Abstract
Oceanic environments are one of the largest sources of bioactive molecules, due to the high degree of biodiversity and the innumerable ecological relationships established between macro and microorganisms found in the different ecosystems of these complex environments. Marine organisms are being studied increasingly because they are considered important producers of biologically active peptides. Peptides extracted from marine sources have different functions and structures, when compared to peptides isolated from terrestrial sources, considering the different adaptive pressures undergone by these organisms throughout the evolutionary process. Most bioactive compounds isolated from marine environments are obtained from symbiont microorganisms. Of these microorganisms, bacteria are an important source of bioactive peptides, isolated by metagenomic studies from complex gene networks expressed under marine conditions. Several peptides have been shown to have biotechnological properties such as antimicrobial, antitumor, antihypertensive, anticoagulant, anti-fouling, and antioxidant activity and can be used in the pharmaceutical and food industries. This review article aims to provide an overview of peptides of biotechnological importance, isolated from different phyla of marine organisms, examining the relationship between structure and function of some of these peptides, as well as the ways of extracting, purifying and prospecting new peptides by traditional methods of isolation or sequence analysis in databases. It also intends to list the peptides that are already being produced and used by the industry, in its various branches, and their current state in the market and in clinical tests.
Introduction
For many decades, terrestrial environments have been the subject of studies regarding the prospecting of natural compounds that could be used in the pharmaceutical industry. Meanwhile, although marine environments represent about 70% of the Earth’s surface, they were not explored for biotechnological purposes (Gerwick and Moore, 2012; Wang et al., 2020). Only in the last decades, with the proven therapeutic and anti-infective potential presented by peptides obtained from marine sources, has a vast field of interest opened up for researchers and companies attracted to prospecting for valuable bioactive compounds produced by marine fauna and flora (Satpute et al., 2010).
Marine organisms are the most promising known source of bioactive peptides, since they represent more than half of the global biodiversity (Mora et al., 2011). The oceans present a set of unique circumstances for the development of life, in terms of variations in pressure, salinity, lighting, and temperature, which allowed the spontaneous appearance of various classes of marine peptides – molecules with 3–20 unique amino acid (AA) sequences – isolated from fish, crustaceans, mollusks, algae, corals, and other marine species (Jimeno et al., 2004; Noyer et al., 2011; Ruiz-Ruiz et al., 2017).
Marine ecosystems are marked by hostile conditions for growth and development of microorganism, plant and animal species (Molinski et al., 2009). In these circumstances, the high degree of competitiveness and antagonism between species and the exposure to pathogenic microorganisms allowed the appearance of peptides with significantly different AA compositions, sequences, structures and conformations from molecules of similar function found in terrestrial organisms (Reen et al., 2015a).
Thus, marine peptides usually present multiple bioactivities, such as antimicrobial, anti-hypertensive, antioxidant, anticoagulant, anti-diabetic, anti-obesity, anti-human immunodeficiency virus, anti-proliferative, calcium-binding and cosmetic properties, among others (Mehbub et al., 2014).
Marine peptides with anti-cancer action are especially relevant and are among the first compounds of this nature to be approved for commercial use, a tradition started in 1969 with cytarabine. This synthetic analog of a C-nucleoside from the sponge Tethya crypta, used to treat acute myelocytic leukemia and non-Hodgkin’s lymphoma became the first marine anticancer compound to be commercialized (Sagar et al., 2010). Brentuximab vedotin (SGN-35), a chimeric anticancer antibody fused to a potent antitubulin agent, dolastatin 10, is another example of a marine compound approved for the treatment of Hodgkin’s lymphoma and anaplastic large cell lymphoma (Katz et al., 2011). In this way, marine life is considered as the most promising source of new and bioactive therapeutic and nutraceutical molecules (Laport et al., 2009; Molinski et al., 2009; Blunt et al., 2018).
Another important aspect of marine peptides is that, in most cases, their strong bioactivities are uncertain and may not be related to their function in situ (Katz et al., 2011). This is a consequence attributed to the complex modulation of metabolic pathways used for the biosynthesis of marine molecules. Therefore, some sea peptides may have a specific role for the survival of the native organism and another different biotechnological activity to be explored ex situ (Reen et al., 2015b; Sable et al., 2017).
Although a large number of bioactive peptides have already been isolated from marine life, it is believed that many special molecules are yet to be discovered in the oceans (Laport et al., 2009; Mehbub et al., 2014). Some of the bottlenecks for the isolation and characterization of marine peptides are the need to improve the protocols for the extraction and purification of these molecules from their primary sources and the restricted existing manufacturing techniques that limit the use of a greater number of marine peptides for human health and nutrition (Romano et al., 2017).
Despite these bottlenecks, advances in genomics, proteomics and mainly metagenomics have been expanding the possibilities for the isolation of bioactive peptides (Wilson and Piel, 2013). The full biotechnological potential of previously uncultivated recalcitrant marine microorganisms is being revealed with increasing speed, as the information about gene expression profiles from symbionts is now increasingly accessible, thanks to cultivation-independent metagenomic approaches (Wilson and Piel, 2013; Bashir et al., 2014).
The opening of the metagenomic padlock has revealed that many marine symbionts are capable of synthesizing even more bioactive peptides than previously thought or isolated. Another interesting conclusion is that the clusters of genes in these microbes tend to be downregulated under conventional culture conditions (Chiang et al., 2011).
This review provides an overview of the structure and function of bioactive peptides isolated from marine organisms, as well as the new prospecting and isolation strategies and the relationship between peptide structure and function. Given the large number of active compounds already isolated from marine organisms, we will list here some of the marine peptides characterized in recent decades. Figure 1 illustrates the relationship between the activity of the peptides and the phyla from which they were isolated. As seen in Figure 1, the phyla Annelida, Arthropoda, Echinodermata, and Chordata stand out for the large number of antimicrobial peptides (AMPs) isolated from the organisms in these groups. Peptides isolated from marine organisms can also be immunomodulators, probiotics, anti-obesity, analgesics, anti-fouling, photo-protective, insecticidal, antidiabetic and human food supplements, seen in Figure 1 as “Other.” Supplementary Table 1 describes the peptides isolated from marine organisms which showed the best results on minimal inhibitory concentrations and cytotoxicity.
FIGURE 1
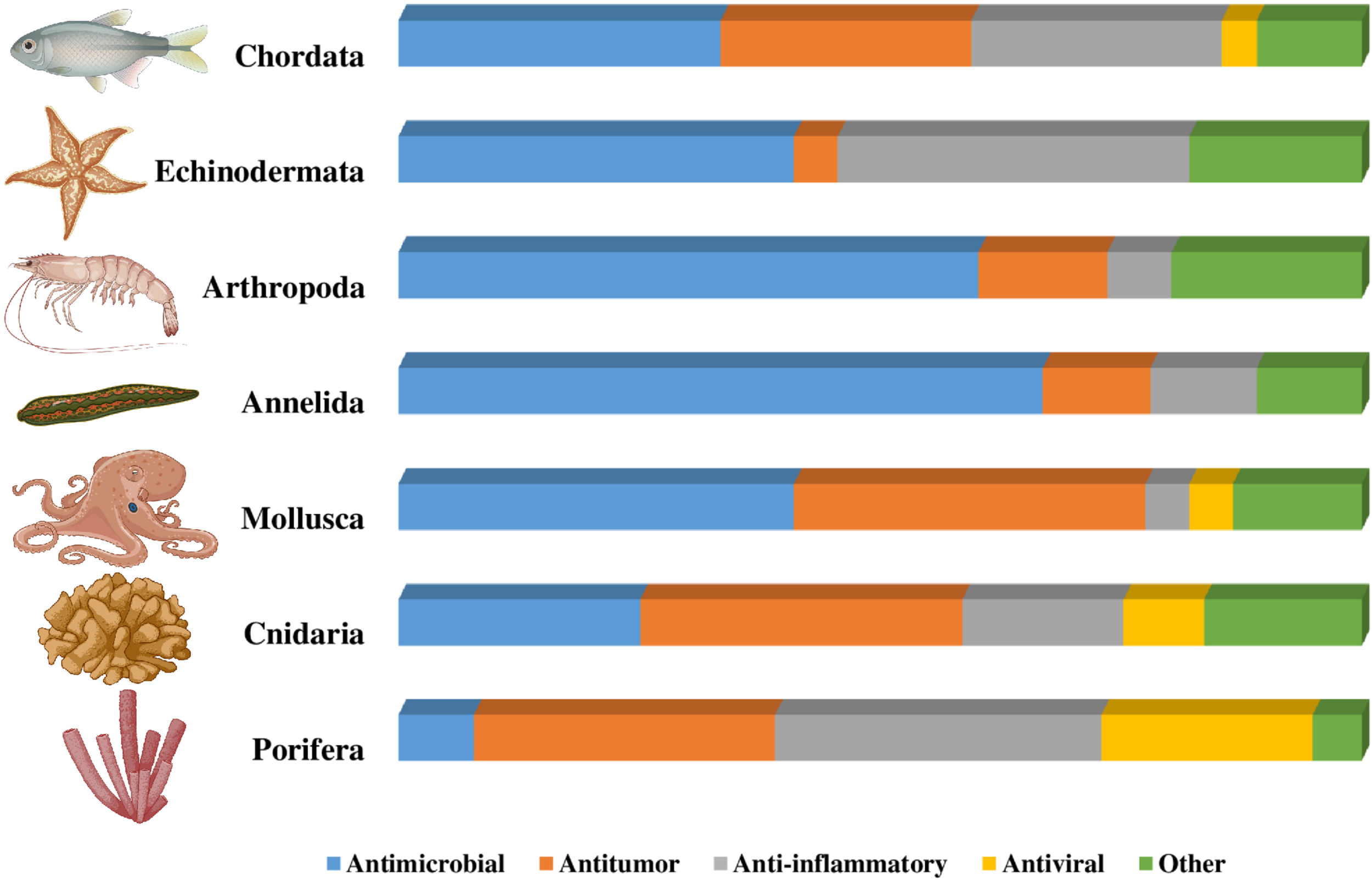
Distribution of peptides according to phylum and function. The figure shows the distribution of the peptides mentioned in this review according to the phyla described and the functionality of the peptides isolated from marine organisms. The bar marked “Other” includes peptides with analgesic, anti-fouling, photo-protective, insecticidal, anti-diabetic, immunomodulatory, anti-obesity, and protiotic activity, and human food supplements.
Prospecting Bioactive Peptides From Marine Organisms
Bioactive molecules have been cited in almost all marine organisms, which can be synthesized by the organism itself or by symbiotic microorganisms (Noyer et al., 2011). Marine sponges appear to be one of the most promising targets for prospecting for bioactive peptides. In the past five decades, experimental evidence has shown that many peptides isolated from sponges have been isolated from symbiotic microorganisms (Schneemann et al., 2011), notably bacteria (Blunt et al., 2012; Teta et al., 2013). An exceptional bacterial biodiversity, represented by about 47 bacterial phyla associated with the body of the sponges, showed that approximately one third of these bacterial isolates were peptide producers with antimicrobial action against Bacillus subtilis (Noyer et al., 2011; Sacristán-Soriano et al., 2011; Reveillaud et al., 2014).
Symbionts producing bioactive peptides are also associated with other marine organisms (notably mollusks and cnidarians) and also with plants and algae, with a fruitful production of these molecules (Laport et al., 2009). Between 2000 and 2010, the annual number of new bioactive peptides with a wide range of activities, isolated from marine symbionts, surpassed the incredible mark of 200 compounds, confirming the relevance of these microbes as the main sources of molecules with biotechnological potential (Laport et al., 2009; Qian et al., 2015).
The success of discovering bioactive peptides from the marine environment may be affected by several factors, such as the availability of organisms to collect, seasonal variations, geographic localization, organism life stage, age, sex, and health condition (Sperstad et al., 2011; Smith et al., 2019). The limited number of specimens that can be collected associated with a low concentration of active compounds in tissues makes this task a challenge. Another option is the use of commercial marine organisms for prospecting (Sharma et al., 2009; Zhong et al., 2013; Zhang et al., 2014, 2015; Pavlicevic et al., 2020).
The purification process is necessary for the characterization of a new bioactive peptide, which involves structure elucidation and activity tests. That requires adequate amounts of the pure peptide. Due to the high diversity of marine organism sources and marine peptides, there is no standardized extraction protocol, directly depending on the source (Sperstad et al., 2011; Pavlicevic et al., 2020).
After collection, a sample preparation may include tissue separation, trituration or cell disruption. After that, an initial treatment may be performed by solid-liquid extraction using organic solvents like methanol, butanol, or ethyl acetate, followed by concentration and solvent partitioning. Precipitation, centrifugation, and membrane filtration may be combined for initial separations and can remove particulate material and large biomolecules, such as proteins and polymers. Size exclusion chromatography, ion-exchange chromatography, and solid-phase extraction can be employed to remove inorganic molecules and fatty acids from extracts (Cheung et al., 2015; Wang X. et al., 2017).
Additionally, high-performance liquid chromatography using reverse phase (RP-HPLC) is typically used as a final purification step, which can separate active from inactive peptides, guided by activity assays. There are advantages and disadvantages in each purification technique. Figure 2 summarizes the main strategies to prospect bioactive peptides from marine organisms discussed below (Sperstad et al., 2011; Cheung et al., 2015; Wang X. et al., 2017).
FIGURE 2
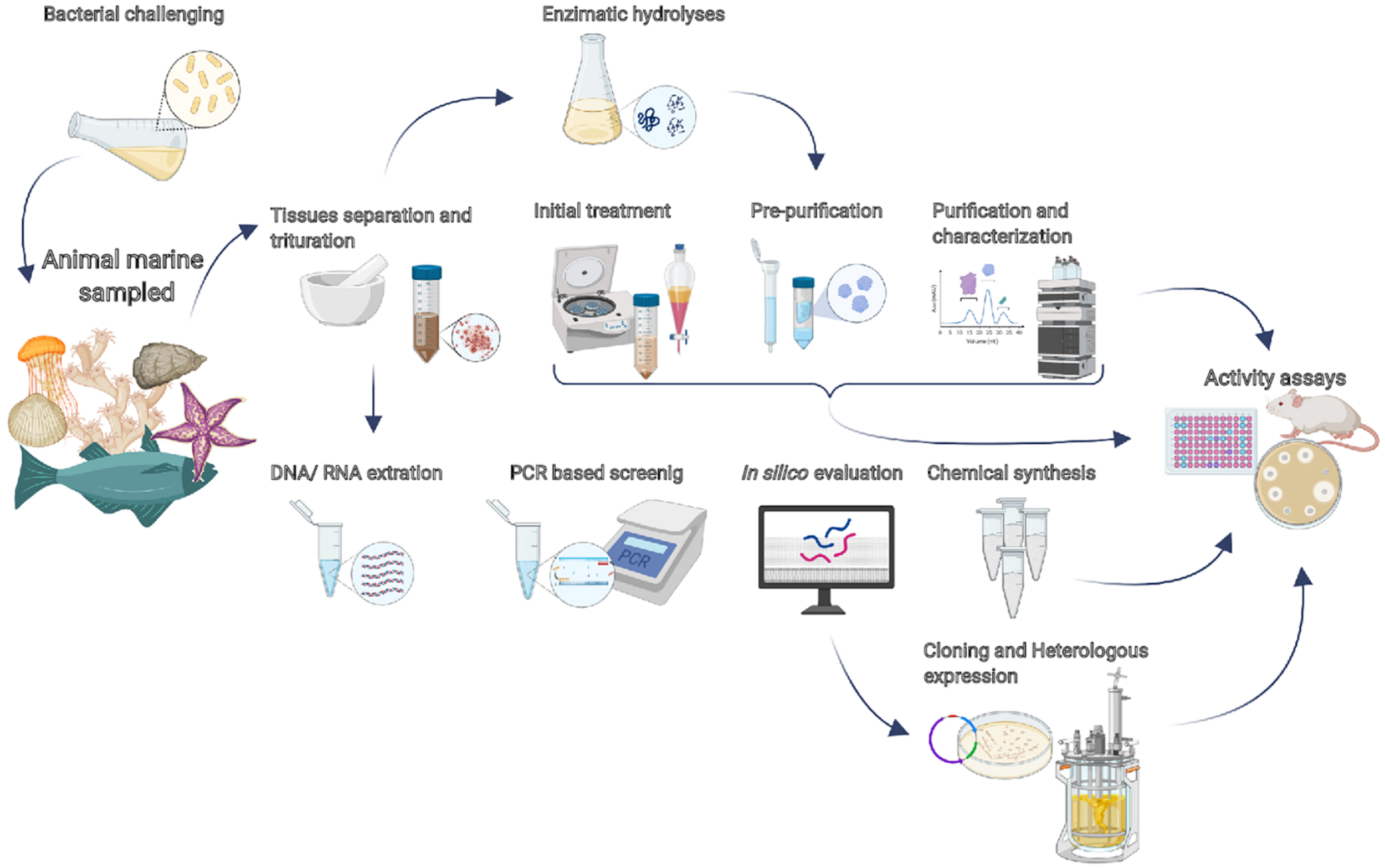
Current strategies for prospecting bioactive peptides from marine organisms. A basic approach is the separation and trituration of the tissues, followed by initial treatment (organic extraction, concentration, and/or partitioning). The pre-purification step may employ size exclusion chromatography, ion-exchange chromatography, and/or solid-phase extraction. All those steps can be guided by activity assays. RP-HPLC is typically used as a final purification step for obtaining the purified peptide that will be able to be characterized. Alternatively, after tissue separation, enzymatic hydrolysis could be employed to prospect bioactive peptides encrypted into proteins. Additionally, the search for bioinspired sequences is performed with PCR-based screening. The bacterial challenge consists of submitting the organisms to pathogenic bacteria, which promote the production of AMPs.
A traditional approach for the discovery of active peptides consists firstly in producing an extract from the organism and screening for specific bioactivity, such as antibacterial, antifungal, antitumoral, etc. In the next step, the bioactive extracts are fractionated using a bioassay-guided approach and finally purified to a single active peptide (Wang X. et al., 2017). A purification approach could include solid-phase extraction followed by reverse-phase high-performance liquid chromatography (Rajanbabu et al., 2015). However, some works have reported the direct purification from plasm by RP-HPLC system (Liao et al., 2013; Qin et al., 2014).
Another approach for prospecting AMPs from marine organisms is the search for bioinspired sequences using specific primers. This method was used by Wei et al. (2015) to identify cathelicidins from sea snakes. Those molecules have never been reported by this group. Therefore, in this work, the total RNA of the snake’s venom glands was extracted using a PCR-based screening method. After analyzing the peptide sequences in silico, the peptides were synthesized and their antimicrobial potential assessed (Wei et al., 2015). Another bioinspired AMP is clavanin MO, a variant of the amphipathic alpha-helical peptide clavanin A. Initially isolated from the tunicate Styela clava, this variant was successfully expressed in Pichia pastoris. The addition of 5 AA residues in the original sequence has improved its antimicrobial activity. Moreover, clavanin MO has also presented antiviral, antitumor, and immunomodulatory activities (Mulder et al., 2015).
