Abstract
Research in cell biology greatly relies on cell-based in vitro assays and models that facilitate the investigation and understanding of specific biological events and processes under different conditions. The quality of such experimental models and particularly the level at which they represent cell behavior in the native tissue, is of critical importance for our understanding of cell interactions within tissues and organs. Conventionally, in vitro models are based on experimental manipulation of mammalian cells, grown as monolayers on flat, two-dimensional (2D) substrates. Despite the amazing progress and discoveries achieved with flat biology models, our ability to translate biological insights has been limited, since the 2D environment does not reflect the physiological behavior of cells in real tissues. Advances in 3D cell biology and engineering have led to the development of a new generation of cell culture formats that can better recapitulate the in vivo microenvironment, allowing us to examine cells and their interactions in a more biomimetic context. Modern biomedical research has at its disposal novel technological approaches that promote development of more sophisticated and robust tissue engineering in vitro models, including scaffold- or hydrogel-based formats, organotypic cultures, and organs-on-chips. Even though such systems are necessarily simplified to capture a particular range of physiology, their ability to model specific processes of human biology is greatly valued for their potential to close the gap between conventional animal studies and human (patho-) physiology. Here, we review recent advances in 3D biomimetic cultures, focusing on the technological bricks available to develop more physiologically relevant in vitro models of human tissues. By highlighting applications and examples of several physiological and disease models, we identify the limitations and challenges which the field needs to address in order to more effectively incorporate synthetic biomimetic culture platforms into biomedical research.
Introduction
Cell culture systems represent an indispensable tool for a wide range of biomedical studies. Harrison's first experiments, early in the twentieth century, on development of frog nerve fibers in a dish, the establishment of aseptic technique and subculture methods by Carrell and Ebeling in 1920s and the successful isolation and maintenance of the first immortalized human cell line (HeLa cells) by Gey in the 1950s, made it possible to grow cells in artificial environments, laying the foundation for cell and molecular biology (Taylor and Taylor, 2014; Jedrzejczak-Silicka, 2017; Simian and Bissell, 2017). Cell culture has come a long way since then and is now a vital and invaluable tool for a vast array of applications, both in academic and industrial settings, including drug development, cancer research and tissue engineering (Przyborski, 2017; Kapałczyńska et al., 2018). In such studies, two-dimensional (2D) cell culture systems dominate, continuing to improve our perception and understanding of cell biology. These cell systems rely mainly on adherent cultures, where cells grow as a monolayer attached to a plastic or glass substrate. Although easy and convenient, 2D cultures exhibit numerous disadvantages. Firstly, they are simplistic imitations of the in vivo situation, where cells grow within a complex three-dimensional (3D) microenvironment. The lack of this environmental context and structural architecture excludes physical cues for cell-cell and cell-matrix communication, critical for several cellular processes (e.g., mitosis, self-renewal, and differentiation). These physical constraints also impede cells from organizing naturally and spreading vertically, forcing them to flatten out and grow as monolayers (Fitzgerald et al., 2015; Przyborski, 2017). In turn, gene expression, production of proteins and cytoskeletal structure are altered, resulting in loss of the diverse cell phenotype and thus of the physiological cellular behavior and function (Birgersdotter et al., 2005; Luca et al., 2013; Fontoura et al., 2020). In addition, the absence of oxygen and nutrient gradients in monolayer cultures disrupts cell response to physiological stimuli, further inhibiting basic cellular processes and intercellular crosstalk, while the lack of a heterogeneous cell population in 2D models hinders their potential to form more complex tissue- or organ-like structures. These inherent limitations and shortcomings of 2D cell systems ultimately lead to failures in understanding cell behavior in healthy or diseased states (Duval et al., 2017). The research community is now beginning to seek alternative technologies that will facilitate development of models able to more closely mimic the complexity of whole tissues in vitro (Fitzgerald et al., 2015; Przyborski, 2017; Kapałczyńska et al., 2018). To this end, 3D cell cultures can provide a well-controlled in vivo-like microenvironment specifically tailored to each application (Chen, 2016; Koledova, 2017; Owens et al., 2017; Przyborski, 2017; Kapałczyńska et al., 2018; Jensen and Teng, 2020).
Although it is thought that the inception of 3D biology was in the 1970s (Schwarz and Bissell, 1977; Bissell, 1981; Bissell and Barcellos-Hoff, 1987), where cells were cultivated in floating collagen gels or agar, in fact, the phrase “three-dimensional cell culture models” was first coined in the studies of Barchelos-Hoff in 1989 and Petersen in 1992, who developed assays to distinguish between healthy and malignant breast epithelial cells grown in laminin-rich matrices (Hamburger and Salmon, 1977; Bissell, 2017; Simian and Bissell, 2017). These studies were followed by a body of research on new technologies focused on enhancing the morphological and physiological relevance of cell culture systems. The increasing number of publications since then, utilizing such cell culture platforms, or suggesting new ones, highlights the transition of the field into 3D cell culture in order to improve the capabilities of experiments performed in vitro (Bissell, 2017; Przyborski, 2017; Devarasetty et al., 2020). Over the years, 3D cell culture has become a generalized term, often used to point out the disparities between conventional and new cell culture technologies. Therefore, it is important to clearly define what is meant—or should be meant—by this term. Broadly speaking, we define 3D cell culture as an in vitro tissue-specific microenvironment that enables individual cells to grow, maintaining their 3D shape and functions, as well as to interact with their surroundings and a heterogeneous population of neighboring cells, establishing sufficient signaling networks. In this environment exogenous interference and support should be minimized (e.g., automated media perfusion) to reduce stress and unnatural cell responses and rather enable growth of different cell types to foster the development of more realistic culture systems (Abbott, 2003; Huh et al., 2011; Shamir and Ewald, 2014). 3D cell culture is also used to describe tissue- and organ-like structures emerging from the combination of 3D cell biology with Tissue Engineering (TE) principles. In these studies, researchers are focused on reconstructing organ structure and function ex vivo (Figure 1), to produce more reliable and physiologically relevant human-like 3D in vitro models (Khademhosseini and Langer, 2016; Caddeo et al., 2017), following the basic TE premise: the appropriate cell types (e.g., primary cells, stem cells) are seeded in biodegradable structures fabricated to mimic the target organ or tissue (i.e., scaffolds) and are supplied with the appropriate cocktail of substances essential for tissue generation (e.g., growth factors and signaling molecules; Langer and Vacanti, 1993). Such tissue-engineered human equivalents represent a promising alternative to the current state-of-the-art and particularly to animal models, which often fail to recapitulate human conditions due to differences in the overall physiology and in the molecular and signaling mechanisms involved in the onset and progression of diseases. This is evidenced by the high failure rates of drugs and therapies to enter clinical trials to get approval from regulatory agencies despite successful tests in animals, underlying the challenges in translating such data to human systems. Besides their greater translational relevance, the development of tissue-engineered in vitro models has recently taken off thanks to ethical and economic arguments (Rouwkema et al., 2011; Caddeo et al., 2017). Although the use and welfare of animals in science is protected by national and international legislation (e.g., the principle of 3Rs), there are still vibrant discussions and room for improvements as well as strong encouragement for reduction and replacement. This is also encouraged by an economic point of view, since the actual costs for drug or treatment candidates to become clinical products are huge and the process is time-consuming and labor intensive (Rouwkema et al., 2011; Fitzgerald et al., 2015).
Figure 1
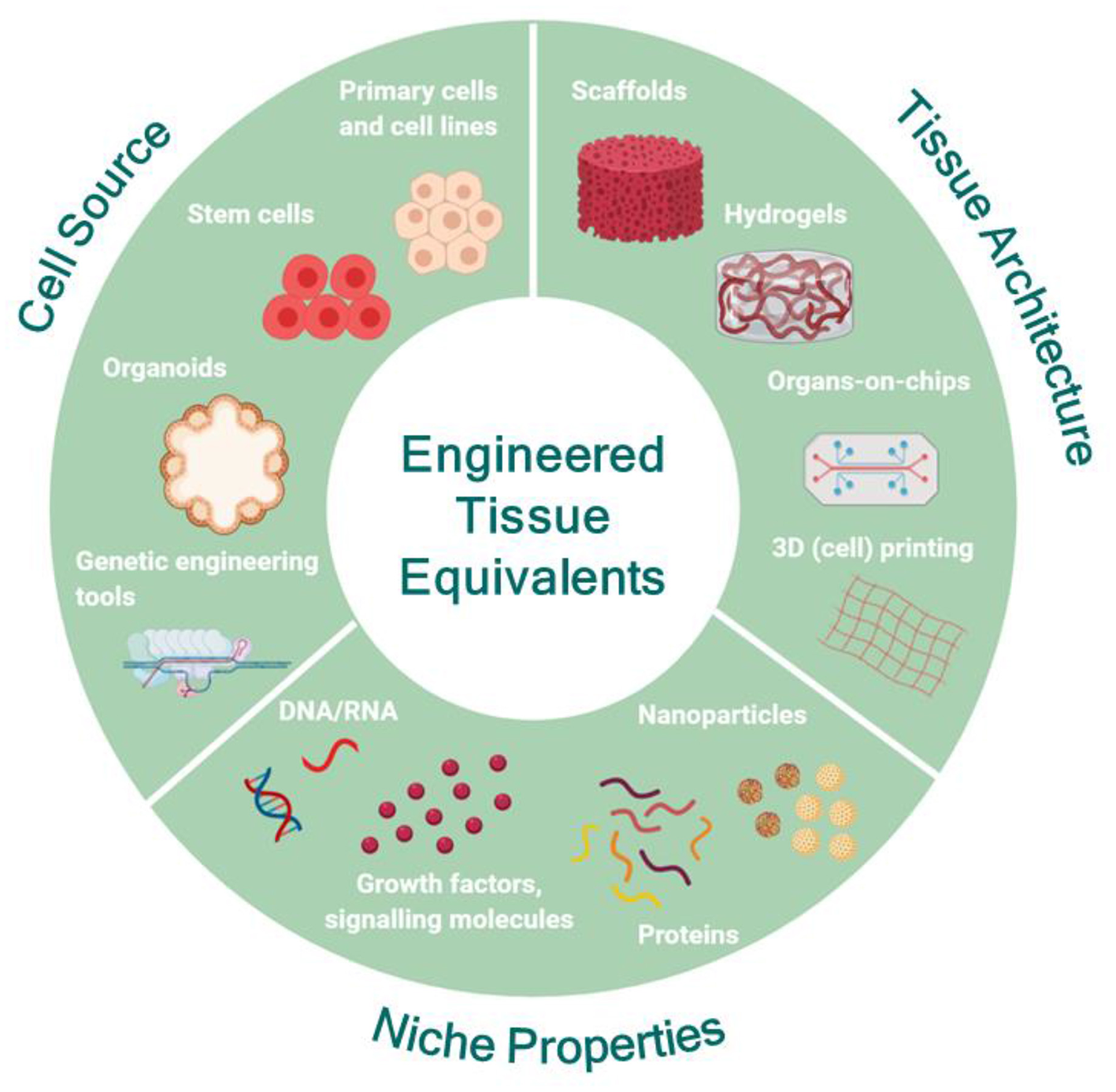
Engineering human tissue equivalents in vitro: the main premise for the successful development of tissue equivalents is to understand the structural and functional role of each counterpart of the native tissue and to carefully choose the range of features necessary to recapitulate the specific characteristics of the native tissue for each application. Then, the appropriate source of cells can be identified and modulated, if necessary, to capture the desired functionality. In parallel, the most appropriate substrate format can be designed and engineered to match the physicochemical properties and architecture of the native tissue under the conditions of interest and to facilitate coupling with the appropriate biochemical and biophysical cues mimicking the in vivo niche. Created with BioRender.com.
In this review, we focus on bioengineering approaches that seek to integrate TE with 3D cell biology toward more sophisticated and reliable 3D human tissue equivalents, with the potential to (i) enhance the predictive value of preclinical studies, (ii) improve the way we study physiology and pathology and thus to address biological questions that so far necessitated animal models, and (iii) bridge the gap between current (pre-)clinical research tools and human systems by assisting and advancing drug development processes in terms of science, bioethics and economy (Rouwkema et al., 2011; Fitzgerald et al., 2015; Caddeo et al., 2017). Current trends in the field suggest that the choice of the culture format/technology/device that will support the tissue equivalent should take into account the constituent parts of the organ(s) to be modeled and the extent to which the in vivo complexity will be recapitulated. Various techniques and culture formats have been developed to meet these requirements, however a single format/technology/device that meets the requirements/needs of all 3D cell culture assays does not exist, and indeed should not exist, given the diversity in the morphology and functions of all the different organs/tissues researchers are emulating for different applications (Shamir and Ewald, 2014; Knight and Przyborski, 2015). Based on the format, 3D cell culture platforms can generally be categorized as scaffold-free or scaffold-based systems. Scaffold-based approaches utilize natural or synthetic materials to provide support in the form of a matrix that creates the desired tissue-specific microenvironment for optimal cell growth and differentiation and natural ECM deposition, while preserving the native tissue architecture (Fitzgerald et al., 2015; Przyborski, 2017). Decellularised scaffolds have also been used to culture cells in vitro by removing cells from whole tissues/organs or from the scaffold biomaterial surface after culturing cells on it for sufficient time for native ECM deposition. This way the structural and functional matrix proteins (e.g., collagen, fibronectin, hyaluronic acid, and laminin) remain intact while the exact composition varies according to the origin of tissue/organ or the cells seeded (Fitzgerald et al., 2015). Another category where cell growth and differentiation is supported by biomimetic matrices is hydrogels, which are networks of cross-linked hydrophilic polymers with the unique capability to absorb and retain copious amounts of water without dissolving but rather swelling. Similar to scaffolds, hydrogels can be made from natural or synthetic materials and cross-linked by either physical or chemical means. Due to their particularly soft nature, they are well-suited for soft tissue in vitro models (Fitzgerald et al., 2015; Przyborski, 2017). In contrast, scaffold-free culture systems seek the formation of multicellular masses without exogenous input as a framework, but rather by encouraging cells to form aggregates, secrete their own ECM and then self-assemble into 3D microstructures (Przyborski, 2017). This categorization however is quite generalized since the progress in, and convergence of, related disciplines have made possible the generation of new, improved, and more sophisticated tools for 3D biomimetic cultures. Currently, novel biofabrication methods (e.g., 3D printing) and microscale technologies (e.g., soft lithography), can be combined with advanced biological systems such as organoids and stem cells, resulting in more complex culture systems, tailored for specific applications. In the following sections, we first provide an overview of the current technological bricks available to develop tissue-engineered human models by summarizing the advancements in cell biology, materials science and bioengineering. Then, through examples of the current state-of-the-art, we identify and discuss the advantages, limitations and challenges the field needs to address in order for in vitro TE models to be successfully implemented in biomedical research.
Building Blocks for Developing Human Tissue Equivalents
Cell Sources for in vitro Tissue Engineering
To successfully design and develop tissue equivalents, it is useful to understand the anatomical and functional characteristics of the tissue of interest, as well as the role and interaction of its constituent parts (Caddeo et al., 2017). Although the end goal is to create organ and tissue equivalents with enhanced biomimicry in the lab, it is important to acknowledge the reductionist nature of these models. In fact, 3D human tissue equivalents are intentionally and necessarily reductionist, carefully designed to capture a specific range of the in vivo physiology over time, fit for a specific application. Therefore, a crucial step in the design process is to identify the appropriate factors that must be incorporated in order to model different in vivo situations (Chen, 2016). To this end, the source and the number of cells must be carefully chosen as this will determine the ability of the model to capture in vitro the desired characteristics of the native tissue at the cell culture level and then establish what the system might or might not recapitulate and to what extent (Chen, 2016; Caddeo et al., 2017).
Stem Cells
Until recently, TE approaches relied almost exclusively on established cell lines and primary cells. Despite the advantages of using cell lines (e.g., easy to use, inexpensive, unlimited availability, reproducibility, no need of ethics approval), they are not considered ideal sources for modeling human conditions since they do not exhibit normal features, often drifting from the genetic and phenotypic profile of the tissue of origin (Carter and Shieh, 2015a). Primary cells are more representative of the morphological and functional features of the tissue they are derived from, but they can be difficult to obtain and maintain for long-term experiments. Moreover, they have low proliferation rates and must be used in early passage stages because they lose their structural, functional, and self-renewal properties as they undergo senescence processes. Reproducibility of results is an additional issue when using primary cells and donor-to-donor variations must be taken into account (Benam et al., 2015; Caddeo et al., 2017).
To overcome these limitations, stem cells are now being employed for reconstructing tissue/organ structure and function in vitro, due to their unique capabilities to self-renew (stemness) and to differentiate toward one or more specialized cell types (potency), representing a versatile source of cellular substitutes for a wide range of applications (Avior et al., 2016; McKee and Chaudhry, 2017; Rowe and Daley, 2019). Until recently, the only source of stem cells for biomedical research was human-derived embryonic stem cell lines (ESCs). However, the discovery of human induced pluripotent stem cells (iPSCs) in 2007 (Takahashi et al., 2007) substantially altered the field of biomedical research. iPSCs are engineered stem cells generated directly from adult (differentiated) somatic cells by introducing a set of pluripotency-associated genes into cells, or through chemical reprogramming or protein delivery (Khademhosseini and Langer, 2016; Caddeo et al., 2017). These cells exhibit similar stemness and potency characteristics as ESCs and, under certain conditions and depending on their origin, they can differentiate toward various cell types. Among other applications, human iPSCs offer an unlimited supply of cells for in vitro TE, disease modeling, cell therapy and pharmaceutical applications. Importantly, as these stem cells can be derived from patients with specific pathology, patient-derived hiPSCs are now used to more accurately model disease and to improve diagnostics and drug discovery, laying the foundations for novel methods of personalized medicine (Cyranoski, 2018; Rowe and Daley, 2019). Despite the great potential of iPSCs to bridge the gap between preclinical studies, animal models and clinical studies, it is important to note that currently most iPSC-derived cell types exhibit immature phenotypes, while some pathologies cause such damage in adult somatic cells that the iPSCs derived from such tissues would not be informative (Benam et al., 2015; Cyranoski, 2018). Ongoing research on establishing protocols for the maturation of iPSC-derived specialized cell types will likely address these challenges via applying different biochemical and biophysical stimuli. TE strategies could be useful in these efforts by providing the appropriate microenvironment (e.g., biomimetic scaffolds) and cues (e.g., ECM rich in growth factors) to guide differentiation and maturation (Benam et al., 2015).
Finally, another type of stem cells that have gained a lot of attention as a cell source for TE are mesenchymal stem/stromal cells (MSCs), mainly due to their therapeutic potential (Rosenbaum et al., 2008; Khademhosseini and Langer, 2016). MSCs are a specific subtype of multipotent stem cells, diversely distributed in the human body including bone marrow, adipose, perinatal tissues, blood, periodontal ligament, and skeletal muscles, from most of which they can be isolated (Rosenbaum et al., 2008; McKee and Chaudhry, 2017; Ullah et al., 2019). Despite having the same capabilities as pluripotent stem cells, MSCs can differentiate toward only a few specific cell types, such as osteogenic, chondrogenic, and adipogenic cell types, depending on the nature and maturity of the tissue of origin (Rosenbaum et al., 2008; McKee and Chaudhry, 2017). Their homing capability has made them very attractive candidates for a wide breadth of preclinical and clinical applications, including tissue regeneration, wound healing, and treatment of autoimmune diseases (Khademhosseini and Langer, 2016; Ullah et al., 2019). Upon injury, MSCs are naturally released in the circulation and migrate to the damaged tissue where they secrete a pool of cytokines, growth factors and other bioactive molecules with immunomodulatory and angiogenic effects, thus creating a microenvironment that promotes tissue repair and regeneration (Ullah et al., 2019). For in vitro TE, MSCs represent a useful resource, mainly due to their ease in isolation, manipulation and differentiation, compared to the longer and more elaborate iPSCs protocols.
Organoids
The advent of human pluripotent stem cells marked the starting point of the development of “organs in a dish,” also known as organotypic cultures or organoids, a major breakthrough of the past decade (Dutta et al., 2017; Lancaster and Huch, 2019). The term “organoid” is not new; it was used in the 1950s and 1960s to describe structures in cell culture systems that resembled organs (Duryee and Doherty, 1954; Schneider et al., 1963; Wolter, 1967) and more recently in studies where 3D cell aggregates, called spheroids, were defined as organoids despite the fact that they were not fully representative of the native tissue (Dutta et al., 2017). The term has become popularized in in vitro biology and evolved to generally refer to tissues or structures that resemble an organ, losing its precision (Dutta et al., 2017; Lancaster and Huch, 2019). A more specific working definition that fulfills the basic definition of organoids was recently proposed, along with several criteria: “(1) a 3D structure containing cells that establish or retain the identity of the organ being modeled; (2) the presence of multiple cell types, as in the organ itself; (3) the tissue exhibits some aspect of the specialized function of the organ; and (4) self-organization according to the same intrinsic organizing principles as in the organ itself” (Lancaster and Huch, 2019). These properties render organoids suitable formats/tools for modeling organ architecture in vitro. Because iPSC-derived organoids follow in vivo like development, their morphology closely recapitulates the native organ structure, making them particularly apt for studies looking at developmental organogenesis, while tissue-specific adult stem cell organoids are mostly suited for studying tissue homeostasis and maintenance, since naturally in the body they are key players in these processes (Fatehullah et al., 2016; Yin et al., 2016; Lancaster and Huch, 2019).
Currently, several protocols exist for the development of organoids for various organs, derived either from pluripotent stem cells (ESCs and iPSCs) or from organ-specific adult stem cells (ASCs) and progenitor cells (Takebe and Wells, 2019). The establishment of a protocol for a long-term, well-defined, and stable culture of murine intestinal organoids in 2009 by Sato et al. (2009) and subsequent adaptation of the protocol and modifications of the growth factor cocktail in the original organoid culture medium allowed the generation of human organoids from various tissues, such as stomach (Bartfeld et al., 2015), liver (Takebe et al., 2013), esophagus (Li et al., 2018), lung (Dye et al., 2015), and ovaries (Kessler et al., 2015; see Kim et al., 2020, for an extended review). Organoids now represent a powerful tool for a wide spectrum of biomedical applications ranging from basic cell biology studies, organogenesis and tissue homeostasis to disease modeling (see Lancaster and Huch, 2019, for an extended review), drug/therapy development, and regenerative medicine (Schweiger and Jensen, 2016; Brassard and Lutolf, 2019). However, there are several general shortcomings and challenges in the development and application of organoids, as well as in the interpretation and translation of the derived data. An important issue is the reproducibility and consistency of organoids from batch to batch. The initial culture conditions and the environment in which organoids grow are of paramount importance for their self-organization and the development of the desired emergent tissue (Brassard and Lutolf, 2019). To date, most organoid systems rely on animal-derived ECMs, such as Matrigel, supplemented with growth factors and endogenous signaling molecules (e.g., Wnt, Noggin, and R-spondin). Even though Matrigel works as an artificial niche, mimicking the native tissue environmental cues, its poorly understood composition, heterogenous nature, and batch-to-batch variability hinders the reproducibility and robustness of the organoid systems, often leading to heterogeneity in size, shape, and viability, even between organoids in the same culture (Fatehullah et al., 2016; De Souza, 2018; Brassard and Lutolf, 2019; Lancaster and Huch, 2019; Kim et al., 2020). In addition, although these organotypic cultures are highly biologically relevant, they alone do not necessarily recapitulate the dynamics present in the human system. For example, the majority of organoid culture systems lack essential components of their living counterparts, such as the enteric nervous system, the immune system, as well as luminal flow and peristalsis (In et al., 2016; Tsakmaki et al., 2017; De Souza, 2018; Kim et al., 2020). Also, lack of vascularization in organoid cultures means that their growth and development depends on diffusion of nutrients from the surrounding media. While this might not be an issue for small organoids, in cases of some larger organoids the diffusion of nutrients is limited, resulting in dramatic necrosis in their interior and hence compromising the long-term viability of the system and the validity of the results (McMurtrey, 2016; Grebenyuk and Ranga, 2019; Lancaster and Huch, 2019). To overcome this limitation, culturing organoids under flow within microfluidic chips has been proposed recently, as a means to induce vascularization and hence to improve the morphological and functional characteristics of the bioengineered tissues (Homan et al., 2019).
The tremendous potential of organoid culture systems could not go unnoticed by tissue engineers. Firstly, since organoid systems are designed and developed to recapitulate the environment and properties of the stem cell niche and the tissue progeny with indefinite culture potential, they represent an alternative, more accessible, and scalable source for harnessing stem cells (Yin et al., 2016). Employment of molecular technology and organoids in the service of in vitro TE can further enhance the potential of these systems in mimicking the in vivo conditions. In particular, genes within organoids can be manipulated using tools, such as CRISPR/Cas-9, to either correct mutations and restore physiological function or to introduce mutations and model various disease phenotypes (Dutta et al., 2017; Lancaster and Huch, 2019; Kim et al., 2020), again providing an alternative source of cells with broad applicability and amenable to manipulation. Additionally, since organoids may contain more than one tissue representative cell type, they can be used as a single cellular input for tissue-engineered equivalents, allowing them to better capture the cellular diversity of the living counterparts (Kasendra et al., 2018).
Stem cells and organoids are a versatile source of cells for in vitro TE applications, thus the convergence of these fields can be mutually beneficial. On one hand the use of human-derived cellular parts enhances the relevance of tissue-engineered equivalents, both in terms of mimicry and data interpretation. On the other hand, tissue engineering provides a broad toolbox to study stem cells and organoids and to address challenges related to engineering the appropriate niche for controlling the culture conditions. It also provides the extrinsic instruction patterns to robustly and elaborately direct self-organization processes. This way TE approaches can aid stem cells and organoids in realizing their full potential as in vitro tools for biomedical research (Benam et al., 2015; Chen, 2016; McKee and Chaudhry, 2017; Brassard and Lutolf, 2019).
Materials for in vitro Tissue Engineering
Alongside the appropriate cell type(s), another fundamental element for effectively engineering tissue equivalents is the choice of the suitable biomaterial(s). In the classic TE paradigm, pre-engineered 3D supports/scaffolds, made from natural or synthetic materials, are used as templates for cell attachment, growth and differentiation toward functional living constructs (Vacanti, 2006). As scaffolds act as a synthetic ECM, modifying the biomaterial building blocks to mimic the native tissue ECM is a major challenge. In the body, ECM is a 3D network that consists of various macromolecules, including proteins and polysaccharides, responsible for tissue support and maintenance, cell-cell and cell-ECM communication, diffusion of nutrients metabolites and growth factors. In addition, ECM mediates signaling pathways from soluble factors and other sources, regulating various cellular behaviors, such as migration, adhesion, proliferation, and differentiation (Frantz et al., 2010; Theocharis et al., 2016; Afewerki et al., 2019; Figure 2). In this context, the role of biomaterials for tissue engineered equivalents is to provide cells with the appropriate framework/template to adhere, proliferate, differentiate, maturate, secrete ECM and form the necessary cell-cell, and cell-scaffold interactions that will enable them to auto-organize as they would in vivo (Przyborski, 2017; Afewerki et al., 2019; Figure 3). The choice of biomaterial depends on the application and the physiological or pathophysiological conditions the tissue model aims to emulate (Caddeo et al., 2017). However, there are many other properties that need to be considered when selecting biomaterials, including biocompatibility, strength and elasticity, porosity, molecular gradients and mass transport of nutrients and growth factors, oxygenation, adhesion or signaling sites, surface roughness, shape, type (e.g., scaffold or hydrogel), and source (Sitarski et al., 2018), in order to more realistically recapitulate the cues naturally occurring in the native tissue.
