- 1Laboratory of Aquaculture and Breeding, Freshwater Fisheries Research Institute of Fujian, Fuzhou, China
- 2Faculty of Animal Science and Technology, Yunnan Agricultural University, Kunming, China
- 3Institute of Environment and Ecology, School of the Environmental and Safety Engineering, Jiangsu University, Zhenjiang, China
- 4Jiangsu Collaborative Innovation Center of Technology and Material of Water Treatment, Suzhou University of Science and Technology, Suzhou, China
Differences in growth rates are commonly observed within cultured populations of the Opsariichthys bidens. In this study, we investigated the role of the intestinal microbiota in the growth and development of O. bidens by comparing two populations from a full-sib family: one with rapid growth (the OW group) and another with slow growth (the UW group). The intestinal contents were analyzed using 16S rRNA sequencing, and histological staining was used to observe intestinal morphology. Digestive enzyme activities and gene expression related to growth and metabolism were also examined. The OW group was dominated by Burkholderia-caballeronia-Paraburkholderia species, and Cetobacterium dominated the UW group. The development of villi and digestive enzyme activities in the OW group were greater than those in the UW group. Transcriptome analysis indicated elevated expression of anabolic genes in the OW group. The correlations coefficients between intestinal microbiota and indicators such as the growth phenotype, intestinal morphological structure, digestive enzyme activity, and gene transcription in O. bidens were all positive. This study investigated the correlation between the intestinal microbiota of O. bidens and various growth indicators, offering guidance and practical applications for the selection and breeding of fast-growing aquaculture varieties.
Introduction
The freshwater fish industry has become an economic pillar of China’s aquaculture. However, as in most forms of aquaculture, the industry has experienced problems such as disease outbreaks and germplasm degradation (Li et al., 2012). Inconsistent growth rates of aquaculture species can lead to problems such as decreased production efficiency, extension of the aquaculture cycle, and increased costs, all of which affect production yield (Liang et al., 2010). Opsariichthys bidens belongs to the family Leuciscinae, genus Opsariichthys (Zhong, 1985). The minnow is also known as flower slits, broad-stomates, and peach blossoms. O. bidens is distributed in China’s plateau area and several other provinces, as well as in Taiwan. The meat of O. bidens is prized by consumers. Its crude protein content is 18.78%, and its crude fat content is 1.79%. In addition, it is rich in seven essential amino acids and polyunsaturated fatty acids. In recent years, O. bidens has become the primary economically important fish species in certain mountainous and hilly areas of southern China (Yao, 1988). O. bidens has formed the basis of aquaculture industries in regions such as Fujian, Zhejiang, Guangdong, Jiangxi, Hubei, and Hunan. The annual breeding of seedlings is about 500 million to over one billion. The price at the pond remains stable at 30–50 yuan per kilogram throughout the year, and the industrial scale has reached about one billion yuan. O. bidens has become the most rapidly developing species representative among native stream fish. However, O. bidens has experienced significant variation in growth and development. For example, in places such as Anhua, Hunan Province, Sandaohe, Hubei Province, and Qingyijiang, Anhui Province, there are differences in the biology and growth characteristics between wild and pond farmed O. bidens (Su et al., 1993; Zhan, 2011; Liu et al., 2015). Previous studies have shown that the main factors affecting the growth of individual fish can be divided into exogenous and endogenous factors. Among these, exogenous factors include the water temperature, pH, light, and feed, while endogenous factors include the intestinal microbial structure, genetic inheritance, digestive enzyme activity, and autoimmunity. These factors may be correlated (Li, 2014; Kim et al., 2023; Mao et al., 2023). Over the past several years, the phenomenon of growth difference has become increasingly severe, restricting the development of the O. bidens aquaculture industry. Therefore, it is essential to determine the reasons for this excessive difference in growth and explore methods for obtaining balanced and rapid growth of cultured O. bidens.
The microbiota of the fish gut is composed of archaea, bacteria, protists, fungi, and viruses (Suo et al., 2017; Zang et al., 2019). Bacteria are the main group, and the dominant phyla are Proteobacteria, Fusobacteria, Firmicutes, Bacteroidetes, Actinobacteria, Verrucomicrobia, and others (Nayak, 2010; Li et al., 2018). With the establishment and development of metagenomics sequencing technology, the functions of the intestinal microbiota and the mutualistic-symbiotic relationships between intestinal microbiota and their hosts have gradually been explored. Intestinal microbiota are important factors affecting the health and nutritional status of the host (Wang et al., 2018), and the complex microbial community in the digestive tract plays an important role in the regulation of the immune and digestive systems of fish (Tran et al., 2018; Valdes et al., 2018). These functions include aiding digestion, promoting mucosal phylogeny, and resisting the invasion of foreign pathogens (Ringø et al., 2007). The composition and structure of the intestinal microbial community vary among fish species, and the intestinal microbial community of fishes has a unique core compared with the microbiota of the environment (Kuang et al., 2020). The relationship between the fish intestinal microbiota and the host is becoming increasingly well characterized; for example, the intestinal microbiota produce a variety of digestive enzymes such as amylase, lipase, and trypsin (Francesco et al., 2021). An imbalance in intestinal flora may lead to acute gastroenteritis, cardiovascular disease, metabolic abnormalities, and other diseases (Blumberg and Powrie, 2012).
There are several individual differences important in the breeding process of O. bidens. The miniaturization of individuals, the slow growth rate, and the large individual differences within the same batch of O. bidens have hindered the development of the O. bidens breeding industry in China. Thus, it is urgent to characterize the mechanisms underlying these differences. The structure and function of the intestinal flora are closely related to fish growth and metabolism and thus are potential factors affecting differences in the growth characteristics of O. bidens. Full-sib family selection is a form of selection in which analysis of phenotypes and tracing of pedigrees of each individual are combined to select quantitative traits, obtain new varieties or new lines, and improve breeding efficiency through multi-generation directional selection. The advantage of this technique is that it can eliminate the interference to the genetic background caused by different parents by comparing genetic information among full-sib families (Gjerde and Korsvoll, 1999). Our group’s previous research and an industry follow-up survey found significant differences in the growth rate of offspring in the large-scale production of O. bidens. At the same time, the intestinal microbial community structure varied between individuals of different sizes within full-sib families. To further explore the regulatory relationship between growth differences and intestinal flora, this study employed a full-sib family of O. bidens as experimental subjects, aiming to minimize the influence of genetic factors. In this study, high-throughput sequencing of the 16S rRNA gene and transcriptomics technology were employed to analyze the variation in microflora, metabolic pathways, and differentially expressed genes in the intestinal tract of O. bidens, aiming to explain the correlations between intestinal microflora, host co-metabolism, and growth functions. Our primary goal was to verify the functional role of intestinal microbiota in regulating individual growth and development. The ultimate goal of this study is to investigate the microecological regulation pathways for enhancing the yield of O. bidens and promoting the sustainable development of the aquaculture industry.
Materials and methods
Culture of O. bidens
The experimental subjects were a full-sib family of O. bidens preserved in the cooperative base of the Freshwater Fisheries Research Institute of Fujian. The research group used wild adult horse mouth fish parents for artificial oxytocin and one-to-one pair breeding. Wild mature parents of O. bidens were artificially induced to ovulate using oxytocin (DOM 3 mg, LRH-A2 5 μg, HCG 800 IU), and paired insemination was performed according to a previous study (Chen et al., 2023). Fertilized eggs from the same parent were cultured under the same environmental conditions. After the larvae hatched, the entire process was isolated in a factory breeding tank. The culture conditions were as follows: the temperature of the aquaculture water body was 24 ± 0.5°C; feed was given twice per day, and about 50% of the water was replaced twice per week.
After continuous cultivation for three months, a total of 100 healthy fish (i.e., complete coverlet scales, a smooth surface, apparent vitality, and no parasites) were randomly selected from the full-sib family. The fish were ranked by body weight, and those in the top 30 were classified as the overweight (OW) group, while those in the bottom 30 were classified as the underweight (UW) group for detection and analysis.
Morphometry
The biological indexes in the OW and UW groups were determined. According to the method of Ichthyology Experimental Guidance (Meng et al., 1995), the whole body length, body length, head length, trunk length, body height, tail length, tail stalk height, and tail fin length of test fish were measured with Vernier calipers (accurate to 0.01 mm), and weight was measured with an electronic scale (accurate to 0.01 g).
Preparation of intestinal tissue sections
After anesthesia with MS222, the intestinal tissues of individuals from the OW and UW groups were dissected; the intestinal attachments were removed, and the midgut was excised and stored in 4% paraformaldehyde for paraffin sectioning. The test samples were transferred to 75% anhydrous ethanol for gradient dehydration, and xylene was applied twice for transparency, followed by three rounds of wax penetration. This was followed by embedding, wax block dressing, slicing (thickness = 5 μm), unfolding, and baking. The sections were stained with hematoxylin-eosin (HE) and sealed with neutral gum. An optical microscope (Olympus BX53) was used for microscopic observation, and the image acquisition software CCD (Olympus DP80) was used for photography. The villus height, villus width, and muscle thickness were measured using the NDP.view 2 software micrometer.