The technique known as bacterial challenge includes exposing the organism directly to the pathogen which induces the production of AMPs (Zhao et al., 2010; Bo et al., 2011; Li et al., 2013; Zhang et al., 2015). Tissues are then collected, and a cDNA library is constructed. Sequencing using specific primers allows candidate AMP sequences to be identified (Shike et al., 2002). Through the bacterial challenge, Li et al. (2013) evaluated the gene expression of hepcidin (an AMP) in the liver, spleen, and head kidney of common carp (Cyprinus carpio L.) infected by Vibrio anguillarum (Li et al., 2013). Other works report the antimicrobial activity of chemically synthesized AMPs isolated by bacterial challenge (Zhang et al., 2014; Liu et al., 2018). The work of Chen et al. (2020) reports the heterologous expression of a hepcidin gene (CiHep) from grass carp (Ctenopharyngodon idella) in E. coli BL21 and purification by affinity chromatography (Chen et al., 2020).
A different strategy is the search for so-called intragenic or encrypted peptides, which are internal sequences of proteins that present biological activity (Lemes et al., 2016; Mariano et al., 2021). The prospection of those molecules could consist of an in silico alignment of conserved sequences of encrypted peptides against the peptides of interest (Brand et al., 2019; Mariano et al., 2021) or by enzymatic hydrolyses of a secretion, tissue, or a crude protein extract of an organism followed by activity assays and fractionation (Lemes et al., 2016; Lee and Hur, 2017). The latter technique was used to identify two antiproliferative peptides from tuna dark muscle after its hydrolyses with papain and protease XXIII. The hydrolysates were purified by gel filtration and HPLC and showed two peptides that were active against the human breast cancer cell line (MCF-7) with the half maximal inhibitory concentration (IC50) values of 8.1 and 8.8 μM, respectively (Hsu et al., 2011). Another example was the isolation of antioxidative and angiotensin I convert enzyme (ACE) inhibitory peptides from the small red scorpionfish (Scorpaena notate). The heads were cooked and hydrolyzed with a protease from the fungus Penicillium digitatum. After ultra-filtration, the peptides were purified by size exclusion chromatography followed by RP-HPLC (Aissaoui et al., 2017).
Thus, although the strategy for obtaining bioactive encrypted peptides from protein hydrolysate has been widely reported in marine organisms (Sila and Bougatef, 2016; Lee and Hur, 2017), the guided search for potential peptides in silico has been limited, probably due to factors such as the low availability of these organisms’ genome and proteome sequences in databases (Suarez-Jimenez et al., 2012; Robert et al., 2014).
Developments in sequencing technologies, allowing access to a huge amount of data at an increasingly low cost, can also be a good strategy for the prospection of bioactive peptides (Woodhouse et al., 2013; Trindade et al., 2015). Furthermore, metagenomics techniques provide means to explore the potential bioactive compounds from any mixed population of microbes, including the access of data provided by investigation of microorganisms that still can‘t be cultivated in vitro culture-dependent techniques are unable to access (Giordano et al., 2018; Mahapatra et al., 2020).
Microbiota studies of marine symbionts allow direct access to the so-called total “environmental” DNA (eDNA) (Wilson and Piel, 2013; Romano et al., 2017). In this context, the metagenomic approach for the discovery of bioactive marine peptides is eminently applied to bacteria and fungi, relatively easily, divided into three major stages: (i) obtaining samples from the natural environment; (ii) the in vitro enrichment of the target microorganisms, and (iii) the massive extraction of genomic DNA from the selected microorganisms, followed by bioinformatics analyses (Xiong et al., 2013). These analyses can be performed by high-throughput sequencing, assembly of contigs, genomic annotation, in silico identification of biosynthetic pathways and by overexpression of peptides in a heterologous manner. Another approach is the direct screening of functional biomolecules from the construction of metagenomic libraries or by the identification of sequences by homology-based comparison (Giordano et al., 2018; Mahapatra et al., 2020).
Transcriptomics can profile the transcriptome of a marine organism and has several advantages over sequencing the whole genome. This method is highly sensitive, cost-effective and allows deeper sequence coverage. Also, there is much sequencing data from various marine organisms currently available at the National Center for Biotechnology Information (NCBI) database.
A transcriptomics project can be described as a pipeline (Figure 3), starting with RNA isolation from the marine organism. This initial step is often the bottleneck in the whole process, as difficulties may be encountered to obtain pure material with the minimum quantity required for sequencing. Next, the RNA is reversely transcribed into cDNA and sent for sequencing. The choice of the Next Generation Sequencing (NGS) platform will depend on the size of the transcriptome and also on the objective of the study. When low-expressed peptides are desired, there is a need for higher sequencing coverage. Paired-end sequencing should be the preferred choice, as it allows a better assembly of the transcriptome, especially by covering repetitive regions. Then, after sequencing, the raw data need to be preprocessed to remove low quality sequences. In the following step, the transcriptome is assembled and then the final reads are aligned (using BLASTx) to protein databases, such as NCBI RefSeq. Finally, the mapped sequences are functionally annotated. A search can be made in this final table using the names of the peptides of interest as queries.
FIGURE 3
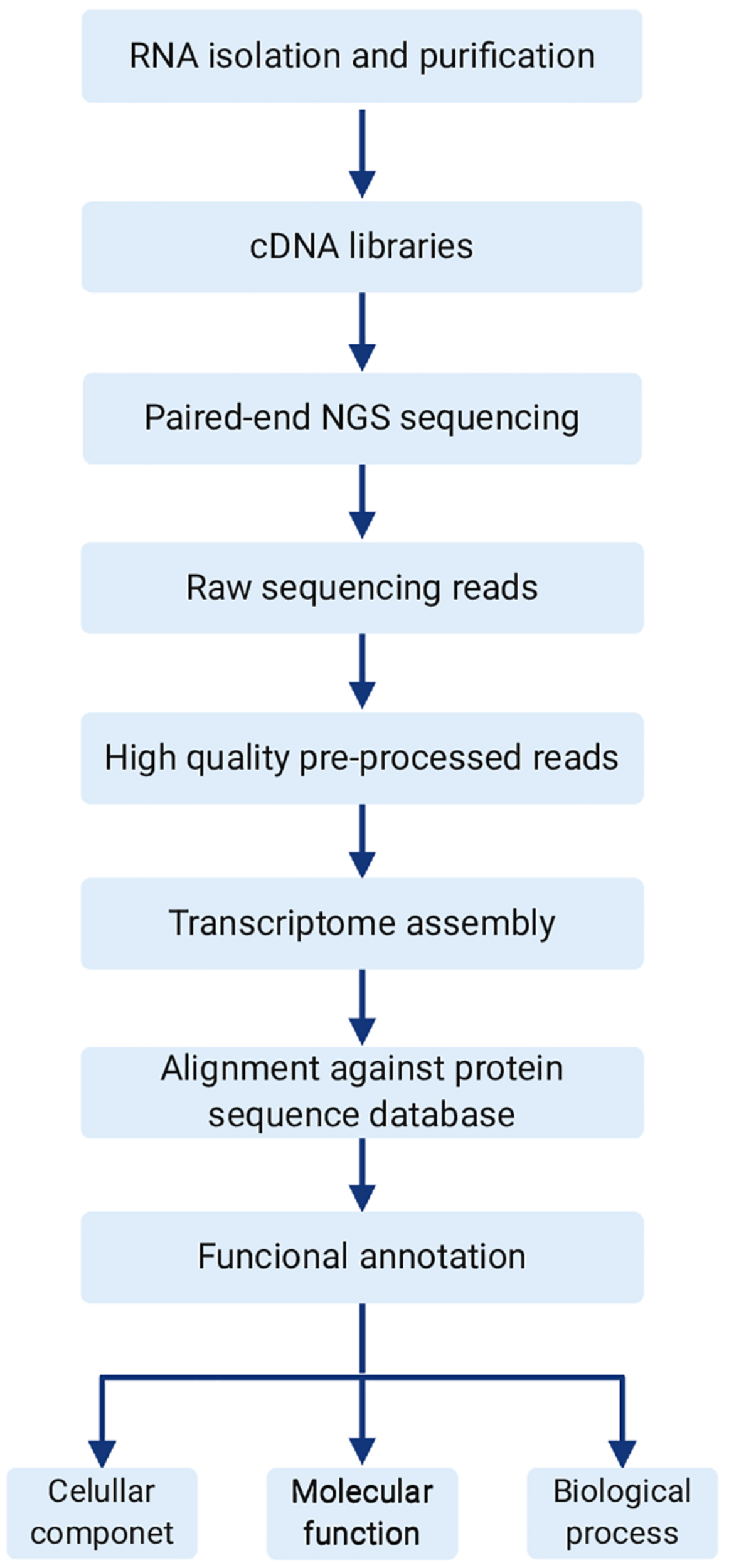
General transcriptome pipeline. The workflow typically starts with total RNA extraction and purification. Then the cDNA library is constructed using an appropriate kit, prior to NGS paired-end sequencing. Next is pre-processing of raw data in order to obtain high quality reads for transcriptome assembly. Finally, assembled transcripts are aligned to a protein database of choice, and functionally annotated.
Koch and Grimmelikhuijzen (2020) used the assembled genomes and transcriptomes from several hexacoral and ceriantharian species to compare neuropeptide preprohormones and trace their origin. This allowed the identification of three cnidarian neuropeptide families (X1PRX2 amides, GRFamides and GLWamides) that likely originated in the common cnidarian ancestor. Zhang et al. also identified and characterized neuropeptides in Patinopecten yessoensis, contributing to a complete understanding of the roles of these molecules in endocrine regulation in bivalve mollusks (Zhang et al., 2018). Besides increasing our knowledge of neuropeptide evolution, these studies also contributed to the creation of peptide inventories, useful resources for subsequent experiments.
The combination of transcriptomics and proteomics research over the past few years has also resulted in significant advances in the identification of potential drugs from marine toxins, while improvements in liquid chromatography-tandem mass spectrometry made this task more achievable (Xie et al., 2017). The importance of research in this area can be exemplified by the identification of bioproducts capable of interfering with specific pathways, allowing the development of anti-cancer treatments (Rodrigo et al., 2021). Rodrigo et al. (2021) discovered that a cocktail of proteins from Eulalia sp. is more toxic toward an ovarian cell line (A2780), supporting the potential of mixed proteins in venoms against aggressively proliferating cells.
However, although the biodiversity of the oceans represents a promising source of bioactive peptides, much information about the sequences of these molecules remains scattered around NCBI sequence repositories. Therefore, further advances in this field depend on the development of toxin databases for marine venomous species, containing reference sequences for alignment with newly sequenced transcriptomes. To this end, Sun et al. (2016) have released the Fish-T1K Project, which aims to sequence the transcriptomes of 1,000 fish species and make the database available to the community. From the website, it is possible to make BLAST searches with sequences of interest and identify the unknown peptides from the annotated sequences.
Thereby, several strategies of prospection are available today. Research and planning concerning the target molecule, collection site, and sampled organism are of great importance and must be carefully considered. Table 1 shows examples of peptides obtained from marine organisms over the last 10 years, using the main strategies for prospecting discussed here.
TABLE 1
| Strategy of prospection | Source | Origin | Peptide | Activity | Reference |
| The crude mucus was lyophilized and resuspended in PBS Gel filtration chromatography followed by RP-HPLC | Achatina fulica (Mollusca) | Mucus | Mytimacin-AF | Antimicrobial | Zhong et al., 2013 |
| Plasm was directly purified in a RP-HPLC system | Mytilus coruscus (Mollusca) | Hemolymph | Myticusin-1 | Antimicrobial | Liao et al., 2013 |
| Mytilus coruscus (Mollusca) | Hemolymph | Mytichitin-CB | Antimicrobial | Qin et al., 2014 | |
| Heterologous expression | Venerupis philippinarum (Mollusca) | Hemocytes | Clam defensin (VpDef) | Antibacterial | Zhang et al., 2015 |
| Heterologous expression/chemical synthesis | Scatophagus argus (Chordata) | Liver | Hepcidin (SA-hepcidin1 and 2) | Antibacterial and antiviral | Gui et al., 2016 |
| Bio inspiration/chemical synthesis | Hydrophis cyanocinctus (Chordata) | Venom gland | Cathelicidin | Antimicrobial and anti-inflammatory | Wei et al., 2015 |
| Scylla paramamosain (Arthropoda) | Hemocytes | SpHyastatin | Antimicrobial | Shan et al., 2016 | |
| Bio inspiration/heterologous expression | Styela clava (Chordata) | Hemocytes | Clavanin MO | Antimicrobial | Mulder et al., 2015 |
| Bacterial challenge | Scophthalmus maximus (Chordata) | Liver | Hepcidin (SM HEP1p and SM HEP2p) | Antimicrobial | Chen et al., 2007; Zhang et al., 2014 |
| Cyprinus carpio L (Chordata) | Liver | Hepcidin | Antimicrobial | Li et al., 2013 | |
| Platichthys stellatus (Chordata) | Liver | PsHepcidin | Antimicrobial | Liu et al., 2018 | |
| Bacterial challenge/Heterologous expression | Ctenopharyngodon idella (Chordata) | – | CiHepsidin | Antimicrobial | Wei et al., 2018; Chen et al., 2020 |
| Enzymatic hydrolysis | Penaeus vannamei (Arthropoda) | Head | PAB2 and PRB2 | Antiproliferative | Hsu et al., 2011 |
| Scorpaena notate (Chordata) | Head | D-I, F-I and E-I | Antioxidative and ACE inhibitory | Aissaoui et al., 2017 |
Strategies to obtain bioactive peptides from marine organisms.
Bioactive Peptides Isolated From Marine Organisms
Porifera Peptides
The organisms belonging to the phylum Porifera, known as sponges, are the oldest remnants among the metazoan group. They are found in large numbers in fresh and saltwater habitats, thanks to the visible ability to adapt to drastic environmental and biological changes. These organisms have a simple level of organization, although their cells are specialized in various functions, but not organized to form tissues and organs (van Soest et al., 2012). Among marine organisms, sponges are the most studied, regarding the production of active peptides, a fact justified by the ease of access and collection of these organisms (van Soest et al., 2012). The early appearance and the sessile way of life of these organisms allowed the development of a group of chemical compounds for communication and modulation of cellular functions and defenses (Braekman and Daloze, 1986). In particular, sponges have become a rich source of new AMPs (Pushpanathan et al., 2013).
In terms of antimicrobial activities (e.g., antibacterial, antifungal and anti-HIV), peptides produced by organisms from the phylum Porifera have been divided into two main classes, including depsipeptides and proline-rich cyclic peptides (Vitali, 2018). These classes differ mainly due to the exclusive cyclic nature of proline-rich cyclic peptide members with numerous proline repetitions that trigger high structural rigidity in these peptides (Wang et al., 2014a). By contrast, depsipeptides can present both linear and cyclic (called cyclic depsipeptides) scaffolds (Plaza et al., 2010). Cyclic depsipeptides with anti-viral activities include callipeltins (Figure 4), mirabamides, celebesides, neamphamide A and papuamides (Cyclodepsipeptides from marine sponges: natural agents for drug research.). These peptides share aliphatic tails, a tyrosine-free hydroxyl group, the presence of a 3,4-dimethylglutamine residue, unusual AA repetitions and N-terminal polyketide-derived moieties (Andjelic et al., 2008; Giordano et al., 2018).
FIGURE 4
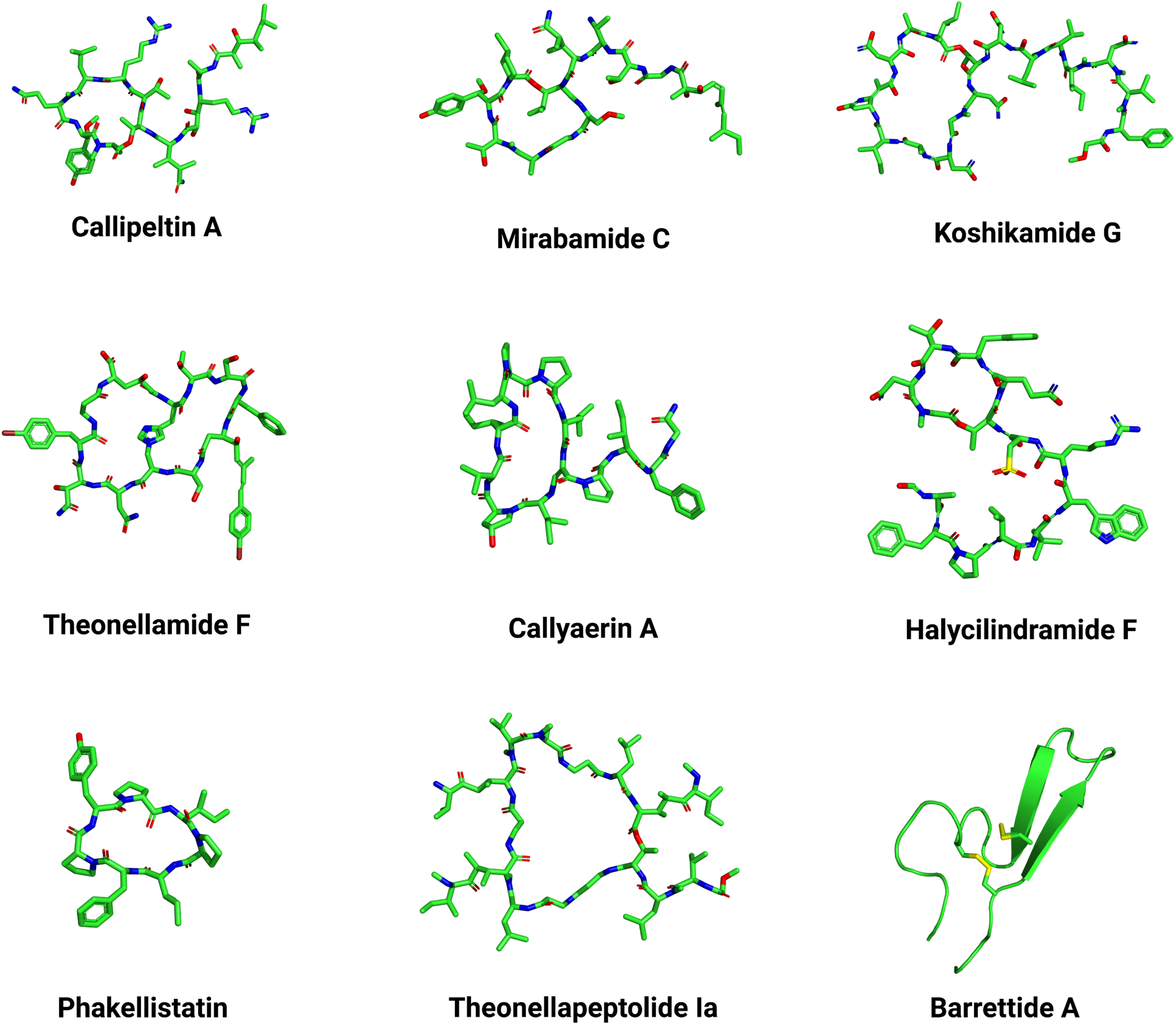
Three-dimensional structures of Porifera peptides. Representative structures with different scaffolds were selected based on the Porifera peptide classes described in the main text, including callipetin A (PubChem ID: 70678682), mirabamide C (PubChem ID: 24179849), koshikamide G (PubChem ID: 101518086), theonellamide F (PubChem ID: 45382310), callyaerin A (PubChem ID: 52941212), halycilindramide F (PubChem ID: 132596267), phakellistatin (PubChem ID: 11765991), theonellapeptolide Ia (PubChem ID: 101601088), and barrettide A (PDB ID: 6CFB). The PubChem.sdf files were parameterized using Maestro v. 11.6.013 (Schrodinger) and further visualized on PyMOL v. 1.8. The three-dimensional structure images were generated on PyMOL v. 1.8.
