- 1 Laboratory of Neuro Imaging, Department of Neurology, University of California Los Angeles, Los Angeles, CA, USA
- 2 Brain Injury Research Center, Departments of Neurology and Neurosurgery, University of California Los Angeles, Los Angeles, CA, USA
- 3 Department of Radiology, David Geffen School of Medicine, University of California Los Angeles, Los Angeles, CA, USA
- 4 Scientific Computing and Imaging Institute, University of Utah, Salt Lake City, UT, USA
- 5 Surgery Planning Laboratory, Department of Radiology, Brigham and Women’s Hospital, Harvard Medical School, Boston, MA, USA
Available approaches to the investigation of traumatic brain injury (TBI) are frequently hampered, to some extent, by the unsatisfactory abilities of existing methodologies to efficiently define and represent affected structural connectivity and functional mechanisms underlying TBI-related pathology. In this paper, we describe a patient-tailored framework which allows mapping and characterization of TBI-related structural damage to the brain via multimodal neuroimaging and personalized connectomics. Specifically, we introduce a graphically driven approach for the assessment of trauma-related atrophy of white matter connections between cortical structures, with relevance to the quantification of TBI chronic case evolution. This approach allows one to inform the formulation of graphical neurophysiological and neuropsychological TBI profiles based on the particular structural deficits of the affected patient. In addition, it allows one to relate the findings supplied by our workflow to the existing body of research that focuses on the functional roles of the cortical structures being targeted. A graphical means for representing patient TBI status is relevant to the emerging field of personalized medicine and to the investigation of neural atrophy.
Introduction
Much promising research motivated by the enormous burden of traumatic brain injury (TBI) upon healthcare worldwide has made progress in recent years, yet the formulation of novel and useful methodologies for the characterization of TBI sequelae remains highly desirable and widely needed (Chen and D’Esposito, 2010). Although TBI-related diffuse axonal injury (DAI) and cortical damage can contribute significantly to both neurobehavioral outcome and functional impairment, the effects of these phenomena can be difficult to assess well and to associate with affected function in a satisfactory manner (Sanz-Arigita et al., 2010). Partly for this reason, the current state of the art in TBI connectomics (the mapping of neural connections in the nervous system) would benefit from further improvement in the ability to represent, describe, and isolate potential impairment. This could assist the clinician to (1) reduce the rates of atrophy, (2) stimulate brain recovery and repair where possible, and (3) lead to neuropsychological outcome improvement.
The advent of methods that can offer original concise but informative insights into the effect of injury upon white matter (WM) atrophy and upon affected brain function would be welcome for several reasons. Firstly, more progress is needed to understand the impact of brain injuries upon the neuroanatomical, behavioral, and cognitive profiles of patients (Chesnut, 1998). Secondly, the relationships between brain dysfunction and neuropsychological impairment can be challenging to establish due to insufficient understanding of WM connectivity function (Bates et al., 2003). Consequently, a wide range of efforts aimed at the amelioration of patient dysfunction would greatly benefit from the introduction of new approaches for mapping WM atrophy in TBI.
The recent interest in the in vivo mapping of human connectomics is evident in the launching of the Human Connectome Project1 (HCP) by the National Institutes of Mental Health and Aging in 2010. Since then, considerable effort has been dedicated to the development of techniques that graphically represent the connectivity of neural structures (Bullmore and Sporns, 2009). Nevertheless, few methodologies are available that allow clinicians to explore patient-tailored atrophy profiles at the connectome-level. In this context, the aim of this paper is to introduce a patient-tailored approach to the graphical representation of WM change over time. The potential use of the method is illustrated using data from three TBI cases, which offers the ability to produce detailed non-invasive characterization of TBI-related GM/WM abnormality and atrophy in vivo. This allows the visualization of brain connections affected by pathology as obviated by MRI and to relate patient injury profiles to the existing body of scientific and clinical knowledge on affected cortical structure function. The approach includes the ability to quantify the neural atrophy of WM tracts for personalized connectomics. Consequently, it offers the ability to integrate such knowledge with other clinical case information so as to inform clinicians on specific neuroplasticity and neurodegeneration patterns that occur in the TBI brain. Using this approach, we hope to contribute to the clinical care of TBI patients, maximizing the utility of modern neuroimaging technologies, and having possible implications for the improvement of outcome in this important clinical population.
Methods
Patients
Anonymized neuroimaging data were acquired at the UCLA Brain Injury Research Center from three adult TBI patients using a protocol designed and carried out according to the Declaration of Helsinki. The study was approved by the UCLA Institutional Review Board and by the patients via signed informed consent. Three sample trauma cases are discussed in this study. The first of these is a 25-year-old male with TBI caused by blunt injury, with a Glasgow coma score (GCS) of 14 (mild) upon hospitalization, and a Glasgow outcome score (GOS) of 3 (lower severe disability) upon leaving the neurointensive care unit. Figure 1A shows sample MR images from this patient displayed in radiological convention, demonstrating focal temporo-parietal hemorrhage in the left hemisphere (LH) at the acute time point (red arrows), especially in the acute gradient recalled echo (GRE) T2 image. A decrease in the volume of the left ventricle is apparent, whereas fluid attenuated inversion recovery (FLAIR) hyperintensities suggest the presence of edema surrounding this hemorrhage. Extra-cerebral inflammation and bleeding are also apparent. Non-hemorrhagic edema is indicated in the FLAIR image by the green arrow. The GRE T2 image shows frontal LH bleeding (blue arrows). At chronic follow-up, bilateral ventricular and cerebral symmetry are improved. Figure 2A illustrates edema (green) and hemorrhage (red) superposed upon a transparent brain model. Red, green, and blue arrows identify the locations of three different insults. Figure 2A also illustrates the bilateral presence of antero-medial edema in the frontal lobes as well as bleeding in the left temporal lobe. Edema in the right temporal and parietal lobes may be associated with contre-coup injuries.
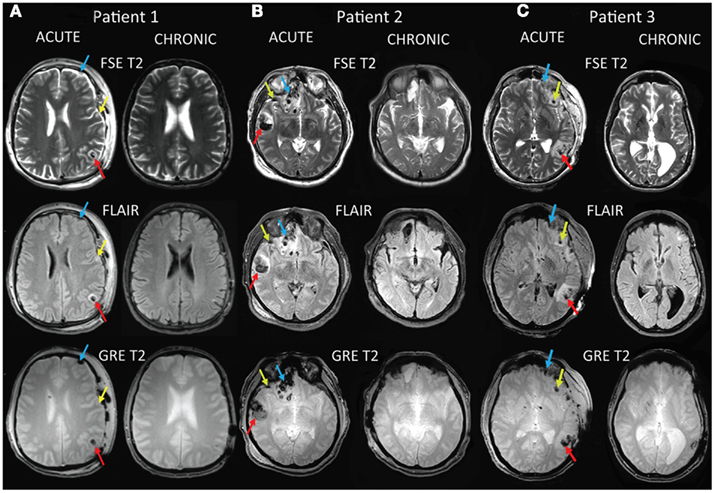
Figure 1. Sample MR images for three TBI cases labeled as patient 1 (A), patient 2 (B), and patient 3 (C). Images are displayed in radiological convention. The sequence types shown include T2, GRE T2, and FLAIR. Red, green, and blue arrows identify the locations of three different insults.
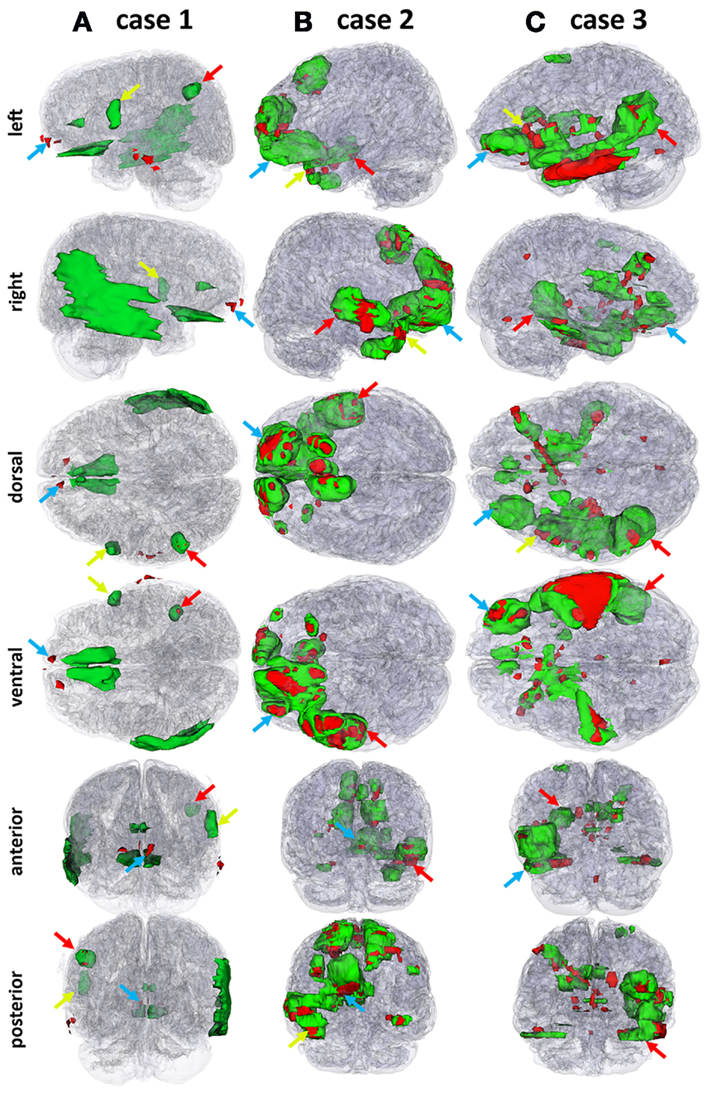
Figure 2. Three-dimensional models of automatically segmented TBI pathology superposed on transparent models of the brain for each patient. Edema and hemorrhage are shown in green and red, respectively. To guide the eye in localizing three distinct lesions in the 3D models, color-coded (red, green, and blue) arrows are provided to identify the locations of the insults indicated in Figure 1 using corresponding colors. Each column (A, B, and C) corresponds to one of the three cases (1, 2 or 3, respectively).
Patient 2 is a 45-year-old male with TBI caused by a closed head injury, with a GCS of 14 (mild) upon hospitalization, and a GOS of 3 (lower severe disability) upon transfer from the NICU. Whereas the GCS and GOS for this patient are the same as those for patient 1, both MR imaging (Figure 1B) and three-dimensional (3D) models of the pathology (Figure 2B) reveal more extensive GM/WM lesions compared to patient 1. Specifically, this patient exhibits large fronto-temporal hemorrhages and edema, both of which also extend fronto-medially and fronto-dorsally (Figures 1B and 2B). To guide the eye in localizing three distinct lesions in the 3D model, color-coded arrows are provided in both figures in order to easily relate the pathology in Figure 1 to the 3D reconstructions afforded by our automated segmentation framework.
The third patient included in this study (Figures 1C and 2C) is a 62-year-old male admitted to the NICU with TBI caused by a bicycle fall. His GCS score upon hospitalization was 7, while the GOS upon transfer from the NICU was 2. Patient 3 exhibits large fronto-temporal edemas and hemorrhages particularly in the LH, although trauma is also present contralaterally to a significant extent. Some microbleeds are also visible throughout the posterior portion of the brain. Comparison of the MR images and 3D models from the three patients reveals, in progressive order from patient 1 to patient 3, a pattern of increasingly poorer combinations of pathology extent and outcome scores, with patient 1 having the mildest and patient 3 the most severe of these.