Figure 2
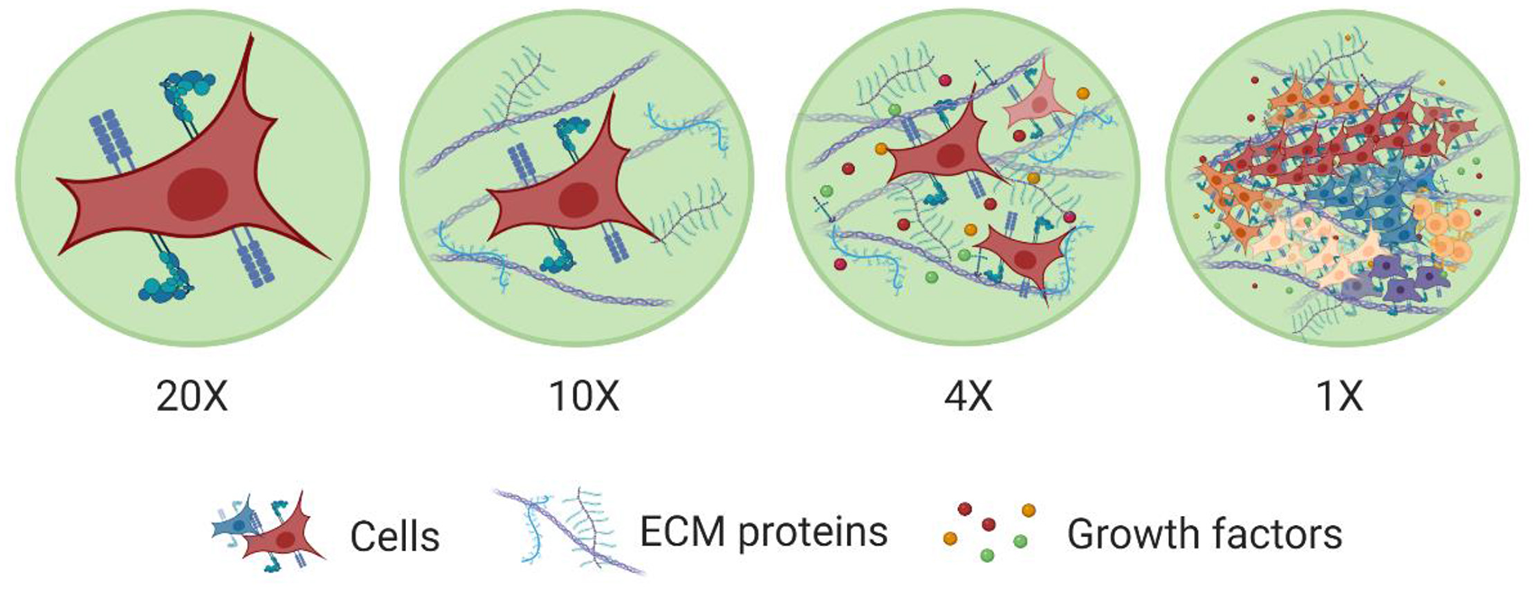
Levels of organization within tissues. The major building block of tissues are cells. Cell membranes are equipped with a wide range of proteins that help them sense and respond to cues in their microenvironment. Cells also interact with other cells and the surrounding ECM network. This ECM network comprises of various macromolecules (e.g., proteins, polysaccharides) and soluble factors (e.g., growth factors) immobilized in its structure, which promote cell-cell and cell-ECM interactions. This way ECM establishes a favorable niche for cells to grow, spread, differentiate, and perform various functions and also to work together with other cells to form more complex structures toward tissues with specific functionalities. Created with BioRender.com.
Figure 3
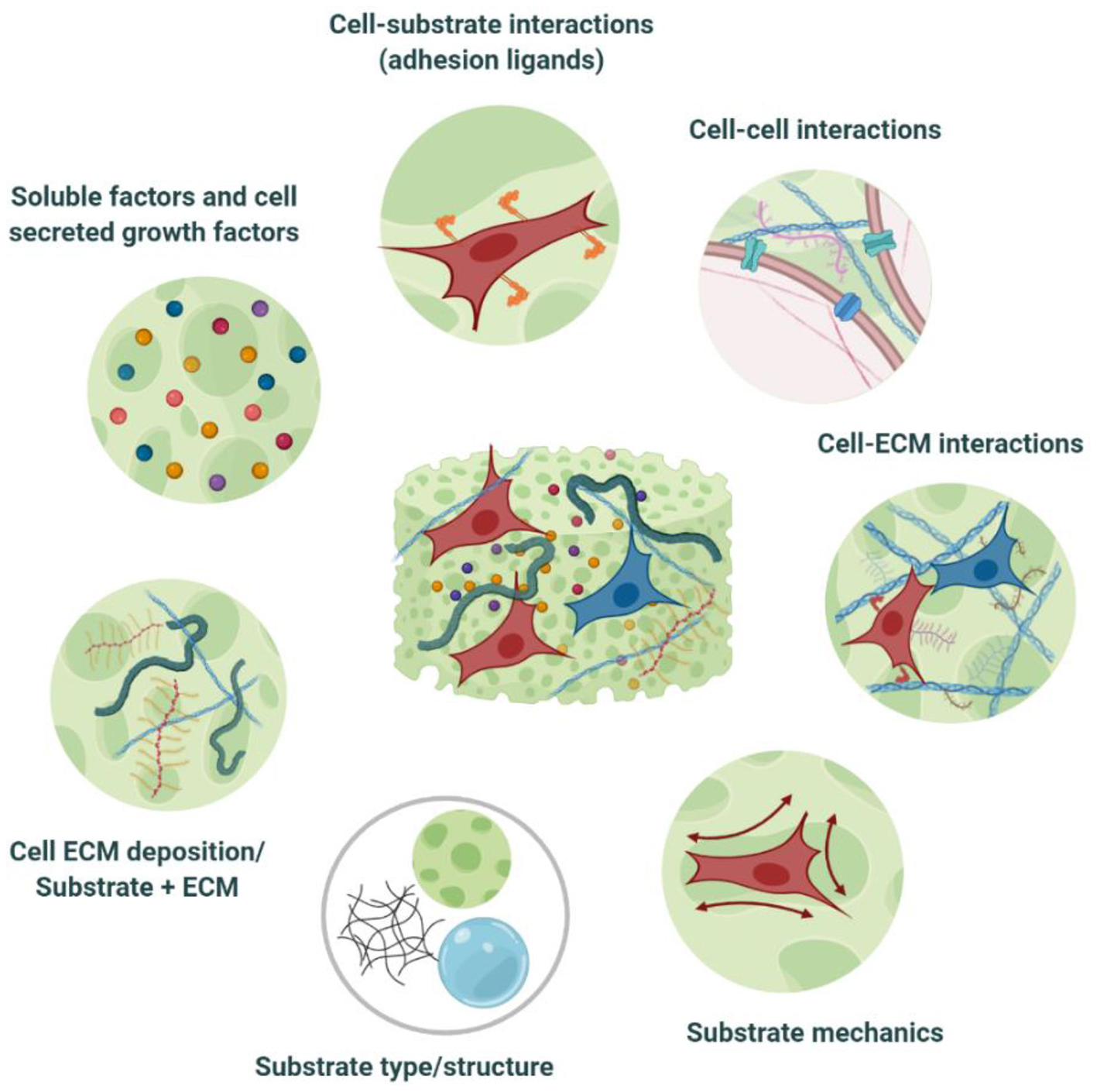
Cell-biomaterial interactions within tissue engineered equivalents. TE substrates constitute an artificial ECM, encompassing in vivo-like, tissue-specific biochemical, and biophysical cues. This niche provides a favorable microenvironment for cells to adhere, proliferate, differentiate, maturate, and deposit their own ECM, as well as to communicate and to establish the necessary cell-cell, and cell-ECM interactions that will enable them to auto-organize as they would in vivo. Modulation of the niche properties can also guide cell bahaviour toward the desired phenotypic output. Created with BioRender.com.
A broad range of materials is now available for the fabrication of various types of substrates, the properties of which can be tailored at the micro- and nano-scale to match the requirements of specific applications (Huang et al., 2017; Nikolova and Chavali, 2019; Cembran et al., 2020). A characteristic property of TE scaffolds that particularly affects primary cell-matrix interactions as well as cell behavior and fate upon seeding, is its topography. It is well-known that micro-topography (1–100 μm) is responsible for cell recruitment, adhesion, orientation and gene expression, while the submicron and nano- features strongly influence the cytoskeletal arrangement (Hayes and Richards, 2010). Tailoring surface topography of biomaterials has been shown to support and enhance differentiation of MSCs toward specific lineages (Abagnale et al., 2015). Mechanobiology studies have revealed that, amongst other cues, cells are also responsive to the material stiffness (Discher et al., 2005), which can affect intracellular signaling cascades that trigger cell adhesion, phenotype maintenance, cytoskeletal reconstruction, and even stem cell differentiation (Engler et al., 2006; Mao et al., 2016; Cao et al., 2017; Kumar et al., 2017; Vining and Mooney, 2017; Darnell et al., 2018). Therefore, selection of biomaterials with the appropriate stiffness is not only important for matching the native tissue mechanical properties, but can be also used as a tool to control cell phenotypes and thus modulate cell behavior (Khademhosseini and Langer, 2016; Ledo et al., 2020b). Furthermore, materials can undergo several chemical modifications, to improve their physicochemical properties as well as to enable incorporation of biologically relevant molecules and signals necessary for guiding and regulating cell response. For example, proteins from the ECM (e.g., hyaluronic acid, collagen, fibronectin etc.) can be blended or grafted to the material surface to improve cell behavior, by acting as matrix-associated biological cues, regulating cell attachment (e.g., via integrin-mediated binding), as well as proliferation and infiltration within the scaffold (Baker and Chen, 2012). For instance, incorporation of well-known cell binding motifs from ECM-derived proteins, such as RGD (arginine-glycine-aspartate) peptide, can enhance cell spreading, and viability in hydrogels (Gallagher et al., 2020).
Biomaterials may be prepared of natural polymers such as collagen, laminin, and hyaluronic acid, or from synthetic materials such as polyethylene glycol (PEG), propylene glycol diacetate (PGDA), polyvinylidene fluoride (PVDF), or co-polymers (Afewerki et al., 2019; Nikolova and Chavali, 2019). Thus, they can be categorized as natural, synthetic, or hybrid biomaterials (see Table 1 for examples of biomaterials and applications). In most cases, naturally derived biomaterials are amino acid-based or sugar-based biopolymers which can be components of the natural ECM (e.g., collagen, laminin, elastin, and fibrinogen) or not (e.g., chitin, silk fibroin, chitosan, and alginate; Silva et al., 2017; Ahadian et al., 2018). Such materials represent an attractive source for in vitro TE applications, due to their microstructure, stability, biocompatibility, and ability to present cells with natural adhesion sites, as well as due to the possibility to tailor and control their properties via physical or chemical treatments (i.e., cross-linking) or by blending them with other biopolymers (Guarino et al., 2016; Ullah and Chen, 2020) to better recapitulate in vitro the physiological milieu. A commonly used natural biomaterial in 3D biomimetic cultures and tissues is collagen, as it is a major component of the natural ECM and it is among the main structural proteins of most connective tissues (Ahadian et al., 2018; Sorushanova et al., 2019). The prevalence of collagen in human tissues, its excellent properties (e.g., low immunogenicity, biocompatibility, biodegradability, hydrophilicity, easy processing, good encapsulation response, etc.; Ahadian et al., 2018; Liu X. et al., 2019) and the advances in preparation and cross-linking methods to boost its physicochemical properties have enabled the fabrication of various types of collagen-based bioactive substrates (e.g., scaffolds, gels, fibers, and sponges; Uchino et al., 2009; Yip and Cho, 2013; Patel et al., 2019; Sorushanova et al., 2019; Ferro et al., 2020). Skin substitute studies, for instance, have extensively employed this natural material for wound healing purposes (Min et al., 2018), while collagen hydrogels have also been shown to effectively support and enhance the growth and survival of primary cortical neurons in a 3D mimetic environment (Evans et al., 2019).
Table 1
| Material(s) and format | Fabrication method | Cell source | References |
|---|---|---|---|
 Skin biomimetic cultures
Skin biomimetic cultures
|
|||
| Collagen hydrogel | Gelation/3D bioprinting | Fibroblasts, melanocytes (MCs), and keratinocytes (KCs) | Min et al., 2018 |
| Fibroblasts and KCs | Lee et al., 2009 | ||
| Alginate/Carboxymethyl cellulose/Nanofibrillated cellulose (ALG/CMC/NFC) bioinks | Human skin fibroblasts (hSF) | Zidarič et al., 2020 | |
| Collagen I scaffolds | Gelation/vitrification | Normal human skin fibroblasts (NhSF), Normal human dendritic cells (NHDC), Normal human epidermal keratinocytes (NHEK) | Uchino et al., 2009 |
| Silk Fibroin/Collagen (SF/COL) scaffolds | Freeze-drying | Primary neonatal foreskin fibroblast (NH), hiNSCs | Vidal et al., 2019 |
| Polycaprolactone (PCL), Polycaprolactone/Collagen (PCL/COL), Polycaprolactone/Poly (L-lactic acid) (PCL/PLLA), Polycaprolactone/Poly (L-lactic acid)/Collagen (PCL/PLLA/COL) scaffolds | MSCs | Rahmani et al., 2018 | |
| Polycaprolactone/Aloe vera (PCL/AV), Polycaprolactone/Curcumin (PCL/CUR), Polycaprolactone/Aloe Vera/Tetracycline hydrochloride (PCL/AV/TCH) scaffolds | Electrospinning | Human dermal fibroblasts | Ezhilarasu et al., 2019 |
| Gelatin methacrylate/Nanofibrils (GelMa/NF), Gelatin/Nanofibrils | Rnjak et al., 2009 | ||
| Milk protein/Polycaprolactone (MP/PCL) scaffolds | Human keratinocytes | Hewitt et al., 2019 | |
| Synthetic Elastin (SE) hydrogels | Human dermal fibroblasts, Human keratinocytes | Mao et al., 2018 | |
 Brain biomimetic cultures
Brain biomimetic cultures
|
|||
| Collagen gels | Gelation | Neuroblastoma cell line (SK-N-BE) | Villasante et al., 2017 |
| Alginate/Collagen hydrogels | hiPSCs | Moxon et al., 2019 | |
| Gelatin hydrogels coated with Collagen IV/Fibronectin | hiPSC, human brain microvascular endothelial cells (BMECs) | Faley et al., 2019 | |
| Silk fibroin scaffold/Collagen I hydrogels | Silk extraction/Salt-leaching | hiPSCs | Rouleau et al., 2020 |
| Liquid crystal elastomers (LCE) scaffolds | Salt-leaching | SH-SY5Y | Prévô et al., 2018 |
| Silk fibroin (SF) scaffolds | Freeze-drying | hiNSCs | Cairns et al., 2020 |
| Pol(vinyl alcohol)/Sodium alginate (PVA/SA) fibers | Multilayer Coaxial Laminar Flow | hiPSCs | Zhu et al., 2017 |
| Sodium Alginate/Gelatin (SA/Gel) based bioinks | Gelation/3D bioprinting | SH-SY5Y, hiPSCs | Fantini et al., 2019 |
| Gelatin Methacrylate (GelMa), Glycidyl/Methacrylate/Hyaluronic acid (GM/HA) hydrogels | Neural Stem Cells (NSCs) | Tang et al., 2020 | |
| Methacrylated Alginate (AlgMA) hydrogels | Neuroblastoma cell line (SK-N-BE) | Monferrer et al., 2020 | |
| Poly(desaminotyrosyl tyrosine ethyl ester carbonate) (pDTEc) nanofibers | Electrospinning | Neural reprogrammed stem cells (RN-iPS) | Carlson et al., 2016 |
| Polycaprolactone (PCL) scaffolds | Human neural progenitor stem cells (hNPCs) | Jakobsson et al., 2017 | |
| Polyethylene diacrylate (PEGDA) scaffolds | UV polymerization | Murphy et al., 2020 | |
 Heart biomimetic cultures
Heart biomimetic cultures
|
|||
| Collagen nanofibers | Electrospinning | Human Bone marrow mesenchymal stem cells (hBM-MSC) | Joshi et al., 2018 |
| Poly(vinylidene fluoride)/Trifluoroethylene (PVDF/TrFE) scaffolds | hiPSCs, Cardiomyocytes (CMs) | Adadi et al., 2020 | |
| Collagen fibers | Melt electro-wiring | Human umbilical cord vein smooth muscle cells (HUVSMCs) | Saidy et al., 2019 |
| Alginate (Alg) hydrogel nanofibers, Alginate/Gelatin (Alg/GelF/MA) hydrogel nanofibers | Wet-electrospinning | Mesenchymal stem cells enhanced with enhanced green fluorescent protein (hEGFP-MSCs) | Majidi et al., 2018 |
| Gelatin hydrogels | Gelation/3D bioprinting | hMSCs | Tijore et al., 2020 |
| Hyaluronic Acid/Arginin-Glycine-Aspartic Acid (HA/RGD) hydrogels | Gallagher et al., 2020 | ||
| Gelatin Methacrylate (GelMa) bioinks | HUVECs, CMs | Zhang et al., 2016b | |
| Gelatin Methacrylate/Polyethylene diacrylate (GelMa-PEGDA) bioinks | Induced multipotent stem cells (iMSCSs) | Nachlas et al., 2020 | |
| Gelatin/Gellan Gum (GG) hydrogels | hiPSC-CMs | Koivisto et al., 2019 | |
| Poly(vinyl alcohol) (PVA) scaffolds | Freeze-drying | Dattola et al., 2019 | |
| Polycaprolactone films (MacPCL) | Layer-by-layer assembly, Laser perforation | hMSCs | Zhang et al., 2019 |
| Matrigel coated fiber matrices | Two-photon polymerization | hiPSC-CMs | Wang C. et al., 2020 |
 Lung biomimetic cultures
Lung biomimetic cultures
|
|||
| Collagen hyaluronate (CHyA-B) scaffolds | Freeze-drying | Bronchial epithelium cells (Calu-3) | O'Leary et al., 2016 |
| Poly(L-lactide-co-glycolide)/Gelatin (PLGA/Gel), Poly(L-lactide-co-glycolide)/Sodium bicarbonate (PLGA-SBC) microparticles | Lung adenocarcinoma cells (A549) | Kuriakose et al., 2019 | |
| Hyaluronic Acid-Furan/ Modified methylcellulose with reactive thiols (HA-Furan/MC-SH) hydrogels | Gelation | Smooth muscle cells (SMCs) | Tam et al., 2019 |
| Poly(vinyl chloride)(PVC) sheets | Lung adenocarcinoma cells (A549) | Simon et al., 2016 | |
| Polyethylene terephthalate (PET) nanofibers | Electrospinning | Human airway smooth muscle (HASM) | Morris et al., 2014 |
| Polyethylene terephthalate (PET) scaffolds | Lung fibroblasts (MRC5) | Htwe et al., 2015 | |
| Polyurethanes/polyhedral oligomeric silsesquioxane (PU/POSS) scaffolds | 3D bioprinting | Human bone marrow mesenchymal stem cells (hBM-MSCs) | Wu et al., 2020 |
 Intestine biomimetic cultures
Intestine biomimetic cultures
|
|||
| Collagen scaffolds | Gelation | Caco-2 | Kim et al., 2014 |
| Propylacrylamide (pNIPAM) hydrogels | Caco-2, HT29-MTX, hiNSCs | Dosh et al., 2017 | |
| Silk fibroin (SF) scaffolds | Freeze-drying | Shaban et al., 2018 | |
| Manousiouthakis et al., 2019 | |||
| Human colonoid culture | Roh et al., 2019 | ||
| Collagen scaffolds | Curing/gelation | Caco-2 | Yu et al., 2012 |
| Polyethylene diacrylate (PEGDA)/Acrylic acid/Fibronectin and composite scaffolds | 3D bioprinting | Creff et al., 2019 | |
| Polyethylene diacrylate/Alginate acid (PEGDA/AA) scaffolds | UV photo-polymerization | Castaño et al., 2019 | |
| Polyethylene terephthalate (PET) nanofibrous scaffolds | Electrospinning | Patient et al., 2019 | |
 Bone biomimetic cultures
Bone biomimetic cultures
|
|||
| Ulvan/gelatin (UG) scaffolds | Gelation/Freeze-drying | Human adipose-derived mesenchymal stem cells (hADMSCs) | Tziveleka et al., 2020 |
| Alginate/Gelatin (Alg/Gel) scaffolds | 3D bioprinting | hMSCs | Zhang et al., 2020 |
| Polycaprolactone/Calcium-polyphosphate (PCL/Ca-polyP) microspheres | Human osteoblast-like cells (SAOS-2) | Neufurth et al., 2017 | |
| Alginate/Gelatin (Alg/Gel) scaffolds coated with graphene oxide (GO) | hADSCs | Li et al., 2020 | |
| Polycaprolactone/Poly(L-lactic acid)/Hyaluronic Acid (PLA/PCL/HA) scaffolds | Human osteocarcinoma cell line (MG63) | Hassanajili et al., 2019 | |
| Gelatin Methacrylate (GelMA)-VEGF hydrogels | MSCs | Byambaa et al., 2017 | |
| Keratin sponges | Casting | Human osteoblast-like cells (SAOS-2) | Bloise et al., 2020 |
| Poly(L-lactic acid)/Dimethyl sulphoxide (PCL/DMSO) scaffolds | 3-Dimensional plotting system (3DPS) | Human bone marrow stromal cells (hBMSCs) | Seok et al., 2020 |
| Poly(L-lactic acid) (PCL), Poly(L-lactic acid)/Silicate-containing hydroxyapatite (PCL-siHA) scaffolds | Electrospinning | hMSCs | Shkarina et al., 2018 |
| Poly(3,4- ethylene dioxythiophene)/Collagen (PEDOT/COLL) scaffolds | Freeze-drying | Neural crest stem cells (NCSCs) | Iandolo et al., 2020 |
| Polypyrrole crosslinked (PPY/XCS) scaffolds | hADMSCs /HUVECs | Zhang et al., 2018 | |
| Tricalcium phosphate/Alginic acid/Graphene Oxide (TCP/AA/GO) scaffolds | Polymerization/3D bioprinting | Human osteoblast cells (hOB) | Boga et al., 2018 |
| Magnesium-β-Tricalcium Phosphate Composite (Mg-TCP) scaffolds | Gelation/ 3D bioprinting | hMSCs, HUVECs | Gu et al., 2019 |
 Liver biomimetic cultures
Liver biomimetic cultures
|
|||
| Collagen gels | Gelation | Human hepatocarcinoma cells (HepG2) | Yip and Cho, 2013 |
| Glycyrrhizin /Alginate/Calcium (GL/Alg/Ca) hydrogels | Gelation/Freeze-drying | Tong et al., 2018 | |
| Chitosan/Gelatin (CS/Gel) scaffolds | Zhang et al., 2016c | ||
| Bioprinting/Gelation/ Freeze-drying | Gong et al., 2014 | ||
| Decellularized extracellular matrix (dECM)based hydrogels | 3D bioprinting | Ma et al., 2018 | |
| Gelatin Methacrylate (GelMA) hydrogels | Human hepatocarcinoma cells (HepG2/C3A)- HUVECs | Massa et al., 2017 | |
| Gelatin Methacrylate/Decellularized extracellular matrix (GelMa/dECM) bioinks | Human induced hepatocytes (hiHep) | Mao et al., 2020 | |
| Collagen/Hyaluronic Acid (COL/HA) bioinks | Activated hepatic stellate cells (aHSC) | Mazzocchi et al., 2019 | |
| Poly(L-lactide-co-glycolide)/Collagen I (PLGA/COL I) nanofibrous scaffolds | Electrospinning | Primary human hepatocytes | Brown et al., 2018 |
| Poly(ethylene glycol)/Alginate/Gelatin (PAG) cryogel matrices | Cryogelation | Human hepatocarcinoma (Huh-7), HepG2 | Kumari et al., 2016 |
| Inverted colloidal crystal (ICC) hydrogel scaffolds | UV photo-polymerization | Huh-7 | Kim M. H. et al., 2016 |
| Polycaprolactone (PCL) micro-scaffolds | Selective Laser Sintering | HepG2, HUVECs | Pang et al., 2020 |
Examples of biomaterials, fabrication methods and cell sources for skin, brain, heart, lung, intestine, bone, and liver biomimetic cultures.
Created with BioRender.com.
Blends of natural materials with other biomolecules or synthetic polymers are also commonly used for the recapitulation of the tissue milieu, as a means to overcome drawbacks related with the poor mechanical properties of some natural polymers [e.g., collagen (Ullah et al., 2018), gelatin (Han et al., 2014)], the low solubility in water [e.g., keratin (Wang et al., 2016)], their limited biostability (Pedron et al., 2013; Ryan and O'Brien, 2015), as well as source availability, and their uncontrollable biodegradation (Pradhan et al., 2020). In fact, such blends allow for enhancement of the mechanical properties and for better control over the biochemical properties of the engineered substrates according to the requirements of the tissue under development. For example, addition of elastin to porous collagen scaffolds was shown to reduce stiffness and enhance viscoelasticity, while inducing a more contractile-like smooth muscle cell phenotype (Ryan and O'Brien, 2015). Combination of collagen with HA was shown recently to yield a bioink/hydrogel suitable for 3D printing liver tissue constructs containing primary human hepatocytes and liver stellate cells, that were viable and functional for over 2 weeks and able to respond to drugs (Mazzocchi et al., 2019). In addition, PLGA nanofibrous scaffolds treated with type I collagen or fibronectin, as the minimal essential ECM components of the liver microenvironment, were able to accommodate long-term in vitro support, maintenance, and function of primary human hepatocytes (Brown et al., 2018).
Another commonly used material is gelatin-polysaccharide composite hydrogels (Afewerki et al., 2019). The chemical similarities of gelatin to the native tissue ECM, its biocompatibility, low antigenicity, cost-effectiveness, and combination with polysaccharides have been shown to produce composite hydrogels with enhanced ECM biomimicry levels, increased mechanical resilience (Afewerki et al., 2019); hydrophilicity (Jansen et al., 2005); and antimicrobial and anti-inflammatory properties (Wang et al., 2007), thus highly promising materials for 3D cell culture and TE applications (Afewerki et al., 2019). This was exemplified by the study of Guan et al., who showed that porous gelatin-chitosan scaffolds, loaded with hyaluronic acid and heparan sulfate, offer a valuable option for neural tissue engineering as they form a suitable 3D microenvironment for the adhesion, growth and differentiation of neural stem and progenitor cells (Guan et al., 2013).
