- 1Marine Biology Research Group, Department of Biology, Ghent University, Ghent, Belgium
- 2Laboratoire Environnement Profond, Institut Français de Recherche pour l’Exploitation de la MER, Plouzané, France
Cold seeps occur globally in areas where gases escape from the seafloor, occasionally resulting in the formation of topographic depressions (pockmarks), characterised by unique physicochemical conditions such as anoxic and sulphuric sediments. Free-living marine nematodes tend to dominate the meiofaunal component in such environments, often occurring at extremely high densities and low richness; the mechanisms defining community assembly in areas of fluid seepage, however, have received little attention. Here we focus on a low-activity pockmark at 789 m in the Mozambique Channel (MC). We assessed the diversity, co-occurrence patterns and phylogenetic community structure of nematodes at this bathyal site to that of a nearby reference area as well as abyssal sediments using metabarcoding. In addition, we compared our molecularly-derived diversity estimates to replicate samples identified morphologically. Overall, nematode Amplicon Sequence Variants (ASVs) and generic richness were similar between Pockmark and Abyssal sediments, but lower compared to the Reference area. Although more than half the genera were shared, over 80% of ASVs were unique within each area and even within each replicate core. Even though both methodologies differentiated the Pockmark from the Reference and Abyssal sites, there was little overlap between the molecularly and morphologically identified taxa, highlighting the deficit of reference sequences for deep-sea nematodes in public databases. Phylogenetic community structure at higher taxonomic levels was clustered and did not differ between the three areas yet analysis within three shared and dominant genera (Acantholaimus, Desmoscolex, Halalaimus), revealed randomness with respect to phylogeny as well as co-occurrence which was exclusive to the Pockmark area. These patterns point to the influence of neutral dynamics at this locality resulting from the stochastic sampling of early colonizing taxa, the successional stage at sampling and/or the functional redundancy within the investigated genera.
Introduction
Cold seeps occur in sedimentary habitats worldwide in areas where methane and reduced sulfur are discharged from the seafloor. Topographic depressions (pockmarks) or elevations (mud volcanoes) resulting from the escaping fluids are often associated with these ecosystems while the surrounding sediments show low oxygen and high hydrogen sulfide concentrations (Levin, 2005; Zeppilli et al., 2018; and references therein). Free-living nematodes which are found globally in marine sediments, appear to be well-adapted to pockmarks where they are the dominant meiofaunal taxon (Vanreusel et al.,2010a,b; Moens et al., 2014; Zeppilli et al., 2018; and references therein). The effect of methane seepage on nematode communities often results in elevated densities and biomass, as well as near monospecific assemblages (Vanreusel et al., 2010b). Dominance by taxa which are typical of shallow-water environments and found in low abundance in non-seep deep-sea sediments (e.g., Halomonhystera, Desmodora, Sabatieria, Terschellingia), has been commonly observed in cold seeps, although no strict affinities have yet been documented (Gaever et al., 2009; Vanreusel et al., 2010b). The community structure of free-living marine nematodes with respect to co-occurrence and phylogeny in a cold seep ecosystem has yet to be investigated.
Prompted by the widespread application of DNA-based methodologies and the availability of phylogenetic trees, the merging of community ecology and phylogenetic biology has allowed for a better understanding of the relative dominance of neutral, historical and niche-related processes in community assembly (Webb et al., 2002). Neutral dynamics assume ecological equivalence between species, the persistence of which is thus determined by the stochastic element inherent to processes such as offspring birth/death and dispersal (Hubbel, 2001). Historical processes refer to patterns of speciation and dispersal which supersede local dynamics. Finally, the niche-based perspective proposes that communities assemble in accordance to fundamental rules (e.g., Diamond, 1975) defined by local biotic and abiotic conditions (Cavender-Bares et al., 2009). The phylogenetic composition of a community represents (at least in part) the mechanisms of assembly because taxa interact via their phenotypes which are non-randomly distributed phylogenetically (Vamosi et al., 2009). Thus, assemblages exhibiting relatedness that is higher than expected of a randomly structured community, is suggestive of environmental filtering due to shared conserved trait(s) (Webb et al., 2002). Conversely, lower relatedness than expected by chance is indicative of convergent evolution of the traits defined by the environmental filter and/or competitive exclusion of conserved traits, although the phylogenetic influence of competition has been disputed (Mayfield and Levine, 2010). Furthermore, it has been proposed that competitive exclusion would manifest as an excess of forbidden taxon pairs (i.e., “checkerboard pairs,” CPs) leading to aggregated assemblages (Diamond, 1975).
A recent metabarcoding study demonstrated that free-living nematode Amplicon Sequence Variants (ASVs) from different sites in a polymetallic nodule-bearing abyssal plain exhibited both higher relatedness as well as higher co-occurrences than would be expected by chance (Macheriotou et al., 2020). As a highly diversified and dominant deep-sea sediment-dwelling eukaryotic taxon, nematodes provide an excellent model group for testing the importance of environmental filtering on the basis of aggregation and clustering in different ecosystems. The Mozambique Channel (MC, Figure 1) between the island of Madagascar to the East and Mozambique to the West, is characterized by complex oceanographic conditions due to its shape and location, making it one of the most turbulent areas in the world. The MC is of unique ecological value, not only for its high biodiversity of coral reefs (Obura, 2012) and near pristine ecosystems (e.g., Scattered Islands), but also due to its connection to the climatologically important Agulhas current and its contribution to inter-ocean exchanges (Ternon et al., 2014). The MC is characterized by submarine canyons, guyots, seamounts, and associated cold fluid emissions, while the deep-sea benthic biota have only in recent years began to be explored (Olu, 2014; Jouet and Deville, 2015; Fontanier et al., 2016).
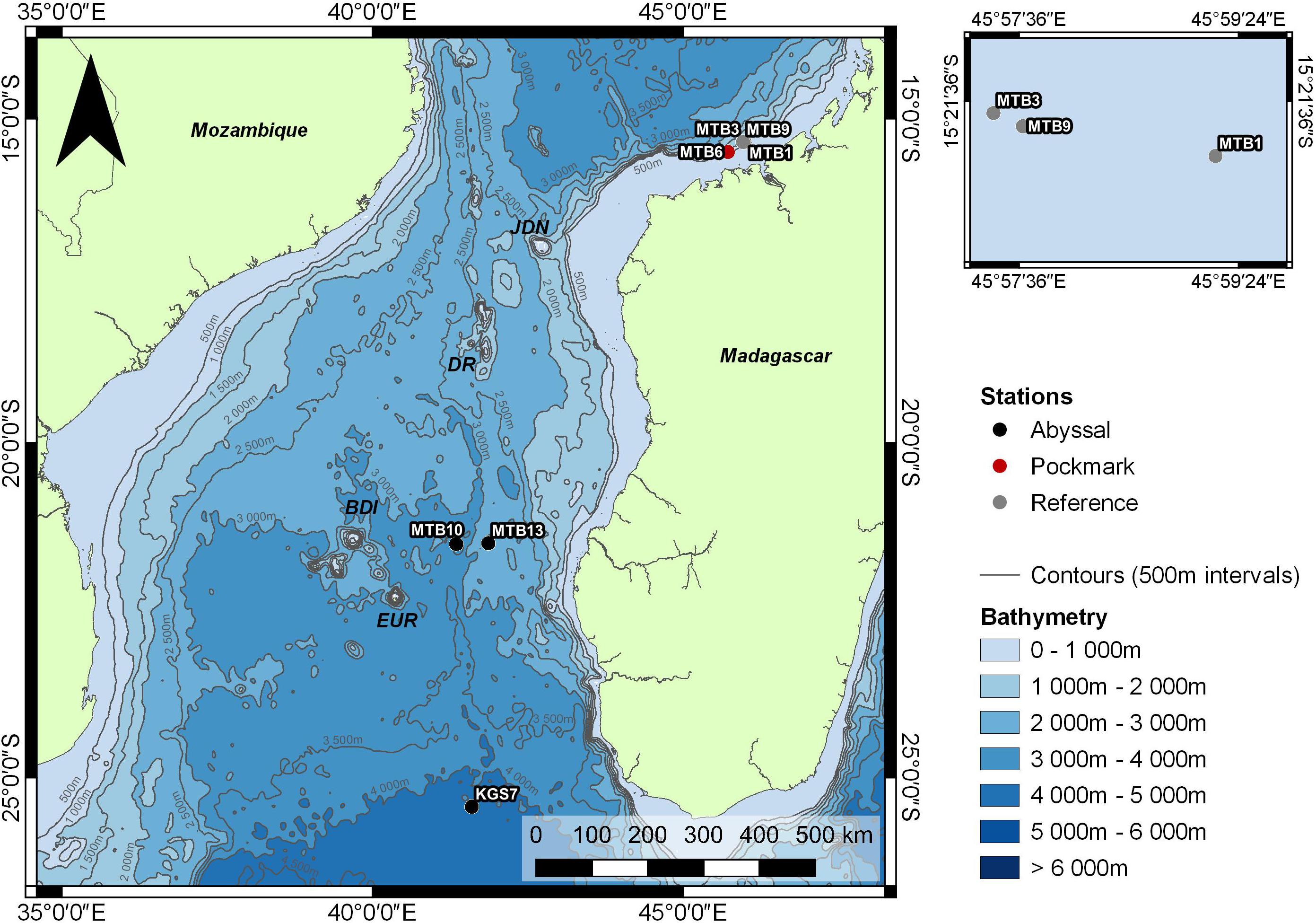
Figure 1. Areas sampled in the Mozambique Channel. Circles indicate sampling sites in the Abyssal (black, MTB10, MTB13, KGS7), Pockmark (red, MTB6) and Reference (gray, MTB1, MTB3, MTB9) areas.
Here, we apply a metabarcoding approach to compare co-occurrence patterns, phylogenetic community structure and diversity of free-living nematode assemblages in a low-activity bathyal pockmark with a nearby area outside any seepage influence as well as abyssal sediments in the Mozambique Channel. Given the aforementioned dominance of single/few taxa, we hypothesize that phylogenetic diversity will be lower in the pockmark. Furthermore, we expect that nematode communities from areas with active emissions are assembled differently than those of surrounding non-seep sediments due to the presence of strong chemical gradients and sporadic disturbance events by fluid discharge; thus we hypothesize that co-occurrence and phylogenetic community patterns will differ between seep and non-seep habitats.
Materials and Methods
Sample Collection and Meiofauna Extraction
Sediment cores were collected during the PAssive Margins Exploration Laboratories PAMELA-Moz01 cruise on board the RV “L’Atalante” (Olu, 2014; PAMELA-MOZ01 cruise, RV L’Atalante1) from 25/09-29/10/2014. Seven sites were sampled using a multitube (MTB, Supplementary Table 1) and one site by sub-sampling cores from a USNEL Boxcore (KGS), representing three different environments: a bathyal Pockmark (depth: 789 m, n = 1), Reference sediments near the Pockmark area (∼30 km from Pockmark, depth: 529–757 m, n = 3) and Abyssal sediments (∼800 km from Reference and ∼1000 km from Pockmark, depth: 3–4 km, n = 3). Environmental parameters derived from a single sediment core showed that the Pockmark differed substantially from the Abyssal area in terms of lower oxygen penetration depth (∼15 mm, ∼85 mm, respectively; Fontanier et al., 2016). In addition, the Pockmark was characterized by elevated elemental sulfur content compared to the Reference area (0–5 cm average: 0.804%, 0.127% respectively; Sanchez et al., 2020), suggesting recent seepage at this location. Gas bubbling had also been recorded above the pockmark confirming current emissions (Dupré et al., unpublished data) Three cores (A, B, C; inner diameter: 62 mm) were obtained from each site, sliced into five 1 cm intervals (Note: MTB1 lacking interval designation). Half of each sediment slice was frozen at −20°C while the other preserved in 4% formalin. Meiofauna was extracted by density-gradient centrifugation (3 × 12 min, 1905 rcf) with the colloidal silica polymer Ludox® HS-40 as a flotation medium (specific density 1.18 g/cm).
