- 1Centre for Microbial Ecology and Genomics, University of Pretoria, Pretoria, South Africa
- 2Department of Microbiology, University of Pretoria, Pretoria, South Africa
- 3Forestry and Agricultural Biotechnology Institute, University of Pretoria, Pretoria, South Africa
- 4Bioinformatics and Systems Biology, Justus Liebig University Giessen, Giessen, Germany
- 5Environmental Genomics and Systems Biology Research Group, Institute of Natural Resource Sciences, Zürich University of Applied Sciences, Wädenswil, Switzerland
- 6Department of Genetics, University of Pretoria, Pretoria, South Africa
Pantoea ananatis is a highly versatile enterobacterium isolated from diverse environmental sources. The ecological diversity of this species may be attributed, in part, to the acquisition of mobile genetic elements. One such element is an Integrative and Conjugative Element (ICE). By means of in silico analyses the ICE elements belonging to a novel family, ICEPan, were identified in the genome sequences of five P. ananatis strains and characterized. PCR screening showed that ICEPan is prevalent among P. ananatis strains isolated from different environmental sources and geographic locations. Members of the ICEPan family share a common origin with ICEs of other enterobacteria, as well as conjugative plasmids of Erwinia spp. Aside from core modules for ICEPan integration, maintenance and dissemination, the ICEPan contain extensive non-conserved islands coding for proteins that may contribute toward various phenotypes such as stress response and antibiosis, and the highly diverse ICEPan thus plays a major role in the diversification of P. ananatis. An island is furthermore integrated within an ICEPan DNA repair-encoding locus umuDC and we postulate its role in stress-induced dissemination and/or expression of the genes on this island.
Introduction
Pantoea ananatis is a ubiquitous and versatile enterobacterial species, with strains isolated globally from a wide range of environmental sources. Most commonly isolated from plants, P. ananatis has been identified as the causative agent of diseases on a wide range of host plants, including agronomically important crops such as rice, corn, onion, and Eucalyptus (Coutinho and Venter, 2009). Other isolates represent non-pathogenic endo- or epiphytes, while a plant growth promoting P. ananatis strain has recently been characterized and patented (Coutinho and Venter, 2009; Kim et al., 2012). The potential of P. ananatis as an effective biological control agent of a number of phytopathogenic bacteria and fungi is also being investigated (Gasser et al., 2012). Furthermore, P. ananatis has been associated with human disease (De Baere et al., 2004). The wide ecological distribution and versatile lifestyles of P. ananatis suggests that this bacterial species has undergone extensive genetic adaptation in order to effectively occupy and exploit its various ecological niches.
A key driver of rapid bacterial adaptation is the acquisition of genetic material through the horizontal exchange of mobile genetic elements. These elements, which include plasmids, phages, transposons and Integrative and Conjugative Elements (ICEs), influence bacterial fitness and allow microorganisms to occupy novel niches (Wozniak and Waldor, 2010; Aminov, 2011). ICE elements are a class of self-transmissible integrative elements found in numerous Gram-positive and Gram-negative bacterial taxa (Bi et al., 2012). At present, only a limited number of ICEs have been classified into ICE families, including the well-known SXT/R391 family found in Vibrio and Providencia spp. (Wozniak and Waldor, 2010). With the exponential increase of available genome sequences, it can be envisaged that many novel integrative and conjugative elements will be identified and novel ICE families described.
Typically, ICEs comprise of three core modules required for functioning of the element. The first module includes an integrase gene (xerC) which ensures the site-specific chromosomal integration of the ICE as well as effective excision of the element, where it may be aided by an excisionase or recombination directionality factor (Burrus et al., 2006; Wozniak and Waldor, 2010). Following excision, the ICE forms a circular extrachromosomal element through recombination between identical sequences at both ends (Burrus et al., 2006). The second core module is involved in the conjugative transfer of the circularized ICE. This module generally comprises a Type IV secretion system (T4SS), which ensures intimate contact between the donor and recipient for dissemination of the ICE (Wozniak and Waldor, 2010). The third core module is involved in ICE maintenance and includes regulatory proteins as well as toxin–antitoxin and partition systems which ensure that the ICE is vertically transmitted within a bacterial lineage (Wozniak and Waldor, 2009). Aside from core modules, ICEs contain an extensive array of cargo genes, which may contribute to diverse phenotypes (Wozniak and Waldor, 2010). These include genes coding for factors involved in pathogenesis, metabolic adaptation, the production of secondary metabolites and resistance to antibiotics and heavy metals (Burrus et al., 2002). Homologous recombination between ICEs may also occur in ICE recipients, leading to the formation of hybrid ICEs, which contribute to the diversity of ICE elements and potential accessory factors they encode (Wozniak and Waldor, 2010).
Previous comparative genomic analyses (De Maayer et al., 2014) revealed the presence of a large genomic island in five of eight compared P. ananatis genomes. Here we have characterized these genomic islands and show that they represent ICE elements, which are prevalent among strains of the species. The ICEPan elements comprise core modules interspersed with divergent cargo regions. Proteins encoded in these cargo regions may contribute to stress-response and production of antibiotic secondary metabolites. Finally, we identified a non-conserved island situated within the umuC gene involved in DNA damage repair and postulate its potential role in the dissemination or expression of the cargo genes.
Methods
In Silico Characterization of the ICEPan Elements
ICE elements were identified on the genomes of five P. ananatis strains (Table 1) by localized tBlastN analysis with protein coding sequences from known ICE element core modules using BioEdit v 7.1.11 (Hall, 1999). The complete genome sequences of P. ananatis AJ13355 (NC_017531.1), LMG5342 (NC_016816.1) and PA13 (CP003085.1), as well as the draft genomes of P. ananatis B1-9 (CAEI00000000) and BD442 (JMJL00000000) are publically available on the NCBI database under the given accession numbers. The full extent of the ICEPan elements was elucidated by BlastN with the full tRNA-Phe nucleotide sequence of the ICE-negative P. ananatis LMG20103 (PANA_t0061) (De Maayer et al., 2010) to determine the ICEPan integration site. The sizes and G+C contents of the ICEPan Island (IR) and Core (CR) regions and other sequence manipulations, such as sequence alignments and localized Blast comparisons were performed using BioEdit (Hall, 1999). The CDS sets encoded on the ICEPan elements were standardized using the FgenesB ORF prediction server (http://www.softberry.com) and reciprocal best hit (RBH) BlastP analysis (Altschul et al., 1990; Moreno-Hagelsieb and Latimer, 2008). Core module CDSs in ICEPan elements were predicted by localized BlastP of the protein products with orthologs from the ICEberg server (Bi et al., 2012; http://db-mml.sjtu.edu.cn/ICEberg/). The core module CDSs were further used to identify ICE-like elements in closely related organisms by BlastP comparison against the NCBI non-redundant protein sequences (nr) database. Average amino acid identities were calculated on the basis of localized RBH BlastP analyses (Moreno-Hagelsieb and Latimer, 2008), where the sum of the number of identities was divided by the sum of the total lengths of the aligned regions. Orthology of the CDSs encoded on the ICEPan IR regions to proteins encoded on other genomes was determined by BlastP comparison of the translated protein products against the NCBI nr database, while conserved domains were predicted using the Batch CD-search tool and the Conserved Domain Database (Marchler-Bauer and Bryant, 2004; Marchler-Bauer et al., 2009).
Phylogenetic Tree Construction
Phylogenetic trees were constructed, using the amino acid sequences of the chromosomal house-keeping markers AtpD, GyrB, InfB, and RpoB; ICEPan core CDSs and enterobacterial ICE core CDSs (ICEEpirCFBP5888, ICEEclATCC13047, YAPI, Ctnscr94, and ICEPwaWPP163); ICEPan core CDSs and conjugative plasmid core CDSs (pEI70 and pEb102) and umuDC island CDSs. The amino acid sequences were concatenated and aligned using the MAFFT web server (Katoh and Standley, 2013). Tree construction was performed using MEGA v 5.2 (Tamura et al., 2011) with the Neighbor-joining approach, with complete gap deletion, Poisson correction and bootstrapping (n = 1000).
