- 1Department of Biology, Pennsylvania State University, University Park, PA, United States
- 2Department of Biology, Florida International University, Miami, FL, United States
- 3Florida Keys Marine Lab, Long Key, FL, United States
- 4Department of Marine and Environmental Sciences, Nova Southeastern University, Fort Lauderdale, FL, United States
- 5Florida Fish and Wildlife Conservation Commission, Marathon, FL, United States
With ongoing changes in climate, rare and ecologically specialized species are at increased risk of extinction. In sessile foundation fauna that reproduce asexually via fragmentation of existing colonies, the number of colonies does not reflect the number of genets and thus can obscure genotypic diversity. Colonies that are the product of fragmentation are not visually distinguishable from colonies that stem from sexual recruits. For this reason, molecular markers are necessary to assess genotypic variation and population structure in clonal organisms such as reef-building corals and their endosymbiotic dinoflagellates. For the rare Caribbean pillar coral, Dendrogyra cylindrus, and its endosymbiotic dinoflagellate, Breviolum dendrogyrum, we use de novo microsatellite markers to infer past demographic changes, describe modern population structure, and quantify the frequency of asexual reproduction. Our analyses show that D. cylindrus comprises three distinct populations across the Greater Caribbean whereas the symbiont could be differentiated into four populations, indicating barriers to gene flow differ between host and symbiont. In Florida, host and symbiont populations reproduced mainly asexually, yielding lower genotypic diversity than predicted from census size. When multiple coral ramets were present, they often associated with the same clonal strain of B. dendrogyrum, pointing to the high fidelity of this relationship. Models of past demographic events revealed no evidence for historical changes in population sizes, consistent with the paleontological record of D. cylindrus indicating it has been rare for hundreds of thousands of years. The most recent global thermal stress event likely triggered a severe disease outbreak among D. cylindrus in Florida, resulting in a severe population decline. Projections indicate a high likelihood that this species will become extinct in the Northern Greater Caribbean within a few decades. The ecosystem consequences of losing rare coral species and their symbionts with increasingly frequent extreme warming events are not known but require urgent study.
Introduction
Ecologically rare species are predicted to be more vulnerable to environmental change and extinction risk with shifts in climate (Caughley, 1994; McKinney, 1997; Davies et al., 2004). Species are rare because they may inhabit narrow geographic ranges (Solórzano et al., 2016), occupy few specific habitats (Deák et al., 2018), and/or exhibit low abundance in nature (Rabinowitz, 1981). There are obvious consequences to having low population densities, including difficulties finding a mate (Stephens and Sutherland, 1999) and vulnerability to genetic drift (Ellstrand and Elam, 1993). Populations of rare species may become fragmented more easily due to the increased local extinction of small populations (Matthies et al., 2004; Blanquer and Uriz, 2010). The persistence of rare species is challenged when they are obligate partners of specific symbionts. For example, the endangered terrestrial orchid Caladenia huegelii associates with a specific mycorrhizal fungus throughout its range (Swarts et al., 2010). This specificity between partners has caused C. huegelii to be rare due to the limited suitable environmental conditions of the mycorrhiza. Consequentially, the strict niche characteristics of one partner in a symbiosis may drive the scarcity of the other. Further, intra-specific diversity in both partners and fidelity of genotype–genotype associations can play a role in how the symbiosis responds to changing conditions (Baums et al., 2014b; Parkinson and Baums, 2014; Parkinson et al., 2015). However, there are many rare species that are adapted to low population densities (de Lange and Norton, 2004; Flather and Sieg, 2007; Blanquer and Uriz, 2010), and these species persist for long periods of evolutionary time.
Rarity is sometimes obscured by the life history of the species, specifically in reference to the prevalence of asexual reproduction. Sessile organisms often reproduce by asexual fragmentation and while some are hermaphroditic, these may be self-incompatible (Scobie and Wilcock, 2009; Gitzendanner et al., 2012; Baums et al., 2013). As a consequence, large clonal stands of a rare species may exist that are unable to reproduce via sexual reproduction. Clonal populations of the rare dwarf shrub Linnaea borealis had low reproductive success due to a lack of nearby conspecifics, leading to increased geitonogamy (Scobie and Wilcock, 2009). Likewise, the aquatic plant Decodon verticillatus had reduced sexual reproduction in clonal populations at the northern range limit (Dorken and Eckert, 2001).
However, extensive clonal reproduction can also be beneficial, especially for rare species, by maintaining population sizes despite a lack of sexual recruitment. In other words, clonal propagation can provide a “storage effect” by increasing the persistence of individual genotypes when the influx of sexual recruits is low or absent due to unfavorable environmental conditions (Warner and Chesson, 1985; Boulay et al., 2014; Dubé et al., 2017). This storage effect is most pronounced when adults are long-lived, reproductive output is high, and population densities are low – such as in some mass-spawning corals – and thus may play an important role in maintaining intraspecies genetic diversity (Baums et al., 2006, 2014a; Boulay et al., 2014).
Census sizes of many Caribbean reef-building corals have declined dramatically over the past few decades. Yet, in recent coral surveys throughout the Caribbean, some species were consistently more abundant (e.g., Orbicella faveolata), while others were rare, such as the pillar coral Dendrogyra cylindrus (Edmunds et al., 1990; Ward et al., 2006; Steiner and Kerr, 2008; Miller et al., 2013). Dendrogyra cylindrus (Ehrenburg, 1834) is a conspicuous Caribbean coral in the family Meandrinidae. It is the only species in the genus, and it occurs between 1 and 25 m depths (Goreau and Wells, 1967). This species was originally characterized as a gonochoric broadcast spawner (Szmant, 1986), however, recent spawning observations in the Florida Keys have revealed that D. cylindrus can be hermaphroditic (Neely et al., 2018). D. cylindrus has been classified as vulnerable on the International Union for Conservation of Nature (IUCN) Redlist (Aronson et al., 2008), and was listed as threatened under the United States Endangered Species Act (NMFS, 2014). The fossil record supports the relative rarity of D. cylindrus through time, although there is localized evidence that pillar corals were more prevalent on Pleistocene reefs (Hunter and Jones, 1996). Corals in the genus Dendrogyra only appear 29 times in the Paleobiology Database, whereas there are 278 Acropora palmata fossils, 111 Orbicella faveolata fossils, and 160 Meandrina (a sister genus to Dendrogyra) fossils1 (accessed February 18, 2018). Despite the growing concern over the persistence of this rare coral, no studies have described genotypic diversity, population connectivity, or past changes in population size in D. cylindrus.
Dendrogyra cylindrus forms an obligate symbiosis with another rare species, Breviolum (formerly Symbiodinium Clade B) dendrogyrum (LaJeunesse et al., 2018; Lewis et al., 2018). In recent surveys of Caribbean corals, B. dendrogyrum (formerly ITS2 type B1k) was only found to associate with D. cylindrus, indicating that it has a narrow habitat range (Finney et al., 2010). Assessing levels of within-species diversity is imperative to understanding the maintenance of and threats to the D. cylindrus-B. dendrogyrum mutualism. If co-dispersal between partners is occurring, then the population structure of both symbiotic species will be congruent (Werth and Scheidegger, 2012) and facilitate local adaptation (Baums et al., 2014b). In this study, we used de novo and existing microsatellite markers for host and symbiont to assess the null hypotheses of no population structure and no asexual reproduction in either species. We further hypothesized that coral colonies harbor B. dendrogyrum throughout the range of the host. Finally, we used a maximum-likelihood demographic model to assess the null hypothesis of no past changes in population size in D. cylindrus.
Materials and Methods
Sample Collection
Samples of D. cylindrus were collected from 51 sites along the Florida Reef Tract (n = 217), three sites in Curaçao (n = 24), six sites in the U.S. Virgin Islands (n = 40), three sites in Belize (n = 10), and four sites in the Turks and Caicos Islands (n = 17) (Figure 1). Regions were visited in different years, but all samples were collected between 2013 and 2017. Because of the rarity of D. cylindrus, we targeted sites with known occurrences of the coral instead of attempting a random sampling scheme. Sampling intensity was higher in Florida, but not exhaustive due to the threatened status of this species. At Florida sites containing many colonies, samples were collected from colonies throughout the site. When sites contained only a few colonies, all were sampled. Also, within-colony sampling (top, middle, and base) was done in Florida to determine if symbiont genotypes were represented consistently within a colony. D. cylindrus samples from all other regions were not collected in a spatially explicit way, rather, colonies were sampled haphazardly. Sampling was accomplished using a combination of clipping small pieces of tissue and conducting a biopsy of two or three polyps using the syringe technique (Kemp et al., 2008). All samples were preserved in 95% non-denatured ethanol. Global positioning system (GPS) coordinates were collected for the Florida colonies so that genetic and clonal diversity estimates could be related to geographic distance among colonies.
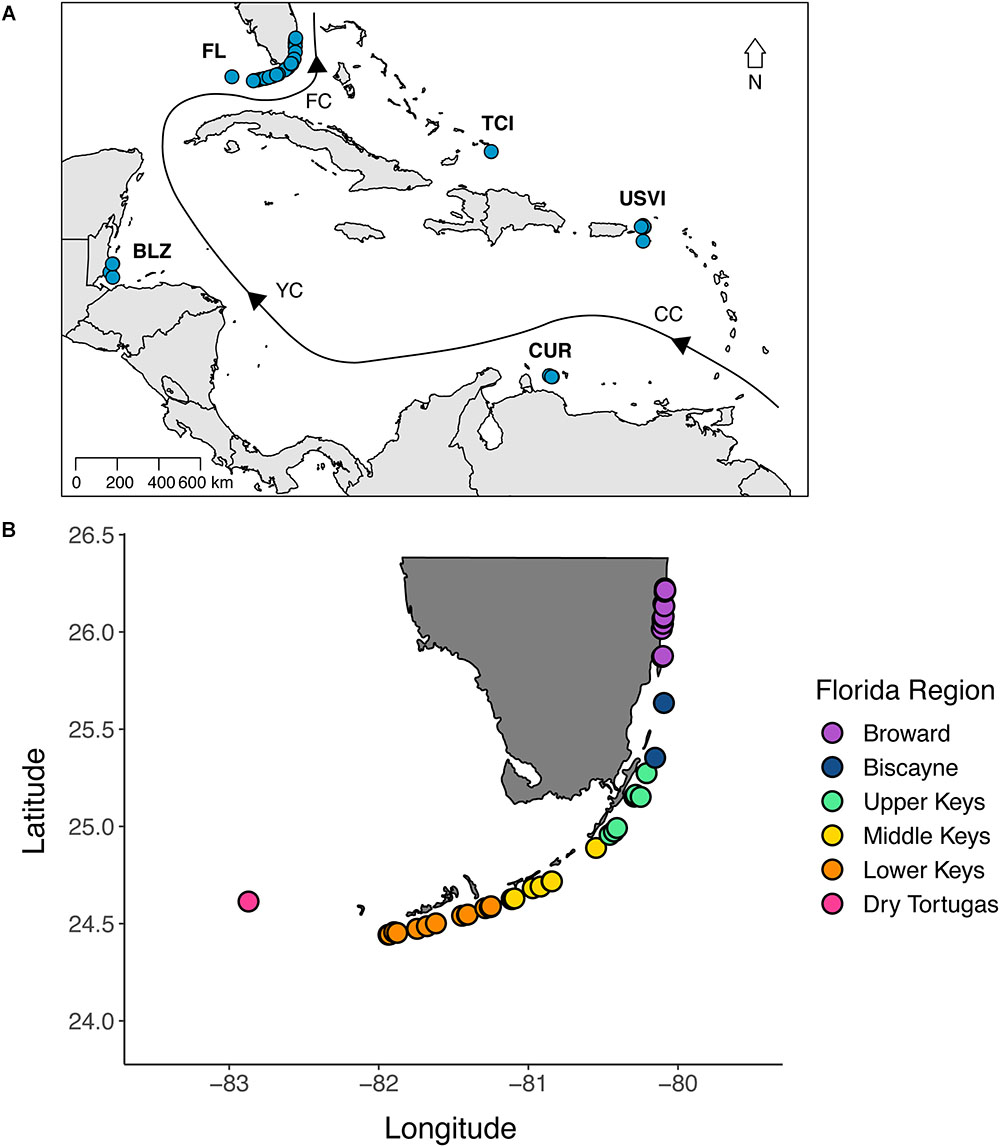
Figure 1. (A) A map of the sites where tissue samples of Dendrogyra cylindrus were collected. Samples were collected from 51 sites along the Florida Reef Tract (FL), three sites in Curaçao (CUR), six sites in the U.S. Virgin Islands (USVI), three sites in Belize (BLZ), and four sites in the Turks and Caicos Islands (TCI). The black line with arrows indicates the direction of ocean currents. CC, Caribbean Current; YC, Yucatan Current; FC, Florida Current. See text for sample sizes. (B) A map of the Florida collection sites, colored by region. More samples were collected from Florida to enable the description of clonal structure and to delineate the relationship between genetic and geographic distance.
Microsatellite Design and Amplification
DNA was extracted from one sample from Florida using the illustra Nucleon PhytoPure Genomic DNA Extraction kit (GE Healthcare) and used as the starting material for generating molecular markers. Because this sample was from an adult colony, DNA from the coral host and dinoflagellate symbiont was sequenced on the Roche GS 454 FLX+ utilizing the Titanium Sequencing Kit (Roche, one half plate). Raw data is available from Penn State’s ScholarSphere2. See Supplementary Materials for detail in primer design. After testing primers for successful amplification, host specificity, and variability, 11 markers were retained (Supplementary Table 1). These markers were combined into four multiplex reactions (labeled A through D, Supplementary Table 1) using the Multiplex Manager software3. Only markers labeled with different colored fluorescent dyes (Applied Biosystems) were combined. Multiplex reactions consisted of 1 μl of template DNA, 1.01x Reaction Buffer (New England Biolabs), 2.53 mM MgCl (New England Biolabs), 0.005 mg BSA (New England Biolabs), 0.202 mM dNTPs (Bioline), 0.051 μM of forward and reverse primer (Applied Biosystems), 0.75 U Taq polymerase (New England Biolabs), and nuclease-free water in a total reaction volume of 9.9 μl. Thermocycler parameters included an initial denaturation at 94°C for 5 min, 30 cycles of denaturing at 94°C for 20 s, annealing at 55°C (or 54°C) for 20 s, and extension at 72°C for 30 s, and a final extension at 72°C for 30 min. All PCR products were visualized using an ABI3730 (Applied Biosystems) automated DNA sequencer with an internal size standard (Gene Scan 500-Liz, Applied Biosystems) for accurate sizing. Electropherograms were analyzed using GeneMapper Software 5.0 (Applied Biosystems).
