- 1 Laboratory of Molecular Genetics and Histocompatibility, Inserm U613, Brest, France
- 2 Laboratory of Molecular Genetics and Histocompatibility, Centre Hospitalier Universitaire Brest, Brest, France
- 3 Faculté de Médecine de Brest et des Sciences de la Santé, IFR148, Brest University, Brest, France
- 4 Department of Cell and Developmental Biology, University College London, London, UK
Mutations in the homeobox transcription factor ARX have been found to be responsible for a wide spectrum of disorders extending from phenotypes with severe neuronal migration defects, such as lissencephaly, to mild forms of intellectual disabilities without apparent brain abnormalities, but with associated features of dystonia and epilepsy. Arx expression is mainly restricted to populations of GABA-containing neurons. Studies of the effects of ARX loss of function, either in humans or mutant mice, revealed varying defects, suggesting multiple roles of this gene in brain patterning, neuronal proliferation and migration, cell maturation and differentiation, as well as axonal outgrowth and connectivity. However, to date, little is known about how Arx functions as a transcription factor or which genes it binds and regulates. Recently, we combined chromatin immunoprecipitation and mRNA expression with microarray analysis and identified approximately 1000 gene promoters bound by Arx in transfected neuroblastoma N2a cells and mouse embryonic brain. To narrow the analysis of Arx targets to those most likely to control cortical interneuron migration and/or differentiation, we compare here our data to previously published studies searching for genes enriched or down-regulated in cortical interneurons between E13.5 and E15.5. We thus identified 14 Arx-target genes enriched (Cxcr7, Meis1, Ppap2a, Slc12a5, Ets2, Phlda1, Egr1, Igf1, Lmo3, Sema6, Lgi1, Alk, Tgfb3, and Napb) and 5 genes specifically down-regulated (Hmgn3, Lmo1, Ebf3, Rasgef1b, and Slit2) in cortical migrating neurons. In this review, we present these genes and discuss how their possible regulation by Arx may lead to the dysfunction of GABAergic neurons, resulting in mental retardation and epilepsy.
Introduction
The cerebral cortex is formed of two broad classes of neurons: excitatory projection neurons, which primarily use glutamate as a neurotransmitter, and inhibitory GABA-containing interneurons. Whereas projection neurons are generated directly from radial glial cells in the germinal ventricular zone (VZ; reviewed in Nadarajah and Parnavelas, 2002; Kriegstein and Noctor, 2004) or indirectly from intermediate progenitors in the subventricular zone (SVZ; Noctor et al., 2001, 2004), the majority of interneurons are generated in the ventral forebrain and reach the cortex by tangential migration along well-defined streams, guided by a combination of chemoattractive and repulsive cues (reviewed by Marín and Rubenstein, 2003; Métin et al., 2006). In rodents, cortical interneurons have been reported to arise predominantly from themedial ganglionic eminences (MGE) orcaudal ganglionic eminences (CGE) and, to a smaller extent, from the embryonic preoptic area (Lavdas et al., 1999; Nery et al., 2002; Yozu et al., 2005; Gelman et al., 2009). They then, display diverse migratory behaviors before establishing themselves in an inside-out manner in the cortical plate (CP), similar to pyramidal neurons (Corbin et al., 2001; Ang Jr. et al., 2003; Tanaka et al., 2006, 2009). In addition, a few studies have suggested that, at least in primates and humans, a significant number of cortical GABAergic neurons may also be generated from progenitors of the VZ and SVZ in the dorsal forebrain (Letinic et al., 2002; Petanjek et al., 2009; Jakovcevski et al., 2011; Zecevic et al., 2011). Over the last 15 years, mutant mouse analysis and RNAi-mediated gene knock-down studies have identified several genes encoding transcription factors such as Dlx1, Dlx2, Dlx5 (Distal-less), Arx (Aristaless-related homeobox), Nkx2.1 (NK2 homeobox 1), or Lhx6 (LIM homeobox protein 6) important for the correct generation, migration or differentiation of these GABAergic neurons (reviewed by Hernandez-Miranda et al., 2010).
Over the past decade, numerous studies have demonstrated that a dysfunction of the GABAergic system is responsible for a large variety of neurodevelopmental disorders (Baulac et al., 2001; Levitt et al., 2004; Di Cristo, 2007). In particular, it is now well accepted that there is a connection between altered neuronal positioning, usually due to defects in migration, and susceptibility to epilepsy. Moreover, subtle alterations in interneuron position and/or function have been reported to contribute to disorders such as dyslexia, schizophrenia, autism, and mental retardation (Galaburda et al., 2006; Nakazawa et al., 2011). It is thus important to better characterize the molecular mechanisms involved in interneuron generation, migration and differentiation through, for example, the analysis of genes important for interneuron development.
The ARX gene was first identified in 2002 as being responsible for a rare and severe cortical malformation in human, the X-linked lissencephaly associated with abnormal genitalia (XLAG), typically characterized by severe congenital or postnatal microcephaly, complete disorganization of cortical layers (lissencephaly), agenesis of the corpus callosum, midbrain malformations, and neonatal-onset intractable epilepsy (Kitamura et al., 2002). Interestingly, a complete absence of interneurons was described in the cortex of these patients (Bonneau et al., 2002; Forman et al., 2005; Okazaki et al., 2008; Marcorelles et al., 2010). Similarly, aberrant migration and differentiation of GABAergic interneurons in the ganglionic eminences and neocortex were described in male embryonic Arx mutant mice (Kitamuraet al.,2002; Colombo et al., 2007). Since then, ARX has been associated with no less than 10 different syndromes ranging from phenotypes characterized by severe neuronal migration defects, to mild or moderate forms of intellectual disability without apparent brain abnormalities, but often with dystonia and epilepsy (reviewed in Friocourt and Parnavelas, 2010; Shoubridge et al., 2010).
Although Arx is expressed in several structures including the brain, pancreas, developing testes, heart, skelet al muscle, and liver (Bienvenu et al., 2002; Collombat et al., 2003; Biressi et al., 2008), the most striking consequences of its loss of function concern the brain and testes in both mouse and human. During development, Arx is expressed early in telencephalic structures, and more specifically in the mantle zones of the developing lateral ganglionic eminences (LGE) and MGE in the basal forebrain. In the developing cortex, its expression is observed in progenitor cells of the VZ as well as in migrating interneurons, but not in radially migrating cells (Colombo et al., 2004; Poirier et al., 2004; Friocourt et al., 2006). The extensive cellular co-localization between Arx and GABA in mouse and human brain, as well as the absence of interneurons documented in the cortex of XLAG patients and Arx mutant mice have led to propose “interneuronopathy” as a new term to describe the group of pathologies ARX is responsible for (Kato and Dobyns, 2005).
Arx encodes a homeobox transcription factor that has been found to contribute to most fundamental processes of brain development: patterning, neuroblast proliferation, neuronal migration and differentiation as well as axonal outgrowth and connectivity (Kitamura et al., 2002; Cobos et al., 2005; Colombo et al., 2007; Colasante et al., 2008; Friocourt et al., 2008), but the signaling pathways controlled by this gene are still unknown. Although two gene expression profile analyses comparing E14.5 wild-type and Arx mutant ventral telencephalic tissues have been published in mouse, very few targets for this transcription factor have been described, and only three (Lmo1, Ebf3, and Shox2) were found to be direct (Fulp et al., 2008; Colasante et al., 2009). For this reason, we recently performed chromatin immunoprecipitation in Arx-transfected neuroblastoma cells (N2a) or E15.5 mouse embryonic brain, followed by hybridization to mouse promoter arrays (ChIP-chip) in order to identify new direct targets of Arx (Quillé et al., 2011). We then examined by transcriptomic experiments whether these genes were differently regulated following the ectopic expression of Arx in N2a cells or its knock-down in mouse ventral telencephalon (Quillé et al., 2011). For the latter, we used the publicly available microarray data generated by Fulp et al. (2008) and Colasante et al. (2009), comparing gene expression between basal telencephalon of E14.5 Arx knock-out and wild-type mice. Out of a total of 1006 genes which promoters were found to be enriched in Arx-immunoprecipitates, approximately 24% showed expression changes following Arx overexpression or knock-down (Quillé et al., 2011).
In order to provide novel insights into genetic networks regulated by Arx and specifically controlling the development of GABAergic neurons, we took advantage of previously published studies comparing the level of expression of genes between, on one hand, cortical interneurons (Dlx5/6-derived cells) and cortical non-interneurons (non-Dlx5/6-derived cells) in the telencephalon of E13.5, E14.5, and E15.5 mice (Batista-Brito et al., 2008; Marsh et al., 2008) and, on the other hand, genes expressed by GABAergic neurons (Gad67-positive cells) in the ganglionic eminence versus those migrating in the cortex at the same stages (Marsh et al., 2008; Faux et al., 2010). As Arx is strongly expressed in cortical migrating interneurons, we decided to compare these datasets with ours and present here a list of ChIP-positive (Arx-bound) and regulated genes (Quillé et al., 2011) that are enriched or down-regulated in migrating cortical interneurons and, thus, are good candidates to control, positively or negatively, molecular mechanisms involved in interneuron migration and/or differentiation (Table 1 ).