HTS Library Preparation
Metabarcoding samples were preserved in sterile Mili-Q water during the first two centrifugation cycles and finally in CTAB at −20°C after the third. Genomic DNA was extracted by adding 6 μL proteinase-K [10 mg/mL] and centrifuged 5 min at 14000 rpm at room temperature (RT). The pellet was ground, bead-beaten for 2 min at 30 cycles/second and incubated at 60°C for 1 h. Ammonium acetate (250 μL, 7.5 M) was added and tubes centrifuged for 10 min at 14000 rpm (RT). The supernatant (750 μL) was transferred into a new sterile tube; 750 μL of cold 80% isopropanol was added, mixed, incubated for 30 min at RT, and centrifuged for 15 min at 14000 rpm (4°C). The supernatant was removed, 1 mL washing buffer (76% EtOH 10 mM ammonium acetate) was added, and tubes were incubated for 30 min on ice and centrifuged for 5 min at 14000 rpm (4°C). Finally, the supernatant was removed, and 20 μL of sterile water was added. The 18S (V1-V2 region) ribosomal locus was amplified using the primers SSU_F_04-SSU/22_R (GCTTGTCTCAAAGATTAAGCC, TCCAAGGAAG GCAGCAGGC, respectively) which were constructed with Illumina overhang adapters as described in “16S Metagenomic Sequencing Library Preparation.” Each sample was amplified in triplicate with the following PCR conditions: 95°C 2 min, 30 × (95°C 1 min, 57°C 45 s, 72°C 1 min), 72°C 10 min. The mix consisted of 8.4 μL PCR-grade H2O, 4 μL Phusion Buffer, 4 μL Dye, 0.4 μL dNTP [10 mM], 1 μL forward and reverse primer [10 μM], 0.2 μL Phusion Hot Start II High Fidelity Polymerase (New England BioLabs, United States) and 1 μL DNA template (diluted 1/10). In the event of failed amplification, DNA templates were diluted 1/50 in PCR-grade H2O and/or 2 μL template were used. PCR products were run on a 1% agarose electrophoresis gel, triplicates were pooled, purified using Agencourt AMPure XP beads and run on Bioanalyzer 2100 High Sensitivity to confirm length and size distribution of the PCR fragments. Library indexing was completed using the FC131-1002 NexteraXT Index Kit (Illumina, United States) and Kapa High Fidelity PCR kit (Kapa Biosystems, United States). The mix consisted of 11.25 μL PCR-grade H2O, 5 μL Buffer, 0.75 μL dNTP [10 mM], 2.5 μL Index1 and Index2, 0.5 μL Kapa Hot Start High Fidelity Polymerase and 2.5 μL PCR product. These were then purified using Agencourt AMPure XP beads and 11 randomly chosen samples were run on Bioanalyzer 2100 High Sensitivity to confirm successful indexing. DNA was quantified using Qubit® dsDNA High Sensitivity Assay Kit in all samples for pooling. Finally, biological replicates were distributed over three pooled libraries and sequenced at Edinburgh Genomics on three Illumina MiSeq-v3 2 × 300 bp paired-end read runs2.
Morphological Analysis
Morphological samples were preserved in 4% formalin throughout and were stained with Bengal Rose (0.5 g/L). Total meiofaunal taxa abundances were counted in each interval while nematode specimens from the 0–1 cm layer from each core excluding KGS7 were also identified morphologically. To achieve this, ca. 100 nematodes were picked randomly from each 0–1 cm slice, transferred to anhydrous glycerol and mounted on permanent glass slides (Seinhorst, 1959). Genus-level identification was completed using a Leica DMLS compound microscope (1000 × magnification) according to pictorial keys (Platt et al., 1988; Warwick et al., 1998; Handbook of Zoology 2013, Ed. by Schmidt-Rhaesa, Andreas) as well as the information contained in the NeMYS database (Bezerra et al., 2021).
Sequence Data Analysis
Gene-specific adapter sequences were truncated from the 5′ and 3′ read ends using Cutadapt v1.12 (Martin, 2014); Amplicon Sequence Variants (ASVs) were generated following the default DADA2 workflow (Callahan et al., 2016) with truncation of forward and reverse reads at 225 and 200 bp, respectively (ESM, Bioinformatics + DADA2 output). Taxonomic assignment of ASVs was completed in QIIME1 (assign_taxonomy.py) (Caporaso et al., 2010) with the Naive Bayesian RDP classifier (Wang et al., 2007) at a confidence estimate of 0.80 in two steps: first, a large eukaryotic reference training set was used (Silva release 123 for QIIME1, 99% OTUs and UGent nematode Sanger sequences; n = 20201). Second, all ASVs with “Nematoda” label were extracted; taxonomy was re-assigned to these ASVs using a smaller, nematode-exclusive training set (2178 sequences). This two-step approach has been shown to deliver the highest number of taxonomic assignments (Macheriotou et al., 2019). Samples were rarefied to the lowest sequence count (n = 26480) for subsequent analyses (QIIME1, alpha_rarefaction.py). Three samples were excluded from the analysis due to low read numbers (MTB1.B.10, MTB10.A.0-1cm, MTB9.C.3-4cm, MTB9.C.3-4cm).
Diversity Analysis
Alpha-diversity metrics [Shannon (H′), Simpson (1- D), occurrence-based Index of Relative Rarity, Leroy et al., 2013] of the rarefied sequence data were calculated at the level of Nematoda ASVs (Nematoda), of the genera these were assigned to (Genus-assigned) as well as the morphological samples (0–1 cm only). Data normality and homoscedasticity were assessed with the Shapiro–Wilk and Levene test respectively; statistically significant differences in diversity along the sediment vertical profile by Area were determined via a two-way Analysis of Variance (ANOVA) with fixed factors Area and Interval. Significant differences by Area where evaluated using one-way ANOVA for the morphological dataset; the Kruskal–Wallis test was used when homoscedasticity was violated. Phylogenetic beta-diversity was calculated with the unweighted UniFrac (Lozupone and Knight, 2005) distance for Nematoda (n = 14197), Genus-assigned (n = 6643), Unassigned (n = 7554), the three most abundant genera Acantholaimus (n = 895), Desmoscolex (n = 2489), Halalaimus (n = 1020) and Other Genera (all genus-assigned ASVs excluding Acantholaimus, Desmoscolex, Halalaimus; n = 2239) ASVs. ASVs lacking a taxonomic assignment beyond Nematoda (“Unassigned”) were included in our analyses to investigate whether these (presumably) unknown taxa exhibit patterns similar to those identified at genus-level. Similarly, we included the level “Other Genera” to assess whether subdominant genera exhibit similar patterns to dominant ones. The Bray–Curtis distance was used for morphological and ASV samples in the 0–1 cm layer. Distances were visualized by Principal Coordinates Analysis (PCoA). Data normality and homoscedasticity were assessed by analysis of multivariate homogeneity of group dispersions (PERMDISP) and two-way permutational analysis of variance (PERMANOVA, 9999 permutations). Pairwise comparisons between group levels with Bonferroni corrections for multiple testing were completed in the event of a significant main test and lack of interaction between the two factors. Phylogenetic diversity (PD, Faith, 1992) was quantified for each of the aforementioned ASV sets. The following R packages were used: BiodiversityR, stats, Rarity, RVAideMemoire, UpSetR, vegan.
Community Structure Analysis
Co-occurrence patterns were analyzed with “Checkerboard pairs” (CPs) and C-score, (Diamond, 1975; Stone and Roberts, 1990). Standard effect sizes (ses, explained below) against a null model (sim2, 10000 replicates) were calculated in “EcoSimR” (Gotelli et al., 2015). The sim2 algorithm randomizes a presence-absence matrix by reshuffling elements within each row equiprobably. The C-score quantifies the average number of CPs across all possible paired combinations; in a matrix with sites as columns and taxa as rows, for each unique pair, the C-score is equal to Cij = (Ri-S)(Rj-S) where Ri and Rj are row sums of taxa i, j and S is the number of shared sites in which both i and j are present. For any particular species pair, the C-score is a numerical index that ranges from a minimum of zero (maximally aggregated) to a maximum of Ri × Rj (maximally segregated with no shared sites). Departures from randomly co-occurring assemblages were assessed via upper- and lower-tail one-sample t-tests. A positive or negative mean value for ses signifies taxon co-occurrences which are lower or higher than expected by chance, respectively. These analyses were computationally feasible only for Acantholaimus, Desmoscolex, and Halalaimus ASVs. Additionally, the ses of the following phylogenetic metrics were calculated with “picante” (Webb et al., 2010):
Phylogenetic Diversity – PD (Faith, 1992): sum of branch lengths between the root and all species in a sample phylogeny.
Mean Pairwise Distance – MPD (Webb et al., 2002): mean pairwise phylogenetic distance between all taxa in a local assemblage (pres/abs).
Mean Nearest Taxon Distance – MNTD (Webb et al., 2002): mean phylogenetic distance between each taxon and its nearest neighbor on the phylogenetic tree with which it co-occurs in the local assemblage (pres/abs).
The observed metric (obsMetric) was compared to that obtained from 999 randomizations (nullMetric) of the assemblage generated using all null models (i.e., taxa.labels, richness, frequency, sample.pool, phylogeny.pool, independentswap, trialswap, Gotelli, 2000; Miklós and Podani, 2004, see Supplementary Table 20 for full description). The ses was calculated as follows: ses.Metric=(obsMetric–nullMetric)/ stdev.nullMetric where stdev equals the standard deviation. Significant departures from a zero mean for ses.PD/ses.MPD/ses.MNTD were tested with a one-sample two-tailed t-test when data were normally distributed as determined by the Shapiro–Wilk test. Alternatively, the Wilcoxon signed rank test was used. A positive or negative mean value for ses signifies phylogenetic relatedness which is higher or lower than expected by chance, respectively. Statistical significance was set at α = 0.05. For these analyses, tree construction was completed by aligning reads with “DECIPHER” (Wright, 2016) using a chained guide tree; alignments were subsequently adjusted with the adjust.alignment command. The software “ModelTest-ng” (Darriba et al., 2019) was used to determine the appropriate model of nucleotide evolution for tree construction for each ASV set. In all instances the General Time Reversible (GTR) model was selected as one of the best-scoring models according to the either the Bayesian or the Akaike Information Criterion. Approximately-maximum-likelihood phylogenetic trees were constructed using FastTree (Price et al., 2010) under the GTR+CAT model; midpoint rooting was completed with “phangorn” (Schliep, 2011). All analyses were completed in RStudio© v1.1.456 unless noted otherwise. Note: Samples belonging to MTB1 (Reference) were removed for vertical profile analysis (unlabelled intervals) resulting in a total of 1421, 7301, and 6112 ASVs in the Abyssal, Pockmark, and Reference areas, respectively.
