- 1Lee Kong Chian School of Medicine, Nanyang Technological University, Singapore, Singapore
- 2Faculty of Medicine, University of British Columbia, Vancouver, BC, Canada
Here we report our perspective on applying GapmeR technology in combination with recombinant angiotensin-converting enzyme 2 (ACE2) in the treatment of COVID-19 patients. GapmeR is a cell-permeating antisense single-stranded DNA molecule that can be designed to specifically target intracellular severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Once internalized into host cells, such as lung alveolar cells, GapmeR molecules can bind to the viral RNA. This RNA/DNA hybrid will then be degraded by the RNase H enzyme abundantly present in the host cells. GapmeRs can be delivered to COVID-19 patients through inhalation or via nebulization. SARS-CoV-2-targeted GapmeR can also be given to frontline healthcare workers as a prophylactic protection. The recombinant ACE2 protein, the efficacy of which is being evaluated in clinical trials, will bind to the spike (S) glycoprotein of extracellular SARS-CoV-2 and potentially block viral infectivity. We propose that combining inhalable SARS-CoV-2-targeted GapmeRs with recombinant ACE2 could provide a viable and rapidly implementable more effective therapeutic approach for eradicating SARS-CoV-2 and save millions of lives.
Introduction
COVID-19 or the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) continues to threaten human health worldwide. As of 15th July 2020, there are over 13.4 million confirmed cases and over 580,000 deaths worldwide1. In an accelerated effort to develop therapeutic modalities and vaccines to manage this devastating pandemic, collaborative teams of regulatory bodies, health professionals, clinicians, and scientists in both academic and industrial settings have been engaged and integrated. Although there has been some success in developing new vaccines, repurposing previous coronavirus antibodies, and other potential therapeutic modalities, there is no effective therapy available for the ongoing COVID-19 pandemic. Through expedited regulatory processes, the US Food and Drug Administration (FDA) has granted Emergency Use Authorization (EUA) for remdesivir (an investigational drug, originally developed to tackle Ebola) on 1st May 2020 to treat severe COVID-19 cases (Media). Surprisingly, there has been relatively little exploration of directly targeting the SARS-CoV-2 genome.
SARS-CoV-2
We now know the complete genome of SARS-CoV-2, which is a single-stranded positive-sense RNA of about ∼30 kb and is encapsulated within a membranous envelope. As with other coronavirus, SARS-CoV-2 contains four main structural proteins – spike (S) glycoprotein, membrane (M) protein, envelope (E) protein, and nucleocapsid (N) protein (Wu et al., 2020). The infection is established by virus particles following their entry through airway and binding of viral S protein to the receptor angiotensin-converting enzyme 2 (ACE2) expressed on type II alveolar cells of lung (Zou et al., 2020). The S-ACE2 complex then internalizes into alveolar cells by endocytosis (Yan et al., 2020). The ACE2 protein is also expressed on esophagus epithelial cells, cardiac muscle cells, ocular cells, kidney proximal tubules, urothelial cells, colon, and on the mucosa of oral cavity (Ma et al., 2020; Xu et al., 2020).
Silencing the Viral Genome
RNA interference (RNAi) is a powerful biological process inherently applied by the host cells to destroy intracellular RNA viruses. For the past two decades, researchers have been widely capitalizing on RNAi mechanisms to silence cellular gene expression in vitro and in vivo. Studies have validated the use of small interfering RNA (siRNA) to safeguard host from viral infection by inhibiting the replication of viral genome, by blocking the expression of viral antigens and accessory genes and by interfering with the assembly of viral particles (Lenartowicz et al., 2016; Levanova and Poranen, 2018). Previous studies have demonstrated successful intratracheal administration of siRNA using pressurized metered dose inhalers or nebulizers (Miwata et al., 2018; Ng et al., 2019). Indeed, OliX Pharmaceuticals, Sirnaomics, and a collaborative team of Alnylam Pharmaceuticals and Vir Biotechnology have initiated the development of siRNA therapeutics to treat COVID-19 patients. However, siRNA-based therapeutic approach is challenging mainly because of poor stability of siRNA molecules in biological environment and problems associated with delivery into patients’ cells and tissues.