From studies carried out with the deep-water species Theonella cupola and T. swinhoei, the linear peptides koshikamides C-E and cyclic depsipeptides koshikamides F-H (Figure 4) were isolated. It was reported that koshikamides F and H inhibited the entry of HIV-1 in low concentrations, while their linear forms (koshikamides C and E) were inactive. The authors suggested that HIV-1 inhibition is due to the presence of the lactone ring with 10 AA residues that forms the structure of koshikamides F and H, but is absent in the linear forms of these peptides (Plaza et al., 2010). Celebesides A-C, cyclic depsipeptides that contain the unusual AAs phosphoserine and 3-carbamoyl threonine, were isolated from Siliquariaspongia mirabilis, and both are considered new marine products. The AA phosphoserine was correlated with the presence of anti-HIV activity in celebesides (Plaza et al., 2009; Giordano et al., 2018). Other peptides with anti-HIV activity isolated from S. mirabilis are mirabamides A–D (Figure 4 and Supplementary Table 1) (Plaza et al., 2007; Giordano et al., 2018). New mirabamides (E–H) were isolated from a sponge of the genus Stelletta, and these compounds also showed anti-HIV action (Lu et al., 2011). The peptides stellettapeptins A and B, isolated from Stelletta sp., are cyclic depsipeptides with anti-HIV activity. These compounds were tested in the human T-cell line CEM-SS infected with HIV-1RF, in which the anti-HIV activity was evaluated in an XTT-based cell viability assay. Stellettapeptins A and B showed a potent inhibition effect on HIV-1 infection half-maximal effective concentration (EC50) values of 23 and 27 nM, respectively, which means they can be considered as a family of anti-HIV peptides (Shin et al., 2015).
All these cyclic depsipeptides’ features have led to a broad spectrum of biological targets, including viral surface components. For instance, many studies have reported the influence of phosphatidylserine phospholipids on the viral surface for cyclicdepsitepeptides’ anti-viral effects (Andjelic et al., 2008). As mentioned above, these peptides present an atyrosine hydroxyl group that interacts with cholesterol membrane, whereas the aliphatic tails favor the peptides’ insertion into the viral membrane. Although this mechanism has been well described for papuamide A, the potency of other papuamide and miramabide peptides can differ depending on the unusual AA residues present in their structures (Plaza et al., 2007). For instance, it has been shown that the rhamnosylated β-methyltyrosine residue in mirabamide A has a direct influence on the higher potency of this peptide, compared to mirabamide B, C, and D (Plaza et al., 2007). Moreover, mirabamide B was found to be the least potent variant, as this is the only mirabamide peptide lacking a 2,3-diaminobutanoic acid residue (Plaza et al., 2007). Interestingly, although such differences in the constitution of unusual AA residues do not deeply interfere with the overall structural profile of these peptides, they greatly affect their anti-viral effects. Additionally, divergent anti-viral effects have also been correlated with peptide cyclization, as is the case of koshikamides, which are non-active in their linear form (Plaza et al., 2010). This same correlation was observed when testing linear and cyclic koshikamides against fungi, including C. albicans (Plaza et al., 2010).
The marine sponge Theonella swinhoei has been studied since the 1990s as a source of different active peptides such as motuporins (De Silva et al., 1992), theonellapeptolides (Figure 4) (Kobayashi et al., 1994), theonegramides (Bewley and Faulkner, 1994; Giordano et al., 2018), cyclolithistide A (Clark et al., 1998) and polytheonamides (Hamada et al., 1994, 2005). One of these compounds, the bicyclic peptide theonellamide F (Figure 4), demonstrated antifungal activity against Aspergillus sp., Candida sp., and Trichophyton sp. in concentrations of 3–12 μg/ml (Matsunaga et al., 1989). The mode of action of this peptide is related to the binding of the histidine-alanine bridge present in its structure that binds to 3-β-hydroxisterols, causing damage to the plasma membrane (Nishimura et al., 2010; Giordano et al., 2018). Theonellamide G (Supplementary Table 1), a glycopeptide also isolated from this sponge, demonstrated cytotoxicity when tested against human colon adenocarcinoma cell lines (HCT-16) with an IC50 of 6.0 μM. Besides, it also showed high antifungal activity against the wild strain (ATCC 32354) and the amphotericin-resistant strain (ATCC 90873) of Candida albicans, with an IC50 of 4.5 and 2.0 μM respectively (Youssef et al., 2014).
Structure-activity correlations have also been observed for Porifera-derived peptides with antifungal, antibacterial and anticancer properties (Figure 1). For instance, callyaerins represent Callyspongia aerizusa sponge-derived peptides with cyclic and linear regions in their structures. Among the callyaerins described by Daletos et al. (2015), callyaerin A (Figure 4) and B share most AA residues in their cyclic region (e.g., isoleucine, proline, leucine, valine, and 4-hydroxyproline) and were the most effective variants against Mycobacterium tuberculosis. In particular, the unusual AA 4-hydroxyproline provides more rigidity to the cyclic ring in these peptides, which has been correlated with possible selective protein binding. Moreover, the differences in the overall hydrophobicity in the linear region between those peptides indicate that callyaerin B is more cytotoxic than callyrarin B (Daletos et al., 2015). Callyaerins A–F and H were assessed against the HeLa (human cervical carcinoma), PC12 (rat brain tumor), and tumor cell line L5178Y (mouse lymphoma), in which callyaerin E was the most active and callyaerin F was the least active (Ibrahim et al., 2010; Giordano et al., 2018). Callyaerin B also showed greater cytotoxicity when tested in the THP-1 cell line (acute human monocytic leukemia) or MRC-5 (human fetal pulmonary fibroblast) (Daletos et al., 2015). Similarly, studies with halycilindramide (Figure 4) have shown that the presence of a macrocyclic lactone ring is required for the antifungal and cytotoxic properties of these peptides (Li et al., 1995). One year later, the identification of halycilindramide E was demonstrated and, instead of presenting a macrocyclic lactone ring at the N-terminus region, it revealed a truncated linear structure with amidated C-terminus. Interestingly, this structural profile resulted in the dissociation of antifungal and cytotoxic activities in halycilindramide E (Li et al., 1996).
Cytotoxic effects have also been reported for marine peptides toward cancer cell lines. Proline-rich cyclic peptides from sponges, including euryjanicin A, carteritins, phakellistatin, and hymenamides, are known to compromise the viability of the human colon cancer cell line (HCT116), human hepatoma (BEL-7402), human lung carcinoma cells (A-549) and HeLA cancer cells (Li Y. et al., 2018; Anand et al., 2019). In general, these peptides present constrained cyclic scaffolds with L-AAs, D-AAs and unnatural AAs in their composition. Interestingly, studies describing the total chemical synthesis of these proline-rich cyclic peptides, although proving their structural and chemical equivalence with their natural counterparts, have shown reduced anticancer effects (Li Y. et al., 2018). This has been attributed to a possible preference of these peptides for a natural, impure condition in which to exert their full activities (Li Y. et al., 2018).
In a study carried out by Tran et al. (2014), four new peptides with antiproliferative and cytotoxic activity were isolated and characterized from Pipestela candelabra, and named milnamides E-G and hemiasterlin D. Another eight known peptides were tested, all isolated from this sponge. These peptides include milnamide A-D (Chevallier et al., 2003; Sonnenschein et al., 2004), hemiasterlin A (Gamble et al., 1999), and geodiamolide D-F (Coleman et al., 1999). All the peptides proved to be efficient against the proliferation of human prostate cancer cells (PC3), and they showed selectivity among cancer cells and human neonatal foreskin fibroblast non-cancer cells (NFF). The results showed that milnamide A and milnamides E–G exhibited greater activity, with IC50 = 11.0 nM (Supplementary Table 1).
Phakellistatins are a group of cycloheptapeptides isolated from sponges of the genus Phakellia. These peptides have been studied for the past two decades due to their strong cytotoxic activity against murine leukemia cells (Meli et al., 2017). The study carried out by Wu et al. (2019) isolated four new cycloheptapeptides from Phakellia fusca. Fuscasins A–D are composed of seven AA residues, including at least one proline residue as well as phakellistatin (Figure 4) family peptides. These four peptides were tested against six human cancer cell lines, but only fuscasin A showed cytotoxic activity against the human cancer cell line HepG2, and it did not show cytotoxicity against non-malignant cells. This result suggests that fuscasin A has the potential to be a targeted antitumor drug.
Compounds isolated from marine sponges were also studied due to their anti-inflammatory activity (Figure 1). This is the case of characellides A and B isolated from Characella pachastrelloides. These peptides are composed of a central tripeptide attached to an alkyl chain ending with a 2,3-dimethyltetrahydropyran and a rare sugar unit. The ability to reduce the production of reactive oxygen species (ROS) in inflammatory conditions by characellides A and B was assessed in microglia BV-2 cells stimulated with lipopolysaccharide (LPS). The microglia-mediated inflammatory process is known for instigating oxidative damage through ROS release. The difference between antioxidant cellular defenses and the production of ROS results in oxidative damage and, consequently, in inflammation. The results showed a 50% reduction in ROS production in the microglia BV-2 cell line stimulated with LPS when treated with characellides (Afoullouss et al., 2019).
A Stylissa massa proline-rich peptide, named stylissatin A (SA), containing both cis- and trans-proline residues in its structure (Kita et al., 2013) showed anti-inflammatory activities in LPS-stimulated murine RAW264.7 macrophages through the inhibitory effects on nitrogen oxide (NO production) (Zhang et al., 2019). This peptide was also assessed for potential inhibition of pre-adipocyte differentiation in Murine 3T3-L1 fibroblasts. Another 23 synthetic SA derivatives were evaluated, and the results showed that SA and its derivatives have potent anti-inflammatory action and inhibit the differentiation of preadipocytes, with the D-Tyr1-tBuSA (EC50 = 12 μM) derivative being six times more potent than SA (EC50 = 73 μM) (Supplementary Table 1). Moreover, diverse stylissatin A has been reported presenting a macrocyclic scaffold, but with key AA substitutions, including the use of D-tyrosine, D-phenylalanine, D-proline, and D-allo-isoleucine (Zhang et al., 2019). D-AAs were submitted to different chemical modifications, including the addition of tert-butyl ether (tBu), propargyl, methyl, and benzyl ethers and D-glucosyl. In terms of structural profile, these modifications revealed that the macrocyclic scaffolds containing L- and D-AAs differ in their conformations and hydrogen bond patterns, thus influencing their anti-inflammatory effects (Zhang et al., 2019).
In addition, peptides isolated from organisms of the phylum Porifera may also have antifouling activity. This is the case of the compounds barettin and 8,9-dihydrobarettin, isolated from the marine sponge Geodia barretti. These cyclic dipeptides were tested for antifouling activity against Balanus improvisus barn larvae, and they were shown to be compatible with the currently used antifouling agents, with EC50 values of 0.9 and 7.9 μM, respectively, but have the advantage of being non-toxic, unlike the compounds already in use (Sjögren et al., 2004, 2006). The compounds barrettides A (Figure 4) and B, also isolated from G. barretti, are disulfide-containing peptides of amphipathic and hydrophobic nature. Both proved to be efficient in terms of antifouling activity for B. improvisus larvae, with EC50 of 0.6 and 6 μM, respectively (Carstens et al., 2015).
Cnidaria Peptides
Organisms such as anemones, jellyfish, and corals make up a numerous, diverse, and ecologically important group of marine invertebrates, the phylum Cnidaria, which houses more than 13,000 described species (Turk and Kem, 2009). The cnidarians have radial symmetry, formed by two layers of cells, the ectoderm and the endoderm, separated by a non-cellular matrix, the mesoglea. These organisms have already been extensively studied for their capacity to produce toxins and poisons (Turk and Kem, 2009), but recent research shows that they are capable of producing interesting active molecules, with more than 3,000 products described as starting from this phylum (Daly et al., 2007).
Peptides isolated from invertebrate marine organisms from the phyla Cnidaria, Mollusca, Annelida, Arthropoda, and Echinodermata have also shown antimicrobial properties, mainly against bacteria (Figure 1). Interestingly, as observed for bioactive peptides from the phylum Porifera, constrained structural profiles are commonly found in those organisms. For instance, the antibacterial peptide aurelin (Figure 5) from marine cnidarian Aurelia aurita consists of a defensin-like peptide with six cysteine residues forming three disulfide bonds, and also presenting an unusual link for structure stabilization involving cysteine, aspartic acid, and lysine residues (Ovchinnikova et al., 2006). A study carried out with arelin showed antimicrobial activity against Gram-positive bacteria [Listeria monocytogenes, with minimum inhibitory concentration (MIC) 22.6 μg/ml] and Gram-negative bacteria (Escherichia coli, strain ML-35p, MIC 7.6 μg/ml) (Ovchinnikova et al., 2006). The set of six cysteines present in this peptide forms a domain identified in other proteins isolated from vertebrates and invertebrates. Although the function of this domain is not so clear, it is estimated that it acts as an extracellular ligand or as a toxin, blocking the K + channel (Ovchinnikova et al., 2006; Shenkarev et al., 2012). Similarly, two mollusk-derived peptides, named mytichitin-CB and RpdefB, are also effective in countering bacterial growth, which has been attributed to their constrained defensin-like structures (Novoa et al., 2016; Yang et al., 2018). This cysteine-rich structure profile has also been demonstrated for peptides from the phylum Echinodermata, including the peptide PpCrAMP (Kim et al., 2018).
FIGURE 5
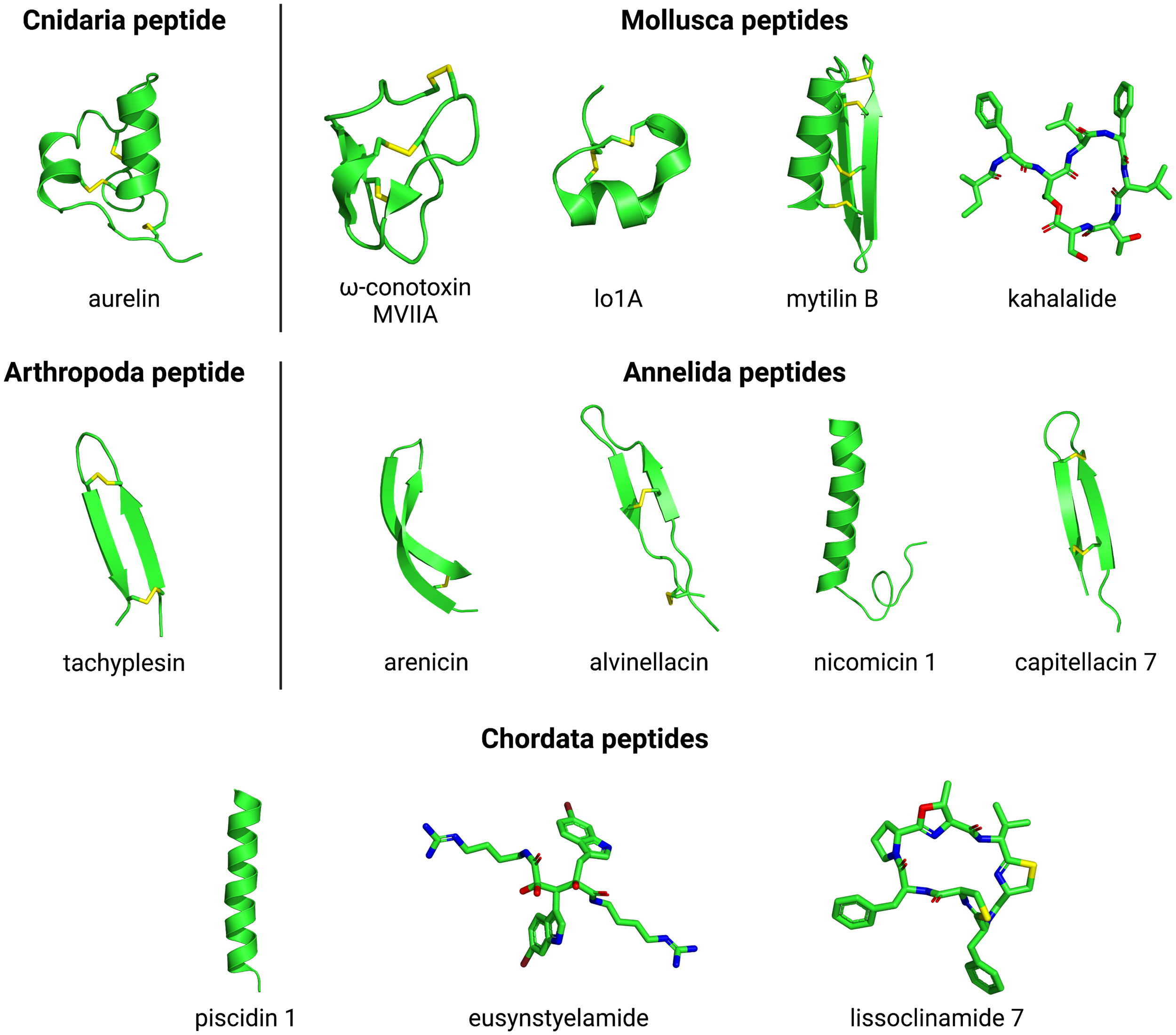
Three-dimensional structures of Cnidaria, Mollusca, Arthropoda, Annelida, and Chordata peptides. The evaluated structures include aurelin (PDB ID: 2LG4) from Cnidaria; omega-conotoxin MVIIA (PDB ID: 1MVI), lo1A (PDB ID: 2MD6), mytilin B (PDB ID: 2EEM) and kahalalide (PubChem ID: 72942) from Mollusca; tachyplesin (PDB ID: 2RTV) from Arthropoda; arencin (PDB ID: 2JSB), alvinellacin (PDB ID: 2LLR), nicomicin 1 (PDB ID: 6HN9) and capitellacin 7 (PDB ID: 7ALD) from Annelida; and piscidin 1 (PDB ID: 2MCV), eusynstyelamide (PubChem ID: 23427997) and lissoclinamide 7 (PubChem ID: 390558) from Chordata. The PubChem.sdf files were parameterized using Maestro v. 11.6.013 (Schrodinger) and further visualized on PyMOL v. 1.8. The three-dimensional structure images were generated on PyMOL v. 1.8.
In research carried out by Lima et al. (2013), a screening of six cnidarians (Carijoa riisei, Muriceopsis sulphurea, Neospongodes atlantica, P. caribeorum, Plexaurella grandiflora, and Phyllogorgia dilatata) was carried out. The crude extract of these cnidarians showed inhibition that varied from 33 to 94% against the bacteria that cause hospital infections Staphylococcus aureus, Proteus mirabilis, Klebsiella pneumoniae, Shigella flexneri, E. coli, Streptococcus pyogenes, and Salmonella typhimurium. From the P. dilatata extract, the peptide Pd-AMP1 was extracted and purified, which presented the best result against S. aureus (Supplementary Table 1).