Results
Nematode Assemblages: Alpha and Beta Diversity
The rarefied data consisted of 24517 ASVs belonging to 40 phyla (Supplementary Figure 1); an average of 58% were identified as nematodes while 21% were Unassigned at phylum-level. These Unassigned ASVs highlight a deficiency in the number reference sequences as well as in the proficiency of the taxonomic assignment algorithm. Within Nematoda ASVs (n = 14197), 71 genera were identified, representing half of the community, with the remainder Unassigned at genus-level (Figure 2). Over half of these genera were shared by all three areas (n = 39); Reference and Abyssal shared an additional 16 genera while three were shared by the Abyssal and Pockmark (Figure 3). Two genera were unique to the Abyssal (Pareurystomina, Setostephanolaimus) and Pockmark (Aegialoalaimus, Atrochromadora) ASVs respectively, while nine were only found in Reference. The most abundant genus was Desmoscolex with over 10% relative abundance in all three areas, followed by Halalaimus, which constituted over 7% of the assemblage throughout. The genus Acantholaimus was dominant (>5% relative abundance) only in the Abyssal and Reference samples (Supplementary Table 2).
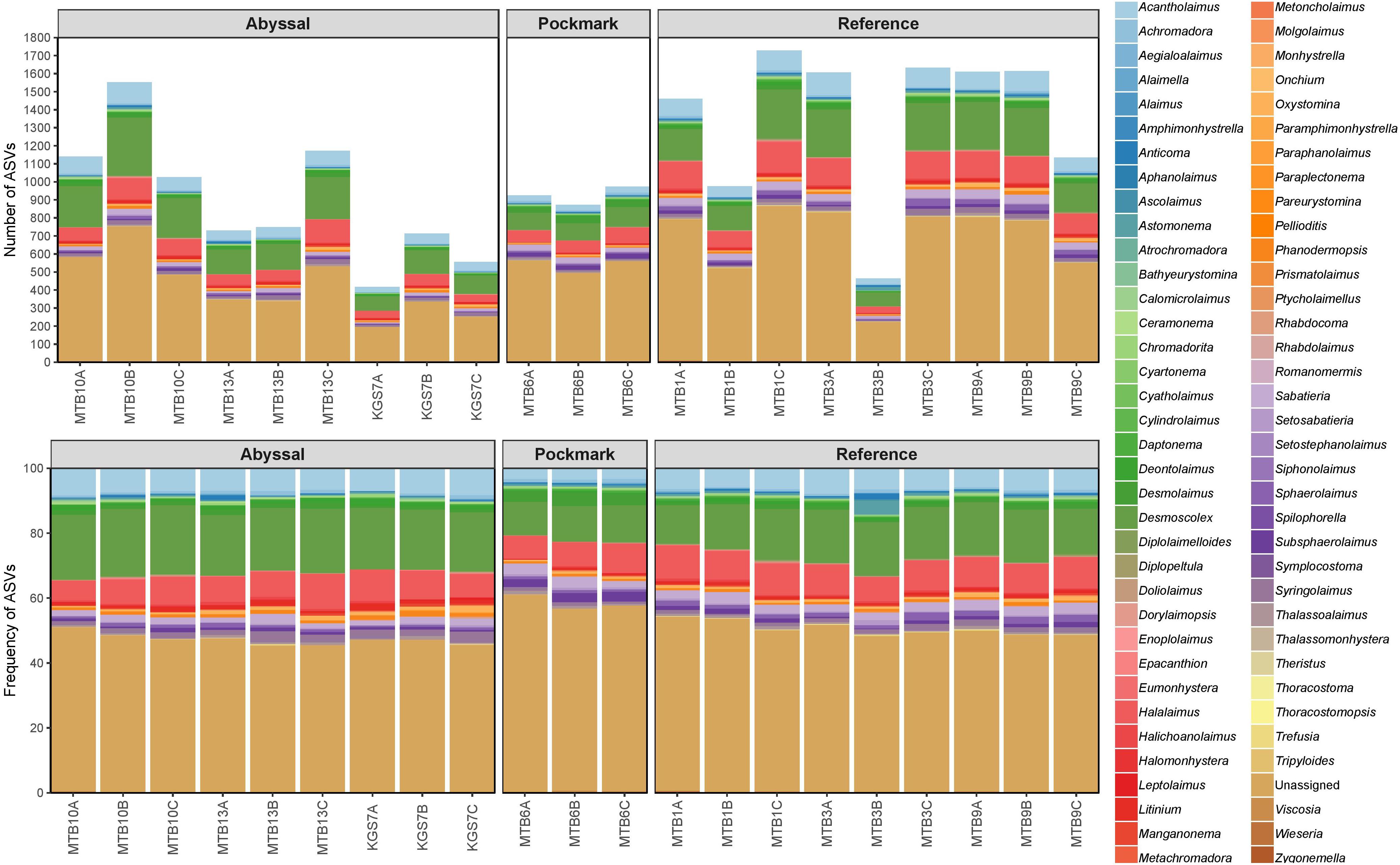
Figure 2. Absolute (top panel) and relative (bottom panel) abundance of nematode genera in the Abyssal, Pockmark, and Reference areas.
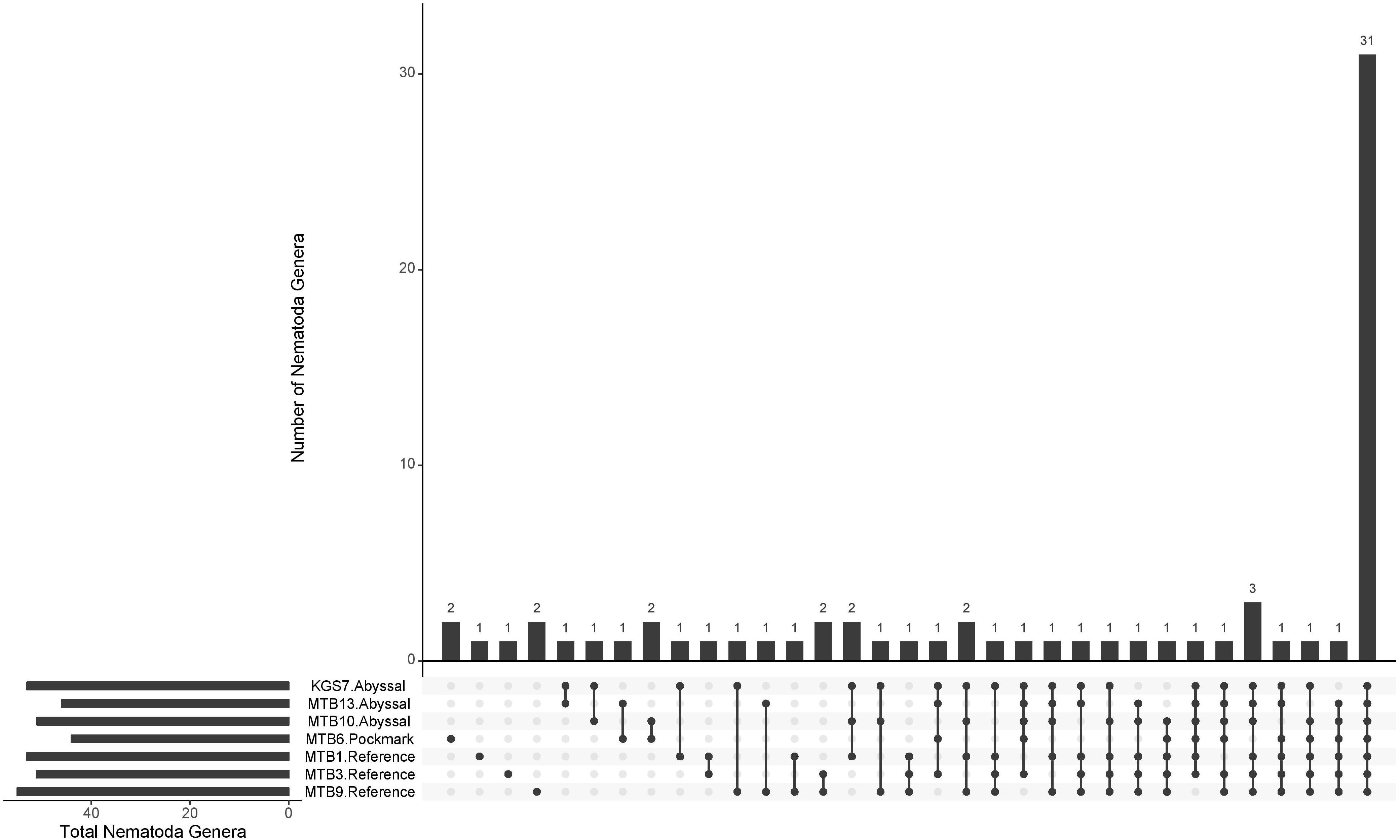
Figure 3. Unique and shared genera identified in Nematoda ASVs. Main panel: numbers above bars indicate the number of nematode genera found in the ASVs of the replicates marked by filled circles. Bottom-left graph: total number of nematode genera in each of the Abyssal, Pockmark, and Reference area replicates (sediment intervals pooled).
The majority of ASVs (>80%) were unique to each area and each replicate within it, with just 57 ASVs (0.4%) shared by all three areas (Supplementary Figure 2). The Pockmark shared 47 and 126 ASVs with Abyssal and Reference respectively, while the latter two had 350 ASVs in common. There was no significant interaction between factors Area and Interval in the two-way ANOVA for alpha-diversity metrics. ASV richness, and that of the genera these were assigned to, was highest in the Reference area (p = 0.001, <0.001, Figure 4 and Supplementary Tables 3–5). The first interval (0–1 cm) contained a larger number of ASVs and genera compared to 3–4 cm and 4–5 cm. Shannon and Simpson values at ASV-level were not significantly different between areas yet significantly lower in the deepest layer (4–5 cm), compared to 0–1 cm and 1–2 cm (Supplementary Table 5). Rarity of ASVs was lowest in the Pockmark area and in the 4–5 cm interval (p < 0.001, = 0.048). Shannon and Simpson indices at the generic-level were lowest in the Abyssal area (p < 0.001) and not significantly different between sediment intervals. Rarity of genera was highest in the Reference ASVs and in the surface sediment layers compared to 4–5 cm.
