- 1Department of Chemistry, University of California, Davis, Davis, CA, United States
- 2Plant Biology Graduate Group, University of California, Davis, Davis, CA, United States
Biological chemical production has gained traction in recent years as a promising renewable alternative to traditional petrochemical based synthesis. Of particular interest in the field of metabolic engineering are photosynthetic microorganisms capable of sequestering atmospheric carbon dioxide. CO2 levels have continued to rise at alarming rates leading to an increasingly uncertain climate. CO2 can be sequestered by engineered photosynthetic microorganisms and used for chemical production, representing a renewable production method for valuable chemical commodities such as biofuels, plastics, and food additives. The main challenges in using photosynthetic microorganisms for chemical production stem from the seemingly inherent limitations of carbon fixation and photosynthesis resulting in slower growth and lower average product titers compared to heterotrophic organisms. Recently, there has been an increase in research around improving photosynthetic microorganisms as renewable chemical production hosts. This review will discuss the various efforts to overcome the intrinsic inefficiencies of carbon fixation and photosynthesis, including rewiring carbon fixation and photosynthesis, investigating alternative carbon fixation pathways, installing sugar catabolism to supplement carbon fixation, investigating newly discovered fast growing photosynthetic species, and using new synthetic biology tools such as CRISPR to radically alter metabolism.
Introduction
It is well established that rising atmospheric CO2 levels are the primary cause for unprecedented climate change impacting the globe (Solomon et al., 2009). Despite this, chemical production still relies mostly on petroleum-based synthesis (Levi and Cullen, 2018). In response to the growing concern over greenhouse gasses, research with a focus on more sustainable chemical production has become high priority. The fields of synthetic biology and metabolic engineering aim to achieve a more sustainable method for chemical production using engineered organisms. These efforts include the use of both heterotrophic and photosynthetic microorganisms. Heterotrophic chemical production involves a carbon input of a sugar feedstock to a microorganism to generate a biochemical product as an output. Alternatively, using photosynthetic microorganisms, such as cyanobacteria, offers the advantage of eliminating the need for sugar feedstocks and the ability to generate valuable chemical commodities from CO2 and sunlight. The two predominant categories of photosynthetic microorganisms being investigated for chemical production are microalgae and cyanobacteria. Microalgae are a diverse group of photosynthetic eukaryotes that have been shown to be viable production hosts for a wide array of useful chemical commodities ranging from biofuels to lipids and vitamins (Sproles et al., 2021). Cyanobacteria are a group of prokaryotic microorganisms with some of the fastest carboxylation rates present in photosynthetic organisms (Flamholz et al., 2019). The focus of this review will be on recent synthetic biology research in cyanobacteria which have garnered interest as efficient photosynthetic chemical production hosts.
Despite the burgeoning interest around these photosynthetic microorganisms, the field still faces many challenges that have yet to be addressed. The primary concern is the inefficient nature of photosynthesis and CO2 fixation. Attempts to improve upon the central carbon fixation enzyme ribulose-1,5-bisphosphate carboxylase-oxygenase (RuBisCO) have been met with little success (Erb and Zarzycki, 2018; Flamholz et al., 2019). Part of the challenge with RuBisCO is its inability to distinguish between CO2 and O2 with high specificity and notably, the oxygenase activity of RuBisCO results in an energetically costly pathway known as photorespiration which has widespread effects on the growth and metabolic needs of many species of photosynthetic organisms (Hagemann and Bauwe, 2016). Recent research suggests that photorespiration is a symptom of RuBisCO evolving in a high CO2 environment where enzymatic specificity was not as vital, with this in mind, recent studies are investigating the possibilities of by reviving ancestral forms of the protein and subjecting it to new environments in the hopes of generating biologically important variants (Shih et al., 2016). Other efforts are looking towards natural adaptations of CO2 fixation for inspiration with a focus on the carboxysome, a bacterial microcompartment that acts to localize RuBisCO with high concentrations of CO2 (Kerfeld and Melnicki, 2016). These studies aim to avoid photorespiration by engineering synthetic protein structures to mimic cyanobacterial carboxysomes to concentrate CO2 near RuBisCO and competitively inhibit the reaction with oxygen (Borden and Savage, 2021). Other efforts to further improve these photosynthetic organisms as chemical production hosts include engineering superior light delivery systems for bioreactors and engineering the light harvesting complexes to take advantage of the entire visible light spectrum (Stephens et al., 2021).