Many antitumor peptides have been isolated from cnidarians (Figure 1). The Anthopleura anjunae antitumor peptide (AAP-H) was obtained by enzymatic hydrolysis of the alkaline protease of the sea anemone Anthopleura anjunae. It is a pentapeptide with an AA sequence of Tyr-Val-Pro-Gly-Pr. This peptide was investigated for inhibitory effects on the proliferation of DU-145 cells in prostate cancer, and it showed antiproliferative activity (with an IC50 concentration of 9.6, 7.9, and 2.3 mM in 24, 48, and 72 h, respectively), inducing apoptosis, causing a decrease in cancer cell growth, and reduced the percentage of viable cells. It also caused morphological changes in DU-145 cells for DNA fragmentation, the formation of bubbles in the membrane and cell shrinkage (Wu et al., 2018). Another study showed that the antitumor mechanism of this peptide in prostate cancer cells may involve the regulation of the PI3K/AKT/mTOR signaling pathway and had no toxic effects for normal fibroblasts. Thus, AAP-H is a promising anti-prostate cancer agent and can be studied as a functional food for the prevention or treatment of this cancer type (Li X. et al., 2018). Three peptides isolated from the coral Sarcophyton glaucum also presented anticancer activity. The peptides AERQ, RDTQ and AGAPGG exhibited cytotoxicity against the human cervical cancer (HeLa) cell line, with EC50 values of 4.9, 5.6, and 8.6 mmol/L, respectively. These compounds manifested low cytotoxicity on non-cancerous Hek293 cells (Supplementary Table 1). Besides that, AERQ, RDTQ and AGAPGG have cytotoxic activity that is 5.8, 5.1, and 3.3 times stronger than that of the anticancer drug 5-fluorouracil, respectively (Quah et al., 2019).
Mollusca Peptides
The phylum Mollusca is composed of aquatic invertebrate organisms, such as mussel, octopus, squid, and oyster, and terrestrial organisms, such as snail (Otero-Gonzáiez et al., 2010; Zhong et al., 2013). Among the known defense mechanisms, these organisms use only physical barriers and the innate immune system as a way of protecting against the most diverse pathogens, since they do not have adaptive immunity mechanisms (Zhong et al., 2013). Organisms in this phylum produce peptides that have different functions, and peptides isolated from different tissues of these organisms are found in the literature (Supplementary Table 1).
Conopeptides or conotoxins belong to the family of peptides isolated from the poison produced by snails from the Conus genus, such as the peptide Lo1A (Figure 5) isolated from Conus longurionis. These are characterized by having a small size and having a composition rich in cysteine, in addition to presenting a unique characteristic because they are highly selective and potent ligands to diverse receptors and ion channels (Lebbe et al., 2014, 2016). The first peptide approved for commercial use was a conopeptide/conotoxin called ω-MVIIA (Bowersox and Luther, 1998; Figure 5), derived from the venom of a fish-hunting cone snail Conus magus, with potent analgesic properties (Rigo et al., 2013). Since then, many peptides with different biological activities have reached different stages of development in the pipeline for the market (Jimeno et al., 2004; Blunt et al., 2018).
Several AMPs have already been isolated and characterized as produced by organisms of this phylum, most abundantly in mussels (Figure 1). The mussel Mytilus coruscus has been the subject of studies for several years, which led to the isolation of AMPs myticusin-1, mytichitin-CB and myticusin-beta. Myticusin-1 is a cysteine-rich peptide with an unusual pattern of the AA. It presented antimicrobial action against Gram-positive bacteria, such as S. aureus (MIC < 5 mM), and Gram-negative, such as E. coli (MIC > 10 mM) (Liao et al., 2013). Mytichitin-CB was isolated from the hemolymph of this mollusk; it is formed by 55 AA residues, with three disulfide bonds. Like myticusin-1, this other peptide has greater antimicrobial activity against Gram-positive bacteria, in addition to antifungal activity, both of which were demonstrated in tests against Bacillus subtilis, S. aureus, Sarcina luteus, and Bacillus megaterium (MIC < 5 mM) and against the fungi C. albicans and Monilia albicans (MIC 6.2–25 mM) (Qin et al., 2014; Oh et al., 2018). Myticusin-beta has an N-terminal sequence similar to myticusin-1, which begins composed of a signal peptide (24 AA residues) and a mature peptide (104 AA residues). This peptide also showed antimicrobial activity against Gram-positive and Gram-negative bacteria, while its recombinant peptides have shown anti-parasitic activity against scuticociliates (Oh et al., 2020). The search for new AMPs in the genome and transcriptome and peptide-derived synthesis is becoming increasingly popular, given the difficulty in extracting peptides naturally produced by organisms. Therefore, genome analysis paves the way for the production of new multifunctional peptides. Octominin, for example, was detected from the analysis of the Octopus minor transcriptome, being modified and synthesized based on the defense protein 3 cDNA sequence from the mollusk. Experimental data demonstrated that this peptide inhibited the growth of C. albicans with minimum fungicidal concentration of 200 μg/mL and (MIC = 50 μg/mL) and a rapid decrease in the number of viable cells (35 and 5% cell viability at 24 h, in treatment with 50 and 100 μg/mL of octominin, respectively). The mechanism of action of octominin is related to the formation of pores in the cell wall and membrane and the accumulation of ROS (Nikapitiya et al., 2020). It also presents anti-inflammatory activity in LPS-activated macrophages by inhibition of pro-inflammatory factors, like cytokines, NO and chemokines (Supplementary Table 1; Sanjeewa et al., 2020).
Some studies on AMP isolation have sought to demonstrate the activity of these compounds against groups of bacteria that affect the health of other marine organisms important to the fishing economy, such as crustaceans, fish, and mollusks (Austin and Zhang, 2006; Liu et al., 2013). This is the case of the peptides RpdefB and VpMacin. RpdefB, identified as being of the defensin type, has been isolated from Ruditapes philippinarum and expressed in the five different tissues of the mollusk. This peptide showed high antibacterial activity against Gram-negative and Gram-positive bacteria, with MIC ranging between 0.5 μM (Vibrio splendidus) and 8.0 μM (S. aureus). RpdefB provided an increase in membrane permeability in an experiment carried out with E. coli, resulting in the death of this microorganism (Yang et al., 2018). VpMacin, a peptide characterized as belonging to the macin family, was isolated from different tissues of Venerupis philippinarum. The recombinant peptide, called rVpMacin, was tested against different bacteria, showing activity against human pathogenic strains and greater activity against strains that are pathogens of marine organisms, such as Vibrio alginolyticus, Vibrio harveyi, and V. splendidus (MIC between 0.5 and 4.0 μM) (Yang et al., 2019).
Sepia ink from mollusks has been known and used for thousands of years in Chinese medicine. It has many properties, such as anti-hypertensive, antibacterial, anti-retrovirus, antioxidant, and antitumor activities. However, the first study to demonstrate the induction of apoptosis in cancer cells of a sepia ink peptide was only published in 2017. The polypeptide SHP was isolated via the hydrolysis of sepia ink from Sepia esculenta and was subjected to a cell proliferation assay on prostate cancer cell line PC-3 (Pca PC-3). The results indicated that SHP inhibited the proliferation of Pca PC-3 in a time- and concentration-dependent (5, 10, 15, 20 mg/ml treated with SHP for 24, 48, and 72 h) and induced apoptosis by upregulation of the Bcl-2/Bax ratio (Huang et al., 2017).
Interestingly, some peptides initially isolated from marine invertebrates can be produced by other organisms (Faircloth and Cuevas, 2006). This is the case of the peptides kahalalides. These peptides were isolated from mollusks Spisula polynyma and Elysia rufescens and they have antitumor action (Gracia et al., 2006; Rao et al., 2008). However, a study by Zan et al. (2019) showed that a bacterial symbiont Candidatus Endobryopsis kahalalidefaciens produced kahalalide (Figure 5) for the chemical defense of the host (alga Bryopsis sp.). Subsequently, this peptide is removed by the mollusk from its food and used for defense.
The peptides mytilin B (Figure 5) and myticin C were isolated from the mussel Mytilus galloprovincialis, with myticin C being the first antiviral peptide against mollusk pathogens. In a study carried out by Novoa et al. (2016), oyster resistance to herpesvirus Ostreide 1 was demonstrated, when mussel hemolymph was introduced during the incubation period of oyster hemocytes, with myticin C the main peptide found in both. Modified myticin C peptides also showed antiviral activity with human herpes simplex viruses 1 and 2, raising the proposal to use it for veterinary and medicinal use (Balseiro et al., 2011).
Photo-protective peptides were also found in mollusk tissues. Peptides from the oyster Crassostrea hongkongensis (OPs) were obtained from the protein hydrolysates and the photo-protective effects investigated by topical application on mouse skin. OPs contained 24 peptide chains (formatted by 4-17 AA) that showed antioxidant activity, inhibiting metalloproteinase secretion and malondialdehyde production. Moreover, topical OP application demonstrated anti-inflammatory activity caused by decreased production of inflammatory cytokines. These results indicate that OPs are photo-protective peptides and can be used as cosmetic or therapeutic products against photoaging (Peng et al., 2020).
Annelida Peptides
Organisms of the phylum Annelida inhabit terrestrial, freshwater, and marine environments and are classified in the classes Polychaeta and Clitellata. The organisms of this phylum have a segmented body with bilateral symmetry, with worms and leeches as the best-known organisms. The Polychaeta class is the largest group, comprising about 14,000 species and most of the diversity within that phylum. They are invertebrates found in almost all marine environments (de Paiva, 2006; Coutinho et al., 2017). Despite the large number of species in this phylum, studies are still few and restricted to a few species. Much of the research is on earthworms (oligochaetes), due to their biological role in the soil, and some species of polychaetes (Schenk and Hoeger, 2020).
Marine polychaetes excel in isolating AMPs (Figure 1). As in other phyla, structural restrictions have also been observed in antibacterial and antifungal peptides from the phylum Annelida, which present structures organized in dimeric β-hairpins, double-stranded antiparallel beta-sheet and α-helix stabilized by disulfide bonds (Kaas et al., 2010; Panteleev et al., 2018), such as the peptide alvinellacin (Figure 5; Lie Tasiemski et al., 2014). The AMPs arenicin-1, 2, and 3 were extracted and purified from the marine polychaete Arenicola marina, both formed by 21 AA residues. The peptides showed antimicrobial activity against Gram-positive and Gram-negative bacteria (S. aureus, Enterococcus faecium, S. epidermidis, E. coli, Pseudomonas aeruginosa, and Listeria monocytogenes), as well as against the fungus C. albicans (Park and Lee, 2009), inhibiting growth in concentrations greater than 25 μg/ml (Ovchinnikova et al., 2004; Lee et al., 2007; Wang et al., 2014b). A study by Andrä et al. (2008) showed that arenicin-1 has potent antimicrobial activity against mutant strains of P. mirabilis resistant to polymyxin B (Supplementary Table 1). The mechanism of action of arenicin 1 and 2 is related to its structure and destabilization of the bacterial plasma membrane (Lee et al., 2007; Cho and Lee, 2011). These factors were studied by Shenkarev et al. (2011). The peptide arenicin-2 has a dimeric β-hairpin hook structure and produces pores in the cell membrane. A recent study proposed several variants of arenicin-1 to assess the role of its structure and symmetry. The results showed that the arrangement of the arenicin-1 β-hairpin (Figure 5) caused an increase in antimicrobial activity against Gram-negative bacteria but a decrease against Gram-negative (Orlov et al., 2019).
Partially cyclic peptides from the phylum Annelida also present anticancer properties, as is the case of nicomicin-1 (Figure 5). This peptide presents a structural arrangement that resembles the Rana-box motif observed in α-helical peptides found in frog skin. It consists of an N-terminus amphipathic α-helix, along with a six-residue loop in a cyclic arrangement and stabilized by a disulfide bond, which has been correlated with this peptide’s antibacterial and anticancer activities. The peptide nicomicin-1 has a unique structure when compared to AMPs isolated from other polychaetes, due to these characteristics. This AMP was isolated from Nicomache minor (Panteleev et al., 2018) and had a BRICHOS domain as a precursor, characterized by participating in post-translational protein processing (Chen et al., 2017). Nicomicin -1 showed activity against Gram-positive bacteria (M. luteus, B. subtilis, Bacillus licheniformis, S. aureus 209P, S. aureus ATCC 29213, Rhodococcus sp.), with MIC varying between 0.062 and 2 μM. The cytotoxicity of nicomycin-1 for cervical adenocarcinoma cells was dose-dependent and reached a percentage of 85% of dead cells in concentrations of 32 μM of the peptide (Panteleev et al., 2018). In the most recent study about AMPs from polychaetes, a new recombinant analog peptide was produced from Capitella teleta. Capitellacin (Figure 5) has a β-hairpin structure, a BRICHOS domain and high homology with the tachyplesins (Figure 5) AMP family. Analysis of the antimicrobial activity of capitellacin demonstrated that this peptide has broad-spectrum activity, but it was up to eightfold lower than that of tachyplesin-1 (a peptide from the tachyplesins family). Unlike other peptides with β-hairpin structure, capithelinacin was inefficient in breaking the cytoplasmic membrane and inhibiting bacterial translation, suggesting other mechanisms of action and targets (Panteleev et al., 2020).
Another much-studied polychaete is Perinereis aibuhitensis. The peptide perinerin was extracted from this organism, a hydrophobic peptide composed of 51 AA residues. This peptide showed antimicrobial activity against the fungus Paecilomyces heliothis (MIC 25 μg/ml), high activity against P. aeruginosa, B. megaterium, and Aerococcus viridans (MIC < 15 μg/ml), moderate activity against E. coli K-12, S. aureus and M. luteus, and lower activity against Proteus vulgaris (MIC > 50 μg/ml) (Pan et al., 2004). Peptides present in protein extracts of Perinereis aibuhitensis also showed antithrombotic (Li et al., 2017) and antihypertensive activities (Chen and Wang, 2017). Jia et al. (2017) isolated the peptide P. aibuhitensis (PAP), a decapeptide with anticancer action that was tested by Jiang et al. (2019) against human lung cancer H1299 cells cultured in vitro, where this peptide showed potential inhibition of malignant lung cancer cell growth and tumor cell apoptosis. This study suggests that PAP could be used for the prevention or treatment of human non-small-cell lung cancer in the future.
A study by Park et al. (2020) isolated and evaluated the peptide NCW from the worm Marphysa sanguinea. NCW was evaluated for anti-inflammatory and antioxidant effects and its cytotoxicity in Raw264.7 cells induced by LPS. The results showed that NCW has a high elimination capacity of 2,2-diphenyl-1-picrilhidrazil (DPPH) when compared to vitamin C (IC50 of 17.5 μg/ml and 2.7 μg/ml, respectively). It also had an anti-inflammatory effect by decreasing NO production stimulated by LPS and inhibiting the secretion of pro-inflammatory cytokines IL-1β and TNF-α, in addition to NCW not showing cytotoxicity in Raw264.7 cells induced by LPS (Supplementary Table 1).
Arthropoda Peptides
When stimulating the mud crab with LPS, Anju et al. (2019) increased the mRNA expression of the Scylla serrata hemocytes and increased the production of the peptide Ss-arasin. Ss-arasin is formed by 65 AAs with a predicted molecular weight of 7 kDa, and it was expressed in a prokaryotic heterologous system. Recombinant rSs-arasin showed significant activity against S. aureus, P. aeruginosa, and E. coli. This recombinant peptide also inhibited the growth of human cervical carcinoma (HeLa) and colon carcinoma (HT-29) cells. These discoveries are of the utmost importance for structure-activity relationships, in order to optimize biological functions for future drug development.
Membrane peptides can be classified as cell-penetrating peptides and AMPs, recognized for playing interchangeable roles. Tachipesin is a peptide isolated from the hemolymph of the horseshoe crab (Tachypleus tridentatus), known for having antimicrobial properties, and investigated for its ability to penetrate cells. Derived from a marine organism, it has internalized cell penetration, being found in plant and mammalian systems. It was established as an efficient non-viral macromolecule nanocarrier (Jain et al., 2015).
Anti-lipopolysaccharides (ALFs) are factors known as important defense molecules for the host of crustaceans. Containing a LPS-binding domain (LBD), some ALFs have strong antimicrobial activity. Zhou et al. (2019) described a group G ALF of the Giant Tiger Shrimp (Penaeus monodon) (ALFPm11). It is a highly amphipathic peptide, with five positively charged residues separated by aromatic residues, normally expressed abundantly in the hepatopancreas of P. monodon. Chemically synthesized ALFPm11-LBD exhibited high inhibitory activity against various bacteria, for example Exiguobacterium sp., Bacillus sp., and Acinetobacter sp., thereby efficiently inducing the agglutination of Gram-positive and Gram-negative bacteria and causing significant permeabilization of the bacterial membrane. Thus, an effective new ALF has been described in P. monodon, demonstrating its antimicrobial mechanism, suggesting potential applications for the future.
Histone H2A participates in the host’s immune defense through the generation of special AMPs in vertebrates and invertebrates. Although thousands of AMPs have been reported in various species, AMPs in crabs are better known (Figure 1). Chen et al. (2015) determined the existence of the peptide sphistin with 38 AA residues and antimicrobial activity based on the analysis of the sequence of an H2A histone identified from Scylla paramamosain. Sphistin exerted high antimicrobial activity against Gram-positive, Gram-negative bacteria and yeasts, including Aeromonas hydrophila, Pseudomonas fluorescens, and Pseudomonas stutzeri, important aquatic pathogens. In the bacterial cells treated with sphistin, leakage of the cell content and rupture of the cell surface were observed. It has been proven that the increased permeability of the E. coli cytoplasmic membrane was caused by sphistin. Synthetic sphistin did not exhibit cytotoxicity to cultured crab hemolymph and mammalian cells, even at a high concentration. This is the first report of a histone-derived sphistin identified from S. paramamosain, which can be promising for application in aquaculture and veterinary medicine (Chen et al., 2015).
An AMP similar to the crustin of giant tiger prawn hemocytes, P. monodon, was partially characterized at the molecular level, and the phylogenetic analysis was performed by Antony et al. (2011). The 299 bp partial coding sequence and 91 deduced AA residues had conserved cysteine residues characteristic of shrimp crustins. The phylogenetic tree and the sequence comparison clearly confirmed the divergence of this crustin-like AMP from other shrimp crustins. The differential expression of a crustin-like AMP in P. monodon in response to the administration of several immunostimulants, namely, two marine yeasts (Candida haemulonii S27 and Candida sake S165) and two β-glucan isolates (extracted from C. haemulonii S27 and C. sake S165), were noted during the study. Responses to the application of two gram-positive probiotic bacteria and Micrococcus were also observed. The immunological profile was recorded pre- and post-challenge with the white spot syndrome virus (WSSV) by semiquantitative RT-PCR. Expressions of seven WSSV genes were also observed to study the intensity of viral infection in experimental organism. A crustin-like AMP has been found to be expressed constitutively in the organism and significant negative regulation can be observed after the WSSV challenge. Notable negative regulation of the gene was observed in organisms fed a pre-challenge immunostimulant followed by a post-challenge WSSV of significant positive regulation. The tissue expression of a crustin-like AMP in the administration of C. haemulonii and Bacillus showed maximum transcriptions in the gills and in the intestine. Marine yeast, C. haemulonii and probiotic bacteria, Bacillus, increased the production of an AMP similar to crustin and conferred significant protection on P. monodon against WSSV infection (Antony et al., 2011).
Antimicrobial peptides studied from hematocyte strata of the red king crab, Paralithodes camtschaticus, are described by Moe et al. (2018). The peptides were isolated by bioassay purification, showing that they are cationic peptides rich in cysteine (Cys), called paralithocins 1–3. The peptides (38–51 AAs in length) share a single Cys motif consisting of eight Cys, forming four disulfide bridges with a Cys1–Cys8 (Cys relative position) connectivity, Cys2–Cys6, Cys3–Cys5, and Cys4–Cys7, a disulfide arrangement that has not been previously reported among AMPs. These peptides are responsible for inhibiting the growth of several marine bacterial strains with minimal inhibitory concentrations in the range of 12.5–100 μM. These findings confirm the hypothesis that marine organisms are a valuable source for the discovery of bioactive peptides with new structural motifs.