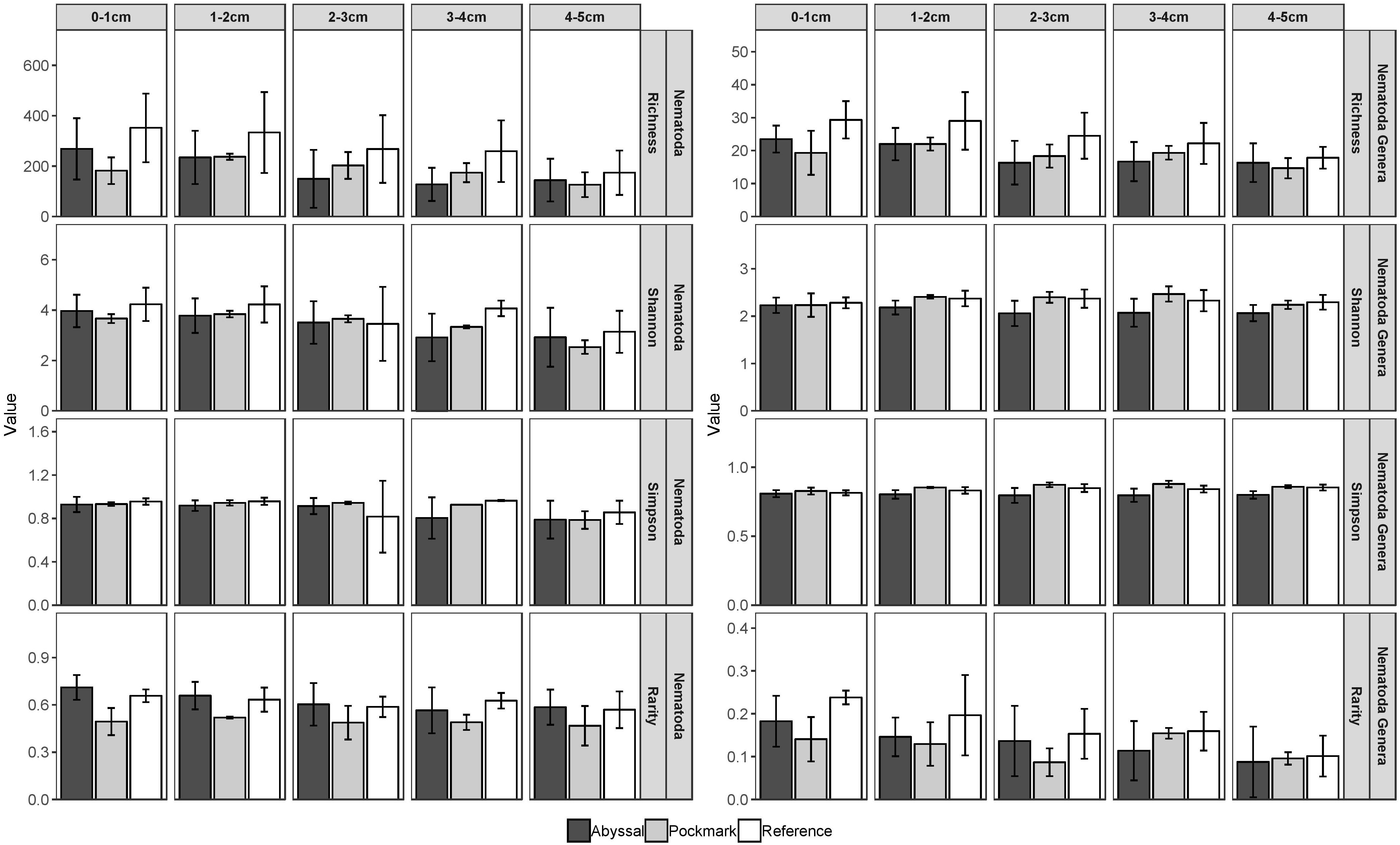
Figure 4. Alpha diversity metrics [Shannon (H0), Simpson (1-D), and occurrence-based index of relative rarity] of rarefied Nematoda ASVs by sediment interval in the Abyssal (dark gray), Pockmark (light gray), and Reference (white) areas. Error bars represent standard deviation.
The morphological samples identified 117 genera, including nine “Unknown” and likely new genera; Desmoscolex and Halalaimus were dominant in the Reference and Abyssal areas while Acantholaimus only in the latter (Supplementary Figure 3 and Supplementary Table 6). The Pockmark was dominated by Desmodora, a genus that was not recorded in the ASVs, most likely due to failed amplification (see section “Discussion”). The highest nematode abundance was observed in the Pockmark (1260 ± sd 1370), followed by the Abyssal (274 ± sd 203) and Reference samples (182 ± sd 83, Supplementary Table 7). The Pockmark exhibited significantly lower generic richness, Shannon and Simpson diversity in the morphological samples (13.32 ± sd 15.50, 1.17 ± sd 1.16, 0.74 ± sd 0.35, respectively) and higher Rarity (0.37 ± sd 0.11) compared to Reference and Abyssal (p < 0.01, Supplementary Figure 4 and Supplementary Tables 8, 9). In the ordination of both the ASVs and the morphologically identified samples (0–1 cm only), Pockmark was strongly segregated from Reference and Abyssal areas, the latter two exhibiting a small amount of overlap (Figure 5).
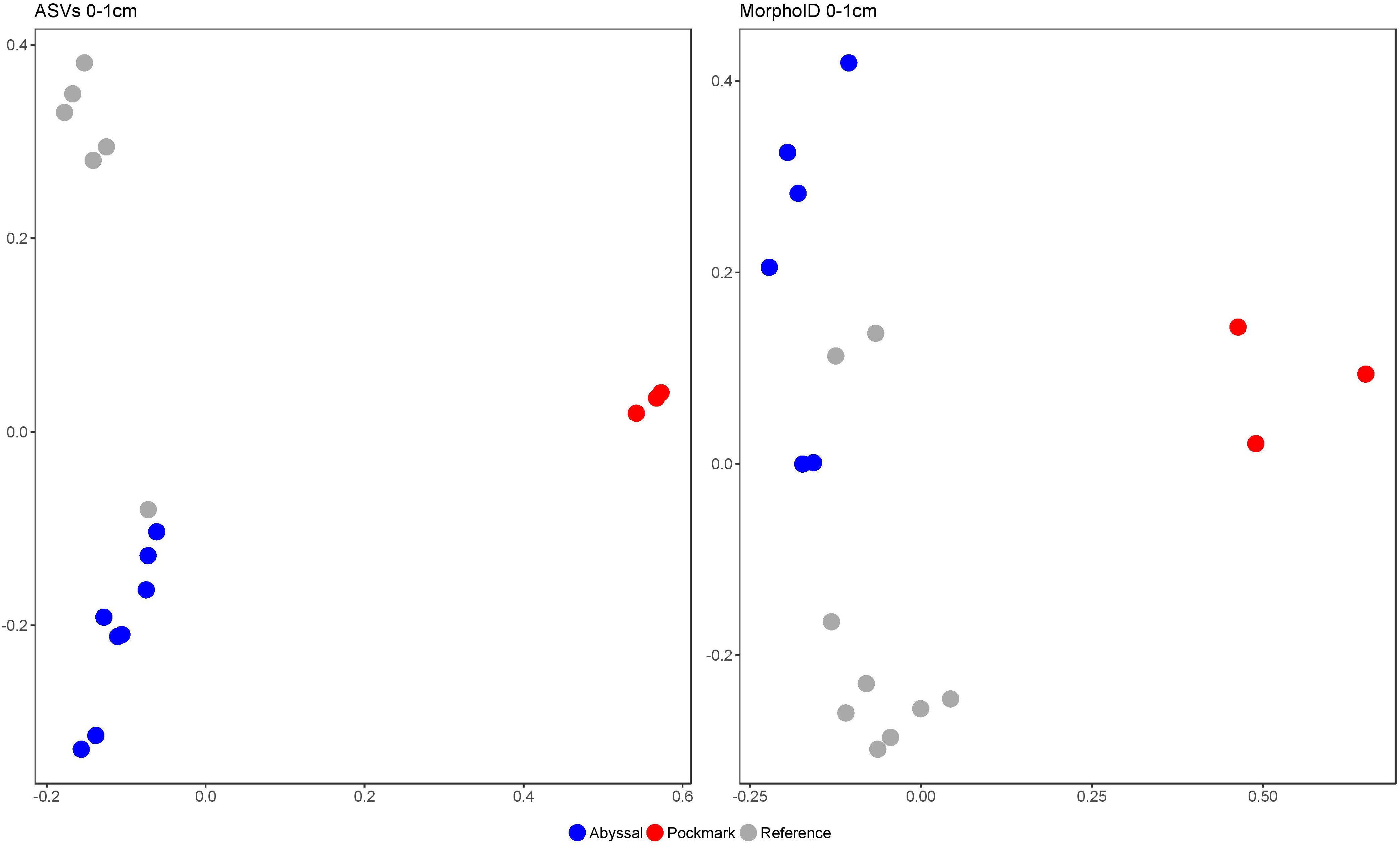
Figure 5. Principal coordinates analysis (PCoA) of Bray-Curtis distance of rarefied ASVs (left panel) and morphologically identified samples (right panel) in the 0-1 cm interval of the Abyssal (blue), Pockmark (red), and Reference (gray) areas.
There was no significant interaction between main factors in the two-way ANOVA of PD; both Area and Interval had a significant effect in all ASV sets. The Reference area contained the highest PD compared to Abyssal and Pockmark in Nematoda, Genus-assigned, Unassigned and Acantholaimus ASVs (Figure 6 and Supplementary Tables 10–12). Desmoscolex ASVs in the Reference area had significantly higher PD than that of the Pockmark; Halalaimus and Other Genera ASVs had higher PD in the Reference area compared to the Abyssal. The first sediment interval contained significantly more PD than 2–3, 3–4, 4–5 cm and the second more than 3–4 and 4–5 cm in Nematoda, Genus-assigned, Desmoscolex and Halalaimus ASVs. The deepest interval contained significantly less PD than the 0–1, 1–2 cm in Unassigned and Other Genera while Acantholaimus ASVs exhibited the least change in PD by depth with significantly lower values in the 4–5 cm compared to the 0–1 cm layer.