Image Acquisition
Three-dimensional scan volumes were acquired at 3 T using a Siemens Trio TIM scanner (Siemens AG, Erlangen, Germany). The MR acquisition protocol included fast spin echo (FSE) T2 [repetition time (TR) = 3330 ms, echo time (TE) = 89 ms, flip angle (FA) = 120°, echo train length (ETL) = 18, slice thickness = 5 mm, phase field of view (FOV) = 100%, sampling = 75%, acquisition type = 2D, matrix dimensions = 512 × 512], GRE T2 (TR = 1500 ms, TE = 7 ms, FA = 20°, ETL = 2, slice thickness = 3 mm, phase FOV = 75%, sampling = 80%, acquisition type = 2D, matrix dimensions = 384 × 512), FLAIR (TR = 8000 ms, TE = 70 ms, TI = 2375, FA = 130°, ETL = 16, slice thickness = 3 mm, phase FOV = 75%, sampling = 75%, acquisition type = 2D, matrix dimensions = 384 × 512), and 32-direction diffusion tensor imaging (DTI; Basser and Jones, 2002); TR = 8000 ms, TE = 95 ms, FA = 90°, ETL = 1, slice thickness = 3 mm, phase FOV = 100%, sampling = 100%, acquisition type = 2D, matrix dimensions = 128 × 128). Sequence parameters are also documented in a previous publication (Irimia et al., 2011a). Scanning sessions were held both 1 day (acute baseline) as well as 7 months (chronic follow-up) after TBI, and the same scanner and sequence parameters were used for both acute and chronic time.
Image Processing
Identification of pathology based on the types of sequences available was performed using standard protocols documented elsewhere (Irimia et al., 2011a). Briefly, non-hemorrhagic lesions were coded as hyperintensities on FLAIR, and segmentation quality was confirmed using GRE imaging as well as TSE T2-weighted volumes. Non-hemorrhagic shearing lesions were defined as hyperintense lesions visible on T2-weighted or FLAIR images (see Figure 1). Following the previous guidelines of Tong et al. (2003), hemorrhagic lesions were defined as hypointense foci that were not compatible with vascular, bone, or artifactual structures on conventional GRE images (see also Irimia et al., 2011a). WM, GM, and pathology were classified using 3D Slicer2, FreeSurfer (FS, freely available from surfer.nmr.mgh.harvard.edu; Dale et al., 1999; Fischl et al., 1999a,b, 2004b), and atlas based classification (ABC; Pohl et al., 2007; Zoellei et al., 2007; Prastawa and Gerig, 2008), the latter having been widely used to detect pathology (Prastawa et al., 2009). In FS, WM was segmented and the estimated GM–WM border was tessellated to provide a topographically correct areal representation. In ABC, multimodal MR volume registration (Maes et al., 1997) was followed by tissue classification (Van Leemput et al., 1999), by lesion segmentation using outlier detection (Prastawa et al., 2004), and by physical model estimation (Prastawa et al., 2003; Prastawa and Gerig, 2008). Co-registration of MR volumes, intensity normalization within and between scans as well as bias field correction were applied. Segmentation results were favorably compared to manual segmentations by a human expert, as described in Wang et al. (2012). Specifically, to quantify the similarity between segmentations, the Dice coefficient – which measures the volumetric overlap of two binary segmentations and lies in (0, 1) – was used. Segmentation error correction was performed using 3D Slicer software under established guidelines (Filippi et al., 1998, see Figure 2 and Results), and Dice coefficients were found to exceed 0.95 for all segmentations, indicating excellent similarity between automatic and user-guided segmentations. Sample MR datasets and segmentations obtained using our protocol are freely available from Irimia et al. (2011a). Diffusion tensors were computed from DWI images and rotationally re-oriented at each voxel. Tensor-valued images were linearly realigned based on trilinear interpolation of log-transformed tensors as described in Chiang et al. (2011) and resampled to isotropic voxel resolution (1.7 mm × 1.7 mm × 1.7 mm). To reconstruct fiber tracts, diffusion gradient data were processed in native space using TrackVis3. This software is freely available and its performance compared to other tractography utilities has been investigated elsewhere (see Wedeen et al., 2008 and references therein). Data processing workflows were created using the LONI Pipeline4. Cortical and subcortical volumetrics were obtained by assigning neuroanatomical labels to voxels based on probabilistic information estimated from a manually labeled training set (Fischl et al., 2002). The method uses (1) the previous probability of a tissue class occurring at a specific atlas location; (2) the likelihood of the image given that tissue class; and (3) the probability of the local spatial configuration of labels given that tissue class. This technique is comparable in accuracy with manual labeling (Fischl et al., 2004a) and was used to calculate WM, GM, amygdalar, and hippocampal volumes. The cortex was parcelated (Fischl et al., 2004a) to compute cortical volume, thickness, surface area, and curvature everywhere on the surface, including gyral and sulcal structures (see Figure 3 and Results). A total of 74 cortical structures and 7 subcortical structures were delineated per hemisphere in addition to the cerebellum. The brain stem was also included. Seven cortical lobes were outlined, with 21 frontal lobe (Fro) parcelations, 8 insular (Ins), 8 limbic (Lim), 11 temporal (Tem), 11 parietal (Par), and 15 occipital (Occ). Five representative statistics were computed for each parcelation: GM volume, surface area, mean cortical thickness, mean curvature, and efferent WM fiber density per unit GM volume (“connectivity density”).
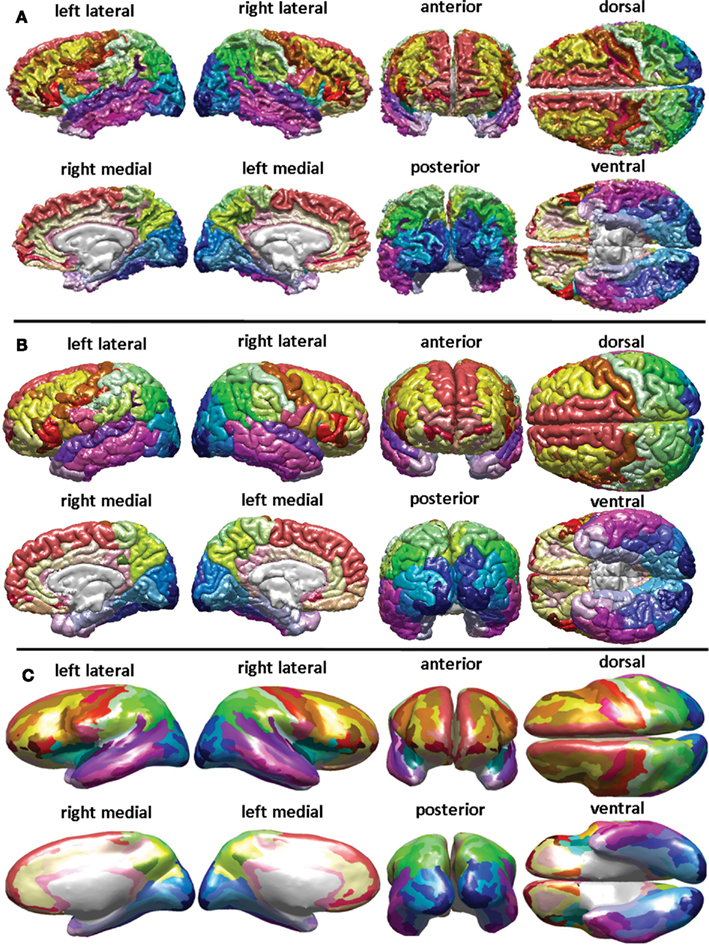
Figure 3. FreeSurfer parcelation of a sample brain. Displayed are the surface corresponding to the WM–GM interface (A), the pial surface (B), and the inflated surface (C), with each parcelated anatomical region displayed in a different color according to the color labeling methodology described in the Section “Methods” (see also Appendix).
Connectogram Design
Connectivity was represented circularly using a framework based on Circos software (Krzywinski et al., 2009). Parcelated regions were displayed as a circle of radially aligned elements (a “connectogram”) representing the left and right hemispheres positioned symmetrically on the corresponding side of the vertical axis (see Figures 4–6 and Results). Parcelated regions were assigned unique RGB colors as shown in Figure 3, and all RGB codes are listed in Table A1 in Appendix. Arrangement of parcelations within each lobe of the connectogram (Figure 3) was performed in the order of their locations along the antero-posterior axis of the cortical surface associated with the published FS normal population atlas (Destrieux et al., 2010). Cortical lobes were assigned unique color schemes: black to red to yellow (Fro), charlotte to turquoise to forest green (Ins), primrose to lavender rose (Lim), etc. (Figure 3). Subcortical structures were colored light gray to black. An unambiguous abbreviation scheme was created to label each parcelation, as summarized in Tables A1 and A2 in Appendix. Within the outermost circle which represents cortical parcelations, five circular heat maps were created to encode one of the five structural measures associated with the corresponding parcelation. Proceeding toward the center of the circle, these measures are total GM volume, total area of the surface associated with parcelation, mean cortical thickness, mean curvature, and connectivity per unit volume. This latter measure was calculated as the density of fibers with endings within that parcelation divided by the parcelation’s total GM volume. The value of each structural measure was encoded as a color using a color scheme mapping that ranged from the minimum to the maximum of the data set. Specifically, the cortical thickness t with values in the interval (tmin, tmax) was normalized as t1 = (t − tmin)/(tmax − tmin). The value of t1 was associated with a unique color; for example, nuances at the extremities of the color map correspond to tmin and tmax, as required. For the brain stem, cerebellum, and subcortical structures, values for area, thickness, and curvature were unavailable from FS and their appropriate heat map entries were drawn in white. The reader is referred to Figure 4 and to the Section “Results” for a sample connectogram incorporating the features above. Software for generating connectograms is being made freely available via the LONI Pipeline (see text footnote 4; see Irimia et al., 2012 for further details).