Determination of intestinal enzyme activity
The midgut tissue of the test fish was excised, and the gut contents were cleaned with normal saline and pre-cooled to 4°C. The fat, mesentery, and other intestinal impurities were removed, and excess water was absorbed with filter paper. The intestinal tissue was accurately weighed, and a corresponding volume of normal saline was added at a ratio of 1:9. Mechanical homogenization was carried out in a low-temperature water bath. The homogenate was centrifuged at 2500 rpm for 10 min, and the supernatant was taken and stored at 4°C. The activities of digestive enzymes, including total protein (TP), lipase (LPS), trypsin (TRY), and amylase (AMS), were measured using a kit produced by the Nanjing Jiancheng Bioengineering Institute.
16S rRNA sequencing analysis
The v3–v4 region of a fragment of the 16S rRNA gene in the gut tract of O. bidens was amplified using the primers F: ACTCCTACGGGAGGCAGCAG and R: GGACTACHVGGGTATCTAAT. The total volume of the amplification system was 25 μL, with 12.5 μL of 2× Mix (TaKaRa), 9.5 μL of dd H2O, 0.5 μL of 5′ and 3′ primers, and 2 μL of cDNA template. The PCR amplification procedure comprised predenaturation at 95°C for 30 s, denaturation at 95°C for 10 s, annealing at 58.6°C for 30 s, extension at 72°C for 2 min for 35 cycles, and a final elongation at 72°C for 10 min. The amplified products were extracted using a gel extraction kit, then sequenced and analyzed on an Illumina Nextseq 2000 (Illumina, USA) sequencing platform according to the specifications of Guangzhou Gidio Biotechnology Co., Ltd. The sequencing read length was 300 bp; the criterion was 97% ASVs similarity, and the rarefaction depth was 30, 171.
Total RNA isolation and purification, and cDNA synthesis
Total RNA was extracted from fish muscle tissue using a TRIzol kit (AG, USA). The values of OD260 nm and OD280 nm were determined using a Cary50 ultraviolet spectrophotometer (Varian, USA), and the extraction purity and concentration of RNA were preliminarily determined according to standard curves. The quality of RNA was subsequently detected by 1% agarose gel electrophoresis. cDNA was synthesized by reverse transcription of the extracted RNA using a PrimeScript RT reagent kit (TaKaRa, Japan) and diluted 10 times with ddH2O for subsequent PCR validation and qRT-PCR tests.
Transcriptome sequencing analysis
The total RNA of the sample was extracted and sent to Beijing NOhe Technology Co., Ltd., for cDNA synthesis, cDNA end repair, splicing, and PCR library enrichment and purification. Sequencing, data splicing assembly, and analysis were performed after quality control. The genes were mapped to each term of the GO database (http://www.geneontology.org/), and the number of genes in each term was counted. GO entries that were significantly enriched in differentially expressed genes compared with the whole genome background were identified using hypergeometric tests. All differentially expressed genes were mapped to pathways in the KEGG database (http://www.kegg.jp/), and the number of genes in each pathway was counted.
Transcriptome real-time fluorescence quantitative PCR verification
A set of expressed genes was randomly selected from the transcriptome database, and forward and reverse primers for real-time quantitative fluorescent PCR (qRT-PCR) were designed according to the gene sequences (Supplementary Table 1). The qualified primers were judged by detecting whether the amplification efficiency was within the range of 95–105%. β-actin was used as an internal reference gene to verify the expression levels of genes between different groups. qRT-PCR amplification was carried out using 10× gradient diluted cDNA as a template to construct standard curves. The total volume of the reaction system was 20 μL and contained 10 μL of 2× SYBR green master mix (TaKaRa), 6.8 μL of ddH2O, 0.4 μL of Rox Dye, 0.4 μL each of the 5′ and 3′ primers, and 2 μL of cDNA template. The amplification procedure comprised predenaturation at 95°C for 30 s, denaturation at 95°C for 5 s, and annealing and extension at 58.6°C for 30 s, for a total of 40 cycles. The qRT-PCR amplification was performed on a Step One Plus fluorescence quantitative PCR instrument.
Statistical analysis
The stats package for R (version 3.3.1) and the scipy package for Python (v1.0.0) were used in the statistical analysis. The Tukey–Kramer post-hoc test was employed for multiple comparisons. The criteria |log2 FC| ≥ 1 and P adjust < 0.05 were used to screen for differentially expressed genes. The “ggcor” package for R Studio software version 2022.07.0-548 (2009–2022 RStudio, PBC) was used for Mantel tests to analyze the correlations between morphological data and relevant indicators. A structural equation model (SEM) was constructed using the software IBM SPSS Amos (version 24). Each variable underwent Z-score standardization prior to modelling. The goodness of fit of the model was determined by the χ2 test (χ2/df ratio around 0 to –2 with P > 0.05), the root mean square error of approximation (RMSEA, 0.05 < RMSEA ≤ 0.08 with P > 0.05 defined as an adequate fit), and the root mean square residual (RMR, RMSEA < 0.01 with P > 0.05 as a good fit). SPSS23.0 (IBM, USA) and Excel were used for statistical analysis, and one-way ANOVA was used to compare the differences among groups. P < 0.05 indicated a significant difference, and P < 0.01 indicated a particularly significant difference. GraphPad Prism 8.0 software was used to map and present the results.
Results
Analysis of differences in the growth index
The growth indexes of body weight, total length, and other measures listed above were significantly higher in the OW group than in the UW group (P < 0.01). The mean body weight of fish in the OW group was more than two times higher than in the UW group; the total length of the OW group was more than 30% higher than that of the UW group. The average growth rate of the OW group was 158.55 mg/d, which was 122.28% greater than that of the UW group (Figure 1).
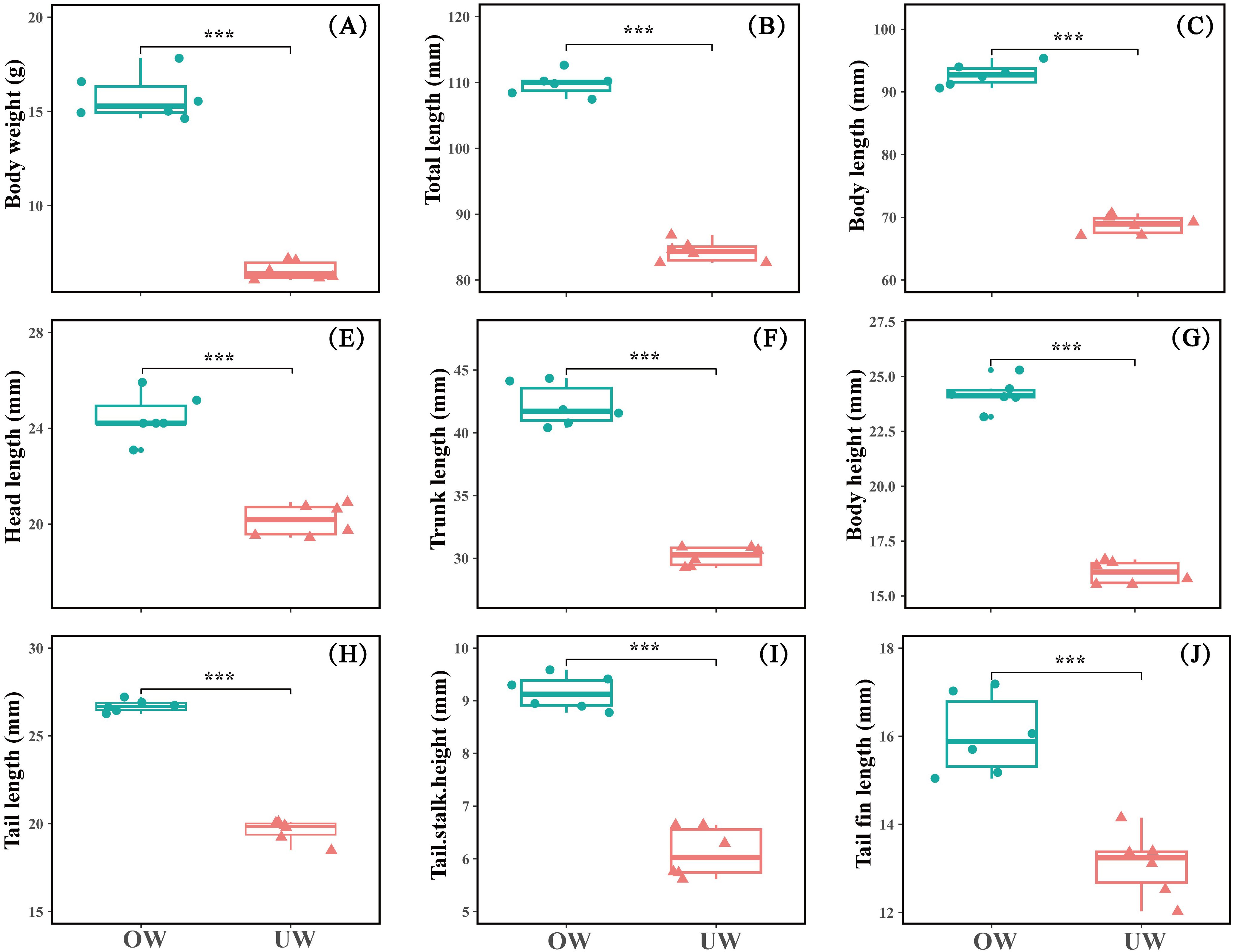
Figure 1. (A-J) Basic growth indicators between the overweight (OW) and underweight (UW) groups of Opsariichthys bidens. Data are presented as mean ± SD, n=6. *** indicates extremely significant difference at the P < 0.001 level. And that with over 99.9% certainty that the result is not accidental.
