Abstract
Graphite oxide (GO) has been used for the immobilization of several classes of enzymes, exhibiting very interesting properties as an immobilization matrix. However, the effect the nanomaterial has on the enzyme cannot be predicted. Herein, the effect GO has on the catalytic behavior of several (S)-selective amine transaminases [(S)-ATAs] has been investigated. These enzymes were the focus of this work as they are homodimers with pyridoxal 5′-phosphate in their active site, significantly more complex systems than other enzymes previously studied. Addition of GO (up to 0.1 mg/ml) in the reaction medium leads to activation (up to 50% improved activity) for most enzymes studied, while they maintain their temperature profile (they perform better between 40 and 45°C) and their stability. However, the effect is not universal and there are enzymes that are negatively influenced by the presence of the nanomaterial. More profound is the effect on the (S)-ATA from Chromobacterium violaceum which loses almost 50% of its activity in the presence of 0.1 mg/ml GO, while the stability was significantly decreased, losing its activity after 2 h incubation at 40°C, in the presence of 25 μg/ml GO. This negative effect seems to rise from minor secondary structure alterations; namely, a loss of α-helices and subsequent increase in random coil (∼3% in the presence of 25 μg/ml GO). We hypothesize that the effect the GO has on (S)-ATAs is correlated to the surface chemistry of the enzymes; the less negatively-charged enzymes are deactivated from the interaction with GO. This insight will aid the rationalization of ATA immobilization onto carbon-based nanomaterials.
Introduction
Αmine transaminases (ATAs) are pyridoxal 5′-phosphate (PLP)-dependent enzymes that attracted significant industrial interest over the last years (Fuchs et al., 2015; Kelly et al., 2018; Kelly et al., 2020). Their excellent enantioselectivity places them among the best routes towards the synthesis of optically pure amines (Savile et al., 2010; Pavlidis et al., 2016; Ferandi and Monti, 2018). For their industrial application, ATAs require operational and thermal stability. Enzyme immobilization has been used as a stabilization technique, while enabling enzyme’s reuse, minimizing thus the cost of the developed bioprocess.
Several materials have been used as immobilization matrices. Among them, nanomaterials are of significant interest for this application (Pavlidis et al., 2014; Liu and Dong 2020; Zahirinejad et al., 2021), due to their high surface area which can lead to high enzyme loading (Min et al., 2012), while their surface can be modified (Kouloumpis et al., 2017; Yüce and Kurt, 2017; Kouloumpis et al., 2018). Among the nanomaterials, carbon-based nanomaterials (CNBs) have attracted significant interest due to their unique electrical mechanical and thermal properties, as well as their biocompatibility (Patila et al., 2017; Potsi et al., 2019; Patila et al., 2020). Graphite Oxide (GO) is a 2D hydrophilic derivative of graphene that has been used as an immobilization matrix, due to its high surface area and the many functional groups present on its surface (i.e. epoxides, hydroxyls, carbonyls) (Bourlinos et al., 2003; Adeel et al., 2018; Tsirka et al., 2018). Moreover, reactivity towards the production of ketones, which are substrates of the ATAs, has been documented, although not being compatible with enzymatic processes (high temperature, high catalyst loading up to 200% w/w) (Dreyer et al., 2010; Navalon et al., 2017).
In the non-covalent immobilization, the interaction of the immobilization support with the protein is of crucial importance. Upon interaction, enzymes’ activity and structure can be altered (Jafarian et al., 2018; Di Giosia et al., 2020), due to surface chemistry, charge, and hydrophobicity of the matrix. So far, despite the immobilization of several enzyme classes onto such nanomaterials (Min et al., 2012; Pavlidis et al., 2012; Patila et al., 2016), we still cannot rationalize the process and predict the catalytic behavior of the biocatalysts. Especially for ATAs, which are active as homodimers and require proper PLP incorporation in the active site, the interactions with 2D materials can be quite intervening and interfere with the dimer formation. To this day, there are scarce or no data about the effect layered nanomaterials have on the activity and on the structure of these enzymes.
Aiming to understand the structure-function relationships of ATAs when interacting with 2D materials, we studied the effect GO has on the activity, stability, and secondary structure of 7 (S)-selective ATAs, expressed and purified in our laboratory; namely, three variants from Ruegeria sp. TM1040 (Rst-1 to Rst-3), which we developed for acceptance of bulkier substrates (Pavlidis et al., 2016; Weiß et al., 2016), an ATA from Sagittula stellata E-37 (Sst) with high identity to Rst-1 (∼90%), and the widely studied ATAs from Vibrio fluvialis (Vfl), Chromobacterium violaceum (Cvi), and Ruegeria pomeroyi (Rpo). The selected enzymes share identity as low as 30%, to cover a broad range of (S)-ATAs. The specific activity of ATAs in the absence and in the presence of GO (up to 0.1 mg/ml) was determined in the reaction of (S)-1-phenylethylamine with pyruvate, while the same reaction was used to monitor their stability and temperature profile. The secondary structure and the melting temperature of the (S)-ATAs was determined via circular dichroism under similar conditions. These results help us to understand the effect of the interaction of GO with fold class I PLP-dependent enzymes, and hopefully they can aid the design of robust and more active immobilized biocatalysts.
Materials and Methods
Sodium pyruvate (≥99%) was purchased from Serva. (S)-α-phenylethylamine [(S)-PEA, 98%], acetophenone (ACP, 99%), and pyridoxal 5′-phosphate (PLP) monohydrate (98%) were purchased from Alfa Aesar. Dimethyl sulfoxide (DMSO, 99.9%) was purchased from Fischer Chemical. Graphite oxide (GO) was synthesized as described in Paragraph 2.2. Enzymes were expressed as described in Paragraph 2.2 and purified as described in Paragraph 2.3. All other reagents and solvents were of analytical grade or superior.