The focus of this review will be on the recent methods employed to overcome the supposed shortcomings of photosynthetic organisms ranging from rewiring carbon metabolism and photosynthesis, introducing additional carbon substrates, generating other chassis organisms capable of superior carbon sequestration, studying faster growing variants of cyanobacteria, and developing new tools via synthetic biology.
Rewiring Photosynthetic Metabolism
Efforts have been made to overcome the intrinsic shortcomings of photosynthetic microorganisms by rewiring metabolism related to carbon fixation and photosynthesis. While efforts to improve RuBisCO have not been met with much success, current research has shifted focus towards rerouting metabolism to improve overall photosynthetic efficiency by focusing on key aspects of the Calvin-Benson cycle or the photosynthetic electron transport chain (PETC). One strategy used to harness the excess energy being lost by the PETC in cyanobacteria involved overexpressing the protein OmcS (Meng et al., 2021). This strategy coupled the excess electrons from the PETC to NADH production and was shown to increase intracellular ATP and NADH allowing for a fourfold improvement of D-lactate production in the cyanobacterium Synechococcus elongatus UTEX 2973 (hereon 2973) (Meng et al., 2021).
An inherent drawback of RuBisCO is its promiscuous nature, when RuBisCO undergoes oxygenase activity a costly side pathway known as photorespiration occurs where the oxygenase product is recycled back into usable metabolism consuming energy and losing CO2 in the process. As much as 30% of energy produced by photosynthesis has been observed to be lost through photorespiration in plants (Hagemann and Bauwe, 2016). Rewiring or preventing photorespiration represents a promising way to improve the overall efficiency of carbon fixation in photosynthetic organisms. Efforts to rewire photorespiration generally involve deleting energetically costly steps, circumventing steps where CO2 is lost, and rerouting metabolites towards central carbon metabolism (Hagemann and Bauwe, 2016). One of the more ambitious efforts to ameliorate the cost of photorespiration was the expression of a synthetic carbon capture pathway to serve as both a photorespiratory bypass and as a supplement to the Calvin-Benson cycle, this was shown to be a viable use of synthetic biology to counteract the costly natural photorespiration pathway (Shih et al., 2014).
It should be noted that photorespiration is not the sole pathway responsible for carbon inefficiencies, many metabolic processes include steps where CO2 is lost to the environment. An important way to engineer microorganisms for sustainability involves carbon conservation, focusing on rerouting metabolism to circumvent decarboxylation reactions (François et al., 2020). Of the more notable strategies is the non-oxidative glycolysis pathway (NOG) which has been shown to function in Escherichia coli and which can effectively conserve all carbon associated with sugar catabolism to acetyl-CoA (Bogorad et al., 2013). While carbon conservation is a powerful methodology for engineering metabolism, the field is still in its infancy and further work is required to evaluate the industrial viability of many carbon conservation strategies. Additionally, de novo carbon fixation pathways, which will be addressed later in this review, are currently being developed and may prove to be a better methodology for the development of sustainable production hosts.
Non-RuBisCO Carbon Fixation
In contrast to research centering on canonical CO2 fixation, investigations into de-novo CO2 fixation pathways have been explored and theorized in recent years as more efficient alternatives to traditional RuBisCO based CO2 assimilation. These pathways may provide advantages in chemical production hosts by offering insight into carboxylation reactions that could work in tandem with RuBisCO. The expression of formate dehydrogenase in the cyanobacterium, Anabaena sp. PCC 7120, was shown to successfully increase intracellular formate concentration, representing an alternative to the photo-reduction of CO2 and can act to supplement natural carbon fixation pathways (Ihara et al., 2013).