Analysis of intestinal morphology and structure
The intestinal tissue morphology and structure of fish in the OW and UW groups were significantly different, and the total intestinal villi size of fish in the OW group was significantly larger than that in the UW group. Further microscopic measurements showed that the intestinal villus height and intestinal muscle thickness in the OW group were significantly greater than those in the UW group (P < 0.05); the villus height was about 1.3 times that of the UW group, and the muscle thickness was about 1.4 times that of the UW group. In addition, the width of the intestinal villi in the OW group was significantly greater than in the UW group (P<0.01), approximately two times that of the UW group (Figures 2, 3).
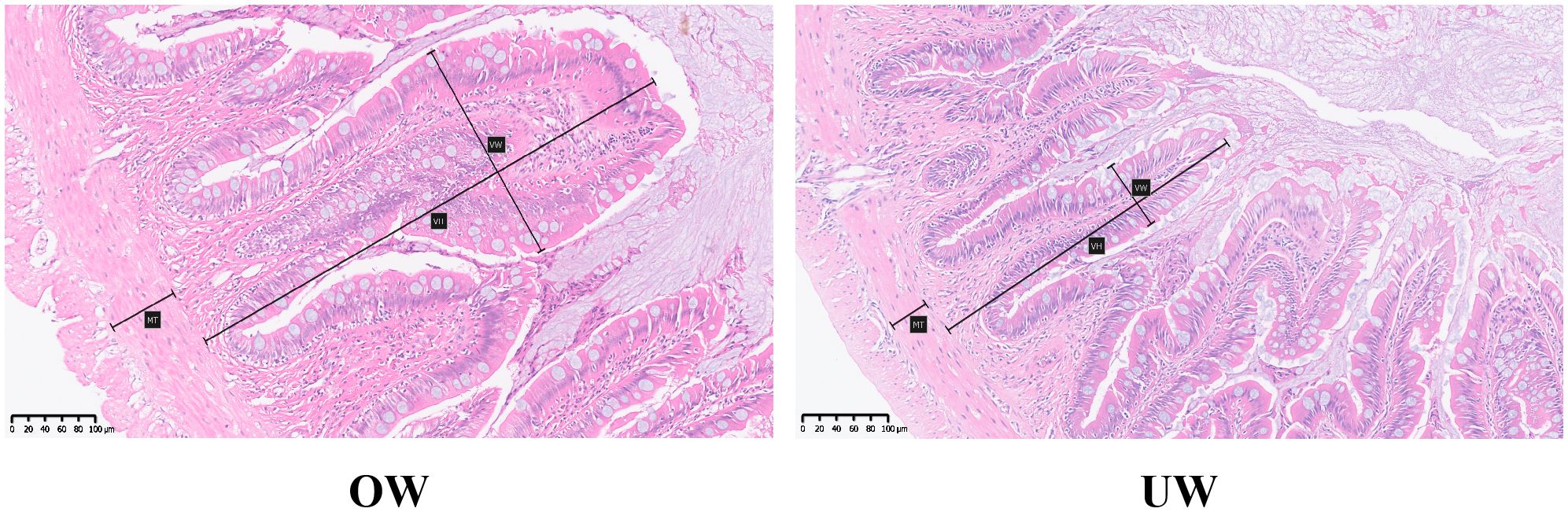
Figure 2. Hematoxylin-eosin stained sections of the intestinal tract between the overweight (OW) and underweight (UW) groups of Opsariichthys bidens. MT represents muscular thickness; VH represents villi height, and VW represents villi width. The scale is 20 μm, and the magnification is 200×.
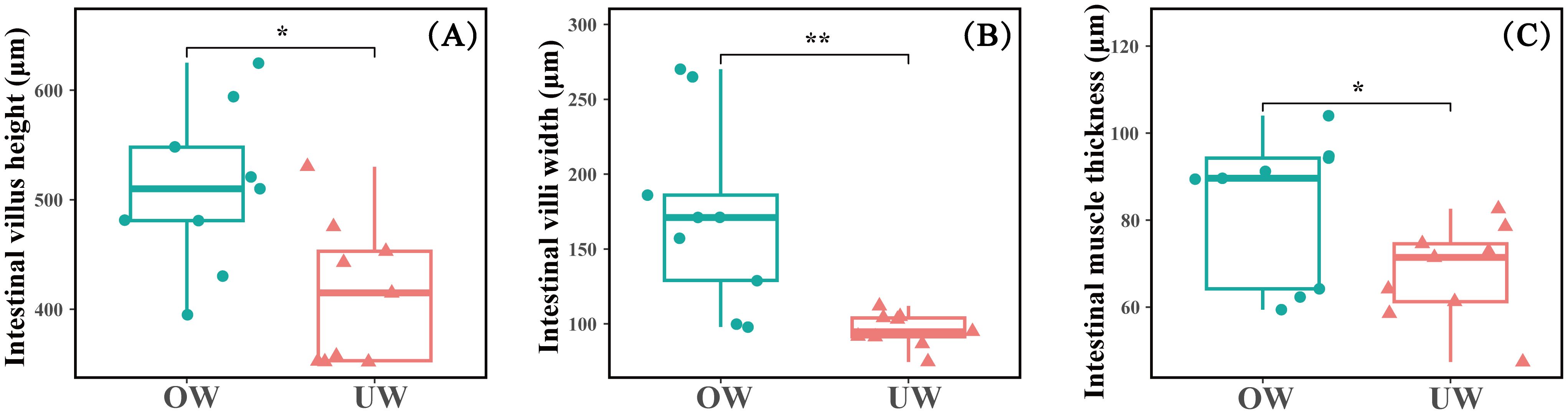
Figure 3. (A-C) The differences in intestinal tissue morphology and structure between overweight (OW) and underweight (UW) groups. Data are presented as mean ± SD, n = 9. * indicates a significant difference at the P < 0.05 level; ** indicates a highly significant difference at the P < 0.01 level. And that with over 99.9% certainty that the result is not accidental.
Analysis of intestinal digestive enzyme activity
The activities of intestinal digestive enzymes were detected, and the activities of lipase and amylase in the OW group were significantly higher than in the UW group (P < 0.05). The lipase activity of the OW was about 1.9 times that of the UW, and the amylase activity was about 2.5 times that of the UW. The total protein content and trypsin activity in the OW group were significantly higher than those in the UW (P < 0.01); the trypsin activity of the OW group was about 17 times that of the UW group (Figure 4).
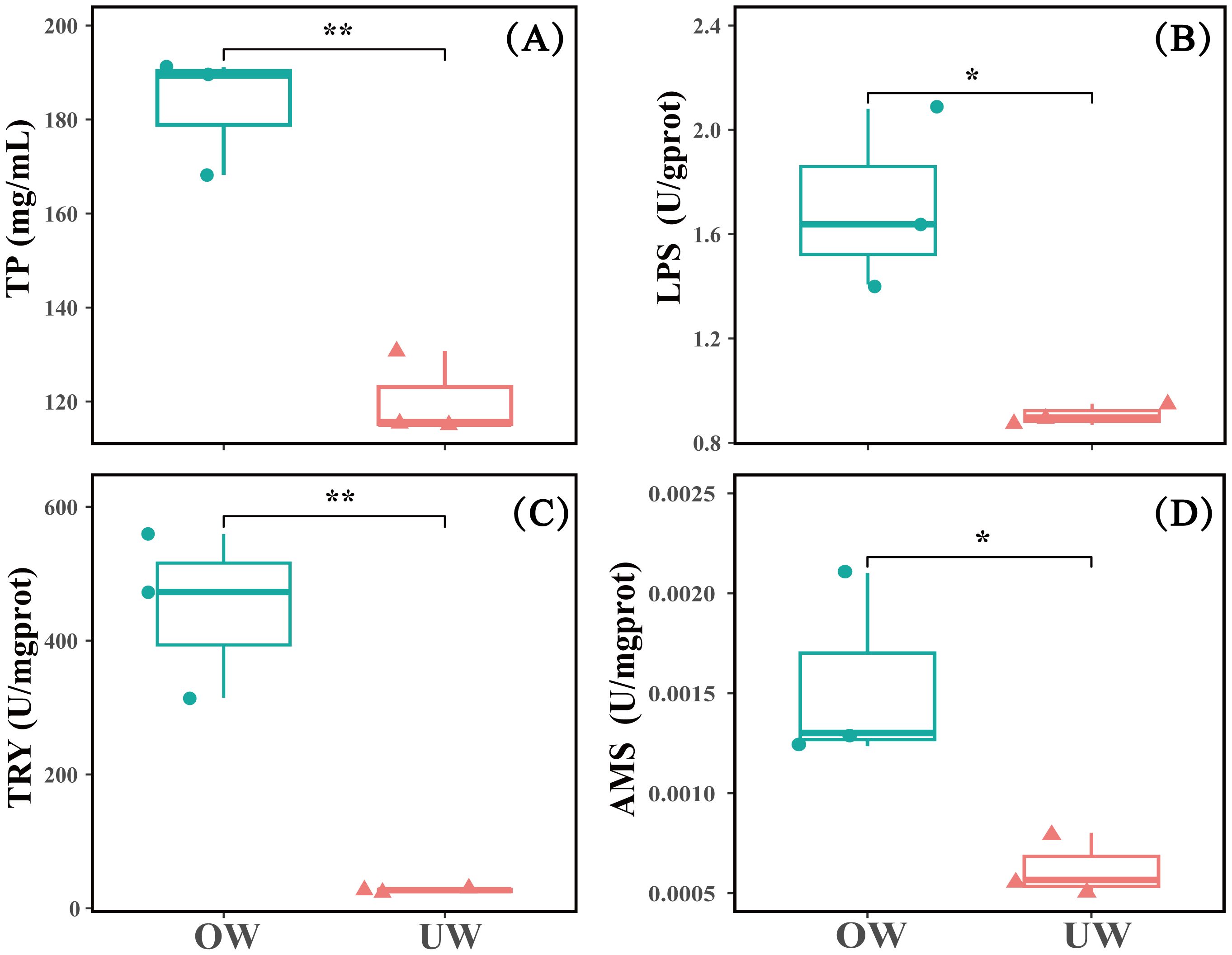
Figure 4. Differences in intestinal digestive enzyme activity among individuals between the overweight (OW) and underweight (UW) groups (n = 3). (A) TP represents total protein content; (B) LPS represents intestinal lipase activity; (C) TRY represents intestinal trypsin activity, and (D) AMS represents intestinal amylase activity. * indicates a significant difference at the P < 0.05 level; ** indicates a highly significant difference at the P < 0.01 level. And that with over 99.9% certainty that the result is not accidental.