Synthetic Procedure of Graphite Oxide
For the synthesis of GO we used a modified Hummers method (Hummers and Offeman, 1958). Via this method, graphite oxide is generated from graphite using strong acids. In detail, 0.5 g of graphite and 0.5 g sodium nitrate were added to a beaker with 23 ml sulfuric acid (96%), and the mixture was stirred for 15 min. The reaction beaker was placed for cooling in an ice bath. After the end of 15 min, 3.2 g of potassium permanganate were added in small portions and the stirring was continued for 1 h more, and at the same time the mixture was heated to 28°C. The temperature was further increased to 95°C and an addition of 40 ml H2O was followed. The reaction mixture was stirred for 30 min before further addition of 100 ml H2O. As the final step, hydrogen peroxide (30 wt%) was slowly added. The mixture was purified by repeated centrifugation and washings several times with H2O. The obtained sample of graphite oxide was dried at room temperature and it was characterized with X-ray diffraction (Supplementary Figure S1) and X-ray photoelectron spectroscopy (Supplementary Figure S2).
Bacterial Strains and Growth Conditions
All ATAs were overexpressed in Escherichia coli BL21(DE3) cells, transformed with the respective plasmids. The amino acid sequences of the respective enzymes are provided in the Supplementary Material. Briefly, 5 ml of an overnight culture of a single colony of E. coli BL21(DE3) with the construct of interest was used to inoculate 500 ml of Lysogeny Broth (LB) medium containing the respective antibiotic (50 μg/ml kanamycin for Vfl and Cvi and 100 μg/ml ampicillin for the rest). The culture was incubated at 37°C with orbital shaking at 300 rpm until the OD at 600 nm (OD600) reached a value of 0.6–0.8. At that point, the expression of the genes of interest was induced with 0.2 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) and left for 16–18 h at 20°C and 300 rpm for the expression. At that point, the biomass was harvested by centrifugation at 6,000 x g for 15 min at 4°C. Supernatants were discarded and the pellet was resuspended in 7–9 ml of phosphate buffer (50 mM, pH 7.5) with 300 mM NaCl, 0.1 mM PLP and 15 mM imidazole. Cells were lysed by sonication using an Ultrasonic sonicator LABSONIC P, at 30% amplitude, 50% pulse, for five cycles of 30 s sonication, following 30 s of resting in ice. The suspension was then centrifuged at 6,000 x g for 15 min at 4°C and the supernatant containing the soluble proteins including the His-tagged ATAs was collected, filtered with 0.22 μm PVDF filters, and used for purification.
Protein Purification
Purification of ATAs was performed by affinity chromatography, using ÄKTA Start Protein Purification System controlled from the UNICORN 7.0 software (GE Healthcare Life Sciences) with an Ni-NTA HisTrap HP 5 ml column. Equilibration of the system was performed using Buffer A (phosphate buffer, 50 mM, pH 7.5 with 300 mM NaCl and 0.1 mM PLP) and Buffer B (Buffer A+ 300 mM imidazole) in a ratio of 95% buffer A and 5% buffer B with 3 ml/min flow. The column was loaded with the sample with 1.5 ml/min flow. Washes were performed using four volumes of 90% buffer A and 10% buffer B while the flow was restored to 3 ml/min. Elution was performed with 100% Buffer B and fractions were collected. Afterwards concentration and desalting of the eluted protein was performed by using an Amicon centrifugal filter with a molecular weight cut off of 30 kDa and Buffer C (phosphate buffer, 50 mM, pH 7.5 with 0.1 mM PLP). The desalted protein was aliquoted and flash frozen with liquid nitrogen, while SDS-PAGE and protein quantification (via Bradford assay, Bradford, 1976) were performed to determine the purity and concentration of the produced ATA. Samples were stored at −20°C until use.
ATAs Activity Assay
For the specific activity measurements, an established photometric assay was employed (Schätzle et al., 2009) using a Multiskan Sky 96-well plate reader (Thermo Fisher). Briefly, 200 μL of a reaction mixture, containing 2 mM sodium pyruvate, 1 mM (S)-PEA, in HEPES buffer (50 mM, pH 8.0) were incubated with free enzyme (final concentrations: Rsp-1, 22.3 μg/ml; Rsp-2, 150 μg/ml; Rsp-3, 91 μg/ml; Sst, 37.4 μg/ml; Vfl, 17.6 μg/ml; Rpo, 39.2 μg/ml; Cvi, 20.8 μg/ml) at 30°C. The increase in absorbance due to ACP production (ε = 14,690 M−1 cm−1) was monitored for 5 min, with 15 s time interval, at 245 nm using Greiner UV-Star 96-well plates. One unit of activity was defined as the amount of enzyme that was required to produce 1 μmol of ACP per minute at the assayed conditions. For the temperature profile measurements, all conditions were the same; however, the reaction medium was preheated before measurements at the temperature of interest for each experiment. All measurements were at least in triplicate. Herein we would like to state that the experiments were performed at pH 8.0 as at this pH all studied (S)-ATAs exhibit high stability; nevertheless, some ATAs perform better at more alkaline pH (data not shown).
Operational Stability Measurements
The operational stability experiments were performed with the same conditions as described in Paragraph 2.5. The enzymes were incubated at 40°C in HEPES buffer (50 mM, pH 8.0) with sodium pyruvate (2 mM final concentration), at 400–600 rpm. For the samples with GO, 25 μg/ml of GO were present in the solution. Samples were removed at specific time intervals, and they were incubated for 10 s on ice and for 5 min at 30°C in the plate reader to reach the temperature of the activity assay. Reactions were initiated with the addition of (S)-PEA (1 mM) and the specific activity was measured as described in Paragraph 2.5.
Circular Dichroism Experiments
All circular dichroism (CD) measurements were performed with A Jasco J-810 spectropolarimeter, equipped with a 150 W xenon lamp and a Peltier temperature controller. In all measurements, a 1 cm pathlength quartz cuvette was used, while the ATAs were diluted in phosphate buffer (5 mM, pH 7.5), at concentrations that allow the measurement (see Paragraph 2.7.1 and 2.7.2 for exact concentrations). Wherever GO was used, the experiments was conducted under the same conditions, with the only alteration being the addition of 25 μg/ml GO (starting from a stock of 1 mg/ml in the respective buffer).
