESeq2” R package (13). Mann–Whitney U test was utilized to compare the FTO expression between the OA group and controls. These computational and statistical analyses were performed using the R software.1
The odd ratios (ORs) and 95% confidence intervals (CIs) were estimated by the random effects model (DerSimonian and Laird methods) to evaluate the strength of correlation between FTO gene polymorphism and OA risk. Stratification analyses were carried out by ethnicity and gender. P < 0.05 was considered statistically significant. Sensitivity analysis was performed by repeating analysis after omitting one study each time to estimate the impact on the overall effects. Heterogeneity was assessed by Q statistic with P-value and I2 statistic (14). Potential publication bias will be examined by Egger’s test if more than 10 studies were included (15). These data analyses were performed in Stata 16.0 (Stata Corp, College Station, TX, USA).
Results
Fat mass and obesity-related expression between osteoarthritis and control
A total of 208 records were derived after incipient search. GSE169077 (USA), GSE117999 (USA), GSE11400 (USA), E-MTAB-6266 (UK), and PRJNA505578 (China) datasets were included. Our results (Figure 1) revealed that FTO demonstrated a significantly increased differential expression (P < 0.001) in the UK OA population but not for the USA or Chinese population (P > 0.05).
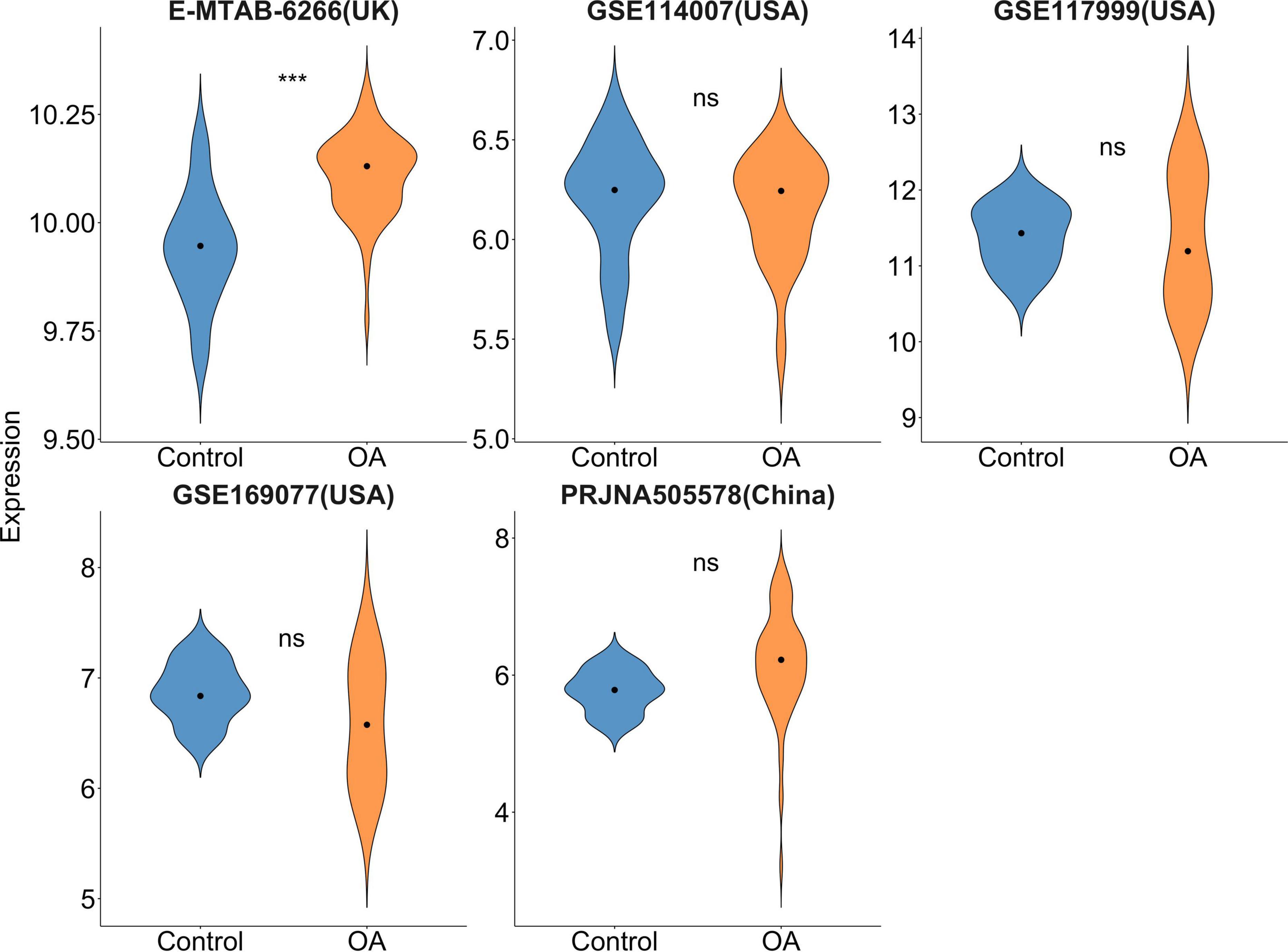
Figure 1. Violin plot of FTO gene expression in microarray and RNA-sequencing data. Dots mean median. ***P < 0.001.