Inhalable GapmeR as a Therapeutic Option for COVID-19
At this juncture, we propose to explore the utility of an alternative and possibly more effective approach using inhalable antisense DNA molecules, such as GapmeRs. Typically, GapmeR molecules have a central stretch of chemically modified DNA “gap” flanked by locked nucleic acids (LNA), which increase the binding affinity of GapmeR to the target RNA. In addition to specificity and highly efficient gene silencing, we have shown that GapmeRs can easily internalize into target cells (e.g., human primary and cultured T-cells) by macropinocytosis or gymnosis (Fazil et al., 2016; Verma et al., 2017; Ong et al., 2018); thus requiring no additional delivery assistance or transfection agents. GapmeR molecules targeting specific regions of SARS-CoV-2 RNA can easily be designed using publicly available genome sequence information and in silico antisense design tools (Kurreck et al., 2002). Care must be taken to avoid potential off-target effects by searching selected sequences of GapmeRs against the databases of human transcripts and filtering out cross-reacting sequences. Several other stringent parameters, including target accessibility, binding affinity and a series of assessment criteria as recommended by the Oligonucleotide Safety Working Group (Marlowe et al., 2017; Yoshida et al., 2019) should be taken into consideration.
We have already designed a number of GapmeR molecules targeting the highly conserved regions of the SARS-CoV-2, including the RNA-dependent RNA polymerase (RdRP), S protein and M protein. While designing SARS-CoV-2 targeting GapmeRs, we have considered multiple parameters, including RNA secondary structure, accessibility of target region, and avoidance of off-target effects using robust