PCR Screening for ICEPan Elements in P. ananatis Strains
Forty-six P. ananatis strains of different geographic origins and sources of isolation (Table 2) were incubated overnight at 28°C and maintained on Luria Bertani (LB) agar. Genomic DNA was extracted using the ZR Fungal/Bacterial DNA Microprep™ kit (Zymo Research Corporation, California, USA), as per the manufacturer's instructions. Strains were confirmed as belonging to the species P. ananatis by PCR amplification and Sanger sequencing of the 16S rRNA gene. The chromosomal DNAs were PCR amplified using the following primer sets: sit1F-sit1R and sit2F-sit2R, covering the hyp-tRNA-Phe-yjdC region and hyp-tRNA-Phe-rbsU regions of ICEPan-negative P. ananatis strains for which genome sequences are available, respectively; xerC1F-xerC1R and xerC2F-xerC2R, within the XerC1 and XerC2 integrases of site 1 and site 2-integrated ICEPan elements, respectively; topBF-topBR, pilVF-pilVR, and traIF-traIR, designed on the nucleotide sequences of topB, pilV, and traI core module genes, respectively; mae1F-mae1R and ssfDF-ssfDR, designed from the nucleotide sequences of ssfD in IR-4A and mae1 in IR-4B, respectively (Table S1).
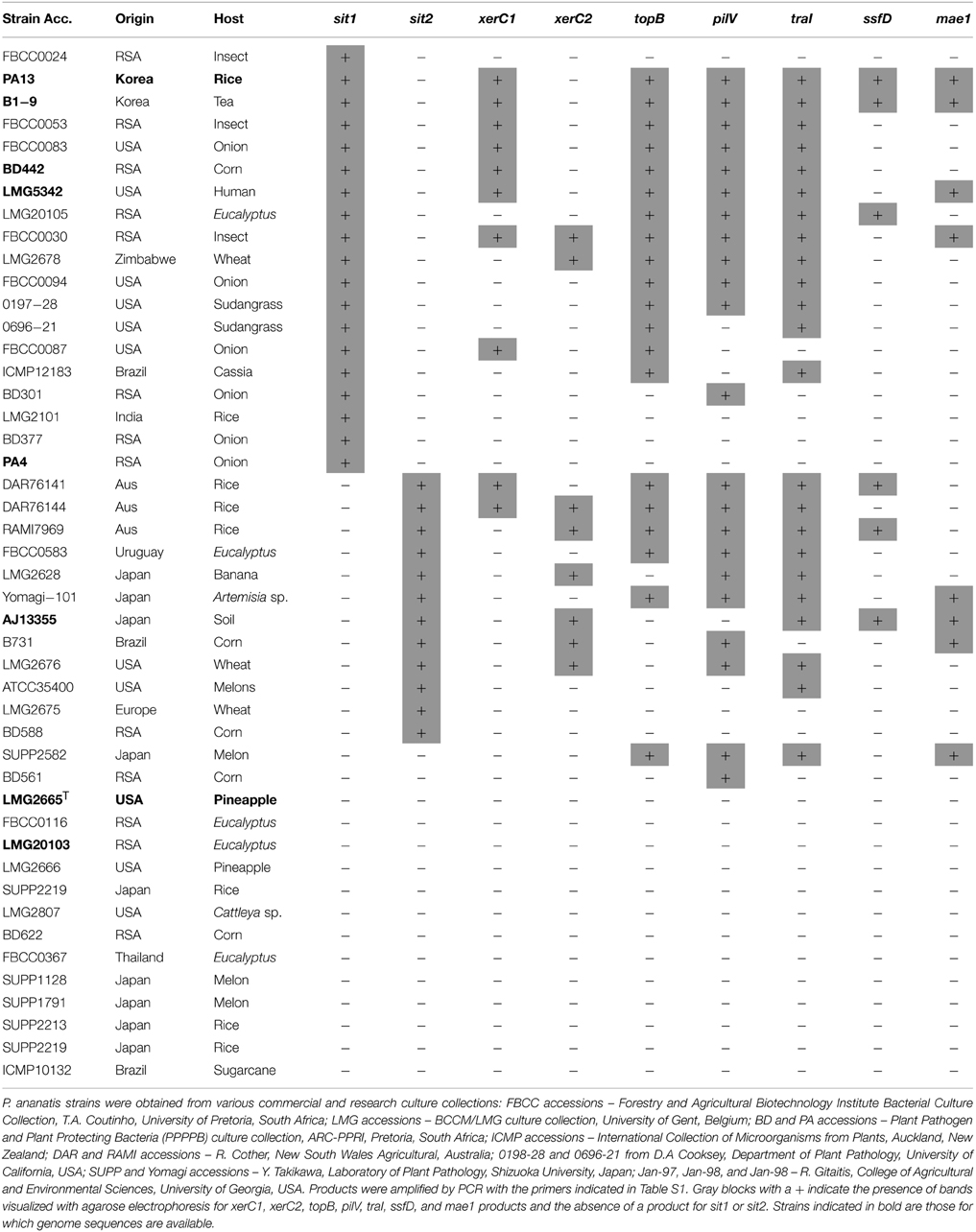
Table 2. The presence or absence of ICEPan in P. ananatis strain isolated from different ecological sources.
PCR amplification was undertaken with a standardized PCR program (94°C for 5 min; 30× {94°C for 1 min, 55°C for 1 min, 72°C for 1 min}; 72°C for 5 min), and the resultant amplicons were visualized, after agarose gel electrophoresis with GelRed™ nucleic acids stain (Biotium, California, USA), using a UV transilluminator. The presence of an insert within tRNA-Phe site 1 or site 2 was confirmed by the absence of a band for Sit1 and/or Sit2, while the presence of an ICEPan element in these sites was determined on the basis of the presence of bands for the xerC1, xerC2, topB, pilV, and traI gene products. The presence of a putative antibiotic biosynthetic locus and the mae1-containing umuDC island was confirmed through the presence of a band for the ssfD and mae1 gene products, respectively.
Results and Discussion
General Properties of the P. ananatis Integrative Conjugative Elements
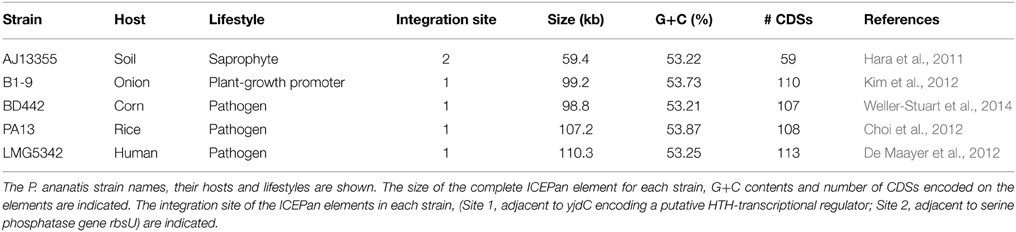
Analysis of eight P. ananatis genomes revealed the presence of integrative and conjugative elements integrated on the chromosomes of five strains (Table 1; Figure 1). These elements belong to a novel family, ICEPan, named in accordance with the nomenclatural system proposed by Burrus et al. (2002), to reflect the species, P. ananatis in which they were identified, and the strain numbers to distinguish between the different ICEPan elements. The ICEPan elements vary in size from 67.4 (ICEPanAJ13355) to 110 kb (ICEPanLMG5342) and carry between 59 and 113 protein coding sequences (CDSs). The G+C contents of the ICEs (53.2–53.9%) are similar to those of the rest of the chromosome (53.5–53.7% for the complete chromosomes of AJ13355, LMG5342, and PA13). As is the case for the majority of genomic islands, it has been observed that most ICEs are integrated within tRNA genes, which serve as hotspots for recombination (Burrus et al., 2002; Boyd et al., 2009). All five ICEPan elements are integrated into one of two copies of identical phenylalanine-specific tRNAs, pheU/V (Table 1) present on all the genomes. Similarly, the ICEs CTnscr94 of Salmonella enterica subsp. enterica serovar Senftenberg 5494-57, YAPI of Yersinia pseudotuberculosis 32777 and ICEEc2 of Escherichia coli BEN374 (Hochhut et al., 1997; Collyn et al., 2002; Roche et al., 2010) are integrated into tRNA-Phe genes. It has been observed that ICE integrases may have evolved to drive ICE insertion within specific tRNA genes. This targeted integration may ensure that essential loci and highly expressed tRNAs, where ICE integration may affect cellular fitness, are avoided (Boyd et al., 2009). While four of the ICEPan elements were integrated in one tRNA-Phe site (Site 1; adjacent to yjdC encoding a putative HTH-transcriptional regulator), the ICEPanAJ13355 was integrated within the second identical tRNA-Phe gene copy (Site 2; adjacent to serine phosphatase gene rbsU). The ICE-associated integrase (XerC1) of the four ICEPan elements integrated in the first site shared 99.4–100% amino acid identity among them, while the ICEPanAJ13355 integrase (XerC2) shares lower sequence identity (58.5% average amino acid identity) with XerC1. Variability in the integrase sequence may thus dictate the ICEPan integration site.