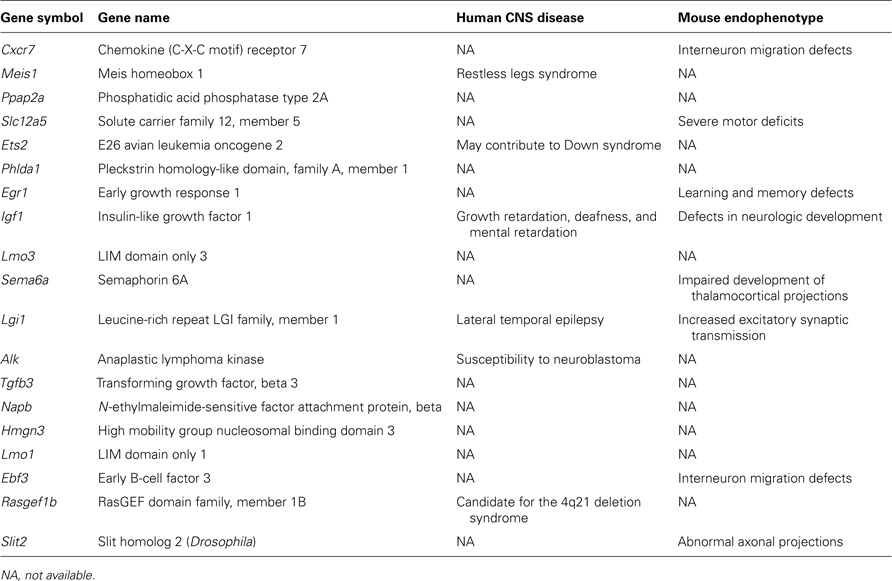
Table 1. List of candidate genes regulated by Arx and controlling cortical interneuron migration and/or differentiation.
Identification of Candidate Genes Positively Regulating Cortical Interneuron Migration and/or Differentiation
As shown in Table 2, we identified 14 ChIP-positive genes (genes whose promoters were enriched in Arx-immunoprecipitates) that showed deregulation, either following Arx overexpression in N2a cells or in Arx mutant subpallium (Quillé et al., 2011), and appeared enriched in migrating cortical interneurons (Batista-Brito et al., 2008; Marsh et al., 2008; Faux et al., 2010). Thus, these genes are good candidates to positively control molecular mechanisms involved in cortical interneuron migration and/or differentiation.
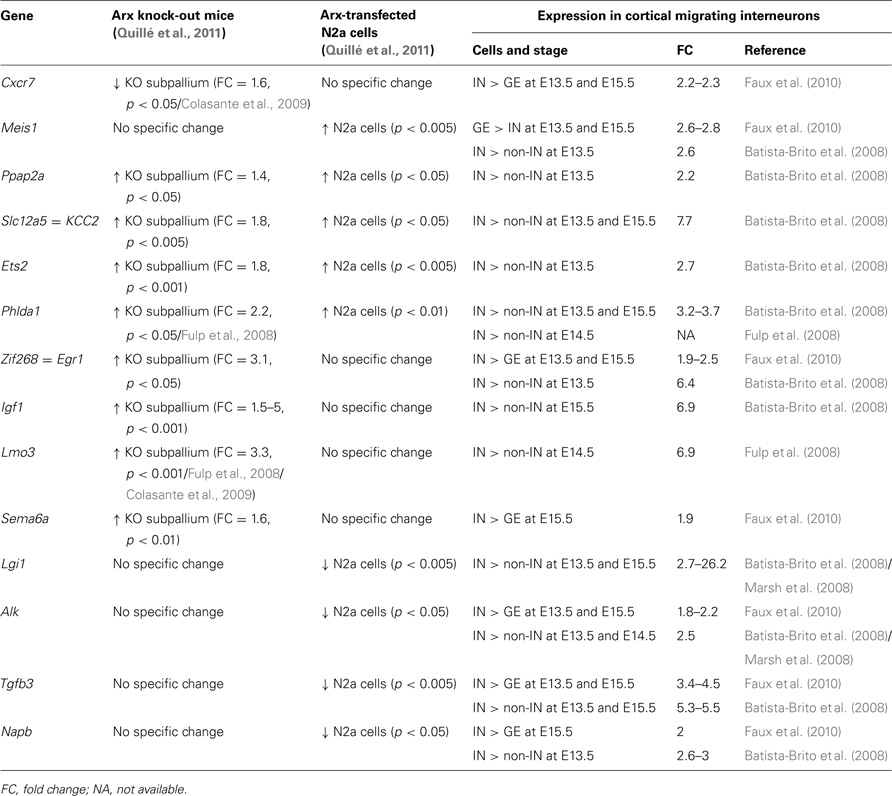
Table 2. Examples of Arx-bound and regulated genes that are enriched in migrating cortical interneurons (IN) compared to neurons in ganglionic eminences (GE) and cortical non-interneurons (non-IN).
In vitro and in vivo studies have previously demonstrated that Arx can act as both transcriptional repressor and activator (Seufert et al., 2005; McKenzie et al., 2007; Fullenkamp and El-Hodiri, 2008). Accordingly, eight of Arx putative direct targets presented in Table 2 (Egr1, Igf1, Lmo3, Sema6a, Lgi1, Alk, Tgfb3, and Napb) show either a significant reduction in expression as a consequence of increased Arx levels or increased expression in Arx knock-out brains (Table 2). However, four genes (Ppap2a, Slc12a5, Ets2, and Phlda1) show similar changes in expression following Arx overexpression or knock-down, thus making it difficult to determine whether Arx normally activates or represses these genes in migrating interneurons. We had previously described similar results for some other genes and suggested that Arx may switch from repressor to activator depending on the developmental stage, the molecular context (posttranslational modifications such as phosphorylation or the presence of cofactors and/or chromatin-modifying enzymes) or the cellular model used (Quillé et al., 2011). On the contrary, two genes look positively regulated by Arx (Table 2).
One such gene positively regulated by Arx is Cxcr7 (CXC chemokine receptor 7), which role in interneuron migration has recently been described (Sánchez-Alcañiz et al., 2011; Wang et al., 2011). Cxcr4 and Cxcr7 are receptors for the chemokine Cxcl12 (also called Sdf1, stromal cell-derived factor) expressed in tangentially migrating interneurons (Sánchez-Alcañiz et al., 2011; Wang et al., 2011). Cxcl12 binding to Cxcr4 triggers GαI protein-dependent signaling, whereas Cxcl12 binding to Cxcr7 activates the mitogen-activated protein kinase (MAPK) cascade (Wang et al., 2011). Cxcl12 expression in the marginal zone (MZ) and SVZ of the cortex attracts Cxcr4- and Cxcr7-expressing migrating interneurons, guiding them to tangential streams and controls the timing of the switch from tangential to radial migration to their destinations in the CP (Tiveron et al., 2006; Li et al., 2008; Lopez-Bendito et al., 2008; Wang et al., 2011). Loss of Cxcl12 signaling has been shown to induce defects in both interneuron motility and leading process morphology, as well as a premature entry into the CP, resulting in a significant decrease in their numbers in the migratory streams and an increase in the lower part of the CP (Stumm et al., 2003; Tiveron et al., 2006; Li et al., 2008; Liapi et al., 2008; Lopez-Bendito et al., 2008; Lysko et al., 2011; Sánchez-Alcañiz et al., 2011; Wang et al., 2011).
Interestingly, Cxcr7 mRNA has been shown to be down-regulated in cortical interneurons and the GE of Dlx1/2 knock-out mice (Long et al., 2009a,b). Similarly, Cxcr7 and Cxcr4 receptors are positively regulated by Lhx6 (Zhao et al., 2008). Dlx1, Dlx2, and Lhx6 have all been shown to be important for tangential interneuron migration (Anderson et al., 1997; Alifragis et al., 2004; Liodis et al., 2007). Similarly, we found in previous work that Arx binds Cxcr7 promoter (Quillé et al., 2011) and positively regulates its expression which, in turn, regulates Cxcr4 protein levels (Sánchez-Alcañiz et al., 2011). Accordingly, Cxcr4 expression was found down-regulated in Arx mutant subpallium (Fulp et al., 2008; Colasante et al., 2009). As Arx expression was shown to be regulated by several members of the Dlx family of homeobox proteins, particularly Dlx2 (Cobos et al., 2005), we may hypothesize that interneuron migration defects observed in Dlx1/2, Lhx6, or Arx mutants are, at least in part, due to abnormal Cxcl12 signaling pathway. In addition, Cxcr4-/- and Cxcr7-/- mutants have been reported to have defects in leading process morphology during migration (Wang et al., 2011), which may also be related to the cell morphology defects observed in migrating inter-neurons in the absence of Arx (Colombo et al., 2007; Friocourt et al., 2008).
Another gene enriched in cortical migrating interneurons and positively regulated by Arx is Meis1 (murine ecotropic integration site 1), a transcription factor involved in cell proliferation in retina and hematopoietic stem cell development. Although the function of this gene has not been studied in interneurons, its expression is consistent with a role in GABAergic neuron development. First detected around E10.5 in mouse ventrolateral telencephalon, it is expressed at later stages at high levels in the CGE and developing amygdala and more weakly in the MGE and LGE (Toresson et al., 2000). Interestingly, a recent paper on the effect of Meis1 knock-down in a human precursor B-cell leukemic line RS4;11 reported an impairment of cell migration, possibly due to a defect in the CXCR4/SDF-1 axis (Orlovsky et al., 2011). Further studies should be undertaken in interneurons to assess whether this gene is also important for their migration and/or differentiation. This may be a worthwhile pursuit, as this gene has been linked to the restless legs syndrome (RLS), a common neurological disorder characterized by an irresistible urge to move the legs at night. RLS is generally considered to be a central nervous system-related disorder due to reduced intracortical inhibition and is often treated by benzodiazepines, a class of sedative drugs affecting the GABA receptors.