bioinformatics algorithms and in silico analysis. We believe that once internalized, virus-specific GapmeRs would bind to SARS-CoV-2 RNA within a host cell. RNase H enzymes present in the host cell will then specifically degrade the DNA/RNA hybrid and thus destroy SARS-CoV-2 (Figure 1).
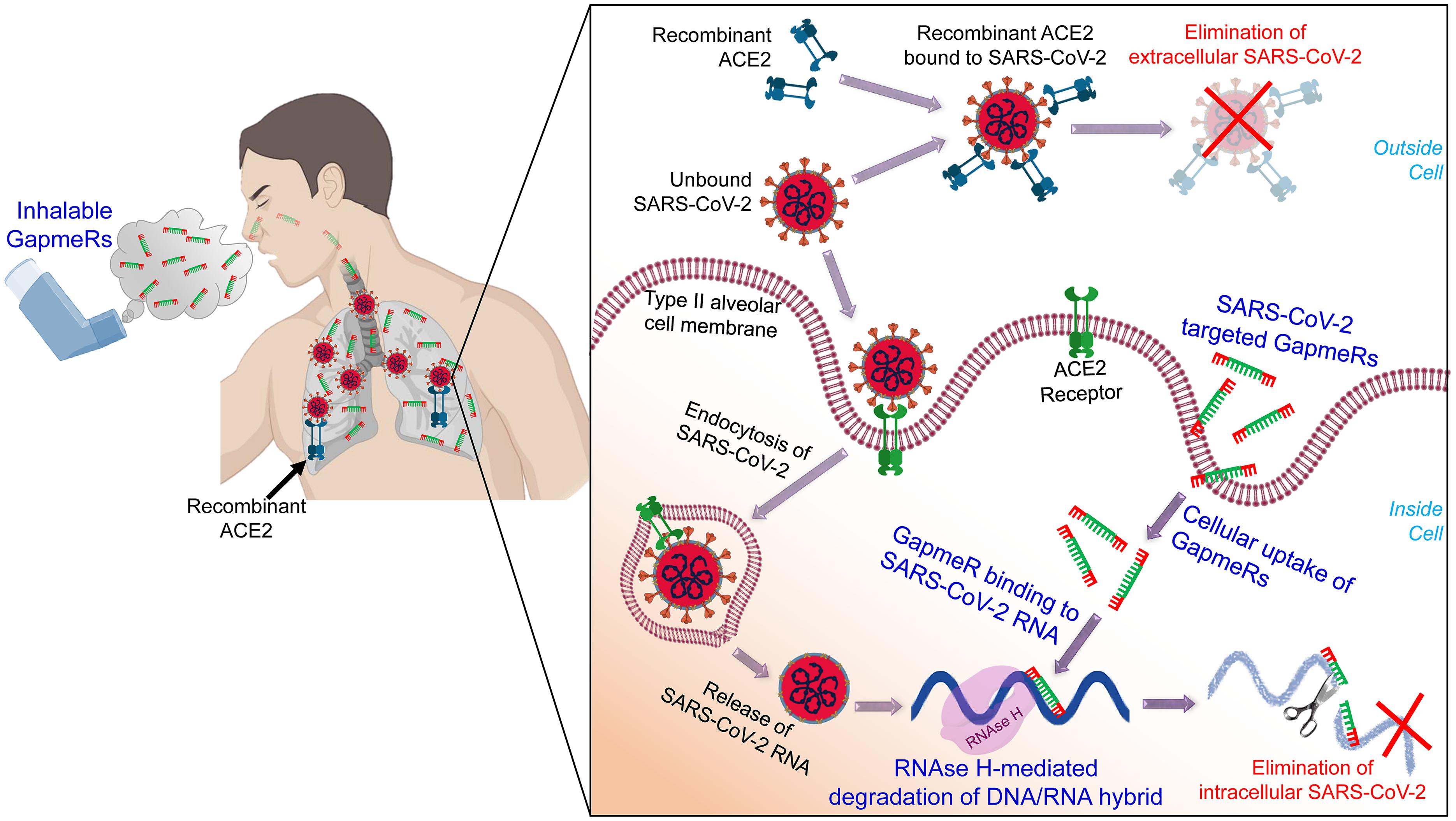
Figure 1. A schematic illustration of administering inhalable GapmeRs and recombinant ACE2 to the lung for treating COVID-19. The proposed mechanism of action of SARS-CoV-2 targeted GapmeR molecules in destroying intracellular viral RNA in type II alveolar cells and recombinant ACE2 in eliminating extracellular virus particles in the lung. Figure prepared using images created with BioRender.
GapmeRs can be administered via inhalation as aerosol or in powder form, either using a customized handheld nebulizer device or through the ventilator. Using either of these simple non-invasive inhalation devices, one can easily achieve high enough concentration of GapmeR molecules in the lung sufficient for type II alveolar cell uptake. GapmeRs could conceivably be administered to frontline healthcare worker dealing with COVID-19 patients to provide them prophylactic protection. Since GapmeR molecules are highly stable in human serum (Grünweller et al., 2003), we would expect less frequent dosing of the SARS-CoV-2-targeted GapmeRs, in this situation. GapmeRs may also remain effective or could be modified as the virus mutates and could have cross-reactivity against related viral strains that may emerge in the future. Other advantages of using GapmeR therapeutics are well-established chemistry and synthesis process, easily achievable manufacturing and bulk production and low storage or shipment cost due to high stability, even at room temperature.