With the recent spread of obesity in developing countries, more attention is focused on the development of appetite-moderating drugs with low side effects. Anti-obesity peptides generally control satiety hormonally. Two important synthetic anti-obesity peptides already characterized are cholecystokinin and gastrin, appetite suppressants with multiple cellular targets (Cheung et al., 2015). Some already characterized marine anti-obesity peptides stimulate the synthesis of cholecystokinin – such as peptides from shrimp head protein hydrolysates, which can stimulate cholecystokinin release in STC-1 cells – or are structurally and mechanistically similar to gastrin/cholecystokin (Sable et al., 2017). The shrimp anti-lipopolysaccharide factor (SALF) was described by Wang L. et al. (2017), an AMP from black tiger prawns that increases the anticancer activity of cisplatin in vitro and inhibits the growth of HeLa cells in nude mice. These peptides also exhibit anti-cancer activity defined in cancer cell lines of cancer and human liver, even when removed from shrimp residue (Wang L. et al., 2017).
Echinodermata Peptides
Echinoderms are marine invertebrates, phylogenetically related to chordates and hemichordates. They have a unique morphology that includes pentameric radial symmetry in their adult forms and bilateral symmetry in larvae and a vascular water system. Echinoderms live in diverse marine environments, rich in fungi, bacteria, viruses and many pathogenic organisms of echinoderms. Thus, these organisms evolved in a complex way, and their immune system is totally innate but highly effective and sophisticated. The complexity of the echinoderm’s immune system can be understood by looking at the large number of AMPs isolated from these organisms (Figure 1).
The first AMP reported from the sea star was the cysteine-rich peptide named PpCrAMP from Patiria pectinifera. It is formed by 38 AA residues with two disulfide bonds and C-terminal amination with a glycine. Synthetic PpCrAMP showed antimicrobial activity against Gram-negative bacteria, S. enterica and Shigella flexneri (MEC of 4.5–31.4 μg/ml), and Gram-positive bacteria, B. subtilis, S. aureus and M. luteus (MEC of 15.6 to >250 μg/ml). The coelomic epithelium extract was also tested and activity against E. coli was detected (Kim et al., 2018).
Strongylocins were the first peptides from echinoderms whose genes were completely sequenced. SdStrongylocin 1 is formed by 48 AA residues while SdStrongylocin 2 has 51 AA residues; both have strong activity against Gram-positive and Gran-negative bacteria (MIC of 1.3–2.5 μM). These peptides were extracted from Strongylocentrotus droebachiensis and were characterized as rich cysteine peptides (Li et al., 2015). From the coelomocytes from S. droebachiensis, the peptides centrocin 1 and 2 were also extracted. These molecules are heterodimeric peptides formed by a chain with 30 AAs (heavy chain) and a chain with 12 AAs (light chain). The heavy chain is linked to the activity against Gram-positive and Gram-negative bacteria (MIC of 1.3–2.5 μM) while the light chain did not show activity or influence in the antimicrobial activity of the heavy chain (Li et al., 2010, 2015).
Three AMPs were isolated and characterized from the coelomic fluid of the sea urchin Echinus esculentus. Two of these peptides are heterodimeric, EeCentrocin 1 and 2, and the other is a cysteine-rich peptide, EeStrongylocin 2. EeCentrocin 1 demonstrated potent antimicrobial activity against Corynebacterium glutamicum and S. aureus (MIC = 0.78 μM against both) and E. coli and P. aeruginosa (MIC = 0.1 and 0.78 μM respectively) (Supplementary Table 1). The other two peptides could not be tested in isolation due to the difficulty of purification, but the mix of these peptides also demonstrated antimicrobial activity (Solstad et al., 2016). Another study made modifications in the structure of EeCentrocin 1 to produce a shorter drug peptide using its heavy chain. The new peptide named P6 was formed of 12 original AA residues of the N-terminal from EeCentrocin 1, two modifications of the AA and one C-terminal amidation. P6 showed greater antibacterial and antifungal activity than EeCentrocin 1, in addition to presenting lower hemolytic activity and no cytotoxic activity (Solstad et al., 2020).
Several peptides with inhibitory natural angiotensin-converting enzyme (ACE) activity have been isolated from diverse marine organisms. These peptides can be used as antihypertensive drugs. Peptides NVA and NLG with ACE inhibitory action were extracted from Acaudina molpadioidea protein hydrolysates, present in the body wall of this sea cucumber (Zhu et al., 2011; Shen et al., 2014). They were modified with the addition of exogenous proline, increasing the inactivation of ACE, with IC50 values of 8.2 ± 0.2 μM for PNVA and 13.2 ± 0.4 μM for PNLG (Li J. et al., 2018). Compounds with this mechanism of action have been studied as a possible treatment for diseases that present hypertension, as is the case of COVID19. SARS-CoV-2 enters cells via interactions between the Angiotensin-converting enzyme 2 (ACE2) and the Spike viral protein. Marine peptides with antiviral and anti-ACE2 activity can be an alternative tool in solving this problem (Festa et al., 2020). From this echinoderm, four peptides (Fa.2-A, Fa.2-B, Fa.2-C, and Fa.2-D) were also isolated from the hydrolysis of collagen. The peptide Fa.2-C showed the highest scavenging activity for 2,2-diphenyl-1-picrylhydrazyl (DPPH) at a concentration of 0.2 mg/mL (Supplementary Table 1). This indicates a reduction in oxidative damage by converting the DPPH radical to less harmful substances. Thus, it can be considered a promising antioxidant collagen peptide and used in the food and drug industry (Jin et al., 2019).
Studies indicate that peptides with chelating action can be used in the production of human food supplements. This is the case of the zinc chelating peptide (ZCP), isolated from the edible sea cucumber Stichopus japonicas. ZCP is composed of eight AA residues (WLTPTYPE) and has a molecular weight of 1005.5 Da, with a zinc chelating capacity equal to 33.3%. Researchers point out that future work can better analyze the mechanism of action of ZCP and the effects regarding the intensification of intestinal zinc absorption (Liu et al., 2019a, b). This sea cucumber was also studied for its capacity to release antidiabetic peptides. The study involved simulating digestion processes by in vitro model, where 58 peptides with lower molecular weight (<3 KDa) were found and characterized. These peptides showed the highest inhibitory activity of the enzyme dipeptidyl peptidase IV (DPP-IV) when compared to anagliptin, a DPP-IV inhibitor. This indicates that S. japonicas can be used to release antidiabetic peptides in gastrointestinal digestion as a functional food (Gong et al., 2020).
The small-molecule oligopeptides (SOP) isolated from the sea cucumber Codonopsis pilosula were studied for their potential immuno-modulatory effects. The assays performed with mice showed that these molecules appeared to increase innate and adaptive immune responses when SOPs were orally administered. This response occurs by NK cell action, macrophage phagocytosis, and humoral and cell-mediated immunity (Supplementary Table 1). Thus, these oligopeptides could be used for cancer patients and immunocompromised persons, to increase immune functions (He et al., 2016). In another study, the SOP isolated from sea cucumber showed healing activity in wounds in diabetic mice. This action can be explained by the capacity for increased angiogenesis, synthesis, and deposition of collagen, which enhanced the antioxidant levels and reduced the inflammatory response (Li D. et al., 2018).
Chordata Peptides
Compared to other phyla, it is worth noting that, when studying bioactive peptides from Chordata phylum organisms (vertebrates), a very different picture is observed. Marine peptides isolated from these organisms are usually composed only of natural AAs in a linear structure that transits from random coil to α-helix and beta-sheet when in contact with bacterial and fungi surfaces. Among the peptides from Chordata are piscidins and marine cathelicidins (Broekman et al., 2011; Mihailescu et al., 2019).
Piscidin (Figure 5) pools are important components of the innate (non-specific) immune system in fish, and they have potent and broad-spectrum antimicrobial and antiparasitic activities (Figure 1). A new antimicrobial cationic peptide from Pseudosciaena crocea is described by Niu et al. (2013), which exhibits a genomic organization (three exons and two introns) and propeptide (signal peptide, mature peptide, and prodomain), conserved signal peptide (22 AAs) and ratio for consensus I-X5-H-X4-IH. In fish piscidins, Pc-pis demonstrated a relatively low general conservation with other known piscidins, which was obviously incorporated in the AA composition of the peptide. Pc-pis is surprisingly rich in glycine residues (27.3%), which interrupted the amphipathic structure of the peptide. Relative quantitative real-time PCR revealed that Pc-pis is typically expressed by gills. The analysis of the sequence, structural characteristics and tissue distribution suggested that Pc-pis was genetically related to the piscidine family and may be a new AMP similar to piscidine. Quantitative PCR analysis revealed that the expression of Pc-pis in the spleen, head-kidney, liver, intestine, skin and gills can be regulated during infection by Cryptocaryon irritans and after the decrease of C. irritans, implying a role for Pc-pis in the immune defense system against C. irritans and secondary bacterial infections. Pc-pis exhibited broad-spectrum activity against bacteria, fungi and C. irritans in parasitic stages. These results provided evidence of the antiparasitic activity of piscidins against ectoparasites of marine fish, namely C. irritans trophons, and further indicated that these data provided new insights into the innate immunity of P. crocea against external protozoan parasites and microbial infections and facilitate the assessment of Pc-pis as a therapeutic agent against the invasion of pathogens (Niu et al., 2013).
Cathelicidins are a group of AMPs that have antibacterial, antifungal, antiviral, immunostimulatory, and immunomodulatory properties. The cathelicidin family is characterized by a highly conserved N-terminal cathelin domain and a variable C-terminal antimicrobial domain that can be extracted by the action of proteinases. Cathelicidin works together with defensins and high concentrations of cathelicidin are found at sites of inflammation, where they appear to destroy microorganisms and act as a bridge between innate and late immunities. The observation that AMPs are also produced in the epithelium supports the hypothesis that they are part of a type of primary defense system. Cathelicidins are among the best characterized AMPs and play an important role in mammalian innate immunity. A new mature cathelicidin peptide (codCath) was isolated by Broekman et al. (2011) from Atlantic cod and the function of codCath was characterized. The peptide showed sensitivity to salt with an annulment of activity at physiological salt concentrations. In a medium of low ionic strength, it showed activity against marine and non-marine Gram-negative bacteria, weak activity against a Gram-positive bacterium and antifungal activity. CodCath’s actions are rapid death accompanied by pronounced cell lysis. Extracellular products (E) from three marine bacteria caused the peptide to break down into smaller fragments, and the cleaved peptide lost its antibacterial activity. Peptide proteolysis, on the other hand, was abolished by previous heat treatment of ECPs, suggesting protease involvement. No peptide cytotoxicity was observed in fish cells, and the selectivity of activity was confirmed with bacterial and mammalian membrane mimetics. In this way, the potent broad-spectrum activity of codCath suggests a role of the peptide in the immune defense of cod (Broekman et al., 2011).
Another AMP (Salmo salar) – which acts as an activation signal for the marine parasite copepod Lepeophtheirus salmonis. L. salmonis – was exposed to 0, 7, 70, and 700 ppb of Cath-2 (cathelicidin-2) and the neural activity, swimming behavior and gene expression profiles of organisms in response to the peptide were evaluated. The neurophysiological, behavioral and transcriptomic results were consistent: L. salmonis detects Cath-2 as a water-soluble peptide released from the skin of the salmon, triggering the chemosensory sensorineural activity associated with altered swimming behavior of copepodids exposed to the peptide, and genes related to chemosensorials increased in copepodids exposed to the peptide. L. salmonis are activated by Cath-2, indicating a strong link between this peptide and the chemosensory system of the salmon louse (Núñez-Acuña et al., 2018).
Antimicrobial peptides play a defense role against microbial invasion. Liver Expressed Antimicrobial Peptides-2 (LEAP-2) is an AMP that belongs to a group of peptides rich in cysteine. Hwang et al. (2019) identified and characterized the molecular properties of LEAP-2 in sea bream (Diplodus sargus). The expression levels of LEAP-2 (RbLEAP-2) were evaluated in 12 different healthy fish tissues and the expression pattern of RbLEAP-2 after infections by Edwardsiella piscicida, Streptococcus iniae and iridovirus from (RSIV) was observed. These data show that RbLEAP-2 plays an important role in innate immunity when infected with a pathogen.
Marine peptides that are known to induce cell death by DNA fragmentation, nucleic shrinkage and swelling of the cell membrane have been described by Zheng et al. (2011). Didemnins belong to a family of depsipeptides that have antitumor, antiviral and immunosuppressive activities, and they were first isolated from Caribbean marine organism tunicates by Rinehart et al. (1981). Didemnin B, which is a branched N-methylated cyclic peptide, was isolated from the genus Trididemnum, and it has the ability to induce the death of a variety of cells by DNA fragmentation within the cytosol. Further studies showed that didemnins are produced by symbiotic bacteria in tunicates. Didemnins are produced by the marine α-proteobacteria Tistrella bauzanensis and Tistrella mobilis (Tsukimoto et al., 2011). These bacteria synthesize the non-ribosomal peptide synthetase-polyketide synthase enzyme complex responsible for the production of didemnin B precursors (Xu et al., 2012; Zan et al., 2019; Supplementary Table 1).
A derivative of the shark cartilage stratum is described as neovastat (AE-941), a liquid extract with defined standardization comprising a <500 kDa fraction of shark cartilage; this product of the mud shark Squalus acanthias was seen to be a direct inhibitor of the growth of tumor cells and angiogenesis. Mice that were treated with neovastat showed a significant reduction in the inflammatory cell count in BAL liquid, and those mice that were treated with neovastat showed a significant reduction in the expression of VEGF and HIF2 alpha in lung tissue (Zheng et al., 2011).
Styelin D, eusynstyelamide (Figure 5), botrilamides A – D (1–4), lissoclinamides and molamidas were isolated from ascidians, presenting a potential for cytotoxicity, but studies still have not provided precise knowledge about this mechanism. Zheng et al. (2011) described the peptide styelin D, as quite cytotoxic and hemolytic for eukaryotic cells. Isolated from blood cells of ascidia Styela clava, this peptide has antimicrobial activity, with 32 residues C-terminal amidate (Supplementary Table 1). Lissoclinamides are cyclic peptides that have shown significant antineoplastic activity, as well as other pharmacological properties against human fibroblasts and bladder carcinoma cell lines and normal lymphocytes, after being isolated from the aplousobranch Lysoclinum patella. Among them, the most potent is lissoclinamide 7 (Figure 5), containing two thiazoline rings, which rivals didemnin B for in vitro cytotoxicity.
Industrial Use of Peptides Isolated From Marine Organisms
Some of the marine peptides already described in the literature are of industrial interest. The nutraceutical, pharmaceutical and cosmetic industries are the sectors that most invest in the discovery and characterization of marine peptides (Cheung et al., 2015). In general, industrial marine peptides have a broad spectrum of bioactivities, such as antimicrobial, antiviral, antioxidative, antihypertensive, cardioprotective, antiatherosclerotic, anticoagulant, immunomodulatory, antitumor, analgesic, anti-diabetic, anxiolytic, neuroprotective and appetite suppressing (Sable et al., 2017). In addition to naturally occurring peptides, some marine peptides are modified/designed by the industry from natural backbones, to increase their potential in the treatment or prevention of diseases. These peptides are called “derivatives” and represent the vanguard of molecules with high added commercial value.
The main classes of bioactive industrial peptides isolated from marine fauna and flora are pharmaceutical peptides and nutraceutical peptides. While pharmaceutical bioactive peptides are those that only have activity in promoting health by preventing or fighting diseases, nutraceutical peptides are products naturally found in foods or food derivatives after enzymatic modification in the gastrointestinal tract, with double commercial value: they have both nutritional and pharmaceutical value (Zhang et al., 2005; Simmons et al., 2008; Ngo et al., 2012; Kang et al., 2015; Sable et al., 2017; Wang et al., 2018).
There are only two pharmaceutical peptides of marine origin reaching the status of marketed products, both chemically synthetic. The first is the natural marine peptide ziconotide (Prialt®), isolated from the snail Conus magus and with analgesic effect, approved by the FDA in 2004 (Bowersox and Luther, 1998; Wang X. et al., 2017). The second is the derivative antitumor peptide brentuximab vedotin (Adcetris®) from dolastatin 10, a linear pentapeptide which was originally isolated from extracts from the marine mollusk, Dolabella auricularia. Dolastatin 10 is also synthesized by the marine cyanobacterium Symploca sp. (Pettit et al., 1987; Luesch et al., 2001). Brentuximab vedotin was approved for the treatment of patients with systemic anaplastic large cell lymphoma and patients with Hodgkin’s lymphoma by the FDA in 2011 (Younes et al., 2012; Giordano et al., 2018). Despite the restricted number of pharmaceutical marine peptides approved for the market, the number of nutraceutical peptides marketed in the United States and Europe is considerably higher (Sable et al., 2017). The fact that these molecules are part of food complexes and not active drug reagents is the main factor that explains the greater number of nutraceutical peptides on the market (Sable et al., 2017).
Limitations inherent to the isolation process, prospecting, structural modification and low rates of biosynthesis in natural conditions of marine peptides have restricted the arrival of a few of these molecules on the market. However, a large number of them have reached different phases of the clinical and preclinical pipeline (Table 2).
TABLE 2
| Molecule | Main characteristic/source | Nature | Applications | Status |
| Plitidepsin | Cyclic depsipeptide/Aplidium albicans | Natural product | Cancer treatment | Phase I/II clinical trials |
| Glembatumumab vedotin | Dolastatin 10/Symploca sp. | Derivative | Cancer treatment | Phase I/II clinical trials |
| Elisidepsin | Kahalalide F/Elysia rufescens | Derivative | Cancer treatment | Phase I clinical trials |
| Kahalalide F | Cyclic ridecapeptide/Elysia rufescens | Natural product | Cancer treatment | Phase I clinical trials |
| HTI-286 | Hemiasterlin/Hemiasterella minor | Derivative | Cancer treatment | Preclinical trials |
| Brentuximab vedotin | Dolastatin 10/Symploca sp. | Derivative | Cancer treatment | Marketed/FDA approved |
| Ziconotide | Ω-conotoxin toxin/C. Magus | Natural product | Analgesic | Marketed/FDA approved |
| Dermochlorella® | Oligopeptide extract/Chlorella vulgaris | Natural product | Skin toner | Sold as component of skin care formula |
| Fragments of fish gelatin | Hydrolysate of fish collagen and gelatin | Natural product | Nutrient supplement | Marketed as nutraceutical |
| Katsuobushi oligopeptide | Pentapeptide from hydrolysate of dried bonito | Natural product | Antihypertensive | Marketed as nutraceutical |
| Gabolysat PC60® /Stabilium® /Protizen® /Procalm® | Hydrolysate of fish protein | Natural product | Anxiolytic | Marketed as nutraceutical |
| Nutripeptin® /Hydro MN Peptide | Hydrolysate of fish protein | Natural product | Postprandial blood glucose control | Marketed as nutraceutical |
| Seacure® | Hydrolysate of fish protein | Natural product | Intestinal health mediator | Marketed as nutraceutical |
The current status of marine peptides on the market and under clinical trials.