Engineering Methods to Reconstitute Tissue Architecture in vitro
The choice of the most suitable biomaterial is coupled with the fabrication method, as this can also influence the final properties of the 3D matrix supporting the engineered tissue (Mabrouk et al., 2020). Various fabrication methods have been utilized so far, spanning from freeze-drying (Mabrouk et al., 2020) and physical/chemical cross-linking reactions (Hu et al., 2019), for scaffold and hydrogel preparation, respectively, to microscale technologies, such as soft-lithography for microfluidic channel fabrication (Khademhosseini et al., 2006).
Conventional scaffold fabrication approaches have relied on techniques such as freeze-drying (Figure 4A), solvent casting/particulate leaching (SCPL) (Figure 4B), melt molding and gas foaming with which different porosity levels can be achieved (Wang et al., 2006). Due to the simplicity of established protocols and the relatively low-cost and tunability of scaffold porosity and geometry, these techniques are the standard for fabricating scaffolds (Mabrouk et al., 2020). However, several drawbacks have been reported, including cases of low interconnectivity of the porous network, irregular pore sizes, use of organic solvents with possible toxic effects and lack of precise control over the overall mechanical properties (El-Kady et al., 2012; Hribar et al., 2014). In an attempt to overcome these limitations, several groups employed electrospinning (Figure 4C) to fabricate various 3D matrices from both natural or synthetic materials and blends, as this method has been shown to provide better control over the mechanical properties (e.g., porosity and tensile strength), the geometry and the micro- and nano-topography of the fibrous scaffolds (Cui et al., 2010). Htwe et al. (2015) fabricated electrospun polyethylene terephthalate (PET)-based nanofibrous scaffolds, with similar geometry to human lung extracellular matrix, to form 3D cultures of lung fibroblasts as a biologically relevant tool for the investigation of such cells in the pathogenesis of lung inflammation via activating the NF-κB signaling pathway (Htwe et al., 2015). In addition, electrospun PET-based 3D nanofibrous scaffolds, coated with collagen and mimicking the basement membrane structure, were shown to successfully support an in vitro model of the human intestinal barrier that exhibited superior performance as a drug-testing platform compared to conventional models (Patient et al., 2019).
Figure 4
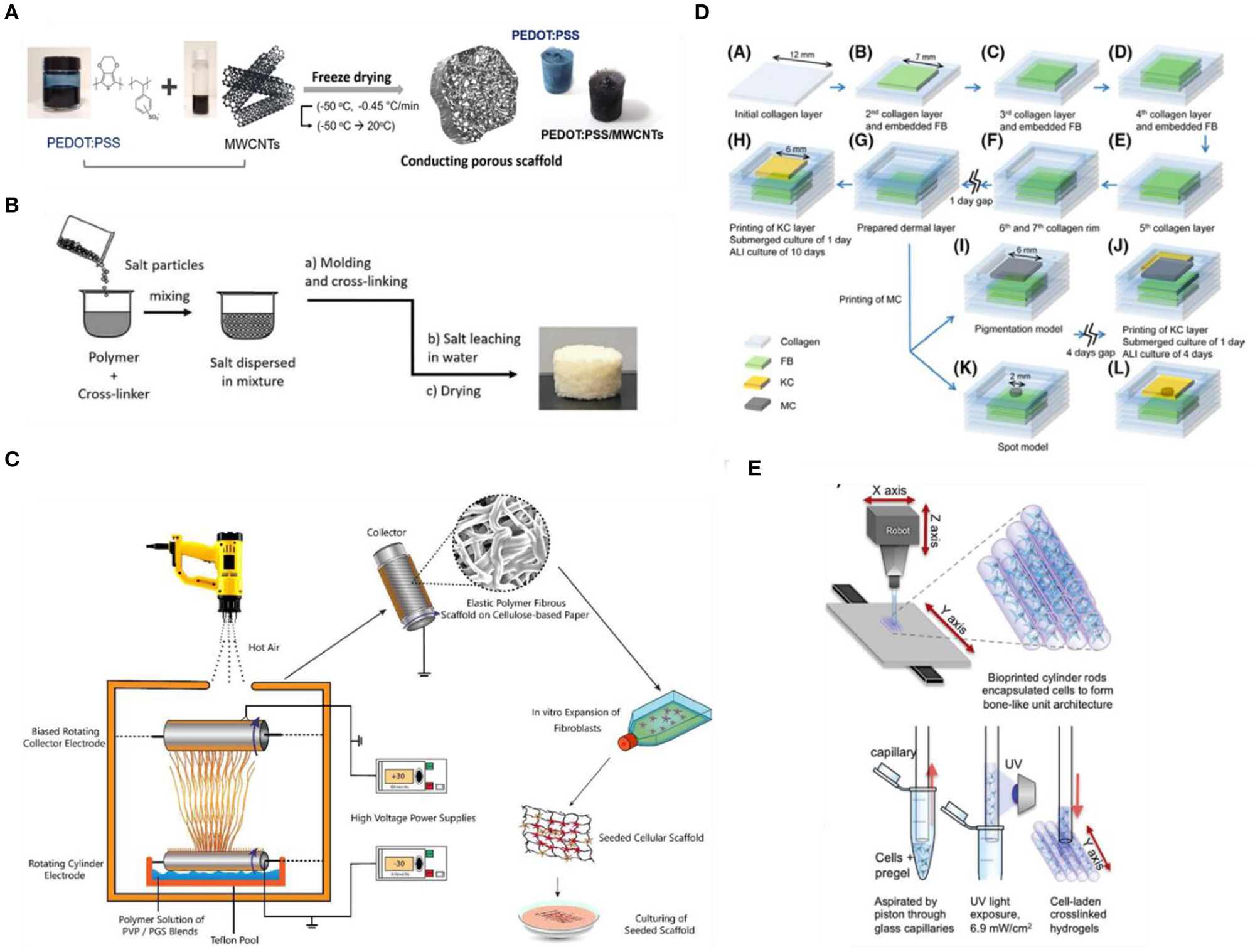
Fabrication methods for engineering 3D scaffold-based biomimetic models in vitro. (A) Fabrication process of 3D hybrid scaffolds based on PEDOT:PSS and multi-walled carbon nanotubes (MWCNT) composites via freeze-drying. Reproduced from Jayaram et al. (2019) under the Creative Commons Attribution license. (B) Preparation of porous Liquid Crystal Elastomers (LCE), as biodegradable brain tissue scaffolds, via salt-leaching. Reproduced from Prévô et al. (2018). (C) Electrospinning of Polyvinylpyrrolidone/Poly(glycerol sebacate) fibrous scaffolds for skin tissue engineering. Reproduced from Keirouz et al. (2019). 3DP techniques for engineering (D) skin and (E) bone tissue equivalents. Reproduced from Byambaa et al. (2017) and Min et al. (2018), respectively.
Over the last few years, additive manufacturing (AD) approaches have also gained a lot of attention as an alternative fabrication route to organize cells in 3D due to their potential to provide precise spatiotemporal control over biophysical and biochemical cues necessary to reproduce a biomimetic microenvironment (Murr, 2016; Bose et al., 2020; Nikolaev et al., 2020; Sun et al., 2020). AD approaches include techniques such as three-dimensional printing (3DP) (Figures 4D,E), light-assisted bioprinting (Trautmann et al., 2018), fused deposition modeling (FDM), selective laser sintering (SLS), that, along with advances in biomaterials and bioinks, enable precise deposition of materials into custom shapes and patterns to replicate complex tissue architectures, not possible using conventional techniques, and with high control and reproducibility (Melchels et al., 2012; Cui et al., 2017). Moreover, AD approaches, and specifically 3DP techniques, based on coupling multimaterial printing with high performance bioinks (i.e., hydrogel solutions that act both as cell carriers, and structural components to control and direct cell activity and fate; Chimene et al., 2020) and biomolecules have been developed to obtain highly customisable, biofunctional, and mechanically compliant scaffolds (Chimene et al., 2016). The potential of bioprinting for building such biorelevant models is highlighted in various recent studies, seeking to develop highly biomimetic and functional tissues for disease modeling and drug testing (Kolesky et al., 2014; Horvath et al., 2015; Lee A. et al., 2019; Lee H. et al., 2019; Theodoridis et al., 2019; Daly et al., 2020). However, there are still challenges and limitations to be addressed before this novel approach is fully adopted by researchers. For example, not all biomaterials are compatible with AD fabrication methods for the recapitulation of some complex micro- and nano-features, while printing modules and parameters, such as print speed, print pressure, and temperature as well as cell density in the bioink, can influence the cell-material dynamics during the printing process (Zhang and Wang, 2019). Nevertheless, as new methods for modulating biomaterial properties (e.g., new cross-linking mechanisms), along with advances in printing technologies—both software and hardware—are developed, we expect to see advances in the convergence of AD manufacturing and in vitro TE with the potential to leverage physicochemical cues and hence facilitate the development of more robust tissue equivalents.
Finally, a highly promising and popular method for building tissue equivalents, which has been favored by the emergence of the aforementioned AD manufacturing technology, is modular tissue engineering (Ouyang et al., 2020). Contrary to the top-down approach of the traditional TE paradigm, this bottom-up approach is based on fabricating living building blocks using cells (optionally together with biomolecules and/or biomaterials) which are then assembled to create more biomimetic customized tissue models (Nichol and Khademhosseini, 2009; Ouyang et al., 2020). Various methods of building and assembling these modular tissue blocks are being explored, including 3DP/bioprinting (Graham et al., 2017; Liu T. et al., 2019; Subbiah et al., 2020), micropatterning (De Gregorio et al., 2018), microfabrication of cell-ladden hydrogels (Onoe et al., 2013; Jeon et al., 2019; Figure 4E) and scaffolds (de Rutte et al., 2019), and self-assembly (Kato-Negishi et al., 2013; see Ouyang et al., 2020, for an extended review). Among other advantages, bottom-up tissue engineering approaches have gained a lot of attention because they offer a new means to generate vascularized tissues via modular assembly of pre-formed vascularized tissue blocks, a major challenge for tissue engineering at present (Nichol and Khademhosseini, 2009; Marga et al., 2012; Ouyang et al., 2020). In some cases, the versatility of these living tissue blocks allows not only for the modular assembly of vascularized tissues and organs (Homan et al., 2019), but also for the assembly of other complex tissue types (Miller et al., 2012; Magnan et al., 2020). It would be interesting to see if this strategy for vascularization of tissues could be also applied to promote innervation of biofabricated tissues and organs (Das et al., 2020).
Engineering Methods to Recapitulate the Physicochemical Properties of the Native Niche
As discussed above, progress in materials science and engineering has enabled the development of functional/smart (bio)materials and platforms for TE applications by aiding the reconstruction and control of an environment that mimics key features of the natural niche. In the in vivo situation, cells have the capacity to generate, sense, integrate, and respond to systemic and local mechano-chemical cues and through interactions with neighboring cells and the surrounding ECM, they collectively generate tissues/organs with impressive structure and functionality (Brassard and Lutolf, 2019). Therefore, besides the appropriate source of cells and the materials with the appropriate physicochemical properties and microstructure, the exposure to biochemical and biophysical cues is of paramount importance for engineering tissues in vitro (Caddeo et al., 2017; Bao et al., 2018; Chen et al., 2019). Biophysical cues include bulk properties (e.g., viscoelasticity, stiffness, and porosity), as well as surface properties (e.g., roughness, guidance cues, charge, and wetting characteristics), while biochemical cues, besides the chemical structure, and composition of the materials, also refer to the presence of gradients of nutrients, signaling molecules, or even reprogramming factors, such as mRNA (Ledo et al., 2020a). In the following section, we discuss recent advances in reconstructing several niche properties and key aspects of the natural biochemical and mechanical signals, known to influence fundamental cellular processes, as well as methods of delivering such stimulants, to simulate the chemical signaling and biological pathways of the native environment and thus promote physiological cell growth and differentiation within the engineered tissue models.
Chemical/Biochemical Stimuli
In vivo, cellular responses are influenced by various spatiotemporal biochemical signals (Caddeo et al., 2017; Park et al., 2019). Within tissues, concentration gradients for soluble components, nutrients, metabolites as well as oxygen and pH exist and are essential for exerting pressures that can stimulate or inhibit basic cellular processes (Przyborski, 2017). The proximity of vasculature and blood vessels, the diffusion of molecules through the surrounding ECM and the metabolic activity of the organ/tissue, which regulate oxygen tension, nutrient consumption and cellular waste secretion and removal, affect these natural gradients and in turn the maintenance of physiological levels of chemotaxis and homeostasis (Langhans, 2018).
Since most tissue-engineered constructs lack a vascular network, cells rely for their survival on diffusion of nutrients and oxygen through the construct (Rouwkema et al., 2009). Along with cell culture media, the engineered (bio-)materials and ECM components of the 3D tissue equivalent can act as a reservoir of such molecules, as well as for soluble components that can enhance, stimulate or inhibit specific cellular functions and guide cells toward the desired output (Caddeo et al., 2017; Afewerki et al., 2019). Essentially, cell culture media is a cocktail of molecules and compounds that range from basic nutrients necessary for cell growth to biochemical stimulants with more specialized role, depending on the needs of the cell line(s) in culture (Table 2). For example, glucose is widely used as the main source of energy for cell metabolism, while serum (e.g., fetal bovine serum) forms the basis of culture media as the cocktail of hormones, growth factors and protease inhibitors, supplemented with buffering systems, inorganic salts, amino acids and proteins/peptides that promote cell growth and viability, while also control pH and osmolality of the cell culture environment (Yang and Xiong, 2012; Carter and Shieh, 2015b; Salazar et al., 2016). A type of biomolecule in media formulations with more specialized role is growth factors (GFs). GFs are used as culture media additives due to their key role in multiple signaling pathways between cells and their environment as well as in fundamental cellular processes. For instance, bone morphogenetic proteins (BMPs) stimulate bone cell differentiation, vascular endothelial growth factors (VEGFs) stimulate blood vessel differentiation (angiogenesis), while GFs, such as epidermal growth factor (EGF), regulate a wide variety of functions in both epithelial and mesenchymal cells (De et al., 2013; Yao and Asayama, 2017). In addition to providing a biochemical link for enhancing cell communication (Gonçalves et al., 2013), addition of GFs in the cellular microenvironment is necessary for the reconstruction of the native tissue niche, where they are part of an extensive cross-talk between cell membrane receptors and ECM components (Brizzi et al., 2012). This is particularly important in the case of stem cells as well as for the derivation and maintenance of various types of organoids (Urbischek et al., 2019) the self-organization and maturation processes of which require a spatially homogeneous cocktail of specific growth factors (e.g., R-spondins and Noggin) and other signaling molecules (Brassard and Lutolf, 2019).
Table 2
| Types of biomolecules | Role | Example |
|---|---|---|
| Carbohydrates | Source of energy | Glucose Galactose |
| Amino acids | Protein synthesis, Secondary source of energy, Regulation of cell proliferation and density, Stimulation of growth and enhancement of cell viability |
L-glutamine L-cysteine L-Lysine |
| Proteins and peptides | Binding of water, salts, free fatty acids, hormones, and vitamins Removal of toxic substances, Protection against proteolysis, Promotion of cell attachment |
Albumin Transferrin Fibronectin Aprotinin |
| Growth factors | Cell signaling and communication, Mediation of processes such as proliferation, differentiation, wound healing, and tissue maturation |
Bone morphogenetic proteins (BMPs) Vascular endothelial growth factors (VEGFs) Epidermal growth factor (EGF) |
| Cytokines | Cell Signaling and communication Stimulation of cells toward differentiation pathways Modeling hematoimmune response of tissues |
Interleukins (IL) Tumor necrosis factor a (TNF-a) |
| Vitamins | Cell growth and proliferation Enzyme co-factors | Vitamin B group |
Common biomolecules in conventional cell culture media.
Another class of biomolecules used to engineer the in vitro niche and mimic native signaling networks is cytokines. Apart from their modulatory role in the hematoimmune system, cytokines produced by a broad range of cells (e.g., lymphocytes, endothelial cells, and fibroblasts),—depending on the type and state of cell—, have also been found immobilized in the ECM, forming a complex functional network within the body, exerting systemic effects that go beyond their immunomodulatory role (Morán et al., 2013). Hence, they are now being employed to engineer the microenvironment of tissue equivalents, not only as agents that enrich the cell culture media, but also as components of the (bio)material blends used to support the 3D culture system. For example, in a perfusion-based bioreactor model of human bone marrow, addition of hematopoietic cytokines (i.e., thrombopoietin, stem cell factor, and Fms-related tyrosine kinase 3 ligand) significantly aided the establishment of a xeno-free environment that in turn favored the expansion of hematopoietic stem cells (Bourgine et al., 2018), while controlled release of BMP and VEGF blended in bone-mimetic substrates was shown to exert a synergistic effect on stimulation of osteoblasts (Bao et al., 2017). Finally, it is worth noting the efforts on substituting animal derived-sera with human (Muraglia et al., 2017; Heger et al., 2018) or synthetic serum (Ejiri et al., 2015; Patel et al., 2015), as these have been shown to better support cell growth and behavior in 3D without compromising the results (Heger et al., 2018). In addition, standardization of such sera formulations will facilitate the development of completely animal-free cell systems and tissue equivalents, better capturing the native biochemical environment of specific cell types (Ejiri et al., 2015).
Physical/Biophysical Stimuli
Besides biochemical cues, each cell within the native tissu e is subject to a unique mechanical environment defined by gradients of intracellular and extracellular forces, the interactions with neighboring cells and the surrounding ECM (Caddeo et al., 2017; Brassard and Lutolf, 2019). Via mechanotransduction mechanisms, cells respond to these biophysical stresses and transduce the mechanical stimuli into biochemical signals, modifying their behavior (e.g., proliferation rate, shape, and migration). In addition, cells rearrange their cytoskeleton and cell membrane positioning and produce and exert endogenous contractile forces in the surrounding microenvironment, remodeling the ECM. This dynamic reciprocity of biophysical cues is constantly reshaping cells and the native niche structure and is associated with different cellular functions and tissue homeostasis (Xu et al., 2009; Humphrey et al., 2014; Kratochvil et al., 2019). Along with the physical properties and microarchitectural features of the tissue engineering materials, it has now been established that experimental platforms of tissue equivalents should acknowledge and incorporate the physiological biophysical variables to successfully imitate in vitro the dynamic interplay between cells and their exterior (Humphrey et al., 2014; Przyborski, 2017).
Over the last couple of decades, efforts are focusing on better understanding the effects of mechanical stimuli on cells and on addressing the challenges of reconstituting biophysical cues of physiological and diseased conditions in vitro. Of particular interest is the application of fluid shear stress that several tissues and cells within the body experience (Delon et al., 2019). For example, blood flow and pressure exert on endothelial cells one of the greatest forces within mammalian organisms (1–5 Pa) (Baeyens et al., 2016). In response to these forces, endothelial cells alter their morphology and orientation, which in turn regulates vessel physiology, function, and remodeling activity accordingly. In addition, endothelial cells transduce the frictional blood flow force into biochemical signals via specialized mechanisms that shape the ability of the vascular system to effectively perfuse all tissues. Alterations in the nature of these forces or in the mechanotransduction mechanism have been shown to contribute to major vasculature diseases (Kamiya and Ando, 1996; Kadohama et al., 2007; Baeyens et al., 2016; Chistiakov et al., 2017). Epithelial cells also experience fluid shear stress (e.g., peristalsis in the intestine), which affects both their structure and function. Several in vitro studies have revealed the importance of this mechanical cue in the formation of microvilli in the apical surface of various epithelial cell types, including intestinal (Delon et al., 2019), lung (Stucki et al., 2018), and placental (Miura et al., 2015), highlighting the importance of incorporating such biophysical cues in bioengineering applications.
The effects of biophysical cues in advancing the relevance of in vitro cellular and tissue models has also been explored in the context of stimulating cells to enhance their functional and phenotypical characteristics or to trigger the differentiation of stem cells toward the desired lineage. Mechanical stretching has long been an attractive experimental strategy for controlling cell growth, gene expression, lineage commitment, and differentiation and thus successfully engineering mechanically functional tissues, such as cardiac, lung, vasculature, and bone (Diederichs et al., 2010; Riehl et al., 2012). For example, cyclic stretch was shown to enhance the viability and functional maturation of 3D cardiac tissue constructs based on human embryonic stem cell-derived cardiomyocytes seeded on gelatin-based scaffolds (Mihic et al., 2014). In another study, Fang, et al. highlight the potential of mechanical stretch for enhancing stem cell behavior and regulating their fate. By applying cyclic stretch to human adipose-derived stem cells (hADSCs), the authors found that stretching significantly promoted the proliferation, adhesion, and migration of hADSCs, it suppressed apoptosis and adipogenesis, while it enhanced osteogenesis (Fang et al., 2019).
Besides mechanical cues, biophysical cues also involve electrical or magnetic fields, ultrasound stimulation and photostimulation (Ding et al., 2017; Chen et al., 2019). Due to the strong presence of bioelectricity (e.g., cell membrane potential, trans-epithelial potential found in all types of epithelial tissues) and its effects on in vivo systems, electrical stimulation has drawn a lot of attention for its potential benefits in tissue engineering (Balint et al., 2013; see McCaig et al., 2005, for an extended review on bioelectricity). Several in vitro studies have revealed the effects of electrical stimulation on various biological events both on cellular and tissue level, spanning from improved cellular migration and differentiation to enhanced wound healing and nerve regeneration (Vodovnik et al., 1992; Llucià-Valldeperas et al., 2015; Snyder et al., 2017; Srirussamee et al., 2019). In the recent years, these effects have also been explored in 3D culture setups. For example, Kumar et al., studied the effect of external dynamic electric field as a guiding cue for osteoblasts seeded on 3D printed porous titanium alloy scaffolds. Their findings suggest that the presence of electric field, under dynamic conditions, had a positive effect on proliferation, growth, and expression level of prominent adhesion and cytoskeletal proteins, as well as on cell-cell interactions (Kumar et al., 2016; Iandolo et al., 2020). However, the emergence of electroactive polymers as a new class of smart materials has brought to the fore the potential of combining materials suitable for TE with electrical stimulation. Early work on 2D cell culture assays based on polypyrrole, for example, has shown that such materials can support the growth, proliferation, and differentiation of mammalian cells (Zelikin et al., 2002) as well as stimulation of neurite outgrowth (Schmidt et al., 1997) or enhancement of osteogenic commitment of bone marrow stromal cells (Shastri et al., 1999) upon application of electrical fields. Moreover, carbon nanotubes (CNTs) have been shown to promote cardiomyocyte maturation (Martinelli et al., 2012) as well as to enhance the performance of engineered neurons and neural networks (Cellot et al., 2009; Fabbro et al., 2012), among other applications. Nowadays, it is well established that such materials can act both as substrates for cell attachment and tissue growth and as bioactive elements for regulating cellular activities within 3D tissue culture systems (Balint et al., 2013; Chen et al., 2019). The use of conducting materials in regulating stem cell function through electrical stimulation in 3D microenvironments was exemplified recently by co-culturing human adipose-derived MSCs (hADMSCs) and umbilical vein endothelial cells (HUVECs) in an electrically conducting polypyrrole/chitosan scaffold, demonstrating enhanced autocrine signaling, promoting the cellular functions of the co-culture system (Zhang et al., 2018). In another study, Zhu et al. developed carbon nanofibrous scaffolds with enhanced electrical conductivity and mechanical flexibility and demonstrated that sufficient support of stem cell-derived neuron-like cells, while application of biphasic electrical stimulation enhanced differentiation and maturation of these cells, as evidenced by the upregulation of the relevant neuronal biomarkers (Zhu et al., 2018). Finally, earlier this year, Iandolo et al. developed highly porous electroactive PEDOT:PSS and collagen type I composite scaffolds that supported neural crest-derived stem cell (NCSC) culture and osteogenic differentiation, without the need for scaffold pre-conditioning. The modulation of mechanical and electrical properties induced by collagen blending provided a new means for directing cell fate and response, as well as a tool for cell-based monitoring and stimulation applications (Iandolo et al., 2020).
Spatiotemporal Delivery and Control of Biochemical and Biophysical Cues in Engineered Tissue Equivalents
As discussed in previous sections, advancements in fabrication technologies and material engineering have enabled the development of tissue engineering substrates that can present cells with the necessary cues to finely elicit a plethora of cellular functions and signaling mechanisms (Leijten et al., 2017). However, our understanding of cell-material interactions so far has been based mainly on static culture systems, while the in vivo array of biochemical and biophysical signals changes over space and time. Even though mimicking natural gradients in 3D culture platforms is possible, the thickness of the culture construct, as well as the competition of cells, can limit the diffusion of nutrients, oxygen, growth factors, and other signaling molecules and due to inhomogeneous distribution, cells located in the middle of the construct might not have access to sufficient supply of those molecules and thus behave differently from cells that are closer to the engineered tissue surface (Levorson et al., 2011; Caddeo et al., 2017). In turn, this results in non-uniform cell proliferation and matrix deposition and in inhomogeneous tissue formation (Gholipourmalekabadi et al., 2016). Therefore, a lot of focus has now shifted to developing culture platforms that can dynamically recapitulate in vitro the native tissue spatio-temporal variation of signals. Such engineered platforms have the potential to facilitate better understanding and to provide more degrees of flexibility and control over cell function and fate and thus to eventually build tissues that better emulate the dynamics of the in vivo conditions (Leijten et al., 2017).
From the materials engineering point of view, even though numerous matrix-based techniques for delivering physicochemical cues, such as blending cell-adhesive ligands within scaffold materials (Gallagher et al., 2020) or micropatterning growth factors in hydrogels (Jeon et al., 2018), have been shown to improve cell behavior, this approach lacks the possibility to fine-tune and precisely control the timing of delivery, which is also important for cell survival and fate. In the case of MSCs, for instance, it has been shown that the RGD cell-adhesive motif is essential for stem cell survival at the early stages of 3D culture in PEG hydrogels, while removal of ligands at later stages does not compromise the viability of cells, but rather improves their differentiation (Kloxin et al., 2009). To overcome limitations in delivering natural cues in a spatiotemporal manner the development of dynamic biomaterials that allow for reversible modulation of the physicochemical properties and on-demand release of the desired molecules via either cell-mediated or user-mediated mechanisms, represents a promising strategy (Willerth et al., 2008; Leijten et al., 2017; Schneeberger et al., 2017; Cimmino et al., 2018; Kratochvil et al., 2019; Xu et al., 2020). A characteristic example where the delivery of physicochemical cues is mediated by cell activity is remodelable 3D hydrogels. Madl et al. showed recently that in elastin-like protein hydrogels, prior to chemically-induced differentiation, a critical amount of matrix remodeling is necessary to maintain the stemness and to enhance the differentiation capacity of neural progenitor cells into astrocytes and mature neurotransmitter-responsive neurons, via a mechanism regulated by cadherin cell–cell contacts and catenin-mediated activation of Yes-Associated Protein (YAP) expression (Madl et al., 2019). These findings highlight the potential of bioresponsive materials as attractive tissue engineering platforms that enable both expansion and subsequent differentiation of stem cells toward the desired tissue within a single cell culture setup. On the other hand, materials can be engineered to respond to user-directed stimuli as a means to fine-tune the presentation of spatiotemporal cues to cells and thus to direct and modulate interactions within the biological system (Cimmino et al., 2018). An example of this approach is the use of protease-cleavable peptides in 3D materials, as shown in a recent publication by Guo et al. The researchers, combining bio-orthogonal click chemistry and protein engineering, developed PEG-based multi-layered hydrogels with spatially-defined regions of immobilized proteins, where exogenous application of enzymes for triggering the temporal release and removal of select proteins was shown to be a promising tool for controlling cellular microenvironments (Guo et al., 2017). Due to their spatiotemporal tunability, materials responsive to light, temperature or electrical fields have also been called for, to allow triggering of the release of ECM-presented cues from a 3D substrate to the cell culture microenvironment, thus directing advanced cellular fates within tissue engineering platforms (Wylie et al., 2011; Deforest and Tirrell, 2015; Cimmino et al., 2018; Ruskowitz and Deforest, 2018; Kratochvil et al., 2019; Shadish et al., 2019).