Secondary Structure Measurements
The spectra for the secondary structure estimation were recorded in the far-UV region (200–250 nm) with a response time of 8 s and scan speed of 50 nm min−1, at 30°C. The concentrations used for the enzymes were: Rsp-1, 22.3 μg/ml; Rsp-2, 22.5 μg/ml, Rsp-3, 37 μg/ml; Sst, 28.6 μg/ml; Vfl, 27.9 μg/ml; Rpo, 39.2 μg/ml; Cvi, 27.8 μg/ml. Measurements below 200 nm were not possible due to high background absorbance. Three scans were accumulated and averaged for each spectrum. A spectrum of the buffer was measured in the same way, and it was subtracted afterwards in the analysis from the spectra of the enzymes. All ellipticity (mdeg) data collected were converted into mean molecular ellipticity data using the following formula:where MME: mean molecular ellipticity per amino acid (deg cm2 dmol−1), CD: circular dichroism (ellipticity, deg), l: optical pathlength (cm), c: molar protein concentration (M) and n: number of protein residues. Lastly, analysis was conducted using the online platform Dichroweb (Miles et al., 2021) using neural network algorithm K2D (Andrade et al., 1993).
Melting Temperature Measurements
Melting temperature (Tm) of ATAs was determined by measuring the ellipticity change at 222 nm which corresponds to α-helix content with increase of temperature. The final enzymes concentrations used were: Rsp-1, 22.3 μg/ml, Rsp-2, 15.0 μg/ml; Cvi, 20.8 μg/ml. Temperature was increased at a rate of 2°C/ min and measurements were acquired every 0.1°C. In order to calculate the Tm, the obtained experimental measurements were fitted to a 5-parameter sigmoidal curve.
Bioinformatic Analysis of Surface
The bioinformatic analysis was performed with YASARA Structure software (v.21.8.27). The structures used were: for the Rsp variants, the PDB ID: 3fcr (crystal structure of the wild type), for Vfl the PDB ID: 3nui, for Rpo the PDB ID: 3hmu, and for Cvi the PDB ID: 6snu (crystal structure of the mutant W60C). Although the crystal structures used for Rsp and Cvi are not exactly the same as the enzymes used in the present work, they are selected for this bioinformatic analysis, as (1) we would like to avoid using models and (2) the mutations are not on the surface and thus their influence in the analysis is expected to be negligible. For each structure the water molecules and any ligands co-crystallized, apart from PLP, were removed, the structures were cleaned from errors and the dimer was prepared, in the cases that the structure was provided in a different oligomerization state. The solvent accessible surface area of the dimer was then calculated, as well as the electrostatic potential, using the Poisson-Boltzmann Solver. The force field YASARA2 was used in pH 7.5 to calculate the total charge of the dimers.
Results
Effect of GO on Catalytic Activity and Operational Stability of ATAs
Initially, we studied the effect the presence of GO (0—100 μg/ml) has on the activity of 7 (S)-ATAs using the established photometric protocol of Schätzle et al. (2009), with (S)-PEA and pyruvate as substrates, at 30°C. As seen in Figure 1A, the presence of GO does not have a universal effect in the activity of the investigated ATAs. Specifically, the presence of GO leads to an increase of the enzymatic activity of all Rsp variants up to 40%. The fact that all variants exhibit the same effect has to do with the fact that the mutations are located in the active site and not in the surface, thus no significant changes on the interaction with the GO are expected, rather than intrinsic behavioral changes (such as stability of the variants). The increase in activity does not seem to be linearly correlated with the concentration of the GO. For instance, in the case of Rsp-1 the lower concentration of GO used (25 μg/ml) was the optimal for the activity. The activity of Sst and Rpo does not alter significantly, while the activity of Vfl and, more dramatically, of Cvi is significantly decreased in the presence of GO, in a dose-dependent way; in the case of Cvi, even 25 μg/ml of GO led to loss of 30% of its activity, while 100 μg/ml leads to the loss of half of the activity.
FIGURE 1
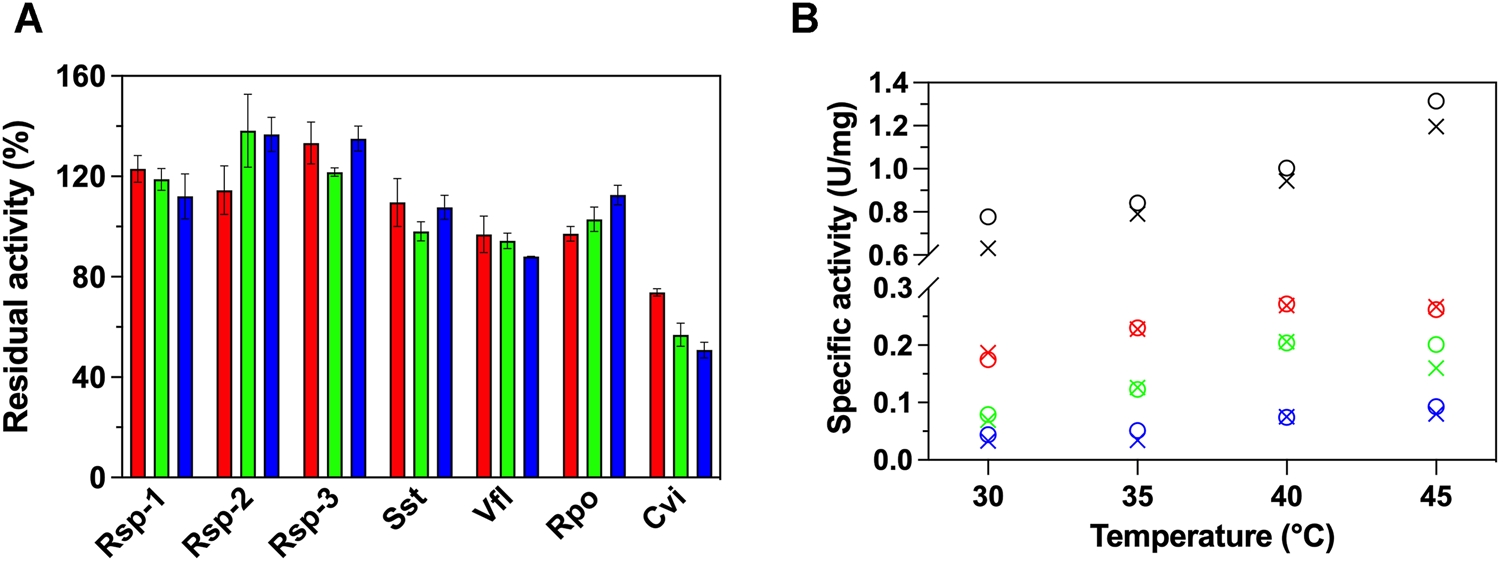
(A) Relative activity (%) of 7 ATAs in the presence of variable GO concentrations (25 μg/ml, red; 50 μg/ml, green; 100 μg/ml, blue) in HEPES buffer (50 mM, pH 7.5) at 30°C and (B) temperature profile of Rsp-1 (black), Rsp-2 (green), Rsp-3 (blue), and Cvi (red), in the presence (25 μg/ml, circle) or in the absence of GO (X mark), in HEPES buffer (50 mM, pH 7.5), at 30–45°C. All activity determinations were performed in the transamination reaction of (S)-PEA (1 mM) with pyruvate (2 mM), as described in Paragraph 2.5. Standard deviation is omitted for clarity; in all cases, it was less than 8%.