Among the genes that appear negatively regulated by Arx is Egr1 (early growth response 1), a transcription factor that has been implicated in synaptic plasticity underlying learning and memory in mouse (Bozon et al.,2003). A recent study has reported that Egr1 overexpression in rat brain hippocampus and primary cultures of neurons regulates cytoskeleton dynamics by inactivating the phosphatase PP1 and activates cyclin-dependent kinase 5 (Cdk5) to promote phosphorylation of Tau, a microtubule-associated protein (Lu et al., 2011). This is of particular interest, as both Cdk5 and PP1 are involved in neuronal migration (Gilmore et al., 1998; Shmueli et al., 2006; Rakić et al., 2009). The observed enrichment of this gene in cortical migrating interneurons suggests that it may positively control interneuron migration and/or differentiation. Accordingly, angiopoietin-1 (Ang-1) was found to stimulate cell migration in endothelial cells by increasing expression of Egr1 (Abdel-Malak et al., 2009). In addition, it has been reported that Reelin, produced by the mitral cells of the olfactory bulb (OB), activates the MAPK/extracellular signal-regulated kinase (ERK) pathway and induces Egr-1 transcription, which in turn promotes the shift from tangential/chain to radial/individual neuronal migration, and the detachment of neurons in the rostral migratory stream (RMS; Simó et al., 2007). These findings are particularly relevant as Reelin, produced by Cajal–Retzius cells, is also important for neuronal migration in the cortex. It is thus possible that fine regulation of Egr1 expression may be necessary to control cortical interneuron migration and the switch from tangential to radial mode into the CP.
Interestingly, Egr1 expression was found up-regulated in Arx mutant mice (Table 2), suggesting that Arx normally represses its expression inthe subpallium. Thisissomewhat contradictorywith its putative positive role in cell migration. This discrepancy may be explained by the fact that Arx is expressed in different cell types in the forebrain during development (cortical neuronal progenitors, subpallial postmitotic cells, and cortical migrating interneurons) and may thus differentially regulate the same gene depending on the developmental stage, the biological context or the molecular environment. It is thus possible that Arx normally down-regulates Egr1 in the subpallium in order to keep these cells undifferentiated, but up-regulates it in cortical migrating interneurons. Alternatively, Egr1 up-regulation in Arx mutant mice may just be the result of the accumulation of cells that failed to exit the subpallium in the absence of Arx.
Another gene that may have a similar role is Igf1 (Insulin-like growth factor-1), which encodes a growth factor highly expressed in embryonic brain. It has been implicated in several processes such as projection neuron growth, dendritic arborization, synaptogenesis, and adult hippocampal neurogenesis. In human, mutations in this gene are responsible for intrauterine and postnatal growth retardation with sensorineural deafness and intellectual deficit (Woods et al., 1996). In mouse, disruption of Igf1 results in reduced brain size, hypomyelination, and loss of hippocampal granule and striatal parvalbumin-containing neurons (Beck et al., 1995). More recently, this gene was found to promote neuronal migration and positioning in theOB and the incorporationof neuroblasts from the SVZ to the RMS (Hurtado-Chong et al., 2009). Igf1-/- mice display altered pattern of OB layering with, in particular, a misplacement of glutamatergic neurons in the mitral cell layer (ML) and the external plexiform layer (EPL), a depletion of several populations of interneurons in the glomerular layer and the EPL, and an accumulation of neuroblasts in the postnatal and adult SVZ (Hurtado-Chong et al., 2009). Interestingly, these authors observed that a number of misplaced glutamatergic neurons in the ML had failed to radially re-orient their cell bodies, maintaining a relatively tangential orientation. These results suggest that, in the absence of Igf1, there was a defect in the switch from tangential to radial migration in the ML of the OB (Hurtado-Chong et al., 2009). In addition, they showed that Dab1 activity was necessary for Igf1 action suggesting that, similar to Egr1, its role in interneuron migration may, at least partly, intersect with the Reelin signaling pathway (Hurtado-Chong et al., 2009). Finally, a role for Igf1 has also been demonstrated in the GABAergic differentiation of neural precursors through the activation of the Pdk1/Akt pathway, which induces Mash1 expression (Oishi et al., 2009). The fact that Pdk1 was found repressed in Arx-overexpressing N2a cells (Quillé et al., 2011) confirms the involvement of Arx in this pathway and may explain the reason why some GABAergic neurons, and especially NPY+ and ChAT+ populations, were found strongly reduced in Arx mutant forebrains (Kitamura et al., 2002, 2009; Colombo et al., 2007; Price et al., 2009).
Similar to Egr1and Igf1, Arx knock-out mice display increased expression of the Sema6a (semaphorin 6a) gene in the subpallium (Table 2). Like other semaphorins, Sema6a is better known for its function in thalamocortical axon guidance, but it has also been reported to play a role in cell migration during cerebellum development. This protein is selectively expressed by postmitotic granule cells during their tangential migration in the deep external granule cell layer, but not during their radial migration (Kerjan et al., 2005). The analysis of granule cell migration and neurite outgrowth in cerebellar explants of mutant mice for Sema6a, or for its receptor Plexin a2, revealed that Sema6a controls the switch from tangential to radial migration by regulating centrosome–nucleus coupling and cellular translocation (Kerjan et al., 2005; Renaud et al., 2008). The role of Sema6a in cortical interneuron migration has never been studied, but this gene is probably a good candidate as other members of the semaphorin family have been shown to be important for the guidance of cortical interneurons as they migrate in the ventral telencephalon (Marín et al., 2001; Hernandez-Miranda et al., 2011).
Although no change of Lgi1 (leucine-rich, glioma-inactivated) was detected in Arx mutant subpallium, this gene was found to be down-regulated following Arx transfection in N2a cells. Interestingly, mutations of this gene result in autosomal dominant lateral temporal epilepsy (Kalachikov et al., 2002). Enhanced excitatory synaptic transmission through increased release of glutamate has been suggested as a basis for the seizure phenotype. Lgi1, which encodes a secreted protein involved in postnatal glutamatergic synapse development, has not been studied in interneurons, but its strong enrichment in these cells compared to cortical non-interneurons (Table 2, fold change = 26.2 at E13.5), suggests an important role in interneuron development and possibly migration, as Lgi1 has been shown to regulate cell mobility through the formation of stress fibers in glioblastoma cells (Kunapuli et al., 2010).
At the start of corticogenesis, GABAergic interneurons enter the neocortex at the level of the preplate layer (PPL) and IZ. Loss of the upper stream(s) was reported at E14.5 and E18.5 in Arx knock-out mice, but the stream in the IZ/SVZ looked intact (Kitamura et al., 2002; Colombo et al., 2007). Interestingly, Lgi1 has recently been found up-regulated in cortical migrating interneurons of the PPL compared to the IZ layer at E14.5 (Antypa et al., 2011). It is thus possible that regulation of Lgi1 expression by Arx may explain the loss of the PPL migratory stream, but not the IZ stream in Arx mutant cortex (Kitamura et al., 2002).
Tgfb3 (transforming growth factor, beta 3) encodes a member of the TGF-β family of proteins. They act by stimulating specific membrane serine/threonine receptor complexes resulting in the phosphorylation and activation of Smad transcription factors (regulatorySmad, R-smad). Smad1,Smad5,and Smad8 are mainly activated by bone morphogenetic protein (BMP) and growth differentiation protein (GDF) receptors, while Smad2 and Smad3 are substrates for TGF-β, activin, and Nodal receptors. Once activated, the R-Smads accumulate in the nucleus where they associate with Smad4, a common partner for all R-Smads to form transcription complexes. Interestingly, in addition to Tgfb3, we found that Arx binds to Smad1 and Smad4 regulatory sequences in both neuroblastoma cells and E15.5 mouse embryonic brain, and that Arx overexpression in N2a cells results in Smad1 down-regulation (Quillé et al., 2011), confirming a possible involvement of Arx in the TGF-β signaling pathway. Maira et al. (2010) have recently investigated a possible role for the TGF-β superfamily pathway in telencephalic GABAergic neuron development. They first showed that Smad1, Smad2, Smad4, and Smad5 are expressed in the sub-pallium at E15.5, where they exhibit expression levels similar to those of Dlx genes. They also observed the presence of activated Smad1 and Smad2 in the developing basal ganglia and in cortical migrating interneurons. In addition, inhibition of TGF-β signaling by the use of dominant-negative forms of Smad proteins in the basal ganglia, impairs tangential migration of cortical interneurons: Smad1 or Smad2 dominant negative mutants only partially blocked migration, whereas inhibition of Smad4 completely abrogated the migration of electroporated cells to the cortex, thus providing evidence that both branches of the TGF-β signaling pathway are important for cortical interneuron migration (Maira et al., 2010).
It is interesting to note that Arx transfection of N2a cells induces down-regulation of Smad1, but that the level of expression of Smad1 and Smad4 was unchanged in Arx knock-out, similar to Dlx1/2 mutant mice (Mairaet al.,2010). It had thus been suggested that expression of most TGF-β superfamily genes was not downstream of Dlx. Interestingly, we found that Arx up-regulates two more components of the TGF-β superfamily, Smad3 and Bmper (BMP binding endothelial regulator). The latter was recently identified as a Dlx5 direct target in inner ear (Sajan et al., 2011). As Arx expression is ectopically induced by forced expression of Dlx1, Dlx2, or Dlx5 in mouse dorsal thalamus (Cobos et al., 2005), it appears very likely that the Dlx and Arx genes act through the same components of the TGF-β pathway to regulate interneuron migration and/or differentiation.