Sophisticated bioreactor systems can be used to fine-tune culture parameters and to reproduce an in vitro tissue-specific physiological microenvironment capable of overcoming diffusion limits and oxygenation for better control or even coupled delivery of chemical and mechanical cues (Levorson et al., 2011; Hansmann et al., 2013; Schmid et al., 2018). For example, a bioreactor system developed by Zohar et al. enabled the investigation of the direct flow-induced shear stress on vascularization of poly(L-lactic acid)/poly(lactic-co-glycolic acid) (PLLA/PLGA) scaffolds and showed that flow conditions enhance vascular network formation and maturation (Zohar et al., 2018). Charoensook et al. created a bioreactor-based functional in vitro model of the neuromuscular junction by cultivation of transdifferentiated myocytes and stem cell-derived motoneurons, where electrical stimulation resulted in improved maturation and function of motoneurons and myocytes, as well as exhibiting physiological response to drugs, thus suggesting its potential as a pharmacological screening platform and controlled studies of neuromuscular diseases (Charoensook et al., 2017). Furthermore, an all-in-one bioreactor approach facilitated the reconstruction and control of a more physiologically relevant 3D cardiac tissue microenvironment by combining, within a single chamber, electrical stimulation of the cardiac tissue, bidirectional interstitial fluid flow and on-line monitoring, and analysis of tissue functionality during culture (Visone et al., 2018).
One of the most promising technologies for bridging the gap between in vitro and in vivo systems is organs-on-chips (OOCs), alternatively called microphysiological in vitro models. OOCs technology has emerged from the combination of recent advances in microengineering and fluidic physics with trends in growing cells in 3D, allowing for the development of models that more faithfully recapitulate key features of specific human tissues and their interactions (Ramadan and Zourob, 2020). The design of the vast majority of OOC models are based on (micro-)fluidic devices, fabricated by soft-lithographic techniques, with continuously perfused chambers inhabited by living cells arranged in a biomimetic manner, while facilitating precise control over delivery of nutrients and spatiotemporal tuning of oxygen and pH gradients (Bein et al., 2018; Ronaldson-Bouchard and Vunjak-Novakovic, 2018). Among the added benefits of using such systems are the continuous supply of nutrients and removal of waste, the unparalleled, independent control over multiple key factors of the cell system, the possibility for in situ, high precision and automated monitoring and sample analysis, as well as the ability to interface different cellular compartments for enhanced cell cross-talk and exchange of signaling molecules and growth factors. But, what creates an enormous potential for enhancing the physiological relevance of the in vitro cell systems, spurring new, unforeseeable applications of this technology, is the combination of a biomimetic niche with accurate, precise, and coupled delivery of more complex biochemical and biophysical cues (Bein et al., 2018; Ramadan and Zourob, 2020). Even though most of the attention OOCs have gained is focused on pharmacology and pre-clinical drug screening applications, as low-cost and animal-free alternative tool (Ramadan and Zourob, 2020), it is clear that the principle behind this technology lines perfectly with the TE paradigm and scope: convergence of cells with the advanced chip technology-biomaterials and delivery of physiologically relevant cues toward more robust tissue equivalents. As a result, various research groups around the world, both in the academic and industrial sectors, have developed a broad range of OOCs, mimicking the human gut (Ramadan and Jing, 2016; Kasendra et al., 2018; Shin et al., 2019), liver (Delalat et al., 2018; Jang et al., 2019), kidney (Jang et al., 2013; Chang et al., 2017; Yin et al., 2020), lung (Huh et al., 2012; Stucki et al., 2018; Felder et al., 2019), blood-brain barrier (Kilic et al., 2016; Wevers et al., 2018), bone (Marturano-Kruik et al., 2018), and vasculature (Schimek et al., 2013; Jeon et al., 2015) in both healthy and pathophysiological conditions, such as infection (Villenave et al., 2017; Ortega-Prieto et al., 2018) and cancer (Ayuso et al., 2016; Hassell et al., 2017; Hao et al., 2018; Carvalho et al., 2019), as well as for the interaction of multiple organs, as first showcased by Shuler et al. and more recently by others, toward multi-organ and whole-body microsystems (Miller and Shuler, 2016; Vernetti et al., 2017; Edington et al., 2018; Herland et al., 2020), to study collective responses to drugs or disease inducing agents and inter-organ communication.
Applications of 3D Biomimetic Cultures and Tissue Equivalents
Modeling Human Health and Disease
The transformation of the field toward more in-vivo-like human models is still ongoing, facilitated by the technological advancements in 3D cell platforms. Their deployment in early biomedical applications has already given prominence to the usefulness and validity of such models, leading to a paradigm shift in our understanding of human cell and tissue biology (Chen, 2016; Przyborski, 2017). Several studies over the past decades have showcased the powerful potential of 3D in vitro models in addressing questions specific to the human biology that are challenging, if not impossible, to answer with animal models and conventional biological assays. Additionally, increasing evidence suggests that such models are highly valuable for modeling pathophysiological conditions to study the disease onset and progression mechanisms, to identify pathogenic factors and potential therapeutics in the pre-clinical and clinical level (Chen, 2016).
As discussed earlier, the advent of human organoids as systems strikingly similar to the in vivo environment has provided researchers with the unique opportunity to recreate the human physiology and architecture in vitro (Kim et al., 2020). A study that laid the foundation for the use of organoids for biomedical applications, was the generation of cerebral organoids in 2013. Lancaster and Knoblich used hiPSCs to establish an organoid culture system with various discrete, independent brain regions, including the cerebral cortex with progenitor populations, that give rise to mature and functional cortical neuron subtypes, recapitulating the formation of neural tissue in human brain development stages. In addition to this healthy brain tissue, they used patient-specific iPSCs and developed cerebral organoids that bore clear characteristics of the microcephaly neurodevelopmental disorder. This system helped them to address questions about the disease in a way that would not have been possible by growing neurons in flat surfaces or by engineering mouse models, due to lack of the necessary niche and interspecies differences, respectively (Lancaster et al., 2013; Figure 5A). Since then, organoids have played a prominent role in enhancing our understanding of various biological phenomena, including development and organogenesis (Karzbrun et al., 2018; Trisno et al., 2018; Vyas et al., 2018; Rossi et al., 2019; Shi et al., 2020), infectious biology (Forbester et al., 2015; Leslie et al., 2015; Karve et al., 2017; Heo et al., 2018; Lamers et al., 2020), cancer (Li et al., 2018; Nagle et al., 2018; Fusco et al., 2019; Ooft et al., 2019), and other diseases. Among others, the recent study by Sachs et al. on long-term expanding lung organoids clearly shows the versatility of these experimental tissue formats in faithfully recapitulating the adult epithelial airway structure and function from both healthy individuals and from patients with cystic fibrosis, lung cancer, and viral infections (Sachs et al., 2019). Finally, the valuable contribution of organoids in disease modeling is also showcased by the great efforts to build novel biomedical resources where samples from minimal amounts of tissue biopsies are used to generate patient-derived organoids, stored for future research, known as living biobanks (Sachs et al., 2018; Yan et al., 2018; Kim et al., 2020; Nelson et al., 2020; Simpson et al., 2020). The potential of such biomedical resources for modeling disease and investigating therapeutic strategies, even at the personalized level, is nicely exemplified in a study published earlier this year that presents methods for generating and biobanking patient-derived glioblastoma organoids (GBOs) that recapitulate and preserve the cellular and mutational diversity of the corresponding glioblastomas, reflecting inter- and intra-tumoral heterogeneity, holding great promise as a precision medicine tool for diagnosis and treatment (Jacob et al., 2020).
Figure 5
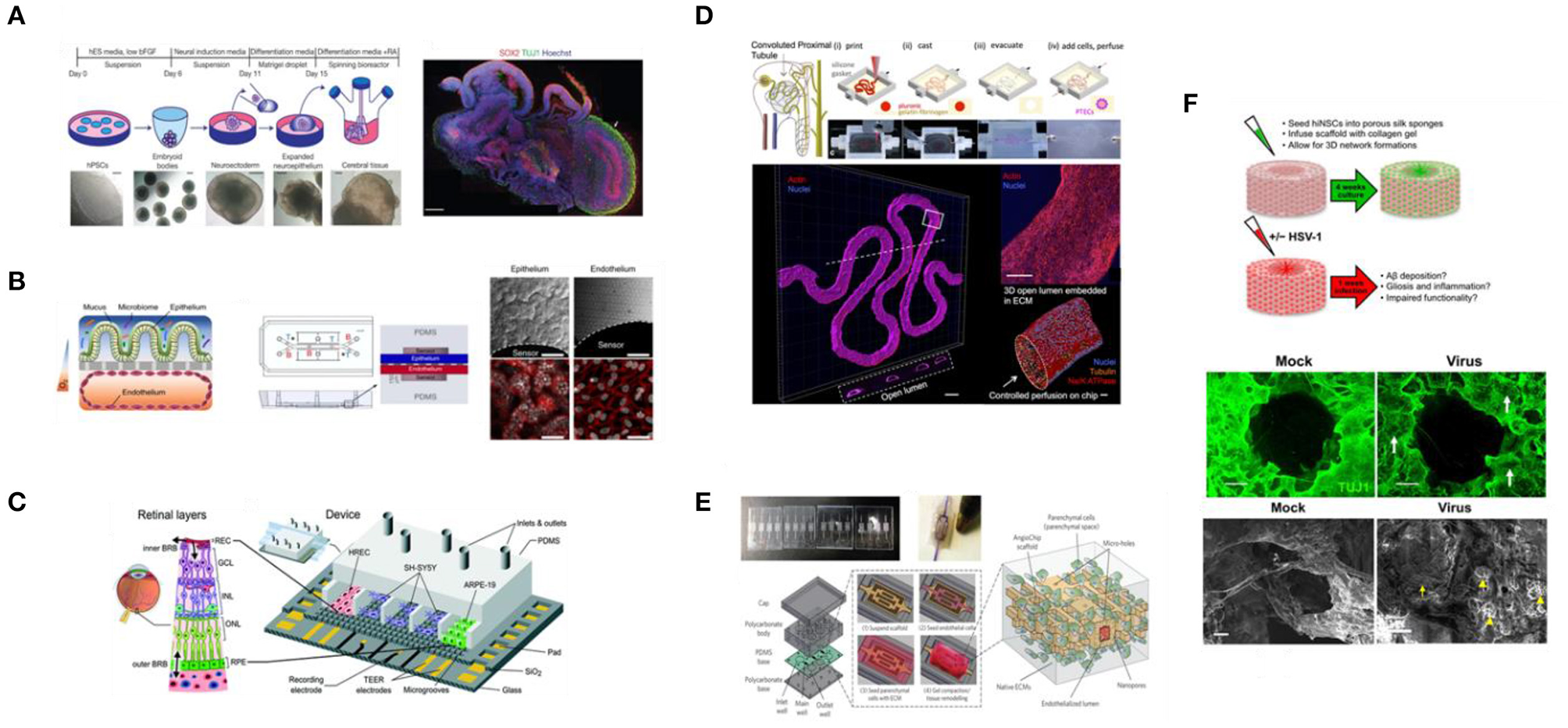
Biomimetic models of human (patho-)physiology. (A) Schematic representation of the cerebral organoid generation method (left) and immunohisto- chemistry image of a cerebral organoid section, revealing its complex morphology with regions containing neural progenitors (SOX2, red) and neurons (TUJ1, green) (right). Adapted from Lancaster et al. (2013). (B) Schematic illustration of the two-channel microfluidic Organ Chip device with an oxygen gradient (color scale) (left), schematic representation of the Intestine Chip with its embedded oxygen sensing unit (middle), and microscopy images of the Intestine Chip showing the morphology of the intestinal epithelium and endothelium morphology in the respective channels (right). Adapted from Jalili-Firoozinezhad et al. (2019). (C) Schematic illustration of a microfluidic chip culture device for modeling the blood-retinal barrier, with integrated TEER measurement electrodes. Reproduced from Yeste et al. (2018). (D) Schematic of the nephron convoluted proximal tubule and illustration of the bioprinting process steps, with corresponding images, in the fabrication of 3D convoluted perfusable proximal tubules (top), confocal microscopy 3D renderings of the bioprinted convoluted renal proximal tubule (bottom). Reproduced from Homan et al. (2016) under the Creative Commons CC BY license. (E) Images of the AngioChip scaffold (top) and schematics of the assembly of the bioreactor and of the vascularized tissue (bottom and right). Reproduced from Zhang et al. (2016a). (F) illustration of a 3D human brain-like model for modeling HSV infection (top), Images of the brain model showing b-III tubulin (TUJI1), and beta amyloid (Ab) immunostaining (middle), and scanning electron micrographs revealing the effects of HSV-1 infection on the brain-like tissue constructs (bottom). Adapted from Cairns et al. (2020) under the Creative Commons Attribution-Non Commercial license.
In parallel, the growing body of literature over the past years reflects the tremendous effort and progress in bioengineering tissue equivalents that allow for interfacing with vasculature and enable more in-depth investigation of tissue structure, homeostasis and pathophysiology, as well as communication between tissues and their surroundings. As discussed in the previous sections, successful engineering of tissues in vitro requires careful consideration of the cell sources, the type of material and the fabrication method as well as of the integration of the relevant biophysical and biochemical cues, all of which highly depend on the native tissue structure and function and on the biological question to be addressed with the model. For example, the delineation of tissue compartments and the regulation of the passage of ions and solutes by epithelia and endothelia are of particular importance for the recreation of such barrier tissues. While early strategies, relied on permeable supports to separate the apical and basal compartments of the barrier-forming monolayers (Lea, 2015; Pearce et al., 2018), microphysiological systems and OOC approaches are taking this concept a step further, by exploiting novel bioengineering techniques to create compartmentalized barrier models with a more biomimetic interface between endothelial and epithelial tissues, combined with delivery of essential environmental cues (Bein et al., 2018). The work of Huh et al. was among the first attempts to this end. The researchers created a “lung-on-a-chip” that models human lung function in both normal and disease states, by co-culturing microvascular endothelial cells (i.e., bottom, microvascular compartment) and alveolar epithelial cells (i.e., top, alveolar compartment) in parallel microchannels separated by a thin semipermeable membrane, that also enabled the establishment of an air-liquid interface environment, where cell culture medium was perfused via the microvascular channels while the alveolar channel was filled with air. Cyclic stretch of the tissue via application of vacuum to the side compartments of the parallel channels was shown to successfully mimic the breathing motion of the lung, while concurrent administration of the cytokine interleukin-2 (IL-2) to the microvascular channel was shown to compromise the barrier, reproducing the pulmonary leakage, which is a characteristic symptom of the pulmonary oedema (Huh et al., 2012). Versions of human gut-on-a-chip were also developed with the same device, where the side vacuum chambers and the application of fluid flow were exploited to recreate the peristaltic motions, but were also shown to induce spontaneous formation of the characteristic intestinal villus-crypts structures (Kim and Ingber, 2013). More recently though, this system was modified to accommodate the development of a co-culture of mucus-producing human primary intestinal epithelium in the same compartment with stable communities of the human gut microbiota by applying a hypoxia gradient across the endothelium-epithelium interface, while simultaneously monitoring oxygen levels and intestinal barrier function (Jalili-Firoozinezhad et al., 2019; Figure 5B). In a different approach, Trietsch et al. developed their intestine-on-a-chip model in a membrane-free manner, where a functional intestinal barrier is formed in a lateral channel of a microfluidic chip, at the interface with an ECM gel that supports the basal side of the epithelium and facilitates access to the perfusion channel on the opposite side of the intestinal tubules (Trietsch et al., 2017). Another way to achieve compartmentalization and tissue-tissue interfaces within OOC systems, bypassing the use of membranes, was proposed by Yeste et al. who developed a microfluidic device where cells grow in parallel compartments that are highly interconnected via a grid of microgrooves, facilitating heterotypic cell-cell contact, and paracrine signaling, while integration of electrodes allows for in-line monitoring of the cell barrier integrity in each compartment. The device successfully supported the generation, maintenance and monitoring of a blood-retinal-barrier model, based on the co-culture of primary retinal endothelial cells (HREC), a human neuroblastoma cell line (SH-SY5Y), and a human retinal pigment epithelial cell line (ARPE-19) (Figure 5C), highlighting the necessity of compartmentalization in enhancing the robustness of barrier models as well as the added benefits of integrating in-line monitoring units (Yeste et al., 2018). OOC technology has also significantly contributed in dissecting in vitro the mechanisms behind various (patho-)physiological mechanisms of the brain (Haring et al., 2017). Herland et al., for instance, were able to study human neurovascular function and inflammation in a 3D model of the human blood-brain barrier (BBB) within a microfluidic chip and to reveal the distinct contributions of astrocytes and pericytes to neuroinflammation (Herland et al., 2016). Kilic et al. proposed a brain-on-chip model suitable for studying the migration of human neural progenitors in response to chemotactic cues (Kilic et al., 2016), while Park et al., proposed a microfluidic device for the brain-like interstitial perfusion of neurospheroids and tested the toxicity of amyloid-β, showcasing the validity of brain-on-chips in modeling and studying neurodegenerative diseases as well (Park et al., 2015).
Although OOC models have been shown to better capture the in vivo situation, compared to conventional 2D culture formats, often such models are quasi-3D, forming an intermediate stage between 2D and 3D cell culture microenvironments rather than truly biomimetic tissues. To overcome this limitation, OOC technologies are now going beyond 2D, utilizing gels and scaffolds as tissue growth templates, toward generating 3D tissue equivalents of high biomimicry (Terrell et al., 2020), as shown recently in reports on the fabrication of 3D convoluted, luminal tissue architectures (Massa et al., 2017; Manousiouthakis et al., 2019; Wang X. et al., 2020). For instance, 3D human renal proximal tubules were engineered via combined bioprinting with 3D cell culture and OOC principles. The tissue construct was housed in perfusable chips and embedded within an extracellular matrix that supported the active perfusion, growth, differentiation, and maintenance of the proximal tubule epithelium for over two months, during which the nephron-like tissue exhibited significantly enhanced epithelial morphology and functional properties (e.g., brush border, basement membrane protein deposition, basolateral interdigitations, enhanced cell height, megalin expression, and albumin uptake), as well as in-vivo-like response upon delivery of nephrotoxin and cyclosporine A (Homan et al., 2016; Figure 5D). Robust 3D vascular models that more faithfully capture the natural milieu have also been called for interfacing with different tissue equivalents (Morgan et al., 2013; Kolesky et al., 2016). In an attempt to overcome the challenges in the choice of material for vasculature engineering as well as to address the challenge of co-cultivating parenchymal cells with vasculature in 3D, Zhang et al. created the AngioChip. This is a stable biodegradable scaffold that consists of a perfusable 3D, branched, luminal microchannel network with thin, flexible but yet mechanically compliant walls, lined by endothelial cells, and surrounded by a tunable matrix that supports the assembly of parenchymal cells. In addition, the walls feature nano-pores and micro-holes that were shown to enhance permeability, to facilitate efficient molecular exchange, intercellular crosstalk (Figure 5E), as well as extravasation of monocytes and endothelial cells upon biomolecular stimulation, as showcased via the successful vascularization of hepatic and cardiac tissues. The precise placement of endothelial and parenchymal cells, in a simple to operate format, the control of the initial architecture of the vasculature as well as the potential to fine-tune the vessel permeability to match the requirements of different organ models, highlight the AngioChip as a versatile tool for cultivating the vasculature in tissue engineering platforms (Zhang et al., 2016a). Progress in modeling the human brain has also been made with the emergence of in vitro TE technology. Researchers have long studied the multi-layered, hierarchical brain architecture and complex physiology, but recapitulating the entire brain in vitro still remains a huge challenge, in part because of this complexity but also due to the complications associated with the available technology (Lozano et al., 2015). Novel cell sources combined with new materials and technological platforms have yielded new tools for building functional brain-like tissues, pushing further the borders of our understanding of the human brain. This is also particularly important for neurodevelopmental and neurodegenerative studies where translation of findings to the clinic is hindered by interspecies differences (e.g., cognition), among other factors (Hackam and Redelmeier, 2006; Hartung, 2013). For instance, in a pioneering study Tang-Schomer et al. developed a 3D brain-like cortical tissue construct using primary cortical neurons in a silk fibroin/collagen gel composite scaffold as a support for the 3D axon connections, that was able to reproduce the compartmentalization of gray and white matter as well as the in vivo relevant biochemical and electrophysiological outcomes necessary for the assessment of both brain physiology and brain disorders (Tang-Schomer et al., 2014). Optimization of this silk-fibroin scaffolds, along with the development of a technique to generate expandable and rapidly differentiating human-induced neural stem cell (hiNSC) lines, has enabled this group to further advance their approach in building 3D brain tissue equivalents that allow for long term studies of neural tissue in various conditions, such as neurodegeneration, brain tumors and injury (Chwalek et al., 2015; Cairns et al., 2016; Cantley et al., 2018; Sood et al., 2019; Rouleau et al., 2020). A remarkable application of this system though was reported earlier this year where the implication of herpes simplex virus type I (HSV-1) as a causative agent of Alzheimer's Disease was investigated. The 3D bioengineered brain model was able to reproduce the herpes-induced AD pathophysiology, encompassing features of the in vivo physiological human disease, including Aβ plaque formation, neuronal loss, reactive gliosis, neuroinflammation, and diminished neural network functionality, reflecting the great progress and the tremendous potential of 3D TE approaches to address the critical need in building robust and physiologically relevant 3D human tissues (Cairns et al., 2020; Figure 5F).
Drug and Treatment Development
3D biomimetic systems technology facilitates not only elucidation of disease biology and deeper understanding of the onset and progression mechanisms, but also identification and screening of potential drug candidates and therapies. Implementation of these systems in various stages of drug discovery and development is considered as a powerful alternative for addressing the challenges associated with the poor predictive power of existing preclinical models that, more often than not, is responsible for the high attrition rates of clinical trials (Roth and Singer, 2014; Caddeo et al., 2017; Ronaldson-Bouchard and Vunjak-Novakovic, 2018). 3D biomimetic cultures in the format of spheroids, organoids, hydrogels and scaffolds that more closely capture the disease physiology have already demonstrated the validity of this technology for drug development, screening, and toxicology assays (Sobrino et al., 2016; Wan et al., 2016; Villasante et al., 2017; Plummer et al., 2019; Han et al., 2020). However, OOCs in particular have gained a lot of attention for their potential to produce more predictive and accurate data by resolving the discrepancies in drug safety and efficacy observed between models currently used in preclinical and clinical stages of drug testing and hence accelerate the translational process (Haring et al., 2017; Low and Tagle, 2017; Ronaldson-Bouchard and Vunjak-Novakovic, 2018). The precise control of multicellular activities, the spatiotemporal distribution/delivery of relevant cues and the interface between different tissue types that OOCs offer has been exploited for the evaluation of novel anticancer therapies (Sontheimer-Phelps et al., 2019), among others (Ribas et al., 2016; Mittal et al., 2018). For example, Bai et al. demonstrated that a microfluidic platform, interfacing lung or bladder carcinoma aggregates with vessel-like structures, can serve as an in vivo-like surrogate for anti-invasive and anti-metastatic drug screening, revealing the role of signaling pathways involved in the drug action mechanism (Bai et al., 2015). More recently, a-tumor-on-a-chip platform, where the efficiency and toxicity of gemcitabine-loaded nanoparticles on Matrigel-embedded human colon cancer cells in contact with a 3D vessel-like colonic endothelium was successfully evaluated, was proposed as a precision onco-nanomedicine tool (Carvalho et al., 2019; Figure 6A). Despite the fact that oxygenation, a highly important parameter for the cancer microenvironment, is not taken into account in terms of monitoring and evaluation, both studies reflect the important developments made toward more accurate and precise screening of cancer therapeutics utilizing OOCs.
Figure 6
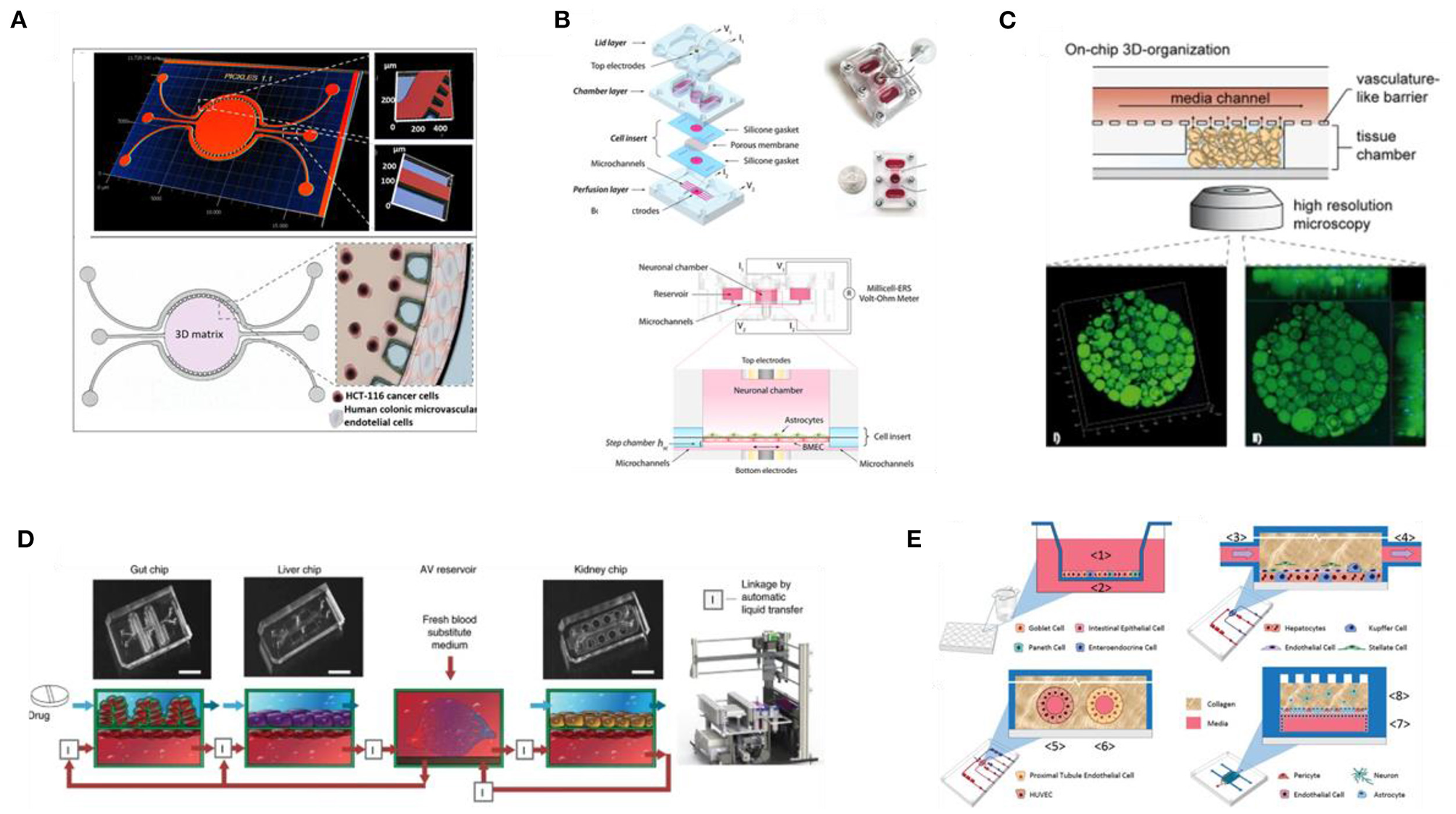
Novel in vitro human models for drug screening and testing applications. (A) 3D map image of the colorectal tumor-on-a-chip system features (top) and schematic of the chip design and culture setup (bottom). Adapted from Carvalho et al. (2019) under the Creative Commons Attribution-Non Commercial license. (B) Schematic illustration of a BBB-on-a-chip platform and assembly steps (top left), image of the assembled device (top right), cross-section of the assembled platform (middle), and zoom-in panel illustrating the device and cell culture setup (bottom). Reproduced from Wang et al. (2017). (C) Design of the human WAT-on-a-chip confocal image of the tissue construct generated in the device. Reproduced from Rogal et al. (2020) under the Creative Commons CC BY license. (D) Design and photographs of a first pass multi-organ chip system (top) and diagram of the fluidic coupling of the gut, liver, and kidney chips, containing vasculature compartments, and of the arteriovenous (AV) reservoir via an automated liquid handling instrument. Reproduced from Herland et al. (2020). (E) Schematic representation of the four organ systems used for the functional coupling of a human microphysiology system: intestine, liver, vascularized kidney, and BBB, including the neurovascular unit. Reproduced from Vernetti et al. (2017) under the Creative Commons Attribution International License.