Existing Symbiodiniaceae B1 primers (Santos and Coffroth, 2003; Pettay and LaJeunesse, 2007; Andras et al., 2009) were tested on D. cylindrus samples from Florida and Curaçao, of which eight were variable and amplified the target product a majority of the time (Supplementary Table 2). These were combined into multiplex reactions (Supplementary Tables 2, 3). Thermocycler parameters (e.g., annealing temperatures) varied by multiplex (Supplementary Table 3). However, after conducting preliminary analyses using this set of markers, it became clear that additional markers would be necessary to increase our power of identifying clonal strains. See the supplement for details on de novo primer design and amplification conditions. The combined set of 15 microsatellite markers (Supplementary Table 2) was used to assess clonal and population structure of the algal symbiont.
Analysis of Population and Clonal Structure
Matching multilocus genotypes were identified using the Data Subset option in GENALEX v. 6.5 (Peakall and Smouse, 2006). Samples with exact allele size matches at all microsatellite loci were considered to be asexually produced clones. The Multilocus Matches option (GENALEX) was used to identify possible misidentified alleles, which were checked and corrected if necessary. Linkage disequilibrium tests were conducted using GENEPOP ON THE WEB 4.0 (Rousset, 2008) Option 2.1. Tests for Hardy–Weinberg equilibrium were conducted in GENALEX v. 6.5 (Peakall and Smouse, 2006) and confirmed using exact tests in GENEPOP ON THE WEB Option 1.1 (Supplementary Table 5). The probability of identity (PI) describes the power of the microsatellite markers to distinguish closely related colonies (such as siblings) from colonies that are identical because they are the result of asexual reproduction (fragmentation). For the diploid D. cylindrus, PI was calculated in GENALEX v. 6.5. The PI for the haploid symbiont was calculated as the sum of each allele’s frequency squared in each population and multiplied across loci, in accordance with previous studies (Baums et al., 2014b).
Genotypic diversity indices for the coral host and symbiont, which included genotypic richness, diversity, and evenness, were calculated as in Baums et al. (2006) using the entire data set of multilocus genotypes (including clones) with complete allele calls at 10 microsatellite markers (excluding marker D48). Briefly, genotypic richness (Ng/N) was calculated by dividing the number of unique multilocus genotypes (Ng) by the total number of samples genotyped (N) and varies from near 0 (highly clonal) to 1 (no clones). Genotypic diversity (Go/Ge) is the ratio of observed to expected genotypic diversity, and varies from 0 (mostly asexual population) to 1 (mostly sexual population). Expected genotypic diversity (Ge) for a solely sexually reproducing population was assumed to be equal to the sample size N. The final metric – genotypic evenness (Go/Ng) – provides information about the longevity of different genotypes, and is the observed genotypic diversity (Go) divided by the number of unique multilocus genotypes (Ng). Genotypic evenness ranges from near 0 (population dominated by few genets with many ramets) to 1 (population contains multiple genets with equal ramets). However, in sites with only one genet, genotypic evenness is no longer an informative metric since it automatically yields a value of one.
Unique multilocus genotypes (without clonal replicates) were clustered using the program STRUCTURE v. 2.3.4 to estimate the number of different populations of host and symbiont in the dataset (Pritchard et al., 2000). After initial testing, the admixture model with no location prior was used with correlated allele frequencies. The prior number of populations (K) was set from 1 to 6 with five replicate runs per K, a burnin of 100,000 and 1,000,000 Markov Chain Monte Carlo repetitions after the burnin using the package ‘PARALLELSTRUCTURE’ (Besnier and Glover, 2013) in R v. 3.4 (R Development Core Team, 2017). The optimal value for K was identified using the ΔK method (Evanno et al., 2005) implemented in the online program STRUCTURE HARVESTER (Earl, 2012). CLUMPAK was used to identify the consensus of inferred clusters for the different replicates of each K-value and to visualize the results (Kopelman et al., 2015). Additional estimators of the number of optimal populations (posterior probability, MedMeaK, MaxMeaK, MedMedK, and MaxMedK) were applied (Pritchard et al., 2009; Puechmaille, 2016). Because uneven sampling has been shown to underestimate the true K (Puechmaille, 2016), larger populations were randomly subsampled to create more even sample sizes across the five regions (see Results). The K value chosen by the most estimators was identified as the optimal number of populations. Global FST, or the proportion of genetic variation between subpopulations (Wright, 1951), was calculated using GENODIVE (Meirmans and Van Tienderen, 2004) and the dataset with clonal replicates removed. The FST value adjusted for the maximum amount of within-population diversity (F′ST) was also obtained from GENODIVE. Pairwise FST values between each of the five regions were calculated using GENALEX v. 6.5. The congruence of population structure and clonal reproduction in D. cylindrus and B. dendrogyrum, as well as the extent of clonal propagation, were compared. In cases where the symbiont population clusters identified by STRUCTURE did not correspond to geography, the symbiont species identity was confirmed by sequencing the Si15 microsatellite flanker region (Finney et al., 2010; Lewis et al., 2018).
Analysis of molecular variance (AMOVA) was employed to test the null hypotheses of an absence of population structure for the host and symbiont (Excoffier et al., 1992). Samples were grouped by region (identified using STRUCTURE and the optimal K value) and location (Belize, Florida, Turks and Caicos Islands, USVI, and Curaçao). The AMOVA option in GENALEX v. 6.5 was used to partition genetic variation from a distance matrix between regions and locations, using 9,999 permutations and assuming an Infinite Allele Model.
Global positioning system coordinates for all Florida colonies were used to create a geographic distance matrix, which was compared to the genetic distance matrix for D. cylindrus. Mantel tests of isolation by distance and spatial autocorrelation analyses were conducted in GENALEX v. 6.5.
Because of the small number of samples, a finding of no population structure could be due to a lack of power. Thus, simulations were conducted in POWSIM v. 4.1 to assess the power of the microsatellite data set to detect low levels of population differentiation. POWSIM estimates the lowest level of differentiation (FST) that can be detected in a simulated population with a minimum of 90% accuracy (Ryman and Palm, 2006). Significance was determined using Fisher’s exact test.
Population Demographic Modeling
The modeling software MIGRAINE v. 0.5.24 was used to test for past changes in population size in the Florida D. cylindrus population. Samples from other locations were not grouped with Florida because population structure within a dataset yields inaccurate demographic parameter inference (Leblois et al., 2014). Additional unique multilocus genotypes were included for this analysis to increase the power of detecting historical population size changes (n = 95 total). The null model assuming a constant population size (OnePop) was run to estimate pGSM, which was close to 0.3. This parameter describes the geometric distribution of mutation step size under a generalized stepwise mutation model (GSM), which is most appropriate for microsatellite markers (Leblois et al., 2014). When we ran the model assuming one past change in population size (OnePopVarSize), we fixed pGSM at 0.3 and inferred the current (2Nμ) and ancestral (2Nancμ) population sizes, as well as the duration of the contraction or expansion event (Dg/2N) and the ratio of current to ancestral population size (Nratio). We assumed a mutation rate (μ) of 5 × 10-4 per locus per generation, a value typical for microsatellite loci (Estoup and Angers, 1998). In addition, we also ran the OnePopFounderFlush model, which assumes two past changes in population size and infers a founder population size (2Nfounderμ) and two additional population size ratios (NcurNfounderratio and NfounderNancratio). Significant population size changes were detected using 95% confidence interval estimates of the population size ratios. If the value 1 (indicating the current and ancestral population sizes were identical) was outside of the 95% confidence interval, then the size change was significant. MIGRAINE uses sequential importance sampling algorithms (De Iorio and Griffiths, 2004a,b) to estimate parameters of past demographic changes from population genetic data. We used 2,000 points, 2,000 trees, and four iterations per run for the models with population size changes. The null model was run using 2,000 points, 100 trees, and three iterations. Two independent runs were conducted per model by changing the estimation seed. Additional model settings are included in the supplement (Supplementary Table 4).
Through the course of this study, we witnessed a severe decline in Florida D. cylindrus (Neely et al., unpublished data). During the summers of 2014 and 2015, water temperatures rose above 30.5°C in the Florida Keys5, triggering a multi-year thermal stress event (Manzello, 2015; Lewis et al., 2017). After some coral colonies bleached in 2014, many contracted a tissue loss disease (Precht et al., 2016). This thermal stress-related disease outbreak led to a massive loss of Florida pillar corals. Some areas experienced higher disease prevalence than others (Neely et al., unpublished data). In Broward County (Figure 1B), 86% of the known colonies were lost in 2 years (Kabay, 2016). Severe hyperthermal events such as this one are expected to occur annually by 2042 on average under Representative Concentration Pathway (RCP) 8.5 (van Hooidonk et al., 2017), although there is projected heterogeneity along the Florida Reef Tract (van Hooidonk et al., 2015).
Prior to the most recent event, extensive coral bleaching from hyperthermal stress in the Caribbean occurred in 1997–1998, 2005, and 2010 (Eakin et al., 2010; Heron et al., 2016). If we assume that thermal stress events such as the 2014–2015 event in Florida occur twice per decade up until the estimated timing of annual severe bleaching (2042), we can estimate how many years are left before local extinction of D. cylindrus for each of three representative percent decline scenarios (80%, 50%, and 20%).
If we extrapolate our data on the number of coral genotypes and symbiont strains in our sample of 161 D. cylindrus colonies to the estimated total Florida population of 610 colonies (Lunz et al., 2016), we would expect to find 212 coral genotypes and 110 symbiont strains. Using our data on the D. cylindrus genotype frequencies and B. dendrogyrum strain frequencies, we simulated the decline of Florida D. cylindrus with future thermal stress-related disease events. We generated 610 D. cylindrus colonies using the statistical software R and assigned each colony a coral genotype and symbiont strain based off of the frequency distribution we observed in our sample of 161 colonies. For simplicity, we only assigned one strain per colony whereas in reality some colonies host more than one strain.
Because we do not know the actual rate of decline from the most recent thermal stress event and associated disease losses for all of Florida, we simulated a range of scenarios where 80%, 50%, and 20% of colonies survive each hyperthermal event. This model assumes that there is no sexual reproduction (a valid assumption for D. cylindrus, see section “Discussion”), no establishment of new clonal fragments, and no successful restoration. We also assume that each high temperature event is equally damaging, resulting in the same percent loss of colonies as the previous event. After running 100 simulations for each of the three rates of decline, we identified the number of thermal stress events that would cause local extinction of D. cylindrus in Florida. D. cylindrus was considered to be locally extinct once the average number of colonies (across the 100 simulations) remaining after a stress event was below one.
Results
Genetic Diversity of de novo Dendrogyra cylindrus Microsatellite Markers
The microsatellite markers developed for Dendrogyra cylindrus ranged in allelic diversity from 6 to 15 and in effective allelic diversity from 2.047 to 6.849 (Table 1). Observed heterozygosity levels ranged from 0.399 to 0.954, expected heterozygosity within subpopulations (e.g., Florida, USVI) ranged from 0.533 to 0.888, and total heterozygosity ranged from 0.686 to 0.895 (Table 1). The total heterozygosity adjusted for sampling a limited number of populations ranged from 0.724 to 0.897 (Table 1). The heterozygosity values for all markers were relatively high (closer to the maximum value of 1), indicating high genetic variability for our samples (Table 1). Variability among inbreeding measures for the 11 markers were between -0.19 and 0.361, with most values close to zero (Table 1). The markers differed in their genetic variability, with D15 showing the lowest differentiation across all four measures (Table 1). D716 and D48 showed the most variability. Markers were tested for deviations from Hardy–Weinberg equilibrium and linkage disequilibrium, if applicable (see Supplementary Text).
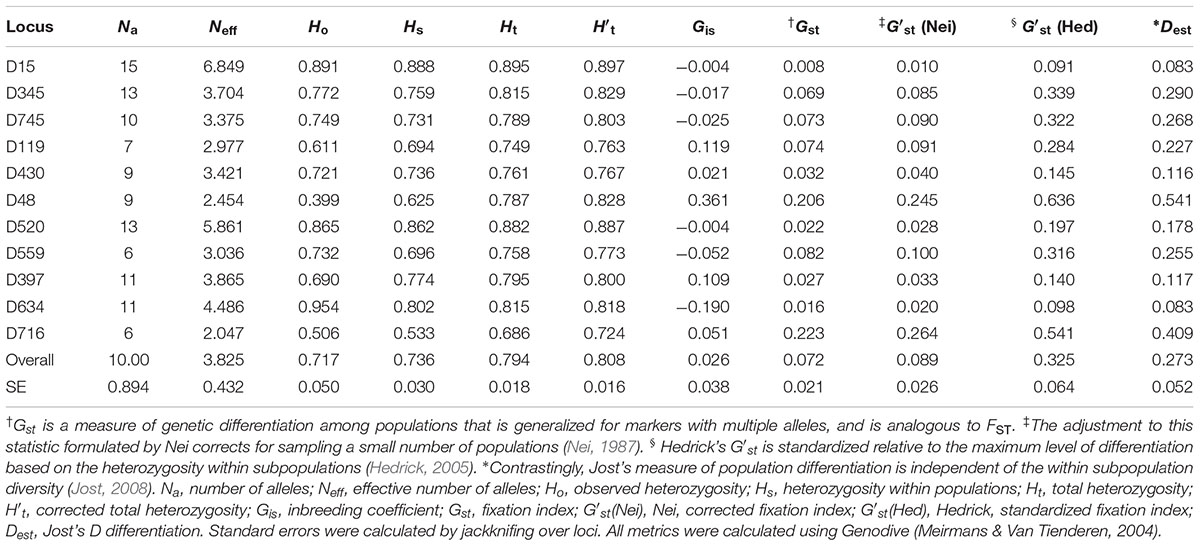
Table 1. Summary statistics per locus for 11 de novo microsatellite markers for Dendrogyra cylindrus.