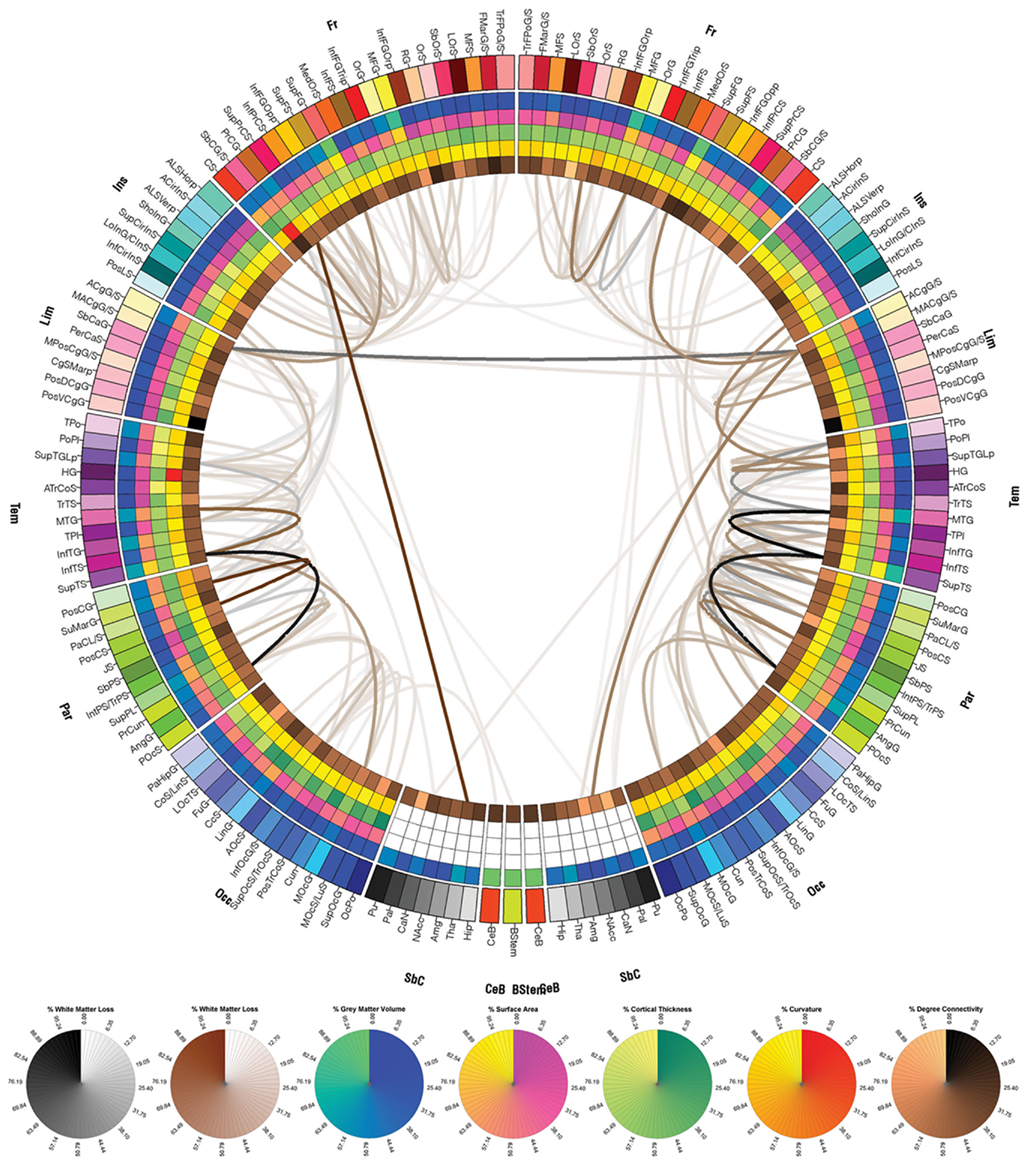
Figure 4. The connectogram (reconstructed connectomic profile) of patient 1. Each cortical structure is assigned a unique RGB color, as explained in the Section “Methods,” and the colors assigned to the parcelated regions on the outermost ring of the connectogram are identical to those of the corresponding cortical structures in Figure 3. Brown links between connectogram nodes indicate WM fibers between a region affected by pathology and another that was not affected, based on the automatic segmentations of pathology shown in Figure 2. Similarly, gray links indicate WM fibers between two regions that were both affected. Circular color maps are displayed below the connectogram for links (first two maps, white to brown and white to black) and for the metrics encoded on the five innermost rings of the connectogram. The range for each of these metrics is from their minimum to their maximum assumed value.
Connectivity Representation
To calculate inter-region connectivity, the location of each fiber tract extremity was identified and each fiber extremity was associated with the appropriate parcelation. This was performed for all fibers that both originated and ended within any two distinct parcelations. Appropriate connectivity matrix entries were updated as necessary to reflect fiber count increments (Hagmann et al., 2008, 2010). To validate the accuracy of the connectivity matrix algorithm, its results were reproduced by individually counting in 3D Slicer all fibers that had exactly one extremity within the volumetric extent of some given parcelation, and by repeating this procedure over several test subjects. Each entry in the connectivity matrix was normalized by the total number of fibers in the entire brain based on the MR volume acquired at each time point and the resulting normalized fiber counts were used for ulterior calculations. Whenever fibers existed between a cortical region affected by pathology and another that was unaffected, the color of the corresponding link was brown. Similarly, links between parcelations that were both affected by pathology were drawn in gray. Thus, one can distinguish between connections that involve only one affected region (brown links) or two affected regions (gray links). Above and throughout, regions affected by pathology are understood to represent regions that contain pathological tissue of any type as obviated by MRI. For connectograms displaying WM atrophy that occurred between the acute and chronic time points, links associated with atrophy (decrease in fiber density) were drawn in blue. The transparency level was directly proportional to the percentage change in fiber density (see also section below). In the case of connectograms displaying atrophy measures (Figures 7–9), the colors on the five inner rings of each connectogram indicate percentage differences in each measure m (where m can be cortical area, cortical thickness, curvature, or GM volume), computed between the acute and chronic time points (t1 and t2, respectively). Percentage differences were computed using the formula [m(t2) − m(t1)] × 100/m(t1), i.e., as percentage changes with respect to the acute baseline time point.
Calculation and Representation of Neural Atrophy
For each pair of cortical regions, the change Δ in the fiber density D between t1 and t2 was computed as a percentage of the fiber count at acute baseline based on the multimodal imaging data acquired at the two time points. (1) In the first step of the analysis, the histogram (empirical distribution function) of fiber counts was computed for all connections, and the fibers belonging to the top 5% of the histogram distribution were selected for further analysis. Although the threshold selected is admittedly somewhat arbitrary, its value can be adjusted by the user to either more conservative or more liberal values for the purpose of data exploration or for reasons related to statistical significance criteria. Here, a value of 5% is selected for illustration. It should be noted here that the procedure described above does not involve the statistical problem of multiple comparisons because no statistical tests are being applied. Instead, tracts are singled out for display based on whether they exhibit atrophy above a user-specified threshold. (2) In the second step, the percentage change in the count of the fibers selected in the previous step was computed for each connection, using the formula Δ = [D(t2) − D(t1)]/D(t1). (3) In the third step, those fibers that had computed changes in fiber density with absolute values greater than 20% (|Δ| > 20%, i.e., the top four-fifths of the distribution of percentage changes) were displayed on a separate connectogram (see Results and Figures 7–9). Because no increases in fiber densities were found between t1 and t2 in any subject, Δ will henceforth be assumed to be negative. Link transparency in the corresponding connectograms (Figures 7–9) encode the quantity [|Δ| − min{|Δ|}]/[max{|Δ|} − min{|Δ|}], i.e., the range of transparency values corresponds to the range [min{|Δ}}, max{|Δ|}], with larger changes (more negative values of Δ) being encoded by more opaque hues of blue (see caption to Figure 7). The quantity [|Δ| − min{|Δ|}]/[max{|Δ|} − min{|Δ|}] is simply the percentage change computed in step (2), scaled such that (A) the lowest color opacity corresponds to the smallest absolute value of the percentage change that is greater than the threshold of 20%, and (B) the highest opacity corresponds to the maximum absolute value of the change in fiber density. This three-step methodology for the calculation of significant atrophy was adopted in order to avoid several pitfalls associated with the process of comparing WM fiber densities as extracted from DTI volumes. Firstly, step (1) above ensures that only fiber bundles with the greatest density (top 5% of the fiber density distribution) are selected for further processing. This eliminates the potential problem of computing WM atrophy for connections with very low fiber densities whose presence in the connectivity matrix may in fact be artifactual. Specifically, the presence of the latter type of links can be due to the unavoidable effects of limited DTI spatial resolution upon tractography algorithms, which can lead to computational inaccuracies and to the generation of artifactual tracts with very low densities. To ensure that atrophy calculation results would be trustworthy, it was therefore considered appropriate to take into account only the largest extracted fibers (top 5%) for further processing. Another precautionary measure was that, in step (3) above, only fibers with densities in the top 5% of the distribution which also had absolute values of the change in fiber densities greater than 20% from t1 to t2 were plotted in Figures 7–9. This restriction was imposed because small percentage changes in fiber densities may not in fact be meaningful, which is the case due to methodological limitations similar to those described above for step (1). In conclusion, our combination of conservative restrictions upon the selection of atrophied fibers allows one to confidently identify fibers undergoing a large amount of atrophy. This is achieved while also taking into account the limitations of DTI as a neuroimaging technique and of tractography as a computational framework.
Connectomic Profile Interpretation
The functional roles of the cortical regions connected by WM fibers affected by either TBI or by subsequent atrophy were identified from available literature on Google Scholar5 using queries whose search parameters had been automatically generated within our analysis environment. Terms included in each search included the name of the cortical structure being investigated as well as topic-related keywords such as “rehabilitation,” “treatment,” “recovery,” and “regeneration,” resulting in URL’s such as http://scholar.google.com/scholar? q = orbital + sulcus + rehabilitation, where the search terms are included after the “=” sign and separated by “+” signs. Similarly, the chronic atrophy profile of each subject was used to identify WM connections that had suffered from atrophy between the acute and chronic time points that was appreciable compared to that in the rest of the brain. Connectograms of WM atrophy may be of interest to clinicians and by other medical professionals to gain in sight on the effect of TBI upon a patient’s clinical picture as well as to examine atrophy trajectories.
Results
Connectomic Profiles of Pathology
Exploration of the connectomic profile of each patient requires reference to Figure 3, which shows the FS parcelation of a sample brain. Displayed there are the surface corresponding to the WM–GM interface (Figure 3A), the pial surface (Figure 3B), and the inflated surface (Figure 3C), with each parcelated anatomical region displayed in a different color according to the color labeling methodology described previously. Figures 4–6 display the connectograms (i.e., there constructed connectomic profiles) of Patients 1–3, respectively. Each cortical structure is assigned a unique RGB color, as explained in Section “Methods,” and the colors assigned to the parcelated regions on the outermost rings of the connectograms in Figures 4–6 are identical to those of the corresponding cortical structures in Figure 3 (see also Methods). Brown links between connectogram nodes in Figure 4 indicate WM fibers between a region affected by pathology and another that was not affected, based on the automatic segmentations of pathology shown in Figure 2. Similarly, gray links indicate WM fibers between two regions that were both affected.
As shown in Figure 4 for patient 1, cortical connections linking the temporal and parietal lobes bilaterally to the rest of the brain appear to have been significantly affected by pathology, as expected. Structures directly affected in both hemispheres include, among other structures, the anterior cingulate gyrus and sulcus and angular gyrus. Structures directly affected only in the LH include the inferior occipital gyrus and sulcus and other structures, while structures directly affected only in the RH include the inferior temporal gyrus. As shown, although the lesion load of Patient 1 appears to be smaller than that of the other two patients, the number of cortical pathways linking affected regions to other parts of the brain is fairly large. One advantage of our visualization method is that it allows one to identify such connections and to explore them systematically.
Figure 5 shows the cortical connections associated with pathology in Patient 2. As expected based on the information provided in Figures 1B and 2B, the right fronto-temporal profile of the lesions in this patient is confirmed by the large number of cortical connections that are affected in these areas. For Patient 3, the numerous cortical connections that appear to have been affected involve all lobes of both hemispheres, which is expectable given the severe lesion load of this patient.