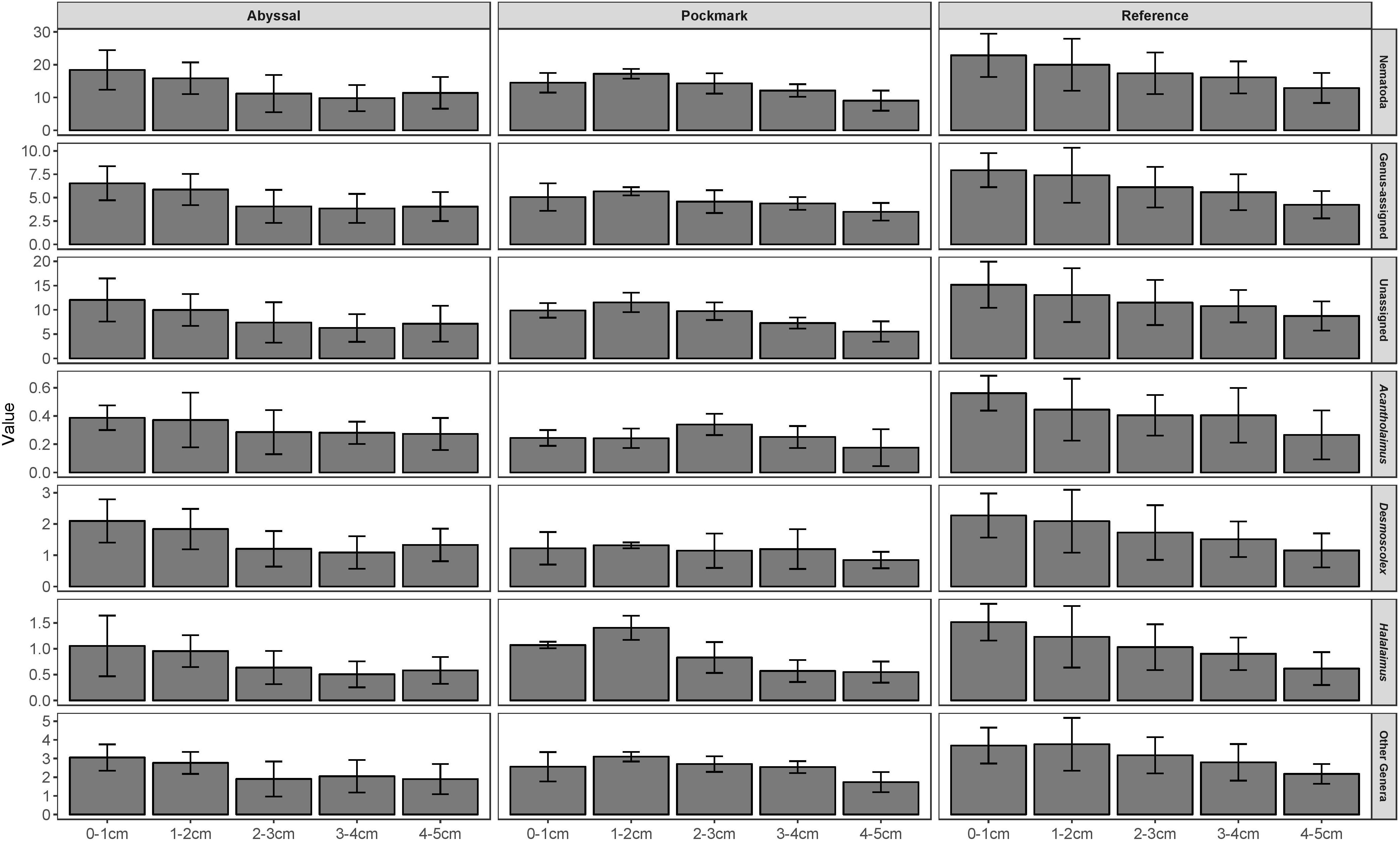

Figure 6. Phylogenetic diversity (PD) of Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in the Abyssal, Pockmark, and Reference areas by sediment interval. Error bars represent standard deviation.
The interaction between main factors Area and Interval was not statistically significant for the UniFrac distances in Genus-assigned, Unassigned, Acantholaimus and Desmoscolex ASVs, for which all areas differed from one another (Supplementary Tables 13, 14). Differences by depth were most pronounced in the latter where the 0–1 cm interval differed from 2–3, 3–4, 4–5 cm and 1–2 cm from 3–4 and 4–5 cm. In Genus-assigned, 0–1 cm differed from 2–3, 3–4, 4–5, and 1–2 cm from 4–5 cm. The 4–5 cm interval also differed from 0–1 cm in Acantholaimus ASVs.
Patterns of Co-occurrence
The co-occurrence analysis demonstrated patterns that were consistent between metrics as well as across Acantholaimus, Desmoscolex, and Halalaimus ASVs (Figure 7). Assemblages showed significantly higher co-occurrences than expected by chance in the Abyssal and Reference areas while these were always random in the Pockmark (p < 0.001, Supplementary Table 15). When sediment intervals were considered separately, all Reference and nearly all Abyssal ASVs (excluding Acantholaimus 0–1 cm, Halalaimus 3–4 cm) exhibited statistically significant aggregation, while in the Pockmark the majority of these were random (Supplementary Table 16). Notably, Desmoscolex ASVs were predominantly aggregated (excluding Pockmark 1–2 cm) while random co-occurrences were more common in Acantholaimus and Halalaimus ASVs.
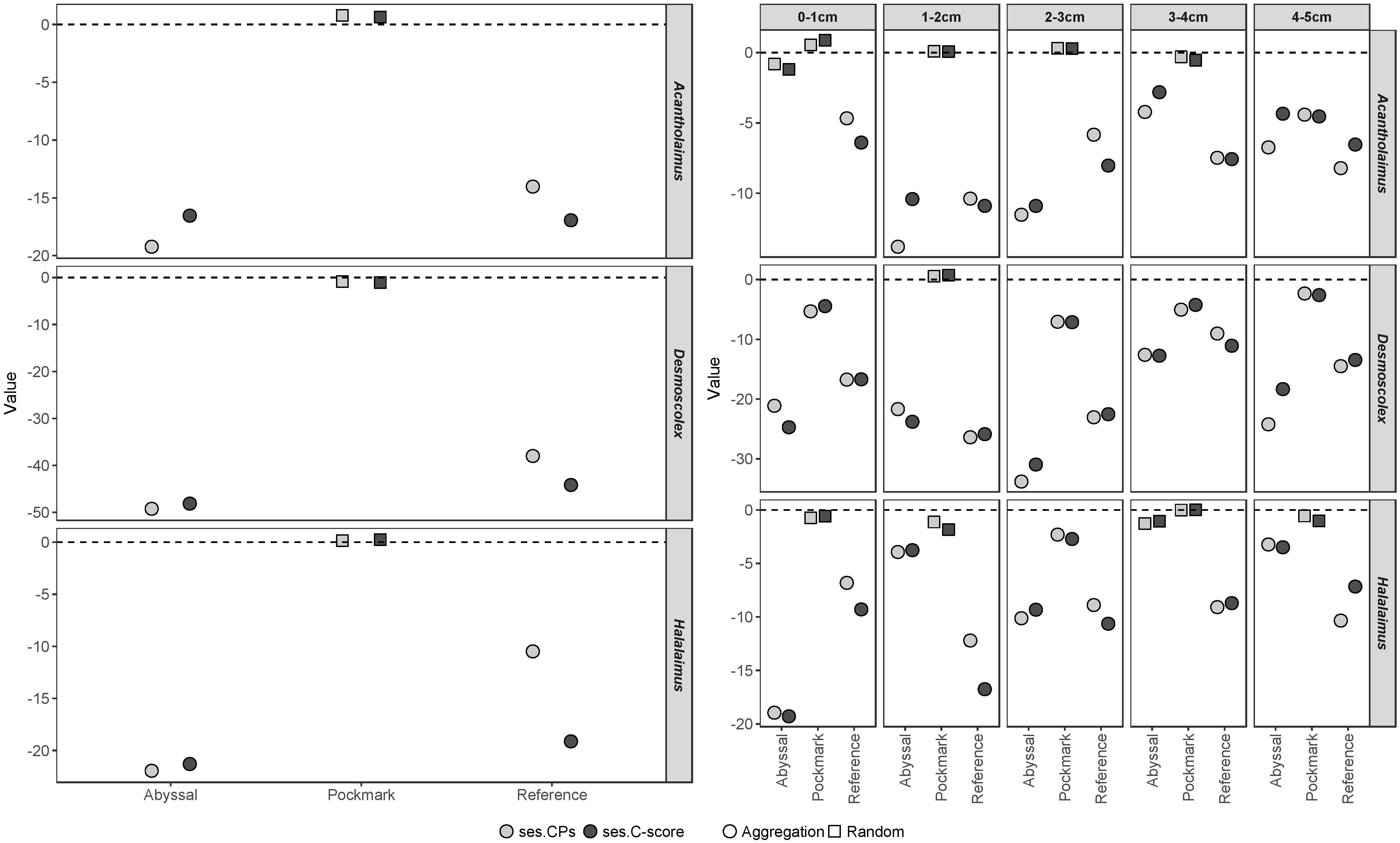
Figure 7. Standard effect size (ses) of Checkerboard Pairs (ses.CPs, light gray) and C-score (ses.C-score, dark gray) of Acantholaimus, Desmoscolex, and Halalaimus ASVs in the Abyssal, Pockmark and Reference areas with sediment intervals pooled (left panel) and by sediment interval (right panel). Symbols represent outcome of the relevant t-test; circle and square indicate statistically significant aggregation and random co-occurrences respectively.
Phylogenetic Community Structure
When all sediment intervals were pooled, the larger ASV sets (Nematoda, Genus-assigned, Unassigned, Other Genera) exhibited phylogenetic clustering in all areas and across all three metrics with the exception of ses.MPD for Pockmark in Other Genera ASVs which were phylogenetically random (Figure 8 and Supplementary Table 17). Differences by area were evident at the generic level; ses.PD, ses.MNTD, and ses.MPD indicated clustering in Acantholaimus, Desmoscolex, and Halalaimus ASVs in the Abyssal and Reference areas while Pockmark contained phylogenetically random assemblages. When sediment intervals were considered separately, the Abyssal samples were typically clustered in all ASV sets for all three metrics and sediment intervals with a few exceptions of randomness in the first and deepest layer for Desmoscolex and Genus-assigned, respectively (Figure 9 and Supplementary Table 18). The Reference ASVs followed a similar pattern yet with more instances of randomness in the first three intervals, giving away to clustering in the last two. The Pockmark samples were most commonly random with exceptional instances of clustering in the deeper sediment layers in Nematoda and Other Genera ASVs. Overall, random phylogenetic structure was most prevalent in the Pockmark while clustering was characteristic of both the Abyssal and Reference ASVs. Overdispersion was found in just two instances; ses.MPD in the Reference area for Genus-assigned and ses.MNTD for Other Genera ASVs in Pockmark.

Figure 8. Standard effect size (ses) of Phylogenetic Diversity (ses.PD, top panel), Mean Nearest Taxon Distance (ses.MNTD, middle panel), and Mean Pairwise Distance (ses.MPD, bottom panel) of Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in the Abyssal (dark gray), Pockmark (light gray) and Reference (white) areas. Symbols represent the outcome of the relevant t-test: triangle and circle indicate statistically significant clustering and random phylogenetic structure, respectively.