Clonal Structure and Spatial Analyses Along the Florida Reef Tract
Clonal structure analyses for the coral host revealed that the sampled colonies of D. cylindrus along the Florida Reef Tract were highly clonal when multiple colonies were present at a site. After genotyping 161 colonies from 51 different sites, we only found 56 unique multilocus genotypes. The Florida population of the coral host also had low values for genotypic richness, diversity, and evenness (Figure 2). The majority of sites contained only one colony. High genotypic diversity at a site usually occurred only when there were few colonies to sample at that site (Figure 3A). The sites where D. cylindrus was locally abundant (at least 10 colonies) have genotypic diversity values very close to zero. This indicates that within each site, the Florida coral colonies were predominantly the product of asexual reproduction. Genotypic evenness values calculated for sites with at least five genotyped samples revealed that most sites had values equal to 1 (Figure 3B). Most of these sites yielded an evenness of 1 because all sampled ramets belonged to the same genet. The host probability of identity values for all sampling regions were less than 1.0 × 10-10, indicating a low probability of misidentifying closely related individuals (full siblings) as clonemates. The probability of identity values for the symbiont were all reasonably low (Waits et al., 2001), with the exception of the Turks and Caicos Islands region (Curaçao: 0.0036, USVI: 0.015, Florida: 1.5 × 10-5, Belize: 0.003, Turks and Caicos: 0.092). Because sampling was biased toward increasing the number of unique genotypes in all regions except Florida, clonal structure could not be described elsewhere.
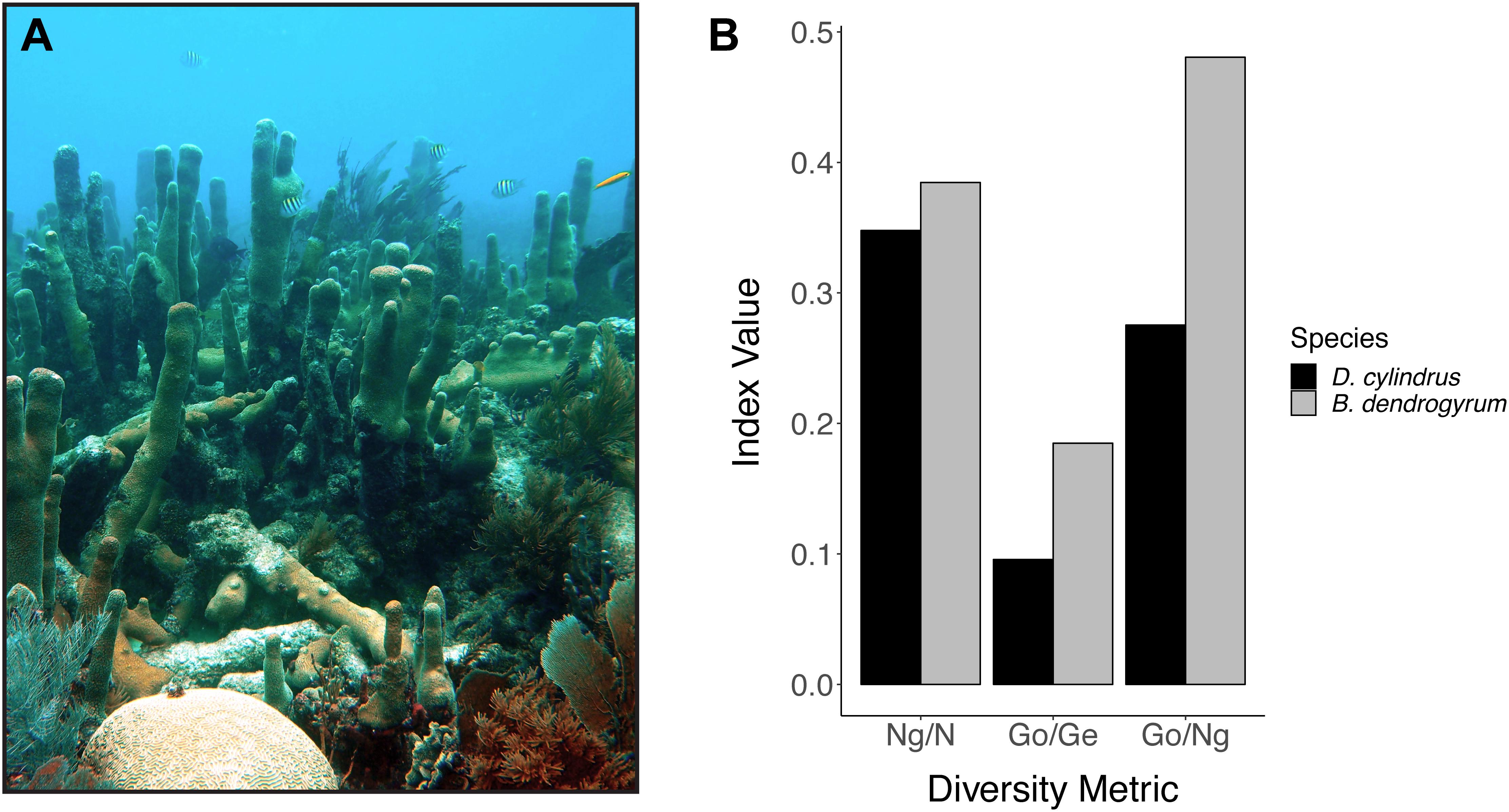
Figure 2. (A) Photo of multiple colonies of Dendrogyra cylindrus, including a recently fragmented colony with new pillar growth in the foreground (photo credit: Iliana Baums). (B) Summary of the clonal structure of Dendrogyra cylindrus and its dominant algal symbiont Breviolum dendrogyrum in Florida. Genotypic richness (Ng/N), diversity (Go/Ge), and evenness (Go/Ng) are plotted on the x-axis and the value of the diversity index is plotted on the y-axis. Ng, number of genotypes; N, number of colonies sampled; Go, observed genotypic diversity; Ge, expected genotypic diversity.
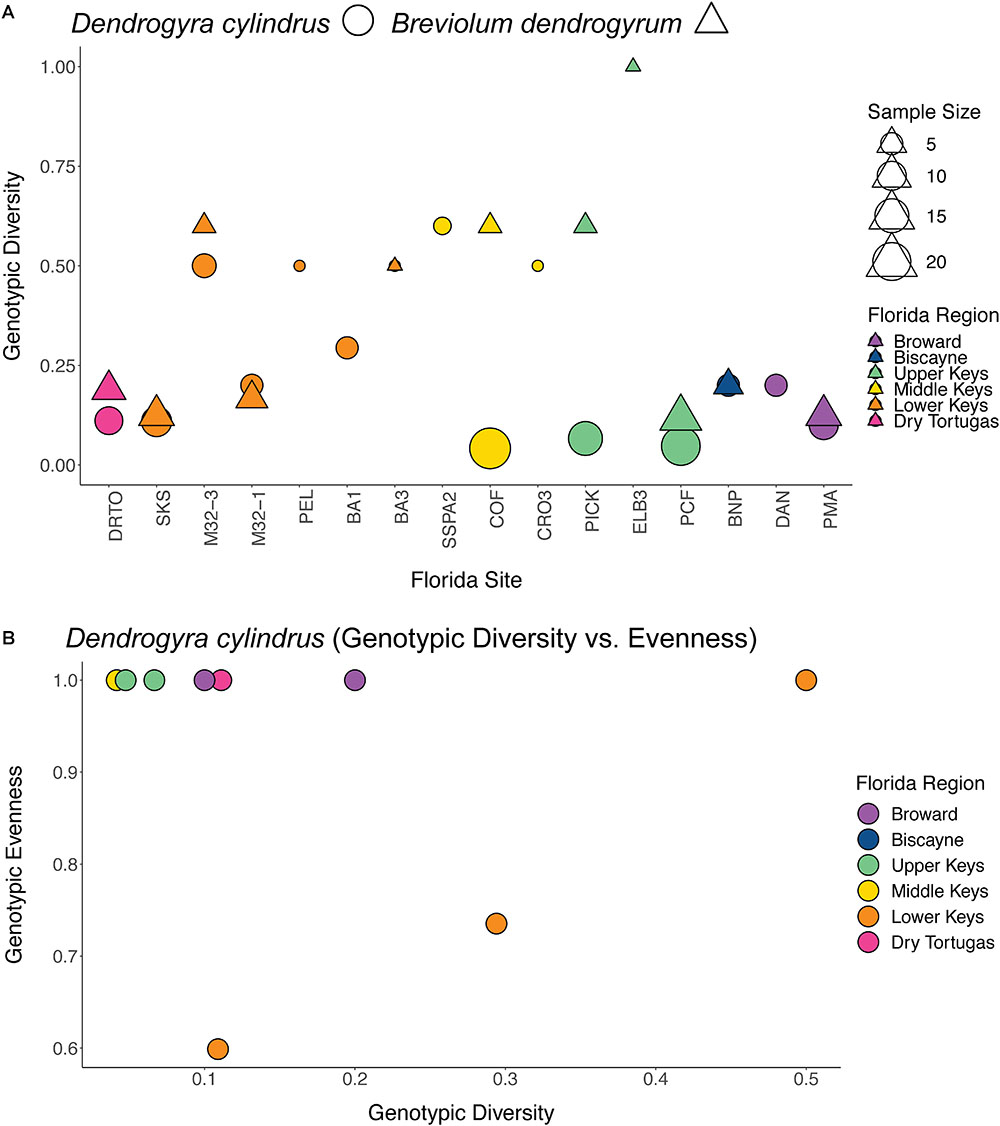
Figure 3. Plots of genotypic diversity by Florida site for the coral host (circles) and symbiont (triangles) (A), and a plot of genotypic diversity vs. evenness for the coral host (B). Only sites with multiple genotyped colonies were included, with symbols scaled to sample sizes in (A). For panel (B), only sites with at least five samples genotyped were included. Sites are colored by region as in Figure 1.
Breviolum dendrogyrum is likely haploid, and thus all samples with more than one allele were considered to be multiple infections. There was no spatial pattern in samples with single versus multiple infections along the Florida Reef Tract (Supplementary Figure 1). However, we could not assign a strain identity in samples with multiple alleles per locus or use these samples in population genetic analyses. After removing samples with multiple infections (n = 98) and samples that failed in one or more of the 15 microsatellite markers (n = 97), we obtained a subset of samples (n = 111) with complete multi-locus genotypes. Only 30 unique genotypes were found in Florida, with most strains confined to single reefs. The values for genotypic richness standardized to sample size (Ng/N), genotypic diversity (Go/Ge), and genotypic evenness (Go/Ng) were all higher in the algal symbiont compared to the host (Figure 2B). The richness values were the most similar, indicating that we found a similar number of unique genotypes of host and symbiont relative to the number of samples we genotyped for each species. The values for genotypic diversity were low in both host and symbiont (albeit, slightly higher in the symbiont), demonstrating that both species reproduced mostly asexually within a site. As was true for the coral host, due to the rarity of D. cylindrus and the frequency of multiple infections, few symbiont strains were genotyped per site. When examining genotypic diversity by site (Figure 3A), the sites with larger sample sizes had the lowest diversities. This again points to the dominance of asexual reproduction within a site. Genotypic evenness was higher in the symbiont relative to the host (Figure 2), meaning that the host population was dominated by relatively few highly replicated coral genotypes. Fewer samples with the same strain of B. dendrogyrum were identified compared to the host, partially because a strain genotype could not be called for many samples due to multiple infections and failures in one or more microsatellite markers.
For the coral host, all of the ramets belonging to the same genet were contained within a single collection site. This indicated that the dispersal abilities of asexual fragments of D. cylindrus were limited, with only one genet extending beyond 80 m (Figure 4). A single coral genet dominated most of our sites, however, three sites had two genets and one site contained three different genets (Supplementary Figure 2A).
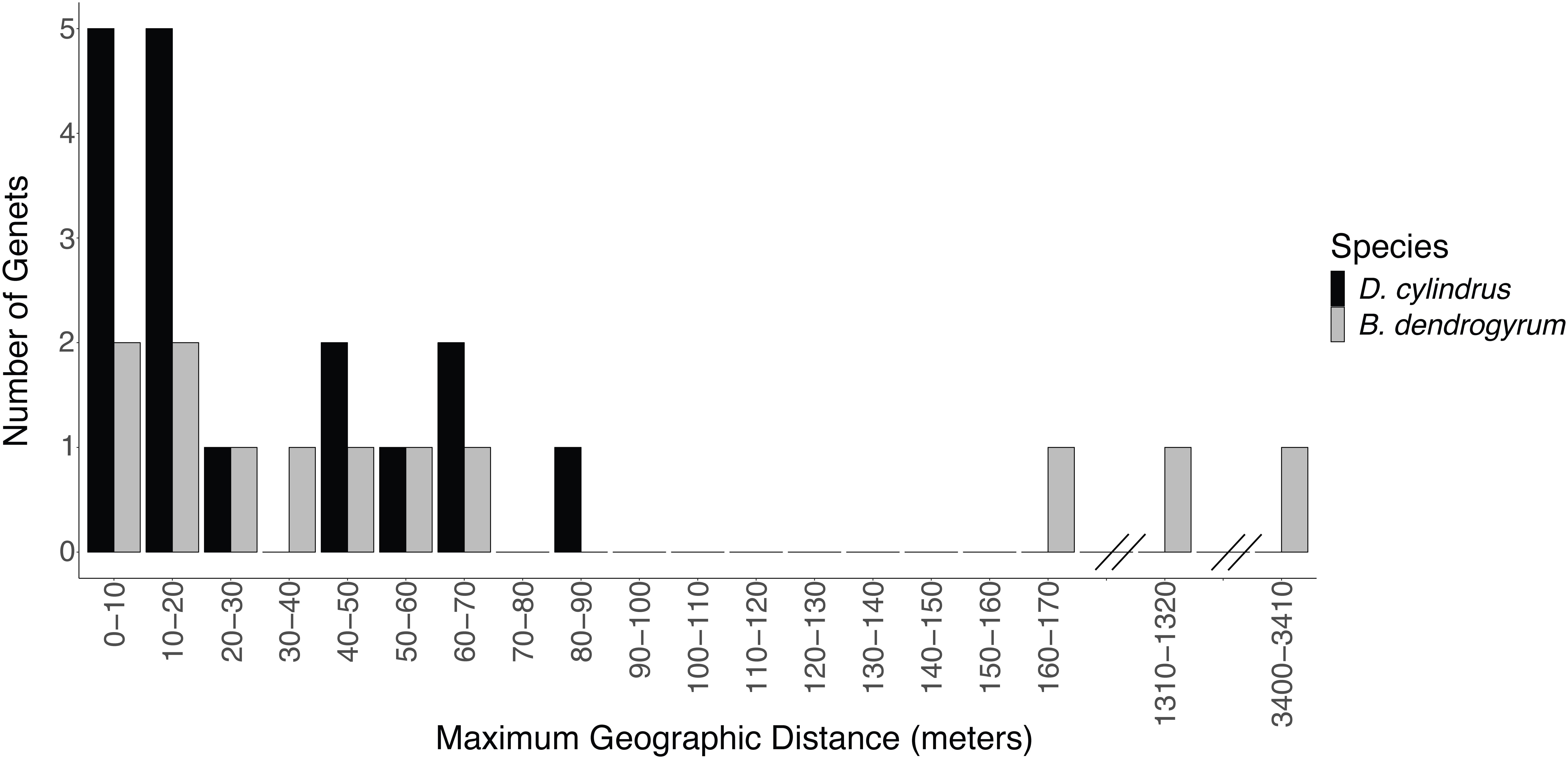
Figure 4. The maximum geographic distance between ramets of the same genet of Dendrogyra cylindrus (black bars) and Breviolum dendrogyrum (gray bars). While no coral genets were shared between sites, symbiont strains were occasionally shared between sites. Note the discontinuous x-axis.