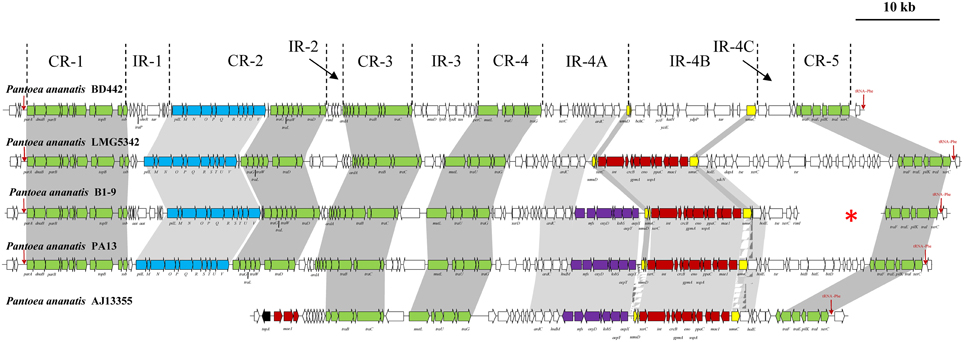
Figure 1. Schematic diagram of the P. ananatis integrative conjugative elements ICEPan. Core regions CR1-5 and island regions (IR1-4) are indicated. Green arrows indicate the conserved genes in the CRs, blue arrows represent the pilLMNOPQRSTUV locus, red arrows the conserved IR-4B genes in ICEPanAJ13355, ICEPanB1-9, ICEPanLMG5342 and ICEPanPA13, while the putative antibiotic biosynthetic locus is indicated by purple arrows. The red arrows at the 5′ and 3′ ends indicate the tRNA-Phe integration site for ICEPan, while the transposase flanking the ICEPanAJ13355 is represented by a black arrow. The red star shows the non-contiguous ends of the contigs that make up ICEPanB1-9.
The ICEPan Elements Contain Core Modules for Integration, Maintenance, and Conjugative Transfer
Comparison of the ICEPan elements of the five strains revealed that they comprise of four (ICEPanAJ13355) or five (ICEPanB1-9, ICEPanBD442, ICEPanLMG5342, and ICEPanPA13) conserved syntenic blocks, CR 1–5, with regions containing non-conserved genes, IR 1–4, interspersed between these blocks (Figure 1). The protein complements of the ICEPan CRs were compared to characterized ICEs to identify the three core functional modules within the CR blocks of the ICEPan elements.
The tyrosine recombinase (XerC), which drives both the integration and excision of various ICEs including those of the SXT/R391 family (Boyd et al., 2009), is localized in CR-5 of the ICEPan. A large number of proteins with putative roles in the conjugative transfer of ICEPan are interspersed within the CRs. These include orthologs of the relaxase TraI (CR-5), Type IV coupling factor TraG (CR-4), conjugative transfer ATPase TraC (CR-3) and putative conjugative transfer proteins TraE and TraF (CR-5). Located within CR-2 are genes encoding orthologs of another conjugation coupling factor TraD, exported transglycosylase TraL and putative conjugative transfer protein TraW. CR-2 also includes a ~11 kb locus encoding 11 proteins, PilL-V, sharing 50 and 46.2% average amino acid identity with the Pil proteins of the ICEs CTnscr94 in S. enterica (CAX68107.1-CAX68117.1) and YAPI in Y. pseudotuberculosis (CAF28485.1-CAF28494.1), respectively (Figure 1). The encoded type IV pilus was initially thought to function as a virulence factor, with deletion of the Y. pseudotuberculosis pil locus resulting in decreased pathogenicity in mice which were infected orally (Collyn et al., 2002). However, more recent analysis of E. coli ICEEc2 has shown that this pilus plays a role in mating pair formation and conjugal transfer, and the decreased pathogenicity of the Y. pseudotuberculosis pil mutant may rather result from accessory elements on YAPI (Collyn et al., 2006; Roche et al., 2010). The chromosome partitioning protein ParA/Soj and plasmid maintenance protein ParB have been demonstrated to play a role in the maintenance and vertical transmission of the excised ICE (Wozniak and Waldor, 2009). Genes encoding orthologs of these proteins are localized within ICEPan CR-1. Our analyses thus revealed that the core modules for integration and excision, maintenance and conjugative dissemination are present in the ICEPan regions of four of the five sequenced ICEPan-containing strains, suggesting that they represent functional and transmissible ICEs. The absence of CR-1 and -2, which carry genes for ICE dissemination in the other strains, suggests that ICEPanAJ13355 has lost its functionality as a transmissible element. The absence of CR-1 is puzzling, as this region is involved in the vertical maintenance of the ICE element during cell division and replication. One possibility is that the distinct chromosomal integration site and/or the presence of integrase XerC2 within the ICEPanAJ13355 prevent its excision. In its un-excised form it may then be replicated as part of the chromosome.
The ICEPan Family Elements are Closely Related to other Enterobacterial ICEs and Share a Common Origin with Conjugative Plasmids of Erwinia amylovora and Erwinia billingiae
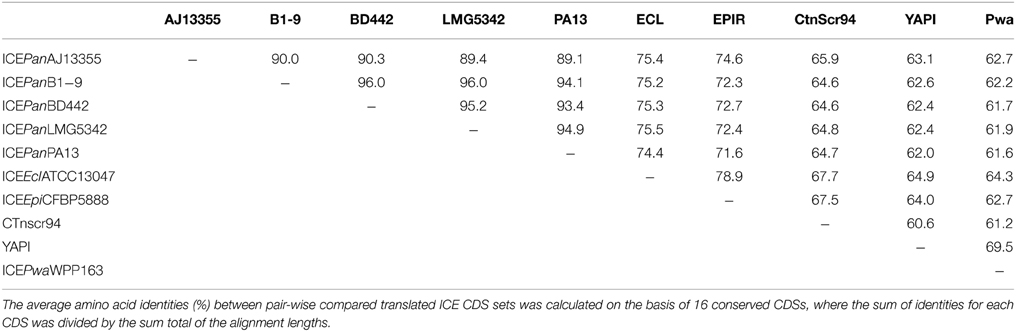
Nineteen CDSs conserved in all five ICEPan elements showed significant sequence identity to proteins encoded on various characterized enterobacterial ICEs, including Ctnscr94 of S. enterica subsp. enterica serovar Senftenberg 5494-57 and YAPI of Y. pseudotuberculosis 32777 (Table 3). The conserved ICEPan CDSs showed highest sequence similarity to those encoded on uncharacterized ICEs identified in the genomes of Enterobacter cloacae ATCC13047 (ICEEclATCC13047) and Erwinia piriflorinigrans CFBP5888T (ICEEpiCFBP5888) (Table 3) (Ren et al., 2010; Smits et al., 2013). Alignment of the ICE loci (Figure 2) also indicated a high level of synteny between ICEPan and the enterobacterial ICEs. A phylogeny based on the concatenated protein products of the conserved ICE CDSs showed similar clustering to a phylogeny constructed on the basis of the concatenated amino acid sequences of four chromosomal house-keeping markers (Figure 3). This suggests that ICEPan and the compared enterobacterial ICEs were derived from a common ancestor, and have undergone subsequent divergence in parallel with the bacterial chromosome.
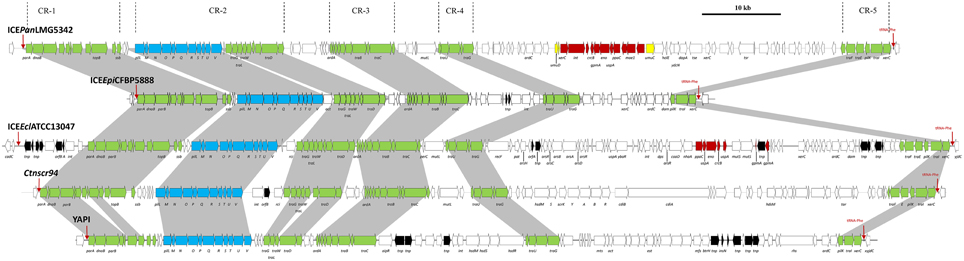
Figure 2. Schematic diagram of ICEPanLMG5342 and closely related ICEs showing the extensive synteny that exists between the CRs. ICEEpiCFBP5888 and ICEEclATCC13047 were identified from the genome sequences of E. piriflorinigrans CFBP 5888T and E. cloacae subsp. cloacae ATCC13047T, respectively. The Genbank files for CTnscr94 of S. enterica subsp. enterica serovar Senftenberg 5494-57 and YAPI of Y. pseudotuberculosis 32777 were obtained from the ICEberg database (Bi et al., 2012). Dark gray shaded regions and green arrows indicate the conserved CRs between the ICEs. Black arrows represent transposases.