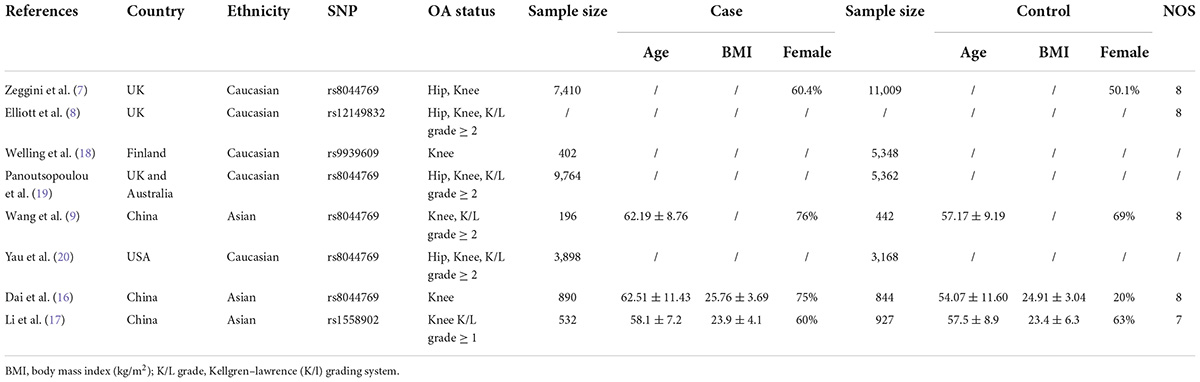
Characteristics of the included studies for meta-analysis
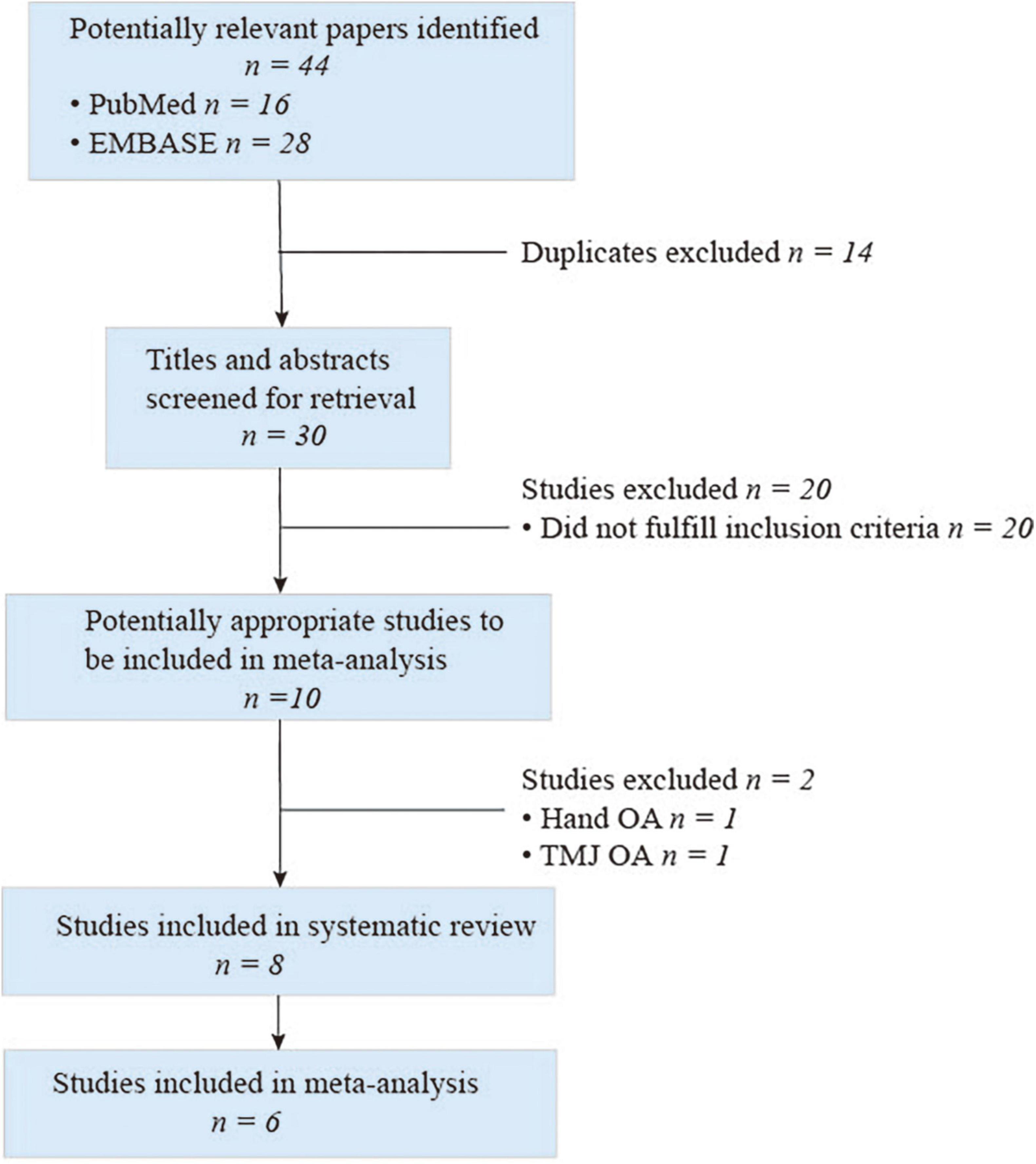
Selection for qualified studies was presented in Figure 2. Our initial computerized literature search identified a total of 44 citations. Among these results, 14 records were duplication, and 20 records did not meet our inclusion criteria following a thorough review of the titles and abstracts. Ten citations were retrieved for further full-text review; two out of the 10 studies investigated the association of FTO polymorphism with hand or temporomandibular joint (TMJ) OA, respectively. Eventually, we identified eight eligible citations for systematic review (7–9, 16–20) and six studies for meta-analysis (7–9, 16–18). The characteristics and quality of these included studies are summarized in Table 1. These available cohort studies were conducted in three countries (number of studies): the UK (2), Finland (1); and China (3) for meta-analysis, and the other two studies synthesized rs8044769 SNP and OA risk in different independent study cohorts (19, 20). Four FTO polymorphisms rs8044769 (7, 9, 16, 19, 20), rs12149832 (8), rs9939609 (18), rs1558902 (17) were investigated in this meta-analysis and systematic review. These results of quality assessment were not performed as one studies was from abstract (18) and two studies from several cohorts (19, 20).