The Slc12a5 gene (solute carrier family 12, member 5, also called KCC2, potassium/chloride cotransporter 2) was found up-regulated in both Arx mutant subpallium and transfected N2a cells, confirming that it is a direct target of Arx. Up-regulation of Slc12a5 by migrating interneurons results in the termination of their tangential migration and allows the radial sorting of different populations of tangentially migrating interneurons, resulting in their layer-specific integration into the emerging cortical network (Bortone and Polleux, 2009; Miyoshi and Fishell, 2011). It is thus likely that Arx, through Slc12a5 expression, regulates the timing of migration and the final organization of interneurons in cortical layers.
There is little information available concerning the other genes found enriched in migrating cortical interneurons. As Slc12a5, the Phlda1 (pleckstrin homology-like domain, family A, member 1) gene was found up-regulated in both Arx mutant subpallium and transfected N2a cells (Table 2 ). This gene encodes a proline– histidine rich nuclear protein which is specifically induced by Igf1. Although it has never been studied in interneuron development, it has recently been reported that siRNA-mediated suppression of Phlda1 in colon cancer cells inhibited cell migration (Sakthianandeswaren et al., 2011). Similarly, Ppap2a (phosphatidic acid phosphatase type 2A) encodes an enzyme that converts phosphatidic acid to diacylglycerol and functions in de novo synthesis of glycerolipids. Ppap2a has been shown to reduce platelet-derived growth factor (PDGF)- and lysophosphatidic acid-induced migration of embryonic fibroblasts (Long et al., 2006). Concerning Ets2 (E26 avian leukemia oncogene 2), this gene encodes a transcription factor located on chromosome 21 which has been suggested to contribute to Down syndrome’s phenotype. Its role is not yet well understood. Finally, the Lmo3 (LIM-only protein 3) gene was consistently found up-regulated in Arx mutant subpallium (Colasante et al., 2008; Fulp et al., 2008; this study). Forced expression of this gene in the MGE of E14.5 brain slices has no effect on neuronal migration (Colasante et al., 2009), suggesting that Lmo3 expression is not required for the initiation of migration. However, to exclude a definite role in migration, it would be interesting to knock down this gene specifically during migration. In addition, Lmo3 has been reported to form a complex with HEN2 and induce Mash1 expression (Isogai et al., 2011). Although these observations were made in neuroblastoma cells, this is potentially relevant as Mash1 is known to be specifically expressed in the ventral part of the telencephalon and contributes to the generation of GABAergic neurons.
In contrast, the Alk (Anaplastic lymphoma kinase) gene seems to be repressed by Arx. This gene encodes a receptor tyrosine kinase (RTK) that has been shown to concentrate in postsynaptic domains. Its role in interneuron development has never been studied. Similarly, Napb encodes soluble N-ethylmaleimide-sensitive factor (NSF) attachment protein beta (beta Snap) involved in SNAP receptor (SNARE)-mediated vesicle trafficking and synapse formation. The mode of migration used by cortical interneurons is very similar to that of other types of migrating cells. They first extend a leading process in the direction of migration, followed by nuclear translocation and the retraction of the trailing process (Métin et al., 2006). During migration, interneurons show highly dynamic branching with changes in the orientation of the leading process, migration in all directions within streams and the use of different substrates. The role of vesicular trafficking in processes such as endocytosis to regulate substrate attachment or detachment, or membrane cycling from the leading and trailing process is thus very important (Skalski et al., 2010; Shieh et al., 2011).
Identification of Candidate Genes Negatively Regulating Cortical Interneuron Migration and/or Differentiation
We found five Arx-bound genes specifically down-regulated in migrating cortical interneurons compared to neurons of the GE or cortical non-interneurons (Table 3). Thus, these genes may be necessary for cortical progenitor proliferation and/or pyramidal neuron development. Alternatively, as Arx negatively regulates most of these genes, they may repress interneuron migration and/or differentiation.
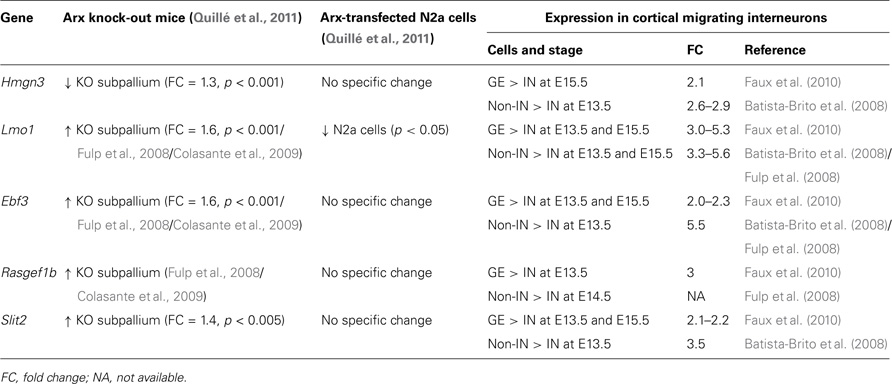
Table 3. Examples of Arx-bound and regulated genes that are down-regulated in migrating cortical interneurons (IN) compared to neurons in the ganglionic eminences (GE) and cortical non-interneurons (non-IN).
Only one gene, Hmgn3 (high mobility group nucleosomal binding domain 3) appears to be positively regulated by Arx (Table 3). This gene encodes a nucleosome binding protein that affects chromatin function. It has been reported to induce the expression of the glycine transporter 1 gene (Glyt1), a membrane transporter that regulates glycine concentration in synaptic junctions (West et al., 2004). In addition, Hmgn3 expression is very similar to that of the glial fibrillary acidic protein (GFAP), suggesting that it might play a role in astrocyte function.
The four other genes seem to be repressed by Arx (Table 3) and may thus normally inhibit cortical interneuron migration and/or differentiation. Accordingly, a recent study has shown that ectopic expression of one of these genes, Ebf3(early B-cell factor 3), in basal ganglia severely interferes with tangential migration of GABAergic interneurons to the cortex. On the opposite, knock-down experiments targeting this gene in Arx mutant mice were found to rescue, at least partially, neuronal migration defects (Colasante et al., 2009). These results confirm the implication of this transcription factor in neuronal migration, although the underlying mechanisms are still unknown. Ebf3 is also known to be expressed in the developing hindbrain and spinal cord where it promotes neuronal differentiation and radial migration (Garcia-Dominguez et al., 2003). This gene is normally not (or only marginally) expressed in the developing telencephalon, but it was found strongly misexpressed in the MGE and present in the LGE of Arx mutant mice (Colasante et al., 2009). Interestingly, it was also found up-regulated in Dlx1/2 mutant subpallium (Fulp et al., 2008), suggesting that Ebf3 may belong to a pathway that is common to Dlx and Arx genes (Cobos et al., 2005).
The Rasgef1b (RasGEF domain family, member1b)gene encodes a guanine nucleotide exchange factor (GEF) for Ras family proteins, and several members have been reported as key regulators of actin and microtubule dynamics during dendrite or spine structural plasticity. GEF proteins stimulate the intrinsic GDP/GTP exchange activity of Ras and promote the formation of active Ras-GTP, which in turn controls diverse signaling networks important for the regulation of cell proliferation, survival, differentiation, vesicular trafficking, or gene expression. Rasgef1b has been shown to function as a very specific exchange factor for Rap2, which is implicated in the regulation of cell adhesion, the establishment of cell morphology, and the modulation of synapses in neurons (Yaman et al., 2009). Normally located in the differentiated striatal mantle, Rasgef1b was found misexpressed in the SVZ of the basal ganglia of Arx mutant mice (Colasante et al., 2009). Interestingly, it is present in some migrating cortical interneurons, at least in the IZ layer, compared to the PPL (Antypa et al., 2011) suggesting that, contrary to Ebf3, the expression of this gene is compatible with migration. In addition, whereas it was found up-regulated in Arx mutant subpallium, it appeared down-regulated in Dlx1/2 mutants (Fulp et al., 2008), suggesting that the role of this gene in neuronal migration may be more complex than just the control of cell motility. However, since this gene has been linked to severe mental retardation, absent speech, distinctive facial features, and severe growth delay (Bonnet et al., 2010), the study of this gene is probably worth pursuing.
Similarly, Lmo1 (LIM domain only 1) gene, consistently found repressed by Arx in several previous studies (Fulp et al., 2008; Colasante et al., 2009; Quillé et al., 2011), is found down-regulated in migrating cortical interneurons (Table 3). Interestingly, we found that Arx also regulates other members of this family of LIM-containing transcription factors. Arx directly binds to Lmo3 and Lmo4 promoters and down-regulates their expression in N2a-transfected cells as shown by gene expression profile analysis and quantitative RT-PCR (Quillé et al., 2011 and data not shown). Accordingly, Lmo1, Lmo3, and Lmo4 genes were found up-regulated in Arx mutant subpallium (Fulp et al., 2008; Colasante et al., 2009). These three genes are normally never, or only marginally, transcribed in the ventral telencephalon, but were found strongly expressed in Arx mutant MGE and LGE/CGE (Colasante et al., 2009). It is thus possible that they play important roles in interneuron differentiation.