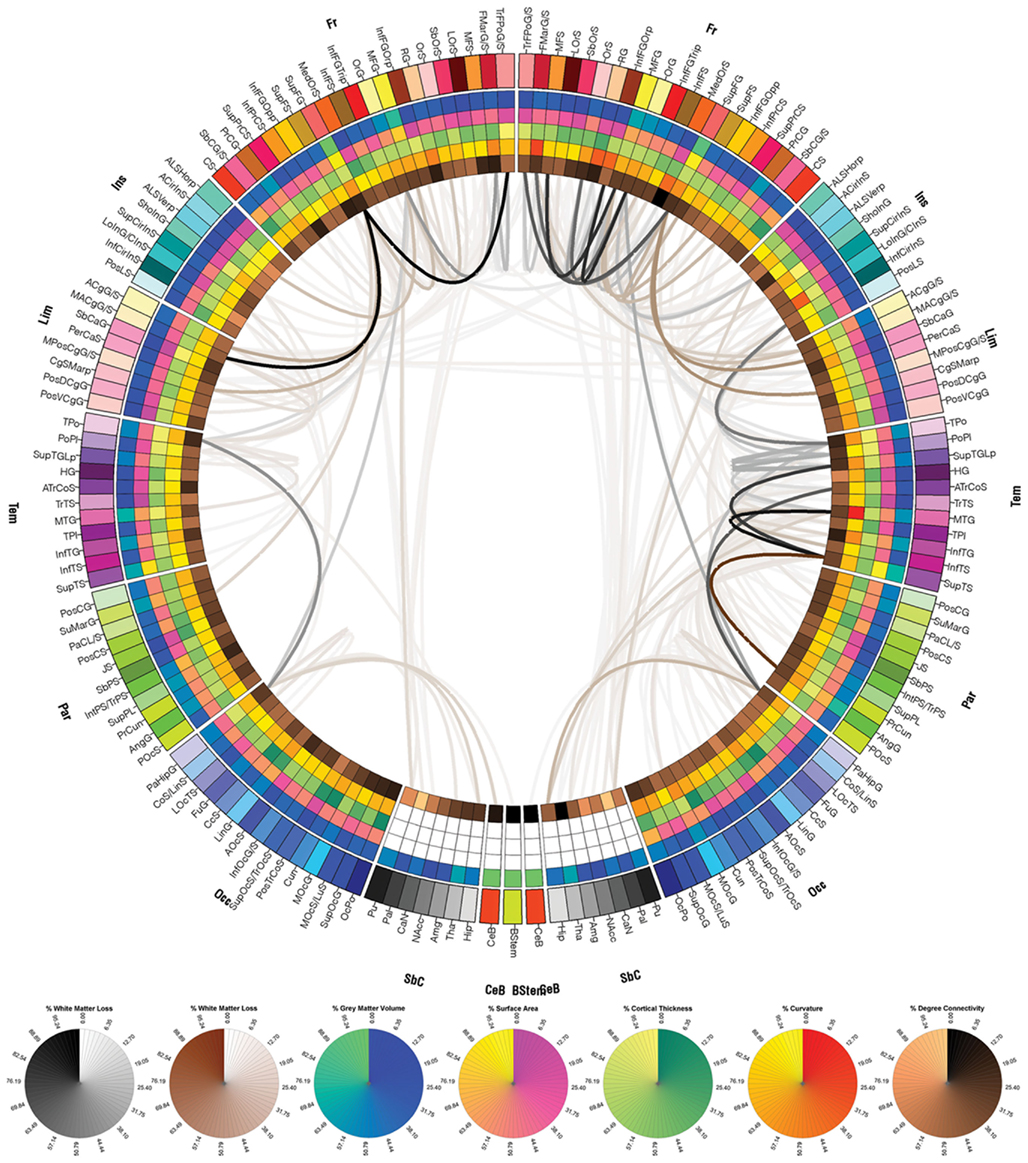
Figure 5. As in Figure 4, for patient 2.
It is worth noting that case severity in these three patients (as indicated by lesion extent and by GCS/GOS) appears to be confirmed by how widespread pathology-affected connections are in each patient’s connectogram. Consequently, the methodology proposed here may be useful to investigate the relationship between the former three measures and cortical network properties in TBI patients, with possible relevance for further quantification of their pathological state and for correlation/prediction of other outcome variables.
Figures 7–9 display the atrophy profile of Patients 1–3, respectively. In all three subjects, it is apparent that the WM connections which have experienced largest atrophy with respect to the acute baseline are, in most cases, connections that were affected by the primary injury, as revealed by comparison of these figures to the corresponding connectograms in Figures 4–6. In Patient 1, for example, the WM connection between the left central sulcus and the left postcentral gyrus is a connection with a large fiber density at acute baseline (Figure 4) which is observed to experience severe atrophy over time (Figure 7). Similar statements can be made for connections between (1) the right superior frontal gyrus and right anterior cingulated gyrus and sulcus, (2) the right middle temporal gyrus and right superior temporal sulcus, (3) the right superior temporal sulcus and the right angular gyrus, as well for a large number of other connections. Inspection and comparison of Figures 4–9 indicate that observations similar to those above can be made for a large number of connections in both patients 2 and 3.
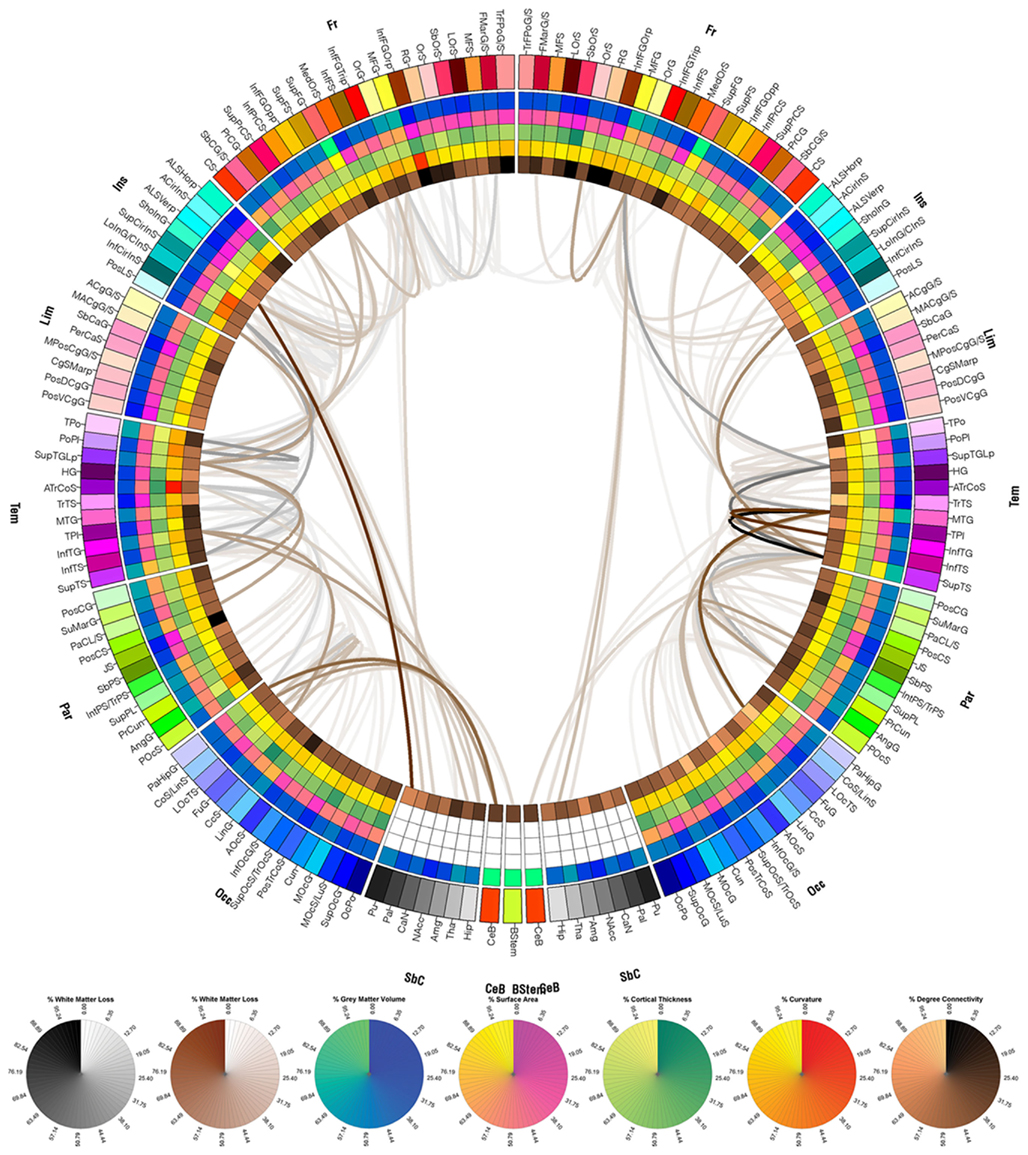
Figure 6. As in Figure 4, for patient 3.
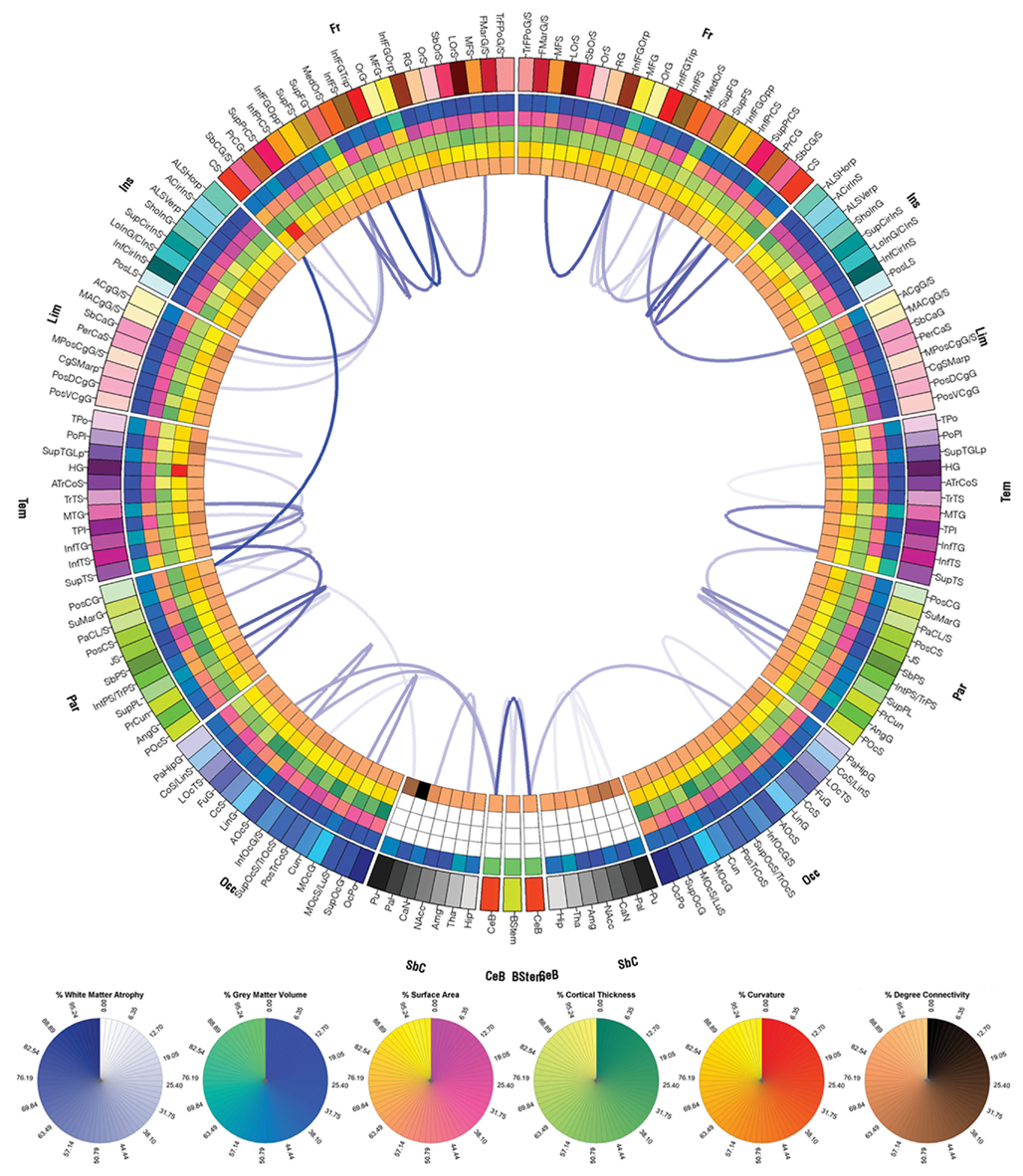
Figure 7. Connectogram of the atrophy profile for patient 1. Links displayed indicate connections that suffered large atrophy from the acute baseline to the chronic follow-up time point. Link transparency encodes the percentage change Δ in fiber density (see Methods), in the range [min{|Δ|}, max{|Δ|}], with larger changes (more negative values of Δ) being encoded by more opaque hues of blue. The lowest color opacity corresponds to the smallest absolute value of the percentage change that is greater than the chosen threshold of 20%, and the highest opacity corresponds to the maximum absolute value of the change in fiber density.