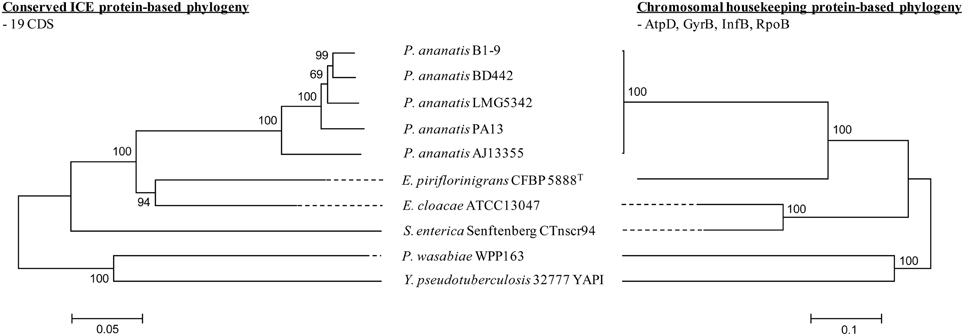
Figure 3. Phylogeny of ICEPan and related ICEs and comparison to chromosomal house-keeping marker phylogeny. Alignments were done with MAFFT (Katoh and Standley, 2013) using the concatenated amino acid sequences of the ICE elements (19 conserved CDSs) and house-keeping markers (AtpD, GyrB, InfB, and RpoB). Phylogenies were constructed using MEGA v 5.2 (Tamura et al., 2011), using the Neighbor-joining algorithm with complete gap deletion, Poisson correction and bootstrap analysis (n = 1000).
BlastP analyses of the translated ICEPan CDSs against the NCBI non-redundant (nr) protein database revealed a number of orthologous CDSs occurring in the conjugative plasmids of E. amylovora ACW56400 (pEI70 – NC_018999.1; ~65 kb) and E. billingiae Eb661 (pEB102 – NC_014304.1; ~102 kb) (Figure 4) (Kube et al., 2010; Llop et al., 2011). Between 36 and 54 of the 114 proteins encoded on pEB102 (60.3% average amino acid identity) and 35–53 of the 70 proteins encoded on pEI70 (60.1% average amino acid identity) shared orthology with proteins encoded on the ICEPan elements. The lower number of orthologous CDSs shared between pEI70/pEb102 and ICEPanAJ13355 is due to the absence of CR-1 and -2 from the latter element. Among the conserved plasmid/ICE CDSs are those in the maintenance (ParAB) and conjugative transfer (TraDEFGILW) modules. With the exception of a shared PilL ortholog, no other orthologs of proteins encoded in the ICEPan pil locus are present in pEb102 or pEI70, suggesting that Pil pilus-associated conjugative transfer is specific to ICE elements. Also absent from the conjugative plasmids is an ortholog of the integration and excision module protein XerC indicative of pEI70 and pEb102 being maintained as plasmids rather than being chromosomally integrated. The presence of a number of orthologous CDSs within the ICEPan loci and the plasmid pEI70 and pEb102 and the presence of these orthologs in syntenous blocks suggest a common origin for the ICEPan family elements and the Erwinia plasmids (Figure 4). A phylogeny constructed on the basis of the amino acid sequences of 21 CDSs conserved in pEI70, pEb102, all five ICEPan elements, ICEEclATCC13047 and ICEEpiCFBP5888 showed that ICEPan is more closely related to the enterobacterial ICE elements than pEb102 and pEI70 (Supplementary Figure S1). This is incongruent with the phylogeny on the basis of the house-keeping markers, which thus suggests that the common ancestor of the ICE/plasmids pre-dates species divergence.
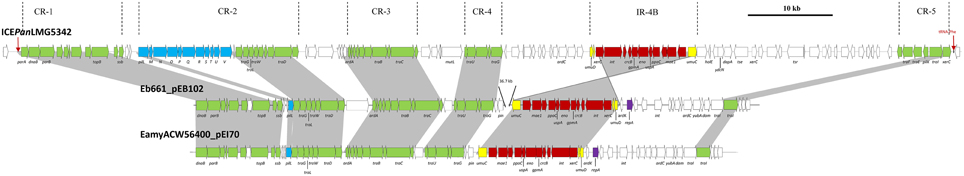
Figure 4. Schematic diagram of ICEPanLMG5342 and the phylogenetically related conjugative plasmids pEI70 of E. amylovora ACW56400 and pEb102 of E. billingiae Eb661. ICEPanLMG5342 is aligned with the plasmids pEI70 of E. amylovora ATCC56400 (NC_018999.1) and pEb102 of E. billingiae Eb661 (NC_014304.1). Dark gray shaded regions and green arrows indicate the conserved CRs between the ICEs. The conserved umuCD island is marked by red arrows, with the umuC fragments and umuD genes colored in yellow, and the plasmid replication gene repA in pEI70 and pEb102 is colored purple.
ICEPan Elements are Prevalent Among P. Ananatis Strains Isolated from Different Sources
The prevalence of ICEs in P. ananatis was determined through PCR amplification with primers designed on the basis of conserved ICE genes and integration sites (Table S1). Initially, two sets of primers sit1F/R and sit2F/R were designed to cover the two genomic tRNA-Phe regions on the basis of conserved flanking sequences in strains which did not have an insertion at these sites. No amplicons were observed in 31 out of 46 strains (67.4% of strains tested), indicating that a large insert is integrated at one or both of the tRNA-Phe sites. Of these, 11 insertions occur in the tRNA-Phe site occupied by ICEPanB1-9, ICEPanBD442, ICEPanLMG5342 and ICEPanPA13 (sit1), nine solely in the tRNA-Phe site of ICEPanAJ13355 (sit2) while 11 strains had large inserts integrated at both sites (Table 2). This would suggest that in the latter strains, a third functional copy of the tRNA-Phe gene exists, or that the tRNA-Phe gene(s) is located at a different site. As these primers only give an indication of the presence of an insertion at a given tRNA-Phe site, further primers were designed on the basis of conserved genes in the ICEPan elements.
PCR amplification with primers designed on the conserved sequences of the ICEPanB1-9, ICEPanBD442, ICEPanLMG5342, and ICEPanPA13 integrase (xerC1F/R) produced amplicons in six additional strains, while those based on the ICEPanAJ13355 integrase (xerC2F/R) amplified products in six more strains. Further primers designed on the basis of the CR genes coding for topisomerase B (topB), type IV pilus shufflon (pilV) and relaxase/helicase (traI) amplified products in 22, 23, and 20 strains, respectively (Table 2). There was some variability in the presence of the conserved ICE genes among the strains. For example, nine strains were positive for one or more of the traI, pilV, topB gene fragments, while they were negative for both the XerC1 and XerC2 integrase primer sets. Similarly, while ICEPanAJ13355 was missing the core regions containing the topB (CR-1) and pilV (CR-2) genes, products were obtained for either or both of the gene fragments in six strains which were positive for the ICEPanAJ13355 integrase XerC2. In addition to the variability observed among ICEPan elements of the sequenced strains, there thus appears to be further variability among the ICE elements of other P. ananatis strains. Variability in the conserved sequences, as was observed for the distinct integrases, may also result in no amplification occurring for conserved ICE genes among P. ananatis strains. Nevertheless, 24 out of 46 (52% of tested strains) P. ananatis strains were positive for two or more of the conserved ICE genes tested (Table 2), which suggests that integrative conjugative elements are relatively common among strains of the species.