A test of isolation by distance including only genets of D. cylindrus along the Florida Reef Tract revealed no significant correlation between genetic and geographic distance (r2 = 0.0001). Spatial autocorrelation analysis using the complete dataset of D. cylindrus genotypes (including clones) in Florida (n = 180) revealed significant positive spatial autocorrelation up to distances of 60 m (Supplementary Figure 3A). However, when clones were removed, there was no significant spatial autocorrelation (Supplementary Figure 3B). These results indicate that the cause of the correlation between genetic distance and geographic distance over small distance classes is a result of asexual reproduction, likely via fragmentation, in D. cylindrus.
The genotype of the algal symbiont often corresponded with the genotype of the coral host when multiple colonies were present at a site, meaning that all of the ramets belonging to the same host genet were often symbiotic with the same clonal strain of B. dendrogyrum (Figure 5). There were exceptions to this observation. For example, in the Pillar Coral Forest site in the Upper Keys, all of the colonies sampled were ramets of the same coral host genet. However, while 10 ramets associated with identical B. dendrogyrum strains, two ramets associated with distinct strains of B. dendrogyrum (Figure 5C and Supplementary Figure 2). The reverse was also observed, with the same symbiont strain found associated with different coral genets, although there was only one example of this pattern found within a site (Figure 5B). More often, the same symbiont strain was found in different coral genets at different sites, with dispersal distances ranging from tens of meters to a maximum of 3.44 km (Figure 4). However, no symbiont strains were shared between Florida regions (Figure 1B). We also sampled the same coral colony in different locations (e.g., top, middle, and base) to assess symbiont diversity. Of the five colonies that were sampled in this way and successfully genotyped, only one contained two different symbiont strains.
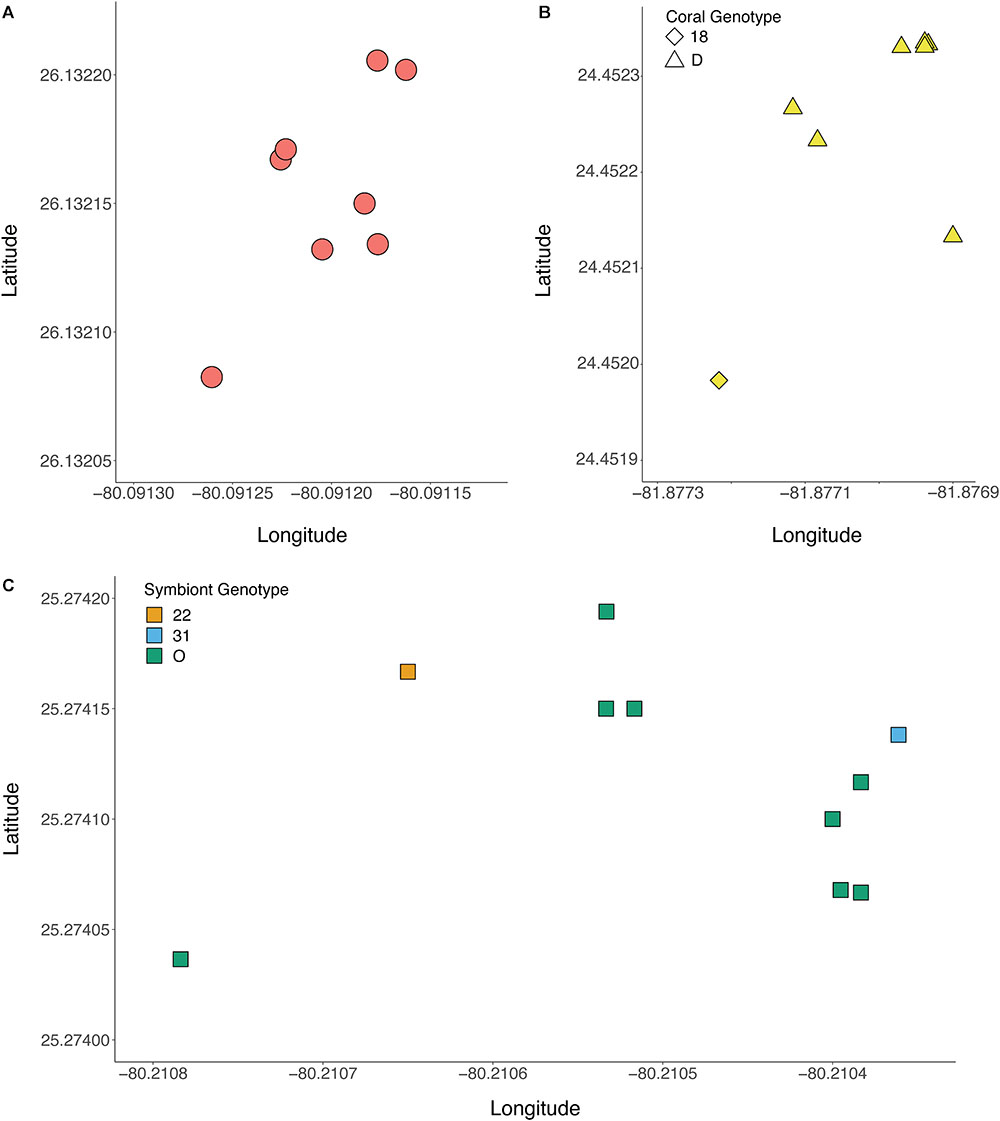
Figure 5. Representative plots showing the spatial arrangement of genotyped colonies of D. cylindrus and associated strains of B. dendrogyrum. Within a site, most ramets of a coral genet associated with the same symbiont strain. (A) Pacific Mako Anchorage, a site in Broward County, (B) Sand Key Spa, a site in the Lower Keys, (C) Pillar Coral Forest, a site in the Upper Keys. The symbol shape corresponds to the coral genotype, and the color corresponds to the symbiont genotype.
Population Structure Analyses
Strong genetic differentiation between populations was found, with a significant global FST value of 0.110 and an F’ST value of 0.414. Bayesian clustering using the entire dataset of unique multilocus genotypes for the coral host revealed the presence of two to four clusters in the dataset. The different estimators for the optimum number of populations yielded different results. For example, the ΔK method (Evanno et al., 2005) yielded a K of 2 and the MaxMeaK with a spurious cluster threshold of 0.5 (a less conservative measurement) yielded a K of 4. In the STRUCTURE plot for K = 2, all of the Florida, Belize, and Turks and Caicos samples have a high probability of membership to the first cluster and all of the Curaçao and U.S. Virgin Islands samples have a high probability of membership to the second cluster (Figure 6A). When the same dataset was run in STRUCTURE setting K = 3 as the a priori number of populations in the dataset, an additional cluster corresponding to the individuals from Curaçao separates from the U.S. Virgin Islands (Figure 6B). The samples from the Turks and Caicos Islands also appear to be admixed. However, our dataset contained locations that were sampled unevenly. Thus, we randomly selected unique multilocus genotypes to a maximum size of 20 to remove the bias associated with uneven sampling (Puechmaille, 2016). When Bayesian clustering analysis was completed for this subsampled dataset, all of the K estimators yielded an optimal cluster number of three. The only exception was the ΔK method, which still indicated that there were two populations.
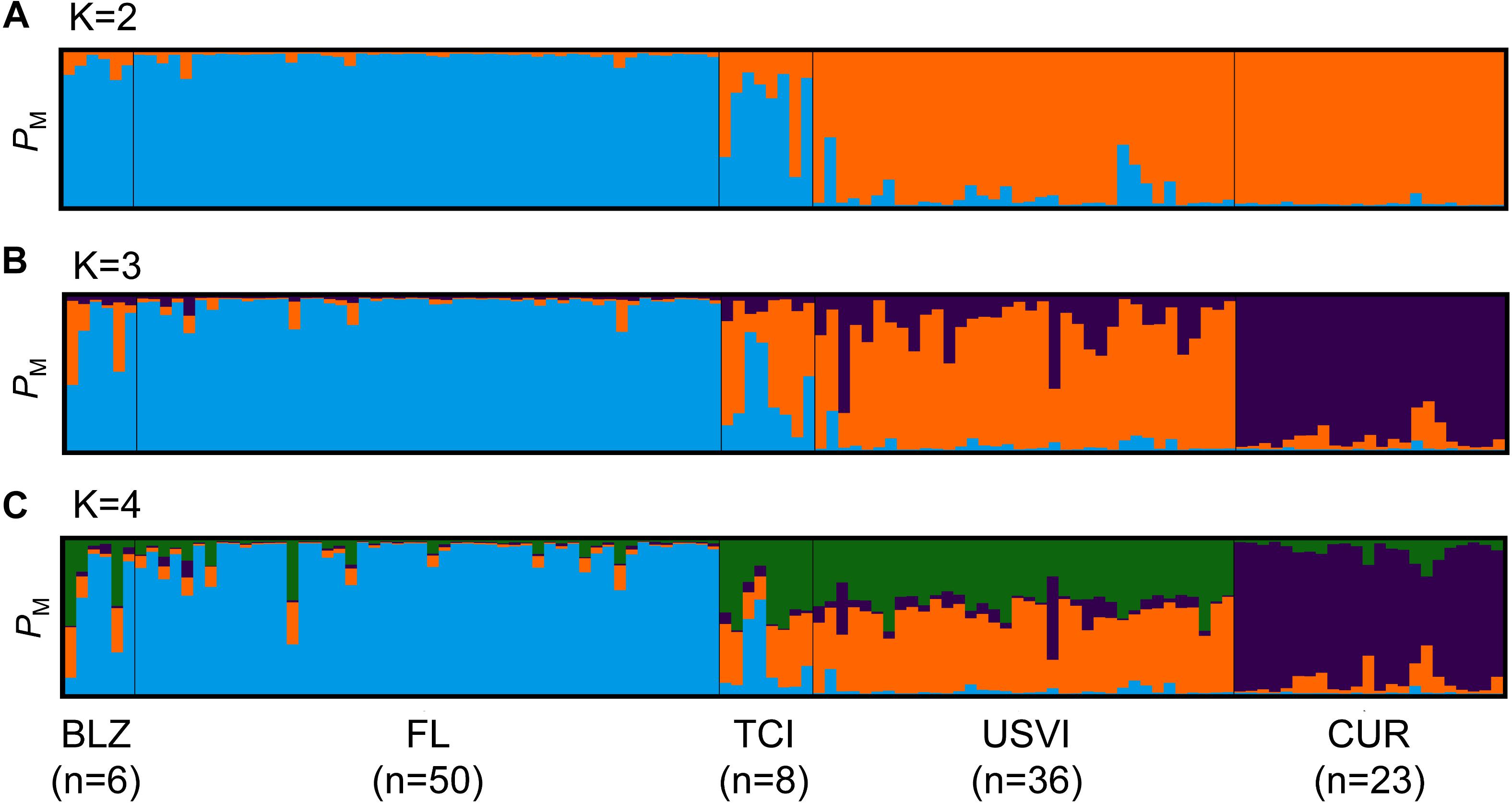
Figure 6. Population structure results for Dendrogyra cylindrus. Each vertical bar represents a unique multilocus genotype. Probability of membership to a cluster is plotted on the y-axis. (A) STRUCTURE results for D. cylindrus revealed two separate clusters (K = 2, indicated by color). (B) STRUCTURE results for D. cylindrus when K = 3. (C) STRUCTURE results for D. cylindrus when K = 4. Individuals are grouped by sample location; abbreviations are the same as in Figure 1.
An AMOVA corroborated these results, with the largest pairwise FST value between Florida and Curaçao (FST = 0.184, p-value < 0.001, Table 2). The U.S. Virgin Islands and Curaçao were significantly differentiated, although less strongly (FST = 0.045, p-value < 0.001). The Turks and Caicos Islands were equally differentiated from Florida and the U.S. Virgin Islands (FST = 0.046, p-value < 0.001), further supporting that this is likely an area of admixture (Table 2).
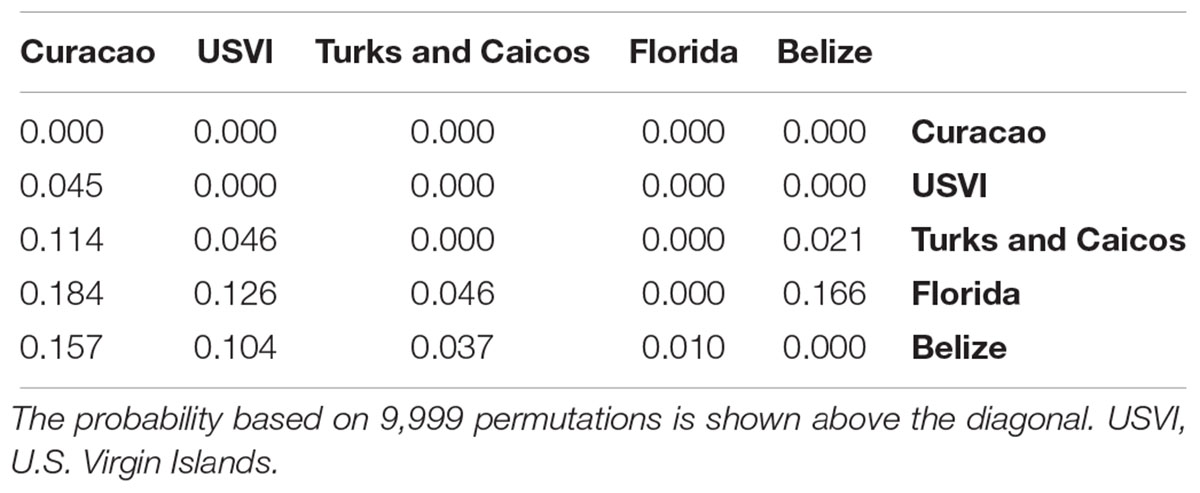
Table 2. Pairwise FST values between Dendrogyra cylindrus samples from five Caribbean regions (below diagonal).