Many of these de novo CO2 fixation pathways have had limited success when installed into model organisms such as E. coli and yeast and it has yet to be shown if these pathways can function effectively in photosynthetic hosts. One pathway of note that has been shown to work in E. coli is the reductive glycine pathway, hereafter RGP (Tashiro et al., 2018). This pathway leverages the native glycine cleavage system in the reverse direction to combine one equivalent of CO2 with 5,10-methylenetetrahydrofolate that has been produced from formate to produce pyruvate. This method allows E. coli to directly assimilate CO2 into central metabolic pathways and is a more efficient method for CO2 fixation than traditional RuBisCO (Bar-Even et al., 2013). This inorganic carbon can then be leveraged for biochemical synthesis. Additional work has also recently shown that the expression of formate dehydrogenase confers further renewable characteristics to strains harboring the RGP by removing the need for glucose supplementation (Bang et al., 2020). Other notable CO2 fixation pathways include the crotonyl-CoA/ethylmalonyl-CoA/hydroxybutyryl-CoA (CETCH) cycle and the tartronyl-CoA (TaCo) pathways (Scheffen et al., 2021). While the RGP has proven to be a viable carbon fixation pathway that was shown to function in E. coli, the growth exhibited by this CO2 fixing E. coli is slower than its traditional heterotrophic phenotype (Tashiro et al., 2018; Bang et al., 2020). Advancements in modeling and metabolomics may allow for an increase in the creation of de-novo carbon fixation pathways that may prove to be both more efficient than traditional pathways and capable of functioning in a wider array of chemical production hosts.
Photomixotrophy
Another approach to increase chemical production capacity is to supplement CO2 with carbohydrates as an auxiliary carbon source for the Calvin-Benson cycle, thereby making the organism photomixotrophic. By re-engineering glucose catabolism to direct carbon flux into the Calvin-Benson cycle, more ribulose-1,5-bisphosphate can be supplied to RuBisCO, accelerating CO2 fixation. This ultimately results in faster growth and production of downstream targets, as well as a six-fold increase in titer once metabolism was rewired to accommodate photomixotrophy (Kanno et al., 2017). The addition of a heterotrophic mode also allows for CO2 fixation in darkness, resulting in a 24 h production period under natural diurnal conditions. Glucose can be readily obtained from the acid hydrolysis of agricultural waste products such as corn stover, in conjunction with other sugars: xylose, arabinose, and galacturonic acid (Mourtzinis et al., 2016). By installing catabolic pathways for these non-glucose sugars, these agricultural waste products can be used more efficiently. Xylose, the second most abundant sugar in corn stover lysate, has successfully been used to achieve photomixotrophic production of 2,3-butanediol in light and dark conditions with significant improvements in growth and product titer over the equivalent photoautotrophic organism (McEwen et al., 2016). A similar strategy has also recently been used to improve the production of 3-hydroxypropionic acid by 4.1 fold in cyanobacteria through the installation of a xylose photomixotrophic module along with other modifications to help assimilate the additional carbon source (Yao et al., 2022). It should also be noted that the cyanobacterium Synechocystis sp. PCC 6803 (hereafter 6803) natively possesses photomixotrophic machinery to assimilate glucose. A recent study was able to improve these photomixotrophic conditions through the installation of the NOG pathway along with targeted gene knockouts to increase the intracellular concentration of acetyl-CoA, thereby improving the growth phenotype of 6803 (Song et al., 2021).
Discovery of Fast-Growing Cyanobacteria
Most research done in this field focuses on using just a handful of species that have traditionally been used as model organisms to study the mechanics of photosynthesis. With increasing interest in using photosynthetic organisms for industrial production there have been efforts to uncover new species that are faster growing and more receptive to engineering. In the realm of cyanobacterial chemical production, species like 2973 and S. elongatus PCC 11802 (hereon 11802) have risen in popularity as they are faster growing than the traditional S. elongatus PCC 7942 (hereon 7942) and have been shown to produce higher titers of target chemical products under certain circumstances (Yu et al., 2015; Sengupta et al., 2020). Additionally, these fast-growing organisms are providing inspiration for how to better engineer existing model organisms. The fast-growing cyanobacterium 2973 has relatively little differences genetically when compared to the model cyanobacterium 7942 (Yu et al., 2015). However, a notable difference in 2973 is an increase in the expression levels of PSI, cytochrome b6f, and plastocyanin on a per cell basis which improves the downstream flux of electrons from PSII, which helps the faster growing cyanobacteria to better utilize photosynthetic energy (Ungerer et al., 2018). The discovery of new fast-growing cyanobacteria may enhance our understanding of photosynthesis and characterizing the differences between these new species with current model organisms.