Adapted from Cheung et al. (2015).
Marine peptides that have already arrived on the market and those that are undergoing pre-clinical and clinical tests are well-finished examples of molecules with added value. However, proportionally, there are still a significant number of marine peptides that are in less advanced intermediate stages in the pipeline. These peptides present great biotechnological potential (Supplementary Table 2).
Conclusion and Prospects
Marine ecosystems came to be considered a rich source of active molecules after the discovery of bioactive compounds isolated from porifera phylum organisms. These organisms have become the focus of several studies and are responsible for the largest number of molecules with biotechnological interesting previously described, with the most diverse functions, such as antimicrobial, anti-inflammatory, and antitumor activities. The defense molecules development by sponges may be associated with adaptive pressure and their sessile life form. This last factor makes these organisms a promising source for prospecting bioactive molecules due to their easy access and the wide structural and function diversity of the natural products produced by them and their symbionts.
With the improvement of extraction and purification protocols of marine metabolites, several molecules of high commercial value have already been isolated and characterized in terms of their structures and bioactivities. Some of these molecules have the potential to be used alone or in conjunction with disease prevention therapies.
With two peptides for pharmaceutical use, at least seven peptides marketed as nutraceuticals and many others at advanced stages in the pre-commercial development pipeline, the arrival of the first bioactive peptides on the market attests to the immense marine biodiversity importance preservation as a biotechnological potential molecular rich source.
Furthermore the advances of omics sciences including metagenomics, transcriptomics and proteomics have provided an enhancement in the active molecules discovery indicating the future of bioprospecting compounds with biotechnological interest, especially with regard to pharmaceutical compounds identification. Another possibility consist in the redesigning compounds as bioinspired products from natural marine backbones. Synthetic derivatives of marine peptides tend to have more intense and selective bioactivities than their natural counterparts, making them value added products for biotechnological exploration.
Nevertheless, some pitfalls and challenges are observed, such as limitations in obtaining raw material in high amounts. This fact may be circumvented by marine organisms cultivation in the open sea (mariculture) and also in artificial cultivation conditions (aquaculture). Biosynthesis of recombinant marine peptides in model organisms and fermenters have also shown to be an important tool for large-scale production of biomass and also marine peptides. Thus, with the growing interest from the industrial and academic sectors, research efforts to improve the methods for discovering novel marine biocompounds are increasing, which is desirable to expand the offer of therapeutics, nutraceuticals and cosmetics.
Statements
Author contributions
SD and OF guided the topic selection and conceived the structure of the manuscript. MM undertook the main writing work of the review. NC, JC, RC, SA, and MC assisted in writing. All the authors contributed and approved the submitted version of this article.
Funding
MM received a doctoral scholarship from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES/PNPD protocol 88887.502159/2020-00). RC received a postdoctoral scholarship from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES/PNPD protocol 88882.315167/2019-01). We also thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), the Fundação de Apoio à Pesquisa do Distrito Federal (FAPDF) and the Fundação de Apoio ao Desenvolvimento do Ensino, Ciência e Tecnologia do Estado de Mato Grosso do Sul (FUNDECT) for financial support.
Acknowledgments
Figures in this work were developed with the support of biorender.com.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2021.667764/full#supplementary-material
References
1
Afoullouss S. Calabro K. Genta-Jouve G. Gegunde S. Alfonso A. Nesbitt R. et al (2019). Treasures from the deep: characellides as anti-inflammatory lipoglycotripeptides from the sponge Characella pachastrelloides.Org. Lett.21246–251. 10.1021/acs.orglett.8b03684
2
Aissaoui N. Abidi F. Hardouin J. Abdelkafi Z. Marrakchi N. Jouenne T. et al (2017). ACE inhibitory and antioxidant activities of novel peptides from Scorpaena notata by-product protein hydrolysate.Int. J. Pept. Res. Ther.2313–23. 10.1007/s10989-016-9536-6
3
Anand M. Alagar M. Ranjitha J. Selvaraj V. (2019). Total synthesis and anticancer activity of a cyclic heptapeptide from marine sponge using water soluble peptide coupling agent EDC.Arab. J. Chem.122782–2787.
4
Andjelic C. D. Planelles V. Barrows L. R. (2008). Characterizing the anti-HIV activity of papuamide A.Mar. Drugs6528–549.
5
Andrä J. Jakovkin I. Grötzinger J. Hecht O. Krasnosdembskaya A. D. Goldmann T. et al (2008). Structure and mode of action of the antimicrobial peptide arenicin.Biochem. J.410113–122. 10.1042/BJ20071051
6
Anju A. Smitha C. K. Preetha K. Boobal R. Rosamma P. (2019). Molecular characterization, recombinant expression and bioactivity profile of an antimicrobial peptide, Ss-arasin from the Indian mud crab, Scylla serrata.Fish Shellfish Immunol.88352–358. 10.1016/j.fsi.2019.03.007
7
Antony S. P. Singh I. S. B. Sudheer N. S. Vrinda S. Priyaja P. Philip R. (2011). Molecular characterization of a crustin-like antimicrobial peptide in the giant tiger shrimp, Penaeus monodon, and its expression profile in response to various immunostimulants and challenge with WSSV.Immunobiology216184–194. 10.1016/j.imbio.2010.05.030
8
Austin B. Zhang X.-H. (2006). Vibrio harveyi: a significant pathogen of marine vertebrates and invertebrates.Lett. Appl. Microbiol.43119–124. 10.1111/j.1472-765X.2006.01989.x
9
Balseiro P. Falcó A. Romero A. Dios S. Martínez-López A. Figueras A. et al (2011). Mytilus galloprovincialis myticin c: a chemotactic molecule with antiviral activity and immunoregulatory properties.PLoS ONE6:e23140. 10.1371/journal.pone.0023140
10
Bashir Y. Pradeep Singh S. Kumar Konwar B. (2014). Metagenomics: an application based perspective.Chinese J. Biol.2014:146030. 10.1155/2014/146030
11
Bewley C. A. Faulkner D. J. (1994). Theonegramide, an antifungal glycopeptide from the philippine lithistid sponge Theonella swinhoei.J. Org. Chem.594849–4852. 10.1021/jo00096a028
12
Blunt J. W. Carroll A. R. Copp B. R. Davis R. A. Keyzers R. A. Prinsep M. R. (2018). Marine natural products.Nat. Prod. Rep.358–53. 10.1039/c7np00052a
13
Blunt J. W. Copp B. R. Keyzers R. A. Munro M. H. G. Prinsep M. R. (2012). Marine natural products.Nat. Prod. Rep.29144–222. 10.1039/c2np00090c
14
Bo J. Cai L. Xu J. H. Wang K. J. Au D. W. T. (2011). The marine medaka Oryzias melastigma - A potential marine fish model for innate immune study.Mar. Pollut. Bull.63267–276. 10.1016/j.marpolbul.2011.05.014
15
Bowersox S. S. Luther R. (1998). Pharmacotherapeutic potential of omega-conotoxin MVIIA (SNX-111), an N-type neuronal calcium channel blocker found in the venom of Conus magus.Toxicon361651–1658. 10.1016/S0041-0101(98)00158-5
16
Braekman J. C. Daloze D. (1986). Chemical defence in sponges.Pure Appl. Chem.58357–364. 10.1351/pac198658030357
17
Brand G. D. Ramada M. H. S. Manickchand J. R. Correa R. Ribeiro D. J. S. Santos M. A. et al (2019). Intragenic antimicrobial peptides (IAPs) from human proteins with potent antimicrobial and anti-inflammatory activity.PLoS One14:e0220656. 10.1371/journal.pone.0220656
18
Broekman D. C. Zenz A. Gudmundsdottir B. K. Lohner K. Maier V. H. Gudmundsson G. H. (2011). Functional characterization of codCath, the mature cathelicidin antimicrobial peptide from Atlantic cod (Gadus morhua).Peptides322044–2051. 10.1016/j.peptides.2011.09.012
19
Carstens B. B. Rosengren K. J. Gunasekera S. Schempp S. Bohlin L. Dahlström M. et al (2015). Isolation, characterization, and synthesis of the barrettides: disulfide-containing peptides from the marine sponge Geodia barretti.J. Nat. Prod.781886–1893. 10.1021/ACS.JNATPROD.5B00210
20
Chen B. Fan D. Q. Zhu K. X. Shan Z. G. Chen F. Y. Hou L. et al (2015). Mechanism study on a new antimicrobial peptide Sphistin derived from the N-terminus of crab histone H2A identified in haemolymphs of Scylla paramamosain.Fish Shellfish Immunol.47833–846. 10.1016/j.fsi.2015.10.010
21
Chen G. Abelein A. Nilsson H. E. Leppert A. Andrade-Talavera Y. Tambaro S. et al (2017). Bri2 BRICHOS client specificity and chaperone activity are governed by assembly state.Nat. Commun.81–14. 10.1038/s41467-017-02056-4
22
Chen L. Wang S. (2017). Preparation of an ACE-inhibitory peptide from Perinereis aibuhitensis protein.Biotechnol. Biotechnol. Equip.311231–1236. 10.1080/13102818.2017.1370983
23
Chen S. L. Li W. Meng L. Sha Z. X. Wang Z. J. Ren G. C. (2007). Molecular cloning and expression analysis of a hepcidin antimicrobial peptide gene from turbot (Scophthalmus maximus).Fish Shellfish Immunol.22172–181. 10.1016/j.fsi.2006.04.004
24
Chen T. Zhou J. Qu Z. Zou Q. Liu X. Su J. et al (2020). Administration of dietary recombinant hepcidin on grass carp (Ctenopharyngodon idella) against Flavobacterium columnare infection under cage aquaculture conditions.Fish Shellfish Immunol.9927–34. 10.1016/j.fsi.2020.01.042
25
Cheung R. C. F. Ng T. B. Wong J. H. (2015). Marine peptides: bioactivities and applications.Mar. Drugs134006–4043. 10.3390/md13074006
26
Chevallier C. Richardson A. D. Edler M. C. Hamel E. Harper M. K. Ireland C. M. (2003). A new cytotoxic and tubulin-interactive milnamide derivative from a marine sponge Cymbastela sp.Org. Lett.53737–3739. 10.1021/ol035476c
27
Chiang Y.-M. Chang S.-L. Oakley B. R. Wang C. C. C. (2011). Recent advances in awakening silent biosynthetic gene clusters and linking orphan clusters to natural products in microorganisms.Curr. Opin. Chem. Biol.15137–143. 10.1016/j.cbpa.2010.10.011
28
Cho J. Lee D. G. (2011). The characteristic region of arenicin-1 involved with a bacterial membrane targeting mechanism.Biochem. Biophys. Res. Commun.405422–427. 10.1016/j.bbrc.2011.01.046
29
Clark D. P. Carroll J. Naylor S. Crews P. (1998). An antifungal cyclodepsipeptide, cyclolithistide A, from the sponge Theonella swinhoei.J. Org. Chem.638757–8764. 10.1021/jo980758p
30
Coleman J. E. Van Soest R. Andersen R. J. (1999). New geodiamolides from the sponge Cymbastela sp. collected in Papua New Guinea.J. Nat. Prod.621137–1141. 10.1021/np990155o
31
Coutinho M. C. L. Teixeira V. L. Santos C. S. G. (2017). A review of “Polychaeta” chemicals and their possible ecological role.J. Chem. Ecol.4472–94. 10.1007/s10886-017-0915-z
32
Daletos G. Kalscheuer R. Koliwer-Brandl H. Hartmann R. De Voogd N. J. Wray V. et al (2015). Callyaerins from the marine sponge Callyspongia aerizusa: cyclic peptides with antitubercular activity.J. Nat. Prod.781910–1925. 10.1021/acs.jnatprod.5b00266
33
Daly M. Brugler M. R. Cartwright P. Collins A. G. Dawson M. N. Fautin D. G. et al (2007). The Phylum Cnidaria: A Review of Phylogenetic Patterns and Diversity 300 Years After Linnaeus. Available online at: https://kuscholarworks.ku.edu/handle/1808/13641(accessed April 21, 2020).
34
de Paiva P. C. (2006). “Filo annelida: classe polychaeta,” in Biodiversidade Bentônica da região central da ZonaEconômica Exclusiva brasileira, edsLavradoH. P.IgnacioB. L. (Rio de Janeiro: Museu Nacional), 261–298.
35
De Silva E. D. Williams D. E. Andersen R. J. Klix H. Holmes C. F. B. Allen T. M. (1992). Motuporin, a potent protein phosphatase inhibitor isolated from the papua new guinea sponge Theonella swinhoei Gray.Tetrahedron Lett.331561–1564.
36
Faircloth G. Cuevas C. (2006). Kahalalide F and ES285: potent anticancer agents from marine molluscs.Prog. Mol. Subcell. Biol.43363–379. 10.1007/978-3-540-30880-5_16
37
Festa M. Sansone C. Brunet C. Crocetta F. Di Paola L. Lombardo M. et al (2020). Cardiovascular active peptides of marine origin with ACE inhibitory activities: potential role as anti-hypertensive drugs and in prevention of SARSCoV-2 infection.Int. J. Mol. Sci.211–20. 10.3390/ijms21218364
38
Gamble W. R. Durso N. A. Fuller R. W. Westergaard C. K. Johnson T. R. Sackett D. L. et al (1999). Cytotoxic and tubulin-interactive hemiasterlins from Auletta sp. and Siphonochalina spp. sponges.Bioorganic Med. Chem.71611–1615. 10.1016/S0968-0896(99)00089-9
39
Gerwick W. H. Moore B. S. (2012). Lessons from the past and charting the future of marine natural products drug discovery and chemical biology.Chem. Biol.1985–98. 10.1016/j.chembiol.2011.12.014
40
Giordano D. Costantini M. Coppola D. Lauritano C. Núñez Pons L. Ruocco N. et al (2018). “Biotechnological applications of bioactive peptides from marine sources,” in Advances in Microbial Physiology, ed.PooleR. K. (Cambridge, MA: Academic Press), 171–220.
41
Gong P.-X. Wang B.-K. Wu Y.-C. Li Q.-Y. Qin B.-W. Li H.-J. (2020). Release of antidiabetic peptides from Stichopus japonicas by simulated gastrointestinal digestion.Food Chem.315:126273. 10.1016/j.foodchem.2020.126273
42
Gracia C. Isidro-Llobet A. Cruz L. J. Acosta G. A. Álvarez M. Cuevas C. et al (2006). Convergent approaches for the synthesis of the antitumoral peptide, Kahalalide F. study of orthogonal protecting groups.J. Org. Chem.717196–7204. 10.1021/jo060976f
43
Gui L. Zhang P. Zhang Q. Zhang J. (2016). Fish & Shell fi sh Immunology Two hepcidins from spotted scat (Scatophagus argus) possess antibacterial and antiviral functions in vitro.Fish Shellfish Immunol.50191–199. 10.1016/j.fsi.2016.01.038
44
Hamada T. Matsunaga S. Yano G. Fusetani N. (2005). Polytheonamides A and B, highly cytotoxic, linear polypeptides with unprecedented structural features, from the marine sponge, Theonella swinhoei.J. Am. Chem. Soc.127110–118. 10.1021/ja045749e
45
Hamada T. Sugawara T. Matsunaga S. Fusetani N. (1994). Polytheonamides, unprecedented highly cytotoxic polypeptides from the marine sponge Theonella swinhoei 2. structure elucidation.Tetrahedron Lett.35609–612. 10.1016/S0040-4039(00)75851-5
46
He L. X. Zhang Z. F. Sun B. Chen Q. H. Liu R. Ren J. W. et al (2016). Sea cucumber (Codonopsis pilosula) oligopeptides: immunomodulatory effects based on stimulating Th cells, cytokine secretion and antibody production.Food Funct.71208–1216. 10.1039/c5fo01480h
47
Hsu K. C. Li-Chan E. C. Y. Jao C. L. (2011). Antiproliferative activity of peptides prepared from enzymatic hydrolysates of tuna dark muscle on human breast cancer cell line MCF-7.Food Chem.126617–622. 10.1016/j.foodchem.2010.11.066
48
Huang F. Jing Y. Ding G. Yang Z. (2017). Isolation and purification of Novel peptides derived from Sepia ink: effects on apoptosis of prostate cancer cell PC-3.Mol. Med. Rep.164222–4228. 10.3892/mmr.2017.7068
49
Hwang S. D. Joo M. S. Hwang J. Y. Kwon M. G. Jeong J. M. Seo J. S. et al (2019). Molecular characterization and gene expression data of liver expressed antimicrobial Peptide-2 (LEAP-2) isolated from rock bream (Oplegnathus fasciatus).Data Br.26:104538. 10.1016/j.dib.2019.104538
50
Ibrahim S. R. M. Min C. C. Teuscher F. Ebel R. Kakoschke C. Lin W. et al (2010). Callyaerins A-F and H, new cytotoxic cyclic peptides from the Indonesian marine sponge Callyspongia aerizusa.Bioorganic Med. Chem.184947–4956. 10.1016/j.bmc.2010.06.012
51
Jain A. Yadav B. K. Chugh A. (2015). Marine antimicrobial peptide tachyplesin as an efficient nanocarrier for macromolecule delivery in plant and mammalian cells.FEBS J.282732–745. 10.1111/febs.13178
52
Jia Y. Ding G. Yang Z. Yu F. Zheng Y. Wu Z. et al (2017). Anticancer activity of a novel peptide derived from hydrolysates of Perinereies aibuhitensis against lung cancer A549 Cells.Food Sci.3827–35.
53
Jiang S. Jia Y. Tang Y. Zheng D. Han X. Yu F. et al (2019). Anti-proliferation activity of a decapeptide from Perinereies aibuhitensis toward human lung cancer H1299 cells.Mar. Drugs17:122. 10.3390/md17020122
54
Jimeno J. Faircloth G. Sousa-Faro J. M. F. Scheuer P. Rinehart K. (2004). New marine derived anticancer therapeutics – a journey from the sea to clinical trials.Mar. Drugs214–29.