ICEPan Elements Carry Extensive Non-conserved Cargo Genes with a Potential Role in Fitness, Stress Response, and Antibiosis
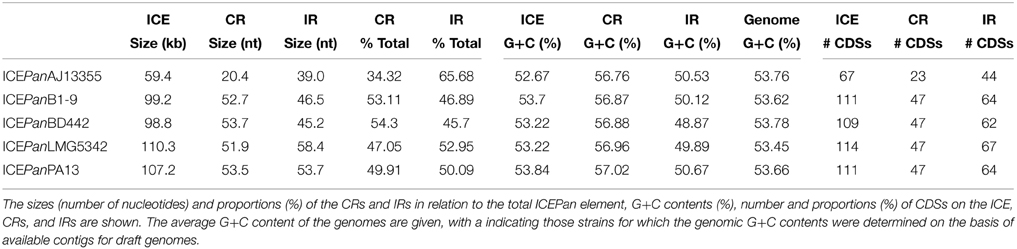
The ICEPan elements of all five strains contain four large non-conserved cargo regions, IR 1–4 (Figure 1). IR4 has further been subdivided into IR-4A and IR-4C, separated by a region flanked by the umuD and partial umuC genes, IR-4B, which is discussed in further detail below. The IRs contribute between 45.7% (ICEPanBD442) and 65.7% (ICEPanAJ13355) of the total ICEPan size (Table 4). The G+C contents of the ICE cargo and core regions were determined. This showed that while the G+C contents of ICEPan (average G+C content: 53.3%) is similar to that of the chromosomes (53.7%), the G+C content of the core regions (56.9%) is somewhat higher and that of the IRs is lower (50.0%) than that of the chromosome (Table 4). This suggests that the current ICEPan structures may have arisen through distinct horizontal acquisition events of the core and island regions. Between 44 (ICEPanAJ13355) and 67 (ICEPanLMG5342) CDSs are encoded in IR1-4. Our previous pan-genome comparison of eight P. ananatis strains revealed a sizeable accessory genome (1690 CDS—30.4% of the pan-genome) for the species (De Maayer et al., 2014). A total of 124 distinct CDSs are encoded in the IRs of the ICEPan elements, suggesting this genomic element contributes substantially to the accessory fraction of the pan-genome and that ICEPan plays a major role in the diversification of P. ananatis.
Several genes within the ICEPan IRs encode orthologs of characterized stress response mechanisms (Table S2). One means of stress response utilized by Gram-negative bacteria is through the production and activation of alternative RNA polymerase σ factors. These regulate the transcription of a large number of different genes that allow the cell to tolerate or counteract various stresses including starvation, osmotic stress, and oxidative and DNA damage (Bougdour and Gottesman, 2007). It is imperative that σ factor concentrations, activity and stability are tightly regulated. Several σ factor stabilizing proteins have been identified including the ATP-dependent protein ClpXP and the anti-adaptor protein IraP which stabilizes σ factor during phosphate starvation (Bougdour and Gottesman, 2007). Genes within the IR-4A of ICEPanPA13 and ICEPanBD442 encode orthologs of ClpXP, while an IraP ortholog is encoded in IR-1 of ICEPanBD442. Another means of stress response is through the expression of Universal Stress Proteins (Usp). UspA of E. coli is produced in response to starvation, osmotic and heat shock, as well as exposure to heavy metals and antimicrobial agents (Kvint et al., 2003). Orthologs of UspA are encoded within the IR-4B of all ICEPan elements with the exception of ICEPanBD442. Bacteria associated with plants are frequently exposed to reactive oxygen species such as superoxide anions and hydroperoxides which form an integral part of plant defense responses. A gene localized within the IR-1 of ICEPanBD442 encodes an ortholog of OhrR, a sensor and regulator of organic hydroperoxide resistance (Panmanee et al., 2006). Within IR-4B of this same strain a gene encoding an ortholog of the σS-regulated manganese-catalase KatN, which has been demonstrated to degrade hydroperoxides into water and oxygen in S. enterica, is present (Robbe-Saule et al., 2001). As is the case for the katN gene in S. enterica, the ICEPanBD442 gene is localized adjacent to genes showing extensive sequence identity to yciEFG, which are likewise thought to play a role in oxidative stress response (Hindupur et al., 2006). Upstream of the ICEPanBD442 katN gene, a gene encoding an ortholog of the oxidoreductase YdeP involved in acid resistance in Shigella flexneri (Oglesby et al., 2005), is present.
Aside from roles in stress response, several ICEPan CDSs may contribute to the fitness of the ICE carrying strains. A locus within IR-4C of ICEPanPA13 (Pagr_3794-Pagr_3798) encodes five proteins sharing 61.8% average amino acid identity with a predicted iron transport system in Agrobacterium tumefaciens 5A (AT5A_20056.1-AT5A_20076.1). This locus may allow P. ananatis PA13 to compete for this limiting nutrient within the plant host. Also interspersed among the IR-1, -4, and -5 of all ICEPan elements with the exception of ICEPanAJ13355 are six genes encoding five distinct methyl-accepting chemotaxis proteins (MCPs). MCPs sense the presence of certain attractants and repellents to determine the direction of bacterial motility, toward or away from these chemical signals (Baker et al., 2006). ICEPan elements may thus play a role in the spread of MCPs for the response to different chemical stimuli, including those they may encounter in novel environmental niches.
Analyses of the ICEs of several microorganisms have shown the presence of loci for the biosynthesis of secondary metabolites such as siderophores and antibiotics on these elements (Ghinet et al., 2011). The ICEPan sequences were compared against the antiSMASH (antibiotics and secondary metabolite analysis shell) server (Blin et al., 2013). A ~7.8 kb locus encoding seven proteins involved in the biosynthesis of a predicted secondary metabolite is present in the IR-4A regions of ICEPanB1-9, ICEPanPA13 and ICEPanAJ13355 (Figure 1; Table S2). The encoded proteins share 99.3–99.9% average amino acid identity among the three strains. Two proteins encode orthologs of the fosmomycin biosynthetic proteins Fom1 and Fom2 of Streptomyces fradiae (ACG70831.1-ACG70832.1—37.6% average amino acid identity). Fom1 encodes a phosphoenolpyruvate (PEP) mutase which catalyzes the first step of fosmomycin biosynthesis, converting PEP into phosphonopyruvate (PnPy), which is consequently converted by PnPy decarboxylase (Fom2) into phosphonoacetaldehyde (Woodyer et al., 2006). Two further proteins share 48.3% average amino acid identity to amidotransferase SsfD (ADE35421.1) and acyl carrier protein SsfC (ADE35420.1) of Streptomyces sp. SF2575 which are required for the production of the malonamate starter unit of tetracyclines (Pickens et al., 2010). Another protein shows weak sequence identity (34.5% average amino acid identity) to StrU of Streptomyces griseus (CAH94303.1), a dehydrogenase involved in the final processing of streptomycin. Also encoded in ICEPan IR-4A of these three strains is an ortholog of the AfsA protein of Streptomyces virginiae MAF10-06014 (BAA23611.1). AfsA is a diffusible γ-butyrolactone autoregulator which regulates the production of secondary metabolites in Streptomyces spp. (Healy et al., 2009). The presence of this locus in the ICEPan IR-4A region suggests that P. ananatis AJ13355, PA13 and B1-9 are capable of producing a secondary metabolite with a potential role in antibiosis. Amplification with primers designed on the basis of the ssfD nucleotide sequences showed that two additional rice-pathogenic strains from Australia and one Eucalyptus-pathogenic strain from South Africa, may potentially encode this putative secondary metabolite biosynthetic locus (Table 2). Within IR-1 of ICEPanB1-9 are two genes encoding proteins with weak sequence identity (31.3% average amino acid identity) to the alveicin bacteriocin A Aat (CDD cl04134: microcin superfamily) of Hafnia alvei and its cognate immunity protein Aai (CDD pfam01024—colicin pore forming domain). Bacteriocins are bacterial toxins that kill closely related bacteria, while the immunity protein provides self-protection against the toxin (Wertz and Riley, 2004). The presence of bacteriocin and putative antibiotic biosynthetic loci within the IR regions of the ICEPan family elements suggest this element may confer a potential competitive advantage over other bacteria and fungi sharing the same environmental niche and thus contribute to the fitness of ICEPan-positive P. ananatis strains.
A Non-conserved Island is Integrated into the umuDC Locus
Localized within IR-4 of the ICEPan of all five strains are genes encoding proteins with extensive sequence identity to the SOS damage response proteins UmuC and UmuD. In response to DNA damage, as a result of genotoxic stresses such as UV irradiation or exposure to mitomycin C, these two proteins effect error-prone replication across the damaged DNA lesion, also known as SOS mutagenesis (Hare et al., 2006). The presence of UmuDC orthologs further supports a role for ICEPan in stress response. BlastN analysis of the ICEPan IR-4B CDS sets revealed that the umuC gene is disrupted by a non-conserved insertion (Figure 5A). A similar disruption of the umuC gene could be observed on the chromosome of Pantoea vagans C9-1 and the Erwinia plasmids pEb102 and pEI70. This insertion is ~11 kb in size in ICEPanB1-9, ICEPanAJ13355, ICEPanLMG5342, and ICEPanPA13 with an average G+C content of 60.3%, substantially higher than the average G+C% for the ICEPan element (53.3%) and the chromosome (Figure 5A). By contrast the umuC insertion in ICEPanBD442 is ~13.7 kb in size with a G+C content of 49.0%. The amino acid sequences of the disrupted umuC genes were aligned (Figure 5B). This showed that the integron is inserted at amino acid position 88 in the UmuC protein of the Erwinia plasmids pEB102, pEI70 and all ICEPan with the exception of ICEPanBD442. In the latter, the insertion is integrated in UmuC after position 37.