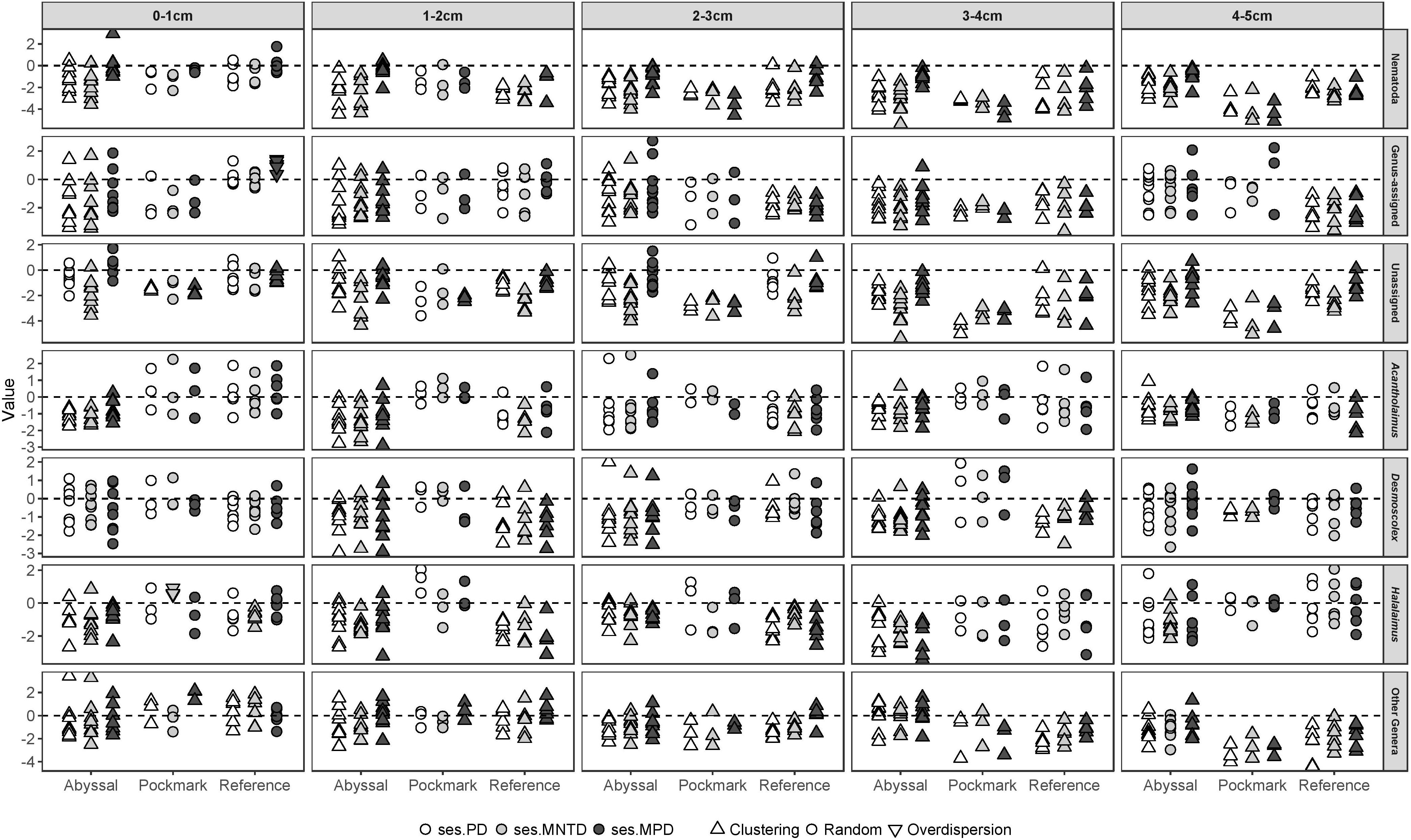

Figure 9. Standard effect size (ses) of Phylogenetic Diversity (ses.PD, white), Mean Nearest Taxon Distance (ses.MNTD, light gray), and Mean Pairwise Distance (ses.MPD, dark gray) by sediment interval of Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in the Abyssal (dark gray), Pockmark (light gray), and Reference (white) areas. Symbols represent the outcome of the relevant t-test: upward triangle, downward triangle, and circle indicate statistically significant clustering, overdispersion, and random phylogenetic structure, respectively.
Discussion
Diversity Patterns in Pockmark, Reference and Abyssal ASVs
The number of genera the Nematoda ASVs were assigned to was comparable across areas while Shannon diversity was higher in the Pockmark and Reference area compared to the Abyssal. Concurrently, evenness was higher in the Pockmark and Reference area than in the Abyssal samples. Rarity at the generic level, as opposed to ASVs (Supplementary Figure 2), was relatively low overall due to a large number of shared genera. Interestingly, the Reference area exhibited the highest Rarity as it contained nine unique genera. The overwhelming majority of ASVs were unique to each area, with less than one percent shared by all three. The Reference area, the most shallow and consequently receiving larger organic carbon inputs, harbored the richest and most diverse assemblages with respect to ASVs and their generic composition, which peaked in the upper two sediment intervals. Although comparable between areas, ASV diversity decreased significantly with sediment layer depth. Nematode ASV densities did not exhibit subsurface maxima as was documented in the morphological samples (Supplementary Figure 5) and at specific Antarctic and south Atlantic pockmarks (Sibuet et al., 2009; Hauquier et al., 2011), but followed the classic pattern of attenuation by depth seen in pockmarks elsewhere (Van Gaever et al., 2004). Moreover, dominance by a small number of taxa, which is characteristic of most morphological assessments of pockmark nematofauna (Van Gaever et al., 2004, 2006; Gaever et al., 2010; Hauquier et al., 2011; Pape et al., 2011; Guilini et al., 2012), did occur in the morphologically identified samples by genus Desmodora (45% ± sd 38%, Supplementary Figures 3, 4) but not in the ASV data.
The ASV and morphological dataset (0–1 cm only) were congruent with respect to lower richness in the Pockmark compared to the Reference and Abyssal samples. Lower Shannon and Simpson values were not observed in both datasets, as dominance by the genus Desmodora was found in the morphological dataset exclusively. Low-diversity assemblages such as those commonly observed at seeps are also known to occur in oxygen minimum zones, where the hypoxic conditions have a homogenizing effect on both seep and non-seep sediments (Guilini et al., 2012). Contrastingly, dominance was low overall in the ASV dataset and highest in the Abyssal area where nearly one fifth of genus-assigned ASVs belonged to Desmoscolex. This genus was also dominant in the Abyssal and Reference areas of the morphological dataset, while it was represented by just a few individuals in the Pockmark samples. This typical deep-sea genus (Vanreusel et al., 2010a) showed the highest relative ASV abundance in both the Pockmark and Reference areas, which is in contrast to the observed trend in previous studies of shallow-water relatives colonizing seep sediments (Vanreusel et al., 2010b). It should be acknowledged, however, that such patterns must interpreted in the context of low replicate samples. The presence of active and/or inactive pockmarks has a prominent effect on sedimentary habitats by increasing environmental heterogeneity and by providing a unique, fluctuating ecological setting, which has been shown to both increase and decrease nematode abundance/diversity (Van Gaever et al., 2006; Zeppilli et al., 2012). It has been proposed that reduced generic richness, diversity and evenness at these food-rich seep areas may be due to the fact that only a small selection of taxa are adapted to oxygen-poor and sulfide-rich environments and/or once established, seep-adapted species dominate via the competitive exclusion of others (Vanreusel et al., 2010b).
The absence of the morphologically dominant genus Desmodora in the Pockmark ASVs is most likely attributable to failed PCR amplification, presumably due to its thick cuticle. We expect this taxon would have been identified as eight Desmodora sequences were included in our reference database and provided its DNA was present. In contrast to the morphological assay, the total number of ASVs was lowest in the Pockmark, pointing to a deficit which may in fact be attributable to the missing Desmodora ASVs. In accordance to our expectations, the Pockmark typically harbored lower PD compared to the Reference area, a pattern consistent with the morphological assessment. However, this needs to be interpreted in the context of the highly abundant Desmodora taxa being absent from the HTS data; lower PD may thus simply be an artifact of the missing ASVs. Concurrently, given that cold seeps have been shown to harbor low diversity and even monospecific assemblages, which would result in only a modest increase in PD, overall lower PD values in the Pockmark could still be valid even if the Desmodora ASVs had been identified. High abundances of Desmodora carry ecological implications with respect to fluid emissions from the seafloor, as this genus is known to be associated with such environments (Gaever et al., 2009; Vanreusel et al., 2010b; Pape et al., 2011; Zeppilli et al., 2011). Some Desmodora species have even evolved symbiotic relationships with chemoautotrophic sulfur-oxidizing bacteria beneath their cuticle in order to capitalize on the strong chemical gradients present in sulphuric sediments, thereby providing a detoxifying mechanism as well as a food source (Ott et al., 1991; Hentschel et al., 1999; Bernhard et al., 2000).
Nematode Community Structure in Areas of Fluid Seepage
Differences in phylogenetic structure between areas with all sediment intervals pooled were only apparent at genus-level. Analysis at increasingly higher taxonomic resolution has been shown to shift from clustering to overdispersion or randomness in plant communities (Cavender-Bares et al., 2006), which was indeed evidenced in the Pockmark ASVs, where higher taxonomic levels were clustered, while Acantholaimus, Desmoscolex, and Halalaimus ASVs were random (Figure 8). This pattern was not observed for the Abyssal and Reference areas, where these dominant genera remained phylogenetically clustered. Analysis by sediment interval confirmed an increased tendency for random structure in the Pockmark, which was generally consistent with increasing depth. Co-occurrence for the aforementioned genera was random in the Pockmark exclusively, while in the Abyssal and Reference areas these were aggregated. Metabarcoding-based investigations of co-occurrence patterns and phylogenetic community structure in deep-sea sediments are few and have typically reported patterns of aggregation and clustering, respectively (Macheriotou et al., 2020); whether randomness as observed in the Pockmark ASVs is uncommon for deep-sea ecosystems, however, remains unknown.
Random phylogenetic community structure is thought to occur when (1) the strength of density-dependent and environmental filtering processes are balanced or weak, (2) species niches are phylogenetically random, (3) species interactions are neutral, or (4) competitive exclusion of convergent traits occurs (Hubbel, 2001; Webb et al., 2002; Kembel and Hubbel, 2006). The specific harsh conditions of the Pockmark would presumably enhance the strength of environmental filtering and density-dependent processes, on which grounds we would dismiss this first line of reasoning. If species niches were phylogenetically random, we would expect this to manifest in the Abyssal and Reference areas also. The fourth interpretation cannot explain randomness in the Pockmark samples for two reasons: first, from the co-occurrence analysis we found little evidence indicating competitive exclusion is a dominant structuring force; second, by definition, convergent traits are those shared between distantly related lineages, which is not applicable within a genus.
The relatively strong influence of the chemical environment at pockmarks seems to preclude Neutral Theory as a mechanism of faunal assembly, supported by the occurrence of phylogenetic clustering in all non-genus-level ASV sets in the Pockmark. Concurrently, one could expect this influence to manifest within the dominant genera of the community as well, which was not observed. Neutrality may be applicable to specific groups, provided they exhibit some degree of functional redundancy and comparable environmental tolerances or habitat requirements (Mora et al., 2010). The morphological samples revealed that nematode genera dominating the Pockmark (Desmodora, Linhystera), are also present in the surrounding sediments at reduced densities. It seems that nematodes do not exhibit genus-specific affinities to seep environments, nor any obvious morphological adaptations to them (Vanreusel et al., 2010b), thus the aforementioned conditions for Neutral Theory could indeed be met. Moreover, Neutral Theory may be applicable to early colonizers; non-endemic vagrant taxa that wander into the pockmark and are sampled at that particular moment (Mora et al., 2010). Although rarely studied, the phylogenetic signature of successional dynamics appears to begin with random assemblages in the pioneer stage, overdispersion during intermediate phases, followed again by randomness (Siles et al., 2009). As randomness can result from competitive exclusion, of which we found no evidence, random co-occurrence and phylogenetic patterns in the Pockmark ASVs of Acantholaimus, Desmoscolex and Halalaimus could be explained by the successional stage of the specific sediment patch; sampling at regular time intervals could serve to further substantiate this hypothesis. Furthermore, repetitive disturbance of the benthos northwest of Madagascar by the deposition of terrestrial sediments rich in organic content over the past 60 years (Fontanier et al., 2018), in combination with the harsh environmental conditions of the Pockmark, may be interfering with the establishment of stable, closely related nematode assemblages, which would simultaneously amplify the importance of neutral dynamics at this locality. Different community assembly models represent a spectrum of non-mutually exclusive hypotheses (Gravel et al., 2006); our results show this to be true and most evident when such analyses are completed at multiple taxonomic levels.