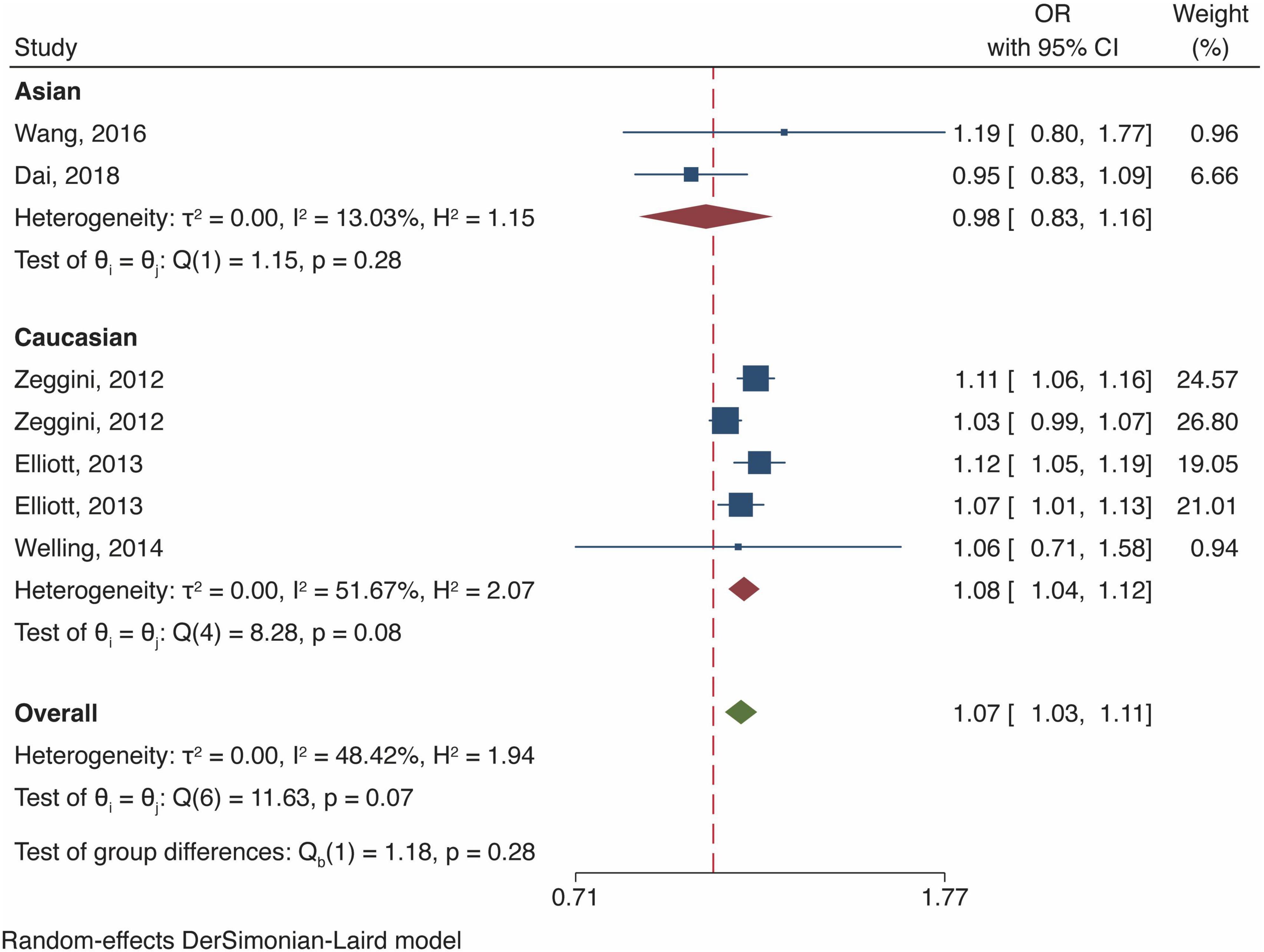
Meta-analysis of fat mass and obesity-related gene polymorphism and osteoarthritis risk in all population
In the general analysis, we found that FTO gene polymorphism increased OA risk [OR and 95% CI, 1.07 (1.03, 1.11), P < 0.001, Figure 3] with accept heterogeneity (I2 = 48.42%). Stratified analysis of ethnicity showed that the risk of OA was considerably elevated by FTO polymorphism thein European Caucasian (OR 1.08 [95% CI 1.04–1.12], P < 0.001, I2 = 51.67%, Figure 3) but did not reveal a statistically significant rise in Asian (OR 0.98 [95% CI 0.83–1.16] P = 0.83, Figure 3) with low heterogeneity (I2 = 13.03%). Meanwhile, Yau et al. documented that FTO polymorphism (rs8044769) increased OA risk in North American Caucasian (OR 1.10 [95% CI 1.03–1.19], P = 0.00613) (20). Four studies investigated the effect of body mass index (BMI) covariate on the OA outcome, consistent herewith, we observed an attenuation of the OA risk after BMI adjustment in the Caucasian population (7, 8, 18, 19). Nonetheless, we did not discover any solid association between FTO polymorphism (rs8044769) and higher BMI in the Chinese population (9, 16).