To further investigate the effect of the GO on the activity of (S)-ATAs, we determined the temperature profile of Rsp variants and Cvi, as the enzymes that were influenced the most from the presence of GO. Our aim was to see if the effect is maintained in higher temperatures. The profile is measured up to 45°C, as at higher temperatures all enzymes investigated lose their activity quickly (1–3 min) and the assay cannot be completed. As seen in Figure 1B, for all ATAs studied, the temperature profile is not significantly affected by the presence of 25 μg/ml GO; all ATAs exhibited an optimal at 40 or 45°C. Only in the case of Rsp-1, the increased activity observed in the presence of GO is maintained at all temperatures. For all other enzymes, the effect the GO has on activity is lost with the increase of temperature, something that hints that the effect of GO on ATAs relies on weak interactions, which can be easily disrupted by an increase in temperature.
The same ΑΤΑs were subjected in stability measurements at 40°C, with (25 μg/ml) and without GO in a time frame of 24 h. As shown in Figure 2, the GO influence significantly the stability of Cvi, and in a lesser way extend the Rsp variants. Cvi is inactivated quite fast at 40°C, as in 1 h it already loses ∼60% of its activity. The negative effect of GO is also reflected in the stability of the enzyme, which loses ∼95% of its activity at the same time. In the case of the Rsp variants, the positive effect of GO is projected in the stability of the enzymes in a lesser extent and the intrinsic stability is the most important parameter influencing the stability. Only in the case of Rsp-1 GO induces a higher hyperactivation of this ATA by 80% in the first hour of incubation, but the GO does not provide a protective effect; this variant quickly loses its activity after the first hour, and after 24 h of incubation it maintains only 17% of its activity, while the one without GO maintains 11% (data not shown). Rsp-2, as described in literature (Pavlidis et al., 2016), is more stable than Rsp-1 and its hyperactivation leads to an increase of activity by ∼40% after 3 h, which is maintained after 24 h, while the enzyme without GO maintains 130% of the activity before incubation. Rsp-3, as it is based on the Rsp-2, is also stable, but it does not exhibit hyperactivation and it retains ∼55% of its activity after 24 h, irrespective of the presence of GO. These effects on the stability of Rsp variants are attributed to the mutations introduced between variants and not to the interaction with the GO, as the differences of the stability with and without GO are low in the long run (<10% after 24 h).
FIGURE 2
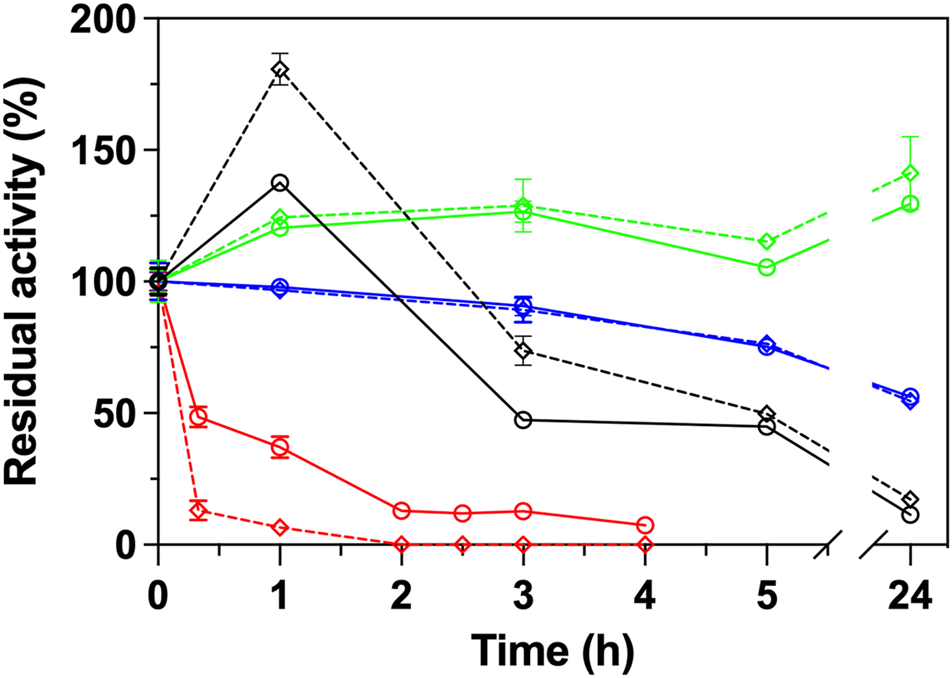
Operational stability of ATAs (Rsp-1, black; Rsp-2, green; Rsp-3, blue; Cvi, red) in the presence (25 μg/ml, dashed line) or in the absence of GO (continuous line) in HEPES buffer (50 mM, pH 8.0) at 40°C. All operational stability experiments were performed in the transamination reaction of (S)-PEA (1 mM) with pyruvate (2 mM) at 30°C, as described in Paragraph 2.6.