55
Jin H. X. Xu H. P. Li Y. Zhang Q. W. Xie H. (2019). Preparation and evaluation of peptides with potential antioxidant activity by microwave assisted enzymatic hydrolysis of collagen from sea cucumber Acaudina molpadioides obtained from Zhejiang province in China.Mar. Drugs17:169. 10.3390/md17030169
56
Kaas Q. Westermann J. C. Craik D. J. (2010). Conopeptide characterization and classifications: an analysis using ConoServer.Toxicon551491–1509. 10.1016/j.toxicon.2010.03.002
57
Kang H. K. Seo C. H. Park Y. (2015). Marine peptides and their anti-infective activities.Mar. Drugs13618–654. 10.3390/md13010618
58
Katz J. Janik J. E. Younes A. (2011). Brentuximab Vedotin (SGN-35).Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res.176428–6436. 10.1158/1078-0432.CCR-11-0488
59
Kim C. H. Go H. J. Oh H. Y. Park J. B. Lee T. K. Seo J. K. et al (2018). Identification of a novel antimicrobial peptide from the sea star Patiria pectinifera.Dev. Comp. Immunol.86203–213. 10.1016/j.dci.2018.05.002
60
Kita M. Gise B. Kawamura A. Kigoshi H. (2013). Stylissatin A, a cyclic peptide that inhibits nitric oxide production from the marine sponge Stylissa massa.Tetrahedron Lett.546826–6828. 10.1016/j.tetlet.2013.10.003
61
Kobayashi M. Kanzaki K. Katayama S. Ohashi K. Okada H. Ikegami S. et al (1994). Marine natural products. XXXIII.1) theonellapeptolide lid, a new tridecapeptide lactone from the okinawan marine sponge Theonella swinhoei.Chem. Pharm. Bull.421410–1415. 10.1248/cpb.42.1410
62
Koch T. L. Grimmelikhuijzen C. J. P. (2020). A comparative genomics study of neuropeptide genes in the cnidarian subclasses Hexacorallia and Ceriantharia.BMC Genomics21:666. 10.1186/s12864-020-06945-9
63
Laport M. S. Santos O. C. S. Muricy G. (2009). Marine sponges: potential sources of new antimicrobial drugs.Curr. Pharm. Biotechnol.1086–105. 10.2174/138920109787048625
64
Lebbe E. K. M. Ghequire M. G. K. Peigneur S. Mille B. G. Devi P. Ravichandran S. et al (2016). Novel conopeptides of largely unexplored indo Pacific Conus sp.Mar. Drugs14:199. 10.3390/md14110199
65
Lebbe E. K. M. Peigneur S. Maiti M. Devi P. Ravichandran S. Lescrinier E. et al (2014). Structure-function elucidation of a new α-conotoxin, Lo1a, from Conus longurionis.J. Biol. Chem.2899573–9583. 10.1074/jbc.M114.556175
66
Lee J. U. Kang D. I. L. Zhu W. L. Shin S. Y. Hahm K. S. Kim Y. (2007). Solution structures and biological functions of the antimicrobial peptide, arenicin-1, and its linear derivative.Pept. Sci. Orig. Res. Biomol.88208–216. 10.1002/bip.20700
67
Lee S. Y. Hur S. J. (2017). Antihypertensive peptides from animal products, marine organisms, and plants.Food Chem.228506–517. 10.1016/j.foodchem.2017.02.039
68
Lemes A. Sala L. Ores J. Braga A. Egea M. Fernandes K. (2016). A review of the latest advances in encrypted bioactive peptides from protein-rich waste.Int. J. Mol. Sci.17:950. 10.3390/ijms17060950
69
Li C. Blencke H. M. Haug T. Stensvåg K. (2015). Antimicrobial peptides in echinoderm host defense.Dev. Comp. Immunol.49190–197. 10.1016/j.dci.2014.11.002
70
Li C. Haug T. Moe M. K. Styrvold O. B. Stensvåg K. (2010). Centrocins: isolation and characterization of novel dimeric antimicrobial peptides from the green sea urchin, Strongylocentrotus droebachiensis.Dev. Comp. Immunol.34959–968. 10.1016/j.dci.2010.04.004
71
Li D. Li L. Xu T. Wang T. Ren J. Liu X. et al (2018). Effect of low molecular weight oligopeptides isolated from sea cucumber on diabetic wound healing in db/db mice.Mar. Drugs16:16. 10.3390/md16010016
72
Li H. Matsunaga S. Fusetani N. (1995). Halicylindramides A-C, antifungal and cytotoxic depsipeptides from the marine sponge halichondria cylindratal.J Med Chem.338–343.
73
Li H. Zhang F. Guo H. Zhu Y. Yuan J. Yang G. et al (2013). Molecular characterization of hepcidin gene in common carp (Cyprinus carpio L.) and its expression pattern responding to bacterial challenge.Fish Shellfish Immunol.351030–1038. 10.1016/j.fsi.2013.07.001
74
Li H. Y. Matsunaga S. Fusetani N. (1996). Halicylindramides D and E, antifungal peptides from the marine sponge Halichondria cylindrata.J. Nat. Prod.59163–166. 10.1021/np9600309
75
Li J. Liu Z. Zhao Y. Zhu X. Yu R. Dong S. et al (2018). Novel natural angiotensin converting enzyme (ACE)-inhibitory peptides derived from sea cucumber-modified hydrolysates by adding exogenous proline and a study of their structure↓activity relationship.Mar. Drugs16:271. 10.3390/md16080271
76
Li X. Tang Y. Yu F. Sun Y. Huang F. Chen Y. et al (2018). Inhibition of prostate cancer DU-145 cells proliferation by anthopleura anjunae oligopeptide (YVPGP) via PI3K/AKT/mTOR signaling pathway.Mar. Drugs16325. 10.3390/md16090325
77
Li Y. Chang Q. Wu M. Zhao X. (2018). Total synthesis of five proline-enriched cyclic heptapeptides from the marine sponge Stylissa carteri.Tetrahedron Lett.591828–1831. 10.1016/j.tetlet.2018.03.083
78
Li Y. Li J. Liu T. Wang Y. Zhou Z. Cheng F. et al (2017). Preparation and antithrombotic activity identification of Perinereis aibuhitensis extract: a high temperature and wide pH range stable biological agent.Food Funct.83533–3541. 10.1039/c7fo00987a
79
Liao Z. Wang X. Liu H. Fan M. Sun J. Shen W. (2013). Molecular characterization of a novel antimicrobial peptide from Mytilus coruscus.Fish Shellfish Immunol.34610–616. 10.1016/j.fsi.2012.11.030
80
Lie Tasiemski A. Jung S. Line Boidin-Wichlacz C. Jollivet D. Cuvillier-Hot V. Pradillon F. et al (2014). Characterization and function of the first antibiotic isolated from a vent organism: the extremophile metazoan Alvinella pompejana.PLoS One9:e95737. 10.1371/journal.pone.0095737
81
Lima L. Migliolo L. Castro C. Pires D. Lopez-Abarrategui C. Goncalves E. et al (2013). Identification of a novel antimicrobial peptide from brazilian coast coral Phyllogorgia dilatata.Protein Pept. Lett.201153–1158. 10.2174/0929866511320100010
82
Liu R. Qiu L. Yu Z. Zi J. Yue F. Wang L. et al (2013). Identification and characterisation of pathogenic Vibrio splendidus from Yesso scallop (Patinopecten yessoensis) cultured in a low temperature environment.J. Invertebr. Pathol.114144–150. 10.1016/j.jip.2013.07.005
83
Liu X. Wang Z. Yin F. Liu Y. Qin N. Nakamura Y. et al (2019a). Zinc-chelating mechanism of sea cucumber (Stichopus japonicus)-derived synthetic peptides.Mar. Drugs17:438. 10.3390/md17080438
84
Liu X. Wang Z. Zhang J. Song L. Li D. Wu Z. et al (2019b). Isolation and identification of zinc-chelating peptides from sea cucumber (Stichopus japonicus) protein hydrolysate.J. Sci. Food Agric.996400–6407. 10.1002/jsfa.9919
85
Liu Z. M. Chen J. Lv Y. P. Hu Z. H. Dai Q. M. Fan X. L. (2018). Molecular characterization of a hepcidin homologue in starry flounder (Platichthys stellatus) and its synergistic interaction with antibiotics.Fish Shellfish Immunol.8345–51. 10.1016/j.fsi.2018.09.019
86
Lu Z. Van Wagoner R. M. Harper M. K. Baker H. L. Hooper J. N. A. Bewley C. A. et al (2011). Mirabamides E-H, HIV-inhibitory depsipeptides from the sponge Stelletta clavosa.J. Nat. Prod.74185–193. 10.1021/np100613p
87
Luesch H. Moore R. E. Paul V. J. Mooberry S. L. Corbett T. H. (2001). Isolation of dolastatin 10 from the marine cyanobacterium Symploca species VP642 and total stereochemistry and biological evaluation of its analogue symplostatin 1.J. Nat. Prod.64907–910. 10.1021/np010049y
88
Mahapatra G. P. Raman S. Nayak S. Gouda S. Das G. Patra J. K. (2020). Metagenomics approaches in discovery and development of new bioactive compounds from marine actinomycetes.Curr. Microbiol.77645–656. 10.1007/s00284-019-01698-5
89
Mariano G. H. Gomes de Sá L. G. Carmo da Silva E. M. Santos M. A. Cardozo Fh J. L. Lira B. O. V. et al (2021). Characterization of novel human intragenic antimicrobial peptides, incorporation and release studies from ureasil-polyether hybrid matrix.Mater. Sci. Eng. C119:111581. 10.1016/j.msec.2020.111581
90
Matsunaga S. Fusetani N. Hashimoto K. Walchli M. (1989). Theonellamide F. A novel antifungal bicyclic peptide from a marine sponge Theonella sp.J. Am. Chem. Soc.1112582–2588. 10.1021/ja00189a035
91
Mehbub M. F. Lei J. Franco C. Zhang W. (2014). Marine sponge derived natural products between 2001 and 2010: trends and opportunities for discovery of bioactives.Mar. Drugs124539–4577. 10.3390/md12084539
92
Meli A. Tedesco C. Della Sala G. Schettini R. Albericio F. De Riccardis F. et al (2017). Phakellistatins: an underwater unsolved puzzle.Mar. Drugs15:78. 10.3390/md15030078
93
Mihailescu M. Sorci M. Seckute J. Silin V. I. Hammer J. Perrin B. S. et al (2019). Structure and function in antimicrobial piscidins: histidine position, directionality of membrane insertion, and pH-dependent permeabilization.J. Am. Chem. Soc.1419837–9853. 10.1021/jacs.9b00440
94
Moe M. K. Haug T. Sydnes M. O. Sperstad S. V. Li C. Vaagsfjord L. C. et al (2018). Paralithocins, antimicrobial peptides with unusual disulfide connectivity from the red king crab, Paralithodes camtschaticus.J. Nat. Prod.81140–150. 10.1021/acs.jnatprod.7b00780
95
Molinski T. F. Dalisay D. S. Lievens S. L. Saludes J. P. (2009). Drug development from marine natural products.Nat. Rev. Drug Discov.869–85. 10.1038/nrd2487
96
Mora C. Tittensor D. P. Adl S. Simpson A. G. B. Worm B. (2011). How many species are there on Earth and in the ocean?PLoS Biol.9:e1001127. 10.1371/journal.pbio.1001127
97
Mulder K. C. de Lima L. A. Aguiar P. S. Carneiro F. C. Franco O. L. Dias S. C. et al (2015). Production of a modified peptide clavanin in Pichia pastoris: cloning, expression, purification and in vitro activities.AMB Express51–8. 10.1186/s13568-015-0129-0
98
Ngo D. H. Vo T. S. Ngo D. N. Wijesekara I. Kim S. K. (2012). Biological activities and potential health benefits of bioactive peptides derived from marine organisms.Int. J. Biol. Macromol.51378–383. 10.1016/j.ijbiomac.2012.06.001
99
Nikapitiya C. Dananjaya S. H. S. Chandrarathna H. P. S. U. de Zoysa M. Whang I. (2020). Octominin: a novel synthetic anticandidal peptide derived from defense protein of octopus minor.Mar. Drugs18:56. 10.3390/md18010056
100
Nishimura S. Arita Y. Honda M. Iwamoto K. Matsuyama A. Shirai A. et al (2010). Marine antifungal theonellamides target 3beta-hydroxysterol to activate Rho1 signaling.Nat. Chem. Biol.6519–526. 10.1038/nchembio.387
101
Niu S. F. Jin Y. Xu X. Qiao Y. Wu Y. Mao Y. et al (2013). Characterization of a novel piscidin-like antimicrobial peptide from Pseudosciaena crocea and its immune response to Cryptocaryon irritans.Fish Shellfish Immunol.35513–524. 10.1016/j.fsi.2013.05.007
102
Novoa B. Romero A. Álvarez ÁL. Moreira R. Pereiro P. Costa M. M. et al (2016). Antiviral activity of myticin C peptide from mussel: an ancient defense against herpesviruses.J. Virol.907692–7702. 10.1128/JVI.00591-16
103
Noyer C. Thomas O. P. Becerro M. A. (2011). Patterns of chemical diversity in the Mediterranean sponge Spongia lamella.PLoS One6:e20844. 10.1371/journal.pone.0020844
104
Núñez-Acuña G. Gallardo-Escárate C. Fields D. M. Shema S. Skiftesvik A. B. Ormazábal I. et al (2018). The Atlantic salmon (Salmo salar) antimicrobial peptide cathelicidin-2 is a molecular host-associated cue for the salmon louse (Lepeophtheirus salmonis).Sci. Rep.8:13738. 10.1038/s41598-018-31885-6
105
Oh R. Lee M. J. Kim Y.-O. Nam B.-H. Kong H. J. Kim J.-W. et al (2018). Purification and characterization of an antimicrobial peptide mytichitin-chitin binding domain from the hard-shelled mussel, {Mytilus} coruscus.Fish Shellfish Immunol.83425–435. 10.1016/j.fsi.2018.09.009
106
Oh R. Lee M. J. Kim Y. O. Nam B. H. Kong H. J. Kim J. W. et al (2020). Myticusin-beta, antimicrobial peptide from the marine bivalve, Mytilus coruscus.Fish Shellfish Immunol.99342–352. 10.1016/j.fsi.2020.02.020
107
Orlov D. S. Shamova O. V. Eliseev I. E. Zharkova M. S. Chakchir O. B. Antcheva N. et al (2019). Redesigning arenicin-1, an antimicrobial peptide from the marine polychaeta arenicola marina, by strand rearrangement or branching, substitution of specific residues, and backbone linearization or cyclization.Mar. Drugs17:376. 10.3390/md17060376
108
Otero-Gonzáiez A. J. Magalhães B. S. Garcia-Villarino M. López-Abarrategui C. Sousa D. A. Dias S. C. et al (2010). Antimicrobial peptides from marine invertebrates as a new frontier for microbial infection control.FASEB J.241320–1334. 10.1096/fj.09-143388
109
Ovchinnikova T. V. Aleshina G. M. Balandin S. V. Krasnosdembskaya A. D. Markelov M. L. Frolova E. I. et al (2004). Purification and primary structure of two isoforms of arenicin, a novel antimicrobial peptide from marine polychaeta Arenicola marina.FEBS Lett.577209–214. 10.1016/j.febslet.2004.10.012
110
Ovchinnikova T. V. Balandin S. V. Aleshina G. M. Tagaev A. A. Leonova Y. F. Krasnodembsky E. D. et al (2006). Aurelin, a novel antimicrobial peptide from jellyfish Aurelia aurita with structural features of defensins and channel-blocking toxins.Biochem. Biophys. Res. Commun.348514–523. 10.1016/j.bbrc.2006.07.078
111
Pan W. Liu X. Ge F. Han J. Zheng T. (2004). Perinerin, a novel antimicrobial peptide purified from the clamworm Perinereis aibuhitensis grube and its partial characterization.J. Biochem.135297–304. 10.1093/jb/mvh036
112
Panteleev P. V. Tsarev A. V. Bolosov I. A. Paramonov A. S. Marggraf M. B. Sychev S. V. et al (2018). Novel antimicrobial peptides from the arctic polychaeta nicomache minor provide new molecular insight into biological role of the BRICHOS domain.Mar. Drugs16:401. 10.3390/md16110401
113
Panteleev P. V. Tsarev A. V. Safronova V. N. Reznikova O. V. Bolosov I. A. Sychev S. V. et al (2020). Structure elucidation and functional studies of a novel β-hairpin antimicrobial peptide from the marine polychaeta Capitella teleta.Mar. Drugs18:620. 10.3390/md18120620
114
Park C. Lee D. G. (2009). Fungicidal effect of antimicrobial peptide arenicin-1.Biochim. Biophys. Acta Biomembr.17881790–1796. 10.1016/j.bbamem.2009.06.008
115
Park Y. R. Park C. I. L. Soh Y. (2020). Antioxidant and anti-inflammatory effects of NCW peptide from clam worm (Marphysa sanguinea).J. Microbiol. Biotechnol.301387–1394. 10.4014/jmb.2003.03050
116
Pavlicevic M. Maestri E. Marmiroli M. (2020). Marine bioactive peptides↔an overview of generation, structure and application with a focus on food sources.Mar. Drugs18:424. 10.3390/MD18080424
117
Peng Z. Chen B. Zheng Q. Zhu G. Cao W. Qin X. et al (2020). Ameliorative effects of peptides from the oyster (Crassostrea hongkongensis) protein hydrolysates against UVB-induced skin photodamage in mice.Mar. Drugs18:288. 10.3390/md18060288
118
Pettit G. R. Kamano Y. Herald C. L. Tuinman A. A. Boettner F. E. Kizu H. et al (1987). The isolation and structure of a remarkable marine animal antineoplastic constituent: dolastatin 10.J. Am. Chem. Soc.1096883–6885. 10.1021/ja00256a070
119
Plaza A. Bifulco G. Keffer J. L. Lloyd J. R. Baker H. L. Bewley C. A. (2009). Celebesides A-C and theopapuamides B-D, depsipeptides from an Indonesian sponge that inhibit HIV-1 entry.J. Org. Chem.74504–512. 10.1021/jo802232u
120
Plaza A. Bifulco G. Masullo M. Lloyd J. R. Keffer J. L. Colin P. L. et al (2010). Mutremdamide A and Koshikamides C-H, peptide inhibitors of HIV-1 entry from different Theonella species.J. Org. Chem.754344–4355. 10.1021/jo100076g
121
Plaza A. Gustchina E. Baker H. L. Kelly M. Bewley C. A. (2007). Mirabamides A-D, depsipeptides from the sponge Siliquariaspongia mirabilis that inhibit HIV-1 fusion.J. Nat. Prod.701753–1760. 10.1021/np070306k
122
Pushpanathan M. Gunasekaran P. Rajendhran J. (2013). Antimicrobial peptides: versatile biological properties.Int. J. Pept.2013:675391. 10.1155/2013/675391
123
Qian P.-Y. Li Z. Xu Y. Li Y. Fusetani N. (2015). Mini-review: marine natural products and their synthetic analogs as antifouling compounds: 2009-2014.Biofouling31101–122. 10.1080/08927014.2014.997226
124
Qin C. Huang W. Zhou S. Wang X. Liu H. Fan M. et al (2014). Characterization of a novel antimicrobial peptide with chitin-biding domain from Mytilus coruscus.Fish Shellfish Immunol.41362–370. 10.1016/j.fsi.2014.09.019
125
Quah Y. Mohd Ismail N. I. Ooi J. L. S. Affendi Y. A. Abd Manan F. Teh L. K. et al (2019). Purification and identification of novel cytotoxic oligopeptides from soft coral Sarcophyton glaucum.J. Zhejiang Univ. Sci. B2059–70. 10.1631/jzus.B1700586
126
Rajanbabu V. Chen J. Y. Wu J. L. (2015). Antimicrobial Peptides from Marine Organisms.Berlin: Springer, 747–758.