The last gene found to be down-regulated in migrating cortical interneurons is Slit 2 (slit homolog 2, Drosophila). This chemorepulsive ligand and its receptors of the Robo family are expressed in the developing and adult brain and have been reported to regulate commissural axon guidance and axonal branching as well as cell migration. Double labeling experiments have shown that the vast majority of cortical interneurons express Robo1 and Robo2 and that Slit proteins are present in a complementary manner in the proliferative zones of the ventral telencephalon and in the septum during early and mid-phases of corticogenesis (Andrews et al., 2008). Evidence from in vitro experiments indicates that migration of Robo-expressing interneurons is initiated by the chemorepulsive activity of Slit secreted from the VZ of the LGE. In the developing cortex, Slit1 is robustly expressed in the CP, Slit3 is restricted to the MZ, while Slit2 is weakly expressed in the VZ. The presence of a putative Slit gradient along the interneuron migratory routes suggests that Slit/Robo signaling may also play a role in the positioning of the different tangential migratory paths within the developing cortex. However, it has been reported that migration of cortical interneurons is normal in Slit1/Slit2 double knock-out mice, prompting speculation that Slits do not play a major role in tangential migration, although they seem to be important regulators of neuronal positioning within the basal telencephalon (Marín et al., 2003). Nevertheless, the Slit/Robo signaling pathway has also been shown to regulate cell division and interneuron morphology (Andrews et al., 2008).
Conclusion
The Arx gene has been shown to play different roles in brain development: patterning, cell proliferation, migration and differentiation as well as axonal outgrowth and connectivity (Kitamura et al., 2002; Cobos et al., 2005; Colombo et al., 2007; Colasante et al., 2008; Friocourt et al., 2008). Accordingly, we recently identified a high number of putative direct targets for this transcription factor (Quillé et al., 2011), several of them being involved in cell proliferation, cytoskeleton dynamic regulation, axonal guidance, or neurotransmission. To narrow the analysis of Arx targets to those most likely to control cortical interneuron migration, we compared our data to previously published studies in search of genes enriched or down-regulated in cortical migrating interneurons. We found several Arx-bound and regulated genes that may be good candidates to explain the severe defects of interneuron migration observed in the absence of Arx in both human and mice. In particular, we identified a few genes (Igf1, Egr1, Cxcr7, Sema6a, and Slc12a5) involved in the control of the mode of migration or the switch from tangential to radial migration. This is of particular interest in the light of our previous observation, that Arx overexpression in radially migrating cells promotes tangentially orientated migration in the SVZ and lower IZ, although these cells do not express GABAergic markers (Friocourt et al., 2008; Friocourt and Parnavelas, 2010).
As previously reported (Marsh et al., 2008), several of the genes that were enriched in migrating cortical interneurons are involved in synaptic transmission (Egr1, Igf1, Lgi1, Alk, Napb, Glyt1,…), probably due to the fact that migration is known to be partly controlled by paracrine transmitter release requiring synaptic vesicle proteins. One of these genes, Slc12a5, has been identified as a direct target for Arx (Quillé et al., 2011), thus implicating Arx in the control of the capacity for migrating cells to respond to GABA and switch from depolarization to hyperpolarization (Ben-Ari, 2002), resulting in a voltage-sensitive, calcium-mediated reduction of interneuron motility. Interestingly, we identified several putative direct targets for Arx which are related to calcium release (Caln1, Calu, Cacng4, Camkv, Cacna2d1, the latter being specifically down-regulated in cortical migrating interneurons) that may be good candidates for a role during interneuron migration.
It is not always clear whether the genes we identified are normally activated or repressed by Arx as a few genes in Tables 2 and 3 showed the same the type of regulation in Arx-overexpressing N2a cells and in knock-out brains. These apparent discrepancies may have several explanations. First, it is important to note that the microarray experiments on Arx knock-out brains were performed from ventral telencephalon (Fulp et al., 2008; Colasante et al., 2009), thus leaving out possibly different gene expression changes during interneuron migration and differentiation in the cortex. Second, some changes in gene expression may be too low to be detected in microarray experiments. We previously observed that some Arx-bound genes such as Gabrb3, Lmo3, or Cdh2 showed a change of expression in qRT-PCR experiments following Arx expression in N2a cells but not on microarrays (Quillé et al., 2011). Third, Arx is expressed in different cell types during brain development and in adult, and it is thus possible that, depending on the presence or the absence of specific cofactors it may activate one gene at some stage and repress the same gene at a different time or in a different cell. Fourth, changes or the absence of change in a given gene expression may be the result of some compensatory effect of Arx knock-out or overexpression.
In addition, the disadvantage of using transfected N2a cells is that, as Arx is normally not expressed in these cells, it may lack binding partners and/or cofactors necessary to regulate the expression of certain genes. On the other hand, as we performed gene expression analysis relatively shortly (2 days) after Arx transfection in N2a cells, it is likely that the changes we see are a direct effect of Arx expression. For example, important genes for cortical development such as Dab1 or Dclk1 were found repressed following Arx transfection in N2a cells (Quillé et al., 2011). Dab1 is an intracellular adaptor which is expressed in cells that respond to Reelin and is required for correct radial migration of cortical pyramidal neurons (Howell et al., 1997; Ware et al., 1997). Expression of Dab1 in early-born interneurons has also been shown to be required for correct cortical layering (Hammond et al., 2006). Similarly, Dclk1 (doublecortin-like kinase 1) is necessary for correct migration of both pyramidal cells and interneurons (Deuel et al., 2006; Koizumi et al., 2006; Friocourt et al., 2007). Thus, gene expression in transfected N2a cells brings new and complementary information to expression studies performed in Arx mutant subpallium (Fulp et al., 2008; Colasante et al., 2009).
In conclusion, we identified here a set of candidate genes important for cortical interneuron migration and/or differentiation. Some of these genes have already been described to play a role in neuronal migration, but others have never been investigated in this context. Further studies will definitely bring new information about their function and how their possible regulation byArx may lead to the dysfunction of GABAergic neurons, resulting in mental retardation and epilepsy.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We wish to thank Drs C. Faux, W. Andrews, M. Antypa, R. Batista-Brito, and G. Fishell for accepting to share their microarray data. We also acknowledge the Inserm, la Fondation Jérome Lejeune, la Fondation Gaetan Saleun, and le Fonds Européen de Développement Régional for support of the work on ARX.
References
Abdel-Malak, N. A., Mofarrahi, M., Mayaki, D., Khachigian, L. M., and Hussain, S. N. (2009). Early growth response-1 regulates angiopoietin-1-induced endothelial cell proliferation, migration, and differentiation. Arterioscler. Thromb. Vasc. Biol. 29, 209–216.
Alifragis, P., Liapi, A., and Parnavelas, J. G. (2004). Lhx6 regulates the migration of cortical interneurons from the ventral telencephalon but does not specify their GABA phenotype. J. Neurosci. 24, 5643–5648.
Anderson, S. A., Eisenstat, D. D., Shi, L., and Rubenstein, J. L. (1997). Interneuron migration from basal forebrain to neocortex: dependence on Dlx genes. Science 278, 474–476.
Andrews, W., Barber, M., Hernadez-Miranda, L. R., Xian, J., Rakic, S., Sundaresan, V., Rabbitts, T. H., Pannell, R., Rabbitts, P., Thompson, H., Erskine, L., Murakami, F., and Par-navelas, J. G. (2008). The role of Slit-Robo signaling in the generation, migration and morphological differentiation of cortical interneurons. Dev. Biol. 313, 648–658.
Ang, E. S. Jr., Haydar, T. F., Gluncic, V., and Rakic, P. (2003). Four-dimensional migratory coordinates of GABAergic interneurons in the developing mouse cortex. J. Neurosci. 23, 5805–5815.
Antypa, M., Faux, C., Eichele, G., Parnavelas, J. G., and Andrews, W. (2011). Differential gene expression in migratory streams of cortical interneurons. Eur. J. Neurosci. 34, 1584–1594.
Batista-Brito, R., Machold, R., Klein, C., and Fishell, G. (2008). Gene expression in cortical interneuron precursors is prescient of their mature function. Cereb. Cortex 18, 2306–2317.
Baulac, S., Huberfeld, G., Gourfinkel-An, I., Mitropoulou, G., Beranger, A., Prud’homme, J. F., Baulac, M., Brice, A., Bruzzone R., and LeGuern, E. (2001). First genetic evidence of GABA(A) receptor dysfunction in epilepsy: a mutation in the gamma2-subunit gene. Nat. Genet. 28, 46–48.
Beck, K. D., Powell-Braxton, L., Widmer, H. R., Valverde, J., and Hefti F. (1995). Igf1 gene disruption results in reduced brain size, CNS hypomyelination, and loss of hippocampal granule and striatal parvalbumincontaining neurons. Neuron 14, 717– 730.
Ben-Ari, Y. (2002). Excitatory actions of GABA during development: the nature of the nurture. Nat. Rev. Neurosci. 3, 728–739.
Bienvenu, T., Poirier, K., Friocourt, G., Bahi, N., Beaumont, D., Fauchereau, F., Ben Jeema, L., Zemni, R., Vinet, M. C., Francis, F., Couvert, P., Gomot, M., Moraine, C., van Bokhoven, H., Kalscheuer, V., Frints, S., Gecz, J., Ohzaki, K., Chaabouni, H., Fryns, J. P., des Portes, V., Beldjord, C., and Chelly, J. (2002). ARX, a novel Prd-class-homeobox gene highly expressed in the telencephalon, is mutated in X-linked mental retardation. Hum. Mol. Genet. 11, 981–991.
Biressi, S., Messina, G., Collombat, P., Tagliafico, E., Monteverde, S., Benedetti, L., Cusella De Angelis, M. G., Mansouri, A., Ferrari, S., Tajbakhsh, S., Broccoli, V., and Cossu, G. (2008). The homeobox gene Arx is a novel positive regulator of embryonic myogenesis. Cell Death Differ. 15, 94–104.