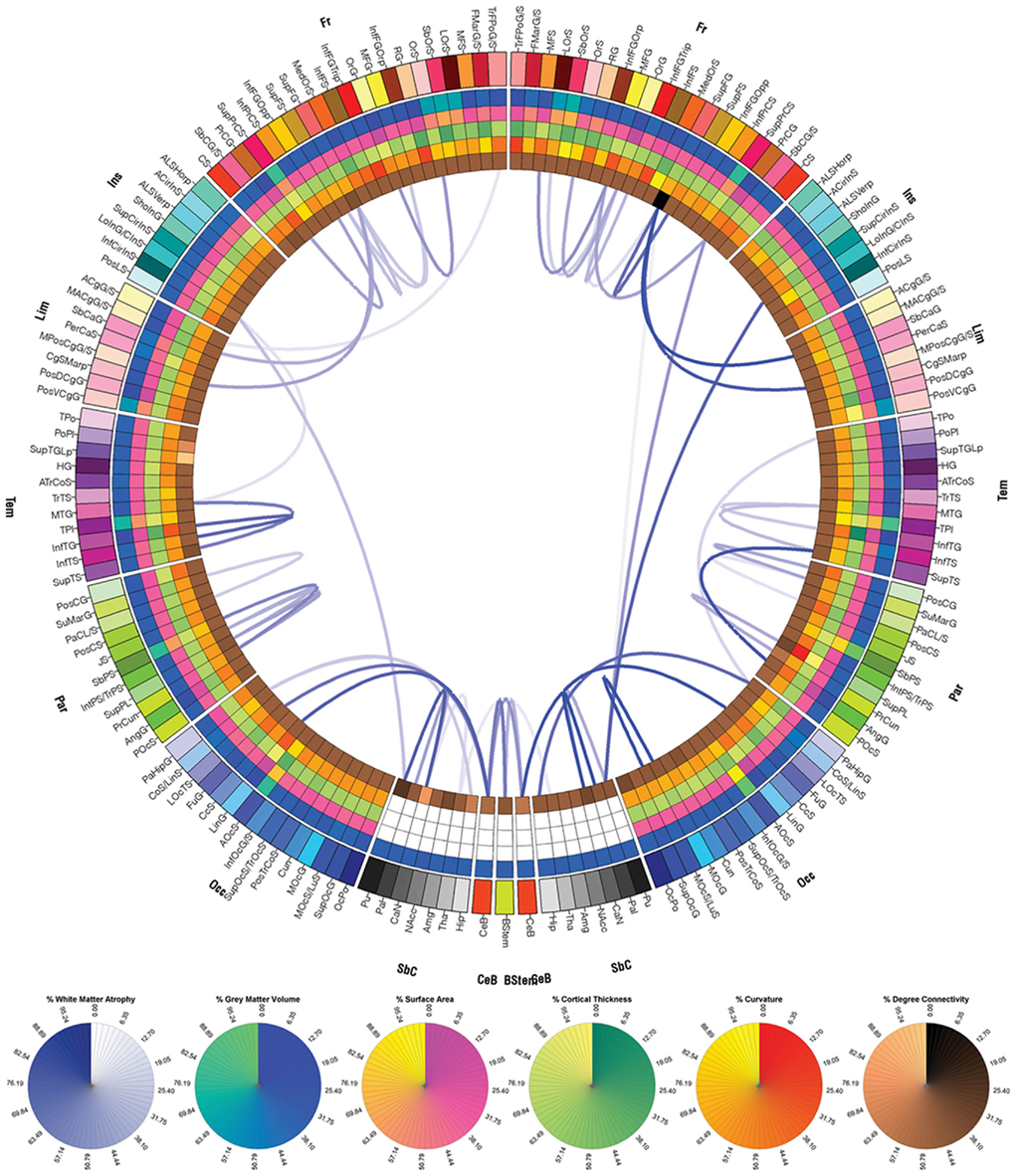
Figure 8. As in Figure 7, for patient 2.
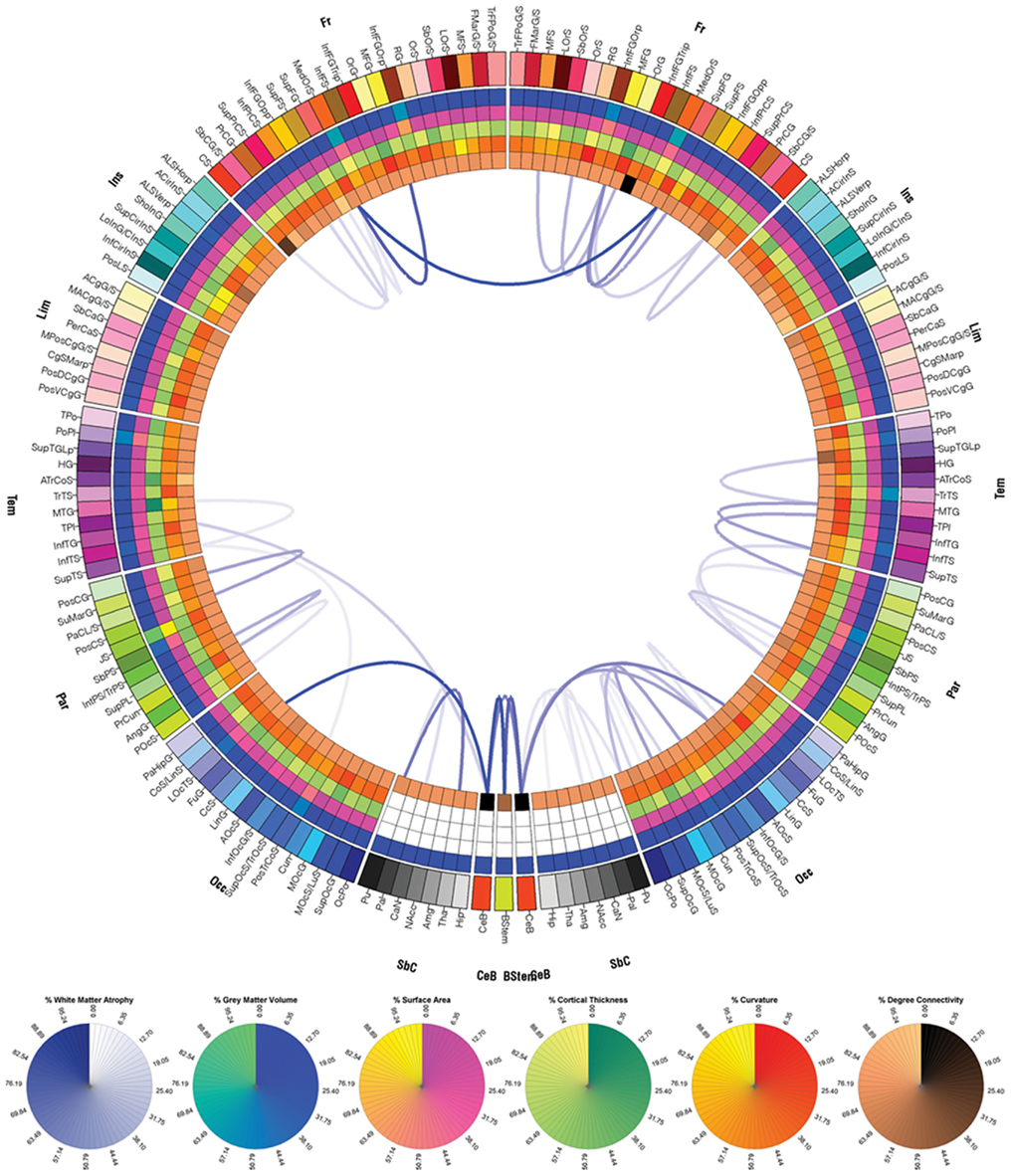
Figure 9. As in Figure 7, for patient 3.
Finally, it is useful to note a group of connections which do not appear to have been affected by primary injuries in any of the three case subjects, yet which experience dramatic atrophy over time. These connections involve primarily cortical structures, on the one hand, and subcortical structures, on the other hand. In particular, inter-cerebellar connections consistently appear to be the object of notable atrophy in all three subjects. This finding is consistent with a number of existing studies (Park et al., 2006; Spanos et al., 2007; Caeyenberghs et al., 2011) which have found large cerebellar atrophy in chronic TBI despite the lack of involvement of the cerebellum itself in the primary acute injury. Although these studies indicate that the causes and dynamics of such atrophy patterns are likely insufficiently understood, the ability of our analysis framework to replicate these findings in individual cases of TBI points to the insight that might be gained from its use. It also indicates the possibility that our method could be used in uncovering useful information on how TBI affects the human brain.
Relationship between TBI Lesion Profiles and Neurophysiological Function
Table 1 lists a selection of cortical regions affected in at least one of the three patients, in addition to (1) the functional description of those regions as identified by various studies from the current literature and to (2) a list of clinical studies with possible relevance to improving the level of insight and detail associated with each TBI lesion profile. For example, the anterior cingulate gyrus, which was affected by TBI in patients 1 and 3, has been involved in error detection (Bush et al., 2000), task anticipation, attention, motivation, and modulation of emotional responses (Luu and Pederson, 2004). Consequently, the lesion profiles suggest that the cognitive–motor tasks outlined in the meta-analyses of Picard and Strick (1996) and Bush et al. (1999) can be informative due to the abilities of these tasks to selectively recruit cortical activity in this brain region, and to increase it in proportion to task difficulty (Bush et al., 2000). The studies associated with this cortical structure in Table 1 describe tasks which include two types of tasks. The first of these are cognitively demanding exercises that involve stimulus–response selection in the face of competing streams of information, including divided-attention tasks, verbal- and motor-response selection tasks. The second type are affect-related tasks involving emotional processing and symptom provocation tasks.