OOC platforms are also highly relevant for testing drug permeability and transport across the blood-brain-barrier (BBB). This distinctive tissue structure is made up of neurons, astrocytes, oligodendrocytes, microglia, smooth muscle cells, brain epithelial and endothelial cells, and pericytes embedded in the brain extra-cellular matrix (ECM), and bears the responsibility of maintaining brain homeostasis by supporting the neuronal activity and by tightly regulating the passage of metabolites, drugs, and other solutes from the peripheral blood into the central nervous system (Griep et al., 2013; Wang et al., 2017; Vatine et al., 2019). As OOC approaches allow for better compartmentalization and efficient assembly of the corresponding cellular components and environmental cues, thus more realistic reconstitution of the native tissue (Booth and Kim, 2012; Adriani et al., 2017; Wang et al., 2017; Campisi et al., 2018; Vatine et al., 2019), their utility in modeling various features of the BBB and their implementation in preclinical drug evaluation studies has gained a lot of attention recently, particularly for testing whether a drug designed to treat neurological diseases can actually cross the BBB to reach its target (Ronaldson-Bouchard and Vunjak-Novakovic, 2018). This was showcased by Wang et al. who proposed the fabrication of a BBB–on-a-chip with integrated TEER sensors for in situ monitoring of the barrier tissue integrity (Figure 6B). Even though the authors utilized primary astrocytes derived from rats, in a co-culture with human iPSC-derived brain microvascular endothelial cells (BMECs), their BBB microfluidic chip was able to generate drug permeability data comparable with in vivo values (Wang et al., 2017). In a different application, a compartmentalized, multilayer OOC device was shown to successfully maintain and monitor functional human white adipose tissue and fatty acid metabolism while also being applicable for testing tissue responsiveness to therapeutic compounds, useful for diabetes, and obesity studies (Rogal et al., 2020; Figure 6C).
Ultimately, what renders 3D biomimetic tissues and particularly OOCs even more attractive alternatives for drug and therapy development is their ability to be linked via their endothelium/vascular channel, in a way that mimics the drug distribution within the body, toward multi-organ systems for simulating pharmacokinetic and pharmacodynamic drug responses (Prantil-Baun et al., 2018; Ronaldson-Bouchard and Vunjak-Novakovic, 2018). This possibility has been recently explored by various studies, where different tissues were fluidically coupled to model the in-vivo-like organ interactions (Zhang et al., 2009; Maschmeyer et al., 2015a,b; Tsamandouras et al., 2017). A very recent characteristic example of such a multi-organ system was reported earlier this year, comprising organ-on-chip models of the gut, liver and kidney. The models were linked by their endothelium-lined channels through a robotic system circulating a common blood substitute that represents the systemic circulation, stored in a fluid-mixer reservoir that represents the arteriovenous (Figure 6D). The system was then used to model first-pass absorption, distribution, metabolism, excretion and toxicity of nicotine and cic-platin and quantitatively predict pharmacodynamics, and pharmacokinetic parameters, with results matching clinical data (Herland et al., 2020). In the same concept, Vernetti et al. coupled human MPS models representing the major absorption, metabolism and clearance organs (the jejunum, liver, and kidney) along with skeletal muscle and neurovascular models and evaluated the organ-specific processing, pharmacokinetic, and toxicity of terfenadine, trimethylamine (TMA)—as a potentially toxic microbiome metabolite—and vitamin D3. Their findings were consistent with clinical data, while they also discovered that trimethylamine-N-oxide (TMAO) can pass through the BBB (Vernetti et al., 2017; Figure 6E).
Without a doubt, this technology has the potential to transform drug discovery and development. However, there are still several challenges that should be addressed before the field realizes its tremendous potential. For example, with a few exceptions, most scaffold/hydrogel-based 3D models and OOCs primarily utilize cell lines, which in some cases may drift from the normal genotypic and phenotypic profile of the tissue of origin (Carter and Shieh, 2015a). Populating next-generation models with more relevant cells, such as iPSCs and organoid-derived patient specific cells can help overcome this limitation and better capture the tissue phenotype and thus to better mimic drug responses. This should be in line with the biological question the model is trying to address, recapitulating the specific features of the pathophysiology of interest. In addition, current OOC and MPS systems are fabricated predominantly with polydimethylsiloxane (PDMS), due to its ease-of-use, elasticity, optical transparency, and low-cost microfabrication. But issues related to the absorption of small hydrophobic molecules by this material severely compromise the validity of such systems in drug screening studies, pushing for transition to alternative, non-absorptive materials (Campbell et al., 2020). Also, 3D tissue equivalents and OOCs, as more sophisticated and multi-parametric models, require close control and synchronism of these different parameters to achieve the necessary functionality, particularly when long-term viability is the case (Sontheimer-Phelps et al., 2019). Moreover, in multi-organ systems, besides in-vivo-like sequential coupling of each counterpart, scaling must also be taken into account, to match flow volume and rate with cultured tissue mass in order to mimic the native conditions and to achieve the required level to support functionality (Wikswo et al., 2013; Bhatia and Ingber, 2014; Ronaldson-Bouchard and Vunjak-Novakovic, 2018; Ramadan and Zourob, 2020). Finally, for this technology to live up to its potential and to be successfully implemented in the drug development process, it is necessary to move from the proof-of-concept laboratory models toward more widely available prototypes to facilitate high-throughput screening and further validation that the models effectively mimic in vivo drug responses. Since OOCs is a multidisciplinary field, the overall progress of this technology and its implementation in drug development, among other applications, heavily relies on parallel advancements in the field of cell biology, materials, microengineering, and microfluidics. Toward this direction, the introduction of commercially available systems is already taking place, promoting the production of more automated, user-friendly OOCs, and will definitely help resolve current limitations (Zhang and Radisic, 2017). Nevertheless, to more efficiently employ this technology, more deep understanding of cell-material and cell-cell interactions and functions, as well as better understanding of the effects of biochemical and biophysical stimuli on the overall tissue structure and function is required.
Sensing and Monitoring
As discussed during the previous sections, novel technologies for 3D TE in vitro offer unprecedented control over various parameters for the development and maintenance of the tissue equivalent of interest. In addition to fine-tuning the biological system parameters, this technology allows for the integration of in-line sensors that report not only on system parameters (e.g., flow rate, O2 levels, and pH), but also provide feedback on cellular activity, thus facilitating the study of a broad range of physiological phenomena (Bhatia and Ingber, 2014). Despite their inherent limitations, the majority of current approaches rely heavily on optical transducers, such as fluorescence microscopy, which is more a qualitative assay, that requires labeling (Kim S. et al., 2016; Sobrino et al., 2016), combined with downstream analysis of effluents to detect changes in gene expression and metabolite production (Curto et al., 2017; Roh et al., 2019). The use of invasive probes and the terminal nature of these assays, which sometimes requires harsh and lengthy protocols for sample preparation prior to imaging and analyzing, can be bypassed by advanced imaging techniques. For example, Raman spectroscopy has emerged as a suitable tool for non-invasive, in situ quality control of cells and substrates, as well as for real-time monitoring of physiologically relevant metabolites (Pudlas et al., 2011; Zbinden et al., 2020). Two-photon excitation microscopy is also being employed now as an alternative to confocal microscopy for 3D and deep tissue imaging, obviating the need for sectioning of 3D tissue-engineered constructs that sometimes can compromise the sample quality (Gioiella et al., 2016; Hume et al., 2018).
Besides new imaging assays, electrical transducers have been also shown to provide a wealth of real-time information through non-invasive and dynamic interfacing with biological systems (Rivnay et al., 2018). A well-established tool for rapid monitoring of cells in vitro, especially for drug toxicology studies, is Electrochemical Impedance Spectroscopy (EIS), with which it is possible to obtain information about cell adhesion, proliferation, and differentiation over time. In fact, EIS is widely used for monitoring Transepithelial/Transendothelial Electrical Resistance (TEER), a commonly used parameter to quantitatively characterize the function and integrity of tissue barriers with fast, non-invasive measurements (Benson et al., 2013; De León et al., 2020). van der Helm et al. recently developed an intestinal OOC with integrated electrodes that allowed for transepithelial barrier function and tissue differentiation monitoring via impedance spectroscopy, while combination with electrical simulation showed that this method can be adapted within any organ-on-chip to better monitor cell activity and to also enable comparisons between different platforms (van der Helm et al., 2019; Figure 7A). Integration of electrical transducers has also been reported recently in 3D biomimetic cultures (Pas et al., 2017; Zhang et al., 2017; Curto et al., 2018; Jahnke et al., 2019; Li et al., 2019). Kalmykov et al., for example, developed an “organ-on-e-chip” where they interfaced human cardiac spheroids with 3D self-roll biosensor arrays, which operated either as microelectrodes for EIS monitoring or as field-effect transistors, enabling acquisition of continuous multiplex recordings that allowed for real-time monitoring of cardiac tissue maturation (Kalmykov et al., 2019; Figure 7B). A high-density multi-electrode array was also proposed for real-time and automated impedimetric monitoring of cell migration out of human breast microtumours (Jahnke et al., 2019; Figure 7C). Despite the great progress made, most of these models fail to achieve intimate electrode-cell coupling which is necessary to accurately record a signal, since they utilize electrodes that are designed for planar culture of cells and are thus ill-adapted for monitoring complex 3D tissues (Inal et al., 2017; Jahnke et al., 2019). To overcome this limitation, conducting polymer scaffolds can be used instead, as evidenced recently by their remarkable performance as tissue building blocks (Wan et al., 2015; Guex et al., 2017; Iandolo et al., 2020). However, what makes these materials more attractive as TE substrates is that in addition to more seamless integration with complex cell cultures, they also allow for more intimate cell-electrode coupling, necessary for accurate signal transduction, and hence more effective monitoring of cell status and tissue formation (Inal et al., 2017; del Agua et al., 2018; Jayaram et al., 2019). We recently explored the potential of these materials for in vitro TE applications and organ-on-chip platforms. In particular, we fabricated tubular 3D macroporous electroactive scaffolds, based on the conducting polymer poly(3,4-ethylenedioxythiophene) doped with poly(styrene sulfonate) (PEDOT:PSS), the electrical, mechanical and biochemical properties of which we were able to fine-tune. We then integrated these tubular PEDOT:PSS scaffolds into a transistor configuration (i.e., transistor in a tube: Tubistor) and showed proof of principle for continuous monitoring of a simple 3D co-culture of mammalian cells over a period of 4 days. The real-time electrical readouts, cross-validated with optical analysis, enabled us to closely monitor cellular activity and even distinguish between cell adhesion and barrier tissue formation in a non-destructive, label-free manner, highlighting the added benefits of integrating in-line sensing components within engineered tissues for building more robust and sophisticated TE models (Pitsalidis et al., 2018; Figure 7D).
Figure 7
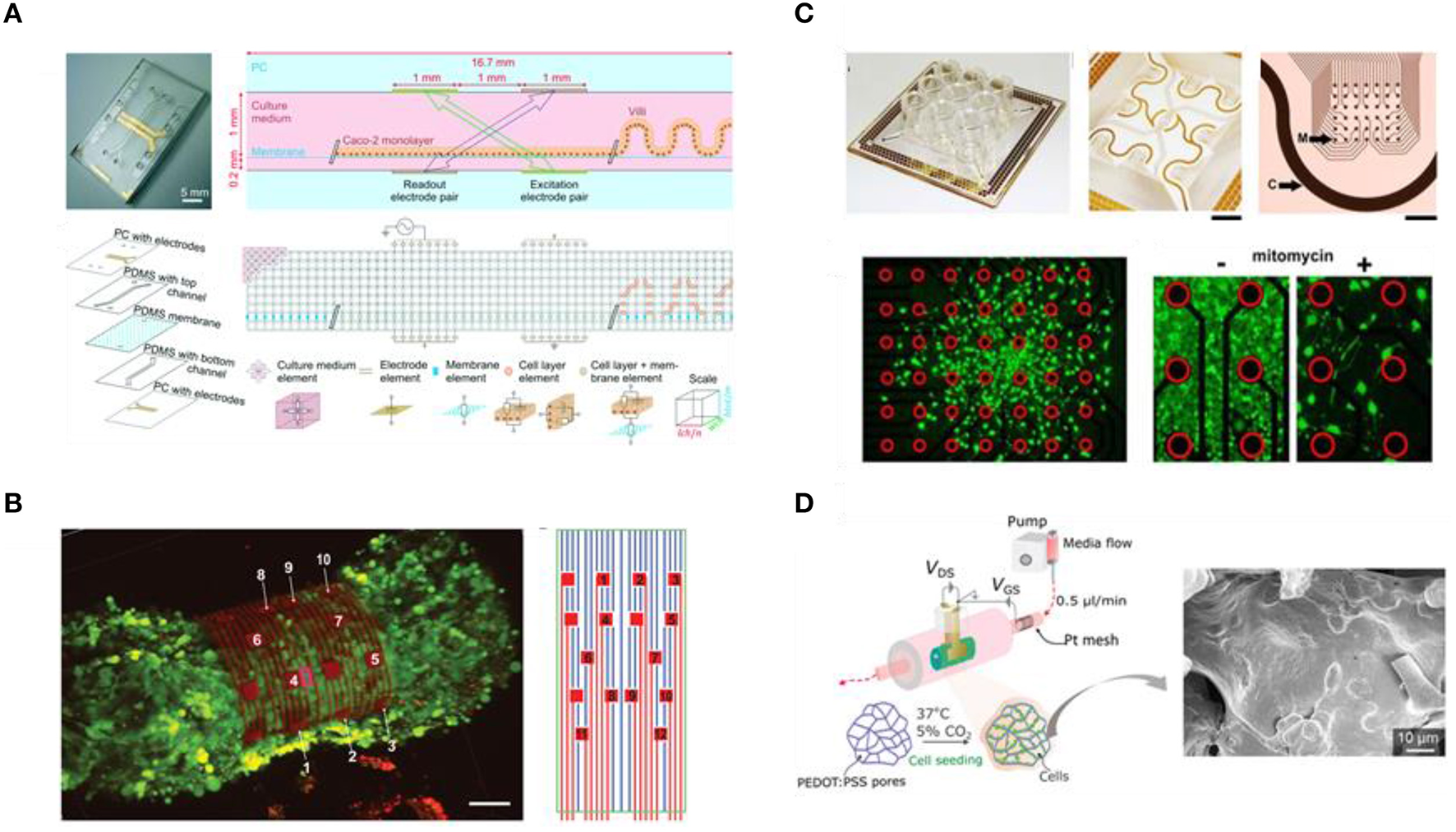
Biomimetic in vitro models of human tissues with integrated sensing and monitoring units. (A) Design and operation principles of a gut-on-chip device, along with the equivalent electrical circuit used for impedance monitoring of the intestinal barrier and the corresponding simulations. Reproduced from van der Helm et al. (2019) under the Creative Commons Attribution-Non Commercial License. (B) A 3D confocal microscopy image of 3D cardiac spheroid labeled with Ca2+ indicator dye (Fluo-4, green fluorescence) encapsulated by a self-roll biosensor array for electrical recordings; organ-on-e-chip (left) and 2D map of the microelectrodes of the biosensor array (right). Reproduced from Kalmykov et al. (2019) under the Creative Commons Attribution-Non Commercial license. (C) Image and design of a novel multi-well high-dense microelectrode array for cell migration studies (top), cell migration pattern for mitomycin C treated human breast cancer cells on the microelectrode array (bottom left), and comparative magnification to mitomycin C untreated cells (red circles mark electrodes) (bottom right). Adapted from Jahnke et al. (2019) under the Creative Commons Attribution International License. (D) Schematic representation of the setup of a 3D transistor in a tube (Tubistor), based on electroactive scaffolds (left), for hosting and concurrently monitoring 3D cell cultures. Reproduced from Pitsalidis et al. (2018) under the Creative Commons Attribution-Non Commercial license.
Conclusions and Outlook
The implementation of TE concepts and methods into biomimetic tissue models, originally directed at regenerative medicine applications, is an accelerating trend. As summarized here, 3D in vitro models of various human tissues and organs have been successfully developed, thanks to the synergistic progress and advances in all disciplines that converge to give rise to more sophisticated tissue and disease models with enhanced structural and functional accuracy. Although many challenges are yet to be resolved, the advent of hiPSCs and organoids has, without a doubt, provided bioengineers with unlimited sources of tissue-specific cells, genetic engineering of which enables further modifications (e.g., insert/delete mutations in healthy/diseased cells) to guide phenotypic behaviors. This provides an excellent opportunity for transforming drug development routes by accelerating the process, and generating data more accurate and relevant to human systems. In addition, the use of patient-specific cells facilitates studies of rare diseases, as well as precision and personalized medicine approaches for the development of drugs and therapies optimized for specific patient biology. Smart biomaterials have now been designed to more faithfully recapitulate the chemical, mechanical and topographical properties of the complex human tissues and their microenvironment. Advanced fabrication methods have additionally enabled arrangement of tissue counterparts and environmental cues with unprecedented control and accuracy. These features have combined to generate complex biological structures of high fidelity, matching the in vivo situation. A key challenge remains however: monitoring and characterization of such models still relies predominantly on end-point, invasive assays, delaying the validation of results, particularly for drug screening, and toxicology studies. Integration of electrical components in tissue engineering platforms can help resolve this limitation. Such tools enable dynamic, non-invasive and continuous monitoring of cells, offering rapid insight into different biological events within these systems. Standardization of components such as media formulation is a non-trivial additional challenge, for maintaining the survival and function of multiple cell types, which combined with spatiotemporal delivery of tissue-specific and application-specific (i.e., homeostatic or pathophysiological conditions) environmental cues, offer a novel means for dynamically controlling and directing tissue generation and maturation in vitro. Without a doubt in vitro tissue engineered systems have shown great promise for revolutionizing many aspects of biomedical research. As more sophisticated and human relevant in vitro models appear in literature, in parallel with commercialization of platforms for hosting such models, we expect to see a paradigm shift in our understanding of human biology as well as in disease diagnosis and treatment.
Statements
Author contributions
C-MM and CB wrote the text and prepared the figures. RMO wrote the text and edited the manuscript and secured the funding. All authors contributed to the article and approved the submitted version.