We did not find significant population structure within Florida for the coral host. POWSIM v. 4.1 with 1000 simulation runs using the allele frequencies from our host marker set was used to test whether this result was due to a lack of power. Fisher’s exact test revealed a high statistical power (1-β = 0.97) of detecting an FST value of 0.0195, and a low type I error rate (α = 0.0410). Thus, our sample size of 50 individuals from Florida was adequate to detect low levels of population structure along the Florida Reef Tract (FRT) if present.
Population structure analyses were also conducted for B. dendrogyrum. Only 58 unique multilocus genotypes were found, after removing samples with multiple infections. Cluster analysis using STRUCTURE and a consideration of multiple estimators for the optimum number of populations revealed that the likely number of populations in the dataset is between 2 and 4 (Figure 7). However, there were locations in this dataset that were sampled more intensely than others, and consequentially we subsampled the locations with the largest sample size to a maximum value of 19. Because we found population structure for B. dendrogyrum within Florida, we subsampled by the six regions (Figure 1). Most of the K estimators indicated that there were likely four populations in the dataset. When the threshold for spurious clusters was set to a strict 0.8, some of the K estimators yielded a K of 2. The ΔK method alone indicated that the optimal number of populations was three. An AMOVA for the haploid B. dendrogyrum revealed that the highest amount of between-group variation was between sample locations (ϕPT = 0.513, p-value = 0.001).
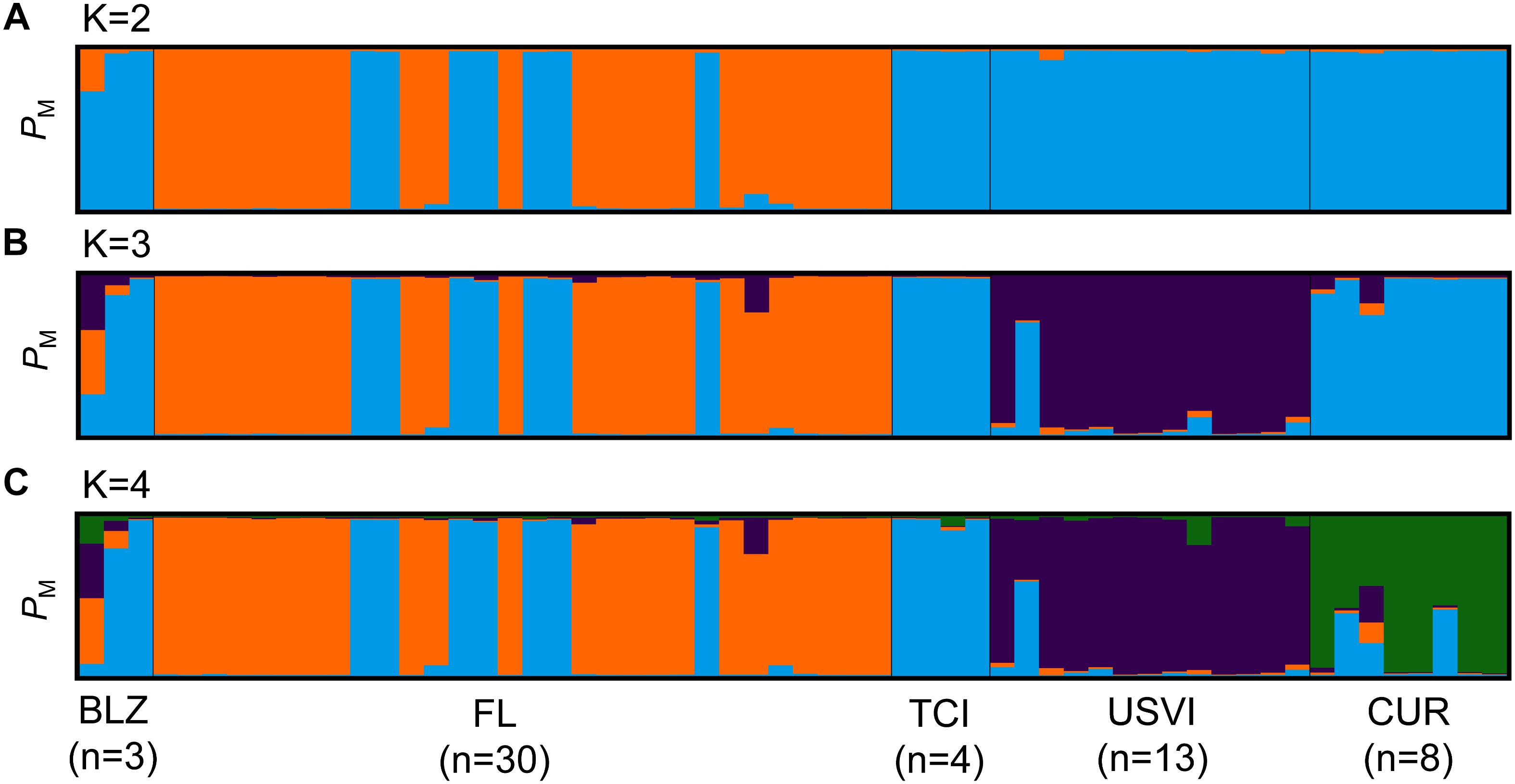
Figure 7. Population structure results for Breviolum dendrogyrum. Each vertical bar represents a unique multilocus genotype. Probability of membership to a cluster is plotted on the y-axis. (A) STRUCTURE results for B. dendrogyrum when K = 2. (B) STRUCTURE results when K was set to 3. (C) STRUCTURE results when K was set to 4. Individuals are grouped by sample location; abbreviations are the same as in Figure 1.
These results indicate that the population structure of the algal symbiont did not match the population structure of the host. Notably, there was population structure within the FRT for the symbiont that was not found in the host. Some samples from Florida clustered more strongly with samples from Belize and the Turks and Caicos Islands than other Florida samples. Because this pattern of population assignment did not correspond with geography, four samples from the unique Florida cluster and three samples from Florida that clustered with the rest of the Caribbean were sequenced using microsatellite flanker sequences (Si15) to confirm that they all belonged to the same species. Comparing the Si15 sequences to representative sequences from Symbiodinium Clade B (Finney et al., 2010) revealed that samples from the two clusters within Florida are indeed the same species (B. dendrogyrum).
Historical and Future Changes in Pillar Coral Population Sizes
Demographic modeling results revealed no evidence for past changes in pillar coral population size (Figure 8). In both the one continuous population size change (OnePopVarSize) and the two population size changes (OnePopFounderFlush) models, the upper limit of the 95% confidence intervals for the ancestral population size (2Nancμ, both models) and the founder population size (2Nfounderμ, OnePopFounderFlush model only) were undefined. The undefined upper limit meant that many possible population sizes for the ancestral and founder population were equally likely, including population sizes that were both larger and smaller than the 95% confidence interval for the current population size (2Nμ). This was further evidenced by the broad profile likelihood ratios for the ancestral and founder population sizes (Figure 8), and the population size ratios that included the value 1, indicating no significant change. To decide which of the three models best described our data, we used the Akaike Information Criterion (AIC) calculated based on the log likelihood and the number of parameters in each model (Anderson, 2007). The AIC values for the OnePop (798.94), OnePopVarSize (799.56), and OnePopFounderFlush (800.6) models were very close to one another. Because lower AIC values indicate better model fit, we conclude that the OnePop model with no past change in population size is the best model. However, the differences in AIC between models were very small, and thus it is possible that all three models do not accurately describe reality. Still, the results from the models assuming past demographic events failed to detect significant changes in population size, supporting our conclusion of no past changes in D. cylindrus populations.
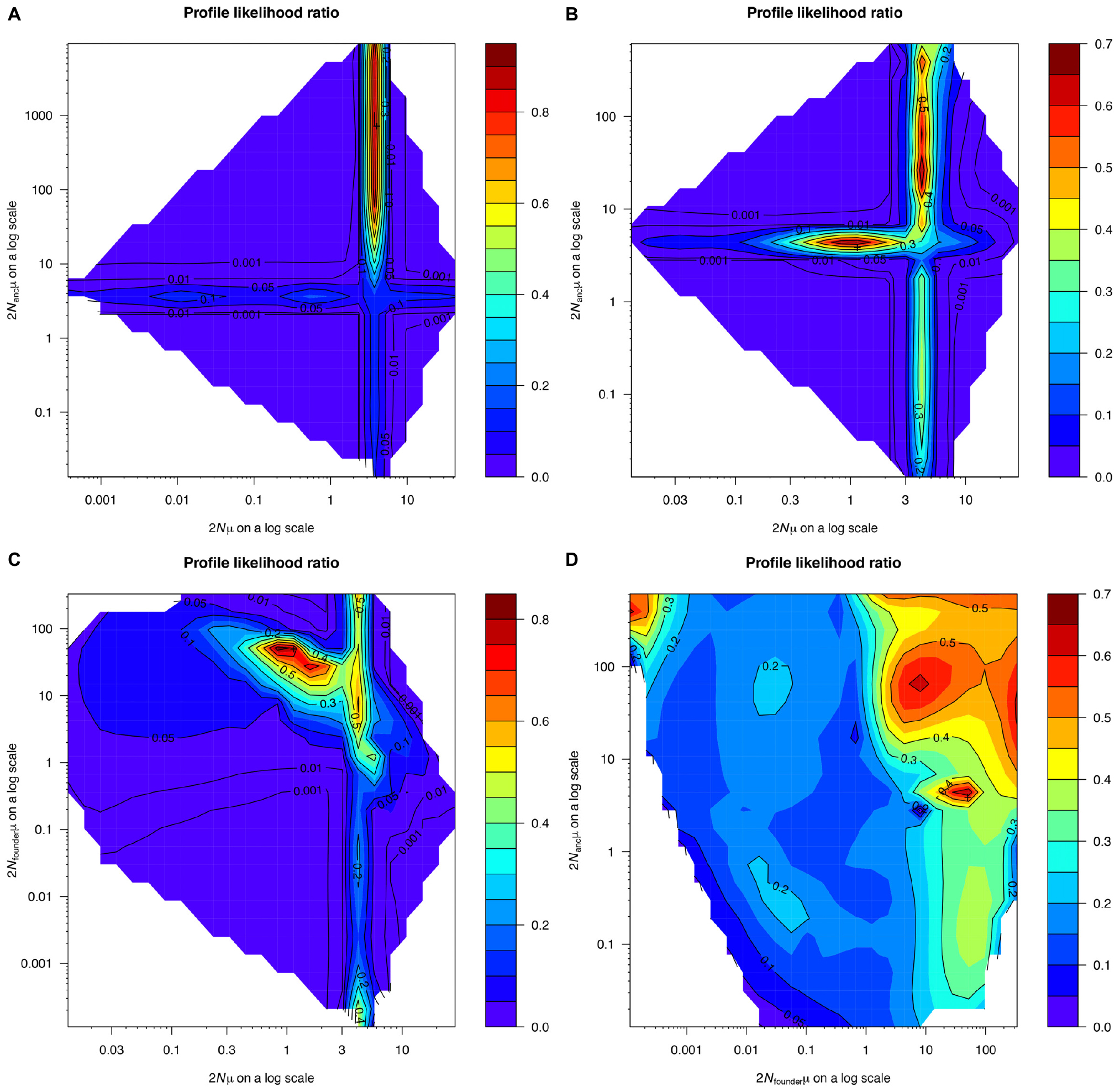
Figure 8. Profile likelihood ratio of the current population size of Dendrogyra cylindrus (x-axis) and the ancestral population size of D. cylindrus (y-axis) from the OnePopVarSize model implemented using the software MIGRAINE (A). Profile likelihood ratio plots for the OnePopFounderFlush model are shown for the current and ancestral populations (B), for the current and founder populations (C), and for the founder and ancestral populations (D).
The three projected rates of decline yielded a range of times to extinction for D. cylindrus. In the most conservative simulated scenario where 80% of colonies survived each hyperthermal event, it took 31 stress events for extirpation of D. cylindrus from the FRT. With 50% and 20% survival, it took 11 and 6 stress events, respectively, for local extinction to occur (Supplementary Figure 4). Because some coral colonies are ramets of the same genet, and these coral ramets often contain clonal strains of B. dendrogyrum, coral colonies are lost at a faster rate than coral genotypes and symbiont strains (Supplementary Figure 4). We then translated our results into years using our estimated frequencies of future hyperthermal stress events. If the rate of survival per thermal stress event throughout the Florida Reef Tract is 80% of colonies, all Florida pillar corals would be lost by 2066. Assuming 50% or 20% survival, we would expect D. cylindrus to become extinct in Florida in 2046 and 2039, respectively.
Discussion
The pillar coral, Dendrogyra cylindrus, and its obligate algal symbiont Breviolum dendrogyrum reproduce mostly via asexual processes within a site, with strains of the algae dispersing over much larger distances than host fragments (Figure 4). Barriers to gene flow were also not congruent between the partners but showed similarities with other coral–algal symbioses in the Caribbean (Baums et al., 2014b). As is often the case for corals (Baums, 2008), while asexual reproduction was prevalent, no signs of inbreeding were detected in the coral host and allelic diversity was greater than or comparable to other Caribbean reef-building coral species (Baums et al., 2005a; Baums et al., 2010; Rippe et al., 2017). Demographic modeling based on molecular data agreed with the geological record that D. cylindrus was historically not a dominant species on Caribbean reefs and yet was able to survive as the sole remaining species in the genus. The forecasted increasing frequency of extreme warm water events in combination with associated disease outbreaks, and the observed absence of sexual recruitment in this species, project a high likelihood that D. cylindrus will become locally extinct in the Florida Keys in modern times (see below). The consequences of the loss of rare coral species and their symbionts are unknown but evidence from other ecosystems indicates that losing rare species can destabilize communities and degrade ecosystem function (Theodose et al., 1996; Lyons and Schwartz, 2001; Mouillot et al., 2014).