Analysis of the intestinal flora structure
Based on 16S rRNA sequencing, we obtained 81,299 high-quality biological sequences from different individuals. The ratio of Valid data to Raw data was above 90%, and the proportion of Q20 data was above 98%. Most of the rarefaction curves for the samples reached saturation, suggesting sufficient sequencing depth for this study. Amplicon sequence variants (ASVs) were defined based on 97% identity. A total of 632 ASVs were obtained from all fish. The total number of ASVs shared by the two groups was 130; 376 ASVs were unique to the OW group (59.49%), and 126 ASVs were unique to the UW group (19.94%). PCA analysis based on ASV composition can reflect the differences and distances between samples. The same treatment groups were closely clustered; the principal components were similar, and the amount of variation was low, indicating good repeatability between samples. The treatment groups were clearly separated, and the degree of dispersion was high, indicating a large difference between the samples. The results also indicated that the composition of intestinal flora in small fish bodies was dissimilar (Figure 5A). Further analysis of the bacterial community composition at the genus level was conducted, and the top 30 significantly enriched bacterial genera were obtained. The bacterial community composition was similar among samples, but the abundance and dominant bacterial groups differed. The dominant strain in the OW group was Burkholderia-Caballeronia-Paraburkholderia, and the dominant strain in the UW group was Cetobacterium (Figure 5B).
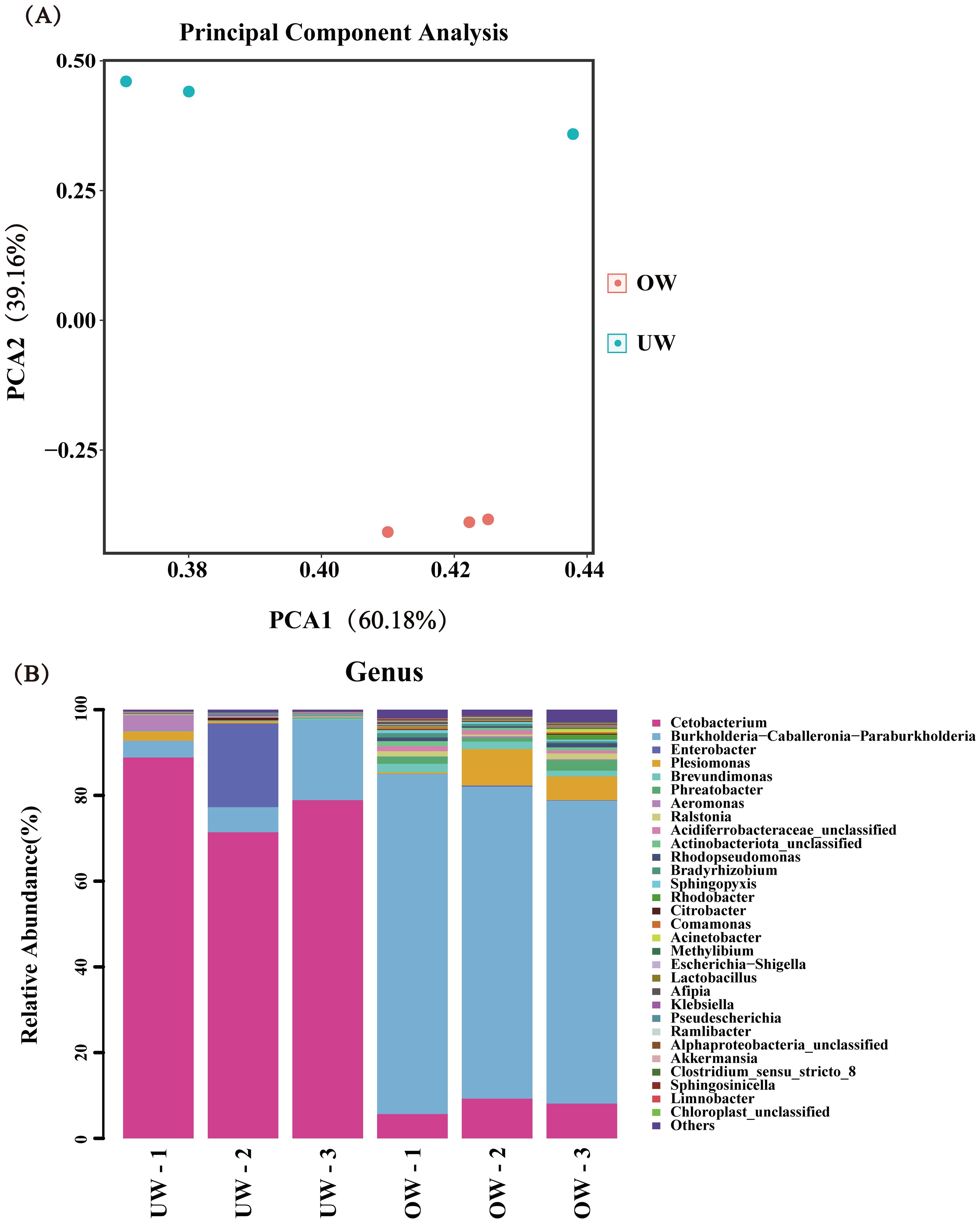
Figure 5. The distribution of dominant intestinal flora between the overweight (OW) and underweight (UW) groups of fish at the genus level. (A) Principal component analysis of intestinal flora in different groups. Different colors represent different groups, and points with the same color represent different samples within a group. The same group is displayed in the form of a circle graph (n = 3) according to the 95% confidence interval. The closer the circles, the higher the similarity between the samples. (B) The distribution of dominant flora at the genus level. Different colors represent different flora.
Transcriptomic analysis
High-throughput sequencing technology was used to conduct transcriptome sequencing, and initial data of clean reads and GC content were obtained after removing low-quality and junction sequences. A total of 71,513 clusters were obtained from the scratch assembly of the clean reads using the Trinity software. To verify the sequencing quality of the transcriptome, 16 genes were randomly selected from the database, and the expression levels of genes in individuals of different sizes were detected by qRT-PCR. In the gene expression pattern and expression profile data, the quantitative results of 12 genes (75%) were consistent with the transcriptome expression profile, indicating high accuracy of the transcriptome data (Figure 6A). A total of 6648 differentially expressed genes were identified in the OW group compared with the UW group, including 2523 upregulated genes and 4125 downregulated genes (Figure 6B). Several growth-related genes were differentially expressed, including growth hormone-releasing hormone and fibroblast growth factor. Analysis of metabolic pathways using the KEGG database showed that the pathways with enriched genes included Biosynthesis of amino acids, 2-Oxocarboxylic acid metabolism, Hematopoietic cell lineage, Glycerolipid metabolism, and the HIF-1 signaling pathway (Figure 6C). To further analyze the expression patterns of growth-related regulatory genes in different individuals of O. bidens, we screened genes related to amino acid metabolism, signal transduction, and other anabolic pathways from the enriched KEGG pathways for expression profile level analysis. The actual expression levels of the related genes on the pathway were verified by qRT-PCR. The results demonstrated that the quantitative data were consistent with the transcriptome expression profiles. The analysis revealed that the expression levels of growth hormone–releasing hormone, vascular endothelial growth factor A, fibroblast growth factor, aminoacylase, serine/threonine protein kinase, mitogen-activated protein kinase, heme oxygenase, and other coding genes in the metabolic pathway were significantly higher in the OW group than in the UW group (Figure 7).