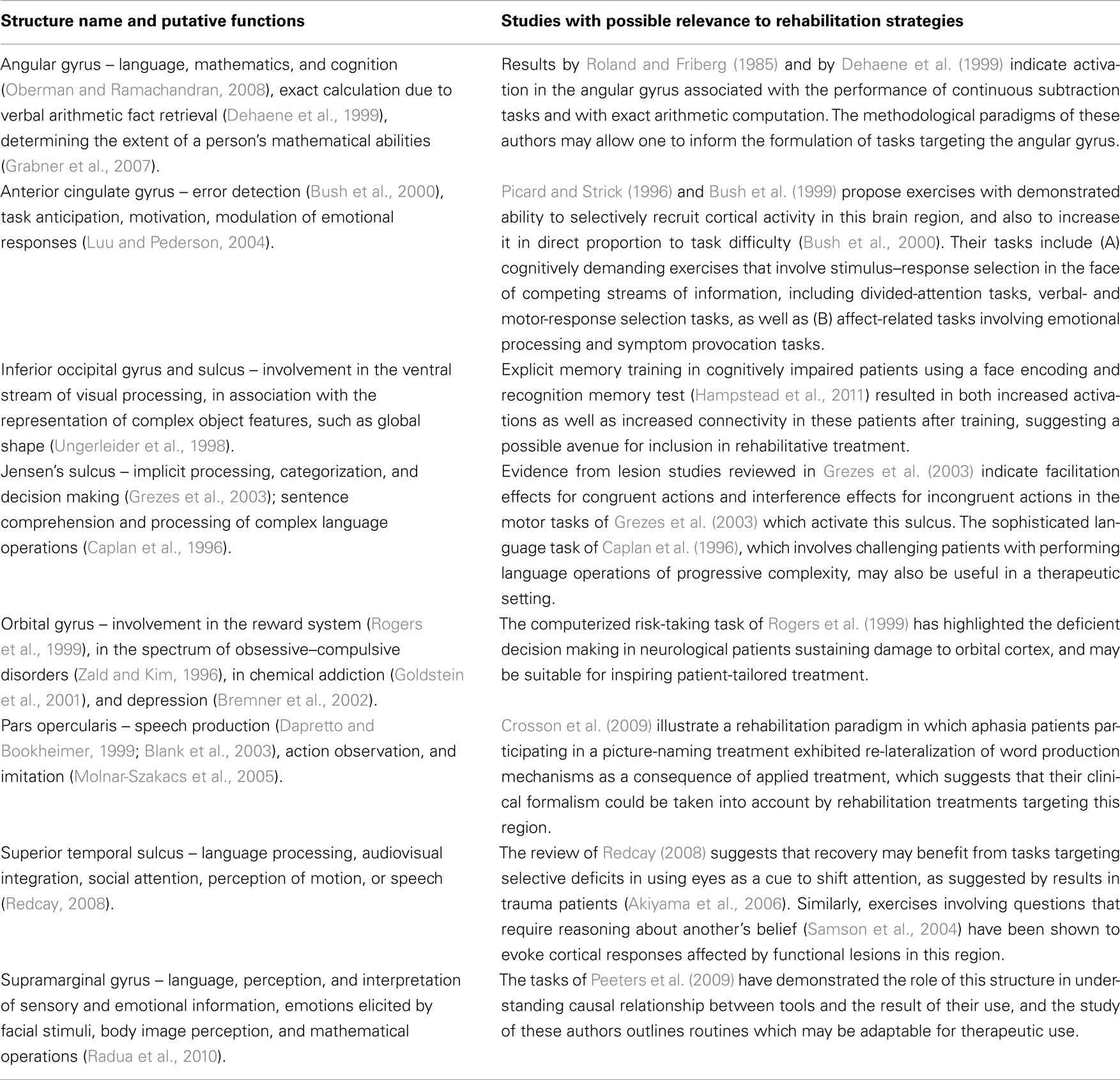
Table 1. Sample selection of cortical regions affected in at least one of the three TBI patients. The first column contains the functional description of each region, while the second column indicates paradigms with possible relevance to rehabilitation strategies.
One cortical region affected in patients 1 and 3 is the superior temporal sulcus, where activations are associated with language processing, audiovisual integration, social attention, and motion, or speech perception (Redcay, 2008). A thorough review (Redcay, 2008) offers useful information concerning the involvement of this structure in tasks targeting selective deficits in using eyes as a cue to shift attention, as also suggested by previous results in trauma patients (Akiyama et al., 2006). Similarly, exercises involving questions that require reasoning about another’s belief (Samson et al., 2004) have been shown to evoke cortical responses affected by functional lesions in this region.
Jensen’s sulcus, also affected in patient 1, has been associated with implicit processing, categorization, and decision making (Grezes et al., 2003), as well as with sentence comprehension and in the processing of syntactically complex language operations (Caplan et al., 1996). Evidence from lesion studies reviewed in Grezes et al. (2003) indicate facilitation effects for congruent actions and interference effects for incongruent actions in the motor tasks of Grezes et al. (2003) that activate this sulcus, while the sophisticated language task of Caplan et al. (1996) involves challenging patients with performing language operations of progressive complexity. Similarly, the supramarginal gyrus has been involved in the perception and interpretation of sensory and emotional information, including emotions elicited by facial stimuli, as well as language, body image perception, and mathematical operations (Radua et al., 2010). The tasks of Peeters et al. (2009) have demonstrated the role of this structure in understanding causal relationship between tools and the result of their use; the study of these authors outlines routines which could be explored to gain further insight into these TBI profiles.
Two other structures affected by TBI in these patients, namely the angular gyrus and of the inferior frontal gyrus, are involved in processes related to language, mathematics, and cognition. Patients with lesions to this structure often have arithmetic deficits and perform poorly in the bouba/kiki task (Oberman and Ramachandran, 2008). These structures are involved in exact calculation due to verbal arithmetic fact retrieval (Dehaene et al., 1999), and in determining the extent of a person’s mathematical abilities (Grabner et al., 2007). Additionally, results by Roland and Friberg (1985) and by Dehaene et al. (1999) indicate activation in the angular gyrus associated with the performance of continuous subtraction tasks and with exact arithmetic computation.
The orbital gyri have been affected in all three case patients discussed in this study. These structures have been repeatedly linked to the reward system (Rogers et al., 1999), and orbitofrontal pathology has been associated with obsessive–compulsive disorder (Zald and Kim, 1996), chemical addiction (Goldstein et al., 2001), and depression (Bremner et al., 2002). The computerized risk-taking task of Rogers et al. (1999) has also highlighted the deficient decision making in neurological patients sustaining damage to orbital cortex. The right pars opercularis of the inferior frontal gyrus has long been known to be involved in speech production (Dapretto and Bookheimer, 1999; Blank et al., 2003), action observation, and imitation (Molnar-Szakacs et al., 2005). The study of Crosson et al. (2009) illustrates a rehabilitation paradigm in which aphasic patients participating in a picture-naming treatment exhibited re-lateralization of word production mechanisms as a consequence of applied treatment. This suggests that the clinical formalism of Crosson et al. may also be suitable for rehabilitation in patients with damage to this area. Finally, the inferior occipital gyrus and sulcus, as well as the inferior temporal gyrus are known to belong to the ventral stream of visual processing (Ungerleider et al., 1998). These regions have been associated with the representation of complex object features, such as global shape. Explicit memory training in cognitively impaired patients using a face encoding and recognition memory test (Hampstead et al., 2011) resulted in both increased activations as well as increased connectivity in these patients after training, suggesting a treatment option for lesions affecting these areas.
Discussion
Rehabilitation of TBI is an exceedingly important public health goal not only because neurotrauma-related activity limitations can have significant impact upon life roles, but also because it affects interpersonal communication, as well as social participation in personal activities of daily living. In this context, our results and methodology hold implications for the systematic mapping of human neural impairment caused by this condition. Firstly, this work is of potential clinical relevance to the study of neural atrophy changes. Aside from identifying and describing connectomic patient profiles, the presented method can be used to generate suggestions for informing and guiding clinical interventions designed to ameliorate recuperation. Rapidly visualizing the longitudinal evolution of individual TBI cases using our diagrammatic tools can reveal how deficit patterns are influenced by lesion site, by relative sparing and redundancy within the distributed cortical system under scrutiny, as well as by the neural plastic changes that can occur with recovery.
Because of the high level of neuroanatomic information presented regarding the structural atrophy and reorganization of the brain in these depictions, they can be exploited to quickly delineate the function of specific WM fibers or cortical (sub-)regions. Thus, in addition to its relevance to the clinical field, our approach has potential applications to the formulation, validation, or information of basic science theories concerning perceptual learning and neural plasticity. It could also complement and extend information already gained from previous animal and human lesion studies. The detailed level of structural impairment description afforded by our technique can be combined with the task selection mechanism of our paradigm to construct more effective patient interventions. Such strategies can be used to explain occupational performance difficulties as well as to shed light upon existing or emerging compensatory rehabilitation techniques.
One possible caveat related to our method is that the presence of hemorrhage or certain other forms of pathology can modify local water diffusivity measures to an extent that may possibly obscure the presence of WM tracts in affected regions, as extracted using DTI tractography. However, it is not foreseeable that this drawback can influence our computed atrophy rates; this is because tissue permeation by blood in the acute TBI phase would be associated with decreased FA compared to the chronic phase. Consequently, acute hemorrhage may result in inappropriate (over-)estimation of WM recovery (due to increases in FA between the two time points), but not atrophy (which is related to decreases in FA). Thus, because our method focuses on the calculation of atrophy rates, the presence of hemorrhage is not expected to influence or results substantially. To state this in another way, the presence of hemorrhage results in decreased FA for some voxels at baseline. This, in turn, results in decreased fiber counts in hemorrhagic regions at acute baseline compared to the chronic stage (when hemorrhage is no longer present). Consequently, it is reasonable to expect the hemorrhage artifact-related percentage change in normalized fiber counts from acute baseline to chronic follow-up to be positive. However, because our technique is concerned with atrophy (i.e., negative percentage changes in normalized fiber counts), the effect of hemorrhage is unlikely to influence our results substantially.
Most methods for connectivity visualization rely on variations of graph theory to position network nodes, represent nodes using shapes, and to modulate edge properties (weight, color, thickness, etc.) according to metrics of connectivity. One popular method for node representation (Bassett et al., 2011; Gerhard et al., 2011; Yan et al., 2011) involves positioning nodes at the 3D locations of brain regions. When viewed in three dimensions, this method allows one to associate network nodes with neuroanatomic landmarks. However, one disadvantage of this representation is that it can have so many nodes and edges that the relationships between them cannot be discerned (Sanz-Arigita et al., 2010). By contrast, the approach proposed here orders nodes along the antero-posterior axis, thereby making all nodes equally visible to the viewer. Although 2D visualization of brain networks is also possible if the cortical surface is spread out as a 2D sheet (Palva et al., 2010; Gerhard et al., 2011), it remains somewhat difficult to associate locations on 2D cortical maps to 3D brain topology.