127
Rao K. V. Na M. K. Cook J. C. Peng J. Matsumoto R. Hamann M. T. (2008). Kahalalides V-Y isolated from a Hawaiian collection of the sacoglossan mollusk Elysia rufescens.J. Nat. Prod.71772–778. 10.1021/np070508g
128
Reen F. J. Gutiérrez-Barranquero J. A. Dobson A. D. W. Adams C. O’Gara F. (2015a). Emerging concepts promising new horizons for marine biodiscovery and synthetic biology.Mar. Drugs132924–2954. 10.3390/md13052924
129
Reen F. J. Romano S. Dobson A. D. W. O’Gara F. (2015b). The sound of silence: activating silent biosynthetic gene clusters in marine microorganisms.Mar. Drugs134754–4783. 10.3390/md13084754
130
Reveillaud J. Maignien L. Murat Eren A. Huber J. A. Apprill A. Sogin M. L. et al (2014). Host-specificity among abundant and rare taxa in the sponge microbiome.ISME J.81198–1209. 10.1038/ismej.2013.227
131
Rigo F. K. Dalmolin G. D. Trevisan G. Tonello R. Silva M. A. Rossato M. F. et al (2013). Effect of ω-conotoxin MVIIA and Phα1β on paclitaxel-induced acute and chronic pain.Pharmacol. Biochem. Behav.1116–22. 10.1016/j.pbb.2013.10.014
132
Rinehart K. L. Gloer J. B. Hughes R. G. Renis H. E. McGovren J. P. Swynenberg E. B. et al (1981). Didemnins: antiviral and antitumor depsipeptides from a Caribbean tunicate.Science212933–935. 10.1126/science.7233187
133
Robert M. Zatylny-Gaudin C. Fournier V. Corre E. Le Corguillé G. Bernay B. et al (2014). Transcriptomic and peptidomic analysis of protein hydrolysates from the white shrimp (L. vannamei).J. Biotechnol.18630–37. 10.1016/j.jbiotec.2014.06.020
134
Rodrigo A. P. Mendes V. M. Manadas B. Grosso A. R. de Matos A. P. A. Baptista P. V. et al (2021). Specific antiproliferative properties of proteinaceous toxin secretions from the marine annelid Eulalia sp. onto Ovarian Cancer Cells.Mar. Drugs19:31. 10.3390/md19010031
135
Romano G. Costantini M. Sansone C. Lauritano C. Ruocco N. Ianora A. (2017). Marine microorganisms as a promising and sustainable source of bioactive molecules.Mar. Environ. Res.12858–69. 10.1016/j.marenvres.2016.05.002
136
Ruiz-Ruiz F. Mancera-Andrade E. I. Iqbal H. M. N. (2017). Marine-derived bioactive peptides for biomedical sectors: a review.Protein Pept. Lett.24109–117. 10.2174/0929866523666160802155347
137
Sable R. Parajuli P. Jois S. (2017). Peptides, peptidomimetics, and polypeptides from marine sources: a wealth of natural sources for pharmaceutical applications.Mar. Drugs15:124. 10.3390/md15040124
138
Sacristán-Soriano O. Banaigs B. Casamayor E. O. Becerro M. A. (2011). Exploring the links between natural products and bacterial assemblages in the sponge Aplysina aerophoba.Appl. Environ. Microbiol.77862–870. 10.1128/AEM.00100-10
139
Sagar S. Kaur M. Minneman K. P. (2010). Antiviral lead compounds from marine sponges.Mar. Drugs82619–2638. 10.3390/md8102619
140
Sanjeewa A. K. K. Nagahawatta D. P. Yang H. W. Oh J. Y. Jayawardena T. U. Jeon Y. J. et al (2020). Octominin inhibits LPS-induced chemokine and pro-inflammatory cytokine secretion from raw 264.7 macrophages via blocking TLRS/nf-κb signal transduction.Biomolecules10:511. 10.3390/biom10040511
141
Satpute S. K. Banat I. M. Dhakephalkar P. K. Banpurkar A. G. Chopade B. A. (2010). Biosurfactants, bioemulsifiers and exopolysaccharides from marine microorganisms.Biotechnol. Adv.28436–450. 10.1016/j.biotechadv.2010.02.006
142
Schneemann I. Wiese J. Kunz A. L. Imhoff J. F. (2011). Genetic approach for the fast discovery of phenazine producing bacteria.Mar. Drugs9772–789. 10.3390/md9050772
143
Schenk S. Hoeger U. (2020). “Annelid coelomic fluid proteins,” in Subcellular Biochemistry, ed.RobinH. J. (Cham: Springer), 1–34.
144
Shan Z. Zhu K. Peng H. Chen B. Liu J. Chen F. (2016). The new antimicrobial peptide sphyastatin from the mud crab Scylla paramamosain with multiple antimicrobial mechanisms and high effect on bacterial infection.Front. Microbiol.7:1140. 10.3389/fmicb.2016.01140
145
Sharma S. Chatterji A. Das P. (2009). Effect of different extraction procedures on antimicrobial activity of marine bivalves: a comparison.Pertanika J. Trop. Agric. Sci.3277–83.
146
Shen Q. Zeng M. Univ Y. Z. (2014). Modification of Acaudina molpadioides hydrolysates by plastein reaction and preparation of ACE-inhibitory peptides.Chem. J. Chin. Univ.35965–970.
147
Shenkarev Z. O. Balandin S. V. Trunov K. I. Paramonov A. S. Sukhanov S. V. Barsukov L. I. et al (2011). Molecular mechanism of action of β-Hairpin antimicrobial peptide arenicin: oligomeric structure in dodecylphosphocholine micelles and pore formation in planar lipid bilayers.Biochemistry506255–6265. 10.1021/bi200746t
148
Shenkarev Z. O. Panteleev P. V. Balandin S. V. Gizatullina A. K. Altukhov D. A. Finkina E. I. et al (2012). Recombinant expression and solution structure of antimicrobial peptide aurelin from jellyfish Aurelia aurita.Biochem. Biophys. Res. Commun.42963–69. 10.1016/j.bbrc.2012.10.092
149
Shike H. Lauth X. Westerman M. E. Ostland V. E. Carlberg J. M. Van Olst J. C. et al (2002). Bass hepcidin is a novel antimicrobial peptide induced by bacterial challenge.Eur. J. Biochem.2692232–2237. 10.1046/j.1432-1033.2002.02881.x
150
Shin H. J. Rashid M. A. Cartner L. K. Bokesch H. R. Wilson J. A. McMahon J. B. et al (2015). Stellettapeptins A and B, HIV-inhibitory cyclic depsipeptides from the marine sponge Stelletta sp.Tetrahedron Lett.564215–4219. 10.1016/j.tetlet.2015.05.058
151
Sila A. Bougatef A. (2016). Antioxidant peptides from marine by-products: isolation, identification and application in food systems. A review.J. Funct. Foods2110–26. 10.1016/j.jff.2015.11.007
152
Simmons T. L. Coates R. C. Clark B. R. Engene N. Gonzalez D. Esquenazi E. et al (2008). Biosynthetic origin of natural products isolated from marine microorganism-invertebrate assemblages.Proc. Natl. Acad. Sci. U.S.A.1054587–4594. 10.1073/pnas.0709851105
153
Sjögren M. Göransson U. Johnson A.-L. Dahlström M. Andersson R. Bergman J. et al (2004). Antifouling activity of brominated cyclopeptides from the marine sponge Geodia barretti.J. Nat. Prod.67368–372. 10.1021/np0302403
154
Sjögren M. Johnson A.-L. Hedner E. Dahlström M. Göransson U. Shirani H. et al (2006). Antifouling activity of synthesized peptide analogs of the sponge metabolite barettin.Peptides272058–2064. 10.1016/j.peptides.2006.03.027
155
Smith D. Buddie A. G. Goss R. J. M. Overmann J. Lepleux C. Brönstrup M. et al (2019). Discovery pipelines for marine resources: an ocean of opportunity for biotechnology?World J. Microbiol. Biotechnol.35:107. 10.1007/s11274-019-2685-y
156
Solstad R. G. Johansen C. Stensvåg K. Strøm M. B. Haug T. (2020). Structure-activity relationship studies of shortened analogues of the antimicrobial peptide EeCentrocin 1 from the sea urchin Echinus esculentus.J. Pept. Sci.26:e3233. 10.1002/psc.3233
157
Solstad R. G. Li C. Isaksson J. Johansen J. Svenson J. Stensvåg K. et al (2016). Novel antimicrobial peptides EeCentrocins 1, 2 and EeStrongylocin 2 from the edible sea urchin Echinus esculentus have 6-Br-Trp post-translational modifications.PLoS One11:e0151820. 10.1371/journal.pone.0151820
158
Sonnenschein R. N. Farias J. J. Tenney K. Mooberry S. L. Lobkovsky E. Clardy J. et al (2004). A further study of the cytotoxic constituents of a milnamide-producing sponge.Org. Lett.6779–782. 10.1021/ol036446c
159
Sperstad S. V. Haug T. Blencke H. M. Styrvold O. B. Li C. Stensvåg K. (2011). Antimicrobial peptides from marine invertebrates: challenges and perspectives in marine antimicrobial peptide discovery.Biotechnol. Adv.29519–530. 10.1016/j.biotechadv.2011.05.021
160
Suarez-Jimenez G.-M. Burgos-Hernandez A. Ezquerra-Brauer J.-M. (2012). Bioactive peptides and depsipeptides with anticancer potential: sources from marine animals.Mar. Drugs10963–986. 10.3390/md10050963
161
Sun Y. Huang Y. Li X. Baldwin C. C. Zhou Z. Yan Z. et al (2016). Fish-T1K (Transcriptomes of 1,000 Fishes) Project: large-scale transcriptome data for fish evolution studies.Gigascience5:18. 10.1186/s13742-016-0124-7
162
Teta R. Irollo E. Della Sala G. Pirozzi G. Mangoni A. Costantino V. (2013). Smenamides A and B, chlorinated peptide/polyketide hybrids containing a dolapyrrolidinone unit from the Caribbean sponge Smenospongia aurea. Evaluation of their role as leads in antitumor drug research.Mar. Drugs114451–4463. 10.3390/md11114451
163
Tran T. D. Pham N. B. Fechner G. A. Hooper J. N. A. Quinn R. J. (2014). Potent cytotoxic peptides from the Australian marine sponge Pipestela candelabra.Mar. Drugs123399–3415. 10.3390/md12063399
164
Trindade M. van Zyl L. J. Navarro-Fernández J. Elrazak A. A. (2015). Targeted metagenomics as a tool to tap into marine natural product diversity for the discovery and production of drug candidates.Front. Microbiol.6:890. 10.3389/fmicb.2015.00890
165
Tsukimoto M. Nagaoka M. Shishido Y. Fujimoto J. Nishisaka F. Matsumoto S. et al (2011). Bacterial production of the tunicate-derived antitumor cyclic depsipeptide didemnin B.J. Nat. Prod.742329–2331. 10.1021/np200543z
166
Turk T. Kem W. R. (2009). The phylum Cnidaria and investigations of its toxins and venoms until 1990.Toxicon541031–1037. 10.1016/j.toxicon.2009.06.031
167
van Soest R. W. M. Boury-Esnault N. Vacelet J. Dohrmann M. Erpenbeck D. de Voogd N. J. et al (2012). Global diversity of sponges (Porifera).PLoS One7:e35105. 10.1371/journal.pone.0035105
168
Vitali A. (2018). Antimicrobial peptides derived from marine sponges.Am. J. Clin. Microbiol. Antimicrob.1:12.
169
Wang L. Dong C. Li X. Han W. Su X. (2017). Anticancer potential of bioactive peptides from animal sources (Review).Oncol. Rep.38637–651. 10.3892/or.2017.5778
170
Wang X. Morinaka B. I. Molinski T. F. (2014a). Structures and solution conformational dynamics of stylissamides G and H from the Bahamian Sponge Stylissa caribica.J. Nat. Prod.77625–630. 10.1021/np400891s
171
Wang X. Wang X. Teng D. Zhang Y. Mao R. Xi D. et al (2014b). Candidacidal mechanism of the arenicin-3-derived peptide NZ17074 from Arenicola marina.Appl. Microbiol. Biotechnol.987387–7398. 10.1007/s00253-014-5784-6
172
Wang X. Yu H. Xing R. Li P. (2017). Characterization, preparation, and purification of marine bioactive peptides.Biomed Res. Int.2017:9746720. 10.1155/2017/9746720
173
Wang Y.-N. Meng L.-H. Wang B.-G. (2020). Progress in research on bioactive secondary metabolites from deep-sea derived microorganisms.Mar. Drugs18:614. 10.3390/md18120614
174
Wang Z. Wang X. Wang J. (2018). Recent advances in antibacterial and antiendotoxic peptides or proteins from marine resources.Mar. Drugs16:57. 10.3390/md16020057
175
Wei L. Gao J. Zhang S. Wu S. Xie Z. Ling G. et al (2015). Identification and characterization of the first cathelicidin from sea snakes with potent antimicrobial and anti- inflammatory activity and special mechanism.J. Biol. Chem.29016633–16652. 10.1074/jbc.M115.642645
176
Wei X. Sarath Babu V. Lin L. Hu Y. Zhang Y. Liu X. et al (2018). Hepcidin protects grass carp (Ctenopharyngodon idellus) against Flavobacterium columnare infection via regulating iron distribution and immune gene expression.Fish Shellfish Immunol.75274–283. 10.1016/j.fsi.2018.02.023
177
Wilson M. C. Piel J. (2013). Metagenomic approaches for exploiting uncultivated bacteria as a resource for novel biosynthetic enzymology.Chem. Biol.20636–647. 10.1016/j.chembiol.2013.04.011
178
Woodhouse J. N. Fan L. Brown M. V. Thomas T. Neilan B. A. (2013). Deep sequencing of non-ribosomal peptide synthetases and polyketide synthases from the microbiomes of Australian marine sponges.ISME J.71842–1851. 10.1038/ismej.2013.65
179
Wu Y. Liu L. Chen H. F. Jiao W. H. Sun F. Liu L. Y. et al (2019). Fuscasins A-D, cycloheptapeptides from the marine sponge Phakellia fusca.J. Nat. Prod.82970–979. 10.1021/acs.jnatprod.8b01033
180
Wu Z. Z. Ding G. F. Huang F. F. Yang Z. S. Yu F. M. Tang Y. P. et al (2018). Anticancer activity of Anthopleura anjunae oligopeptides in prostate cancer DU-145 cells.Mar. Drugs16:125. 10.3390/md16040125
181
Xie B. Huang Y. Baumann K. Fry B. G. Shi Q. (2017). From marine venoms to drugs: efficiently supported by a combination of transcriptomics and proteomics.Mar. Drugs15:103. 10.3390/md15040103
182
Xiong Z.-Q. Wang J.-F. Hao Y.-Y. Wang Y. (2013). Recent advances in the discovery and development of marine microbial natural products.Mar. Drugs11700–717. 10.3390/md11030700
183
Xu Y. Kersten R. D. Nam S. J. Lu L. Al-Suwailem A. M. Zheng H. et al (2012). Bacterial biosynthesis and maturation of the didemnin anti-cancer agents.J. Am. Chem. Soc.1348625–8632. 10.1021/ja301735a
184
Yang D. Han Y. Chen L. Cao R. Wang Q. Dong Z. et al (2019). A macin identified from Venerupis philippinarum: investigation on antibacterial activities and action mode.Fish Shellfish Immunol.92897–904. 10.1016/j.fsi.2019.07.031
185
Yang D. Zhang Q. Wang Q. Chen L. Liu Y. Cong M. et al (2018). A defensin-like antimicrobial peptide from the manila clam Ruditapes philippinarum: investigation of the antibacterial activities and mode of action.Fish Shellfish Immunol.80274–280. 10.1016/j.fsi.2018.06.019
186
Younes A. Yasothan U. Kirkpatrick P. (2012). Brentuximab vedotin.Nat. Rev. Drug Discov.11119–20. 10.1038/nrd3629
187
Youssef D. T. A. Shaala L. A. Mohamed G. A. Badr J. M. Bamanie F. H. Ibrahim S. R. M. (2014). Theonellamide G, a potent antifungal and cytotoxic bicyclic glycopeptide from the Red Sea marine sponge Theonella swinhoei.Mar. Drugs121911–1923. 10.3390/md12041911
188
Zan J. Li Z. Diarey Tianero M. Davis J. Hill R. T. Donia M. S. (2019). A microbial factory for defensive kahalalides in a tripartite marine symbiosis.Science364:eaaw6732. 10.1126/science.aaw6732
189
Zhang J. Yu L. Li M. Sun L. (2014). Fish & Shell fi sh Immunology Turbot (Scophthalmus maximus) hepcidin-1 and hepcidin-2 possess antimicrobial activity and promote resistance against bacterial and viral infection.Fish Shellfish Immunol.38127–134. 10.1016/j.fsi.2014.03.011
190
Zhang L. An R. Wang J. Sun N. Zhang S. Hu J. et al (2005). Exploring novel bioactive compounds from marine microbes.Curr. Opin. Microbiol.8276–281. 10.1016/j.mib.2005.04.008
191
Zhang L. Yang D. Wang Q. Yuan Z. Wu H. Pei D. et al (2015). A defensin from clam Venerupis philippinarum: molecular characterization, localization, antibacterial activity, and mechanism of action.Dev. Comp. Immunol.5129–38. 10.1016/j.dci.2015.02.009
192
Zhang M. Sunaba T. Sun Y. Sasaki K. Isoda H. Kigoshi H. et al (2019). Anti-inflammatory marine cyclic peptide stylissatin A and its derivatives inhibit differentiation of murine preadipocytes.Chem. Commun.555471–5474.
193
Zhang M. Wang Y. Li Y. Li W. Li R. Xie X. et al (2018). Identification and characterization of neuropeptides by transcriptome and proteome analyses in a bivalve mollusc Patinopecten yessoensis.Front. Genet.9:197. 10.3389/fgene.2018.00197
194
Zhao J. Li C. Chen A. Li L. Su X. Li T. (2010). Molecular characterization of a novel big defensin from clam Venerupis philippinarum.PLoS One5:e0013480. 10.1371/journal.pone.0013480
195
Zheng L. H. Wang Y. J. Sheng J. Wang F. Zheng Y. Lin X. K. et al (2011). Antitumor peptides from marine organisms.Mar. Drugs91840–1859.
196
Zhong J. Wang W. Yang X. Yan X. Liu R. (2013). A novel cysteine-rich antimicrobial peptide from the mucus of the snail of Achatina fulica.Peptides391–5. 10.1016/j.peptides.2012.09.001
197
Zhou L. Li G. Jiao Y. Huang D. Li A. Chen H. et al (2019). Molecular and antimicrobial characterization of a group G anti-lipopolysaccharide factor (ALF) from Penaeus monodon.Fish Shellfish Immunol.94149–156. 10.1016/j.fsi.2019.08.066
198
Zhu X. J. Zeng M. Y. Ma G. L. Zhao Y. H. Liu Z. Y. (2011). Research on plastein reaction-basic modification of Acaudina molpadioidea protein hydrolysates and its effects on ACE activity.Chin. J. Mar Drugs306–12.
Summary
Keywords
bioactive peptides, pharmaceutical, biotechnological prospecting, multiple activities, marine organisms
Citation
Macedo MWFS, Cunha NBd, Carneiro JA, Costa RAd, Alencar SAd, Cardoso MH, Franco OL and Dias SC (2021) Marine Organisms as a Rich Source of Biologically Active Peptides. Front. Mar. Sci. 8:667764. doi: 10.3389/fmars.2021.667764
Received
14 February 2021
Accepted
17 June 2021
Published
15 July 2021
Volume
8 - 2021
Edited by
Susana P. Gaudêncio, New University of Lisbon, Portugal
Reviewed by
Roland Kersten, University of Michigan, United States; Blanca Hernandez-Ledesma, Consejo Superior de Investigaciones Científicas (CSIC), Spain; Adrianna Ianora, Anton Dohrn Zoological Station, Italy
Updates
Copyright
© 2021 Macedo, Cunha, Carneiro, Costa, Alencar, Cardoso, Franco and Dias.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Octávio Luiz Franco, ocfranco@gmail.com
This article was submitted to Marine Biotechnology, a section of the journal Frontiers in Marine Science
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.