Clonal Structure and Diversity
Clonal diversity indices for D. cylindrus and B. dendrogyrum in Florida were more typical of an asexual population (closer to 0) than a sexual one (closer to 1, Figure 2). Florida sites that contained multiple colonies of D. cylindrus were highly clonal, showed positive spatial autocorrelation, and had low genotypic diversity, indicating that the primary mode of reproduction over this scale is asexual fragmentation (Figures 2, 3 and Supplementary Figure 3). The prevalence of asexual reproduction in the elkhorn coral, Acropora palmata, was attributed to increased physical retention of fragments in certain habitats and low rates of sexual recruitment (Baums et al., 2006). Similar forces, as well as fragmentation from hurricanes, are likely responsible for high asexual reproduction in Florida D. cylindrus. This study was limited to assessing clonal structure in Florida alone, and thus future work should measure genotypic richness, diversity, and evenness in other locations.
Low genotypic diversity as a result of clonal reproduction is often identified in foundation species (Baums et al., 2006; Dubé et al., 2017; Gélin et al., 2017; Miller et al., 2018). Clonal reproduction can maintain high population densities in the absence of sexual reproduction (i.e., a storage effect) by increasing the number of established adults (Warner and Chesson, 1985). In benthic surveys of nine inshore to offshore sites in Key Largo, Florida, no juvenile colonies of D. cylindrus were discovered (Miller et al., 2010), indicating successful sexual recruitment of juvenile pillar corals is non-existent or very low. In addition, the Florida population of D. cylindrus is at the northern edge of this species’ range. Previous work on clonal plants and corals has pointed to the prominence of asexual reproduction at the edge of species’ geographical ranges, enabling marginal populations to persist despite low sexual recruitment (Silvertown, 2008; Boulay et al., 2014). Thus, recent reproduction in the Florida D. cylindrus population may be entirely asexual due to the lack of sexual recruits from neighboring populations and from within Florida.
The absence of successful sexual reproduction in D. cylindrus raises concerns about the longevity of this species, particularly in Florida. Because most sites tended to be occupied by a single coral genotype (Figure 5 and Supplementary Figure 2A), it is possible that Florida D. cylindrus are already experiencing an Allee effect, wherein the density of compatible colonies is too low for successful sexual reproduction and population growth (Courchamp et al., 1999). Recent work describing instances of hermaphroditism in D. cylindrus are promising (Neely et al., 2018), and could increase local larval production if D. cylindrus is self-compatible. Despite the prevalence of asexual reproduction and the small census size, heterozygosity levels remain high, indicating that D. cylindrus is not inbred. Future studies of controlled crosses between different D. cylindrus genets and opposite sex ramets of the same genet are necessary to discern compatibility in this species (Baums et al., 2013).
The Florida population of B. dendrogyrum is also clonal, with the same symbiont strain often found in all of the ramets of a coral genet (Figure 5). Despite the physical damage and transportation of fragments to lower light micro-environments when pillars fragment and fall over, the association between a particular coral genotype and specific symbiont strain appears to be mostly stable. High fidelity between individual genotypes of symbiotic partners has been found previously in the Caribbean elkhorn coral (Acropora palmata) and its algal symbiont (Symbiodinium ‘fitti’) (Baums et al., 2014b). In this system, different combinations of host genotypes with a symbiont strain were shown to possess distinct physiological responses to heat stress (Parkinson et al., 2015) but this is not always the case (Parkinson et al., 2018). It is unknown if different combinations of D. cylindrus genotypes and B. dendrogyrum strains could result in functional variation of the holobiont.
While coral genets were restricted to one site each, four pairs of sites did share symbiont strains. The symbiont is able to disperse asexually over larger distances than its coral host, although successful long-distance dispersal appears to be rare (Figure 4). A typical fragment dispersal distance for D. cylindrus is less than 60 m, as evidenced by the spatial autocorrelation analysis (see below and Supplementary Figure 3), although one genet extended over 80 m (Figure 4). Similarly, the asexual dispersal ability of Symbiodinium fitti was found to be significantly higher than its coral host, A. palmata (Baums et al., 2014b). However, the farthest asexual dispersal distance for B. dendrogyrum (3,440 m) was higher than what was found for S. fitti (greater than 2,000 m). While Symbiodiniaceae cells are constantly being expelled from the coral host, little is known about the persistence of their free-living stage (Thornhill et al., 2017). Despite similar morphologies, different species of Symbiodiniaceae may differ greatly in their abilities to disperse and infect new host corals.
While the storage effect provided by asexually produced colonies can be beneficial in the absence of environmental change, populations of foundation species with low genotypic diversity are more susceptible to disturbances. Increasing genotypic diversity in plots of the seagrass, Zostera marina, yielded higher resistance to disturbance by grazing geese (Hughes and Stachowicz, 2004). High intraspecies variation in crop plants increases plant fitness and agricultural yields, and decreases susceptibility to insect pests (Tooker and Frank, 2012). In addition, high genotypic diversity confers a greater ability for populations to recover from warm temperature stress events (Reusch et al., 2005), which are expected to become more frequent with climate change (Stocker et al., 2013).
Barriers to Gene Flow
Population structure results for D. cylindrus indicate that there are at least two genetic breaks in the Caribbean, resulting in three populations (K of 3 chosen by most K estimators). The first population includes samples from Florida, Belize, and the Turks and Caicos Islands, the second population includes samples from the U.S. Virgin Islands, and the third population comprises samples from Curaçao (Figure 6). Cluster analysis results and pairwise FST analyses revealed that there is little to no gene flow between Florida and either Curaçao or the USVI. A genetic break in this area has been well characterized in other corals (Vollmer and Palumbi, 2006; Baums et al., 2010; Foster et al., 2012; Andras et al., 2013; Rippe et al., 2017).
Dendrogyra cylindrus lacks population structure along the Florida Reef Tract, which was also found in both A. cervicornis and A. palmata using microsatellite markers (Baums et al., 2005b, 2010). POWSIM simulations revealed our set of microsatellite markers had sufficient power to detect an FST value as low as 0.0195. In contrast to the microsatellite results, an analysis of Florida A. cervicornis using SNPs found significant population structure within the Florida Reef Tract (Willing et al., 2012), and thus it is possible that higher resolution markers may resolve weaker population differentiation in D. cylindrus.
The genetic break between D. cylindrus in the USVI and Curaçao was weaker than the break between Florida and these two locations but again consistent with a similar genetic break in A. palmata (Devlin-Durante and Baums, 2017). It is possible that the Caribbean Current is restricting coral larval exchange between these two islands (Figure 1A). This current was shown to affect dispersal in other planktonic marine species (Díaz-Ferguson et al., 2010; Jossart et al., 2017), however, population genetic data for other coral species does not show a break in this area (Andras et al., 2013; Rippe et al., 2017).
After correcting for uneven sampling effort, the results for B. dendrogyrum population structure revealed four populations in the Caribbean (Figure 7). B. dendrogyrum strains in both Curaçao and the U.S. Virgin Islands separate as two populations distinct from the rest of the Caribbean. These genetic breaks are in concordance with the breaks observed in the host coral, D. cylindrus. However, there is population structure within the FRT for the algal symbiont, with some Florida individuals clustering with Belize and the Turks and Caicos Islands. A similar incongruence between the population structure of host and symbiont was found in G. ventalina (Andras et al., 2011, 2013) and A. palmata (Baums et al., 2005b, 2014b). A recent review explained the observed lower connectivity in horizontally transmitted algal symbionts relative to their coral hosts as potentially resulting from restricted dispersal as well as from resident symbionts outcompeting migrants (Thornhill et al., 2017). The inconsistency in population structure between host and symbiont supports that D. cylindrus obtains its symbionts horizontally (from the surrounding environment) and implies that gene flow occurs over different spatial scales in the partners (Baums et al., 2014b). Because such a small sample of symbiont strains was included from both Belize and the Turks and Caicos Islands, it is possible that the cluster containing these samples and some Florida samples is an artifact of STRUCTURE. Underrepresented populations can be incorrectly merged together by STRUCTURE (Puechmaille, 2016), although subsampling our larger populations and using additional K estimators reduced the uneven sampling bias.
Demographic Modeling of Pillar Coral Populations
Demographic modeling results using MIGRAINE revealed no evidence for past changes in population size in Florida D. cylindrus. This is consistent with previous descriptions of D. cylindrus as a naturally rare species (NOAA, 2014), and with the rarity of D. cylindrus in the fossil record (see section “Introduction”). Abundance can be high in localized areas, however, due to asexual fragmentation. This suggests that historically, D. cylindrus did not experience a range-wide decline since its first appearance in the fossil record during the late Pliocene/early Pleistocene (Budd, 2000).
The results from our forward projection model demonstrate that the increased frequencies of hyperthermal stress events will likely lead to extirpation of D. cylindrus in Florida in the near future. While these are simple projections of potential species decline that do not take into account the heterogeneity of abiotic and biotic factors along the Florida Reef Tract, they are still useful for demonstrating the severity of the current situation for pillar corals. Detailed demographic information about the actual loss of Florida D. cylindrus colonies, genotypes, and B. dendrogyrum strains will be crucial for successful species rehabilitation. If no active management strategies such as nursery rearing and propagation are enacted to enhance sexual reproduction and increase genotypic diversity (National Academies of Sciences Engineering and Medicine, 2018), we can expect pillar corals to disappear from Florida within just a few decades. Global change will continue to alter the composition of reefs if greenhouse gas emissions are not curbed.
The Consequences of Rare Species Loss
What are the consequences of losing rare species? Experimental removal of rare species was shown to reduce ecosystem resistance to invasion by an exotic grass (Lyons and Schwartz, 2001). It is possible that less common species significantly contribute to the proper maintenance of ecosystem function. Rare plant species were shown to more significantly impact nutrient cycling and retention in an alpine meadow compared to more abundant species (Theodose et al., 1996). In a comprehensive study of species occurrence datasets from coral reefs, alpine meadows, and tropical forests, rare species were repeatedly shown to predominantly support vulnerable functions (Mouillot et al., 2013). These vulnerable functions were defined as ecosystem roles with low redundancy, in that uncommon species with distinct trait combinations bolstered these particular functions. Despite the high diversity in these ecosystems, abundant species did not insure against the services lost by removing rare species (Mouillot et al., 2014). Therefore, maintaining these uncommon species is essential to overall community functional diversity.
When two rare species are combined in an obligate symbiosis, then the loss of one species would yield coextinction of the other (Koh et al., 2004). It is possible that D. cylindrus sexual recruits can survive by associating with B. meandrinium in the absence of B. dendrogyrum but the latter has never been found in another coral host species (Lewis et al., 2018). It is unknown whether there are free-living strains of B. dendrogyrum. Determining the degree to which the association is obligate for the host-specialist symbiont is beyond the scope of this study. Likewise, the functional roles of this rare coral–algal symbiosis are unknown, although the tall pillar morphology of this species may promote fish aggregating behavior (Shantz et al., 2015). Without knowing the ecological function of rare marine symbioses such as the one between D. cylindrus and B. dendrogyrum, it is prudent to assume important ecosystem contributions when forming conservation strategies (Lyons et al., 2005).
Continued unprecedented global change will cause further loss of biodiversity. Because species that are both rare and specialized are especially at risk of disappearing (Davies et al., 2004), studies of the ecology and evolution of these species are particularly timely and important. Recent work on D. cylindrus (Neely et al., unpublished data), including this study, has highlighted a lack of recent successful sexual recruitment. The ability of coral species to survive climate change hinges upon ongoing sexual reproduction, enabling selection for more resilient genotypes (Matz et al., 2018). Adaptation is thus extremely unlikely in D. cylindrus, necessitating continued ex situ efforts to understand spawning, larval development, and larval settlement in this unique Caribbean coral species (Marhaver et al., 2015). Additionally, more drastic measures, such as assisted gene flow, could become necessary (National Academies of Sciences Engineering and Medicine, 2018).
Coral reefs are currently experiencing global decline due to the loss of important herbivores, pollution and nutrient runoff, disease, and climate change (Bruno et al., 2007; Hoegh-Guldberg et al., 2007; Jackson et al., 2014). Since the 1980s, coral cover on Caribbean reefs has already declined by an average of 80% (Gardner et al., 2003). Losing rare species and their corresponding functions may further reduce reef resilience and thus exacerbate coral reef decline beyond what has been predicted (Hoegh-Guldberg et al., 2017; Hughes et al., 2017).
Ethics Statement
This study was carried out in accordance with the recommendations of Convention on International Trade in Endangered Species of Wild Fauna and Flora Resolution Conf. 11.10 (Rev. CoP15). Coral samples were collected under Florida Keys National Marine Sanctuary permits FKNMS-2013-085-A1, FKNMS-2014-004-A1, FKNMS-2016-062, CITES AN001.12US784243/g, and Indigenous Species Research, Retention, and Export permit DFW16005T.
Author Contributions
AC performed all molecular work and data analysis, and wrote the manuscript. CL and KN collected samples and edited the manuscript. IB directed the study, obtained funding, provided laboratory space, collected samples, and wrote the manuscript.
Funding
This work was supported by the Florida Fish and Wildlife Conservation Commission’s Program, Florida’s Wildlife Legacy Initiative, and the United States Fish and Wildlife Service’s State Wildlife Grants Program (Marine Projects 2012, T-32). AC was supported by the National Science Foundation (NSF) Graduate Research Fellowship Program (Grant No. DGE1255832). The conclusions are those of the authors and do not necessarily reflect the views of the NSF.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank L. Kabay, K. Macaulay, L. Carne, and the Keys Marine Lab AAUS staff divers for outstanding field support. M. Rodriguez-Lanetty (Florida International University) provided some funding and supplies for field sampling. We also thank the staff at the Penn State Genomics Core Facility for running samples as well as M. Devlin-Durante and K. Lunz for help with obtaining funds. We thank S. Kitchen, J. Keller, and R. Leblois for valuable help with bioinformatics analyses. Computational analyses were conducted on the PSU Institute for CyberScience Advanced CyberInfrastructure high-performance computer cluster. This manuscript has been released as a pre-print at bioRxiv (Chan et al., 2018).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2019.00218/full#supplementary-material
Footnotes
- ^paleobiodb.org
- ^https://scholarsphere.psu.edu/concern/parent/31z40kt37x/file_sets/x6969z327w
- ^www.multiplexmanager.com
- ^http://kimura.univ-montp2.fr/∼rousset/Migraine.htm
- ^http://coralreefwatch.noaa.gov
References
Anderson, D. R. (2007). Model Based Inference in the Life Sciences: a Primer on Evidence. Berlin: Springer Science+Business Media.