Effect of GO on Secondary Structure of ATAs
In order to understand the effect of the GO on the activity and stability of the ATAs, we investigated the potential alterations on the secondary structure elements of the enzymes, in the presence of 25 μg/ml and 50 μg/ml GO, using circular dichroism. The algorithm K2D (Andrade et al., 1993) was selected after a prescreening of several algorithms, as it was the one that provided results closer to the known crystal structures (data not shown) and could be used with the restricted spectrum we had, due to the high absorbance of the sample (no measurement possible <200 nm). As seen in Table 1, the presence of 25 μg/ml does not influence significantly the secondary structure of most enzymes. In the case of Cvi and Vfl, the two enzymes that their activity is negatively influenced, the presence of GO leads to increase of random coil, something that indicates unfolding, however the differences are small and in the frame of statistical error. In the case of Cvi, which activity is influenced in a greater extent from the presence of GO, the α-helical content is decreased, which is related with the activity of the enzymes (De Diego et al., 2005; Wu et al., 2014), and thus the structural results seem to correlate with the negative influence on activity. Rsp-1 and Rsp-2 lose a significant amount of the β-sheets (about 3–4% of total structure), which in the case of Rsp-1 is reflected in increase of the random coil content. However, the content of proteins in β-sheets is not related to the activity and thus it is not influencing negatively the activity. The results at 50 μg/ml (data provided in SI) were similar.
TABLE 1
| % α-helix | % β-sheets | % Random coil | ||||
|---|---|---|---|---|---|---|
| Enzyme | w/o GO | With GO | w/o GO | With GO | w/o GO | With GO |
| Rsp-1 | 29 ± 3 | 29 ± 1 | 19 ± 3 | 16 ± 1 | 52 ± 6 | 55 ± 0 |
| Rsp-2 | 29 ± 3 | 31 ± 1 | 20 ± 4 | 16 ± 2 | 51 ± 1 | 53 ± 2 |
| Rsp-3 | 31 ± 1 | 30 ± 1 | 17 ± 1 | 18 ± 2 | 53 ± 1 | 53 ± 2 |
| Sst | 33 ± 1 | 34 ± 1 | 18 ± 0 | 18 ± 1 | 49 ± 1 | 49 ± 2 |
| Rpo | 31 ± 1 | 30 ± 0 | 12 ± 1 | 13 ± 0 | 58 ± 1 | 57 ± 1 |
| Vfl | 31 ± 1 | 30 ± 3 | 18 ± 1 | 15 ± 1 | 51 ± 1 | 55 ± 3 |
| Cvi | 35 ± 3 | 32 ± 2 | 16 ± 2 | 16 ± 1 | 49 ± 3 | 52 ± 2 |
Secondary structure analysis of (S)-selective ATAs in the presence (25 μg/ml) and in the absence of GO in phosphate buffer (5 mM, pH 7.5), at 30°C. Enzyme concentration varied from 22.3 to 39.2 μg/ml. All spectra were measured at least duplicate with three scans per sample, as described in Paragraph 2.7.1.
As mentioned before, the content of α-helix is considered correlated to the enzymatic activity (De Diego et al., 2005; Wu et al., 2014). Thus, to correlate the stability observed for the (S)-ATAs with structural data, we calculated the melting temperature of the enzymes, based on the analysis of the negative peak of ellipticity at 222 nm, which is attributed to α-helix. The increase in temperature results in a sigmoidal curve, where the melting temperature can be calculated (see Supplementary Figure S3). We investigated Rsp-1, Rsp-2, and Cvi, as they provided the most interesting results in stability experiments. Rsp-1 exhibited significant hyperactivation and statistically important improvement of activity at the first hours of incubation. Rsp-2 is also exhibiting hyperactivation, but more importantly has an intrinsic better stability compared to Rsp-1. The activity of Cvi was negatively affected from the presence of GO, both in activity and stability measurements. As seen in Table 2, the melting temperature of Rsp-1 and Rsp-2 were not significantly affected by the presence of GO, as the operational stability experiments indicated. However, it can be observed that the Rsp-2 has an increase of about 2°C in the melting temperature compared to the less stable variant, Rsp-1. This difference is observed regardless of the presence of GO. The most interesting finding was the significant decrease of the melting temperature of Cvi in the presence of GO, as a decrease of more than 10°C is observed. These results are in line with the significant decrease in activity and stability of this enzyme in the presence of the nanomaterial. The melting temperature calculated with this technique is significantly higher compared to the temperature that the enzyme loses its activity, but we need to highlight here that the Tm calculated from circular dichroism refers to the temperature where 50% of the enzymes lose their structure (as calculated from the α-helical content), while Tm calculated from activity measurements relies on the loss of activity. In several cases, small changes in structure are responsible for the loss of activity, way before the protein is unfolded, and thus Tm based on activity is lower than the one based on the structure of the protein (Gall et al., 2014).
TABLE 2
| Enzyme | Tm (°C) | Tm with GO (°C) |
|---|---|---|
| Rsp-1 | 58.3 ± 0.2 | 58.2 ± 0.3 |
| Rsp-2 | 60.5 ± 0.3 | 59.8 ± 1.0 |
| Cvi | 83.9 ± 2.1 | 72.3 ± 2.8 |
Melting temperatures of Rsp-1 (22.3 μg/ml), Rsp-2 (15.0 μg/ml), and Cvi (20.8 μg/ml) in phosphate buffer (5 mM, pH 7.5) in the presence (25 mg/ml) or absence of GO, as calculated by the analysis of the ellipticity at 222 nm. Experimental details are provided in Paragraph 2.7.2. All data are measured at least in duplicate.
Discussion
Graphite oxide interactions with enzymes and peptides has been widely investigated in past studies (Baptista-Pires et al., 2014; Wang et al., 2015; Pandit and De, 2017), mainly in terms of immobilization (Zhang et al., 2010; Adeel et al., 2018), and reviews have been published trying to elucidate the leading interactions present in graphite oxide and biomolecules (Nie et al., 2019; Chaudhary et al., 2021). All aforementioned studies conclude that the forces that reign the nanomaterial-biomolecule interactions are hydrophophic, electrostatic, π-π stacking, and hydrogen bonds. The extent of the interactions depends on many structural factors on both enzymes and nanomaterials. Huang et al. (2020) investigated the interactions of trypsin with GO and found that both secondary and tertiary structure of the enzyme is affected, due to the intramolecular interactions mentioned before, mainly due to hydrophobic interactions. In another study, Zhang and colleagues (2010) immobilized efficiently horseradish peroxidase onto GO and found that the driving force was hydrogen bonds formed with the oxygen functionalities of the nanomaterials. Trypsin has higher affinity to hydrophobic surfaces (Koutsopoulos et al., 2007), while horseradish peroxidase is a more hydrophilic protein (Jia et al., 2010). A deeper physicochemical investigation was performed by Pandit and De (2017), where they explored the interaction of peptides with graphite oxide, in terms of thermodynamics, and their findings suggest that electrostatic interactions with Lys, His, Arg, Phe, and Tyr and π-π stacking with Trp are the ruling forces when peptides and graphite oxide meet. All the above clearly show the complicated way this material interacts with enzymes and their building blocks.