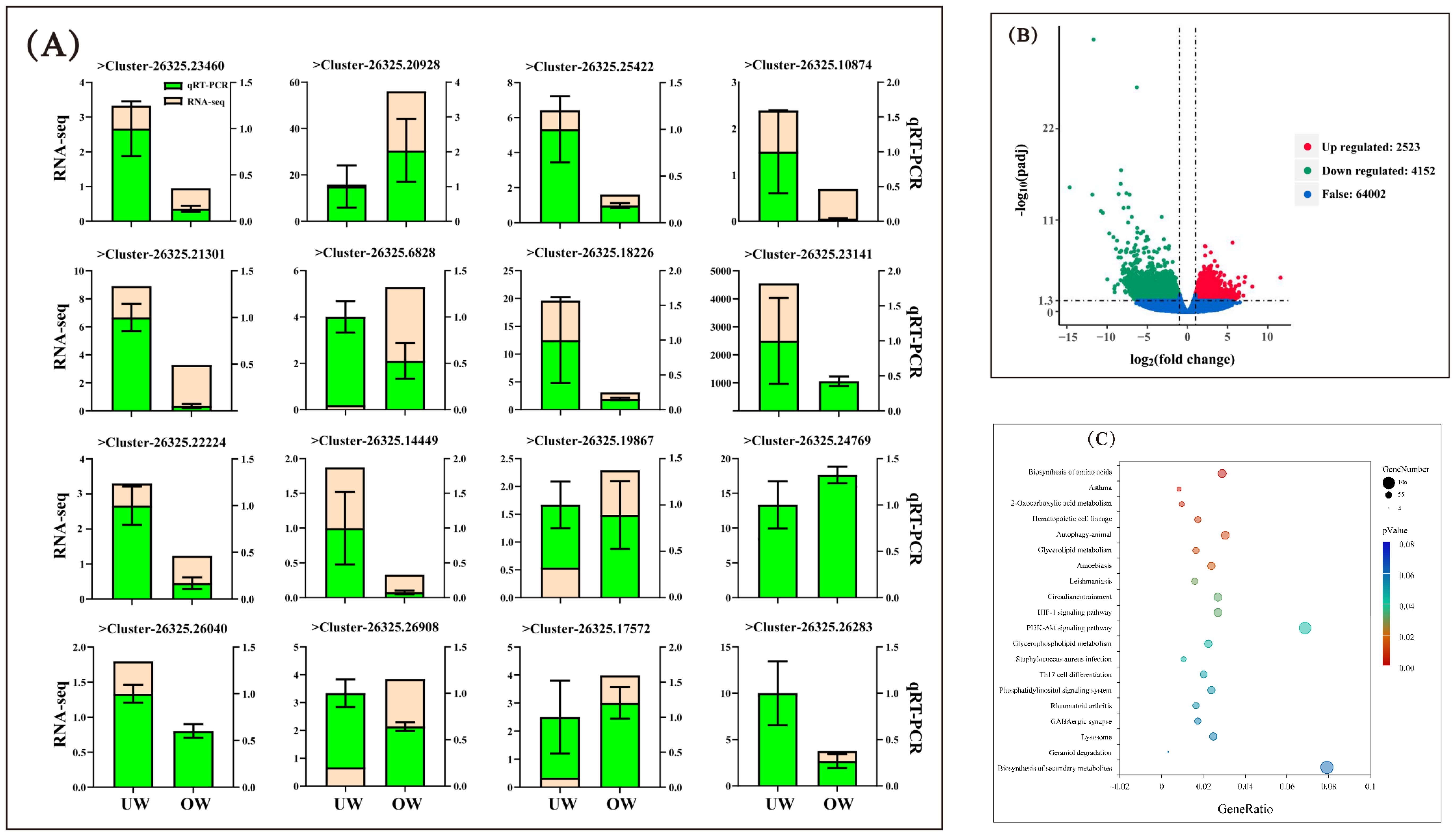
Figure 6. Analysis of differentially expressed genes. (A) The results of qRT-PCR verification of genes in different-sized individuals. The light yellow color indicates the expression level of the gene in the transcriptome expression profile, and the fluorescent green color indicates the expression trend of the gene in different individuals tested by qRT-PCR. The qRT-PCR data are presented as mean ± SD, n = 4. (B) The horizontal coordinate is the log2 Fold Change value. The vertical coordinate is –log10 padj or –log10 P-value, and the dashed blue line represents the threshold line of the differential gene screening criteria. (C) Bubble map of KEGG enrichment of differentially expressed genes. The scatter diagram illustrates KEGG pathway enrichment: the vertical axis represents the pathway name; the horizontal axis represents the Gene Ratio corresponding to the pathway, and the q-value is represented by the color of the dots. The smaller the q-value, the closer the color to red. The number of differential genes contained in each pathway is represented by the size of the dots. The more differential genes, the larger the spot.
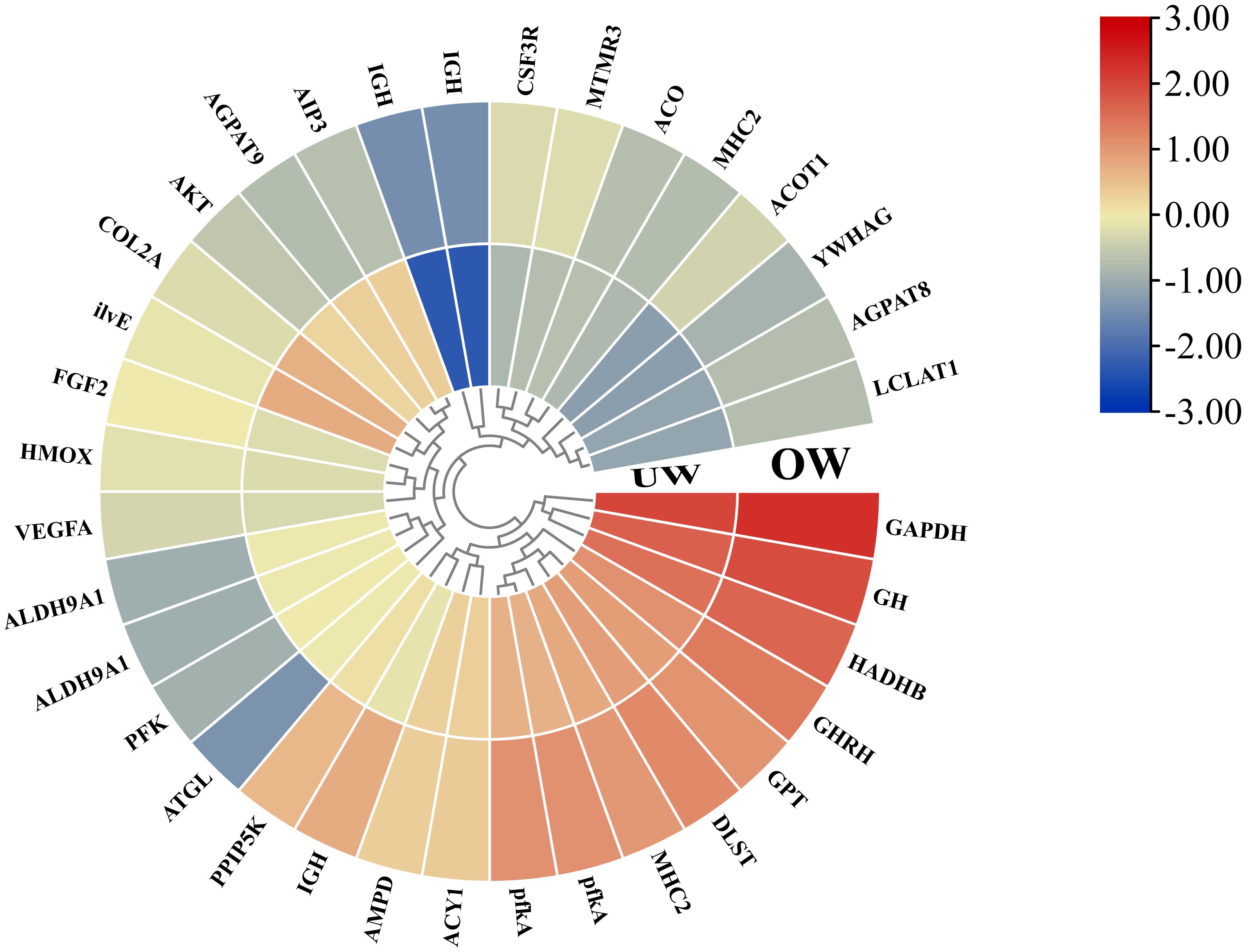
Figure 7. Analysis of gene expression patterns related to anabolic pathways. The gene expression level under each treatment was used to construct a normalized heat map, and the expression level difference was more than twice that of the differential gene.