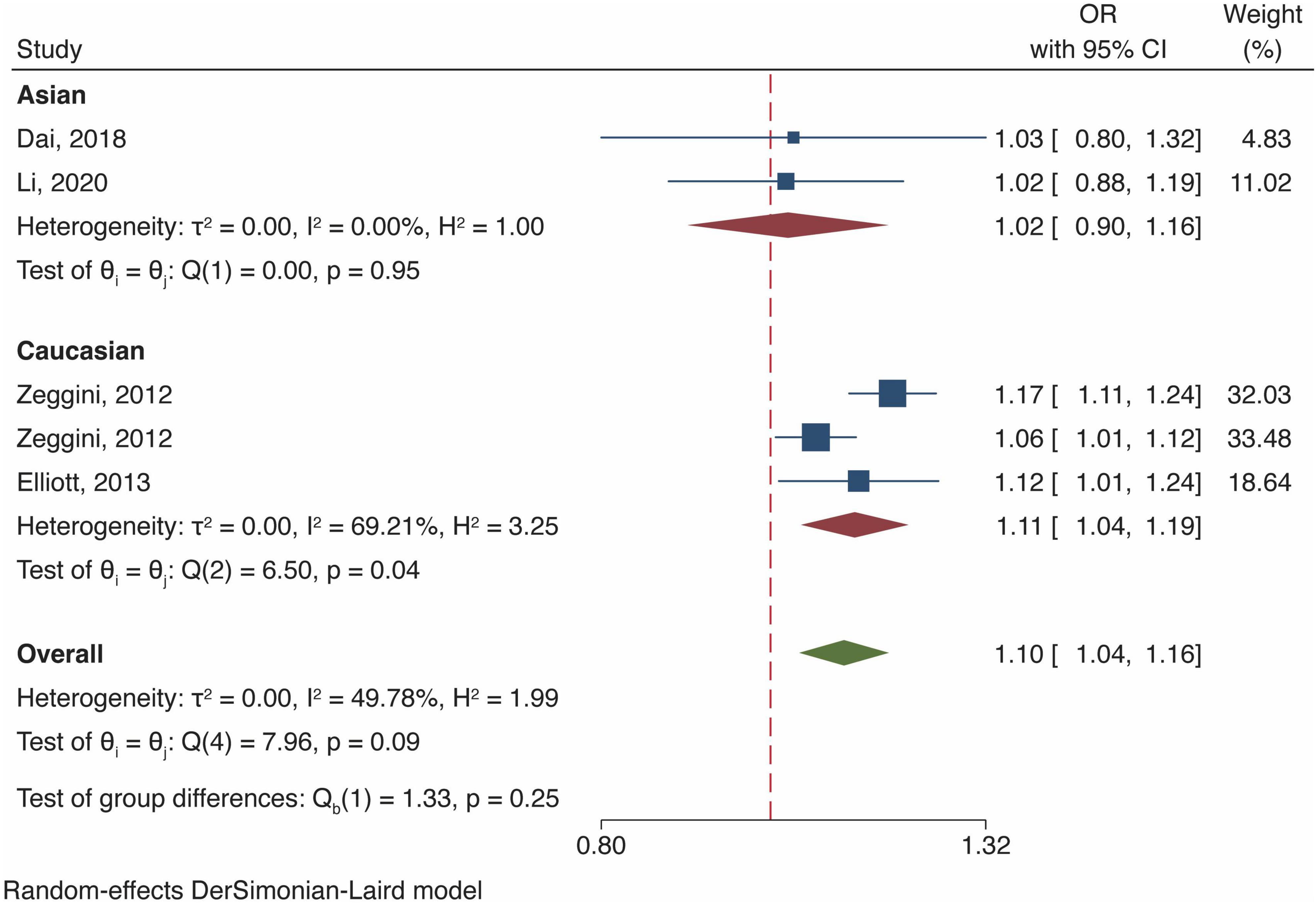
Meta-analysis of fat mass and obesity-related gene polymorphism and osteoarthritis risk in female population
Four studies investigated the association of FTO polymorphism and OA risk in female population, all of which reported results that ascribed the increase in OA risk to FTO gene polymorphism [OR and 95% CI, 1.10 (1.04, 1.16), P < 0.01, Figure 4]. The ethnic-stratified analysis demonstrated that FTO polymorphisms significantly augmented the OA risk in European Caucasian, with Asian as the exception, which was consistent with the overall population (Figure 4).