It is now evident that GapmeR technology is gaining significant interest in therapeutic development (Hagedorn et al., 2018; Pedersen et al., 2020). There has been substantial progress since the first systemically delivered GapmeR therapeutic mipomersen (Kynamro) approved by the FDA in 2013 for reducing cholesterol levels in familial hypercholesterolemia (Parham and Goldberg, 2019). Tominersen (previously known as IONIS-HTTRx) is another GapmeR molecule that has been granted Orphan Drug Designation by FDA in 2016 for the treatment of Huntington’s disease (Rodrigues and Wild, 2018). Nusinersen (Spinraza), an antisense nucleotide with phosphorothioate backbones and 2′-O-methoxyethyl modification, has been approved by the FDA in 2016 for treating spinal muscular atrophy (Aartsma-Rus, 2017). More recently, in December 2019, FDA has granted accelerated approval to an antisense phosphorodiamidate morpholino oligomer, golodirsen, for treating patients with Duchene Muscular Dystrophy (Heo, 2020). GapmeR targeting SOD1 mutation, tofersen (previously known as IONIS SOD1Rx), is currently in phase III clinical trial (NCT02623699) for treating a fatal neurodegenerative disease amyotrophic lateral sclerosis. The continued success of GapmeR therapeutics and tolerable safety profiles in ongoing clinical trials predict a robust outlook for a new effective treatment strategy for COVID-19.
It is worth noting that, like any other drugs, GapmeRs may have unforeseen off-target effects, including for example, unwanted interactions with host proteins (e.g., Toll-Like Receptors). While such interactions between SARS-CoV-2-targeted GapmeRs and host proteins may not be strong, they may trigger unintended signaling pathways, cytokine release, and yield confounding physiological effects. In our previous work using gene expression analysis, we did not find any major immunological off-target effects of a panel of GapmeR molecules that we designed against multiple targets (Fazil et al., 2016). Nevertheless, if toxicity arises in further advanced studies, measures to eliminate or mitigate potential risks could be implemented during pre-clinical testing. Another challenge would be to efficiently deliver GapmeR molecules to type II alveolar cells in the lung. An antisense oligonucleotide eluforsen has been successfully administered via inhalation to treat respiratory symptoms in certain patients with cystic fibrosis and granted fast track designation by the FDA in 2016 (Kim et al., 2019). Single and multiple doses of inhaled eluforsen have been proven to be safe and well-tolerated (Drevinek et al., 2020). It would also be important to adjust the administering frequency and dosing of inhalable GapmeRs to manage varying levels of viremia as well as to treat patients at different stages of symptomatic or non-symptomatic infections. The key to success of translating GapmeR technology to treat COVID-19 patients would be to (i) screen multiple GapmeR designs against highly conserved regions of SARS-CoV-2 genome, (ii) carefully select potent candidates with high stability and no off-target effect, (iii) demonstrate their delivery to the lung using multimodal in vitro, organoids, and animal model systems appropriate for COVID-19 preclinical research (Takayama, 2020), (iv) determine dosing regimens for various infection scenarios, including for example, cases where the virus titer is increasing, and (v) achieve a good therapeutic window with acceptable toxicity profiles while suppressing viral loads in well-controlled experiments.
Possibility of Combining SARS-CoV-2-Targeted GapmeR With Recombinant ACE2
In a recent study, Monteil et al. (2020) have shown the therapeutic potential of human recombinant soluble ACE2 (hrACE2) protein to inhibit the replication of SARS-CoV-2. In addition, hrACE2 protein has been shown to be protective in acute lung injury (Imai et al., 2005). Thus, soluble form of ACE2 would not only block the early entry of SARS-CoV-2 in host cells and hence viral spread, it would also protect the COVID-19 patients form severe acute lung failure (Zhang et al., 2020). Previous phase II trial results confirm that the hrACE2 is well-tolerated (even at high dose up to 0.8 mg/kg) in patients with acute respiratory distress syndrome (Khan et al., 2017) and significantly improves cardiac output and pulmonary vascular resistance in pulmonary arterial hypertension (Hemnes et al., 2018). Treatment of diabetic mice with hrACE2 (intraperitoneal injection of 2 mg/kg hrACE2 for 4 weeks) has been shown to slow the progression of diabetic nephropathy (Oudit et al., 2010). Accordingly, a number of clinical trials, including NCT04375046, NCT04335136, and NCT04382950, to evaluate the recombinant ACE2 therapy for COVID-19 have recently been initiated2. While viral replication is reduced through the use of hrsACE2, SARS-CoV-2 is not eliminated. It should be noted that SARS-CoV-2 can target multiple cell types expressing ACE2 and that, in addition to alveolar epithelial cells, many human tissues and organs express the ACE2 (Xu et al., 2020; Yan et al., 2020; Zou et al., 2020), which might provide possible routes of entry for the SARS-CoV-2. Anticipating successful attenuation of outcomes in these clinical trials for recombinant ACE2, we propose that simultaneously administration SARS-CoV-2-targeted GapmeR through inhalation route could be synergistic in fully inhibiting viral replication. While recombinant ACE2 would block the extracellular virus particles entering host cells, therapeutic GapmeR would destroy intracellular SARS-CoV-2 genome (Figure 1). This combinatorial approach could thus eradicate SARS-CoV-2, effectively reduce the severity of infection and save millions of lives.