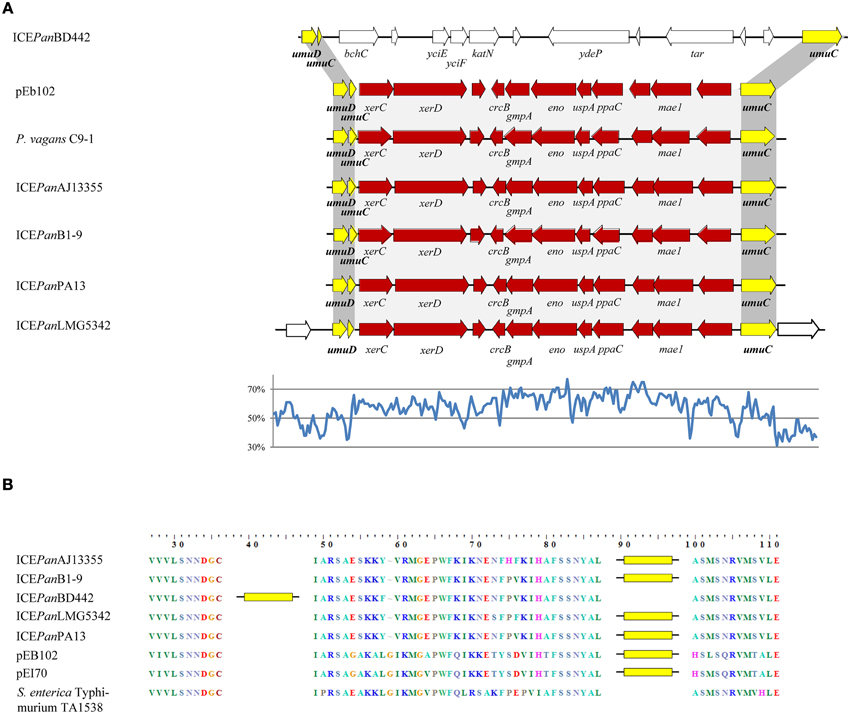
Figure 5. Schematic diagram of the umuDC island in ICEPan and related elements (A) and the integration site within the umuC amino acid sequence (B). (A) The ICEPan umuDC islands are aligned with the umuDC island from the conjugative plasmids of E. amylovora ACW56400 (pEI70) and E. billingiae Eb661 (pEb102) and the chromosomally integrated umuDC island of P. vagans C9-1. A line graph of the G+C contents (%) for ICEPanLMG5342, determined on the basis of G+C contents for 100 nt steps with 50 nt overlap, is shown. (B) The amino acid sequences of the ICEPan UmuC protein were aligned with that of CTnscr94, and the insertions are indicated by yellow bars.
The ICEPanBD442 insertion encodes 11 proteins not encoded in the other ICEPan, while 11 CDSs are conserved among the other four ICEPan (99.2% average amino acid identity) (Table S2). The ICEPanBD442 umuDC island CDSs include the catalase KatN, YciEFG and YdeP orthologs, which may play a role in oxidative and acid stress response as discussed above, as well as a methyl-accepting chemotaxis protein (Figure 5A). Proteins encoded within ICEPanAJ13355, ICEPanB1-9, ICEPanLMG5342, and ICEPanPA13 umuDC region include an ortholog of the Mae1 protein in the yeast Schizosaccharomyces pombe which plays a role in the uptake of the dicarboxylic acid substrates L-malate, succinate, and malonic acid that are central to the tricarboxylic acid (TCA) pathway (Grobler et al., 1995). Also encoded within this island are orthologs of the 2,3-bisphosphoglycerate-independent phosphoglyceratemutase GpmA and enolase Eno which catalyze the reversible conversion of 3-phospho-D-glycerate to PEP, involved in the glycolysis and gluconeogenesis (Nurmohamed et al., 2010). The umuDC island-encoded enzymes thus play a likely role in energy production and conversion. Also included in the region is a gene (ppaC) encoding a manganese-dependent inorganic pyrophosphatase, which removes pyrophosphate generated as a by-product of many biosynthetic and metabolic reactions, as well as a gene encoding the universal stress protein UspA, which is discussed above. Finally, orthologs of the recombinases XerC and XerD are encoded adjacent to the small 5′ fragment of umuC.
The 11 translated CDSs in the ICEPan elements of AJ13355, B1-9, LMG5342, and PA13 share 99.3% average amino acid identity, similar to the average amino acid identity for the core genome CDS sets (99.3%) (De Maayer et al., 2014). By contrast the IR-4B CDSs share much higher average amino acid identity (~92.0%) with orthologs in P. vagans C9-1, E. billingiae Eb661, and E. amylovora ACW56400 than the genomic average (84.8, 76.0, and 76.3%, respectively) (Table S3). Most striking is the comparison of the 11 umuC-inserted CDSs on the chromosome of P. vagans C9-1 and those encoded on pEb102 and pEI70, which share an average amino acid identity of 99.7 and 100%, respectively (Table S3), which is substantially higher than the genomic average amino acid identities (76.4, 77.2, and 81.0% average amino acid identity for P. vagans C9-1 vs. E. billingiae Eb661 and E. amylovora ACW56400, and Eb661 vs. ACW56400, respectively). Alignment of the nucleotide sequences for these regions shows only three nucleotide polymorphisms between the umuDC islands of P. vagans C9-1 and E. amylovora pEI70, while those of C9-1 and E. billingiae Eb661 share 100% nucleotide identity. This suggests very recent inter-genera horizontal exchange of the umuDC island regions between the plasmids of E. amylovora ACW56400 and E. billingiae Eb661, ICEPanAJ13355, ICEPanB1-9, ICEPanLMG5342, and ICEPanPA13 and the chromosome of P. vagans C9-1 (Supplementary Figure S2).
While the functions encoded in the umuDC islands of the ICEPan, P. vagans C9-1 chromosome and Erwinia plasmids, remain to be elucidated, we hypothesize two potential reasons for the integration of this island into the umuC gene. Firstly, as the umuDC is transcriptionally upregulated in response to stress, the insertion into the umuC reading frame may result in the genes on the island being expressed. This hypothesis is supported by observations in Acinetobacter baylyi ADP1, where a truncated umuC is also present and umuDC regulates the expression of DNA-damage inducible gene ddrR, which is located upstream of umuD (Hare et al., 2006). The truncated umuC may thus serve as a DNA-damage inducible regulator for genes on the island which may potentially play a role in stress response. Alternatively, the recombinases encoded by the xerC and xerD genes located directly upstream of the truncated 5′ region of umuC (Figure 5A) may function in a similar fashion to the integration and excision core module tyrosine recombinase XerC in ICEPan CR-5. The stress-induced upregulation of umuC may thus drive the excision and dissemination of the island and its encoded genes. A BlastP analysis with the UmuC and UmuD protein sequences against the ICEberg dataset showed that orthologs were present in 21 out of 270 complete ICE elements. Of these, an island insertion was observed in the umuC gene of 10 ICE elements, integrated mostly in ICEs of Vibrio spp., but also one Pseudomonas aeruginosa and one Streptococcus pyogenes ICE (Table S4). The islands inserted into the umuC gene in the ICE elements of these microorganisms carried genes implicated in mercury resistance and resistance to several antibiotics including tetracycline, chloramphenicol/florfenicol and streptomycin (Beaber et al., 2002; Battle et al., 2008). In the context of the above hypotheses, the umuC island may thus play a major role in the regulation and/or dissemination of important phenotypes from both a clinical and ecological perspective.