Genome Engineering Tools
In cyanobacteria, traditional genomic modifications are a labor-intensive task and limiting in nature due to the polyploidal nature of these organisms and the need for antibiotic resistance markers (Griese et al., 2011). The current methodology for genomic integration involves constructing a plasmid with an antibiotic selection marker in a plasmid host such as E. coli. After introduction of this plasmid to cyanobacteria, several rounds of antibiotic screening are required to ensure complete genome segregation (Golden et al., 1987). This process generally limits the number of modifications that can be performed in a single strain due to the physiological constraints of expressing multiple different antibiotic resistance genes.
The overall task of metabolic engineering in cyanobacteria has been made dramatically more efficient thanks to the advent of CRISPR gene editing which allows for markerless edits (Behler et al., 2018). However, the protein Cas9 is toxic to a number of cyanobacteria species (Wendt et al., 2016). Researchers have recently uncovered other endonucleases that are similarly capable of CRISPR gene editing. The main endonuclease of interest is Cpf1 which, while similar to Cas9, is better tolerated by photosynthetic hosts (Ungerer and Pakrasi, 2016). As the body of research grows around Cpf1, more engineering strategies will be made available in the realm of photosynthetic chemical production and should offer a boon towards the viability of these organisms to begin replacing their non-CO2 fixing brethren in the realm of biochemical production (Bishé et al., 2019; Niu et al., 2019). Additionally, having the ability to perform markerless genomic modifications unlocks the potential to engineer these microorganisms far more ambitiously than what was previously possible.
Other work on CRISPR technologies in cyanobacteria includes the use of CRISPR inhibition (CRISPRi) by using dead Cas9 (dCas9) (Qi et al., 2013). While the endonuclease activity of the intact Cas9 protein seems to be toxic to these production hosts, dCas9 is able to function in the same manner in photosynthetic hosts as it is able to in heterotrophic hosts such as E. coli (Santos et al., 2021). While the use of dCas9 may not be as broadly useful as traditional CRISPR, dCas9 has been shown to be invaluable in certain chemical production applications where more traditional gene knockouts would otherwise be toxic.
Concluding Remarks
While many challenges remain and must be overcome to enable widespread adoption of photosynthetic chemical production hosts, the above studies suggest that there are myriad avenues of research that can get closer to this goal. The renewed interest in the field due to the ongoing climate crisis has spurred efforts to improve and adopt these microorganisms as a sustainable alternative for traditional petroleum-based synthesis. Many of the challenges in this field revolve around the intrinsic inefficiencies of carbon fixation and photosynthesis. While engineering RuBisCO remains an interesting target for improving carbon fixation, it has proven to be highly resistant to traditional engineering and decades of research would suggest that it is next to impossible to improve. Focusing on engineered carbon fixation pathways is a more promising route towards improving the carbon sequestration ability of cyanobacteria. Other research into improving the efficiency of photosynthesis by introducing alternative pathways downstream of the PETC for the production of chemical products is a prime example of how we can engineer these microbes to make full use of excess reducing potential from the PETC. The aforementioned approaches aim to enhance our understanding of the inefficiencies related to carbon fixation and photosynthesis while also representing some of the more novel approaches being undertaken by the field of synthetic biology. The discovery of new synthetic biology tools and investigation into faster growing cyanobacteria is also expanding the field of photosynthetic microbial research to make photosynthetic microorganisms a more viable alternative to petroleum based chemical production.
Of the discussed challenges for synthetic biology in cyanobacteria, improving the rate and efficiency of carbon fixation seems to be the most difficult, however, this task also holds the most promise. While RuBisCO is resistant to direct engineering strategies, adding additional carbon fixation modules can enhance the viability of cyanobacteria as a chemical production chassis. Further research into de novo carbon fixation pathways capable of operating in parallel to the Calvin-Benson cycle and RuBisCO holds great promise for circumventing the inefficiencies of carbon fixation in cyanobacteria. Multiple carbon fixation pathways operating in tandem could exponentially increase the amount of CO2 sequestered by cyanobacteria and greatly enhance growth and product formation. The process of carbon fixation is a highly regulated process, and this strategy will likely face further challenges before successful implementation. Overall, it is highly likely that the optimal route for improving the conversion of CO2 into valuable chemical commodities in cyanobacteria lies in exploiting multiple of the aforementioned strategies contained within this review (Figure 1).