Correlation analysis of growth performance, physiological and biochemical indices, and intestinal microbiota
An association analysis identified correlations between morphological indices, intestinal structure, and digestive enzyme activity, factors closely related to intestinal microbiota but having low correlations with gene transcription (Figure 8A). The results of the SEM were basically consistent with the abovementioned correlation analysis, and the direct and indirect effects between variables were further verified (Figure 8B). Intestinal microbiota positively affected gene transcription (0.477); gene transcription positively affected the intestinal morphological structure (0.416); the intestinal morphological structure positively affected intestinal digestive enzymes (2.714), and intestinal digestive enzymes positively affected the physiological phenotype (0.516). In addition, intestinal microbiota had a positive effect on the intestinal morphological structure (0.733), and intestinal morphological structure had a positive effect on physiological phenotypes (0.513).
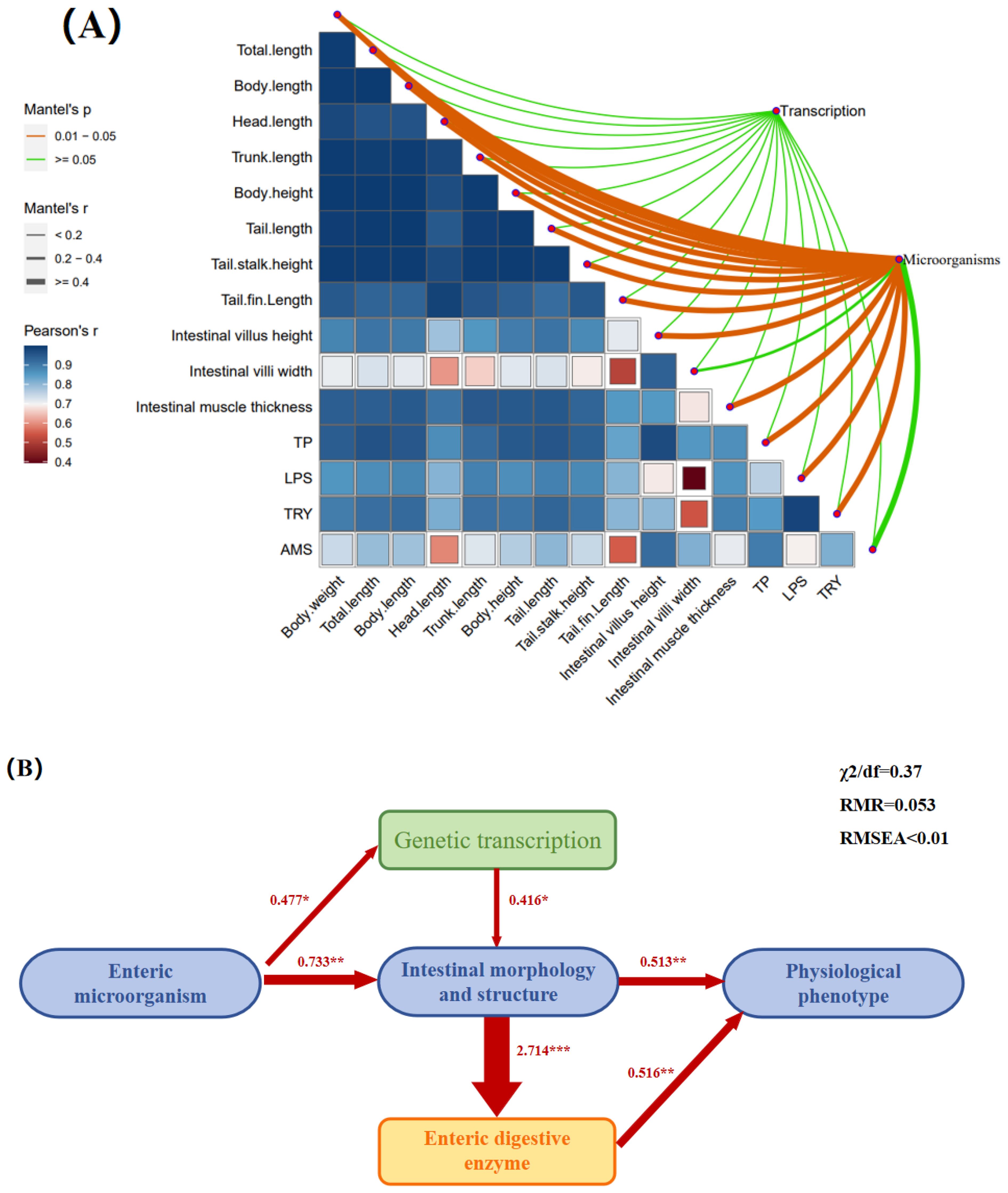
Figure 8. Correlations among different indicators. (A) A heat map of the correlations between body weight, total length, body length, head length, trunk length, body height, tail length, tail stalk height, tail fin length, intestinal muscle thickness, villi length, villi width, total protein, lipase, amylase, and trypsin. (B) Correlation indices of the intestinal microbiome, gene transcription, intestinal morphological structure, intestinal digestive enzyme activity, and physiological phenotype. * indicates a significant difference at the P < 0.05 level; ** indicates a highly significant difference at the P < 0.01 level; *** indicates an extremely significant difference at the P < 0.001 level. And that with over 99.9% certainty that the result is not accidental.
Discussion
Intestinal microbiota promote the development of the intestinal tract to accelerate the growth of O. bidens
The microbiota and the environment are interdependent; the breeding environment is regulated by the microbiota, which are supported by the environment. The health of aquatic organisms is closely tied to their environment. As a crucial component of the internal ecosystem, the intestinal microflora not only support nutritional metabolism and immune defense but also stabilize the internal environment to maintain physiological homeostasis (Blaxter et al., 2005; Wang et al., 2012).
The intestinal tract functions to absorb food nutrients, and its morphological structure determines the digestion and absorption ability of fish. The intestinal morphological structure includes intestinal villus height, intestinal villus width, intestinal muscle thickness, crypt depth, and the pillow–crypt ratio (Zhang et al., 2020). The height and width of the intestinal villi are indicators of the intestinal tract’s ability to absorb nutrients (Sang and Fotedar, 2009). Studies have shown that intestinal microbiota is essential for maintaining normal gut structure and function (Lutgendorff et al., 2008). The soluble protein p40 derived from probiotics can activate EGFR, leading to Akt activation and thereby preventing cytokine-induced apoptosis and structural damage to intestinal epithelial cells (Yan et al., 2011). The growth and health status of fish are reflected in their intestinal microbiota and the presence of probiotics. A higher abundance of probiotics produces a more stable intestinal environment, leading to improved growth status in fish (Yu et al., 2018). Zebrafish without microflora are unable to digest or absorb large protein molecules, and the artificial construction of microbial communities can help zebrafish recover and improve their digestion and absorption of proteins (Bates et al., 2006). In this study, the intestinal flora abundance of the OW group was higher than that of the UW group, and its intestinal environment was relatively more stable, indicating a stronger ability to digest and absorb nutrients. Some species of (Burkholderia-Caballeronia-Paraburkholderia), a common component of the intestinal microbiome, produce growth hormones that can improve the growth rate and health status of the host, and thus the relative abundance of this group is often closely related to the growth and health status of the host (Blandford et al., 2018; Rong et al., 2024). Our results were similar to those of previous studies, as (Burkholderia-Caballeronia-Paraburkholderia) was dominant in the intestinal tract among the experimental groups of O. bidens. The results showed that (Burkholderia-Caballeronia-Paraburkholderia) were the most abundant bacteria in the OW group while being the second most abundant taxon in the UW group, indicating that the intestinal health of the OW group was superior to that of the UW group (Figure 5B). Further analysis found that the height and width of intestinal villi in the OW group were significantly greater than those in the UW group, and the higher and wider intestinal villi increased the contact area between the villi and nutrients, thereby improving the digestive and absorption functions of the intestine. The thickness of the intestinal muscle can also reflect the integrity of the intestinal tissue structure. The muscle thickness of the OW group was about 1.4 times that of the UW group, and the thicker the muscle thickness, the greater the intestinal absorption efficiency (Figures 2 and 3). Previous studies have confirmed that improving the intestinal structure can enhance the growth performance of fish (Zhu, 2021). This study suggests a correlation between the intestinal microbiota and the intestinal morphological structure of O. bidens. The correlation analysis revealed a positive association between intestinal microbiota and intestinal morphological structure, with a correlation coefficient of 0.733 (Figure 8). We hypothesized that the intestinal microbiota may influence intestinal development, thereby facilitating more complete absorption of nutrients and accelerating the growth rate.
Growth-promoting mechanisms of intestinal microbiota on the performance of O. bidens
Intestinal microbiota play a crucial role in regulating digestion and metabolism, as well as growth, development, and immune defense. Symbiotic bacteria in the intestine can directly affect the host’s digestive function. The activity of digestive enzymes in fish intestines is an important index of the ability of fish to digest and absorb nutrients (Chen et al., 2019). Lipase (LPS), amylase (AMS), and trypsin (TRY) are the main digestive enzymes in fish intestines, and their activities not only reflect the absorption and utilization efficiency of nutrients but also play a key role in growth (Qin et al., 2018; Jin et al., 2019). Probiotics can produce the aforementioned digestive enzymes in the intestine, promoting nutrient absorption and the assimilation of food by fish (Douglas, 2014). The enhancement of lipase and protein activities could improve the activity of intestinal proteases, further promoting the growth of fish (Wang et al., 2023). Trypsin, as the most important protease among intestinal digestive enzymes, is also an important indicator of the digestion and absorption ability of fish (Ding and Li, 2011). An increase in intestinal trypsin activity improved the utilization rate of carbohydrates and promoted the growth of groupers (Liu et al., 2023). This study confirmed the results of previous research, finding that the intestinal total protein content of the OW group was much higher than that of the UW group, and the activities of lipase and amylase were two to three times higher than those of the UW group. In addition, the trypsin activity of the OW group was approximately 17 times higher than that of the UW group (Figure 4). The results indicated that intestinal microbiota can enhance intestinal digestive enzyme activity, thereby increasing nutrient absorption and promoting the growth of O. bidens.