Funding
The authors wish to acknowledge funding from the European Research Council (ERC) under the European Union's Horizon 2020 research and innovation programme (grant agreement no. 723951 to RO, C-MM, and CB). The authors also wish to acknowledge funding by the Engineering and Physical Sciences Research Council Centre for Doctoral Training in Sensor Technologies and Applications (EP/L015889/1 to CB).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1
Abagnale G. Steger M. Nguyen V. H. Hersch N. Sechi A. Joussen S. et al . (2015). Surface topography enhances differentiation of mesenchymal stem cells towards osteogenic and adipogenic lineages. Biomaterials61, 316–326. 10.1016/j.biomaterials.2015.05.030
2
Abbott A. (2003). Biology's new dimension. Nature424, 870–872. 10.1038/424870a
3
Adadi N. Yadid M. Gal I. Asulin M. Feiner R. Edri R. et al . (2020). Electrospun fibrous PVDF-TrFe scaffolds for cardiac tissue engineering, differentiation, and maturation. Adv. Mater. Technol.5:1900820. 10.1002/admt.201900820
4
Adriani G. Ma D. Pavesi A. Kamm R. D. Goh E. L. K. (2017). A 3D neurovascular microfluidic model consisting of neurons, astrocytes and cerebral endothelial cells as a blood–brain barrier. Lab Chip17, 448–459. 10.1039/C6LC00638H
5
Afewerki S. Sheikhi A. Kannan S. Ahadian S. Khademhosseini A. (2019). Gelatin-polysaccharide composite scaffolds for 3D cell culture and tissue engineering: towards natural therapeutics. Bioeng. Transl. Med.4, 96–115. 10.1002/btm2.10124
6
Ahadian S. Civitarese R. Bannerman D. Mohammadi M. H. Lu R. Wang E. et al . (2018). Organ-on-a-chip platforms: a convergence of advanced materials, cells, and microscale technologies. Adv. Healthc. Mater.7:1700506. 10.1002/adhm.201700506
7
Avior Y. Sagi I. Benvenisty N. (2016). Pluripotent stem cells in disease modelling and drug discovery. Nat. Rev. Mol. Cell Biol.17, 170–182. 10.1038/nrm.2015.27
8
Ayuso J. M. Virumbrales-Muñoz M. Lacueva A. Lanuza P. M. Checa-Chavarria E. Botella P. et al . (2016). Development and characterization of a microfluidic model of the tumour microenvironment. Sci. Rep.6:36086. 10.1038/srep36086
9
Baeyens N. Bandyopadhyay C. Coon B. G. Yun S. Schwartz M. A. (2016). Endothelial fluid shear stress sensing in vascular health and disease. J. Clin. Invest.126, 821–828. 10.1172/JCI83083
10
Bai J. Tu T. Y. Kim C. Thiery J. P. Kamm R. D. (2015). Identification of drugs as single agents or in combination to prevent carcinoma dissemination in a microfluidic 3D environment. Oncotarget6, 36603–36614. 10.18632/oncotarget.5464
11
Baker B. M. Chen C. S. (2012). Deconstructing the third dimension-how 3D culture microenvironments alter cellular cues. J. Cell Sci.125, 3015–3024. 10.1242/jcs.079509
12
Balint R. Cassidy N. J. Cartmell S. H. (2013). Electrical stimulation: a novel tool for tissue engineering. Tissue Eng. Part B Rev.19, 48–57. 10.1089/ten.teb.2012.0183
13
Bao M. Xie J. Huck W. T. S. (2018). Recent advances in engineering the stem cell microniche in 3D. Adv. Sci.5:1800448. 10.1002/advs.201800448
14
Bao X. Zhu L. Huang X. Tang D. He D. Shi J. et al . (2017). 3D biomimetic artificial bone scaffolds with dual-cytokines spatiotemporal delivery for large weight-bearing bone defect repair. Sci. Rep.7:7814. 10.1038/s41598-017-08412-0
15
Bartfeld S. Bayram T. Van De Wetering M. Huch M. Begthel H. Kujala P. et al . (2015). In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection. Gastroenterology148, 126.e6–136.e6. 10.1053/j.gastro.2014.09.042
16
Bein A. Shin W. Jalili-Firoozinezhad S. Park M. H. Sontheimer-Phelps A. Tovaglieri A. et al . (2018). Microfluidic organ-on-a-chip models of human intestine. Cell. Mol. Gastroenterol. Hepatol.5, 659–668. 10.1016/J.JCMGH.2017.12.010
17
Benam K. H. Dauth S. Hassell B. Herland A. Jain A. Jang K.-J. et al . (2015). Engineered in vitro disease models. Annu. Rev. Pathol. Mech. Dis.10, 195–262. 10.1146/annurev-pathol-012414-040418
18
Benson K. Cramer S. Galla H.-J. (2013). Impedance-based cell monitoring: barrier properties and beyond. Fluids Barriers CNS10:5. 10.1186/2045-8118-10-5
19
Bhatia S. N. Ingber D. E. (2014). Microfluidic organs-on-chips. Nat. Biotechnol.32, 760–772. 10.1038/nbt.2989
20
Birgersdotter A. Sandberg R. Ernberg I. (2005). Gene expression perturbation in vitro- a growing case for three-dimensional (3D) culture systems. Semin. Cancer Biol.15, 405–412. 10.1016/j.semcancer.2005.06.009
21
Bissell M. J. (1981). The differentiated state of normal and malignant cells or how to define a “normal” cell in culture. Int. Rev. Cytol.70, 27–100. 10.1016/S0074-7696(08)61130-4
22
Bissell M. J. (2017). Goodbye flat biology – time for the 3rd and the 4th dimensions. J. Cell Sci.130, 3–5. 10.1242/jcs.200550
23
Bissell M. J. Barcellos-Hoff M. H. (1987). The influence of extracellular matrix on gene expression: is structure the message?J. Cell Sci.1987, 327–343. 10.1242/jcs.1987.supplement_8.18
24
Bloise N. Patrucco A. Bruni G. Montagna G. Caringella R. Fassina L. et al . (2020). In vitro production of calcified bone matrix onto wool keratin scaffolds via osteogenic factors and electromagnetic stimulus. Materials13:3052. 10.3390/ma13143052
25
Boga J. C. Miguel S. P. de Melo-Diogo D. Mendonça A. G. Louro R. O. Correia I. J. (2018). In vitro characterization of 3D printed scaffolds aimed at bone tissue regeneration. Colloids Surfaces B Biointerfaces165, 207–218. 10.1016/j.colsurfb.2018.02.038
26
Booth R. Kim H. (2012). Characterization of a microfluidic in vitro model of the blood-brain barrier (μBBB). Lab Chip12, 1784–1792. 10.1039/c2lc40094d
27
Bose S. Koski C. Vu A. A. (2020). Additive manufacturing of natural biopolymers and composites for bone tissue engineering. Mater. Horizons7, 2011–2027. 10.1039/d0mh00277a
28
Bourgine P. E. Klein T. Paczulla A. M. Shimizu T. Kunz L. Kokkaliaris K. D. et al . (2018). In vitro biomimetic engineering of a human hematopoietic niche with functional properties. Proc. Natl. Acad. Sci. U.S.A.115, E5688–E5695. 10.1073/pnas.1805440115
29
Brassard J. A. Lutolf M. P. (2019). Engineering stem cell self-organization to build better organoids. Cell Stem Cell24, 860–876. 10.1016/j.stem.2019.05.005
30
Brizzi M. F. Tarone G. Defilippi P. (2012). Extracellular matrix, integrins, and growth factors as tailors of the stem cell niche. Curr. Opin. Cell Biol.24, 645–651. 10.1016/j.ceb.2012.07.001
31
Brown J. H. Das P. DiVito M. D. Ivancic D. Tan L. P. Wertheim J. A. (2018). Nanofibrous PLGA electrospun scaffolds modified with type I collagen influence hepatocyte function and support viability in vitro. Acta Biomater.73, 217–227. 10.1016/j.actbio.2018.02.009
32
Byambaa B. Annabi N. Yue K. Trujillo-de Santiago G. Alvarez M. M. Jia W. et al . (2017). Bioprinted osteogenic and vasculogenic patterns for engineering 3D bone tissue. Adv. Healthc. Mater.6:1700015. 10.1002/adhm.201700015
33
Caddeo S. Boffito M. Sartori S. (2017). Tissue engineering approaches in the design of healthy and pathological in vitro tissue models. Front. Bioeng. Biotechnol.5:40. 10.3389/fbioe.2017.00040
34
Cairns D. M. Chwalek K. Moore Y. E. Kelley M. R. Abbott R. D. Moss S. et al . (2016). Expandable and rapidly differentiating human induced neural stem cell lines for multiple tissue engineering applications. Stem Cell Rep.7, 557–570. 10.1016/j.stemcr.2016.07.017
35
Cairns D. M. Rouleau N. Parker R. N. Walsh K. G. Gehrke L. Kaplan D. L. (2020). A 3D human brain–like tissue model of herpes-induced Alzheimer's disease. Sci. Adv.6:eaay8828. 10.1126/sciadv.aay8828
36
Campbell S. B. Wu Q. Yazbeck J. Liu C. Okhovatian S. Radisic M. (2020). Beyond polydimethylsiloxane: alternative materials for fabrication of organ-on-a-chip devices and microphysiological systems. ACS Biomater. Sci. Eng.10.1021/acsbiomaterials.0c00640
37
Campisi M. Shin Y. Osaki T. Hajal C. Chiono V. Kamm R. D. (2018). 3D self-organized microvascular model of the human blood-brain barrier with endothelial cells, pericytes and astrocytes. Biomaterials180, 117–129. 10.1016/j.biomaterials.2018.07.014
38
Cantley W. L. Du C. Lomoio S. Depalma T. Peirent E. Kleinknecht D. et al . (2018). Functional and sustainable 3D human neural network models from pluripotent stem cells. ACS Biomater. Sci. Eng.4, 4278–4288. 10.1021/acsbiomaterials.8b00622
39
Cao X. Ban E. Baker B. M. Lin Y. Burdick J. A. Chen C. S. et al . (2017). Multiscale model predicts increasing focal adhesion size with decreasing stiffness in fibrous matrices. Proc. Natl. Acad. Sci. U.S.A.114, E4549–E4555. 10.1073/pnas.1620486114
40
Carlson A. L. Bennett N. K. Francis N. L. Halikere A. Clarke S. Moore J. C. et al . (2016). Generation and transplantation of reprogrammed human neurons in the brain using 3D microtopographic scaffolds. Nat. Commun.7:10862. 10.1038/ncomms10862
41
Carter M. Shieh J. (2015a). Guide to Research Techniques in Neuroscience. Elsevier. Available online at: https://zu.edu.jo/UploadFile/Library/E_Books/Files/LibraryFile_91258_7.pdf
42
Carter M. Shieh J. (eds.). (2015b). Chapter 14 - cell culture techniques, in Guide to Research Techniques in Neuroscience, 2nd Edn (Academic Press), 295–310. 10.1016/B978-0-12-800511-8.00014-9
43
Carvalho M. R. Barata D. Teixeira L. M. Giselbrecht S. Reis R. L. Oliveira J. M. et al . (2019). Colorectal tumor-on-a-chip system: a 3D tool for precision onco-nanomedicine. Sci. Adv.5:eaaw1317. 10.1126/sciadv.aaw1317
44
Castaño A. G. García-Díaz M. Torras N. Altay G. Comelles J. Martínez E. (2019). Dynamic photopolymerization produces complex microstructures on hydrogels in a moldless approach to generate a 3D intestinal tissue model. Biofabrication11:025007. 10.1088/1758-5090/ab0478
45
Cellot G. Cilia E. Cipollone S. Rancic V. Sucapane A. Giordani S. et al . (2009). Carbon nanotubes might improve neuronal performance by favouring electrical shortcuts. Nat. Nanotechnol.4, 126–133. 10.1038/nnano.2008.374
46
Cembran A. Bruggeman K. F. Williams R. J. Parish C. L. Nisbet D. R. (2020). Biomimetic materials and their utility in modeling the 3-dimensional neural environment. iScience23:100788. 10.1016/j.isci.2019.100788
47
Chang S. Y. Weber E. J. Sidorenko V. S. Chapron A. Yeung C. K. Gao C. et al . (2017). Human liver-kidney model elucidates the mechanisms of aristolochic acid nephrotoxicity. JCI Insight2:e95978. 10.1172/jci.insight.95978
48
Charoensook S. N. Williams D. J. Chakraborty S. Leong K. W. Vunjak-Novakovic G. (2017). Bioreactor model of neuromuscular junction with electrical stimulation for pharmacological potency testing. Integr. Biol.9, 956–967. 10.1039/c7ib00144d
49
Chen C. Bai X. Ding Y. Lee I. S. (2019). Electrical stimulation as a novel tool for regulating cell behavior in tissue engineering. Biomater. Res.23, 1–12. 10.1186/s40824-019-0176-8
50
Chen C. S. (2016). 3D biomimetic cultures: the next platform for cell biology. Trends Cell Biol.26, 798–800. 10.1016/j.tcb.2016.08.008
51
Chimene D. Kaunas R. Gaharwar A. K. (2020). Hydrogel bioink reinforcement for additive manufacturing: a focused review of emerging strategies. Adv. Mater.32:1902026. 10.1002/adma.201902026
52
Chimene D. Lennox K. K. Kaunas R. R. Gaharwar A. K. (2016). Advanced bioinks for 3D printing: a materials science perspective. Ann. Biomed. Eng.44, 2090–2102. 10.1007/s10439-016-1638-y
53
Chistiakov D. A. Orekhov A. N. Bobryshev Y. V. (2017). Effects of shear stress on endothelial cells: go with the flow. Acta Physiol.219, 382–408. 10.1111/apha.12725
54
Chwalek K. Tang-Schomer M. D. Omenetto F. G. Kaplan D. L. (2015). In vitro bioengineered model of cortical brain tissue. Nat. Protoc.10, 1362–1373. 10.1038/nprot.2015.091
55
Cimmino C. Rossano L. Netti P. A. Ventre M. (2018). Spatio-temporal control of cell adhesion: toward programmable platforms to manipulate cell functions and fate. Front. Bioeng. Biotechnol.6:190. 10.3389/fbioe.2018.00190
56
Creff J. Courson R. Mangeat T. Foncy J. Souleille S. Thibault C. et al . (2019). Fabrication of 3D scaffolds reproducing intestinal epithelium topography by high-resolution 3D stereolithography. Biomaterials221:119404. 10.1016/J.BIOMATERIALS.2019.119404
57
Cui H. Nowicki M. Fisher J. P. Zhang L. G. (2017). 3D bioprinting for organ regeneration. Adv. Healthc. Mater.6:1601118. 10.1002/adhm.201601118
58
Cui W. Zhou Y. Chang J. (2010). Electrospun nanofibrous materials for tissue engineering and drug delivery. Sci. Technol. Adv. Mater.11:014108. 10.1088/1468-6996/11/1/014108
59
Curto V. F. Ferro M. P. Mariani F. Scavetta E. Owens R. M. (2018). A planar impedance sensor for 3D spheroids. Lab Chip18, 933–943. 10.1039/c8lc00067k
60
Curto V. F. Marchiori B. Hama A. Pappa A.-M. Ferro M. P. Braendlein M. et al . (2017). Organic transistor platform with integrated microfluidics for in-line multi-parametric in vitro cell monitoring. Microsystems Nanoeng.3:17028. 10.1038/micronano.2017.28
61
Cyranoski D. (2018). How human embryonic stem cells sparked a revolution. Nature. Available online at: https://www.nature.com/articles/d41586-018-03268-4 (accessed September 24, 2020).
62
Daly A. Davidson M. Burdick J. (2020). 3D bioprinting of high cell-density heterogeneous tissue models through spheroid fusion within self-healing hydrogels. bioRxiv10.1101/2020.05.21.103127
63
Darnell M. Gu L. Mooney D. (2018). RNA-seq reveals diverse effects of substrate stiffness on mesenchymal stem cells. Biomaterials181, 182–188. 10.1016/j.biomaterials.2018.07.039
64
Das S. Gordián-Vélez W. J. Ledebur H. C. Mourkioti F. Rompolas P. Chen H. I. et al . (2020). Innervation: the missing link for biofabricated tissues and organs. npj Regen. Med.5:11. 10.1038/s41536-020-0096-1
65
Dattola E. Parrotta E. I. Scalise S. Perozziello G. Limongi T. Candeloro P. et al . (2019). Development of 3D PVA scaffolds for cardiac tissue engineering and cell screening applications. RSC Adv.9, 4246–4257. 10.1039/C8RA08187E
66
De Gregorio V. Imparato G. Urciuolo F. Netti P. A. (2018). Micro-patterned endogenous stroma equivalent induces polarized crypt-villus architecture of human small intestinal epithelium. Acta Biomater.81, 43–59. 10.1016/j.actbio.2018.09.061
67
De León S. E. Pupovac A. McArthur S. L. (2020). Three-Dimensional (3D) cell culture monitoring: opportunities and challenges for impedance spectroscopy. Biotechnol. Bioeng.117, 1230–1240. 10.1002/bit.27270
68
de Rutte J. M. Koh J. Di Carlo D. (2019). Scalable high-throughput production of modular microgels for in situ assembly of microporous tissue scaffolds. Adv. Funct. Mater.29:1900071. 10.1002/adfm.201900071
69
De Souza N. (2018). Organoids. Nat. Methods15:23. 10.1038/nmeth.4576
70
De P. Dey N. Leyland-Jones B. (2013). Growth factor and signaling networks, in Brenner's Encyclopedia of Genetics, 2nd Edn. (Elsevier Inc.), 365–369.
71
Deforest C. A. Tirrell D. A. (2015). A photoreversible protein-patterning approach for guiding stem cell fate in three-dimensional gels. Nat. Mater.14, 523–531. 10.1038/nmat4219
72
del Agua I. Marina S. Pitsalidis C. Mantione D. Ferro M. Iandolo D. et al . (2018). Conducting polymer scaffolds based on poly(3,4-ethylenedioxythiophene) and xanthan gum for live-cell monitoring. ACS Omega3, 7424–7431. 10.1021/acsomega.8b00458
73
Delalat B. Cozzi C. Rasi Ghaemi S. Polito G. Kriel F. H. Michl T. D. et al . (2018). Microengineered bioartificial liver chip for drug toxicity screening. Adv. Funct. Mater.28:1801825. 10.1002/adfm.201801825
74
Delon L. C. Guo Z. Oszmiana A. Chien C. C. Gibson R. Prestidge C. et al . (2019). A systematic investigation of the effect of the fluid shear stress on Caco-2 cells towards the optimization of epithelial organ-on-chip models. Biomaterials225:119521. 10.1016/j.biomaterials.2019.119521
75
Devarasetty M. Forsythe S. D. Shelkey E. Soker S. (2020). In vitro modeling of the tumor microenvironment in tumor organoids. Tissue Eng. Regen. Med.17, 759–771. 10.1007/s13770-020-00258-4
76
Diederichs S. Böhm S. Peterbauer A. Kasper C. Scheper T. van Griensven M. (2010). Application of different strain regimes in two-dimensional and three-dimensional adipose tissue-derived stem cell cultures induces osteogenesis: implications for bone tissue engineering. J. Biomed. Mater. Res. Part A94A, 927–936. 10.1002/jbm.a.32772
77
Ding S. Kingshott P. Thissen H. Pera M. Wang P.-Y. (2017). Modulation of human mesenchymal and pluripotent stem cell behavior using biophysical and biochemical cues: a review. Biotechnol. Bioeng.114, 260–280. 10.1002/bit.26075
78
Discher D. E. Janmey P. Wang Y. L. (2005). Tissue cells feel and respond to the stiffness of their substrate. Science310, 1139–1143. 10.1126/science.1116995
79
Dosh R. H. Essa A. Jordan-Mahy N. Sammon C. Le Maitre C. L. (2017). Use of hydrogel scaffolds to develop an in vitro 3D culture model of human intestinal epithelium. Acta Biomater.62, 128–143. 10.1016/j.actbio.2017.08.035
80
Duryee W. R. Doherty J. K. (1954). Nuclear and cytoplasmic organoids in the living cell. Ann. N. Y. Acad. Sci.58, 1210–1231. 10.1111/j.1749-6632.1954.tb45904.x
81
Dutta D. Heo I. Clevers H. (2017). Disease modeling in stem cell-derived 3D organoid systems. Trends Mol. Med.23, 393–410. 10.1016/j.molmed.2017.02.007
82
Duval K. Grover H. Han L. H. Mou Y. Pegoraro A. F. Fredberg J. et al . (2017). Modeling physiological events in 2D vs. 3D cell culture. Physiology32, 266–277. 10.1152/physiol.00036.2016
83
Dye B. R. Hill D. R. Ferguson M. A. Tsai Y. H. Nagy M. S. Dyal R. et al . (2015). In vitro generation of human pluripotent stem cell derived lung organoids. Elife4:e05098. 10.7554/eLife.05098
84
Edington C. D. Chen W. L. K. Geishecker E. Kassis T. Soenksen L. R. Bhushan B. M. et al . (2018). Interconnected microphysiological systems for quantitative biology and pharmacology studies. Sci. Rep.8:4530. 10.1038/s41598-018-22749-0
85
Ejiri H. Nomura T. Hasegawa M. Tatsumi C. Imai M. Sakakibara S. et al . (2015). Use of synthetic serum-free medium for culture of human dermal fibroblasts to establish an experimental system similar to living dermis. Cytotechnology67, 507–514. 10.1007/s10616-014-9709-0
86
El-Kady A. M. Rizk R. A. Abd El-Hady B. M. Shafaa M. W. Ahmed M. M. (2012). Characterization, and antibacterial properties of novel silver releasing nanocomposite scaffolds fabricated by the gas foaming/salt-leaching technique. J. Genet. Eng. Biotechnol.10, 229–238. 10.1016/j.jgeb.2012.07.002
87
Engler A. J. Sen S. Sweeney H. L. Discher D. E. (2006). Matrix elasticity directs stem cell lineage specification. Cell126, 677–689. 10.1016/j.cell.2006.06.044
88
Evans M. G. Al-Shakli A. Chari D. M. (2019). Electrophysiological properties of neurons grown on soft polymer scaffolds reveal the potential to develop neuromimetic culture environments. Integr. Biol.11, 395–403. 10.1093/intbio/zyz033
89
Ezhilarasu H. Ramalingam R. Dhand C. Lakshminarayanan R. Sadiq A. Gandhimathi C. et al . (2019). Biocompatible aloe vera and tetracycline hydrochloride loaded hybrid nanofibrous scaffolds for skin tissue engineering. Int. J. Mol. Sci.20:5174. 10.3390/ijms20205174
90
Fabbro A. Bosi S. Ballerini L. Prato M. (2012). Carbon nanotubes: artificial nanomaterials to engineer single neurons and neuronal networks. ACS Chem. Neurosci.3, 611–618. 10.1021/cn300048q
91
Faley S. L. Neal E. H. Wang J. X. Bosworth A. M. Weber C. M. Balotin K. M. et al . (2019). iPSC-derived brain endothelium exhibits stable, long-term barrier function in perfused hydrogel scaffolds. Stem Cell Rep.12, 474–487. 10.1016/j.stemcr.2019.01.009
92
Fang B. Liu Y. Zheng D. Shan S. Wang C. Gao Y. et al . (2019). The effects of mechanical stretch on the biological characteristics of human adipose-derived stem cells. J. Cell. Mol. Med.23, 4244–4255. 10.1111/jcmm.14314
93
Fantini V. Bordoni M. Scocozza F. Conti M. Scarian E. Carelli S. et al . (2019). Bioink composition and printing parameters for 3D modeling neural tissue. Cells8:830. 10.3390/cells8080830
94
Fatehullah A. Tan S. H. Barker N. (2016). Organoids as an in vitro model of human development and disease. Nat. Cell Biol.18, 246–254. 10.1038/ncb3312
95
Felder M. Trueeb B. Stucki A. O. Borcard S. Stucki J. D. Schnyder B. et al . (2019). Impaired wound healing of alveolar lung epithelial cells in a breathing lung-on-a-chip. Front. Bioeng. Biotechnol.7:3. 10.3389/fbioe.2019.00003
96
Ferro M. P. Heilshorn S. C. Owens R. M. (2020). Materials for blood brain barrier modeling in vitro. Mater. Sci. Eng. R Rep.140:100522. 10.1016/j.mser.2019.100522
97
Fitzgerald K. A. Malhotra M. Curtin C. M. O'Brien F. J. O'Driscoll C. M. (2015). Life in 3D is never flat: 3D models to optimise drug delivery. J. Control. Release215, 39–54. 10.1016/j.jconrel.2015.07.020
98
Fontoura J. C. Viezzer C. dos Santos F. G. Ligabue R. A. Weinlich R. Puga R. D. et al . (2020). Comparison of 2D and 3D cell culture models for cell growth, gene expression and drug resistance. Mater. Sci. Eng. C107:110264. 10.1016/j.msec.2019.110264
99
Forbester J. L. Goulding D. Vallier L. Hannan N. Hale C. Pickard D. et al . (2015). Interaction of salmonella enterica serovar Typhimurium with intestinal organoids derived from human induced pluripotent stem cells. Infect. Immun.83, 2926–2934. 10.1128/IAI.00161-15
100
Frantz C. Stewart K. M. Weaver V. M. (2010). The extracellular matrix at a glance. J. Cell Sci.123, 4195–4200. 10.1242/jcs.023820
101
Fusco P. Parisatto B. Rampazzo E. Persano L. Frasson C. Di Meglio A. et al . (2019). Patient-derived organoids (PDOs) as a novel in vitro model for neuroblastoma tumours. BMC Cancer19:970. 10.1186/s12885-019-6149-4
102
Gallagher L. B. Dolan E. B. O'Sullivan J. Levey R. Cavanagh B. L. Kovarova L. et al . (2020). Pre-culture of mesenchymal stem cells within RGD-modified hyaluronic acid hydrogel improves their resilience to ischaemic conditions. Acta Biomater.107, 78–90. 10.1016/j.actbio.2020.02.043
103
Gholipourmalekabadi M. Zhao S. Harrison B. S. Mozafari M. Seifalian A. M. (2016). Oxygen-generating biomaterials: a new, viable paradigm for tissue engineering?Trends Biotechnol.34, 1010–1021. 10.1016/j.tibtech.2016.05.012
104
Gioiella F. Urciuolo F. Imparato G. Brancato V. Netti P. A. (2016). An engineered breast cancer model on a chip to replicate ECM-activation in vitro during tumor progression. Adv. Healthc. Mater.5, 3074–3084. 10.1002/adhm.201600772
105
Gonçalves A. I. Rodrigues M. T. Lee S. J. Atala A. Yoo J. J. Reis R. L. et al . (2013). Understanding the role of growth factors in modulating stem cell tenogenesis. PLoS ONE8:e83734. 10.1371/journal.pone.0083734
106
Gong H. Agustin J. Wootton D. Zhou J. G. (2014). Biomimetic design and fabrication of porous chitosan-gelatin liver scaffolds with hierarchical channel network. J. Mater. Sci. Mater. Med.25, 113–120. 10.1007/s10856-013-5061-8
107
Graham A. D. Olof S. N. Burke M. J. Armstrong J. P. K. Mikhailova E. A. Nicholson J. G. et al . (2017). High-resolution patterned cellular constructs by droplet-based 3D printing. Sci. Rep.7:7004. 10.1038/s41598-017-06358-x
108
Grebenyuk S. Ranga A. (2019). Engineering organoid vascularization. Front. Bioeng. Biotechnol.7:39. 10.3389/fbioe.2019.00039
109
Griep L. M. Wolbers F. De Wagenaar B. Ter Braak P. M. Weksler B. B. Romero I. A. et al . (2013). BBB on CHIP: microfluidic platform to mechanically and biochemically modulate blood-brain barrier function. Biomed. Microdevices15, 145–150. 10.1007/s10544-012-9699-7
110
Gu Y. Zhang J. Zhang X. Liang G. Xu T. Niu W. (2019). Three-dimensional printed Mg-Doped β-TCP bone tissue engineering scaffolds: effects of magnesium ion concentration on osteogenesis and angiogenesis in vitro. Tissue Eng. Regen. Med.16, 415–429. 10.1007/s13770-019-00192-0
111
Guan S. Zhang X.-L. Lin X.-M. Liu T.-Q. Ma X.-H. Cui Z.-F. (2013). Chitosan/gelatin porous scaffolds containing hyaluronic acid and heparan sulfate for neural tissue engineering. J. Biomater. Sci. Polym. Ed.24, 999–1014. 10.1080/09205063.2012.731374
112
Guarino V. Cirillo V. Ambrosio L. (2016). Bicomponent electrospun scaffolds to design extracellular matrix tissue analogs. Expert Rev. Med. Devices13, 83–102. 10.1586/17434440.2016.1126505
113
Guex A. G. Puetzer J. L. Armgarth A. Littmann E. Stavrinidou E. Giannelis E. P. et al . (2017). Highly porous scaffolds of PEDOT:PSS for bone tissue engineering. Acta Biomater.62, 91–101. 10.1016/j.actbio.2017.08.045
114
Guo C. Kim H. Ovadia E. M. Mourafetis C. M. Yang M. Chen W. et al . (2017). Bio-orthogonal conjugation and enzymatically triggered release of proteins within multi-layered hydrogels. Acta Biomater.56, 80–90. 10.1016/j.actbio.2017.04.002
115
Hackam D. G. Redelmeier D. A. (2006). Translation of research evidence from animals to humans. J. Am. Med. Assoc.296, 1731–1732. 10.1001/jama.296.14.1731
116
Hamburger A. W. Salmon S. E. (1977). Primary bioassay of human tumor stem cells. Science197, 461–463. 10.1126/science.560061
117
Han F. Dong Y. Su Z. Yin R. Song A. Li S. (2014). Preparation, characteristics and assessment of a novel gelatin-chitosan sponge scaffold as skin tissue engineering material. Int. J. Pharm.476, 124–133. 10.1016/j.ijpharm.2014.09.036
118
Han S. Kim S. Chen Z. Shin H. K. Lee S.-Y. Moon H. E. et al . (2020). 3D bioprinted vascularized tumour for drug testing. Int. J. Mol. Sci.21:2993. 10.3390/ijms21082993
119
Hansmann J. Groeber F. Kahlig A. Kleinhans C. Walles H. (2013). Bioreactors in tissue engineering-principles, applications and commercial constraints. Biotechnol. J.8, 298–307. 10.1002/biot.201200162
120
Hao S. Ha L. Cheng G. Wan Y. Xia Y. Sosnoski D. M. et al . (2018). A spontaneous 3D bone-on-a-chip for bone metastasis study of breast cancer cells. Small14:1702787. 10.1002/smll.201702787
121