To the best of our knowledge, this is the first report on interactions of graphite oxide with amine transaminases. The (S)-selective ATAs form dimers, with the catalytic center located in the interface of the two monomers. The structure of the (S)-ATAs studied in our work is not heavily influenced from the presence of GO, as shown from the CD measurements. Thus, the GO does not seem to intercalate in the dimer formation, something shown also from the fact that all (S)-ATAs remain active upon interaction with the GO. Our findings suggest that the catalytic behavior differences observed are structure dependent, a fact which is in accordance with the aforementioned studies for other enzyme classes. The surface chemistry of enzymes heavily influences the interactions of the enzymes with the nanomaterials, and even the localization of the interacting groups on the surface can be critical. In a previous work we have identified that hydrolases interact with GO and carbon nanotubes both with hydrophobic and electrostatic interactions (Pavlidis et al., 2012). Το understand the reason of the alterations in catalytic behavior of the enzymes studied, we calculated their available surface area, their electrostatic potential, and their total charge (see Supplementary Table S2). As a reminder, the catalytic behavior of Vfl and Cvi, in a larger extent, are negatively influenced from the presence of GO, while the Rsp variants and Rpo are activated. The surface area is not correlated to these findings. On the contrary, the total charge of the proteins investigated seems to have some correlation; the enzymes that are activated are more negatively charged, while the ones whose activity and stability are decreased are also negatively charged but have significantly less total charge. A direct correlation cannot be established, as the localization of the interaction is quite important. For instance, if the interaction happens close to the entrance to the active site, it can lead to diffusion limitations for the reactants. Nevertheless, a visual observation of the 3D structures of the four crystal structures analyzed in Supplementary Table S2 did not reveal hydrophobic or positively charged patches on their surface, and thus facilitate the interactions (data not shown). Nevertheless, we need to state that the correlation observed here does not imply causation, and further studies are required to fully understand the effect. Cvi clearly loses its activity and stability when the nanomaterial is present and does not seem to be a suitable enzyme for immobilization studies with this particular carbon-based nanomaterial. Rsp and Rpo seem more suitable candidates for immobilization studies; however, their negative charge may lead to repulsive forces with the nanomaterial and thus to low enzyme loading.
Conclusion
The effect of nanomaterials on the catalytic behavior of enzymes strongly depends on the interactions developed between the counterparts. Herein we observed that the effect of GO on the activity of several (S)-selective ATAs is not uniform and can greatly vary. The addition of up to 0.1 mg/ml GO in the reaction medium leads to up to 50% improved activity of (S)-ATAs that are not highly negatively charged, while these enzymes maintain their thermal stability. However, in (S)-ATAs with significant negative charge, the effect can be the opposite. The (S)-ATA from Chromobacterium violaceum loses almost 50% of its activity in the presence of 0.1 mg/ml GO. At the same time, its stability was significantly decreased in the presence of 25 μg/ml GO, losing its activity at a double rate compared to its thermal inactivation in the absence of GO. Although the prediction of the effects on the catalytic behavior is not yet possible, we still can relate the changes in activity and stability with some local minor structural changes. For instance, the negative effect observed in the activity and stability of Cvi seems to rise as a loss of α-helices, accompanied with an equal increase in random coil (∼3% in the presence of 25 μg/ml GO). These changes seem to arise from the weak interactions of GO with the ATAs. From this work, a hint is provided that the total charge of the enzyme, at least for the (S)-ATAs studied, may be correlated with the effect on the catalytic behavior, as it is the driving force for the intensity of the interactions between the nanomaterial and the enzyme. Nevertheless, we are still far from predicting the interactions and their effect on the behavior of enzymes, in order to be able to rationally design nanobiocatalytic systems.
Statements
Data availability statement
The datasets presented in this study can be found in FAIRDOMHub, under the SEEK ID: https://fairdomhub.org/publications/637.
Author contributions
IP conceived and initiated the project. PZ, NC, and KS synthesized, and characterized the GO, under the supervision of DG. NK designed and performed the enzymatic activity and CD experiments. NK and IP contributed to data interpretation and manuscript preparation. NK and IP have drafted the first version of the manuscript and all authors and edited and gave their approval to the final version of the manuscript.
Funding
The research project was supported by the Hellenic Foundation for Research and Innovation (H.F.R.I.) under the “1st Call for H.F.R.I. Research Projects to support Faculty Members and Researchers and the Procurement of High-Cost Research Equipment Grant” (Project Number:664).