Conclusion
In conclusion, there is an urgent need to radically rethink current management of the devastating COVID-19 pandemic. Although there are a set of rapidly increasing challenges, logistic, and regulatory hurdles ahead to bring a proposed GapmeR technology to clinics, these can be overcome by engaging collaborative academia-industry partnerships, setting-up joint screening, and testing facilities and making research funding available to support new concepts. We believe that targeted GapmeR in combination with recombinant ACE2 protein could provide a viable rapidly implementable therapeutic approach for COVID-19 and similar infections in the future.
Data Availability Statement
Publicly available datasets were analyzed in this study. This data can be found here: https://www.ncbi.nlm.nih.gov/labs/virus/vssi/#/virus?SeqType_s=Nucleotide&VirusLineage_ss=SARS-CoV-2,%20taxid:2697049&utm_source=gene&utm_medium=referral&utm_campaign=COVID-19.
Author Contributions
NV and DK conceived the idea. NV, MF, SD, and DK wrote the manuscript. All authors approved the final manuscript version.
Funding
NV acknowledges funding support from the Singapore Ministry of Education under its Singapore Ministry of Education (MOE) Academic Research Fund (AcRF) Tier 2 Grant (MOE2017-T2-2-004). DK and SD acknowledge Tiny Ventures Ltd, Canada.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
References
Aartsma-Rus, A. (2017). FDA approval of nusinersen for spinal muscular atrophy makes 2016 the year of splice modulating oligonucleotides. Nucleic Acid Ther. 27, 67–69. doi: 10.1089/nat.2017.0665
Drevinek, P., Pressler, T., Cipolli, M., De Boeck, K., Schwarz, C., Bouisset, F., et al. (2020). Antisense oligonucleotide eluforsen is safe and improves respiratory symptoms in F508DEL cystic fibrosis. J. Cyst. Fibros. 19, 99–107. doi: 10.1016/j.jcf.2019.05.014
Fazil, M. H. U. T., Ong, S. T., Chalasani, M. L., Low, J. H., Kizhakeyil, A., Mamidi, A., et al. (2016). GapmeR cellular internalization by macropinocytosis induces sequence-specific gene silencing in human primary T-cells. Sci. Rep. 6: 37721.
Grünweller, A., Wyszko, E., Bieber, B., Jahnel, R., Erdmann, V. A., and Kurreck, J. (2003). Comparison of different antisense strategies in mammalian cells using locked nucleic acids, 2′-O-methyl RNA, phosphorothioates and small interfering RNA. Nucleic Acids Res. 31, 3185–3193. doi: 10.1093/nar/gkg409
Hagedorn, P. H., Persson, R., Funder, E. D., Albæk, N., Diemer, S. L., Hansen, D. J., et al. (2018). Locked nucleic acid: modality, diversity, and drug discovery. Drug Discov. Today 23, 101–114. doi: 10.1016/j.drudis.2017.09.018
Hemnes, A. R., Rathinasabapathy, A., Austin, E. A., Brittain, E. L., Carrier, E. J., Chen, X., et al. (2018). A potential therapeutic role for angiotensin-converting enzyme 2 in human pulmonary arterial hypertension. Eur. Respir. J. 51:1702638. doi: 10.1183/13993003.02638-2017
Imai, Y., Kuba, K., Rao, S., Huan, Y., Guo, F., Guan, B., et al. (2005). Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature 436, 112–116. doi: 10.1038/nature03712
Khan, A., Benthin, C., Zeno, B., Albertson, T. E., Boyd, J., Christie, J. D., et al. (2017). A pilot clinical trial of recombinant human angiotensin-converting enzyme 2 in acute respiratory distress syndrome. Crit. Care 21:234.