Sensitivity analysis and publication bias
The leave-one-out analysis in all populations revealed that no single study changed pooled ORs (Supplementary Table 3), indicating the statistical robustness of our results. Since only six studies were included, we would not carry out the publication bias.
Discussion
To our best knowledge, this is the first meta-analysis that incorporated FTO gene expression data to evaluate the association between FTO gene polymorphism and OA susceptibility. Our results revealed that FTO polymorphism-induced OA risk increase was significant in European Caucasian but not in Asian populations, which is consistent with the results of FTO exhibiting significant differential expressions in the UK population but not in Chinese population. The association strength of FTO polymorphism and risk of OA attenuated after BMI adjustment in Caucasian population.
Fat mass and obesity-related polymorphism manifests differences between Asian and Caucasian populations. This is possibly owing to the fact that ethnic heritage and geographic localization are major influential factors contributing to genetic polymorphisms, which could render a difference in allele frequency (21). However, this result may also be affected by other confounding factors. The sample size of the Chinese population might not be statistically large enough to reach a convincing conclusion. On the other hand, selection bias in patient enrolment and differences in OA-occurring joint sites could potentially undermine the robustness of the findings. Panoutsopoulou et al. examined the strength of association of rs8044769 with knee or hip OA (adjusted for gender) and detected a distinct association of the FTO variant with knee OA (OR 1.08 [95% CI 1.02–1.14], P = 0.009) rather than hip OA (OR 1.04 [95% CI 0.98–1.11], P = 0.17) in Caucasian populations. For non-weight-bearing joints such as hand and TMJ OA, FTO polymorphism also increased the OA risk (22, 23). These findings require further validation in the future with larger-scale observational studies.
Fat mass and obesity-related is an obesity susceptibility gene, but its mechanism on OA is still controversial. Our results showed that the association signal was fully attenuated after BMI adjustment, insinuating the possibility that the FTO gene exerts its effect on OA through obesity in the Caucasian population. However, in the Asian population, the relationship between FTO gene polymorphism and obesity remained ambiguous. Our results illustrated that there is no solid association between FTO polymorphism and higher BMI in the Chinese population, which is contrary to the result of Chang et al. (24). Consistent herewith, the Japanese studies also failed to demonstrate the association of FTO polymorphism with obesity or BMI in their population (25–27). Since the Asian population, generally, is lighter than the UK and even more the USA one; it is possible the presence of a bias due to this difference in BMI of the different populations. On the other hand, these may be due to the sample selection bias for study subjects or genetic variants in FTO, and further studies are necessary to contemplate the association of FTO with BMI and the risk of OA in the Asian population. Meanwhile, more research needs to fill the gaps in the association between FTO polymorphism and OA risk in the African population. In addition, FTO plays an important role in N6-methyladenosine (m6A) modification. m6A modification affects the stability and function of RNAs through the “writers,” “erasers,” and “readers” proteins (28). Several studies reinforced this concept by proving that METTL3 which is the “writer” of m6A, could limit OA progression by inhibiting m6A expression (29). Herein, FTO, as the “eraser” of m6A, has the ability to remove the m6A modification. As such, FTO should therefore be fully investigated for its role in the onset and progression of OA.
Nevertheless, there are some limitations in this meta-analysis. First, due to limited data, we were unable to conduct further stratification analyses of other potential risk factors, such as age, type of SNPs, BMI, and OA site. On top of that, we could not perform a meta-analysis using a dominant model or recessive model. Second, some studies shared the study subjects of control group, which may lead to bias in the final results albeit the fact that a sensitivity analysis was conducted. Third, our results were predominantly based on unadjusted estimates for confounding factors, which might have affected the final results. Fourth, the exclusive inclusion of articles written in English but no other languages in this study might have introduced selection bias.