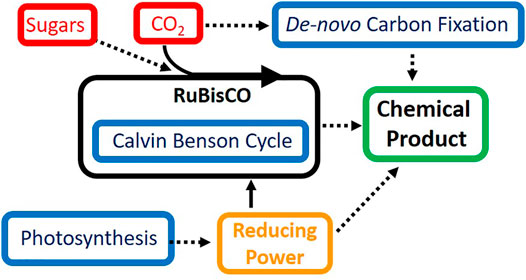
FIGURE 1. Strategies for improving efficiency of carbon fixation and photosynthesis in cyanobacteria. Shown in red are carbon sources for metabolism in cyanobacteria, in blue are pathways of interest to synthetic biology and metabolic engineering, solid lines represent native pathways and dashed lines represent pathways of interest for improvement using synthetic biology.
As new synthetic biology tools become available for cyanobacteria, high throughput screening will allow for rapid progress to be made within this field. The advent of CRISPR technology has had profound effects on research in a wide variety of fields but is relatively new to cyanobacteria. Additionally, faster growing species of cyanobacteria are rising in popularity and more have yet to be discovered. While these newly discovered cyanobacteria are efficient production hosts in their own right, they also inform the field on future targets for modification. Understanding the inherent differences among these organisms is vital to improving our understanding of carbon fixation and photosynthesis. It stands to reason that research into discovering more of these faster growing species, as well as studying the known cyanobacterial variants will provide insight and guidance for future work in this field. While improving photosynthetic production hosts has been historically difficult, the studies described in this work point to a promising future.
Author Contributions
TT, JG, JP, and SA wrote the manuscript.
Funding
This work was supported by the National Science Foundation (CBET-1902014).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Bang, J., Hwang, C. H., Ahn, J. H., Lee, J. A., and Lee, S. Y. (2020). Escherichia coli Is Engineered to Grow on CO2 and Formic Acid. Nat. Microbiol. 5, 1459–1463. doi:10.1038/s41564-020-00793-9
Bar-Even, A., Noor, E., Flamholz, A., and Milo, R. (2013). Design and Analysis of Metabolic Pathways Supporting Formatotrophic Growth for Electricity-dependent Cultivation of Microbes. Biochim. Biophys. Acta Bioenerg. 1827, 1039–1047. doi:10.1016/j.bbabio.2012.10.013
Behler, J., Vijay, D., Hess, W. R., and Akhtar, M. K. (2018). CRISPR-based Technologies for Metabolic Engineering in Cyanobacteria. Trends Biotechnol. 36, 996–1010. doi:10.1016/j.tibtech.2018.05.011
Bishé, B., Taton, A., and Golden, J. W. (2019). Modification of RSF1010-Based Broad-Host-Range Plasmids for Improved Conjugation and Cyanobacterial Bioprospecting. iScience 20, 216–228. doi:10.1016/j.isci.2019.09.002
Bogorad, I. W., Lin, T.-S., and Liao, J. C. (2013). Synthetic Non-oxidative Glycolysis Enables Complete Carbon Conservation. Nature 502, 693–697. doi:10.1038/nature12575
Borden, J. S., and Savage, D. F. (2021). New Discoveries Expand Possibilities for Carboxysome Engineering. Curr. Opin. Microbiol. 61, 58–66. doi:10.1016/j.mib.2021.03.002
Erb, T. J., and Zarzycki, J. (2018). A Short History of RubisCO: the Rise and Fall (?) of Nature's Predominant CO2 Fixing Enzyme. Curr. Opin. Biotechnol. 49, 100–107. doi:10.1016/j.copbio.2017.07.017
Flamholz, A. I., Prywes, N., Moran, U., Davidi, D., Bar-On, Y. M., Oltrogge, L. M., et al. (2019). Revisiting Trade-Offs between Rubisco Kinetic Parameters. Biochemistry 58, 3365–3376. doi:10.1021/acs.biochem.9b00237