Kim, J., Basiri, B., Hassan, C., Punt, C., van der Hage, E., den Besten, C., et al. (2019). Metabolite profiling of the antisense oligonucleotide eluforsen using liquid chromatography-mass spectrometry. Mol. Ther. Nucleic Acids 17, 714–725. doi: 10.1016/j.omtn.2019.07.006
Kurreck, J., Wyszko, E., Gillen, C., and Erdmann, V. A. (2002). Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res. 30, 1911–1918. doi: 10.1093/nar/30.9.1911
Lenartowicz, E., Nogales, A., Kierzek, E., Kierzek, R., Martínez-Sobrido, L., and Turner, D. H. (2016). Antisense oligonucleotides targeting influenza A segment 8 genomic RNA inhibit viral replication. Nucleic Acid Ther. 26, 277–285. doi: 10.1089/nat.2016.0619
Levanova, A., and Poranen, M. M. (2018). RNA interference as a prospective tool for the control of human viral infections. Front. Microbiol. 9:2151. doi: 10.3389/fmicb.2018.02151
Ma, D., Chen, C. B., Jhanji, V., Xu, C., Yuan, X. L., Liang, J. J., et al. (2020). Expression of SARS-CoV-2 receptor ACE2 and TMPRSS2 in human primary conjunctival and pterygium cell lines and in mouse cornea. Eye 34, 1212–1219. doi: 10.1038/s41433-020-0939-4
Marlowe, J. L., Akopian, V., Karmali, P., Kornbrust, D., Lockridge, J., and Semple, S. (2017). Recommendations of the oligonucleotide safety working group’s formulated oligonucleotide subcommittee for the safety assessment of formulated oligonucleotide-based therapeutics. Nucleic Acid Ther. 27, 183–196. doi: 10.1089/nat.2017.0671
Miwata, K., Okamoto, H., Nakashima, T., Ihara, D., Horimasu, Y., Masuda, T., et al. (2018). Intratracheal administration of siRNA dry powder targeting vascular endothelial growth factor inhibits lung tumor growth in mice. Mol. Ther. Nucleic Acids 12, 698–706. doi: 10.1016/j.omtn.2018.07.009
Monteil, V., Kwon, H., Prado, P., Hagelkrüys, A., Wimmer, R. A., Stahl, M., et al. (2020). Inhibition of SARS-CoV-2 infections in engineered human tissues using clinical-grade soluble human ACE2. Cell 181, 905–913. doi: 10.1016/j.cell.2020.04.004
Ng, B., Cash-Mason, T., Wang, Y., Seitzer, J., Burchard, J., Brown, D., et al. (2019). Intratracheal administration of siRNA triggers mRNA silencing in the lung to modulate T cell immune response and lung inflammation. Mol. Ther. Nucleic Acids 16, 194–205. doi: 10.1016/j.omtn.2019.02.013
Ong, S. T., Chalasani, M. L. S., Fazil, M. H. U. T., Prasannan, P., Kizhakeyil, A., Wright, G. D., et al. (2018). Centrosome- and Golgi-localized protein kinase N-associated protein serves as a docking platform for protein kinase A signaling and microtubule nucleation in migrating T-cells. Front. Immunol. 9:397. doi: 10.3389/fmicb.2018.0397
Oudit, G. Y., Liu, G. C., Zhong, J., Basu, R., Chow, F. L., Zhou, J., et al. (2010). Human recombinant ACE2 reduces the progression of diabetic nephropathy. Diabetes 59, 529–538. doi: 10.2337/db09-1218