Conclusion
This study represents the first metabarcoding assay of deep-sea nematodes in the Mozambique Channel. The comparison of communities at a Pockmark area with Reference sediments without seepage from similar depths and an Abyssal area provided an ideal setting for the phylogenetic investigation of nematode community structure. Even though at higher taxonomic levels patterns were identical in all three areas (i.e., phylogenetic clustering), genus-level analysis uncovered the distinct phylogenetic properties of the Pockmark. Random co-occurrence patterns and phylogenetic structure were exclusive to the Pockmark, a pattern consistent with the influence of neutral dynamics. Although the expectation would be that the harsh conditions of a pockmark would promote the dominance of environmental filtering, which was observed in the Reference and Abyssal areas, the relevance of neutrality at this site could be attributable to functional redundancy within the investigated genera, the stochastic sampling of early colonizing taxa and/or the successional stage of the sediment patch at time of sampling. Similar to previous work on seep nematofauna, we found no evidence of seep-specific genera. Finally, there was little overlap between the molecularly and morphologically identified taxa, highlighting the deficit of reference sequences for deep-sea nematodes in public databases and the (ever-present) need for barcoding initiatives.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA623689.
Author Contributions
LM and KO collected the field data. LM and AR carried out the molecular work. DZ contributed identification data for the morphological samples. LM completed all data analysis and composed the manuscript. SD participated in defining the analyses performed. SD and AV participated in the design of the study. KO, AR, DZ, SD, and AV critically revised the manuscript. All the authors contributed to the article and approved the submitted version.
Funding
This work was completed within the “Passive Margins Exploration Laboratories” (PAMELA) Project, co-funded by TOTAL and IFREMER. The PAMELA project is a scientific project lead by IFREMER and TOTAL in collaboration with Université de Bretagne Occidentale, Université Rennes 1, Université Pierre and Marie Curie, CNRS et IFPEN. This study was supported by Fonds Wetenschappelijk Onderzoek (FWO), grant no. 3F015515; The molecular research was carried out with infrastructure funded by EMBRC Belgium (FWO project GOH3817N).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank the Captain, Chief Scientist, scientific staff, and crew on board L’ Atalante during the Pamela-Mozambique01 campaign.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2021.549834/full#supplementary-material
Supplementary Table 1 | Sampling stations latitude, longitude (in decimal degrees), and depth.
Supplementary Table 2 | Average (Avg.) and standard deviation (St. dev.) of the relative abundance of nematode genera of rarefied ASVs in the Abyssal, Pockmark, and Reference areas.
Supplementary Table 3 | Average (Avg.) and standard deviation (St. dev.) for alpha-diversity metrics (Richness, Shannon, Simpson, Rarity) for rarefied Nematoda ASVs and their generic composition (Nematoda Genera) in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile.
Supplementary Table 4 | Results of statistical testing (p-value) for data normality (Shapiro–Wilk, S-W.Ar./S-W.Int. for Area and Interval, respectively), homoscedasticity (Levene) and two-way analysis of variance (ANOVA) for alpha-diversity metrics (Richness, Shannon, Simpson, Rarity) in rarefied Nematoda ASVs and their generic composition (Nematoda Genera) in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 5 | Tukey’s HSD (Honestly Significant Difference) test results (p-value) for pairwise comparisons of alpha-diversity metrics (Richness, Shannon, Simpson, Rarity) in rarefied Nematoda ASVs and their generic composition (Nematoda Genera) in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 6 | Average (Avg.) and standard deviation (St. dev.) of the relative abundance of nematode genera of 0–1 cm interval of the morphological samples in the Abyssal, Pockmark, and Reference areas.
Supplementary Table 7 | Total abundance (Sum), average (Avg.), and standard deviation (St. dev.) of nematodes of 0–1 cm interval of the morphological samples in the Abyssal, Pockmark, and Reference areas.
Supplementary Table 8 | Average (Avg.) and standard deviation (St. dev.) for alpha-diversity metrics (Richness, Shannon, Simpson, Rarity) of nematodes of 0–1 cm interval of the morphological samples in the Abyssal, Pockmark, and Reference areas.
Supplementary Table 9 | Results of statistical testing (p-value) for data normality (Shapiro–Wilk, SW), homoscedasticity (Levene), one-way analysis of variance (ANOVA), Kruskal–Wallis (Kruskal), and Tukey’s HSD (Honestly Significant Difference) for pairwise comparisons of alpha diversity metrics (Richness, Shannon, Simpson, Rarity) of nematodes of 0–1 cm interval of the morphological samples in the Abyssal, Pockmark, and Reference areas. Significant values indicated in gray (α = 0.05).
Supplementary Table 10 | Average (Avg.) and standard deviation (St. dev.) for Phylogenetic Diversity (PD) for Nematoda, Genus-assigned, All Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in Abyssal, Pockmark, and Reference areas along the sediment vertical profile.
Supplementary Table 11 | Results of statistical testing (p-value) for data normality (Shapiro–Wilk, S-W.Ar./S-W.Int. for Area and Interval, respectively), homoscedasticity (Levene) and two-way analysis of variance (ANOVA) for Phylogenetic Diversity (PD) in Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 12 | Tukey’s HSD (Honestly Significant Difference) test results (p-value) for pairwise comparisons of Phylogenetic Diversity (PD) in Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 13 | Results of statistical testing (p-value) for analysis of multivariate homogeneity of group dispersions (PERMDISP) and two-way permutational analysis of variance (PERMANOVA) for unweighted UniFrac distances in Nematoda, Unassigned, Genus-assigned, Acantholaimus, Desmoscolex, and Halalaimus ASVs in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 14 | Results (p-value) of pairwise comparisons between group levels with Bonferroni corrections for multiple testing of unweighted UniFrac distances in Nematoda, Genus-assigned, Acantholaimus, and Desmoscolex ASVs in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 15 | Observed, simulated mean and variance, lower/upper-tail p-values (two-tailed t-test) and standardized effect size (ses) for checkerboard pairs (CPs) and C-score for Acantholaimus, Desmoscolex, and Halalaimus ASVs in the Abyssal, Pockmark, and Reference areas. Significant values indicated in gray (α = 0.05).
Supplementary Table 16 | Observed, simulated mean and variance, lower/upper-tail p-values (two-tailed t-test) and standardized effect size (ses) for checkerboard pairs (CPs) and C-score for Acantholaimus, Desmoscolex, and Halalaimus ASVs in the Abyssal, Pockmark, and Reference areas along the sediment vertical profile. Significant values indicated in gray (α = 0.05).
Supplementary Table 17 | Statistical testing of normality (Shapiro–Wilk, S-W) and non-zero means (t-test) for standardized effect size of Phylogenetic Diversity (ses.PD), Mean Pairwise Distance (ses.MPD), and Mean Nearest Taxon Distance (ses.MNTD) for Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, Halalaimus, and Other Genera ASVs in the Abyssal, Pockmark, and Reference areas. Non-parametric equivalents (Wilcoxon signed-rank test, Wilcox) used when normality assumption was violated. Significant values indicated in grey (α = 0.05).
Supplementary Table 18 | Statistical testing of normality (Shapiro–Wilk, S-W) and non-zero means (t-test) for standardized effect size of Phylogenetic Diversity (ses.PD), Mean Pairwise Distance (ses.MPD), and Mean Nearest Taxon Distance (ses.MNTD) for Nematoda, Genus-assigned, Unassigned, Acantholaimus, Desmoscolex, and Halalaimus ASVs in the Abyssal, Pockmark and Reference areas per 1 cm intervals. Non-parametric equivalents (Wilcoxon signed-rank test, Wilcox) used when normality assumption was violated. Significant values indicated in gray (α = 0.05).
Supplementary Table 19 | Description of null models applied in the calculation of standard effect size (ses) of Phylogenetic Diversity (ses.PD), unweighted Mean Pairwise Distance (ses.MPD), and unweighted Mean Nearest Taxon Distance (ses.MNTD) for Nematoda, Genus-assigned, Unassigned, Desmoscolex, Acantholaimus, and Halalaimus ASVs Abyssal, Pockmark, and Reference areas. Source: R Package “picante” vignette.
Supplementary Table 20 | Patterns and processes of phylogenetic community structure and co-occurrence patterns.
Footnotes
References
Bernhard, J. M., Buck, K. R., Farmer, M. A., and Bowser, S. S. (2000). The santa barbara basin is a symbiosis oasis. Nature 403, 77–80. doi: 10.1038/47476
Bezerra, T.N., Decraemer, W., Eisendle-Flöckner, U., Hodda, M., Holovachov, O., Leduc, D., et al. (2021). Nemys: World Database of Nematodes. Available online at: http://nemys.ugent.be (accessed January 25, 2021).
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P. (2016). DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583. doi: 10.1038/nmeth.3869
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high- throughput community sequencing data. Nature 7, 335–336. doi: 10.1038/nmeth0510-335
Cavender-Bares, J., Keen, A., and Miles, B. (2006). Phylogenetic structure of floridian plant communities depends on taxonomic and spatial scale. Ecology 87, 109–122.
Cavender-Bares, J., Kozak, K. H., Fine, P. V. A., and Kembel, S. W. (2009). The merging of community ecology and phylogenetic biology. Ecol. Lett. 12, 693–715. doi: 10.1111/j.1461-0248.2009.01314.x
Darriba, D., Posada, D., Kozlov, A. M., Stamatakis, A., Morel, B., and Flouri, T. (2019). ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 37, 291–294. doi: 10.1093/molbev/msz189
Diamond, J. M. (1975). “Assembly of species communities,” in Ecology and Evolution of Communities, eds M. L. Cody and J. M. Diamond (Cambridge: Harvard University Press), 342–444.
Faith, D. P. (1992). Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10. doi: 10.1016/0006-3207(92)91201-91203
Fontanier, C., Garnier, E., Brandily, C., Dennielou, B., Bichon, S., Gayet, N., et al. (2016). Living (stained) benthic foraminifera from the Mozambique Channel (eastern Africa): exploring ecology of deep-sea unicellular meiofauna. Deep. Res. Part I Oceanogr. Res. Pap. 115, 159–174. doi: 10.1016/j.dsr.2016.06.007
Fontanier, C., Mamo, B., Toucanne, S., Bayon, G., Schmidt, S., De, B., et al. (2018). Are deep-sea ecosystems surrounding Madagascar threatened by land-use or climate change? Deep. Res. Part I Oceanogr. Res. Pap. 131, 93–100. doi: 10.1016/j.dsr.2017.11.011
Gaever, S., Van, Olu, K., Derycke, S., Vanreusel, A., Van Gaever, S., et al. (2009). Metazoan meiofaunal communities at cold seeps along the Norwegian margin: influence of habitat heterogeneity and evidence for connection with shallow-water habitats. Deep. Res. Part I Oceanogr. Res. Pap. 56, 772–785. doi: 10.1016/j.dsr.2008.12.015
Gaever, S., Van, Raes, M., Pasotti, F., Vanreusel, A., Van Gaever, S., et al. (2010). Spatial scale and habitat-dependent diversity patterns in nematode communities in three seepage related sites along the Norwegian Sea margin. Mar. Ecol. 31, 66–77. doi: 10.1111/j.1439-0485.2009.00314.x
Gotelli, N. J. (2000). Null model analysis of species co-occurrence patterns. Ecology 81, 2606–2621. doi: 10.1890/0012-9658(2000)081[2606:NMAOSC]2.0.CO;2
Gotelli, N. J., Hart, E. M., and Ellison, A. M. (2015). EcoSimR: Null Model Analysis for Ecological Data. Available online at: http://github.com/gotellilab/EcoSimR (accessed September, 2016)
Gravel, D., Canham, C. D., Beaudet, M., Messier, C., and Letters, E. (2006). Reconciling niche and neutrality: the continuum hypothesis. Ecol. Lett. 9, 399–409. doi: 10.1111/j.1461-0248.2006.00884.x
Guilini, K., Levin, L. A., and Vanreusel, A. (2012). Cold seep and oxygen minimum zone associated sources of margin heterogeneity affect benthic assemblages, diversity and nutrition at the Cascadian margin (NE Pacific Ocean). Prog. Oceanogr. 96, 77–92. doi: 10.1016/j.pocean.2011.10.003
Hauquier, F., Ingels, J., Gutt, J., Raes, M., and Vanreusel, A. (2011). Characterisation of the nematode community of a Low-Activity cold seep in the recently Ice-Shelf free Larsen B area, Eastern Antarctic Peninsula. PLoS One 6:e22240. doi: 10.1371/journal.pone.0022240
Hentschel, U., Berger, E. C., Bright, M., Felbeck, H., and Ott, J. A. (1999). Metabolism of nitrogen and sulfur in ectosymbiotic bacteria of marine nematodes (Nematoda. Stilbonematinae). Mar. Ecol. Prog. Ser. 183, 149–158. doi: 10.3354/meps183149
Hubbel, S. P. (2001). A Unified Neutral Theory of Biodiversity and Biogeography. Princeton, NJ: Princeton University Press.