Andras, J. P., Kirk, N. L., Coffroth, M. A., and Harvell, C. (2009). Isolation and characterization of microsatellite loci in Symbiodinium B1/B184, the dinoflagellate symbiont of the Caribbean sea fan coral, Gorgonia ventalina. Mol. Ecol. Resour. 9, 989–993. doi: 10.1111/j.1755-0998.2009.02549.x
Andras, J. P., Kirk, N. L., and Harvell, C. D. (2011). Range-wide population genetic structure of Symbiodinium associated with the Caribbean sea fan coral, Gorgonia ventalina. Mol. Ecol. 20, 2525–2542. doi: 10.1111/j.1365-294X.2011.05115.x
Andras, J. P., Rypien, K. L., and Harvell, C. D. (2013). Range-wide population genetic structure of the Caribbean sea fan coral, Gorgonia ventalina. Mol. Ecol. 22, 56–73. doi: 10.1111/mec.12104
Aronson, R., Bruckner, A., Moore, J., Precht, B., and Weil, E. (2008). Dendrogyra cylindrus. The IUCN Red List Threat. Species 2008:e.T133124A3582471. doi: 10.2305/IUCN.UK.2008.RLTS.T133124A3582471.en
Baums, I. B. (2008). A restoration genetics guide for coral reef conservation. Mol. Ecol. 17, 2796–2811. doi: 10.1111/j.1365-294X.2008.03787.x
Baums, I. B., Devlin-Durante, M. K., Laing, B. A., Feingold, J., Smith, T., Bruckner, A., et al. (2014a). Marginal coral populations: the densest known aggregation of Pocillopora in the Galápagos Archipelago is of asexual origin. Front. Mar. Sci. 1:59. doi: 10.3389/fmars.2014.00059
Baums, I. B., Devlin-Durante, M. K., and LaJeunesse, T. C. (2014b). New insights into the dynamics between reef corals and their associated dinoflagellate endosymbionts from population genetic studies. Mol. Ecol. 23, 4203–4215. doi: 10.1111/mec.12788
Baums, I. B., Devlin-Durante, M., Polato, N., Xu, D., Giri, S., Altman, N., et al. (2013). Genotypic variation influences reproductive success and thermal stress tolerance in the reef building coral, Acropora palmata. Coral Reefs 32, 703–717. doi: 10.1007/s00338-013-1012-6
Baums, I. B., Hughes, C. R., and Hellberg, M. E. (2005a). Mendelian microsatellite loci for the Caribbean coral Acropora palmata. Mar. Ecol. Prog. Ser. 288, 115–127. doi: 10.3354/meps288115
Baums, I. B., Miller, M. W., and Hellberg, M. E. (2005b). Regionally isolated populations of an imperiled Caribbean coral, Acropora palmata. Mol. Ecol. 14, 1377–1390. doi: 10.1111/j.1365-294x.2005.02489.x
Baums, I. B., Johnson, M., Devlin-Durante, M., and Miller, M. (2010). Host population genetic structure and zooxanthellae diversity of two reef-building coral species along the Florida Reef Tract and wider Caribbean. Coral Reefs 29, 835–842. doi: 10.1007/s00338-010-0645-y
Baums, I. B., Miller, M. W., and Hellberg, M. E. (2006). Geographic variation in clonal structure in a reef-building Caribbean coral, Acropora palmata. Ecol. Monogr. 76, 503–519. doi: 10.1890/0012-9615(2006)076
Besnier, F., and Glover, K. A. (2013). ParallelStructure: a R package to distribute parallel runs of the population genetics program STRUCTURE on multi-core computers. PLoS One 8:e70651. doi: 10.1371/journal.pone.0070651
Blanquer, A., and Uriz, M. J. (2010). Population genetics at three spatial scales of a rare sponge living in fragmented habitats. BMC Evol. Biol. 10:13. doi: 10.1186/1471-2148-10-13
Boulay, J. N., Hellberg, M. E., Cortés, J., and Baums, I. B. (2014). Unrecognized coral species diversity masks differences in functional ecology. Proc. R. Soc. Lond. B Biol. Sci. 281:20131580. doi: 10.1098/rspb.2013.1580
Bruno, J. F., Selig, E. R., Casey, K. S., Page, C. A., Willis, B. L., Harvell, C. D., et al. (2007). Thermal stress and coral cover as drivers of coral disease outbreaks. PLoS Biol. 5:e124. doi: 10.1371/journal.pbio.0050124
Budd, A. F. (2000). Diversity and extinction in the Cenozoic history of Caribbean reefs. Coral Reefs 19, 25–35. doi: 10.1007/s003380050222
Chan, A. N., Lewis, C. L., Neely, K. L., and Baums, I. B. (2018). Fallen pillars: the past, present, and future population dynamics of a rare, specialist coral-algal symbiosis. bioRxiv [Preprint].
Courchamp, F., Clutton-Brock, T., and Grenfell, B. (1999). Inverse density dependence and the Allee effect. Trends Ecol. Evol. 14, 405–410. doi: 10.1016/s0169-5347(99)01683-3
Davies, K. F., Margules, C. R., and Lawrence, J. F. (2004). A synergistic effect puts rare, specialized species at greater risk of extinction. Ecology 85, 265–271. doi: 10.1890/03-0110
De Iorio, M., and Griffiths, R. C. (2004a). Importance sampling on coalescent histories. I. Adv. Appl. Probabil. 36, 417–433. doi: 10.1239/aap/1086957579
De Iorio, M., and Griffiths, R. C. (2004b). Importance sampling on coalescent histories. II: subdivided population models. Adv. Appl. Probabil. 36, 434–454. doi: 10.1239/aap/1086957580
de Lange, P. J., and Norton, D. A. (2004). The ecology and conservation of Kunzea sinclairii (Myrtaceae), a naturally rare plant of rhyolitic rock outcrops. Biol. Conserv. 117, 49–59. doi: 10.1016/s0006-3207(03)00262-3
Deák, B., Valkó, O., Török, P., Kelemen, A., Bede,Á., Csathó, A. I., et al. (2018). Landscape and habitat filters jointly drive richness and abundance of specialist plants in terrestrial habitat islands. Landsc. Ecol. 33, 1117–1132. doi: 10.1007/s10980-018-0660-x
Devlin-Durante, M. K., and Baums, I. B. (2017). Genome-wide survey of single-nucleotide polymorphisms reveals fine-scale population structure and signs of selection in the threatened Caribbean elkhorn coral, Acropora palmata. PeerJ 5:e4077. doi: 10.7717/peerj.4077
Díaz-Ferguson, E., Haney, R., Wares, J., and Silliman, B. (2010). Population genetics of a trochid gastropod broadens picture of Caribbean Sea connectivity. PLoS One 5:e12675. doi: 10.1371/journal.pone.0012675
Dorken, M. E., and Eckert, C. G. (2001). Severely reduced sexual reproduction in northern populations of a clonal plant, Decodon verticillatus (Lythraceae). J. Ecol. 89, 339–350. doi: 10.1046/j.1365-2745.2001.00558.x
Dubé, C. E., Boissin, E., Maynard, J. A., and Planes, S. (2017). Fire coral clones demonstrate phenotypic plasticity among reef habitats. Mol. Ecol. 26, 3860–3869. doi: 10.1111/mec.14165
Eakin, C. M., Morgan, J. A., Heron, S. F., Smith, T. B., Liu, G., Alvarez-Filip, L., et al. (2010). Caribbean corals in crisis: record thermal stress, bleaching, and mortality in 2005. PLoS One 5:e13969. doi: 10.1371/journal.pone.0013969
Earl, D. A. (2012). STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361. doi: 10.1007/s12686-011-9548-7
Edmunds, P., Roberts, D., and Singer, R. (1990). Reefs of the northeastern Caribbean I. Scleractinian populations. Bull. Mar. Sci. 46, 780–789. doi: 10.1371/journal.pone.0003718
Ehrenburg, C. (1834). Beitrage zur physiologischen Kenntniss der Corallenthiere im allgemeined, und besonders des rothen Meers, nebst einem Versucch zu physiologishen systematic derselben. Phys. Abh. Konigl. Akad. Wissech. Berlin Jahar. 1832, 225–380.
Ellstrand, N. C., and Elam, D. R. (1993). Population genetic consequences of small population size: implications for plant conservation. Annu. Rev. Ecol. Syst. 24, 217–242. doi: 10.1146/annurev.ecolsys.24.1.217
Estoup, A., and Angers, B. (1998). “Microsatellites and minisatellites for molecular ecology: theoretical and empirical considerations,” in Advances in Molecular Ecology, ed. G. R. Carvalho (Amsterdam: IOS Press).
Evanno, G., Regnaut, S., and Goudet, J. (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620. doi: 10.1111/j.1365-294x.2005.02553.x
Excoffier, L., Smouse, P. E., and Quattro, J. M. (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, 479–491.
Finney, J. C., Pettay, D. T., Sampayo, E. M., Warner, M. E., Oxenford, H. A., and LaJeunesse, T. C. (2010). The relative significance of host–habitat, depth, and geography on the ecology, endemism, and speciation of coral endosymbionts in the genus Symbiodinium. Microb. Ecol. 60, 250–263. doi: 10.1007/s00248-010-9681-y
Flather, C. H., and Sieg, C. H. (2007). “Species rarity: definition, causes and classification,” in Conservation of Rare or Little-Known Species: Biological, Social, and Economic Considerations, eds M. G. Raphael and R. Molina (Washington, DC: Island Press), 40–66.
Foster, N. L., Paris, C. B., Kool, J. T., Baums, I. B., Stevens, J. R., Sanchez, J. A., et al. (2012). Connectivity of Caribbean coral populations: complementary insights from empirical and modelled gene flow. Mol. Ecol. 21, 1143–1157. doi: 10.1111/j.1365-294X.2012.05455.x
Gardner, T. A., Côté, I. M., Gill, J. A., Grant, A., and Watkinson, A. R. (2003). Long-term region-wide declines in Caribbean corals. Science 301, 958–960. doi: 10.1126/science.1086050
Gélin, P., Fauvelot, C., Mehn, V., Bureau, S., Rouzé, H., and Magalon, H. (2017). Superclone expansion, long-distance clonal dispersal and local genetic structuring in the coral Pocillopora damicornis type β in Reunion Island, South Western Indian Ocean. PLoS One 12:e0169692. doi: 10.1371/journal.pone.0169692
Gitzendanner, M. A., Weekley, C. W., Germain-Aubrey, C. C., Soltis, D. E., and Soltis, P. S. (2012). Microsatellite evidence for high clonality and limited genetic diversity in Ziziphus celata (Rhamnaceae), an endangered, self-incompatible shrub endemic to the Lake Wales Ridge, Florida, USA. Conserv. Genet. 13, 223–234. doi: 10.1007/s10592-011-0287-9
Goreau, T. F., and Wells, J. (1967). The shallow-water Scleractinia of Jamaica: revised list of species and their vertical distribution range. Bull. Mar. Sci. 17, 442–453.
Hedrick, P. W. (2005). A standardized genetic differentiation measure. Evolution 59, 1633–1638. doi: 10.1111/j.0014-3820.2005.tb01814.x
Heron, S. F., Maynard, J. A., Van Hooidonk, R., and Eakin, C. M. (2016). Warming trends and bleaching stress of the world’s coral reefs 1985–2012. Sci. Rep. 6:38402. doi: 10.1038/srep38402
Hoegh-Guldberg, O., Mumby, P. J., Hooten, A. J., Steneck, R. S., Greenfield, P., Gomez, E., et al. (2007). Coral reefs under rapid climate change and ocean acidification. Science 318, 1737–1742.
Hoegh-Guldberg, O., Poloczanska, E. S., Skirving, W., and Dove, S. (2017). Coral reef ecosystems under climate change and ocean acidification. Front. Mar. Sci. 4:158. doi: 10.3389/fmars.2017.00158
Hughes, A. R., and Stachowicz, J. J. (2004). Genetic diversity enhances the resistance of a seagrass ecosystem to disturbance. Proc. Natl. Acad. Sci. U.S.A. 101, 8998–9002. doi: 10.1073/pnas.0402642101
Hughes, T. P., Barnes, M. L., Bellwood, D. R., Cinner, J. E., Cumming, G. S., Jackson, J. B., et al. (2017). Coral reefs in the Anthropocene. Nature 546, 82–90.
Hunter, I., and Jones, B. (1996). Coral associations of the pleistocene ironshore formation, grand cayman. Coral Reefs 15, 249–267. doi: 10.1007/bf01787459
Jackson, J., Donovan, M., Cramer, K., and Lam, V. (2014). Status and Trends of Caribbean Coral Reefs: 1970-2012. Townsville: Global Coral Reef Monitoring Network.
Jossart, Q., De Ridder, C., Lessios, H. A., Bauwens, M., Motreuil, S., Rigaud, T., et al. (2017). Highly contrasted population genetic structures in a host–parasite pair in the Caribbean Sea. Ecol. Evol. 7, 9267–9280. doi: 10.1002/ece3.3413
Jost, L. (2008). GST and its relatives do not measure differentiation. Mol. Ecol. 17, 4015–4026. doi: 10.1111/j.1365-294X.2008.03887.x
Kabay, L. (2016). Population Demographics and Sexual Reproduction Potential of the Pillar Coral, Dendrogyra cylindrus, on the Florida Reef Tract. NSUWorks 433. Master’s thesis, Nova Southeastern University, Fort Lauderdale, FL.
Kemp, D., Fitt, W., and Schmidt, G. (2008). A microsampling method for genotyping coral symbionts. Coral Reefs 27, 289–293. doi: 10.1007/s00338-007-0333-8
Koh, L. P., Dunn, R. R., Sodhi, N. S., Colwell, R. K., Proctor, H. C., and Smith, V. S. (2004). Species coextinctions and the biodiversity crisis. Science 305, 1632–1634. doi: 10.1126/science.1101101
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A., and Mayrose, I. (2015). Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191. doi: 10.1111/1755-0998.12387
LaJeunesse, T. C., Parkinson, J. E., Gabrielson, P. W., Jeong, H. J., Reimer, J. D., Voolstra, C. R., et al. (2018). Systematic revision of Symbiodiniaceae highlights the antiquity and diversity of coral endosymbionts. Curr. Biol. 28, 2570–2580.e6. doi: 10.1016/j.cub.2018.07.008
Leblois, R., Pudlo, P., Néron, J., Bertaux, F., Reddy Beeravolu, C., Vitalis, R., et al. (2014). Maximum-likelihood inference of population size contractions from microsatellite data. Mol. Biol. Evol. 31, 2805–2823. doi: 10.1093/molbev/msu212
Lewis, A. M., Chan, A. N., and LaJeunesse, T. C. (2018). New species of closely related endosymbiotic dinoflagellates in the greater caribbean have niches corresponding to host coral phylogeny. J. Eukaryot. Microbiol.