The role of intestinal microbiota extends beyond regulating nutrient absorption in fish to influencing the expression of specific genes. Rawls et al. (2004) studied 79 young zebrafish and found that apolipoprotein, myeloperoxidase, serum amyloprotein, and other genes exhibited strong responses to the presence or absence of ungraded microbiota in the gut and that intestinal microbiota controlled the expression of 212 genes during aseptic basal culture. In this study, the results of gene expression profiles showed that the KEGG pathways of differentially expressed genes were concentrated in cell differentiation, the PI3K-Akt signaling pathway (promoting cell survival), the phosphatidylinositol signaling system, phospholipid metabolism, and biosynthesis of secondary metabolites (Figure 6C). The expression patterns of differentially expressed genes in the enriched pathways were further analyzed, and GH (growth hormone), GHRH (growth hormone-releasing hormone), FGF2 (fibroblast growth factor), pfkA (vascular endothelial growth factor A), ALT (alanine aminotransferase) and other differentially expressed genes directly or indirectly related to growth were screened. Studies have shown that the GH/IGF axis plays a crucial role in regulating the growth and development of fish, with GH and GHRH being key genes in this endocrine axis (Zheng and Ru, 2012). GHRH is an important neuroendocrine factor in fish. GHRH releases GH by stimulating the anterior pituitary, which in turn regulates the growth and development of fish by binding to the growth hormone receptors (GHRs) in liver cells (Nam et al., 2011). Therefore, GH and GHRH play positive roles in regulating the growth and development of fish. In this study, GH and GHRH gene expression in the OW group was significantly higher than in the UW group. In addition, the expression levels of most genes related to positive growth regulation, such as FGF2, pfk2, and ALT, in the OW group were significantly higher than those in the UW group (Figure 7). The correlation analysis of different indicators identified a positive association between intestinal microbiota and gene transcription and expression (Figure 8). We hypothesized that intestinal microbiota may further participate in the regulation of the rapid growth of the OW group by positively regulating genes.
Conclusion
In this study, despite having identical environmental conditions and shared genetic backgrounds, significant variation was observed in the phenotypic growth of O. bidens, which may be attributed to changes in the composition of the endogenous intestinal microbiota. There were correlations between the intestinal microbiota and gene expression related to growth, the intestinal structure, and digestive enzyme activity. We speculate that intestinal microbiota, on the one hand, promote the development of intestinal tissue structure, thereby increasing the surface area for nutrient absorption by enhancing the contact between intestinal villi and food. On the other hand, the microbiota regulate the expression of genes involved in the production and activity of digestive enzymes and transport proteins, ultimately enhancing digestive efficiency. This, in turn, promotes the growth of O. bidens. These results have deepened our understanding of the regulatory mechanisms of the development of O. bidens and provide theoretical guidance for breeding rapidly growing varieties.
Data availability statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA1159450 and PRJNA1159474.
Ethics statement
The animal study was approved by Laboratory Animal Centre, Freshwater Fisheries Research Institute of Fujian. The study was conducted in accordance with the local legislation and institutional requirements.
Author contributions
GZ: Methodology, Writing – original draft, Validation, Writing – review & editing, Investigation. ML: Methodology, Validation, Writing – review & editing. MW: Conceptualization, Supervision, Writing – review & editing. ZHH: Writing – review & editing, Investigation, Validation. ZXH: Validation, Investigation, Writing – review & editing. DC: Writing – review & editing, Methodology. HF: Conceptualization, Methodology, Writing – review & editing. YG: Writing – review & editing, Methodology, Validation. YZ: Data curation, Investigation, Writing – review & editing. MN: Validation, Writing – review & editing. ZD: Writing – original draft, Investigation, Conceptualization, Writing – review & editing. LX: Writing – original draft, Writing – review & editing, Investigation, Methodology. BW: Conceptualization, Writing – original draft, Investigation, Writing – review & editing.
Funding
The author(s) declare financial support was received for the research and/or publication of this article. This work was supported by Fujian Provincial Department of Science and Technology (grant numbers 2021R1014004, 2022R1014004, and 2024R1014001) and Fujian Provincial Department of Ocean and Fisheries (grant numbers 2021FJSCZY04 and FJHY-YYKJ-2024-2-7).
Acknowledgments
We thank LetPub (www.letpub.com.cn) for its linguistic assistance during the preparation of this manuscript.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2025.1635417/full#supplementary-material
References
Bates J. M., Mittge E., Kuhlman J., Baden K. N., Cheesman S. E., and Guillemin K. (2006). Distinct signals from the microbiota promote different aspects of zebrafish gut differentiation. Dev. Biol. 297, 374–386. doi: 10.1016/j.ydbio.2006.05.006
Blandford M. l., Taylor-Brown A., Schlacher T. A., Nowak B., and Polkinghorne A. (2018). Epitheliocystis in fish: An emerging aquaculture disease with a global impact. Transboundary Emerging Dis. 65, 1436–1446. doi: 10.1111/tbed.12908
Blaxter M., Mann J., Chapman T., Thomas F., Whitton C., Floyd R., et al. (2005). Defining operational taxonomic units using DNA barcode data. Philos. Trans. R Soc B. 360, 1935–1943. doi: 10.1098/rstb.2005.1725
Blumberg R. and Powrie F. (2012). Microbiota, disease, and back to health: a metastable journey. Sci. Transl. Med. 4, 137rv7. doi: 10.1126/scitranslmed.3004184
Chen M., Fu G. X., Dong J. E., and He W. F. (2019). Effects of rutin on growth performance, body composition,non-specific immunity and intestinal digestive enzyme activities of grass carp (Ctenopharyngodon idella). Chin. J. Anim. Nutr. 31, 4868–4876. doi: 10.3969/j.issn.1006-267x.2019.10.053
Chen Y. S., Xue L. Z., Chen D. H., Hu Z. X., Liao M. X., Wu M. Y., et al. (2023). Effects of different oxytocin formulae and postpartum care on artificial breeding of Opsariichthys uncirostris bidens Gunther. Hunan Agric. Sci. 09, 33–38. doi: 10.16498/j.cnki.hnnykx.2023.009.007
Ding L. and Li Y. W. (2011). Research progress of trypsin in cold-water fish. Hebei Fisheries 12, 50–54. doi: 10.3969/j.issn.1004-6755.2011.12.015
Douglas A. E. (2014). Multiorganismal insects: diversity and function of resident microorganisms. Annu. Rev. Entomol. 60, 17–34. doi: 10.1146/annurev-ento-010814-020822
Francesco B., Laura G., Marco B., Angela T., Christian C., Cristina B., et al. (2021). Performance and fillet traits of rainbow trout (Oncorhynchus mykiss) fed different levels of Hermetia illucens meal in a low-tech aquaponic system. Aquaculture. 546, 737279. doi: 10.1016/j.aquaculture.2021.737279
Gjerde B. and Korsvoll A. (1999). Realised selection differentials for growth rate and early sexual maturity in Atla Intic salmon. Aquac. Eur. 99, 73–74. doi: 10.1080/13880299909353933
Jin Y. Q., Yu E. M., Zhang K., Li Z. P., Wang G. J., Xie J., et al. (2019). Effects of three feeds on serum enzyme activity, intestinal structure and bacterial flora of Ctenopharyngodon idellus. J. Agric. Biotechnol. 27, 1652–1663. doi: 10.3969/j.issn.1674-7968.2019.09.014
Kim B. H., Hur S. P., Choi S. H., Lee Y. D., and Lee C. H. (2023). Exploring rearing water temperature conditions for inducing body growth without water temperature stress in red-spotted grouper (Epinephelus akaara). Ocean Sci. J. 58, 25. doi: 10.1007/s12601-023-00117-y
Kuang T. X., He A. Y., Lin Y. F., Huang X. D., Liu L., and Zhou L. (2020). Comparative analysis of microbial communities associated with the gill, gut, and habitat of two filter-feeding fish. Aquacult Rep. 18, 100501. doi: 10.1016/j.aqrep.2020.100501
Li W. D. (2014). Effect of Dietary Lipid Level on growth performance, body composition and digestive enzyme activities of Ancherythroculter nigrocauda (Wuhan, China: Wuhan polytechnic University).