Haring A. P. Sontheimer H. Johnson B. N. (2017). Microphysiological human brain and neural systems-on-a-chip: potential alternatives to small animal models and emerging platforms for drug discovery and personalized medicine. Stem Cell Rev. Rep.13, 381–406. 10.1007/s12015-017-9738-0
122
Hartung T. (2013). Food for thought; look back in anger - what clinical studies tell us about preclinical work. ALTEX30, 275–291. 10.14573/altex.2013.3.275
123
Hassanajili S. Karami-Pour A. Oryan A. Talaei-Khozani T. (2019). Preparation and characterization of PLA/PCL/HA composite scaffolds using indirect 3D printing for bone tissue engineering. Mater. Sci. Eng. C104:109960. 10.1016/j.msec.2019.109960
124
Hassell B. A. Goyal G. Lee E. Sontheimer-Phelps A. Levy O. Chen C. S. et al . (2017). Human organ chip models recapitulate orthotopic lung cancer growth, therapeutic responses, and tumor dormancy in vitro. Cell Rep.21, 508–516. 10.1016/j.celrep.2017.09.043
125
Hayes J. S. Richards R. G. (2010). Surfaces to control tissue adhesion for osteosynthesis with metal implants: in vitro and in vivo studies to bring solutions to the patient. Expert Rev. Med. Devices7, 131–142. 10.1586/erd.09.55
126
Heger J. I. Froehlich K. Pastuschek J. Schmidt A. Baer C. Mrowka R. et al . (2018). Human serum alters cell culture behavior and improves spheroid formation in comparison to fetal bovine serum. Exp. Cell Res.365, 57–65. 10.1016/j.yexcr.2018.02.017
127
Heo I. Dutta D. Schaefer D. A. Iakobachvili N. Artegiani B. Sachs N. et al . (2018). Modelling Cryptosporidium infection in human small intestinal and lung organoids. Nat. Microbiol.3, 814–823. 10.1038/s41564-018-0177-8
128
Herland A. Maoz B. M. Das D. Somayaji M. R. Prantil-Baun R. Novak R. et al . (2020). Quantitative prediction of human pharmacokinetic responses to drugs via fluidically coupled vascularized organ chips. Nat. Biomed. Eng.4, 421–436. 10.1038/s41551-019-0498-9
129
Herland A. van der Meer A. D. FitzGerald E. A. Park T.-E. Sleeboom J. J. F. Ingber D. E. (2016). Distinct contributions of astrocytes and pericytes to neuroinflammation identified in a 3D human blood-brain barrier on a chip. PLoS ONE11:e0150360. 10.1371/journal.pone.0150360
130
Hewitt E. Mros S. McConnell M. Cabral J. D. Ali A. (2019). Melt-electrowriting with novel milk protein/PCL biomaterials for skin regeneration. Biomed. Mater.14:055013. 10.1088/1748-605X/ab3344
131
Homan K. A. Gupta N. Kroll K. T. Kolesky D. B. Skylar-Scott M. Miyoshi T. et al . (2019). Flow-enhanced vascularization and maturation of kidney organoids in vitro. Nat. Methods16, 255–262. 10.1038/s41592-019-0325-y
132
Homan K. A. Kolesky D. B. Skylar-Scott M. A. Herrmann J. Obuobi H. Moisan A. et al . (2016). Bioprinting of 3D convoluted renal proximal tubules on perfusable chips. Sci. Rep.6:34845. 10.1038/srep34845
133
Horvath L. Umehara Y. Jud C. Blank F. Petri-Fink A. Rothen-Rutishauser B. (2015). Engineering an in vitro air-blood barrier by 3D bioprinting. Sci. Rep.5:7974. 10.1038/srep07974
134
Hribar K. C. Soman P. Warner J. Chung P. Chen S. (2014). Light-assisted direct-write of 3D functional biomaterials. Lab Chip14, 268–275. 10.1039/c3lc50634g
135
Htwe S. S. Harrington H. Knox A. Rose F. Aylott J. Haycock J. W. et al . (2015). Investigating NF-κB signaling in lung fibroblasts in 2D and 3D culture systems. Respir. Res.16:144. 10.1186/s12931-015-0302-7
136
Hu W. Wang Z. Xiao Y. Zhang S. Wang J. (2019). Advances in crosslinking strategies of biomedical hydrogels. Biomater. Sci.7, 843–855. 10.1039/c8bm01246f
137
Huang G. Li F. Zhao X. Ma Y. Li Y. Lin M. et al . (2017). Functional and biomimetic materials for engineering of the three-dimensional cell microenvironment. Chem. Rev.117, 12764–12850. 10.1021/acs.chemrev.7b00094
138
Huh D. Hamilton G. A. Ingber D. E. (2011). From 3D cell culture to organs-on-chips. Trends Cell Biol.21, 745–754. 10.1016/j.tcb.2011.09.005
139
Huh D. Leslie D. C. Matthews B. D. Fraser J. P. Jurek S. Hamilton G. A. et al . (2012). A human disease model of drug toxicity-induced pulmonary edema in a lung-on-a-chip microdevice. Sci. Transl. Med.4:159ra147. 10.1126/scitranslmed.3004249
140
Hume R. D. Berry L. Reichelt S. D'Angelo M. Gomm J. Cameron R. E. et al . (2018). An engineered human adipose/collagen model for in vitro breast cancer cell migration studies. Tissue Eng. Part A24, 1309–1319. 10.1089/ten.tea.2017.0509
141
Humphrey J. D. Dufresne E. R. Schwartz M. A. (2014). Mechanotransduction and extracellular matrix homeostasis. Nat. Rev. Mol. Cell Biol.15, 802–812. 10.1038/nrm3896
142
Iandolo D. Sheard J. Karavitas Levy G. Pitsalidis C. Tan E. Dennis A. et al . (2020). Biomimetic and electroactive 3D scaffolds for human neural crest-derived stem cell expansion and osteogenic differentiation. MRS Commun.10, 179–187. 10.1557/mrc.2020.10
143
In J. G. Foulke-Abel J. Estes M. K. Zachos N. C. Kovbasnjuk O. Donowitz M. (2016). Human mini-guts: new insights into intestinal physiology and host–pathogen interactions. Nat. Rev. Gastroenterol. Hepatol.13, 633–642. 10.1038/nrgastro.2016.142
144
Inal S. Hama A. Ferro M. Pitsalidis C. Oziat J. Iandolo D. et al . (2017). Conducting polymer scaffolds for hosting and monitoring 3D cell culture. Adv. Biosyst.1:1700052. 10.1002/adbi.201700052
145
Jacob F. Salinas R. D. Zhang D. Y. Nguyen P. T. T. Schnoll J. G. Wong S. Z. H. et al . (2020). A patient-derived glioblastoma organoid model and biobank recapitulates inter- and intra-tumoral heterogeneity. Cell180, 188.e22–204.e22. 10.1016/j.cell.2019.11.036
146
Jahnke H. G. Mewes A. Zitzmann F. D. Schmidt S. Azendorf R. Robitzki A. A. (2019). Electrochemical live monitoring of tumor cell migration out of micro-tumors on an innovative multiwell high-dense microelectrode array. Sci. Rep.9:13875. 10.1038/s41598-019-50326-6
147
Jakobsson A. Ottosson M. Zalis M. C. O'Carroll D. Johansson U. E. Johansson F. (2017). Three-dimensional functional human neuronal networks in uncompressed low-density electrospun fiber scaffolds. Nanomed. Nanotechnol. Biol. Med.13, 1563–1573. 10.1016/j.nano.2016.12.023
148
Jalili-Firoozinezhad S. Gazzaniga F. S. Calamari E. L. Camacho D. M. Fadel C. W. Bein A. et al . (2019). A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nat. Biomed. Eng.3, 520–531. 10.1038/s41551-019-0397-0
149
Jang K.-J. Mehr A. P. Hamilton G. A. McPartlin L. A. Chung S. Suh K.-Y. et al . (2013). Human kidney proximal tubule-on-a-chip for drug transport and nephrotoxicity assessment. Integr. Biol.5, 1119–1129. 10.1039/c3ib40049b
150
Jang K. J. Otieno M. A. Ronxhi J. Lim H. K. Ewart L. Kodella K. R. et al . (2019). Reproducing human and cross-species drug toxicities using a Liver-Chip. Sci. Transl. Med.11:eaax5516. 10.1126/scitranslmed.aax5516
151
Jansen E. J. P. Sladek R. E. J. Bahar H. Yaffe A. Gijbels M. J. Kuijer R. et al . (2005). Hydrophobicity as a design criterion for polymer scaffolds in bone tissue engineering. Biomaterials26, 4423–4431. 10.1016/j.biomaterials.2004.11.011
152
Jayaram A. K. Pitsalidis C. Tan E. Moysidou C.-M. De Volder M. F. L. Kim J.-S. et al . (2019). 3D hybrid scaffolds based on PEDOT:PSS/MWCNT composites. Front. Chem.7:363. 10.3389/fchem.2019.00363
153
Jedrzejczak-Silicka M. (2017). History of cell culture, in New Insights into Cell Culture Technology, ed Thatha GowderS. J. (InTech). Available online at: https://www.intechopen.com/books/new-insights-into-cell-culture-technology/history-of-cell-culture. 10.5772/66905
154
Jensen C. Teng Y. (2020). Is it time to start transitioning from 2D to 3D cell culture?Front. Mol. Biosci.7:33. 10.3389/fmolb.2020.00033
155
Jeon J. S. Bersini S. Gilardi M. Dubini G. Charest J. L. Moretti M. et al . (2015). Human 3D vascularized organotypic microfluidic assays to study breast cancer cell extravasation. Proc. Natl. Acad. Sci. U.S.A.112, 214–219. 10.1073/pnas.1417115112
156
Jeon O. Lee K. Alsberg E. (2018). Spatial micropatterning of growth factors in 3D hydrogels for location-specific regulation of cellular behaviors. Small14:1800579. 10.1002/smll.201800579
157
Jeon O. Lee Y. B. Hinton T. J. Feinberg A. W. Alsberg E. (2019). Cryopreserved cell-laden alginate microgel bioink for 3D bioprinting of living tissues. Mater. Today Chem.12, 61–70. 10.1016/j.mtchem.2018.11.009
158
Joshi J. Brennan D. Beachley V. Kothapalli C. R. (2018). Cardiomyogenic differentiation of human bone marrow-derived mesenchymal stem cell spheroids within electrospun collagen nanofiber mats. J. Biomed. Mater. Res. Part A106, 3303–3312. 10.1002/jbm.a.36530
159
Kadohama T. Nishimura K. Hoshino Y. Sasajima T. Sumpio B. E. (2007). Effects of different types of fluid shear stress on endothelial cell proliferation and survival. J. Cell. Physiol.212, 244–251. 10.1002/jcp.21024
160
Kalmykov A. Huang C. Bliley J. Shiwarski D. Tashman J. Abdullah A. et al . (2019). Organ-on-e-chip: three-dimensional self-rolled biosensor array for electrical interrogations of human electrogenic spheroids. Sci. Adv.5:eaax0729. 10.1126/sciadv.aax0729
161
Kamiya A. Ando J. (1996). Responses of vascular endothelial cells to fluid shear stress: mechanism, in Biomechanics, eds HayashiK.KamiyaA.OnoK. (Tokyo: Springer), 29–56. 10.1007/978-4-431-68317-9_2
162
Kapałczyńska M. Kolenda T. Przybyła W. Zajaczkowska M. Teresiak A. Filas V. et al . (2018). 2D and 3D cell cultures – a comparison of different types of cancer cell cultures. Arch. Med. Sci.14, 910–919. 10.5114/aoms.2016.63743
163
Karve S. S. Pradhan S. Ward D. V. Weiss A. A. (2017). Intestinal organoids model human responses to infection by commensal and Shiga toxin producing Escherichia coli. PLoS ONE12:e0178966. 10.1371/journal.pone.0178966
164
Karzbrun E. Kshirsagar A. Cohen S. R. Hanna J. H. Reiner O. (2018). Human brain organoids on a chip reveal the physics of folding. Nat. Phys.14, 515–522. 10.1038/s41567-018-0046-7
165
Kasendra M. Tovaglieri A. Sontheimer-Phelps A. Jalili-Firoozinezhad S. Bein A. Chalkiadaki A. et al . (2018). Development of a primary human Small Intestine-on-a-Chip using biopsy-derived organoids. Sci. Rep.8:2871. 10.1038/s41598-018-21201-7
166
Kato-Negishi M. Morimoto Y. Onoe H. Takeuchi S. (2013). Millimeter-sized neural building blocks for 3D heterogeneous neural network assembly. Adv. Healthc. Mater.2, 1564–1570. 10.1002/adhm.201300052
167
Keirouz A. Fortunato G. Zhang M. Callanan A. Radacsi N. (2019). Nozzle-free electrospinning of Polyvinylpyrrolidone/Poly(glycerol sebacate) fibrous scaffolds for skin tissue engineering applications. Med. Eng. Phys.71, 56–67. 10.1016/j.medengphy.2019.06.009
168
Kessler M. Hoffmann K. Brinkmann V. Thieck O. Jackisch S. Toelle B. et al . (2015). The Notch and Wnt pathways regulate stemness and differentiation in human fallopian tube organoids. Nat. Commun.6:8989. 10.1038/ncomms9989
169
Khademhosseini A. Langer R. (2016). A decade of progress in tissue engineering. Nat. Protoc.11, 1775–1781. 10.1038/nprot.2016.123
170
Khademhosseini A. Langer R. Borenstein J. Vacanti J. P. (2006). Microscale Technologies for Tissue Engineering and Biology. Available online at: www.pnas.org/cgi/doi/10.1073/pnas.0507681102 (accessed September 25, 2020).
171
Kilic O. Pamies D. Lavell E. Schiapparelli P. Feng Y. Hartung T. et al . (2016). Brain-on-a-chip model enables analysis of human neuronal differentiation and chemotaxis. Lab Chip16, 4152–4162. 10.1039/c6lc00946h
172
Kim H. J. Ingber D. E. (2013). Gut-on-a-Chip microenvironment induces human intestinal cells to undergo villus differentiation. Integr. Biol.5:1130. 10.1039/c3ib40126j
173
Kim J. Koo B. K. Knoblich J. A. (2020). Human organoids: model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol.21, 571–584. 10.1038/s41580-020-0259-3
174
Kim M. H. Kumar S. K. Shirahama H. Seo J. Lee J. H. Zhdanov V. P. et al . (2016). Biofunctionalized hydrogel microscaffolds promote 3D hepatic sheet morphology. Macromol. Biosci.16, 314–321. 10.1002/mabi.201500338
175
Kim S. Chung M. Ahn J. Lee S. Jeon N. L. (2016). Interstitial flow regulates the angiogenic response and phenotype of endothelial cells in a 3D culture model. Lab Chip16, 4189–4199. 10.1039/c6lc00910g
176
Kim S. H. Chi M. Yi B. Kim S. H. Oh S. Kim Y. et al . (2014). Three-dimensional intestinal villi epithelium enhances protection of human intestinal cells from bacterial infection by inducing mucin expression. Integr. Biol.6, 1122–1131. 10.1039/c4ib00157e
177
Kloxin A. M. Kasko A. M. Salinas C. N. Anseth K. S. (2009). Photodegradable hydrogels for dynamic tuning of physical and chemical properties. Science324, 59–63. 10.1126/science.1169494
178
Knight E. Przyborski S. (2015). Advances in 3D cell culture technologies enabling tissue-like structures to be created in vitro. J. Anat.227, 746–756. 10.1111/joa.12257
179
Koivisto J. T. Gering C. Karvinen J. Maria Cherian R. Belay B. Hyttinen J. et al . (2019). Mechanically biomimetic gelatin-gellan gum hydrogels for 3D culture of beating human cardiomyocytes. ACS Appl. Mater. Interfaces11, 20589–20602. 10.1021/acsami.8b22343
180
Koledova Z. (2017). 3D cell culture: an introduction, in Methods in Molecular Biology, ed KiledovaZ. (New York, NY: Humana Press Inc.), 1–11. 10.1007/978-1-4939-7021-6_1
181
Kolesky D. B. Homan K. A. Skylar-Scott M. A. Lewis J. A. (2016). Three-dimensional bioprinting of thick vascularized tissues. Proc. Natl. Acad. Sci. U.S.A.113, 3179–3184. 10.1073/pnas.1521342113
182
Kolesky D. B. Truby R. L. Gladman A. S. Busbee T. A. Homan K. A. Lewis J. A. (2014). 3D bioprinting of vascularized, heterogeneous cell-laden tissue constructs. Adv. Mater.26, 3124–3130. 10.1002/adma.201305506
183
Kratochvil M. J. Seymour A. J. Li T. L. Paşca S. P. Kuo C. J. Heilshorn S. C. (2019). Engineered materials for organoid systems. Nat. Rev. Mater.4, 606–622. 10.1038/s41578-019-0129-9
184
Kumar A. Nune K. C. Misra R. D. K. (2016). Understanding the response of pulsed electric field on osteoblast functions in three-dimensional mesh structures. J. Biomater. Appl.31, 594–605. 10.1177/0885328216658376
185
Kumar A. Placone J. K. Engler A. J. (2017). Understanding the extracellular forces that determine cell fate and maintenance. Development144, 4261–4270. 10.1242/dev.158469
186
Kumari J. Karande A. A. Kumar A. (2016). Combined effect of cryogel matrix and temperature-reversible soluble-insoluble polymer for the development of in vitro human liver tissue. ACS Appl. Mater. Interfaces8, 264–277. 10.1021/acsami.5b08607
187
Kuriakose A. E. Hu W. Nguyen K. T. Menonid J. U. (2019). Scaffold-based lung tumor culture on porous PLGA microparticle substratesPLoS ONE14:e0217640. 10.1371/journal.pone.0217640
188
Lamers M. M. Beumer J. Vaart J. Van Der Knoops K. Puschhof J. Breugem T. I. et al . (2020). SARS-CoV-2 productively infects human gut enterocytes. Science369, 50–54. 10.1126/science.abc1669
189
Lancaster M. A. Huch M. (2019). Disease modelling in human organoids. Dis. Model. Mech.12:dmm039347. 10.1242/dmm.039347
190
Lancaster M. A. Renner M. Martin C.-A. Wenzel D. Bicknell L. S. Hurles M. E. et al . (2013). Cerebral organoids model human brain development and microcephaly. Nature501, 373–379. 10.1038/nature12517
191
Langer R. Vacanti J. P. (1993). Tissue engineering. Science260, 920–926. 10.1126/science.8493529
192
Langhans S. A. (2018). Three-dimensional in vitro cell culture models in drug discovery and drug repositioning. Front. Pharmacol.9:6. 10.3389/fphar.2018.00006
193
Lea T. (2015). Epithelial cell models; general introduction, in The Impact of Food Bioactives on Health: in vitro and ex vivo models, 95–102. Available onlineat: https://link.springer.com/content/pdf/10.1007%2F978-3-319-16104-4_9.pdf (accessed March 27, 2018).
194
Ledo A. M. Senra A. Rilo-Alvarez H. Borrajo E. Vidal A. Alonso M. J. et al . (2020a). mRNA-activated matrices encoding transcription factors as primers of cell differentiation in tissue engineering. Biomaterials247:120016. 10.1016/j.biomaterials.2020.120016
195
Ledo A. M. Vining K. H. Alonso M. J. Garcia-Fuentes M. Mooney D. J. (2020b). Extracellular matrix mechanics regulate transfection and SOX9-directed differentiation of mesenchymal stem cells. Acta Biomater.110, 153–163. 10.1016/j.actbio.2020.04.027
196
Lee A. Hudson A. R. Shiwarski D. J. Tashman J. W. Hinton T. J. Yerneni S. et al . (2019). 3D bioprinting of collagen to rebuild components of the human heart. Science365, 482–487. 10.1126/science.aav9051
197
Lee H. Chae S. Kim J. Y. Han W. Kim J. Choi Y. et al . (2019). Cell-printed 3D liver-on-a-chip possessing a liver microenvironment and biliary system. Biofabrication11:25001. 10.1088/1758-5090/aaf9fa
198
Lee W. Debasitis J. C. Lee V. K. Lee J. H. Fischer K. Edminster K. et al . (2009). Multi-layered culture of human skin fibroblasts and keratinocytes through three-dimensional freeform fabrication. Biomaterials30, 1587–1595. 10.1016/j.biomaterials.2008.12.009
199
Leijten J. Seo J. Yue K. Trujillo-de Santiago G. Tamayol A. Ruiz-Esparza G. U. et al . (2017). Spatially and temporally controlled hydrogels for tissue engineering. Mater. Sci. Eng. R Rep.119, 1–35. 10.1016/j.mser.2017.07.001
200
Leslie J. L. Huang S. Opp J. S. Nagy M. S. Kobayashi M. Young V. B. et al . (2015). Persistence and toxin production by Clostridium difficile within human intestinal organoids result in disruption of epithelial paracellular barrier function. Infect. Immun.83, 138–145. 10.1128/IAI.02561-14
201
Levorson E. J. Kasper F. K. Mikos A. G. (2011). Scaffolds: flow perfusion bioreactor design, in Comprehensive Biomaterials, ed DucheyneP. (Elsevier), 1–11. 10.1016/b978-0-08-055294-1.00154-9
202
Li J. Liu X. Crook J. M. Wallace G. G. (2020). 3D printing of cytocompatible graphene/alginate scaffolds for mimetic tissue constructs. Front. Bioeng. Biotechnol.8:824. 10.3389/fbioe.2020.00824
203
Li Q. Nan K. Le Floch P. Lin Z. Sheng H. Liu J. (2019). Cyborg organoids: implantation of nanoelectronics via organogenesis for tissue-wide electrophysiology. bioRxiv697664. 10.1101/697664
204
Li X. Francies H. E. Secrier M. Perner J. Miremadi A. Galeano-Dalmau N. et al . (2018). Organoid cultures recapitulate esophageal adenocarcinoma heterogeneity providing a model for clonality studies and precision therapeutics. Nat. Commun.9:2983. 10.1038/s41467-018-05190-9
205
Liu T. kun Pang Y. Zhou Z. zhen Yao R. Sun W. (2019). An integrated cell printing system for the construction of heterogeneous tissue models. Acta Biomater.95, 245–257. 10.1016/j.actbio.2019.05.052
206
Liu X. Zheng C. Luo X. Wang X. Jiang H. (2019). Recent advances of collagen-based biomaterials: multi-hierarchical structure, modification and biomedical applications. Mater. Sci. Eng. C99, 1509–1522. 10.1016/j.msec.2019.02.070
207
Llucià-Valldeperas A. Sanchez B. Soler-Botija C. Gálvez-Montón C. Prat-Vidal C. Roura S. et al . (2015). Electrical stimulation of cardiac adipose tissue-derived progenitor cells modulates cell phenotype and genetic machinery. J. Tissue Eng. Regen. Med.9, E76–E83. 10.1002/term.1710
208
Low L. A. Tagle D. A. (2017). Tissue chips-innovative tools for drug development and disease modeling. Lab Chip17, 3026–3036. 10.1039/c7lc00462a
209
Lozano R. Stevens L. Thompson B. C. Gilmore K. J. Gorkin R. Stewart E. M. et al . (2015). 3D printing of layered brain-like structures using peptide modified gellan gum substrates. Biomaterials67, 264–273. 10.1016/j.biomaterials.2015.07.022
210
Luca A. C. Mersch S. Deenen R. Schmidt S. Messner I. Schäfer K.-L. et al . (2013). Impact of the 3D microenvironment on phenotype, gene expression, and EGFR inhibition of colorectal cancer cell lines. PLoS ONE8:e59689. 10.1371/journal.pone.0059689
211
Ma X. Yu C. Wang P. Xu W. Wan X. Lai C. S. E. et al . (2018). Rapid 3D bioprinting of decellularized extracellular matrix with regionally varied mechanical properties and biomimetic microarchitecture. Biomaterials185, 310–321. 10.1016/j.biomaterials.2018.09.026
212
Mabrouk M. Beherei H. H. Das D. B. (2020). Recent progress in the fabrication techniques of 3D scaffolds for tissue engineering. Mater. Sci. Eng. C110:110716. 10.1016/j.msec.2020.110716
213
Madl C. M. LeSavage B. L. Dewi R. E. Lampe K. J. Heilshorn S. C. (2019). Matrix remodeling enhances the differentiation capacity of neural progenitor cells in 3D hydrogels. Adv. Sci.6:1801716. 10.1002/advs.201801716
214
Magnan L. Labrunie G. Fénelon M. Dusserre N. Foulc M. P. Lafourcade M. et al . (2020). Human textiles: a cell-synthesized yarn as a truly “bio” material for tissue engineering applications. Acta Biomater.105, 111–120. 10.1016/j.actbio.2020.01.037
215
Majidi S. S. Slemming-Adamsen P. Hanif M. Zhang Z. Wang Z. Chen M. (2018). Wet electrospun alginate/gelatin hydrogel nanofibers for 3D cell culture. Int. J. Biol. Macromol.118, 1648–1654. 10.1016/j.ijbiomac.2018.07.005
216
Manousiouthakis E. Chen Y. Cairns D. M. Pollard R. Gerlovin K. Dente M. J. et al . (2019). Bioengineered in vitro enteric nervous system. J. Tissue Eng. Regen. Med.13, 1712–1723. 10.1002/term.2926
217
Mao A. S. Shin J. W. Mooney D. J. (2016). Effects of substrate stiffness and cell-cell contact on mesenchymal stem cell differentiation. Biomaterials98, 184–191. 10.1016/j.biomaterials.2016.05.004
218
Mao Q. Wang Y. Li Y. Juengpanich S. Li W. Chen M. et al . (2020). Fabrication of liver microtissue with liver decellularized extracellular matrix (dECM) bioink by digital light processing (DLP) bioprinting. Mater. Sci. Eng. C109:110625. 10.1016/j.msec.2020.110625
219
Mao W. Kang M. K. Shin J. U. Son Y. J. Kim H. S. Yoo H. S. (2018). Coaxial hydro-nanofibrils for self-assembly of cell sheets producing skin bilayers. ACS Appl. Mater. Interfaces10, 43503–43511. 10.1021/acsami.8b17740
220
Marga F. Jakab K. Khatiwala C. Shepherd B. Dorfman S. Hubbard B. et al . (2012). Toward engineering functional organ modules by additive manufacturing. Biofabrication4:022001. 10.1088/1758-5082/4/2/022001
221
Martinelli V. Cellot G. Toma F. M. Long C. S. Caldwell J. H. Zentilin L. et al . (2012). Carbon nanotubes promote growth and spontaneous electrical activity in cultured cardiac myocytes. Nano Lett.12, 1831–1838. 10.1021/nl204064s
222
Marturano-Kruik A. Nava M. M. Yeager K. Chramiec A. Hao L. Robinson S. et al . (2018). Human bone perivascular niche-on-a-chip for studying metastatic colonization. Proc. Natl. Acad. Sci. U.S.A.115, 1256–1261. 10.1073/pnas.1714282115
223
Maschmeyer I. Hasenberg T. Jaenicke A. Lindner M. Lorenz A. K. Zech J. et al . (2015a). Chip-based human liver-intestine and liver-skin co-cultures - a first step toward systemic repeated dose substance testing in vitro. Eur. J. Pharm. Biopharm.95, 77–87. 10.1016/j.ejpb.2015.03.002
224
Maschmeyer I. Lorenz A. K. Schimek K. Hasenberg T. Ramme A. P. Hübner J. et al . (2015b). A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin and kidney equivalents. Lab Chip15, 2688–2699. 10.1039/c5lc00392j
225
Massa S. Sakr M. A. Seo J. Bandaru P. Arneri A. Bersini S. et al . (2017). Bioprinted 3D vascularized tissue model for drug toxicity analysis. Biomicrofluidics11:044109. 10.1063/1.4994708
226
Mazzocchi A. Devarasetty M. Huntwork R. Soker S. (2019). Optimization of collagen type I-hyaluronan hybrid bioink for 3D bioprinted liver microenvironments. Biofabrication11:015003. 10.1088/1758-5090/aae543
227
McCaig C. D. Rajnicek A. M. Song B. Zhao M. (2005). Controlling cell behavior electrically: current views and future potential. Physiol. Rev.85, 943–978. 10.1152/physrev.00020.2004
228
McKee C. Chaudhry G. R. (2017). Advances and challenges in stem cell culture. Colloids Surfaces B Biointerfaces159, 62–77. 10.1016/j.colsurfb.2017.07.051
229
McMurtrey R. J. (2016). Analytic models of oxygen and nutrient diffusion, metabolism dynamics, and architecture optimization in three-dimensional tissue constructs with applications and insights in cerebral organoids. Tissue Eng. Part C Methods22, 221–249. 10.1089/ten.tec.2015.0375
230
Melchels F. P. W. Domingos M. A. N. Klein T. J. Malda J. Bartolo P. J. Hutmacher D. W. (2012). Additive manufacturing of tissues and organs. Prog. Polym. Sci.37, 1079–1104. 10.1016/j.progpolymsci.2011.11.007
231
Mihic A. Li J. Miyagi Y. Gagliardi M. Li S. H. Zu J. et al . (2014). The effect of cyclic stretch on maturation and 3D tissue formation of human embryonic stem cell-derived cardiomyocytes. Biomaterials35, 2798–2808. 10.1016/j.biomaterials.2013.12.052
232
Miller J. S. Stevens K. R. Yang M. T. Baker B. M. Nguyen D. H. T. Cohen D. M. et al . (2012). Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nat. Mater.11, 768–774. 10.1038/nmat3357
233
Miller P. G. Shuler M. L. (2016). Design and demonstration of a pumpless 14 compartment microphysiological system. Biotechnol. Bioeng.113, 2213–2227. 10.1002/bit.25989
234
Min D. Lee W. Bae I. H. Lee T. R. Croce P. Yoo S. S. (2018). Bioprinting of biomimetic skin containing melanocytes. Exp. Dermatol.27, 453–459. 10.1111/exd.13376
235
Mittal R. Woo F. W. Castro C. S. Cohen M. A. Karanxha J. Mittal J. et al . (2018). Organ-on-chip models: Implications in drug discovery and clinical applications. J. Cell. Physiol.234, 8352–8380. 10.1002/jcp.27729
236
Miura S. Sato K. Kato-Negishi M. Teshima T. Takeuchi S. (2015). Fluid shear triggers microvilli formation via mechanosensitive activation of TRPV6. Nat. Commun.6:8871. 10.1038/ncomms9871
237
Monferrer E. Martín-Vañó S. Carretero A. García-Lizarribar A. Burgos-Panadero R. Navarro S. et al . (2020). A three-dimensional bioprinted model to evaluate the effect of stiffness on neuroblastoma cell cluster dynamics and behavior. Sci. Rep.10:35367. 10.1038/s41598-020-62986-w
238
Morán G. A. G. Parra-Medina R. Cardona A. G. Quintero-Ronderos P. Rodríguez É. G. (2013). Cytokines, Chemokines and Growth Factors. Available online at: https://www.ncbi.nlm.nih.gov/books/NBK459450/ (accessed September 24, 2020).