In conclusion, this meta-analysis confirms that FTO gene polymorphism increased OA risk. Stratification analysis of ethnicity revealed that the augmented risk of OA due to FTO polymorphism may exert its effect through obesity in the Caucasian population. Further studies with larger sample size are necessary to validate whether FTO gene polymorphisms contribute to OA susceptibility with an emphasis on studying Asian and African populations.
Data availability statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Author contributions
KZ and PS conceived and designed the meta-analysis. KZ, LN, and PS performed the literature search and analyzed the data. KZ wrote the manuscript and XY revised it. GC polished the language. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the Natural Science Foundation of Zhejiang Province, China (Grant No. LQ23H060024) and the Medical Science and Technology Project of Zhejiang Provincial (Grant No. 2022KY555).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmed.2022.1024750/full#supplementary-material
Footnotes
References
2. Chaganti RK, Lane NE. Risk factors for incident osteoarthritis of the hip and knee. Curr Rev Musculoskelet Med. (2011) 4:99–104. doi: 10.1007/s12178-011-9088-5
3. Blüher M. Metabolically healthy obesity. Endocr Rev. (2020) 41:bnaa004. doi: 10.1210/endrev/bnaa004
4. Junker S, Frommer KW, Krumbholz G, Tsiklauri L, Gerstberger R, Rehart S, et al. Expression of adipokines in osteoarthritis osteophytes and their effect on osteoblasts. Matrix Biol. (2017) 62:75–91. doi: 10.1016/j.matbio.2016.11.005
5. Thijssen E, van Caam A, van der Kraan PM. Obesity and osteoarthritis, more than just wear and tear: pivotal roles for inflamed adipose tissue and dyslipidaemia in obesity-induced osteoarthritis. Rheumatology. (2015) 54:588–600. doi: 10.1093/rheumatology/keu464
6. Loos RJ, Yeo GS. The bigger picture of FTO: the first GWAS-identified obesity gene. Nat Rev Endocrinol. (2014) 10:51–61. doi: 10.1038/nrendo.2013.227
7. Zeggini E, Panoutsopoulou K, Southam L, Rayner NW, Day-Williams AG, Lopes MC, et al. Identification of new susceptibility loci for osteoarthritis (arcOGEN): a genome-wide association study. Lancet. (2012) 380:815–23. doi: 10.1016/S0140-6736(12)60681-3
8. Elliott KS, Chapman K, Day-Williams A, Panoutsopoulou K, Southam L, Lindgren CM, et al. Evaluation of the genetic overlap between osteoarthritis with body mass index and height using genome-wide association scan data. Ann Rheum Dis. (2013) 72:935–41. doi: 10.1136/annrheumdis-2012-202081
9. Wang Y, Chu M, Rong J, Xing B, Zhu L, Zhao Y, et al. No association of the single nucleotide polymorphism rs8044769 in the fat mass and obesity-associated gene with knee osteoarthritis risk and body mass index: a population-based study in China. Bone Joint Res. (2016) 5:169–74. doi: 10.1302/2046-3758.55.2000589
10. Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. (2021) 372:n71. doi: 10.1136/bmj.n71
11. Davis S, Meltzer PS. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics. (2007) 23:1846–7. doi: 10.1093/bioinformatics/btm254
12. Robinson MD, McCarthy DJ, Smyth GK. EdgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. (2010) 26:139–40. doi: 10.1093/bioinformatics/btp616
13. Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. (2014) 15:550. doi: 10.1186/s13059-014-0550-8
14. Higgins JP, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. (2002) 21:1539–58. doi: 10.1002/sim.1186
15. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. (1997) 315:629–34. doi: 10.1136/bmj.315.7109.629