François, J. M., Lachaux, C., and Morin, N. (2020). Synthetic Biology Applied to Carbon Conservative and Carbon Dioxide Recycling Pathways. Front. Bioeng. Biotechnol. 7, 1–16. doi:10.3389/fbioe.2019.00446
Golden, S. S., Brusslan, J., and Haselkorn, R. (1987). Genetic Engineering of the Cyanobacterial Chromosome. Methods Enzymol. 153, 215–231. doi:10.1016/0076-6879(87)53055-5
Griese, M., Lange, C., and Soppa, J. (2011). Ploidy in Cyanobacteria. FEMS Microbiol. Lett. 323, 124–131. doi:10.1111/j.1574-6968.2011.02368.x
Hagemann, M., and Bauwe, H. (2016). Photorespiration and the Potential to Improve Photosynthesis. Curr. Opin. Chem. Biol. 35, 109–116. doi:10.1016/j.cbpa.2016.09.014
Ihara, M., Kawano, Y., Urano, M., and Okabe, A. (2013). Light Driven CO2 Fixation by Using Cyanobacterial Photosystem I and NADPH-dependent Formate Dehydrogenase. PLoS ONE 8, e71581–8. doi:10.1371/journal.pone.0071581
Kanno, M., Carroll, A. L., and Atsumi, S. (2017). Global Metabolic Rewiring for Improved CO2 Fixation and Chemical Production in Cyanobacteria. Nat. Commun. 8, 1–11. doi:10.1038/ncomms14724
Kerfeld, C. A., and Melnicki, M. R. (2016). Assembly, Function and Evolution of Cyanobacterial Carboxysomes. Curr. Opin. Plant Biol. 31, 66–75. doi:10.1016/j.pbi.2016.03.009
Levi, P. G., and Cullen, J. M. (2018). Mapping Global Flows of Chemicals: From Fossil Fuel Feedstocks to Chemical Products. Environ. Sci. Technol. 52, 1725–1734. doi:10.1021/acs.est.7b04573
McEwen, J. T., Kanno, M., and Atsumi, S. (2016). 2,3 Butanediol Production in an Obligate Photoautotrophic Cyanobacterium in Dark Conditions via Diverse Sugar Consumption. Metab. Eng. 36, 28–36. doi:10.1016/j.ymben.2016.03.004
Meng, H., Zhang, W., Zhu, H., Yang, F., Zhang, Y., Zhou, J., et al. (2021). Over-expression of an Electron Transport Protein OmcS Provides Sufficient NADH for D-Lactate Production in Cyanobacterium. Biotechnol. Biofuels 14, 1–16. doi:10.1186/s13068-021-01956-4
Mourtzinis, S., Cantrell, K. B., Arriaga, F. J., Balkcom, K. S., Novak, J. M., Frederick, J. R., et al. (2016). Carbohydrate and Nutrient Composition of Corn stover from Three southeastern USA Locations. Biomass Bioenergy 85, 153–158. doi:10.1016/j.biombioe.2015.11.031
Niu, T.-C., Lin, G.-M., Xie, L.-R., Wang, Z.-Q., Xing, W.-Y., Zhang, J.-Y., et al. (2019). Expanding the Potential of CRISPR-Cpf1-Based Genome Editing Technology in the CyanobacteriumAnabaenaPCC 7120. ACS Synth. Biol. 8, 170–180. doi:10.1021/acssynbio.8b00437
Qi, L. S., Larson, M. H., Gilbert, L. A., Doudna, J. A., Weissman, J. S., Arkin, A. P., et al. (2013). Repurposing CRISPR as an RNA-Guided Platform for Sequence-specific Control of Gene Expression. Cell 152, 1173–1183. doi:10.1016/j.cell.2013.02.022
Santos, M., Pacheco, C. C., Yao, L., Hudson, E. P., and Tamagnini, P. (2021). CRISPRi as a Tool to Repress Multiple Copies of Extracellular Polymeric Substances (EPS)-Related Genes in the Cyanobacterium Synechocystis Sp. PCC 6803. Life 11, 1198. doi:10.3390/life11111198
Scheffen, M., Marchal, D. G., Beneyton, T., Schuller, S. K., Klose, M., Diehl, C., et al. (2021). A New-To-Nature Carboxylation Module to Improve Natural and Synthetic CO2 Fixation. Nat. Catal. 4, 105–115. doi:10.1038/s41929-020-00557-y
Sengupta, S., Jaiswal, D., Sengupta, A., Shah, S., Gadagkar, S., and Wangikar, P. P. (2020). Metabolic Engineering of a Fast-Growing Cyanobacterium Synechococcus Elongatus PCC 11801 for Photoautotrophic Production of Succinic Acid. Biotechnol. Biofuels 13, 1–18. doi:10.1186/s13068-020-01727-7