One difficulty involved in visualizing connectomes is the depiction of nodes using symbols representative of their properties. In Dosenbach et al. (2007), Spoormaker et al. (2010), Chen et al. (2011), Ginestet and Simmons (2011), and elsewhere, colored circles or spheres are used to this end. Although this offers the ability to differentiate groups of edges according to their properties, nodes, and edges can occasionally overlap. This can be partially addressed by changing node transparency (Ginestet and Simmons, 2011), although this may not always be helpful for very dense networks. In our approach, by contrast, all nodes are visible and node properties are represented using concentric color-coded rings. Another difficulty refers to how best to encode network edge properties according to the degree of connectivity between nodes. Edges can be modulated by color (Cao and Slobounov, 2010; van den Heuvel and Pol, 2010) or thickness (He and Evans, 2010; He et al., 2010), and the top-down subdivision of the nervous system can be represented by concentric circles, e.g., brain to cortex to lobes to gyral/sulcal structures, as in (Holten, 2006; Modha and Singh, 2010). In the latter case, edges radiate from the center of a circle (the top structure) to the circumference (the lowest-level structures) and the circular shape of the connectivity graph is appealing and useful for 2D representation of relationships. One challenge that can be addressed by representing connectomes as circular diagrams is that of complexity and dimensionality reduction. In the hierarchical approach (Holten, 2006; Modha and Singh, 2010), this is done via top-down organization of connections; in ours, it can be achieved by displaying connections efferent from one hemisphere, one lobe, or from a single brain structure, thus allowing one to change the complexity of the connectogram.
It is important to emphasize that our workflow allows the user to use connectivity matrices that are generated using segmentation and connectivity matrix calculation methods other than our own. This feature may be preferable to investigators who wish to use some other standard for extracting connectivity information from brain imaging data. Naturally, the extent to which the connectogram functionality being proposed can be useful to investigators depends on how accurate the information contained in the connectivity matrix is. Consequently, although our own method for generating connectivity matrices has been validated, it can be expected that the results of our graphical workflow are only as accurate as the connectivity matrix itself. Unless our own method for generating connectivity matrices is being used, this is a factor beyond our control, which nevertheless leaves undiminished the potential use and novelty of our contribution.
Conclusion
The potential for recovery depends on the patient’s ability for regeneration of structures and on his/her capacity for neural plasticity. Consequently, the ability to provide TBI researchers and rehabilitation professionals with information regarding the longitudinal atrophy/regeneration profile of patients as described using our graphical method is an important advantage of our framework. Firstly, our technique for rapidly visualizing structural WM connectivity evaluation may allow clinicians to compare changes in cortical regions and in connectivity with metrics of patient case evolution. Secondly, this approach can be applied to individual patients as well as used to visualize brain morphometric and connectomics on a population level, as well. Thirdly, the use of our approach to the graphical representation of atrophy profiles may be useful in the context of personalized rehabilitation treatments by informing qualified personnel on recovery prospects and by providing guidance in the process of evaluating the need for long-term care.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We acknowledge the assistance of Marcel Prastawa, Bo Wang, Maria Etcheparre, Steve Pieper, Stephen Aylward, Sonja Pujol, and of the staff at the Laboratory of Neuro Imaging, University of California, Los Angeles. 3D Slicer is a multi-platform, free and open source software package for visualization and medical image computing available from www.slicer.org. This work was supported by the National Alliance for Medical Image Computing (NA-MIC; www.na-mic.org), under NIH Roadmap Initiative grant 2U54EB005149 to Ron Kikinis and sub-award to John D. Van Horn, and by the National Institute of Neurological Disorders and Stroke (NINDS), grant P01NS058489 to Paul M. Vespa.
Footnotes
References
Akiyama, T., Kato, M., Muramatsu, T., Saito, F., Umeda, S., and Kashima, H. (2006). Gaze but not arrows: a dissociative impairment after right superior temporal gyrus damage. Neuropsychologia 44, 1804–1810.
Basser, P. J., and Jones, D. K. (2002). Diffusion-tensor MRI: theory, experimental design and data analysis – a technical review. NMR Biomed. 15, 456–467.
Bassett, D. S., Brown, J. A., Deshpande, V., Carlson, J. M., and Grafton, S. T. (2011). Conserved and variable architecture of human white matter connectivity. Neuroimage 54, 1262–1279.
Bates, E., Wilson, S. M., Saygin, A. P., Dick, F., Sereno, M. I., Knight, R. T., and Dronkers, N. F. (2003). Voxel-based lesion-symptom mapping. Nat. Neurosci. 6, 448–450.
Blank, S. C., Bird, H., Turkheimer, F., and Wise, R. J. (2003). Speech production after stroke: the role of the right pars opercularis. Ann. Neurol. 54, 310–320.
Bremner, J. D., Vythilingam, M., Vermetten, E., Nazeer, A., Adil, J., Khan, S., Staib, L. H., and Charney, D. S. (2002). Reduced volume of orbitofrontal cortex in major depression. Biol. Psychiatry 51, 273–279.
Bullmore, E., and Sporns, O. (2009). Complex brain networks: graph theoretical analysis of structural and functional systems. Nat. Rev. Neurosci. 10, 186–198.
Bush, G., Frazier, J. A., Rauch, S. L., Seidman, L. J., Whalen, P. J., Jenike, M. A., Rosen, B. R., and Biederman, J. (1999). Anterior cingulate cortex dysfunction in attention-deficit/hyperactivity disorder revealed by fMRI and the counting Stroop. Biol. Psychiatry 45, 1542–1552.
Bush, G., Luu, P., and Posner, M. I. (2000). Cognitive and emotional influences in anterior cingulate cortex. Trends Cogn. Sci. (Regul. Ed.) 4, 215–222.
Caeyenberghs, K., Leemans, A., Geurts, M., Linden, C. V., Smits-Engelsman, B. C., Sunaert, S., and Swinnen, S. P. (2011). Correlations between white matter integrity and motor function in traumatic brain injury patients. Neurorehabil. Neural Repair 25, 492–502.
Cao, C., and Slobounov, S. (2010). Alteration of cortical functional connectivity as a result of traumatic brain injury revealed by graph theory, ICA, and sLORETA analyses of EEG signals. IEEE Trans. Neural Syst. Rehabil. Eng. 18, 11–19.
Caplan, D., Hildebrandt, N., and Makris, N. (1996). Location of lesions in stroke patients with deficits in syntactic processing in sentence comprehension. Brain 119 (Pt 3), 933–949.
Chen, A. J., and D’Esposito, M. (2010). Traumatic brain injury: from bench to bedside [corrected] to society. Neuron 66, 11–14.
Chen, H. F., Liao, W., Ding, J. R., Marinazzo, D., Xu, Q. A., Wang, Z. G., Yuan, C. P., Zhang, Z. G., and Lu, G. M. (2011). Small-world directed networks in the human brain: multivariate Granger causality analysis of resting-state fMRI. Neuroimage 54, 2683–2694.
Chesnut, R. M. (1998). Implications of the guidelines for the management of severe head injury for the practicing neurosurgeon. Surg. Neurol. 50, 187–193.
Chiang, M. C., Barysheva, M., Toga, A. W., Medland, S. E., Hansel, N. K., James, M. R., Mcmahon, K. L., De Zubicaray, G. I., Martin, N. G., Wright, M. J., and Thompson, P. M. (2011). BDNF gene effects on brain circuitry replicated in 455 twins. Neuroimage 55, 448–454.
Crosson, B., Moore, A. B., Mcgregor, K. M., Chang, Y. L., Benjamin, M., Gopinath, K., Sherod, M. E., Wierenga, C. E., Peck, K. K., Briggs, R. W., Rothi, L. J. G., and White, K. D. (2009). Regional changes in word-production laterallity after a naming treatment designed to produce a rightward shift in frontal activity. Brain Lang. 111, 73–85.
Dale, A. M., Fischl, B., and Sereno, M. I. (1999). Cortical surface-based analysis – I. Segmentation and surface reconstruction. Neuroimage 9, 179–194.
Dapretto, M., and Bookheimer, S. Y. (1999). Form and content: dissociating syntax and semantics in sentence comprehension. Neuron 24, 427–432.
Dehaene, S., Spelke, E., Pinel, P., Stanescu, R., and Tsivkin, S. (1999). Sources of mathematical thinking: behavioral and brain-imaging evidence. Science 284, 970–974.
Destrieux, C., Fischl, B., Dale, A., and Halgren, E. (2010). Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. Neuroimage 53, 1–15.
Dosenbach, N. U. F., Fair, D. A., Miezin, F. M., Cohen, A. L., Wenger, K. K., Dosenbach, R. A. T., Fox, M. D., Snyder, A. Z., Vincent, J. L., Raichle, M. E., Schlaggar, B. L., and Petersen, S. E. (2007). Distinct brain networks for adaptive and stable task control in humans. Proc. Natl. Acad. Sci. U.S.A. 104, 11073–11078.
Filippi, M., Gawne-Cain, M. L., and Gasterini, C. (1998). Effect of training and different measurement strategies on the reproducibility of brain MRI lesion load measurements in multiple sclerosis. Neurology 50, 238–244.
Fischl, B., Salat, D. H., Busa, E., Albert, M., Dieterich, M., Haselgrove, C., Van Der Kouwe, A., Killiany, R., Kennedy, D., Klaveness, S., Montillo, A., Makris, N., Rosen, B., and Dale, A. M. (2002). Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron 33, 341–355.
Fischl, B., Salat, D. H., Van Der Kouwe, A. J., Makris, N., Segonne, F., Quinn, B. T., and Dale, A. M. (2004a). Sequence-independent segmentation of magnetic resonance images. Neuroimage 23(Suppl. 1), S69–S84.
Fischl, B., Van Der Kouwe, A., Destrieux, C., Halgren, E., Segonne, F., Salat, D. H., Busa, E., Seidman, L. J., Goldstein, J., Kennedy, D., Caviness, V., Makris, N., Rosen, B., and Dale, A. M. (2004b). Automatically parcellating the human cerebral cortex. Cereb. Cortex 14, 11–22.
Fischl, B., Sereno, M. I., and Dale, A. M. (1999a). Cortical surface-based analysis – II: inflation, flattening, and a surface-based coordinate system. Neuroimage 9, 195–207.
Fischl, B., Sereno, M. I., Tootell, R. B. H., and Dale, A. M. (1999b). High-resolution intersubject averaging and a coordinate system for the cortical surface. Hum. Brain Mapp. 8, 272–284.
Gerhard, S., Daducci, A., Lemkaddem, A., Meuli, R., Thiran, J., and Hagmann, P. (2011). The Connectome Viewer Toolkit: an open source framework to manage, analyze, and visualize connectomes. Front. Neuroinform. 5:3. doi: 10.3389/fninf.2011.00003
Ginestet, C. E., and Simmons, A. (2011). Statistical parametric network analysis of functional connectivity dynamics during a working memory task. Neuroimage 55, 688–704.
Goldstein, R. Z., Volkow, N. D., Wang, G. J., Fowler, J. S., and Rajaram, S. (2001). Addiction changes orbitofrontal gyrus function: involvement in response inhibition. Neuroreport 12, 2595–2599.
Grabner, R. H., Ansari, D., Reishofer, G., Stern, E., Ebner, F., and Neuper, C. (2007). Individual differences in mathematical competence predict parietal brain activation during mental calculation. Neuroimage 38, 346–356.
Grezes, J., Tucker, M., Armony, J., Ellis, R., and Passingham, R. E. (2003). Objects automatically potentiate action: an fMRI study of implicit processing. Eur. J. Neurosci. 17, 2735–2740.
Hagmann, P., Cammoun, L., Gigandet, X., Gerhard, S., Grant, P. E., Wedeen, V., Meuli, R., Thiran, J. P., Honey, C. J., and Sporns, O. (2010). MR connectomics: principles and challenges. J. Neurosci. Methods 194, 34–45.