Conclusions
ICEs of the novel ICEPan family are a common feature among P. ananatis strains isolated from various environmental sources and hosts, and with different lifestyles. In silico characterization of the ICEPan elements showed similar ICEs are found in other enterobacteria and that these ICEs, and conjugative plasmids in E. billingiae and E. amylovora, are likely derived from a common ancestor. ICEPan contain the core modules required for their chromosomal integration, maintenance and dissemination. The presence of these core elements, however, does not provide definitive proof that the ICEPan elements can excise, circularize and integrate within the chromosome. Currently, assays are being performed to determine whether ICEPan represent functional integrative and conjugative elements, particularly ICEPanAJ13355, which lacks two of the core modules. Analyses of the ICE cargo genes suggest a likely role in response to various stresses they may encounter in the environment, including oxidative, pH, and genotoxic stresses. ICEPan elements could therefore play a role in facilitating P. ananatis survival under these stresses and provide a potential mechanism for this species to exploit novel ecological niches. With recent interest in the use of P. ananatis as biological control agent against phytopathogenic bacteria and fungi, the ICEPan elements, and the putative antibiotic biosynthetic loci encoded on these elements, provide potential targets for the exploration of P. ananatis antibiosis phenotypes. Perhaps the most pertinent finding in the ICEPan elements is the integration of an island in the umuDC locus, which shows evidence of recent horizontal acquisition between different Pantoea species and related genera. Here, we postulate on a potential role of umuDC in the dissemination and/or expression of phenotypes encoded on this island. Islands integrated into this locus are furthermore found in other bacterial taxa and encode phenotypes such as antibiotic and heavy metal resistance. It can be reasonably expected that similar features will be identified in the ICEs of many other bacterial taxa whose genomes are sequenced. Considering the clinical and ecological relevance of the phenotypes encoded in umuDC, this phenomenon warrants further attention.
Author Contributions
PM, SV, BD, TS, and TC conceived the study. PM, WC, DM, JB, and TS performed experiments and analyses, PM, SV, BD, DC, TS, and TC wrote the original manuscript. All authors contributed to and approved of the final version.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study was partially supported by the University of Pretoria Postdoctoral Fellowship Fund, National Research Foundation (NRF), the Tree Protection Co-operative Programme (TPCP), the NRF/Dept. of Science and Technology Centre of Excellence in Tree Health Biotechnology (CTHB), and the THRIP support program of the Department of Trade and Industry, South Africa, the department of Life Sciences and Facility Management of ZHAW, the Swiss Federal Office for Agriculture (BLW Fire Blight Research – Achilles), and the Swiss Secretariat for Education and Research (SBF C07.0038). The authors wish to acknowledge G. Carstensen for laboratory assistance and the various culture collections that provided P. ananatis strains (Table 2).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.00576/abstract
References
Altschul S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local alignment search tool. J. Mol. Biol. 215, 403–410.
Aminov, R. I. (2011). Horizontal gene exchange in environmental microbiota. Front. Microbiol. 2:158. doi: 10.3389/fmicb.2011.00158
Baker, M. D., Wolanin, P. M., and Stock, J. B. (2006). Signal transduction mechanisms in bacterial chemotaxis. Bioessays 28, 9–22. doi: 10.1002/bies.20343
Battle, S. E., Meyer, F., Rello, J., Kung, V. L., and Hauser, A. R. (2008). Hybrid pathogenicity island PAGI-5 contributes to the highly virulent phenotype of a Pseudomonas aeruginosa isolate in mammals. J. Bacteriol. 190, 7130–7140. doi: 10.1128/JB.00785-08
Beaber, J. W., Hochhut, B., and Waldor, M. K. (2002). Genomic and functional analyses of SXT, an integrating antibiotic resistance gene transfer element derived from Vibrio cholerae. J. Bacteriol. 184, 4259–4269. doi: 10.1128/JB.184.15.4259-4269.2002
Bi, D., Xu, Z., Harrison, E. M., Tai, C., Wei, Y., He, X., et al. (2012). ICEberg: a web-based resource for integrative and conjugative elements found in Bacteria. Nucleic Acids Res. 40, 621–626. doi: 10.1093/nar/gkr846
Blin, K., Medema, M. H., Kazempour, D., Fischbach, M. A., Breitlin, R., Takano, E., et al. (2013). antiSMASH 2.0—a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res. 41, 204–212. doi: 10.1093/nar/gkt449
Bougdour, A., and Gottesman, S. (2007). ppGpp regulation via anti-adaptor protein IraP. Proc. Natl. Acad. Sci. U.S.A. 104, 12896–12901. doi: 10.1073/pnas.0705561104
Boyd, E. F., Almagro-Moreno, S., and Parent, M. A. (2009). Genomic islands are dynamic, ancient integrative elements in bacterial evolution. Trends Microbiol. 17, 47–53. doi: 10.1016/j.tim.2008.11.003
Burrus, V., Marrero, J., and Waldor, M. K. (2006). The current ICE age: biology and evolution of SXT-related integrating conjugative elements. Plasmid 55, 173–183. doi: 10.1016/j.plasmid.2006.01.001
Burrus, V., Pavlovic, G., Decaris, B., and Guédon, G. (2002). Conjugative transposons: the tip of the iceberg. Mol. Microbiol. 46, 601–610. doi: 10.1046/j.1365-2958.2002.03191.x
Choi, O., Lim, J. Y., Seo, Y.-S., Hwang, I., and Kim, J. (2012). Complete genome sequence of the rice pathogen Pantoea ananatis PA13. J. Bacteriol. 194, 531. doi: 10.1128/JB.06450-11
Collyn, F., Guy, L., Marceau, M., Simonet, M., and Roten, C. H. (2006). Describing ancient horizontal gene transfers at the nucleotide and gene levels by comparative pathogenicity island genometrics. Bioinformatics 22, 1072–1079. doi: 10.1093/bioinformatics/bti793
Collyn, F., Léty, M. A., Nair, S., Escuyer, V., Ben Younes, A., Simonet, M., et al. (2002). Yersinia pseudotuberculosis harbours a type IV pilus gene cluster that contributes to pathogenicity. Infect. Immun. 70, 6196–6205. doi: 10.1128/IAI.70.11.6196-6205.2002
Coutinho, T. A., and Venter, S. N. (2009). Pantoea ananatis: an unconventional plant pathogen. Mol. Plant Pathol. 10, 325–335. doi: 10.1111/j.1364-3703.2009.00542.x
De Baere, T., Verhelst, R., Labit, C., Verschraegen, G., Wouters, G., Claeys, G., et al. (2004). Bacteremic infection with Pantoea ananatis. J. Clin. Microbiol. 42, 4393–4395. doi: 10.1128/JCM.42.9.4393-4395.2004
De Maayer, P., Chan, W. Y., Rezzonico, F., Bühlmann, A., Venter, S. N., Blom, J., et al. (2012). Complete genome sequence of clinical isolate Pantoea ananatis LMG5342. J. Bacteriol. 194, 1615–1616. doi: 10.1128/JB.06715-11
De Maayer, P., Chan, W. Y., Rubagotti, E., Venter, S. N., Toth, I. K., Birch, P. R. J., et al. (2014). Analysis of the Pantoea ananatis pan-genome reveals factors underlying its ability to colonize and interact with plant, insect and vertebrate hosts. BMC Genomics 15:404. doi: 10.1186/1471-2164-15-404
De Maayer, P., Chan, W. Y., Venter, S. N., Toth, I. K., Birch, P. R. J., Joubert, F., et al. (2010). Genome sequence of Pantoea ananatis LMG20103, the causative agent of Eucalyptus blight and dieback. J. Bacteriol. 192, 2936–2937. doi: 10.1128/JB.00060-10
Gasser, F., Cardinale, M., Schildberger, B., and Berg, G. (2012). Biocontrol of Botrytis cinerea by successful introduction of Pantoea ananatis in the grapevine phyllosphere. Int. J. Wine Res. 4, 53–63. doi: 10.2147/ijwr.s31339
Ghinet, M. G., Bordeleau, E., Beaudin, J., Brzezinski, R., Roy, S., and Burrus, V. (2011). Uncovering the prevalence and diversity of integrating conjugative elements in Actinobacteria. PLoS ONE 6:e27846. doi: 10.1371/journal.pone.0027846
Grobler, J., Bauer, F., Subden, R. E., and Van Vuuren, H. J. (1995). The mae1 gene of Schizosaccharomyces pombe encodes a permease for malate and other C4 dicarboxylic acids. Yeast 11, 1485–1491. doi: 10.1002/yea.320111503
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98.