Bonneau, D., Toutain, A., Laquerrière, A., Marret, S., Saugier-Veber, P., Barthez, M. A., Radi, S., Biran-Mucignat, V., Rodriguez, D., and Gélot, A. (2002). X-linked lissencephaly with absent corpus callosum and ambiguous genitalia (XLAG): clinical, magnetic resonance imaging, and neuropathological findings. Ann. Neurol. 51, 340–349.
Bonnet, C., Andrieux, J., Béri-Dexheimer, M., Leheup, B., Boute, O., Manouvrier, S., Delobel, B., Copin, H., Receveur, A., Mathieu, M., Thiriez, G., Le Caignec, C., David, A., de Blois, M. C., Malan, V., Philippe, A., Cormier-Daire, V., Colleaux, L., Flori, E., Dollfus, H., Pelletier, V., Thauvin-Robinet, C., Masurel-Paulet, A., Faivre, L., Tardieu, M., Bahi-Buisson, N., Callier, P., Mugneret, F., Edery, P., Jonveaux, P., and Sanlaville, D. (2010). Microdeletion at chromosome 4q21 defines a new emerging syndrome with marked growth restriction, mental retardation and absent or severely delayed speech. J. Med. Genet. 47, 377–384.
Bortone, D., and Polleux, F. (2009). KCC2 expression promotes the termination of cortical interneuron migration in a voltage-sensitive calcium-dependent manner. Neuron 62, 53–71.
Bozon, B., Davis, S., and Laroche, S. (2003). A requirement for the immediate early gene zif268 in reconsolidation of recognition memory after retrieval. Neuron 40, 695–701.
Cobos, I., Broccoli, V., and Rubenstein, J. L. R. (2005). The vertebrateortholog of Aristaless is regulated by Dlx genes in the developing forebrain. J. Comp. Neurol. 483, 292–303.
Colasante, G., Collombat, P., Raimondi, V., Bonanomi, D., Ferrai, C., Maira, M., Yoshikawa, K., Mansouri, A., Valtorta, F., Rubenstein, J. L., and Broccoli, V. (2008). Arx is a direct target of Dlx2 and thereby contributes to the tangential migration of GABAergic interneurons. J Neurosci. 28, 10674–10686.
Colasante, G., Sessa, A., Crispi, S., Calogero, R., Mansouri, A., Collombat, P., and Broccoli, V. (2009). Arx acts as a regional key selector gene in the ventral telencephalon mainly through its transcriptional repression activity. Dev. Biol. 334, 59–71.
Collombat, P., Mansouri, A., Hecksher-Sorensen, J., Serup, P., Krull, J., Gradwohl, G., and Gruss, P. (2003). Opposing actions of Arx and Pax4 in endocrine pancreas development. Genes Dev. 17, 2591–2603.
Colombo, E., Collombat, P., Colasante, G., Bianchi, M., Long, J., Mansouri, A., Rubenstein, J. L., and Broccoli, V. (2007). Inactivation of Arx, the murine ortholog of the X-linked lissencephaly with ambiguous genitalia gene, leads to severe disorganization of the ventral telencephalon with impaired neuronal migration and differentiation. J. Neurosci. 27, 4786–4798.
Colombo, E., Galli, R., Cossu, G., Gécz, J., and Broccoli, V. (2004). Mouse orthologue of ARX, a gene mutated in several X-linked forms of mental retardation and epilepsy, is a marker of adult neural stem cells and forebrain GABAergic neurons. Dev. Dyn. 231, 631–639.
Corbin, J. G., Nery, S., and Fishell, G. (2001). Telencephalic cells take a tangent: non-radial migration in the mammalian forebrain. Nat. Neurosci. 4, 1177–1182.
Deuel, T. A., Liu, J. S., Corbo, J. C., Yoo, S. Y., Rorke-Adams, L. B., and Walsh, C. A. (2006). Genetic interactions between doublecortin and doublecortin-like kinase in neuronal migration and axon outgrowth. Neuron 49, 41–53.
Di Cristo, G. (2007). Development of cortical GABAergic circuits and its implications for neurodevelopmental disorders. Clin. Genet. 72, 1–8.
Faux, C., Rakić, S., Andrews, W., Yanagawa, Y., Obata, K., and Parnavelas, J. G. (2010). Differential gene expression in migrating cortical interneurons during mouse forebrain development. J. Comp. Neurol. 518, 1232–1248.
Forman, M. S., Squier, W., Dobyns, W. B., and Golden, J. A. (2005). Genotypically defined lissencephalies show distinct pathologies. J. Neuropathol. Exp. Neurol. 64, 847–857.
Friocourt, G., Kanatani, S., Tabata, H., Yozu, M., Takahashi, T., Antypa, M., Raguénès, O., Chelly, J., Férec, C., Nakajima, K., and Parnavelas, J. G. (2008). Cell-autonomous roles of ARX in cell proliferation and neuronal migration during corticogenesis. J. Neurosci. 28, 5794–5805.
Friocourt, G., Liu, J. S., Antypa, M., Rakic, S., Walsh, C. A., and Parnavelas, J. G. (2007). Both doublecortin and doublecortin-like kinase play a role in cortical interneuron migration. J. Neurosci. 27, 3875–3883.
Friocourt, G., and Parnavelas, J. G. (2010). Mutations in ARX result in several defects involving GABAergic neurons. Front. Cell. Neurosci. 4:4. doi: 10.3389/fncel.2010.00004
Friocourt, G., Poirier, K., Rakić, S., Parnavelas, J. G., and Chelly, J. (2006). The role of ARX in cortical development. Eur. J. Neurosci. 23, 869–876.
Fullenkamp, A. N., and El-Hodiri, H. M. (2008). The function of the Aristaless-related homeobox (Arx) gene product as a transcriptional repressor is diminished by mutations associated with X-linked mental retardation (XLMR). Biochem. Bio-phys. Res. Commun. 377, 73–78.
Fulp, C. T., Cho, G., Marsh, E. D., Nasrallah, I. M., Labosky, P. A., and Golden, J. A. (2008). Identification of Arx transcriptional targets in the developing basal forebrain. Hum. Mol. Genet. 17, 3740–3760.
Galaburda, A. M., LoTurco, J., Ramus, F., Fitch, R. H., and Rosen, G. D. (2006). From genes to behavior in developmental dyslexia. Nat. Neurosci. 9, 1213–1217.
Garcia-Dominguez, M., Poquet, C., Garel, S., and Charnay, P. (2003). Ebf gene function is required for coupling neuronal differentiation and cell cycle exit. Development 130, 6013–6025.
Gelman, D. M., Martini, F. J., Nóbrega-Pereira, S., Pierani, A., Kessaris, N., and Marín, O. (2009). The embryonic preoptic area is a novel source of cortical GABAergic interneurons. J. Neurosci. 29, 9380–9389.
Gilmore, E. C., Ohshima, T., Goffinet, A. M., Kulkarni, A. B., and Herrup, K. (1998). Cyclin-dependent kinase 5-deficient mice demonstrate novel developmental arrest in cerebral cortex. J. Neurosci. 18, 6370–6377.
Hammond, V., So, E., Gunnersen, J., Valcanis, H., Kalloniatis, M., and Tan, S. S. (2006). Layer positioning of late-born cortical interneurons is dependent on Reelin but not p35 signaling. J. Neurosci. 26, 1646–1655.
Hernandez-Miranda, L. R., Cariboni, A., Faux, C., Ruhrberg, C., Cho, J. H., Cloutier, J. F., Eickholt, B. J., Parnavelas, J. G., and Andrews, W. D. (2011). Robo1 regulates semaphorin signaling to guide the migration of cortical interneurons through the ventral forebrain. J. Neurosci. 31, 6174–6187.
Hernandez-Miranda, L. R., Parnavelas, J. G., and Chiara, F. (2010). Molecules and mechanisms involved in the generation and migration of cortical interneurons. ASN Neuro 2, e00031. doi: 10.1042/AN20090053
Howell, B. W., Hawkes, R., Soriano, P., and Cooper, J. A. (1997). Neuronal position in the developing brain is regulated by mouse disabled-1. Nature 389, 733–737.
Hurtado-Chong, A., Yusta-Boyo, M. J., Vergaño-Vera, E., Bulfone, A., de Pablo, F., and Vicario-Abejón, C. (2009). IGF-I promotes neuronal migration and positioning in the olfactory bulb and the exit of neuroblasts from the subventricular zone. Eur. J. Neurosci. 30, 742–755.
Isogai, E., Ohira, M., Ozaki, T., Oba, S., Nakamura, Y., and Nakagawara, A. (2011). Oncogenic LMO3 collaborates with HEN2 to enhance neuroblastoma cell growth through transactivation of Mash1. PLoS ONE 6, e19297. doi: 10.1371/journal.pone.0019297
Jakovcevski, I., Mayer, N., and Zecevic, N. (2011). Multiple origins of human neocortical interneurons are supported by distinct expression of transcription factors. Cereb. Cortex 21, 1771–1782.
Kalachikov, S., Evgrafov, O., Ross, B., Winawer, M., Barker-Cummings, C., Boneschi, F. M., Choi, C., Morozov, P., Das, K., Teplitskaya, E., Yu, A., Cayanis, E., Penchaszadeh, G., Kottmann, A. H., Pedley, T. A., Hauser, W. A., Ottman, R., and Gilliam, T. C. (2002). Mutations in LGI1 cause autosomal-dominant partial epilepsy with auditory features. Nat. Genet. 30, 335–341.