Parham, J. S., and Goldberg, A. C. (2019). Mipomersen and its use in familial hypercholesterolemia. Expert. Opin. Pharmacother. 20, 127–131. doi: 10.1080/14656566.2018.1550071
Pedersen, L., Hagedorn, P., Vikeså, J., Karlsen, S. T., Særmark, P., Koch, T., et al. (2020). Targeting repeated regions unique to a gene is an effective strategy for discovering potent and efficacious antisense oligonucleotides. Mol. Ther. Nucleic Acids 19, 124–131. doi: 10.1016/j.omtn.2019.10.040
Rodrigues, F. B., and Wild, E. J. (2018). Huntington’s disease clinical trials corner: february 2018. J. Huntingtons. Dis. 7, 89–98. doi: 10.3233/jhd-189001
Takayama, K. (2020). In vitro and animal models for SARS-CoV-2 research. Trends Pharmacol. Sci. 41, 513–517. doi: 10.1016/j.tips.2020.05.005
Verma, N. K., Fazil, M. H. U. T., Ong, S. T., Chalasani, M. L., Low, J. H., and Kelleher, D. (2017). A Lymphocyte Permeating Chimeric Oligonucleotide, Methods And Uses Thereof WO2017184082A1. International Application Published Under the Patent Cooperation Treaty (PCT) on 26 October 2017.
Wu, F., Zhao, S., Yu, B., Chen, Y. M., Wang, W., Song, Z. G., et al. (2020). A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269.
Xu, H., Zhong, L., Deng, J., Peng, J., Dan, H., Zeng, X., et al. (2020). High expression of ACE2 receptor of 2019-nCoV on the epithelial cells of oral mucosa. Int. J. Oral Sci. 12:8. doi: 10.1038/s41368-020-0074-x
Yan, R., Zhang, Y., Li, Y., Xia, L., Guo, Y., and Zhou, Q. (2020). Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448. doi: 10.1126/science.abb2762
Yoshida, T., Naito, Y., Yasuhara, H., Sasaki, K., Kawaji, H., Kawai, J., et al. (2019). Evaluation of off-target effects of gapmer antisense oligonucleotides using human cells. Genes Cells 24, 827–835. doi: 10.1111/gtc.12730
Zhang, H., Penninger, J. M., Li, Y., Zhong, N., and Slutsky, A. S. (2020). Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Inten. Care Med. 46, 586–590. doi: 10.1007/s00134-020-05985-9
Keywords: COVID-19 (2019-nCoV), GapmeR, angiotensin-converting enzyme 2, nasal delivery, therapy
Citation: Verma NK, Fazil MHUT, Duggan SP and Kelleher D (2020) Combination Therapy Using Inhalable GapmeR and Recombinant ACE2 for COVID-19. Front. Mol. Biosci. 7:197. doi: 10.3389/fmolb.2020.00197
Received: 18 May 2020; Accepted: 22 July 2020;
Published: 07 August 2020.
Edited by:
Pier Paolo Piccaluga, University of Bologna, ItalyReviewed by:
Sohrab Zafar Khan, University of Virginia, United StatesVeronika Butin-Israeli, Northwestern University, United States
Copyright © 2020 Verma, Fazil, Duggan and Kelleher. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Navin Kumar Verma, bmt2ZXJtYUBudHUuZWR1LnNn; dmVybWFuQHRjZC5pZQ==
 Navin Kumar Verma
Navin Kumar Verma Mobashar Hussain Urf Turabe Fazil
Mobashar Hussain Urf Turabe Fazil Shane P. Duggan
Shane P. Duggan Dermot Kelleher
Dermot Kelleher