Hara, Y., Kadotani, N., Izui, H., Katashkina, J. I., Kuvaeva, T., Andreeva, I. G., et al. (2011). The complete genome sequence of Pantoea ananatis AJ13355, an organism with great biotechnological potential. Appl. Microbiol. Biotechnol. 93, 331–341. doi: 10.1007/s00253-011-3713-5
Hare, J. M., Perkins, S. N., and Greg-Jolly, L. A. (2006). A constitutively expressed, truncated umuDC operon regulates the recA-dependent DNA damage induction of a gene in Acinetobacter bayli strain ADP1. Appl. Environ. Microbiol. 72, 4036–4043. doi: 10.1128/AEM.02774-05
Healy, F. G., Eaton, K. P., Limsirichai, P., Aldrich, J. F., Plowman, A. K., and King, R. R. (2009). Characterization of γ-butyrolactone autoregulatory signaling gene homologs in the angucyclinone polyketide WS5595B producer Streptomyces acidiscabies. J. Bacteriol. 191, 4786–4797. doi: 10.1128/JB.00437-09
Hindupur, A., Liu, D., Zhao, Y., Bellamy, H. D., White, M. A., and Fox, R. O. (2006). The crystal structure of the E. coli stress protein YciF. Protein Sci. 409, 2605–2611. doi: 10.1110/ps.062307706
Hochhut, B., Jahreis, K., Lengeler, J. W., and Schmid, K. (1997). CTnscr94, a conjugative transposon found in enterobacteria. J. Bacteriol. 179, 2097–2102.
Katoh, K., and Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780. doi: 10.1093/molbev/mst010
Kim, H. J., Lee, J. H., Kang, B. R., Rong, X., McSpadden Gardener, B. B., Ji, H. J., et al. (2012). Draft genome sequence of Pantoea ananatis B1-9, a non-pathogenic plant growth-promoting bacterium. J. Bacteriol. 194, 729. doi: 10.1128/JB.06484-11
Kube, M., Migdoll, A. M., Gehring, I., Heitmann, K., Mayer, Y., Kuhl, H., et al. (2010). Genome comparison of the epiphytic Erwinia billingiae and E. tasmaniensis with the pear pathogen E. pyrifoliae. BMC Genomics 11:393. doi: 10.1186/1471-2164-11-393
Kvint, K., Nachin, L., Diez, A., and Nystro, T. (2003). The bacterial universal stress protein: function and regulation. Curr. Opin. Microbiol. 6, 140–145. doi: 10.1016/S1369-5274(03)00025-0
Llop, P., Cabrefiga, J., Smits, T. H. M., Dreo, T., Barbé, S., Pulawska, J., et al. (2011). Erwinia amylovora novel plasmid pEI70: complete sequence, biogeography, and role in aggressiveness in the fire blight phytopathogen. PLoS ONE 6:e28651. doi: 10.1371/journal.pone.0028651
Marchler-Bauer, A., Anderson, J. B., Chitsaz, F., Derbyshire, M. K., DeWeese-Scott, C., Fong, J. H., et al. (2009). CDD: specific functional annotation with the Conserved Domain Database. Nucleic Acids Res. 37, 205–210. doi: 10.1093/nar/gkn845
Marchler-Bauer, A., and Bryant, S. H. (2004). CD-Search: protein domain annotations on the fly. Nucleic Acids Res. 32, 327–331. doi: 10.1093/nar/gkh454
Moreno-Hagelsieb, G., and Latimer, K. (2008). Choosing BLAST options for better detection of orthologs as reciprocal best hits. Bioinformatics 24, 319–324. doi: 10.1093/bioinformatics/btm585
Nurmohamed, S., McKay, A. R., Robinson, C. V., and Luisi, B. F. (2010). Molecular recognition between Escherichia coli enolase and ribonuclease E. Acta Crystallogr. D Biol. Crystallogr. 66, 1036–1040. doi: 10.1107/S0907444910030015
Oglesby, A. G., Murphy, E. R., Iyer, V. R., and Payne, S. M. (2005). Fur regulates acid resistance in Shigella flexneri via RyhB and ydeP. Mol. Microbiol. 58, 1354–1367. doi: 10.1111/j.1365-2958.2005.04920.x
Panmanee, W., Vattanaviboon, P., Poole, L. B., and Mongkolsuk, S. (2006). Novel organic hydroperoxide-sensing and responding mechanisms for OhrR, a major bacteria sensor and regulator of organic hydroperoxide stress. J. Bacteriol. 188, 1389–1395. doi: 10.1128/JB.188.4.1389-1395.2006
Pickens, L., Kim, W., Wang, P., Zhou, H., Watanabe, K., Gomi, S., et al. (2010). Biochemical analysis of the biosynthetic pathway of an anticancer tetracycline SF2575. J. Am. Chem. Soc. 131, 17677–17689. doi: 10.1021/ja907852c
Ren, Y., Ren, Y., Zhou, Z., Guo, X., Li, Y., Feng, L., et al. (2010). Complete genome sequence of Enterobacter cloacae subsp. cloacae type strain ATCC 13047. J. Bacteriol. 192, 2463–2464. doi: 10.1128/JB.00067-10
Robbe-Saule, V., Coynault, C., Ibanez-Ruiz, M., Hermant, D., and Norel, F. (2001). Identification of a non-haem catalase in Salmonella and its regulation by RpoS (σS). Mol. Microbiol. 39, 1533–1545. doi: 10.1046/j.1365-2958.2001.02340.x
Roche, D., Fléchard, M., Lallier, N., Répérant, M., Brée, A., Pascal, G., et al. (2010). ICEEc2, a new integrative and conjugative element belonging to the pKLC102/PAGI-2 family, identified in Escherichia coli BEN374. J. Bacteriol. 192, 5026–5046. doi: 10.1128/JB.00609-10
Smits, T. H. M., Rezzonico, F., Lopez, M. M., Blom, J., Goesmann, A., Frey, J. E., et al. (2013). Phylogenetic position and virulence apparatus of the pear flower necrosis pathogen Erwinia piriflorinigrans CFBP 5888T as assessed by comparative genomics. Syst. Appl. Microbiol. 36, 449–456. doi: 10.1016/j.syapm.2013.04.003
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., and Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739. doi: 10.1093/molbev/msr121
Weller-Stuart, T., Chan, W. Y., Coutinho, T. A., Venter, S. N., Smits, T. H. M., Duffy, B., et al. (2014). Draft genome sequences of the onion center rot pathogen Pantoea ananatis PA4 and maize brown stalk rot pathogen P. ananatis BD442. Genome Announc. 2:e00750-14. doi: 10.1128/genomeA.00750-14
Wertz, J. E., and Riley, M. A. (2004). Chimeric nature of two plasmids of Hafnia alvei encoding the bacteriocins Alveicins A and B. J. Bacteriol. 186, 1598–1605. doi: 10.1128/JB.186.6.1598-1605.2004
Woodyer, R. D., Shao, Z., Thomas, P. M., Kelleher, N. L., Blodgett, J. A. V., Metcalf, W. W., et al. (2006). Heterologous production of fosfomycin and identification of the minimal biosynthetic gene cluster. Chem. Biol. 13, 1171–1182. doi: 10.1016/j.chembiol.2006.09.007
Wozniak, R. A. F., and Waldor, M. K. (2009). A toxin-antitoxin system promotes the maintenance of an integrative conjugative element. PLoS Genet. 5:e1000439. doi: 10.1371/journal.pgen.1000439
Keywords: Pantoea ananatis, integrative and conjugative element, ICEPan, stress response, antibiosis, umuDC
Citation: De Maayer P, Chan W-Y, Martin DAJ, Blom J, Venter SN, Duffy B, Cowan DA, Smits THM and Coutinho TA (2015) Integrative conjugative elements of the ICEPan family play a potential role in Pantoea ananatis ecological diversification and antibiosis. Front. Microbiol. 6:576. doi: 10.3389/fmicb.2015.00576
Received: 18 March 2015; Accepted: 25 May 2015;
Published: 08 June 2015.
Edited by:
Eric Altermann, AgResearch Ltd., New ZealandReviewed by:
Daniela Ceccarelli, University of Maryland, USAClaudio Palmieri, Polytechnic University of Marche, Italy
Copyright © 2015 De Maayer, Chan, Martin, Blom, Venter, Duffy, Cowan, Smits and Coutinho. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Pieter De Maayer, Centre for Microbial Ecology and Genomics, University of Pretoria, 3-17, Natural Sciences II Building, 0002 Pretoria, South Africa,cGlldGVyLmRlbWFheWVyQHVwLmFjLnph
 Pieter De Maayer
Pieter De Maayer Wai-Yin Chan2,3
Wai-Yin Chan2,3 Douglas A. J. Martin
Douglas A. J. Martin Stephanus N. Venter
Stephanus N. Venter Brion Duffy
Brion Duffy Don A. Cowan
Don A. Cowan Theo H. M. Smits
Theo H. M. Smits