Kato, M., and Dobyns, W. B. (2005). X-linked lissencephaly with abnormal genitalia as a tangential migration disorder causing intractable epilepsy: proposal for a new term, “interneuronopathy”. J. Child Neurol. 20, 392–397.
Kerjan, G., Dolan, J., Haumaitre, C., Schneider-Maunoury, S., Fujisawa, H., Mitchell, K. J., and Chédotal A. (2005). The transmembrane semaphorin Sema6A controls cerebellar granule cell migration. Nat. Neurosci. 8, 1516–1524.
Kitamura, K., Itou, Y., Yanazawa, M., Ohsawa, M., Suzuki-Migishima, R., Umeki, Y., Hohjoh, H., Yanagawa, Y., Shinba, T., Itoh, M., Nakamura, K., and Goto, Y. (2009). Three human ARX mutations cause the lissencephaly-like and mental retardation with epilepsy-like pleiotropic phenotypes in mice. Hum. Mol. Genet. 18, 3708–3724.
Kitamura, K., Yanazawa, M., Sugiyama, N., Miura, H., Iizuka-Kogo, A., Kusaka, M., Omichi, K., Suzuki, R., Kato-Fukui, Y., Kamiirisa, K., Matsuo, M., Kamijo, S. I., Kasahara, M., Yoshioka, H., Ogata, T., Fukuda, T., Kondo, I., Kato, M., Dobyns, W. B., Yokoyama, M., and Morohashi, K. I. (2002). Mutation of ARX causes abnormal development of forebrain and testes in mice and X-linked lissencephaly with abnormal genitalia in humans. Nat. Genet. 32, 359–369.
Koizumi, H., Tanaka, T., and Gleeson, J. G. (2006). Doublecortin-like kinase functions with doublecortin to mediate fiber tract decussation and neuronal migration. Neuron 49, 55–66.
Kriegstein, A. R., and Noctor, S. C. (2004). Patterns of neuronal migration in the embryonic cortex. Trends Neurosci. 27, 392–399.
Kunapuli, P., Lo, K., Hawthorn, L., and Cowell, J. K. (2010). Reexpression of LGI1 in glioma cells results in dysregulation of genes implicated in the canonical axon guidance pathway. Genomics 95, 93–100.
Lavdas, A. A., Grigoriou, M., Pachnis, V., and Parnavelas, J. G. (1999). The medial ganglionic eminence gives rise to a population of early neurons in the developing cerebral cortex. J. Neurosci. 19, 7881–7888.
Letinic, K., Zoncu, R., and Rakic, P. (2002). Origin of GABAergic neurons in the human neocortex. Nature 417, 645–649.
Levitt, P., Eagleson, K. L., and Powell, E. M. (2004). Regulation of neocortical interneuron development and the implications for neurodevelopmental disorders. Trends Neurosci. 27, 400–406.
Li, G., Adesnik, H., Li, J., Long, J., Nicoll, R. A., Rubenstein, J. L., and Pleasure, S. J. (2008). Regional distribution of cortical interneurons and development of inhibitory tone are regulated by cxcl12/cxcr4 signaling. J. Neurosci. 28, 1085–1098.
Liapi, A., Pritchett, J., Jones, O., Fujii, N., Parnavelas, J. G., and Nadarajah, B. (2008). Stromal-derived factor 1 signalling regulates radial and tangential migration in the developing cerebral cortex. Dev. Neurosci. 30, 117–131.
Liodis, P., Denaxa, M., Grigoriou, M., Akufo-Addo, C., Yanagawa, Y., and Pachnis, V. (2007). Lhx6 activity is required for the normal migration and specification of cortical interneuron subtypes. J. Neurosci. 27, 3078–3089.
Long, J. E., Cobos, I., Potter, G. B., and Rubenstein, J. L. (2009a). Dlx1&2 and Mash1 transcription factors control MGE and CGE patterning and differentiation through parallel and overlapping pathways. Cereb. Cortex 19(Suppl. 1), i96–i106.
Long, J. E., Swan, C., Liang, W. S., Cobos, I., Potter, G. B., and Ruben-stein, J. L. (2009b). Dlx1&2 and Mash1 transcription factors control striatal patterning and differentiation through parallel and overlapping pathways. J. Comp. Neurol. 512, 556–572.
Long, J. S., Yokoyama, K., Tigyi, G., Pyne, N. J., and Pyne, S. (2006). Lipid phosphate phosphatase-1 regulates lysophosphatidic acid- and platelet-derived-growth-factor-induced cell migration. Biochem. J. 394, 495–500.
Lopez-Bendito, G., Sánchez-Alcañiz, J. A., Pla, R., Borrell, V., Pico, E., Valdeolmillos, M., and Marin, O. (2008). Chemokine signaling controls intracortical migration and final distribution of gabaergic interneurons. J. Neurosci. 28, 1613–1624.
Lu, Y., Li, T., Qureshi, H. Y., Han, D., and Paudel, H. K. (2011). Early growth response 1 (Egr-1) regulates phosphorylation of microtubule-associated protein tau in mammalian brain. J. Biol. Chem. 286, 20569–20581.
Lysko, D. E., Putt, M., and Golden, J. A. (2011). SDF1 regulates leading process branching and speed of migrating interneurons. J. Neurosci. 31, 1739–1745.
Maira, M., Long, J. E., Lee, A. Y., Rubenstein, J. L., and Stifani, S. (2010). Role for TGF-beta superfamily signaling in telencephalic GABAergic neuron development. J. Neurodev. Disord. 2, 48–60.
Marcorelles, P., Laquerriere, A., Adde-Michel, C., Marret, S., Saugier-Veber, P., Beldjord, C., and Friocourt, G. (2010). Evidence for tangential migration disturbances in human lissencephaly resulting from a defect in LIS1, DCX and ARX genes. Acta Neuropathol. 120, 503–515.
Marín, O., Plump, A. S., Flames, N., Sánchez-Camacho, C., Tessier-Lavigne, M., and Rubenstein, J. L. (2003). Directional guidance of interneuron migration to the cerebral cortex relies on subcortical Slit1/2-independent repulsion and cortical attraction. Development 130, 1889–1901.
Marín, O., and Rubenstein, J. L. (2003). Cell migration in the forebrain. Annu. Rev. Neurosci. 26, 441–483.
Marín, O., Yaron, A., Bagri, A., Tessier-Lavigne, M., and Rubenstein, J. L. (2001). Sorting of striatal and cortical interneurons regulated by semaphorin–neuropilin interactions. Science 293, 872–875.
Marsh, E. D., Minarcik, J., Campbell, K., Brooks-Kayal, A. R., and Golden, J. A. (2008). FACS-array gene expression analysis during early development of mouse telencephalic interneurons. Dev. Neurobiol. 68, 434–445.
McKenzie, O., Ponte, I., Mangelsdorf, M., Finnis, M., Colasante, G., Shoubridge, C., Stifani, S., Gécz, J., and Broccoli V. (2007). Aristaless-related homeobox gene, the gene responsible for West syndrome and related disorders, is a Groucho/transducin-like enhancer of split dependent transcriptional repressor. Neuroscience 146, 236–247.
Métin, C., Baudoin, J. P., Rakić, S., and Parnavelas, J. G. (2006). Cell and molecular mechanisms involved in the migration of cortical interneurons. Eur. J. Neurosci. 23, 894–900.
Miyoshi, G., and Fishell, G. (2011). GABAergic interneuron lineages selectively sort into specific cortical layers during early postnatal development. Cereb. Cortex 21, 845–852.
Nadarajah, B., and Parnavelas, J. G. (2002). Modes of neuronal migration in the developing cerebral cortex. Nat. Rev. Neurosci. 3, 423–432.
Nakazawa, K., Zsiros V., Jiang, Z., Nakao, K., Kolata, S., Zhang, S., and Belforte, J. E. (2011). GABAergic interneuron origin of schizophrenia pathophysiology. Neuropharmacology. doi: 10.1016/j.neuropharm. 2011.01.022 [Epub ahead of print].
Nery, S., Fishell, G., and Corbin, J. G. (2002). The caudal ganglionic eminence is a source of distinct cortical and subcortical cell populations. Nat. Neurosci. 5, 1279–1287.
Noctor, S. C., Flint, A. C., Weissman, T. A., Dammerman, R. S., and Kriegstein, A. R. (2001). Neurons derived from radial glial cells establish radial units in neocortex. Nature 409, 714–720.
Noctor, S. C., Martínez-Cerdeno, V., Ivic, L., and Kriegstein, A. R. (2004). Cortical neurons arise in symmetric and asymmetric division zones and migrate through specific phases. Nat. Neurosci. 7, 136–144.
Oishi, K., Watatani, K., Itoh, Y., Okano, H., Guillemot, F., Nakajima, K., and Gotoh, Y. (2009). Selective induction of neocortical GABAergic neurons by the PDK1-Akt pathway through activation of Mash1. Proc. Natl. Acad. Sci. USA 106, 13064–13069.
Okazaki, S., Ohsawa, M., Kuki, I., Kawawaki, H., Koriyama, T., Ri, S., Ichiba, H., Hai, E., Inoue, T., Nakamura, H., Goto, Y., Tomiwa, K., Yamano, T., Kitamura, K., and Itoh, M. (2008). Aristaless-related homeobox gene disruption leads to abnormal distribution of GABAergic interneurons in human neocortex: evidence based on a case of X-linked lissencephaly with abnormal genitalia (XLAG). Acta Neuropathol. 116, 453–462.