Lewis, C. L., Neely, K. L., Richardson, L. L., and Rodriguez-Lanetty, M. (2017). Temporal dynamics of black band disease affecting pillar coral (Dendrogyra cylindrus) following two consecutive hyperthermal events on the Florida Reef Tract. Coral Reefs 36, 427–431. doi: 10.1007/s00338-017-1545-1
Lunz, K., Neely, K. L., Kabay, L., and Gilliam, D. (2016). Monitoring and Mapping of Dendrogyra cylindrus of the Florida Reef Tract. Final Report 9751-281-1168:39.
Lyons, K. G., Brigham, C., Traut, B., and Schwartz, M. W. (2005). Rare species and ecosystem functioning. Conserv. Biol. 19, 1019–1024. doi: 10.1111/j.1523-1739.2005.00106.x
Lyons, K. G., and Schwartz, M. W. (2001). Rare species loss alters ecosystem function–invasion resistance. Ecol. Lett. 4, 358–365. doi: 10.1046/j.1461-0248.2001.00235.x
Manzello, D. P. (2015). Rapid recent warming of coral reefs in the Florida Keys. Sci. Rep. 5:16762. doi: 10.1038/srep16762
Marhaver, K. L., Vermeij, M. J., and Medina, M. M. (2015). Reproductive natural history and successful juvenile propagation of the threatened Caribbean Pillar Coral Dendrogyra cylindrus. BMC Ecol. 15:9. doi: 10.1186/s12898-015-0039-7
Matthies, D., Bräuer, I., Maibom, W., and Tscharntke, T. (2004). Population size and the risk of local extinction: empirical evidence from rare plants. Oikos 105, 481–488. doi: 10.1111/j.0030-1299.2004.12800.x
Matz, M. V., Treml, E. A., Aglyamova, G. V., and Bay, L. K. (2018). Potential and limits for rapid genetic adaptation to warming in a Great Barrier Reef coral. PLoS Genetics 14:e1007220. doi: 10.1371/journal.pgen.1007220
McKinney, M. L. (1997). Extinction vulnerability and selectivity: combining ecological and paleontological views. Annu. Rev. Ecol. Syst. 28, 495–516. doi: 10.1146/annurev.ecolsys.28.1.495
Meirmans, P. G., and Van Tienderen, P. H. (2004). GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol. Ecol. Notes 4, 792–794. doi: 10.1111/j.1471-8286.2004.00770.x
Miller, M., Baums, I., Pausch, R., Bright, A., Cameron, C., Williams, D., et al. (2018). Clonal structure and variable fertilization success in Florida Keys broadcast-spawning corals. Coral Reefs 37, 239–249. doi: 10.1007/s00338-017-1651-0
Miller, S., Precht, W. F., Rutten, L. M., and Chiappone, M. (2013). Florida Keys Population Abundance Estimates for Nine Coral Species Proposed for Listing Under the US Endangered Species Act. Fort Lauderdale, FL: Nova Southeastern University, 1–82.
Miller, S. L., Chiappone, M., and Rutten, L. M. (2010). Abundance, Distribution and Condition of Benthic Coral Reef Organisms in the Upper Florida Keys National Marine Sanctuary: 2010 Quick Look Report and Data Summary. Wilmington, DE: University of North Carolina Wilmington.
Mouillot, D., Bellwood, D. R., Baraloto, C., Chave, J., Galzin, R., Harmelin-Vivien, M., et al. (2013). Rare species support vulnerable functions in high-diversity ecosystems. PLoS Biol. 11:e1001569. doi: 10.1371/journal.pbio.1001569
Mouillot, D., Villéger, S., Parravicini, V., Kulbicki, M., Arias-González, J. E., Bender, M., et al. (2014). Functional over-redundancy and high functional vulnerability in global fish faunas on tropical reefs. Proc. Natl. Acad. Sci. U.S.A. 111, 13757–13762. doi: 10.1073/pnas.1317625111
National Academies of Sciences Engineering and Medicine (2018). A Research Review of Interventions to Increase the Persistence, and. (Resilience) of Coral Reefs. Washington, DC: The National Academies Press.
Neely, K., Lewis, C., Chan, A., and Baums, I. (2018). Hermaphroditic spawning by the gonochoric pillar coral Dendrogyra cylindrus. Coral Reefs 37, 1087–1092. doi: 10.1007/s00338-018-1730-x
NMFS (2014). Endangered and Threatened Wildlife and Plants: Final Listing Determinations on Proposal to List 66 Reef-Building Coral Species and To Reclassify Elkhorn and Staghorn Corals, Vol. 79. College Park, MD: National Archives and Records Administration, 272.
NOAA. (2014). Endangered and threatened wildlife and plants: final listing determinations on proposal to list 66 reef-building coral species and to reclassify elkhorn and staghorn corals. Fed Reg 75:271.
Parkinson, J. E., Banaszak, A. T., Altman, N. S., LaJeunesse, T. C., and Baums, I. B. (2015). Intraspecific diversity among partners drives functional variation in coral symbioses. Sci. Rep. 5:15667. doi: 10.1038/srep15667
Parkinson, J. E., Bartels, E., Devlin-Durante, M. K., Lustic, C., Nedimyer, K., Schopmeyer, S., et al. (2018). Extensive transcriptional variation poses a challenge to thermal stress biomarker development for endangered corals. Mol. Ecol. 27, 1103–1119. doi: 10.1111/mec.14517
Parkinson, J. E., and Baums, I. B. (2014). The extended phenotypes of marine symbioses: ecological and evolutionary consequences of intraspecific genetic diversity in coral–algal associations. Front. Microbiol. 5:445. doi: 10.3389/fmicb.2014.00445
Peakall, R., and Smouse, P. E. (2006). GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6, 288–295. doi: 10.1111/j.1471-8286.2005.01155.x
Pettay, D. T., and LaJeunesse, T. C. (2007). Microsatellites from clade B Symbiodinium spp. specialized for Caribbean corals in the genus Madracis. Mol. Ecol. Notes 7, 1271–1274. doi: 10.1111/j.1471-8286.2007.01852.x
Precht, W. F., Gintert, B. E., Robbart, M. L., Fura, R., and Van Woesik, R. (2016). Unprecedented disease-related coral mortality in Southeastern Florida. Sci. Rep. 6:31374. doi: 10.1038/srep31374
Pritchard, J. K., Stephens, M., and Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics 155, 945–959.
Pritchard, J. K., Wen, X., and Falush, D. (2009). Documentation for Structure Software: Version 2.3.
Puechmaille, S. J. (2016). The program structure does not reliably recover the correct population structure when sampling is uneven: subsampling and new estimators alleviate the problem. Mol. Ecol. Resour. 16, 608–627. doi: 10.1111/1755-0998.12512
R Development Core Team (2017). R: A Language and Environment for Statistical Computing. Vienna: R Foundation for Statistical Computing. Available at: https://www.R-project.org/
Rabinowitz, D. (1981). “Seven forms of rarity,” in The Biological Aspects of Rare Plants Conservation, ed. H. Synge (Chichester: John Wiley & Sons), 205–217.
Reusch, T. B., Ehlers, A., Hämmerli, A., and Worm, B. (2005). Ecosystem recovery after climatic extremes enhanced by genotypic diversity. Proc. Natl. Acad. Sci. U.S.A. 102, 2826–2831. doi: 10.1073/pnas.0500008102
Rippe, J. P., Matz, M. V., Green, E. A., Medina, M., Khawaja, N. Z., Pongwarin, T., et al. (2017). Population structure and connectivity of the mountainous star coral, Orbicella faveolata, throughout the wider Caribbean region. Ecol. Evol. 7, 9234–9246. doi: 10.1002/ece3.3448
Rousset, F. (2008). genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106. doi: 10.1111/j.1471-8286.2007.01931.x
Ryman, N., and Palm, S. (2006). POWSIM: a computer program for assessing statistical power when testing for genetic differentiation. Mol. Ecol. Notes 6, 600–602. doi: 10.1111/j.1471-8286.2006.01378.x
Santos, S. R., and Coffroth, M. A. (2003). Molecular genetic evidence that dinoflagellates belonging to the genus Symbiodinium Freudenthal are haploid. Biol. Bull. 204, 10–20. doi: 10.2307/1543491
Scobie, A., and Wilcock, C. (2009). Limited mate availability decreases reproductive success of fragmented populations of Linnaea borealis, a rare, clonal self-incompatible plant. Ann. Bot. 103, 835–846. doi: 10.1093/aob/mcp007
Shantz, A. A., Ladd, M. C., Schrack, E., and Burkepile, D. E. (2015). Fish-derived nutrient hotspots shape coral reef benthic communities. Ecol. Appl. 25, 2142–2152. doi: 10.1890/14-2209.1
Silvertown, J. (2008). The evolutionary maintenance of sexual reproduction: evidence from the ecological distribution of asexual reproduction in clonal plants. Int. J. Plant Sci. 169, 157–168. doi: 10.1086/523357
Solórzano, S., Arias, S., and Dávila, P. (2016). Genetics and conservation of plant species of extremely narrow geographic range. Diversity 8:31. doi: 10.3390/d8040031
Steiner, S. C., and Kerr, J. (2008). Stony corals in Dominica during the 2005 bleaching episode and one year later. Rev. Biol. Trop. 56, 139–148.
Stephens, P. A., and Sutherland, W. J. (1999). Consequences of the Allee effect for behaviour, ecology and conservation. Trends Ecol. Evol. 14, 401–405. doi: 10.1016/s0169-5347(99)01684-5
Stocker, T., Qin, D., Plattner, G., Tignor, M., Allen, S., Boschung, J., et al. (2013). “IPCC, 2013: climate change 2013: the physical science basis,” in Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change, eds T. F. Stocker, D. Qin, G.-K. Plattner, M. Tignor, S. K. Allen, J. Boschung, et al. (Cambridge: Cambridge University Press), 1535.
Swarts, N. D., Sinclair, E. A., Francis, A., and Dixon, K. W. (2010). Ecological specialization in mycorrhizal symbiosis leads to rarity in an endangered orchid. Mol. Ecol. 19, 3226–3242. doi: 10.1111/j.1365-294X.2010.04736.x
Szmant, A. M. (1986). Reproductive ecology of Caribbean reef corals. Coral Reefs 5, 43–53. doi: 10.1007/bf00302170
Theodose, T. A., Jaeger, C. H. III, Bowman, W. D., and Schardt, J. C. (1996). Uptake and allocation of 15 N in alpine plants: implications for the importance of competitive ability in predicting community structure in a stressful environment. Oikos 75, 59–66. doi: 10.2307/3546321
Thornhill, D., Howells, E., Wham, D., Steury, T., and Santos, S. (2017). Population genetics of reef coral endosymbionts (Symbiodinium, Dinophyceae). Mol. Ecol. 26, 2640–2659. doi: 10.1111/mec.14055
Tooker, J. F., and Frank, S. D. (2012). Genotypically diverse cultivar mixtures for insect pest management and increased crop yields. J. Appl. Ecol. 49, 974–985. doi: 10.1111/j.1365-2664.2012.02173.x
van Hooidonk, R., Maynard, J., Tamelander, J., Gove, J., Ahmadia, G., Raymundo, L., et al. (2017). Coral Bleaching Futures: Downscaled Projections of Bleaching Conditions for the World’s Coral Reefs, Implications of Climate Policy and Management Responses. Available at: http://hdl.handle.net/20.500.11822/22048 (accessed April 2018).
van Hooidonk, R., Maynard, J. A., Liu, Y., and Lee, S. K. (2015). Downscaled projections of Caribbean coral bleaching that can inform conservation planning. Glob. Chan. Biol. 21, 3389–3401. doi: 10.1111/gcb.12901
Vollmer, S. V., and Palumbi, S. R. (2006). Restricted gene flow in the Caribbean staghorn coral Acropora cervicornis: implications for the recovery of endangered reefs. J. Hered. 98, 40–50. doi: 10.1093/jhered/esl057
Waits, L. P., Luikart, G., and Taberlet, P. (2001). Estimating the probability of identity among genotypes in natural populations: cautions and guidelines. Mol. Ecol. 10, 249–256. doi: 10.1046/j.1365-294X.2001.01185.x
Ward, J., Rypien, K., Bruno, J., Harvell, C., Jordan-Dahlgren, E., Mullen, K., et al. (2006). Coral diversity and disease in Mexico. Dis. Aquat. Org. 69, 23–31.
Warner, R. R., and Chesson, P. L. (1985). Coexistence mediated by recruitment fluctuations: a field guide to the storage effect. Am. Nat. 125, 769–787. doi: 10.1086/284379
Werth, S., and Scheidegger, C. (2012). Congruent genetic structure in the lichen-forming fungus Lobaria pulmonaria and its green-algal photobiont. Mol. Plant Microbe Interact. 25, 220–230. doi: 10.1094/MPMI-03-11-0081
Willing, E.-M., Dreyer, C., and Van Oosterhout, C. (2012). Estimates of genetic differentiation measured by FST do not necessarily require large sample sizes when using many SNP markers. PLoS One 7:e42649. doi: 10.1371/journal.pone.0042649
Keywords: Dendrogyra cylindrus, climate change, population genetics, Symbiodiniaceae, rare species, clonal structure
Citation: Chan AN, Lewis CL, Neely KL and Baums IB (2019) Fallen Pillars: The Past, Present, and Future Population Dynamics of a Rare, Specialist Coral–Algal Symbiosis. Front. Mar. Sci. 6:218. doi: 10.3389/fmars.2019.00218
Received: 14 January 2019; Accepted: 08 April 2019;
Published: 21 May 2019.
Edited by:
Rui Rosa, Universidade de Lisboa, PortugalReviewed by:
Nancy Knowlton, Smithsonian Institution, United StatesCelia Schunter, The University of Hong Kong, Hong Kong
Daniel J. Thornhill, National Science Foundation (NSF), United States
Copyright © 2019 Chan, Lewis, Neely and Baums. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Andrea N. Chan, YW5kcmVhLm4uY2hhbjEzQGdtYWlsLmNvbQ==
 Andrea N. Chan
Andrea N. Chan Cynthia L. Lewis
Cynthia L. Lewis Karen L. Neely
Karen L. Neely Iliana B. Baums
Iliana B. Baums