Li C., Feng W., Qiu L., Xia C., Su X., and Jin C. (2012). Characterization of skin ulceration syndrome associated micrornas in sea cucumber Apostichopus japonicus by deep sequencing. Fish Shellfish Immunol. 33, 436–441. doi: 10.1016/j.fsi.2012.04.013
Li W. H., Liu J. M., Tan H., Yang C. H., Ren L., Liu Q. F., et al. (2018). Genetic Effects on the gut microbiota assemblages of hybrid fish from parents with different feeding habits. Front. Microbiol. 9. doi: 10.3389/fmicb.2018.02972
Liang M., Dong S. L., Gao Q. F., Wang F., and Tian X. L. (2010). Individual variation in growth in sea cucumber Apostichopus japonicus (selenck) housed individually. J. Ocean Univ. China. 9 (003), 291–296. doi: 10.1007/s11802-010-1739-4
Liu H. Y., Pan L., Shen J. F., Tan B. P., Dong X. H., Yang Q. H., et al. (2023). Effects of carbohydrase supplementation on growth performance, intestinal digestive enzymes and flora, glucose metabolism enzymes, and glut2 gene expression of hybrid grouper (Epinephelus fuscoguttatus♀ × E. lanceolatus♂) fed different CHO/L ratio diets. Metabolites. 13, 98. doi: 10.3390/metabo13010098
Liu X. Y., Chen C., Gong W. Q., Chen K. J., Li Q. S., Liu P., et al. (2015). Biological Characteristics of Opsariichthys bidenin Anhua. Journal of Beijing University of Agriculture. 49–52. doi: 10.13473/j.cnki.issn.1002-3186.2015.0024
Lutgendorff F., Akkermans L. M. A., and Söderholm J. D. (2008). The role of microbiota and probiotics in stress-induced gastro-intestinal damage. Curr. Mol. Med. 8, 282–298. doi: 10.2174/156652408784533779
Mao H. D., Wang N., Sui C., Liu Z. Y., Zhang D. Y., and Chen J. H. (2023). Effects of dietary methionine on growth performance of juvenile green-finned filefish (Thamnaconus septentrionalis). Feed Industry (in Chinese). 45, 1–13. doi: 10.13302/j.cnki.fi.2024.06.011
Meng Q. W., Li W. D., and Zhou B. Y. (1995). Experimental guidance of fish science (Beijing, China: China Agriculture Press).
Nam B. H., Moon J. Y., Kim Y., Kong H. J., Kim W. J., Kim K. K., et al. (2011). Molecular and functional analyses of growth hormone-releasing hormone (GHRH) from olive flounder (Paralichthys olivaceus). Comp. Biochem. Physiol Part B: Biochem. Mol. Biol. 152, 84–91. doi: 10.1016/j.cbpb.2011.02.006
Nayak S. K. (2010). Role of gastrointestinal microbiota in fish. Aquacult Res. 41, 1553–1573. doi: 10.1111/j.1365-2109.2010.02546.x
Qin Q., Chen X. H., Jiang G. Z., Wang M. H., Zhang S. Y., Zhong L. Q., et al. (2018). Comparison of growth, body composition and digestive enzyme activities on five families of yellow catfish. Genomics Appl. Biol. 37, 768–773. doi: 10.13417/j.gab.037.000768
Rawls J. F., Samuel B. S., and Gordon J. I. (2004). Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proc. Natl. Acad. Sci. U S A. 101, 4596–4601. doi: 10.1073/pnas.0400706101
Ringø E., Myklebust R., Mayhew T. M., and Olsen R. K. (2007). Bacterial translocation and pathogenesis in the digestive tract of larvae and fry. Aquaculture. 268, 251–264. doi: 10.1016/j.aquaculture.2007.04.047
Rong H., Xie Y. Q., Zhao T., Yin H., Yuan L. Y., Liu X. H., et al. (2024). Comparative analysis of the composition and diversity of the intestinal microflora of four fishes with different feeding habits. Prog. Fishery Sci. 45, 1–12. doi: 10.19663/j.issn2095-9869.20230315001
Sang H. M. and Fotedar R. (2009). Effects of mannan oligosaccharide dietary supplementation on performances of the tropical spiny lobsters juvenile (Panulirus ornatus, Fabricius 1798). Fish Shellfish Immunol. 28, 483–489. doi: 10.1016/j.fsi.2009.12.011
Su J. X., Qiu F. X., Zhou W. X., Xie C. X., Wang W. M., and Chen C. F. (1993). Age, growth, reproduction and feeding habits of Opsariichthys bidens in Sandaohe Reservoir. J. Hydroecology 1, 15–18. Available online at: http://dianda.cqvip.com/Qikan/Article/Detail?id=1173363.
Suo Y. T., Li E. C., Li T. Y., Jia Y. Y., Qin J. G., Gu Z. M., et al. (2017). Response of gut health and microbiota to sulfide exposure in Pacific white shrimp Litopenaeus vannamei. Fish Shellfish Immunol. 63, 87–96. doi: 10.1016/j.fsi.2017.02.008
Tran N. T., Zhang J., Xiong F., Wang G. T., Li W. X., Wu S. G., et al. (2018). Altered gut microbiota associated with intestinal disease in grass carp (Ctenopharyngodon idellus). World J. Microbiol. Biotechnol. 34, 1–9. doi: 10.1007/s11274-018-2447-2
Valdes A., Walter J., Segal E., and Spector T. (2018). Role of the gut microbiota in nutrition and health. BMJ 361, k2179. doi: 10.1136/bmj.k2179
Wang A., Ran C., Ring E., and Zhou Z. (2018). Progress in fish gastrointestinal microbiota research. Rev. Fish Sci. 10, 626–640. doi: 10.1111/raq.12191
Wang Y., Sheng H. F., He Y., Wu J. Y., Jiang Y. X., Tam N. F. Y., et al. (2012). Comparison of the levels of bacterial diversity in freshwater, intertidal wetland, and marine sediments by using millions of illumina tags. Appl. Environ. Microbiol. 78, 8264–8271. doi: 10.1128/aem.01821-12
Wang X., Wan M., Wang Z., Zhang H. T., Zhu S., Cao X. F., et al. (2023). Effects of tributyrin supplementation on growth performance, intestinal digestive enzyme activity, antioxidant capacity, and inflammation-related gene expression of large yellow croaker (Larimichthys crocea) fed with a high level of clostridium autoethanogenum protein. Aquacult Nutr. 1, 2687734. doi: 10.1155/2023/2687734
Yan F., Cao H. W., Cover T. L., Washington M. K., Shi Y., Liu L. S., et al. (2011). Colon-specific delivery of a probiotic-derived soluble protein ameliorates intestinal inflammation in mice through an EGFR-dependent mechanism. J. Clin. Invest. 121, 2242–2253. doi: 10.1172/jci44031
Yu W. N., Dai W. F., Tao Z., and Xiong J. B. (2018). Characterizing the compositional and functional structures of intestinal micro-flora between healthy and diseased Litopenaeus vannamei. J. Fish. China. 42, 399–409. doi: 10.11964/jfc.20170310737
Zang L., Ma Y., Huang W., Ling Y., Sun L., Wang X., et al. (2019). Dietary Lactobacillus plantarum ST-III alleviates the toxic effects of triclosan on zebrafish (Danio rerio) via gut microbiota modulation. Fish Shellfish Immunol. 84, 1157–1169. doi: 10.1016/j.fsi.2018.11.007
Zhan Y. J. (2011). Life history oftwo species of Danioninae in the Qingyi Stream (Anhui, China: Anhui normal university).
Zhang N., Zhou Y. W., Bai Y., Zhou Z. K., and Jiang Y. M. (2020). Research progress in effect of microplastics on intestinal flora. China Plast. 34, 103–110. doi: 10.19491/j.issn.1001-9278.2020.03.015
Zheng Y. and Ru S. G. (2012). Effects of exogenous chemicals on fish growth and GH/IGF-I axis Vol. 42 (China: Period. Ocean Univ), 102–106. doi: 10.16441/j.cnki.hdxb.2012.s1.015
Zhong Z. X. (1985). Preliminary observations on the impact of Opsariichthys uncirostris bidens Günther on reservoir fisheries. Curr. Fisheries 5, 20.
Keywords: Opsariichthys bidens, intestinal microbiota, intestinal structure, digestive enzyme activity, multiomics
Citation: Zeng G, Liao M, Weng M, Hu Z, Hu Z, Chen D, Fan H, Gao Y, Zhang Y, Naz M, Dai Z, Xue L and Wu B (2025) The growth-promoting role of the intestinal microbiota in Opsariichthys bidens: implications for selective breeding. Front. Mar. Sci. 12:1635417. doi: 10.3389/fmars.2025.1635417
Received: 26 May 2025; Accepted: 07 August 2025;
Published: 29 August 2025.
Edited by:
Rodrigo Vidal, University of Santiago, ChileReviewed by:
Zhimin Zhang, Chinese Academy of Sciences (CAS), ChinaXingkun Jin, Hohai University, China
Copyright © 2025 Zeng, Liao, Weng, Hu, Hu, Chen, Fan, Gao, Zhang, Naz, Dai, Xue and Wu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhicong Dai, ZGFpemhpY29uZ0AxNjMuY29t; Lingzhan Xue, eGx6NTg3MkAxNjMuY29t; Bin Wu, d3ViaW5maXJlQDEyNi5jb20=
 Gaoxiong Zeng
Gaoxiong Zeng Mengxiang Liao1
Mengxiang Liao1 Yu Gao
Yu Gao Misbah Naz
Misbah Naz Zhicong Dai
Zhicong Dai