16. Dai J, Ying P, Shi D, Hou H, Sun Y, Xu Z, et al. FTO variant is not associated with osteoarthritis in the Chinese Han population: replication study for a genome-wide association study identified risk loci. J Orthop Surg Res. (2018) 13:65. doi: 10.1186/s13018-018-0769-2
17. Li Y, Liu F, Xu X, Zhang H, Lu M, Gao W, et al. A novel variant near LSP1P3 is associated with knee osteoarthritis in the Chinese population. Clin Rheumatol. (2020) 39:2393–8. doi: 10.1007/s10067-020-04995-8
18. Welling M, Hämäläinen S, Ojajärvi A, Hirvonen A, Heliövaara M, Leino-Arjas P, et al. Knee osteoarthritis genetics in finnish health 2000 survey. Osteoarthr Cartil. (2014) 22:S237–8. doi: 10.1016/j.joca.2014.02.461
19. Panoutsopoulou K, Metrustry S, Doherty SA, Laslett LL, Maciewicz RA, Hart DJ, et al. The effect of FTO variation on increased osteoarthritis risk is mediated through body mass index: a Mendelian randomisation study. Ann Rheum Dis. (2014) 73:2082–6. doi: 10.1136/annrheumdis-2013-203772
20. Yau MS, Yerges-Armstrong LM, Liu Y, Lewis CE, Duggan DJ, Renner JB, et al. Genome-Wide association study of radiographic knee osteoarthritis in North American caucasians. Arthritis Rheumatol. (2017) 69:343–51. doi: 10.1002/art.39932
21. Loughlin J. Genetic contribution to osteoarthritis development: current state of evidence. Curr Opin Rheumatol. (2015) 27:284–8. doi: 10.1097/BOR.0000000000000171
22. Hämäläinen S, Solovieva S, Vehmas T, Leino-Arjas P, Hirvonen A. Adipose tissue associated genes in hand osteoarthritis in finnish women. Osteoarthr Cartil. (2014) 22:S237. doi: 10.1016/j.joca.2014.02.459
23. Takaoka R, Kuyama K, Yatani H, Ishigaki S, Kayashima H, Koishi Y, et al. Involvement of an FTO gene polymorphism in the temporomandibular joint osteoarthritis. Clin Oral Investig. (2021) 26:2965–73. doi: 10.1007/s00784-021-04278-9
24. Chang YC, Liu PH, Lee WJ, Chang TJ, Jiang YD, Li HY, et al. Common variation in the fat mass and obesity-associated (FTO) gene confers risk of obesity and modulates BMI in the Chinese population. Diabetes. (2008) 57:2245–52. doi: 10.2337/db08-0377
25. Horikoshi M, Hara K, Ito C, Shojima N, Nagai R, Ueki K, et al. Variations in the HHEX gene are associated with increased risk of type 2 diabetes in the Japanese population. Diabetologia. (2007) 50:2461–6. doi: 10.1007/s00125-007-0827-5
26. Omori S, Tanaka Y, Takahashi A, Hirose H, Kashiwagi A, Kaku K, et al. Association of CDKAL1, IGF2BP2, CDKN2A/B, HHEX, SLC30A8, and KCNJ11 with susceptibility to type 2 diabetes in a Japanese population. Diabetes. (2008) 57:791–5. doi: 10.2337/db07-0979
27. Hotta K, Nakata Y, Matsuo T, Kamohara S, Kotani K, Komatsu R, et al. Variations in the FTO gene are associated with severe obesity in the Japanese. J Hum Genet. (2008) 53:546–53. doi: 10.1007/s10038-008-0283-1
28. Chen XY, Zhang J, Zhu JS. The role of m(6)A RNA methylation in human cancer. Mol Cancer. (2019) 18:103. doi: 10.1186/s12943-019-1033-z
Keywords: osteoarthritis, FTO, polymorphism, meta-analysis, systematic review
Citation: Zhao K, Nie L, Chin GMJ, Ye X and Sun P (2022) Association between fat mass and obesity-related variant and osteoarthritis risk: Integrated meta-analysis with bioinformatics. Front. Med. 9:1024750. doi: 10.3389/fmed.2022.1024750
Received: 22 August 2022; Accepted: 02 September 2022;
Published: 23 September 2022.
Edited by:
Elisa Belluzzi, University of Padua, ItalyReviewed by:
Alessia Faggian, University of Trento, ItalySwetha Sunkar, Sathyabama Institute of Science and Technology, India
Copyright © 2022 Zhao, Nie, Chin, Ye and Sun. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Peng Sun, c3AxMjBAMTYzLmNvbQ==
 Kun Zhao
Kun Zhao Liuyan Nie
Liuyan Nie Grace Min Jun Chin3
Grace Min Jun Chin3 Peng Sun
Peng Sun