Kembel, S. W., and Hubbel, S. P. (2006). The phylogenetic structure of a neotropical forest tree. Ecology 87, 86–99.
Leroy, B., Canard, A., and Ysnel, F. (2013). Integrating multiple scales in rarity assessments of invertebrate taxa. Divers. Distrib. 19, 794–803. doi: 10.1111/ddi.12040
Levin, L. (2005). Ecology of cold seep sediments. Oceanogr. Mar. Biol. An Annu. Rev. 43, 1–46. doi: 10.1201/9781420037449.ch1
Lozupone, C., and Knight, R. (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235. doi: 10.1128/AEM.71.12.8228-8235.2005
Macheriotou, L., Guilini, K., Tania, Bezerra, N., Tytgat, B., and Dinh et al. (2019). Metabarcoding free-living marine nematodes using curated 18S and CO1 reference sequence databases for species-level taxonomic assignments. Ecol. Evol. 9, 1211–1226. doi: 10.1002/ece3.4814
Macheriotou, L., Rigaux, A., Derycke, S., and Vanreusel, A. (2020). Phylogenetic clustering and rarity imply risk of local species extinction in prospective deep-sea mining areas of the Clarion-Clipperton Fracture Zone. Proc. R. Soc. B Biol. Sci. 287:20192666. doi: 10.1098/rspb.2019.2666
Martin, M. (2014). Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12. doi: 10.14806/ej.17.1.200
Mayfield, M. M., and Levine, J. M. (2010). Opposing effects of competitive exclusion on the phylogenetic structure of communities. Ecol. Lett. 13, 1085–1093. doi: 10.1111/j.1461-0248.2010.01509.x
Miklós, I., and Podani, J. (2004). Randomization of presence-absence matrices: comments and new algorithms. Ecology 85, 86–92. doi: 10.1890/03-0101
Moens, T., Braeckman, U., Derycke, S., Fonseca, G., Gallucci, F., Gingold, R., et al. (2014). “Ecology of free-living marine nematodes,” in Handbook of Zoology Vol. 2: Nematoda, ed. A. Schmidt-Rhaesa 109–152.
Mora, C., Roy, K. O., Cordes, E. E., Cunha, M. R., Gaever, S., and Van et al. (2010). The influence of geological, geochemical, and biogenic habitat heterogeneity on seep biodiversity. Mar. Ecol. 31, 51–65. doi: 10.1111/j.1439-0485.2009.00334.x
Obura, D. (2012). The diversity and biogeography of western indian ocean reef-building corals. PLoS One 7:e45013. doi: 10.1371/journal.pone.0045013
Ott, J. A., Novak, R., Schiemer, F., Hentschel, U., Nebelsick, M., and Polz, M. (1991). Tackling the sulfide gradient: a novel strategy involving marine nematodes and chemoautotrophic ectosymbionts. Mar. Ecol 12, 261–279. doi: 10.1111/j.1439-0485.1991.tb00258.x
Pape, E., Bezerra, T. N., Vanneste, H., Heeschen, K., Moodley, L., Leroux, F., et al. (2011). Community structure and feeding preference of nematodes associated with methane seepage at the Darwin mud volcano (Gulf of Cádiz). Mar. Ecol. Prog. Ser. 438, 71–83. doi: 10.3354/meps09278
Platt, R. M., Warwick, H. M., and Platt, R. M. (1988). “Free-living marine nematodes,” Part II: British Chromadorids. Pictorial Key to World Genera and Notes for the Identification of British Species, Synopses of the British Fauna (New Series), eds D. M. Kermack and R. S. K. Barnes (Cambridge University Press).
Price, M. N., Dehal, P. S., and Arkin, A. P. (2010). FastTree 2 - Approximately maximum-likelihood trees for large alignments. PLoS One 5:e9490. doi: 10.1371/journal.pone.0009490
Sanchez, N., Zeppilli, D., Baldrighi, E., Vanreusel, A., Gasimandova, M., Brandily, C., et al. (2020). A threefold perspective on the role of a pockmark in benthic faunal communities and biodiversity patterns. Deep. Res. Part I. Oceanographic Res. Pap. 167:103425. doi: 10.1016/j.dsr.2020.103425
Schliep, K. P. (2011). phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593. doi: 10.1093/bioinformatics/btq706
Seinhorst, J. W. (1959). A rapid method for the transfer of nematodes from fixative to anhydrous glycerin. Nematologica 4, 67–69. doi: 10.1163/187529259X00381
Sibuet, M., Vanreusel, A., Gaever, S., Van, Van Gaever, S., Galéron, J., et al. (2009). Deep-sea habitat heterogeneity influence on meiofaunal communities in the Gulf of Guinea. Deep. Res. Part II Top. Stud. Oceanogr. 56, 2259–2269. doi: 10.1016/j.dsr2.2009.04.008
Siles, G., Rey, P. J., Alca, J. M., Verdu, M., Verdú, M., Rey, P. J., et al. (2009). Phylogenetic signatures of facilitation and competition in successional communities. J. Ecol. 97, 1171–1180. doi: 10.1111/j.1365-2745.2009.01565.x
Stone, L., and Roberts, A. (1990). The checkerboard score and species distributions. Oecologia 85, 74–79.
Ternon, J. F., Bach, P., Barlow, R., Huggett, J., Jaquemet, S., Marsac, F., et al. (2014). The mozambique channel: from physics to upper trophic levels. Deep. Res. Part II Top. Stud. Oceanogr. 100, 1–9. doi: 10.1016/j.dsr2.2013.10.012
Vamosi, S. M., Heard, S. B., Vamosi, J. C., and Webb, C. O. (2009). Emerging patterns in the comparative analysis of phylogenetic community structure. Mol. Ecol. 18, 572–592. doi: 10.1111/j.1365-294X.2008.04001.x
Van Gaever, S., Moodley, L., Beer, D., De, Vanreusel, A., Van Gaever, S., et al. (2006). Meiobenthos at the arctic håkon mosby mud volcano, with a parental-caring nematode thriving in sulphide-rich sediments. Mar. Ecol. Prog. Ser. 321, 143–155. doi: 10.3354/meps321143
Van Gaever, S., Vanreusel, A., Hughes, J. A. A. O., Bett, B. J. J. O., and Kiriakoulakis, K. P. (2004). The macro- and micro-scale patchiness of meiobenthos associated with the Darwin Mounds (north-east Atlantic). J. Mar. Biol. Assoc. United Kingdom 84, 547–556. doi: 10.1017/s0025315404009555h
Vanreusel, A., Fonseca, G., Danovaro, R., Da Silva, M. C., Esteves, A. M., Ferrero, T., et al. (2010a). The contribution of deep-sea macrohabitat heterogeneity to global nematode diversity. Mar. Ecol. 31, 6–20. doi: 10.1111/j.1439-0485.2009.00352.x
Vanreusel, A., Groote, A., De, Gollner, S., and Bright, M. (2010b). Ecology and Biogeography of free-living nematodes associated with chemosynthetic environments in the deep sea?: a review. PLoS One 5:e12449. doi: 10.1371/journal.pone.0012449
Wang, Q., Garrity, G. M., Tiedje, J. M., and Cole, J. R. (2007). Naıve bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267. doi: 10.1128/AEM.00062-67
Warwick, H. M., Somerfield, P. J., and Platt, R. M. (1998). Free-Living Marine Nematodes Part III – Monhysterids. Pictorial Key to World Genera and Notes for the Identification of British Species.
Webb, C. O., Ackerly, D. D., McPeek, M. A., and Donoghue, M. J. (2002). Phylogenies and community ecology. Annu. Rev. Ecol. Syst. 33, 475–505. doi: 10.1146/annurev.ecolsys.33.010802.150448
Webb, C. O., Cornwell, W. K., Kembel, S. W., Blomberg, S. P., Cowan, P. D., Helmus, M. R., et al. (2010). Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26, 1463–1464. doi: 10.1093/bioinformatics/btq166
Wright, E. S. (2016). Using DECIPHER v2.0 to analyze big biological sequence data in R. R. J. 8, 352–359.
Zeppilli, D., Canals, M., and Danovaro, R. (2012). Pockmarks enhance deep-sea benthic biodiversity: a case study in the western Mediterranean Sea. Divers. Distrib. 18, 832–846. doi: 10.1111/j.1472-4642.2011.00859.x
Zeppilli, D., Leduc, D., Fontanier, C., Fontaneto, D., Fuchs, S., Gooday, A. J., et al. (2018). Characteristics of meiofauna in extreme marine ecosystems: a review. Mar. Biodivers. 48, 35–71. doi: 10.1007/s12526-017-0815-z
Keywords: cold seeps, Nematoda, metabarcoding, community structure, phylogenetic community structure
Citation: Macheriotou L, Rigaux A, Olu K, Zeppilli D, Derycke S and Vanreusel A (2021) Deep-Sea Nematodes of the Mozambique Channel: Evidence of Random Community Assembly Dynamics in Seep Sediments. Front. Mar. Sci. 8:549834. doi: 10.3389/fmars.2021.549834
Received: 07 April 2020; Accepted: 18 January 2021;
Published: 12 February 2021.
Edited by:
Chiara Romano, Center for Advanced Studies of Blanes (CSIC), SpainReviewed by:
Lisa Ann Levin, University of California, San Diego, United StatesXiaoshou Liu, Ocean University of China, China
Copyright © 2021 Macheriotou, Rigaux, Olu, Zeppilli, Derycke and Vanreusel. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lara Macheriotou, bGFyYW1hY2hlcmlvdG91QGdtYWlsLmNvbQ==
 Lara Macheriotou1*
Lara Macheriotou1* Sofie Derycke
Sofie Derycke