Acknowledgments
All authors would like to thank the Department of Biotechnology and Enzyme Catalysis of University of Greifswald (Germany) and especially Prof. Dr. UT Bornscheuer for the constructs of the ATAs, as well as Prof. Dr. Michail Kokkinides for the access to the spectropolarimeter for the CD measurements.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors, and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fctls.2021.803850/full#supplementary-material
References
1
Adeel M. Bilal M. Rasheed T. Sharma A. Iqbal H. M. N. (2018). Graphene and Graphene Oxide: Functionalization and Nano-Bio-Catalytic System for Enzyme Immobilization and Biotechnological Perspective. Int. J. Biol. Macromolecules120, 1430–1440. 10.1016/j.ijbiomac.2018.09.144
2
Andrade M. A. Chacón P. Merelo J. J. Morán F. (1993). Evaluation of Secondary Structure of Proteins from UV Circular Dichroism Spectra Using an Unsupervised Learning Neural Network. Protein Eng. Des. Sel6 (4), 383–390. 10.1093/protein/6.4.383
3
Baptista-Pires L. Pérez-López B. Mayorga-Martinez C. C. Morales-Narváez E. Domingo N. Esplandiu M. J. et al (2014). Electrocatalytic Tuning of Biosensing Response through Electrostatic or Hydrophobic Enzyme-Graphene Oxide Interactions. Biosens. Bioelectron.61, 655–662. 10.1016/j.bios.2014.05.028
4
Bourlinos A. B. Gournis D. Petridis D. Szabó T. Szeri A. Dékány I. (2003). Graphite Oxide: Chemical Reduction to Graphite and Surface Modification with Primary Aliphatic Amines and Amino Acids. Langmuir19 (15), 6050–6055. 10.1021/la026525h
5
Bradford M. M. (1976). A Rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding. Anal. Biochem.72 (1-2), 248–254. 10.1006/abio.1976.9999
6
Chaudhary K. Kumar K. Venkatesu P. Masram D. T. (2021). Protein Immobilization on Graphene Oxide or Reduced Graphene Oxide Surface and Their Applications: Influence over Activity, Structural and thermal Stability of Protein. Adv. Colloid Interf. Sci289, 102367. 10.1016/j.cis.2021.102367
7
De Diego T. Lozano P. Gmouh S. Vaultier M. Iborra J. L. (2005). Understanding Structure−Stability Relationships of Candida Antartica Lipase B in Ionic Liquids. Biomacromolecules6 (3), 1457–1464. 10.1021/bm049259q
8
Di Giosia M. Marforio T. D. Cantelli A. Valle F. Zerbetto F. Su Q. et al (2020). Inhibition of α-chymotrypsin by Pristine Single-wall Carbon Nanotubes: Clogging up the Active Site. J. Colloid Interf. Sci.571, 174–184. 10.1016/j.jcis.2020.03.034
9
Dreyer D. R. Jia H. P. Bielawski C. W. (2010). Graphene Oxide: a Convenient Carbocatalyst for Facilitating Oxidation and Hydration Reactions. Angew. Chem. Int. Ed. Engl.49 (38), 6813–6816. 10.1002/anie.201002160
10
Ferandi E. E. Monti D. (2018). Amine Transaminases in Chiral Amines Synthesis: Recent Advances and Challenges. World J. Microbiol. Biotechnol.34 (1), 13. 10.1007/s11274-017-2395-2
11
Fuchs M. Farnberger J. E. Kroutil W. (20152015). The Industrial Age of Biocatalytic Transamination. Eur. J. Org. Chem.2015 (32), 6965–6982. 10.1002/ejoc.201500852
12
Gall M. G. Nobili A. Pavlidis I. V. Bornscheuer U. T. (2014). Improved Thermostability of a Bacillus Subtilis Esterase by Domain Exchange. Appl. Microbiol. Biotechnol.98, 1719–1726. 10.1007/s00253-013-5053-0
13
Huang S. Li H. Liu Y. Yang L. Wang D. Xiao Q. (2020). Investigations of Conformational Structure and Enzymatic Activity of Trypsin after its Binding Interaction with Graphene Oxide. J. Hazard. Mater.392, 122285. 10.1016/j.jhazmat.2020.122285
14
Hummers W. S. Offeman R. E. (1958). Preparation of Graphitic Oxide. J. Am. Chem. Soc.80 (6), 1339. 10.1021/ja01539a017
15
Jafarian F. Bordbar A.-K. Zare A. Khosropour A. (2018). The Performance of Immobilized Candida Rugosa Lipase on Various Surface Modified Graphene Oxide Nanosheets. Int. J. Biol. Macromolecules111, 1166–1174. 10.1016/j.ijbiomac.2018.01.133
16
Jia W. Schwamborn S. Jin C. Xia W. Muhler M. Schuhmann W. et al (2010). Towards a High Potential Biocathode Based on Direct Bioelectrochemistry between Horseradish Peroxidase and Hierarchically Structured Carbon Nanotubes. Phys. Chem. Chem. Phys.12, 10088–10092. 10.1039/c0cp00349b
17
Kelly S. A. Mix S. Moody T. S. Gilmore B. F. (2020). Transaminases for Industrial Biocatalysis: Novel Enzyme Discovery. Appl. Microbiol. Biotechnol.104 (11), 4781–4794. 10.1007/s00253-020-10585-0
18
Kelly S. A. Pohle S. Wharry S. Mix S. Allen C. C. R. Moody T. S. et al (2018). Application of ω-Transaminases in the Pharmaceutical Industry. Chem. Rev.118 (1), 349–367. 10.1021/acs.chemrev.7b00437
19
Kouloumpis A. Thomou E. Chalmpes N. Dimos K. Spyrou K. Bourlinos A. B. et al (2017). Graphene/Carbon Dot Hybrid Thin Films Prepared by a Modified Langmuir-Schaefer Method. ACS Omega2 (5), 2090–2099. 10.1021/acsomega.7b00107
20
Kouloumpis A. Vourdas N. Zygouri P. Chalmpes N. Potsi G. Kostas V. et al (2018). Controlled Deposition of Fullerene Derivatives within a Graphene Template by Means of a Modified Langmuir-Schaefer Method. J. Colloid Interf. Sci.524, 388–398. 10.1016/j.jcis.2018.04.049
21
Koutsopoulos S. Patzsch K. Bosker W. T. E. Norde W. (2007). Adsorption of Trypsin on Hydrophilic and Hydrophobic Surfaces. Langmuir23 (4), 2000–2006. 10.1021/la062238s
22
Liu D.-M. Dong C. (2020). Recent Advances in Nano-Carrier Immobilized Enzymes and Their Applications. Process Biochem.92, 464–475. 10.1016/j.procbio.2020.02.005
23
Miles A. J. Ramalli S. G. Wallace B. A. (2021). DichroWeb , a Website for Calculating Protein Secondary Structure from Circular Dichroism Spectroscopic Data. Protein Sci. 10.1002/pro.4153