Hagmann, P., Cammoun, L., Gigandet, X., Meuli, R., Honey, C. J., Wedeen, V., and Sporns, O. (2008). Mapping the structural core of human cerebral cortex. PLoS Biol. 6, e159. doi: 10.1371/journal.pbio.0060159
Hampstead, B. M., Stringer, A. Y., Stilla, R. F., Deshpande, G., Hu, X. P., Moore, A. B., and Sathian, K. (2011). Activation and effective connectivity changes following explicit-memory training for face-name pairs in patients with mild cognitive impairment: a Pilot Study. Neurorehabil. Neural Repair 25, 210–222.
He, Y., and Evans, A. (2010). Graph theoretical modeling of brain connectivity. Curr. Opin. Neurol. 23, 341–350.
He, Y., Lo, C. Y., Wang, P. N., Chou, K. H., Wang, J. H., and Lin, C. P. (2010). Diffusion tensor tractography reveals abnormal topological organization in structural cortical networks in Alzheimer’s disease. J. Neurosci. 30, 16876–16885.
Holten, D. (2006). Hierarchical edge bundles: visualization of adjacency relations in hierarchical data. IEEE Trans. Vis. Comput. Graph 12, 741–748.
Irimia, A., Chambers, M. C., Alger, J. R., Filippou, M., Prastawa, M. W., Wang, B., Hovda, D. A., Gerig, G., Toga, A. W., Kikinis, R., Vespa, P. M., and Van Horn, J. D. (2011a). Comparison of acute and chronic traumatic brain injury using semi-automatic multimodal segmentation of MR volumes. J. Neurotrauma (in press).
Irimia, A., Chambers, M. C., and Van Horn, J. D. (2011b). Automatic Segmentation of Traumatic Brain Injury MRI Volumes Using Atlas Based Classification and 3D Slicer. Los Angeles, CA. Available at: http://www.na-mic.org/Wiki/index.php/TBISegmentation_TutorialContestSummer2011 [accessed 30 November, 2011].
Irimia, A., Chambers, M. C., Torgerson, C. M., and Van Horn, J. D. (2012). Circular representation of human cortical networks for subject and population-level connectomic visualization. Neuroimage (in press).
Krzywinski, M., Schein, J., Birol, I., Connors, J., Gascoyne, R., Horsman, D., Jones, S. J., and Marra, M. A. (2009). Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645.
Luu, P., and Pederson, S. M. (2004). “The anterior cingulate cortex: regulating actions in context,” in Cognitive Neuroscience of Attention, ed. M. I. Posner (New York: Guilford Press), 232–244.
Maes, F., Collignon, A., Vandermeulen, D., Marchal, G., and Suetens, P. (1997). Multimodality image registration by maximization of mutual information. IEEE Trans. Med. Imaging 16, 187–198.
Modha, D. S., and Singh, R. (2010). Network architecture of the long-distance pathways in the macaque brain. Proc. Natl. Acad. Sci. U.S.A. 107, 13485–13490.
Molnar-Szakacs, I., Iacoboni, M., Koski, L., and Mazziotta, J. C. (2005). Functional segregation within pars opercularis of the inferior frontal gyrus: evidence from fMRI studies of imitation and action observation. Cereb. Cortex 15, 986–994.
Oberman, L. M., and Ramachandran, V. S. (2008). Preliminary evidence for deficits in multisensory integration in autism spectrum disorders: the mirror neuron hypothesis. Soc. Neurosci. 3, 348–355.
Palva, J. M., Monto, S., Kulashekhar, S., and Palva, S. (2010). Neuronal synchrony reveals working memory networks and predicts individual memory capacity. Proc. Natl. Acad. Sci. U.S.A. 107, 7580–7585.
Park, E., Mcknight, S., Ai, J., and Baker, A. J. (2006). Purkinje cell vulnerability to mild and severe forebrain head trauma. J. Neuropathol. Exp. Neurol. 65, 226–234.
Peeters, R., Simone, L., Nelissen, K., Fabbri-Destro, M., Vanduffel, W., Rizzolatti, G., and Orban, G. A. (2009). The representation of tool use in humans and monkeys: common and uniquely human features. J. Neurosci. 29, 11523–11539.
Picard, N., and Strick, P. L. (1996). Motor areas of the medial wall: a review of their location and functional activation. Cereb. Cortex 6, 342–353.
Pohl, K. M., Bouix, S., Nakamura, M., Rohlfing, T., Mccarley, R. W., Kikinis, R., Grimson, W. E., Shenton, M. E., and Wells, W. M. (2007). A hierarchical algorithm for MR brain image parcellation. IEEE Trans. Med. Imaging 26, 1201–1212.
Prastawa, M., Bullitt, E., and Gerig, G. (2009). Simulation of brain tumors in MR images for evaluation of segmentation efficacy. Med. Image Anal. 13, 297–311.
Prastawa, M., Bullitt, E., Ho, S., and Gerig, G. (2004). A brain tumor segmentation framework based on outlier detection. Med. Image Anal. 8, 275–283.
Prastawa, M., Bullitt, E., Moon, N., Van Leemput, K., and Gerig, G. (2003). Automatic brain tumor segmentation by subject specific modification of atlas priors. Acad. Radiol. 10, 1341–1348.
Prastawa, M., and Gerig, G. (2008). Brain lesion segmentation through physical model estimation. Lecture Notes Comput. Sci. 5358, 562–571.
Radua, J., Phillips, M. L., Russell, T., Lawrence, N., Marshall, N., Kalidindi, S., El-Hage, W., Mcdonald, C., Giampietro, V., Brammer, M. J., David, A. S., and Surguladze, S. A. (2010). Neural response to specific components of fearful faces in healthy and schizophrenic adults. Neuroimage 49, 939–946.
Redcay, E. (2008). The superior temporal sulcus performs a common function for social and speech perception: implications for the emergence of autism. Neurosci. Biobehav. Rev. 32, 123–142.
Rogers, R. D., Owen, A. M., Middleton, H. C., Williams, E. J., Pickard, J. D., Sahakian, B. J., and Robbins, T. W. (1999). Choosing between small, likely rewards and large, unlikely rewards activates inferior and orbital prefrontal cortex. J. Neurosci. 19, 9029–9038.
Roland, P. E., and Friberg, L. (1985). Localization of cortical areas activated by thinking. J. Neurophysiol. 53, 1219–1243.
Samson, D., Apperly, I. A., Chiavarino, C., and Humphreys, G. W. (2004). Left temporoparietal junction is necessary for representing someone else’s belief. Nat. Neurosci. 7, 499–500.
Sanz-Arigita, E. J., Schoonheim, M. M., Damoiseaux, J. S., Rombouts, S. A. R. B., Maris, E., Barkhof, F., Scheltens, P., and Stam, C. J. (2010). Loss of ‘small-world’ networks in Alzheimer’s disease: graph analysis of fMRI resting-state functional connectivity. PLoS ONE 5, e13788. doi: 10.1371/journal.pone.0013788
Spanos, G. K., Wilde, E. A., Bigler, E. D., Cleavinger, H. B., Fearing, M. A., Levin, H. S., Li, X., and Hunter, J. V. (2007). Cerebellar atrophy after moderate-to-severe pediatric traumatic brain injury. AJNR Am. J. Neuroradiol. 28, 537–542.
Spoormaker, V. I., Schroter, M. S., Gleiser, P. M., Andrade, K. C., Dresler, M., Wehrle, R., Samann, P. G., and Czisch, M. (2010). Development of a large-scale functional brain network during human non-rapid eye movement sleep. J. Neurosci. 30, 11379–11387.
Tong, K. A., Ashwal, S., Holshouser, B. A., Shutter, L. A., Herigault, G., Haacke, E. M., and Kido, D. K. (2003). Hemorrhagic shearing lesions in children and adolescents with posttraumatic diffuse axonal injury: improved detection and initial results. Radiology 227, 332–339.
Ungerleider, L. G., Courtney, S. M., and Haxby, J. V. (1998). A neural system for human visual working memory. Proc. Natl. Acad. Sci. U.S.A. 95, 883–890.
van den Heuvel, M. P., and Pol, H. E. H. (2010). Exploring the brain network: a review on resting-state fMRI functional connectivity. Eur. Neuropsychopharmacol. 20, 519–534.
Van Leemput, K., Maes, F., Vandermeulen, D., and Suetens, P. (1999). Automated model-based tissue classification of MR images of the brain. IEEE Trans. Med. Imaging 18, 897–908.
Wang, B., Prastawa, M. W., Awate, S. P., Irimia, A., Chambers, M. C., Vespa, P. M., Van Horn, J. D., and Gerig, G. (2012). “Segmentation of serial MRI of TBI patients using personalized atlas construction and topological change estimation,” in Proceedings of the International Symposium on Biomedical Engineering (ISBI 2012), ed. I. Press (Barcelona: IEEE).
Wedeen, V. J., Wang, R. P., Schmahmann, J. D., Benner, T., Tseng, W. Y. I., Dai, G., Pandya, D. N., Hagmann, P., D’Arceuil, H., and De Crespignya, A. J. (2008). Diffusion spectrum magnetic resonance imaging (DSI) tractography of crossing fibers. Neuroimage 41, 1267–1277.
Yan, C., Gong, G., Wang, J., Wang, D., Liu, D., Zhu, C., Chen, Z. J., Evans, A., Zang, Y., and He, Y. (2011). Sex- and brain size-related small-world structural cortical networks in young adults: a DTI tractography study. Cereb. Cortex 21, 449–458.
Zald, D. H., and Kim, S. W. (1996). Anatomy and function of the orbital frontal cortex, I: anatomy, neurocircuitry; and obsessive-compulsive disorder. J. Neuropsychiatry Clin. Neurosci. 8, 125–138.
Zoellei, L., Shenton, M., Wells, W., and Pohl, K. (2007). “The impact of atlas formation methods on atlas-guided brain segmentation,” in International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), Brisbane.
Appendix

Table A1. Summary of each parcelation’s abbreviation, full description, and FreeSurfer code as defined in the original parcelation scheme (Destrieux et al., 2010), as well as the associated RGB code in the connectogram.
Keywords: connectomics, traumatic brain injury, atrophy, rehabilitation, DTI
Citation: Irimia A, Chambers MC, Torgerson CM, Filippou M, Hovda DA, Alger JR, Gerig G, Toga AW, Vespa PM, Kikinis R and Van Horn JD (2012) Patient-tailored connectomics visualization for the assessment of white matter atrophy in traumatic brain injury. Front. Neur. 3:10. doi: 10.3389/fneur.2012.00010
Received: 04 November 2011; Paper pending published: 28 November 2011;
Accepted: 16 January 2012; Published online: 06 February 2012.
Edited by:
Mattias Sköld, Uppsala University, SwedenReviewed by:
David Krishna Menon, University of Cambridge, UKElna-Marie Larsson, Uppsala University, Sweden
Copyright: © 2012 Irimia, Chambers, Torgerson, Filippou, Hovda, Alger, Gerig, Toga, Vespa, Kikinis and Van Horn. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: John D. Van Horn, Laboratory of Neuro Imaging, Department of Neurology, University of California Los Angeles, 635 Charles E. Young Drive South, Suite 225, Los Angeles, CA 90095, USA. e-mail:amFjay52YW5ob3JuQGxvbmkudWNsYS5lZHU=


 Carinna M. Torgerson1
Carinna M. Torgerson1