Orlovsky, K., Kalinkovich, A., Rozovskaia, T., Shezen, E., Itkin, T., Alder, H., Ozer, H. G., Carramusa, L., Avigdor, A., Volinia, S., Buchberg, A., Mazo, A., Kollet, O., Largman, C., Croce, C. M., Nakamura, T., Lapidot, T., and Canaani, E. (2011). Down-regulation of homeobox genes MEIS1 and HOXA in MLL-rearranged acute leukemia impairs engraftment and reduces proliferation. Proc. Natl. Acad. Sci. USA 108, 7956–7961.
Petanjek, Z., Berger, B., and Esclapez, M. (2009). Origins of cortical GABAergic neurons in the cynomolgus monkey. Cereb. Cortex 19, 249–262.
Poirier, K., Van Esch, H., Friocourt, G., Saillour, Y., Bahi, N., Backer, S., Souil, E., Castelnau-Ptakhine, L., Beldjord, C., Francis, F., Bienvenu, T., and Chelly, J. (2004). Neuroanatomical distribution of ARX in brain and its localisation in GABAergic neurons. Mol. Brain Res. 122, 35–46.
Price, M. G., Yoo, J. W., Burgess, D. L., Deng, F., Hrachovy, R. A., Frost, J. D. Jr., and Noebels, J. L. (2009). A triplet repeat expansion genetic mouse model of infantile spasms syndrome, Arx(GCG)10 + 7, with interneuronopathy, spasms in infancy, persistent seizures, and adult cognitive and behavioral impairment. J. Neurosci. 29, 8752–8763.
Quillé, M. L., Carat, S., Quéméner-Redon, S., Hirchaud, E., Baron, D., Benech, C., Guihot, J., Placet, M., Mignen, O., Férec, C., Houlgatte, R., and Friocourt, G. (2011).High-throughput analysis of promoter occupancy reveals new targets for Arx, a gene mutated in mental retardation and interneuronopathies. PLoS ONE 6, e25181. doi: 10.1371/ journal.pone.0025181
Rakić, S., Yanagawa, Y., Obata, K., Faux, C., Parnavelas, J. G., and Nikolić, M. (2009). Cortical interneurons require p35/Cdk5 for their migration and laminar organization. Cereb. Cortex 19, 1857–1869.
Renaud, J., Kerjan, G., Sumita, I., Zagar, Y., Georget, V., Kim, D., Fouquet, C., Suda, K., Sanbo, M., Suto, F., Ackerman, S. L., Mitchell, K. J., Fujisawa, H., and Chédotal, A. (2008). Plexin-A2 and its ligand, Sema6A, control nucleus–centrosome coupling in migrating granule cells. Nat. Neurosci. 11, 440–449.
Sajan, S. A., Rubenstein, J. L., Warchol, M. E., and Lovett, M. (2011). Identification of direct downstream targets of Dlx5 during early inner ear development. Hum. Mol. Genet. 20, 1262–1273.
Sakthianandeswaren, A., Christie, M., D’Andreti, C., Tsui, C., Jorissen, R. N., Li, S., Fleming, N. I., Gibbs, P., Lipton, L., Malaterre, J., Ramsay, R. G., Phesse, T. J., Ernst, M., Jeffery, R. E., Poulsom, R., Leedham, S. J., Segditsas, S., Tomlinson, I. P., Bernhard, O. K., Simpson, R. J., Walker, F., Faux, M. C., Church, N., Catimel, B., Flanagan, D. J., Vincan, E., and Sieber, O. M. (2011). PHLDA1 expression marks the putative epithelial stem cells and contributes to intestinal tumorigenesis. Cancer Res. 71, 3709–3719.
Sánchez-Alcañiz, J. A., Haege, S., Mueller, W., Pla, R., Mackay, F., Schulz, S., López-Bendito, G., Stumm, R., and Marín, O. (2011). Cxcr7 controls neuronal migration by regulating chemokine responsiveness. Neuron 69, 77–90.
Seufert, D. W., Prescott, N. L., and El-Hodiri, H. M. (2005). Xenopus aristaless-related homeobox (xARX) gene product functions as both a transcriptional activator and repressor in forebrain development. Dev. Dyn. 232, 313–324.
Shieh, J. C., Schaar, B. T., Srinivasan, K., Brodsky, F. M., and McConnell, S. K. (2011). Endocytosis regulates cell soma translocation and the distribution of adhesion proteins in migrating neurons. PLoS ONE 6, e17802. doi: 10.1371/journal.pone. 0017802
Shmueli, A., Gdalyahu, A., Sapoznik, S., Sapir, T., Tsukada, M., and Reiner, O. (2006). Site-specific dephosphorylation of doublecortin (DCX) by protein phosphatase 1 (PP1). Mol. Cell. Neurosci. 32, 15–26.
Shoubridge, C., Fullston, T., and Gécz, J. (2010). ARX spectrum disorders: making inroads into the molecular pathology. Hum. Mutat. 31, 889–900.
Simó, S., Pujadas, L., Segura, M. F., La Torre, A., Del Río, J. A., Ureña, J. M., Comella, J. X., and Soriano, E. (2007). Reelin induces the detachment of postnatal subventricular zone cells and the expression of the Egr-1 through Erk1/2 activation. Cereb. Cortex 17, 294–303.
Skalski, M., Yi, Q., Kean, M. J., Myers, D. W., Williams, K. C., Burtnik, A., and Coppolino, M. G. (2010). Lamellipodium extension and membrane ruffling require different SNARE-mediated trafficking pathways. BMC Cell Biol. 11, 62. doi: 10.1186/1471-2121-11-62
Stumm, R. K., Zhou, C., Ara, T., Lazarini, F., Dubois-Dalcq, M., Nagasawa, T., Höllt, V., and Schulz, S. (2003). CXCR4 regulates interneuron migration in the developing neocortex. J. Neurosci. 23, 5123–5130.
Tanaka, D. H., Maekawa, K., Yanagawa, Y., Obata, K., and Murakami, F. (2006). Multidirectional and multizonal tangential migration of GABAergic interneurons in the developing cerebral cortex. Development 133, 2167–2176.
Tanaka, D. H., Yanagida, M., Zhu, Y., Mikami, S., Nagasawa, T., Miyazaki, J., Yanagawa, Y., Obata, K., and Murakami, F. (2009). Random walk behavior of migrating cortical interneurons in the marginal zone: time-lapse analysis in flat-mount cortex. J. Neurosci. 29, 1300–1311.
Tiveron, M. C., Rossel, M., Moepps, B., Zhang, Y. L., Seidenfaden, R., Favor, J., Konig, N., and Cremer, H. (2006). Molecular interaction between projection neuron precursors and invading interneurons via stromal-derived factor 1 (cxcl12)/cxcr4 signaling in the cortical subventricular zone/intermediate zone. J. Neurosci. 26, 13273–13278.
Toresson, H., Parmar, M., and Campbell, K. (2000). Expression of Meis and Pbx genes and their protein products in the developing telencephalon: implications for regional differentiation. Mech. Dev. 94, 183–187.
Wang, Y., Li, G., Stanco, A., Long, J. E., Crawford, D., Potter, G. B., Pleasure, S. J., Behrens, T., and Rubenstein, J. L. (2011). CXCR4 and CXCR7 have distinct functions in regulating interneuron migration. Neuron 69, 61–76.
Ware, M. L., Fox, J. W., González, J. L., Davis, N. M., Lambert de Rouvroit, C., Russo, C. J., Chua, S. C. Jr., Goffinet, A. M., and Walsh, C. A. (1997). Aberrant splicing of a mouse disabled homolog, mdab1, in the scrambler mouse. Neuron 19, 239–249.
West, K. L., Castellini, M. A., Duncan, M. K., and Bustin, M. (2004). Chromosomal proteins HMGN3a and HMGN3b regulate the expression of glycine transporter 1. Mol. Cell. Biol. 24, 3747–3756.
Woods, K. A., Camacho-Hubner, C., Savage, M. O., and Clark, A. J. L. (1996). Intrauterine growth retardation and postnatal growth failure associated with deletion of the insulin-like growth factor I gene. New Eng. J. Med. 335, 1363–1367.
Yaman, E., Gasper, R., Koerner, C., Wittinghofer, A., and Tazebay, U. H. (2009). RasGEF1A and RasGEF1B are guanine nucleotide exchange factors that discriminate between Rap GTP-binding proteins and mediate Rap2-specific nucleotide exchange. FEBS J. 276, 4607–4616.
Yozu, M., Tabata, H., and Nakajima, K. (2005). The caudal migratory stream: a novel migratory stream of interneurons derived from the caudal ganglionic eminence in the developing mouse forebrain. J. Neurosci. 25, 7268–7277.
Zecevic, N., Hu, F., and Jakovcevski, I. (2011). Interneurons in the developing human neocortex. Dev. Neurobiol. 71, 18–33.
Keywords: ARX, GABA, epilepsy, interneurons, neuronal migration, basal ganglia
Citation: Friocourt G and Parnavelas JG (2011) Identification of Arx targets unveils new candidates for controlling cortical interneuron migration and differentiation. Front. Cell. Neurosci. 5:28. doi: 10.3389/fncel.2011.00028
Received: 14 October 2011; Accepted: 08 December 2011;
Published online: 27 December 2011.
Edited by:
Yehezkel Ben-Ari, Institut National de la Santé et de la Recherche Médicale, FranceCopyright: © 2011 Friocourt and Parnavelas. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Gaëlle Friocourt, Laboratory of Molecular Genetics and Histocompatibility, Inserm U613, 46 rue Félix Le Dantec, CS51819, 29218 Brest Cedex 2, France. e-mail:Z2FlbGxlLmZyaW9jb3VydEB1bml2LWJyZXN0LmZy