24
Min K. Kim J. Park K. Yoo Y. J. (2012). Enzyme Immobilization on Carbon Nanomaterials: Loading Density Investigation and Zeta Potential Analysis. J. Mol. Catal. B: Enzymatic83, 87–93. 10.1016/j.molcatb.2012.07.009
25
Navalon S. Dhakshinamoorthy A. Alvaro M. Antonietti M. García H. (2017). Active Sites on Graphene-Based Materials as Metal-free Catalysts. Chem. Soc. Rev.46 (15), 4501–4529. 10.1039/c7cs00156h
26
Nie C. Ma L. Li S. Fan X. Yang Y. Cheng C. et al (2019). Recent Progresses in Graphene Based Bio-Functional Nanostructures for Advanced Biological and Cellular Interfaces. Nano Today26, 57–97. 10.1016/j.nantod.2019.03.003
27
Pandit S. De M. (2017). Interaction of Amino Acids and Graphene Oxide: Trends in Thermodynamic Properties. J. Phys. Chem. C121, 600–608. 10.1021/acs.jpcc.6b11571
28
Patila M. Chalmpes N. Dounousi E. Stamatis H. Gournis D. (2020). Use of Functionalized Carbon Nanotubes for the Development of Robust Nanobiocatalysts. Methods Enzymol.630, 263–301. 10.1016/bs.mie.2019.10.015
29
Patila M. Orfanakis G. Polydera A. C. Pavlidis I. V. Stamatis H. (2017). “Graphene-Based Nanobiocatalytic Systems,” in Biocatalysis & Nanotechnology. Editor GrunwaldP. (Boca Raton: Pan Stanford Series on Biocatalysis), 243–277. 10.1201/9781315196602-6
30
Patila M. Pavlidis I. V. Kouloumpis A. Dimos K. Spyrou K. Katapodis P. et al (2016). Graphene Oxide Derivatives with Variable Alkyl Chain Length and Terminal Functional Groups as Supports for Stabilization of Cytochrome C. Int. J. Biol. Macromolecules84, 227–235. 10.1016/j.ijbiomac.2015.12.023
31
Pavlidis I. V. Patila M. Bornscheuer U. T. Gournis D. Stamatis H. (2014). Graphene-based Nanobiocatalytic Systems: Recent Advances and Future Prospects. Trends Biotechnol.32 (6), 312–320. 10.1016/j.tibtech.2014.04.004
32
Pavlidis I. V. Vorhaben T. Tsoufis T. Rudolf P. Bornscheuer U. T. Gournis D. et al (2012). Development of Effective Nanobiocatalytic Systems through the Immobilization of Hydrolases on Functionalized Carbon-Based Nanomaterials. Bioresour. Tech.115, 164–171. 10.1016/j.biortech.2011.11.007
33
Pavlidis I. V. Weiß M. S. Genz M. Spurr P. Hanlon S. P. Wirz B. et al (2016). Identification of (S)-selective Transaminases for the Asymmetric Synthesis of Bulky Chiral Amines. Nat. Chem8 (11), 1076–1082. 10.1038/nchem.2578
34
Potsi G. Bourlinos A. B. Mouselimis V. Poláková K. Chalmpes N. Gournis D. et al (2019). Intrinsic Photoluminescence of Amine-Functionalized Graphene Derivatives for Bioimaging Applications. Appl. Mater. Today17, 112–122. 10.1016/j.apmt.2019.08.002
35
Savile C. K. Janey J. M. Mundorff E. C. Moore J. C. Tam S. Jarvis W. R. et al (2010). Biocatalytic Asymmetric Synthesis of Chiral Amines from Ketones Applied to Sitagliptin Manufacture. Science329 (5989), 305–309. 10.1126/science.1188934
36
Schätzle S. Höhne M. Redestad E. Robins K. Bornscheuer U. T. (2009). Rapid and Sensitive Kinetic Assay for Characterization of ω-Transaminases. Anal. Chem.81 (19), 8244–8248. 10.1021/ac901640q
37
Tsirka K. Katsiki A. Champles N. Gournis D. Paipetis A. S. (2018). Mapping of Graphene Oxide and Single Layer Graphene Flakes—Defects Annealing and Healing. Front. Mater.5, 37. 10.3389/fmats.2018.00037
38
Wang Z. Li Y. Li L. Li D. Huang Y. Nie Z. et al (2015). DNA-mediated Supercharged Fluorescent Protein/graphene Oxide Interaction for Label-free Fluorescence Assay of Base Excision Repair Enzyme Activity. Chem. Commun.51 (69), 13373–13376. 10.1039/c5cc04759e
39
Weiß M. S. Pavlidis I. V. Spurr P. Hanlon S. P. Wirz B. Iding H. et al (2016). Protein-engineering of an Amine Transaminase for the Stereoselective Synthesis of a Pharmaceutically Relevant Bicyclic Amine. Org. Biomol. Chem.14, 10249–10254. 10.1039/c6ob02139e
40
Wu B.-P. Wen Q. Xu H. Yang Z. (2014). Insights into the Impact of Deep Eutectic Solvents on Horseradish Peroxidase: Activity, Stability and Structure. J. Mol. Catal. B: Enzymatic101, 101–107. 10.1016/j.molcatb.2014.01.001
41
Yüce M. Kurt H. (2017). How to Make Nanobiosensors: Surface Modification and Characterisation of Nanomaterials for Biosensing Applications. RSC Adv.7 (78), 49386–49403. 10.1039/c7ra10479k
42
Zahirinejad S. Hemmati R. Homaei A. Dinari A. Hosseinkhani S. Mohammadi S. et al (2021). Nano-organic Supports for Enzyme Immobilization: Scopes and Perspectives. Colloids Surf. B204, 111774. 10.1016/j.colsurfb.2021.111774
43
Zhang J. Zhang F. Yang H. Huang X. Liu H. Zhang J. et al (2010). Graphene Oxide as a Matrix for Enzyme Immobilization. Langmuir26 (9), 6083–6085. 10.1021/la904014z
Summary
Keywords
transaminase, graphite oxide (GO), interaction, circular dichroism (CD), function-structure analysis
Citation
Kaloudis N, Zygouri P, Chalmpes N, Spyrou K, Gournis D and Pavlidis IV (2022) Effect of Graphite Oxide on the Catalytic Behavior of (S)-Selective Amine Transaminases. Front. Catal. 1:803850. doi: 10.3389/fctls.2021.803850
Received
28 October 2021
Accepted
06 December 2021
Published
01 February 2022
Volume
1 - 2021
Edited by
Francesca Paradisi, University of Bern, Switzerland
Reviewed by
José Cleiton Sousa dos Santos, University of International Integration of Afro-Brazilian Lusophony, Brazil
Ana Beloqui, Polymat, Spain
Updates
Copyright
© 2022 Kaloudis, Zygouri, Chalmpes, Spyrou, Gournis and Pavlidis.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ioannis V. Pavlidis, ipavlidis@uoc.gr
This article was submitted to Biocatalysis, a section of the journal Frontiers in Catalysis
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.