Shih, P. M., Zarzycki, J., Niyogi, K. K., and Kerfeld, C. A. (2014). Introduction of a Synthetic CO2-fixing Photorespiratory Bypass into a Cyanobacterium. J. Biol. Chem. 289, 9493–9500. doi:10.1074/jbc.C113.543132
Shih, P. M., Occhialini, A., Cameron, J. C., Andralojc, P. J., Parry, M. A. J., and Kerfeld, C. A. (2016). Biochemical Characterization of Predicted Precambrian RuBisCO. Nat. Commun. 7, 1–11. doi:10.1038/ncomms10382
Solomon, S., Plattner, G.-K., Knutti, R., and Friedlingstein, P. (2009). Irreversible Climate Change Due to Carbon Dioxide Emissions. PNAS 106, 1704–1709. doi:10.1073/pnas.0812721106
Song, X., Diao, J., Yao, J., Cui, J., Sun, T., Chen, L., et al. (2021). Engineering a Central Carbon Metabolism Pathway to Increase the Intracellular Acetyl-CoA Pool in Synechocystis Sp. PCC 6803 Grown under Photomixotrophic Conditions. ACS Synth. Biol. 10, 836–846. doi:10.1021/acssynbio.0c00629
Sproles, A. E., Fields, F. J., Smalley, T. N., Le, C. H., Badary, A., and Mayfield, S. P. (2021). Recent Advancements in the Genetic Engineering of Microalgae. Algal Res. 53, 102158. doi:10.1016/j.algal.2020.102158
Stephens, S., Mahadevan, R., and Allen, D. G. (2021). Engineering Photosynthetic Bioprocesses for Sustainable Chemical Production: A Review. Front. Bioeng. Biotechnol. 8, 1–15. doi:10.3389/fbioe.2020.610723
Tashiro, Y., Hirano, S., Matson, M. M., Atsumi, S., and Kondo, A. (2018). Electrical-biological Hybrid System for CO2 Reduction. Metab. Eng. 47, 211–218. doi:10.1016/j.ymben.2018.03.015
Ungerer, J., and Pakrasi, H. B. (2016). Cpf1 Is A Versatile Tool for CRISPR Genome Editing across Diverse Species of Cyanobacteria. Sci. Rep. 6, 1–9. doi:10.1038/srep39681
Ungerer, J., Lin, P.-C., Chen, H.-Y., and Pakrasi, H. B. (2018). Adjustments to Photosystem Stoichiometry and Electron Transfer Proteins Are Key to the Remarkably Fast Growth of the Cyanobacterium Synechococcus Elongatus UTEX 2973. MBio 9, e02327. doi:10.1128/mBio.02327-17
Wendt, K. E., Ungerer, J., Cobb, R. E., Zhao, H., and Pakrasi, H. B. (2016). CRISPR/Cas9 Mediated Targeted Mutagenesis of the Fast Growing Cyanobacterium Synechococcus Elongatus UTEX 2973. Microb. Cel Fact. 15, 1–8. doi:10.1186/s12934-016-0514-7
Yao, J., Wang, J., Ju, Y., Dong, Z., Song, X., Chen, L., et al. (2022). Engineering a Xylose-Utilizing Synechococcus Elongatus UTEX 2973 Chassis for 3-Hydroxypropionic Acid Biosynthesis under Photomixotrophic Conditions. ACS Synth. Biol. 11, 678–688. doi:10.1021/acssynbio.1c00364
Keywords: CO2 fixation, cyanobacteria, photosynthesis, RuBisCO, CRISPR
Citation: Treece TR, Gonzales JN, Pressley JR and Atsumi S (2022) Synthetic Biology Approaches for Improving Chemical Production in Cyanobacteria. Front. Bioeng. Biotechnol. 10:869195. doi: 10.3389/fbioe.2022.869195
Received: 04 February 2022; Accepted: 25 February 2022;
Published: 11 March 2022.
Edited by:
M. Kalim Akhtar, United Arab Emirates University, United Arab EmiratesReviewed by:
Weiwen Zhang, Tianjin University, ChinaCopyright © 2022 Treece, Gonzales, Pressley and Atsumi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Shota Atsumi, c2F0c3VtaUB1Y2RhdmlzLmVkdQ==
 Tanner R. Treece1
Tanner R. Treece1 Joseph R. Pressley
Joseph R. Pressley Shota Atsumi
Shota Atsumi