239
Morgan J. P. Delnero P. F. Zheng Y. Verbridge S. S. Chen J. Craven M. et al . (2013). Formation of microvascular networks in vitro. Nat. Protoc.8, 1820–1836. 10.1038/nprot.2013.110
240
Morris G. E. Bridge J. C. Eltboli O. M. I. Knox A. J. Aylott J. W. Brightling C. E. et al . (2014). Human airway smooth muscle maintain in situ cell orientation and phenotype when cultured on aligned electrospun scaffolds. Am. J. Physiol. Lung Cell. Mol. Physiol.307, L38. 10.1152/ajplung.00318.2013
241
Moxon S. R. Corbett N. J. Fisher K. Potjewyd G. Domingos M. Hooper N. M. (2019). Blended alginate/collagen hydrogels promote neurogenesis and neuronal maturation. Mater. Sci. Eng. C104:109904. 10.1016/j.msec.2019.109904
242
Muraglia A. Nguyen V. T. Nardini M. Mogni M. Coviello D. Dozin B. et al . (2017). Culture medium supplements derived from human platelet and plasma: cell commitment and proliferation support. Front. Bioeng. Biotechnol.5:66. 10.3389/fbioe.2017.00066
243
Murphy A. R. Haynes J. M. Laslett A. L. Cameron N. R. O'Brien C. M. (2020). Three-dimensional differentiation of human pluripotent stem cell-derived neural precursor cells using tailored porous polymer scaffolds. Acta Biomater.101, 102–116. 10.1016/j.actbio.2019.10.017
244
Murr L. E. (2016). Frontiers of 3D printing/additive manufacturing: from human organs to aircraft fabrication. J. Mater. Sci. Technol.32, 987–995. 10.1016/j.jmst.2016.08.011
245
Nachlas A. L. Y. Li S. Streeter B. W. De Jesus Morales K. J. Sulejmani F. Madukauwa-David D. I. et al . (2020). A multilayered valve leaflet promotes cell-laden collagen type I production and aortic valve hemodynamics. Biomaterials240:119838. 10.1016/j.biomaterials.2020.119838
246
Nagle P. W. Plukker J. T. M. Muijs C. T. van Luijk P. Coppes R. P. (2018). Patient-derived tumor organoids for prediction of cancer treatment response. Semin. Cancer Biol.53, 258–264. 10.1016/j.semcancer.2018.06.005
247
Nelson L. Tighe A. Golder A. Littler S. Bakker B. Moralli D. et al . (2020). A living biobank of ovarian cancer ex vivo models reveals profound mitotic heterogeneity. Nat. Commun.11:822. 10.1038/s41467-020-14551-2
248
Neufurth M. Wang X. Wang S. Steffen R. Ackermann M. Haep N. D. et al . (2017). 3D printing of hybrid biomaterials for bone tissue engineering: calcium-polyphosphate microparticles encapsulated by polycaprolactone. Acta Biomater.64, 377–388. 10.1016/j.actbio.2017.09.031
249
Nichol J. W. Khademhosseini A. (2009). Modular tissue engineering: Engineering biological tissues from the bottom up. Soft Matter5, 1312–1319. 10.1039/b814285h
250
Nikolaev M. Mitrofanova O. Broguiere N. Geraldo S. Dutta D. Tabata Y. et al . (2020). Homeostatic mini-intestines through scaffold-guided organoid morphogenesis. Nature585, 574–578. 10.1038/s41586-020-2724-8
251
Nikolova M. P. Chavali M. S. (2019). Recent advances in biomaterials for 3D scaffolds: a review. Bioact. Mater.4, 271–292. 10.1016/j.bioactmat.2019.10.005
252
O'Leary C. Cavanagh B. Unger R. E. Kirkpatrick C. J. O'Dea S. O'Brien F. J. et al . (2016). The development of a tissue-engineered tracheobronchial epithelial model using a bilayered collagen-hyaluronate scaffold. Biomaterials85, 111–127. 10.1016/j.biomaterials.2016.01.065
253
Onoe H. Okitsu T. Itou A. Kato-Negishi M. Gojo R. Kiriya D. et al . (2013). Metre-long cell-laden microfibres exhibit tissue morphologies and functions. Nat. Mater.12, 584–590. 10.1038/nmat3606
254
Ooft S. N. Weeber F. Dijkstra K. K. McLean C. M. Kaing S. van Werkhoven E. et al . (2019). Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer patients. Sci. Transl. Med.11:eaay2574. 10.1126/scitranslmed.aay2574
255
Ortega-Prieto A. M. Skelton J. K. Wai S. N. Large E. Lussignol M. Vizcay-Barrena G. et al . (2018). 3D microfluidic liver cultures as a physiological preclinical tool for hepatitis B virus infection. Nat. Commun.9:682. 10.1038/s41467-018-02969-8
256
Ouyang L. Armstrong J. P. K. Salmeron-Sanchez M. Stevens M. M. (2020). Assembling living building blocks to engineer complex tissues. Adv. Funct. Mater.30:1909009. 10.1002/adfm.201909009
257
Owens R. M. Iandolo D. Wittmann K. (2017). A (bio) materials approach to three-dimensional cell biology. MRS Commun.7, 287–288. 10.1557/mrc.2017.102
258
Pang Y. Sutoko S. Wang Z. Horimoto Y. Montagne K. Horiguchi I. et al . (2020). Organization of liver organoids using Raschig ring-like micro-scaffolds and triple co-culture: Toward modular assembly-based scalable liver tissue engineering. Med. Eng. Phys.76, 69–78. 10.1016/j.medengphy.2019.10.010
259
Park J. Lee B. K. Jeong G. S. Hyun J. K. Lee C. J. Lee S. H. (2015). Three-dimensional brain-on-a-chip with an interstitial level of flow and its application as an in vitro model of Alzheimer's disease. Lab Chip15, 141–150. 10.1039/c4lc00962b
260
Park S. M. Eom S. Hong H. Yoon J. Lee S. J. Kim B. C. et al . (2019). Reconstruction of in vivo-like in vitro model: enabling technologies of microfluidic systems for dynamic biochemical/mechanical stimuli. Microelectron. Eng. 203–204, 6–24. 10.1016/j.mee.2018.10.010
261
Pas J. Pitsalidis C. Koutsouras D. A. Quilichini P. P. Santoro F. Cui B. et al . (2017). Neurospheres on patterned PEDOT:PSS microelectrode arrays enhance electrophysiology recordings. Adv. Biosyst.2:1700164. 10.1002/adbi.201700164
262
Patel A. K. Celiz A. D. Rajamohan D. Anderson D. G. Langer R. Davies M. C. et al . (2015). A defined synthetic substrate for serum-free culture of human stem cell derived cardiomyocytes with improved functional maturity identified using combinatorial materials microarrays. Biomaterials61, 257–265. 10.1016/j.biomaterials.2015.05.019
263
Patel M. Nakaji-Hirabayashi T. Matsumura K. (2019). Effect of dual-drug-releasing micelle–hydrogel composite on wound healing in vivo in full-thickness excision wound rat model. J. Biomed. Mater. Res. Part A107, 1094–1106. 10.1002/jbm.a.36639
264
Patient J. D. Hajiali H. Harris K. Abrahamsson B. Tannergren C. White L. J. et al . (2019). Nanofibrous scaffolds support a 3D in vitro permeability model of the human intestinal epithelium. Front. Pharmacol.10:456. 10.3389/fphar.2019.00456
265
Pearce S. C. Coia H. G. Karl J. P. Pantoja-Feliciano I. G. Zachos N. C. Racicot K. (2018). Intestinal in vitro and ex vivo models to study host-microbiome interactions and acute stressors. Front. Physiol.9:1584. 10.3389/fphys.2018.01584
266
Pedron S. Becka E. Harley B. A. C. (2013). Regulation of glioma cell phenotype in 3D matrices by hyaluronic acid. Biomaterials34, 7408–7417. 10.1016/j.biomaterials.2013.06.024
267
Pitsalidis C. Ferro M. P. Iandolo D. Tzounis L. Inal S. Owens R. M. (2018). Transistor in a tube: a route to three-dimensional bioelectronics. Sci. Adv.4:eaat4253. 10.1126/sciadv.aat4253
268
Plummer S. Wallace S. Ball G. Lloyd R. Schiapparelli P. Quiñones-Hinojosa A. et al . (2019). A Human iPSC-derived 3D platform using primary brain cancer cells to study drug development and personalized medicine. Sci. Rep.9:1407. 10.1038/s41598-018-38130-0
269
Pradhan S. Brooks A. K. Yadavalli V. K. (2020). Nature-derived materials for the fabrication of functional biodevices. Mater. Today Bio.7:100065. 10.1016/j.mtbio.2020.100065
270
Prantil-Baun R. Novak R. Das D. Somayaji M. R. Przekwas A. Ingber D. E. (2018). Physiologically based pharmacokinetic and pharmacodynamic analysis enabled by microfluidically linked organs-on-chips. Annu. Rev. Pharmacol. Toxicol.58, 37–64. 10.1146/annurev-pharmtox-010716-104748
271
Prévô M. E. Andro H. M. Alexander S. L. Ustunel S. Zhu C. Nikolov Z. et al . (2018). Liquid crystal elastomer foams with elastic properties specifically engineered as biodegradable brain tissue scaffolds. Soft Matter14:354. 10.1039/c7sm01949a
272
Przyborski S. (2017). Technology Platforms for 3D Cell Culture. Wiley-Blackwell. Available online at: https://www.wiley.com/en-gb/Technology+Platforms+for+3D+Cell+Culture:+A+User's+Guide-p-9781118851500 (accessed January 8, 2018).
273
Pudlas M. Koch S. Bolwien C. Thude S. Jenne N. Hirth T. et al . (2011). Raman spectroscopy: a noninvasive analysis tool for the discrimination of human skin cells. Tissue Eng. Part C Methods17, 1027–1040. 10.1089/ten.tec.2011.0082
274
Rahmani A. Bakhshayesh D. Mostafavi E. Alizadeh E. Asadi N. Akbarzadeh A. et al . (2018). Fabrication of three-dimensional scaffolds based on nano-biomimetic collagen hybrid constructs for skin tissue engineering. ACS Omega3, 8605–8611. 10.1021/acsomega.8b01219
275
Ramadan Q. Jing L. (2016). Characterization of tight junction disruption and immune response modulation in a miniaturized Caco-2/U937 coculture-based in vitro model of the human intestinal barrier. Biomed. Microdevices18:11. 10.1007/s10544-016-0035-5
276
Ramadan Q. Zourob M. (2020). Organ-on-a-chip engineering: Toward bridging the gap between lab and industry. Biomicrofluidics14:41501. 10.1063/5.0011583
277
Ribas J. Sadeghi H. Manbachi A. Leijten J. Brinegar K. Zhang Y. S. et al . (2016). Cardiovascular organ-on-a-chip platforms for drug discovery and development. Appl. Vitr. Toxicol.2, 82–96. 10.1089/aivt.2016.0002
278
Riehl B. D. Park J. H. Kwon I. K. Lim J. Y. (2012). Mechanical stretching for tissue engineering: two-dimensional and three-dimensional constructs. Tissue Eng. Part B Rev.18, 288–300. 10.1089/ten.teb.2011.0465
279
Rivnay J. Inal S. Salleo A. Owens R. M. Berggren M. Malliaras G. G. (2018). Organic electrochemical transistors. Nat. Rev. Mater.3:17086. 10.1038/natrevmats.2017.86
280
Rnjak J. Li Z. Maitz P. K. M. Wise S. G. Weiss A. S. (2009). Primary human dermal fibroblast interactions with open weave three-dimensional scaffolds prepared from synthetic human elastin. Biomaterials30, 6469–6477. 10.1016/j.biomaterials.2009.08.017
281
Rogal J. Binder C. Kromidas E. Roosz J. Probst C. Schneider S. et al . (2020). WAT-on-a-chip integrating human mature white adipocytes for mechanistic research and pharmaceutical applications. Sci. Rep.10:6666. 10.1038/s41598-020-63710-4
282
Roh T. T. Chen Y. Paul H. T. Guo C. Kaplan D. L. (2019). 3D bioengineered tissue model of the large intestine to study inflammatory bowel disease. Biomaterials225:119517. 10.1016/j.biomaterials.2019.119517
283
Ronaldson-Bouchard K. Vunjak-Novakovic G. (2018). Organs-on-a-chip: a fast track for engineered human tissues in drug development. Cell Stem Cell22, 310–324. 10.1016/j.stem.2018.02.011
284
Rosenbaum A. J. Grande D. A. Dines J. S. (2008). The use of mesenchymal stem cells in tissue engineering: a global assessment. Organogenesis4, 23–27. 10.4161/org.6048
285
Rossi G. Boni A. Guiet R. Girgin M. Kelly R. Lutolf M. (2019). Embryonic organoids recapitulate early heart organogenesis. bioRxiv802181. 10.1101/802181
286
Roth A. Singer T. (2014). The application of 3D cell models to support drug safety assessment: opportunities and challenges. Adv. Drug Deliv. Rev. 69–70, 179–189. 10.1016/j.addr.2013.12.005
287
Rouleau N. Cantley W. L. Liaudanskaya V. Berk A. Du C. Rusk W. et al . (2020). A Long-living bioengineered neural tissue platform to study neurodegeneration. Macromol. Biosci.20:e2000004. 10.1002/mabi.202000004
288
Rouwkema J. Gibbs S. Lutolf M. P. Martin I. Vunjak-Novakovic G. Malda J. (2011). In vitro platforms for tissue engineering: implications for basic research and clinical translation. J. Tissue Eng. Regen. Med.5, e164–e167. 10.1002/term.414
289
Rouwkema J. Koopman B. F. J. M. Blitterswijk C. A. V. Dhert W. J. A. Malda J. (2009). Supply of nutrients to cells in engineered tissues. Biotechnol. Genet. Eng. Rev.26, 163–178. 10.5661/bger-26-163
290
Rowe R. G. Daley G. Q. (2019). Induced pluripotent stem cells in disease modelling and drug discovery. Nat. Rev. Genet.20, 377–388. 10.1038/s41576-019-0100-z
291
Ruskowitz E. R. Deforest C. A. (2018). Photoresponsive biomaterials for targeted drug delivery and 4D cell culture. Nat. Rev. Mater.3:17087. 10.1038/natrevmats.2017.87
292
Ryan A. J. O'Brien F. J. (2015). Insoluble elastin reduces collagen scaffold stiffness, improves viscoelastic properties, and induces a contractile phenotype in smooth muscle cells. Biomaterials73, 296–307. 10.1016/j.biomaterials.2015.09.003
293
Sachs N. de Ligt J. Kopper O. Gogola E. Bounova G. Weeber F. et al . (2018). A living biobank of breast cancer organoids captures disease heterogeneity. Cell172, 373.e10–386.e10. 10.1016/j.cell.2017.11.010
294
Sachs N. Papaspyropoulos A. Zomer-van Ommen D. D. Heo I. Böttinger L. Klay D. et al . (2019). Long-term expanding human airway organoids for disease modeling. EMBO J.38:e100300. 10.15252/embj.2018100300
295
Saidy N. T. Wolf F. Bas O. Keijdener H. Hutmacher D. W. Mela P. et al . (2019). Biologically inspired scaffolds for heart valve tissue engineering via melt electrowriting. Small15:e1900873. 10.1002/smll.201900873
296
Salazar A. Keusgen M. Von Hagen J. (2016). Amino acids in the cultivation of mammalian cells. Amino Acids48, 1161–1171. 10.1007/s00726-016-2181-8
297
Sato T. Vries R. G. Snippert H. J. van de Wetering M. Barker N. Stange D. E. et al . (2009). Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature459, 262–265. 10.1038/nature07935
298
Schimek K. Busek M. Brincker S. Groth B. Hoffmann S. Lauster R. et al . (2013). Integrating biological vasculature into a multi-organ-chip microsystem. Lab Chip13:3588. 10.1039/c3lc50217a
299
Schmid J. Schwarz S. Meier-Staude R. Sudhop S. Clausen-Schaumann H. Schieker M. et al . (2018). A perfusion bioreactor system for cell seeding and oxygen-controlled cultivation of three-dimensional cell cultures. Tissue Eng. Part C Methods24, 585–595. 10.1089/ten.tec.2018.0204
300
Schmidt C. E. Shastri V. R. Vacanti J. P. Langer R. (1997). Stimulation of neurite outgrowth using an electrically conducting polymer. Proc. Natl. Acad. Sci. U.S.A.94, 8948–8953. 10.1073/pnas.94.17.8948
301
Schneeberger K. Spee B. Costa P. Sachs N. Clevers H. Malda J. (2017). Converging biofabrication and organoid technologies: the next frontier in hepatic and intestinal tissue engineering?Biofabrication9:013001. 10.1088/1758-5090/aa6121
302
Schneider H. Muirhead E. E. Zydeck F. A. (1963). Some unusual observations of organoid tissues and blood elements in monolayer cultures. Exp. Cell Res.30, 449–459. 10.1016/0014-4827(63)90322-7
303
Schwarz R. I. Bissell M. J. (1977). Dependence of the differentiated state on the cellular environment: modulation of collagen synthesis in tendon cells. Proc. Natl. Acad. Sci. U.S.A.74, 4453–4457. 10.1073/pnas.74.10.4453
304
Schweiger P. J. Jensen K. B. (2016). Modeling human disease using organotypic cultures. Curr. Opin. Cell Biol.43, 22–29. 10.1016/j.ceb.2016.07.003
305
Seok J. M. Rajangam T. Jeong J. E. Cheong S. Joo S. M. Oh S. J. et al . (2020). Fabrication of 3D plotted scaffold with microporous strands for bone tissue engineering. J. Mater. Chem. B8, 951–960. 10.1039/c9tb02360g
306
Shaban L. Chen Y. Fasciano A. C. Lin Y. Kaplan D. L. Kumamoto C. A. et al . (2018). A 3D intestinal tissue model supports Clostridioides difficile germination, colonization, toxin production and epithelial damage. Anaerobe50, 85–92. 10.1016/j.anaerobe.2018.02.006
307
Shadish J. A. Benuska G. M. DeForest C. A. (2019). Bioactive site-specifically modified proteins for 4D patterning of gel biomaterials. Nat. Mater.18, 1005–1014. 10.1038/s41563-019-0367-7
308
Shamir E. R. Ewald A. J. (2014). Three-dimensional organotypic culture: Experimental models of mammalian biology and disease. Nat. Rev. Mol. Cell Biol.15, 647–664. 10.1038/nrm3873
309
Shastri V. P. Rahman N. Martin I. Langer R. (1999). Application of conductive polymers in bone regeneration. Mater. Res. Soc. Symp. Proc.550, 215–219. 10.1557/proc-550-215
310
Shi Y. Sun L. Wang M. Liu J. Zhong S. Li R. et al . (2020). Vascularized human cortical organoids (vOrganoids) model cortical development in vivo. PLoS Biol.18:e3000705. 10.1371/journal.pbio.3000705
311
Shin W. Wu A. Massidda M. W. Foster C. Thomas N. Lee D.-W. et al . (2019). A Robust longitudinal co-culture of obligate anaerobic gut microbiome with human intestinal epithelium in an anoxic-oxic interface-on-a-chip. Front. Bioeng. Biotechnol.7:13. 10.3389/fbioe.2019.00013
312
Shkarina S. Shkarin R. Weinhardt V. Melnik E. Vacun G. Kluger P. et al . (2018). 3D biodegradable scaffolds of polycaprolactone with silicate-containing hydroxyapatite microparticles for bone tissue engineering: high-resolution tomography and in vitro study. Sci. Rep.8:8907. 10.1038/s41598-018-27097-7
313
Silva S. S. Fernandes E. M. Pina S. Silva-Correia J. Vieira S. Oliveira J. M. et al . (2017). 2.11 polymers of biological origin, in Comprehensive Biomaterials II, ed DucheyneP. (Elsevier), 228-252. Available online at: https://www.sciencedirect.com/referencework/9780081006924/comprehensive-biomaterials-ii
314
Simian M. Bissell M. J. (2017). Organoids: a historical perspective of thinking in three dimensions. J. Cell Biol.216, 31–40. 10.1083/jcb.201610056
315
Simon K. A. Mosadegh B. Minn K. T. Lockett M. R. Mohammady M. R. Boucher D. M. et al . (2016). Metabolic response of lung cancer cells to radiation in a paper-based 3D cell culture system. Biomaterials95, 47–59. 10.1016/j.biomaterials.2016.03.002
316
Simpson K. L. Stoney R. Frese K. K. Simms N. Rowe W. Pearce S. P. et al . (2020). A biobank of small cell lung cancer CDX models elucidates inter- and intratumoral phenotypic heterogeneity. Nat. Cancer1, 437–451. 10.1038/s43018-020-0046-2
317
Sitarski A. M. Fairfield H. Falank C. Reagan M. R. (2018). 3D tissue engineered in vitro models of cancer in bone. ACS Biomater. Sci. Eng.4, 324–336. 10.1021/acsbiomaterials.7b00097
318
Snyder S. DeJulius C. Willits R. K. (2017). Electrical stimulation increases random migration of human dermal fibroblasts. Ann. Biomed. Eng.45, 2049–2060. 10.1007/s10439-017-1849-x
319
Sobrino A. Phan D. T. T. Datta R. Wang X. Hachey S. J. Romero-López M. et al . (2016). 3D microtumors in vitro supported by perfused vascular networks. Sci. Rep.6:31589. 10.1038/srep31589
320
Sontheimer-Phelps A. Hassell B. A. Ingber D. E. (2019). Modelling cancer in microfluidic human organs-on-chips. Nat. Rev. Cancer19, 65–81. 10.1038/s41568-018-0104-6
321
Sood D. Tang-Schomer M. Pouli D. Mizzoni C. Raia N. Tai A. et al . (2019). 3D extracellular matrix microenvironment in bioengineered tissue models of primary pediatric and adult brain tumors. Nat. Commun.10:4529. 10.1038/s41467-019-12420-1
322
Sorushanova A. Delgado L. M. Wu Z. Shologu N. Kshirsagar A. Raghunath R. et al . (2019). The collagen suprafamily: from biosynthesis to advanced biomaterial development. Adv. Mater.31:1801651. 10.1002/adma.201801651
323
Srirussamee K. Mobini S. Cassidy N. J. Cartmell S. H. (2019). Direct electrical stimulation enhances osteogenesis by inducing Bmp2 and Spp1 expressions from macrophages and preosteoblasts. Biotechnol. Bioeng.116, 3421–3432. 10.1002/bit.27142
324
Stucki J. D. Hobi N. Galimov A. Stucki A. O. Schneider-Daum N. Lehr C. M. et al . (2018). Medium throughput breathing human primary cell alveolus-on-chip model. Sci. Rep.8:14359. 10.1038/s41598-018-32523-x
325
Subbiah R. Hipfinger C. Tahayeri A. Athirasala A. Horsophonphong S. Thrivikraman G. et al . (2020). 3D printing of microgel-loaded modular microcages as instructive scaffolds for tissue engineering. Adv. Mater.32:2001736. 10.1002/adma.202001736
326
Sun H. Jia Y. Dong H. Dong D. Zheng J. (2020). Combining additive manufacturing with microfluidics: an emerging method for developing novel organs-on-chips. Curr. Opin. Chem. Eng.28, 1–9. 10.1016/j.coche.2019.10.006
327
Takahashi K. Tanabe K. Ohnuki M. Narita M. Ichisaka T. Tomoda K. et al . (2007). Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell131, 861–872. 10.1016/j.cell.2007.11.019
328
Takebe T. Sekine K. Enomura M. Koike H. Kimura M. Ogaeri T. et al . (2013). Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature499, 481–484. 10.1038/nature12271
329
Takebe T. Wells J. M. (2019). Organoids by design. Science364, 956–959. 10.1126/science.aaw7567
330
Tam R. Y. Yockell-Lelièvre J. Smith L. J. Julian L. M. Baker A. E. G. Choey C. et al . (2019). Rationally designed 3D hydrogels model invasive lung diseases enabling high-content drug screening. Adv. Mater.31:1806214. 10.1002/adma.201806214
331
Tang M. Xie Q. Gimple R. C. Zhong Z. Tam T. Tian J. et al . (2020). Three-dimensional bioprinted glioblastoma microenvironments model cellular dependencies and immune interactions. Cell Res.30, 833–853. 10.1038/s41422-020-0338-1
332
Tang-Schomer M. D. White J. D. Tien L. W. Schmitt L. I. Valentin T. M. Graziano D. J. et al . (2014). Bioengineered functional brain-like cortical tissue. Proc. Natl. Acad. Sci. U.S.A.111, 13811–13816. 10.1073/pnas.1324214111
333
Taylor M. W. Taylor M. W. (2014). A history of cell culture, in Viruses and Man: A History of Interactions (Cham: Springer International Publishing), 41–52. 10.1007/978-3-319-07758-1_3
334
Terrell J. A. Jones C. G. Kabandana G. K. M. Chen C. (2020). From cells-on-a-chip to organs-on-a-chip: Scaffolding materials for 3D cell culture in microfluidics. J. Mater. Chem. B8, 6667–6685. 10.1039/d0tb00718h
335
Theocharis A. D. Skandalis S. S. Gialeli C. Karamanos N. K. (2016). Extracellular matrix structure. Adv. Drug Deliv. Rev.97, 4–27. 10.1016/j.addr.2015.11.001
336
Theodoridis K. Aggelidou E. Vavilis T. Manthou M. E. Tsimponis A. Demiri E. C. et al . (2019). Hyaline cartilage next generation implants from adipose-tissue-derived mesenchymal stem cells: comparative study on 3D-printed polycaprolactone scaffold patterns. J. Tissue Eng. Regen. Med.13, 342–355. 10.1002/term.2798
337
Tijore A. Irvine S. A. Sarig U. (2020). Contact guidance for cardiac tissue engineering using 3D bioprinted gelatin patterned hydrogel. Biofabrication10:025003. 10.1088/1758-5090/aaa15d
338
Tong X. F. Zhao F. Q. Ren Y. Z. Zhang Y. Cui Y. L. Wang Q. S. (2018). Injectable hydrogels based on glycyrrhizin, alginate, and calcium for three-dimensional cell culture in liver tissue engineering. J. Biomed. Mater. Res. Part A106, 3292–3302. 10.1002/jbm.a.36528
339
Trautmann A. Lemke H.-D. Hellmann R. Walther T. Rüth M. (2018). Large 3D direct laser written scaffolds for tissue engineering applications, in Nanophotonics Australasia 2017, eds. ChonJ. W. M.JiaB. (SPIE), 186. 10.1117/12.2282417
340
Trietsch S. J. Naumovska E. Kurek D. Setyawati M. C. Vormann M. K. Wilschut K. J. et al . (2017). Membrane-free culture and real-time barrier integrity assessment of perfused intestinal epithelium tubes. Nat. Commun.8:262. 10.1038/s41467-017-00259-3
341
Trisno S. L. Philo K. E. D. McCracken K. W. Catá E. M. Ruiz-Torres S. Rankin S. A. et al . (2018). Esophageal organoids from human pluripotent stem cells delineate Sox2 functions during esophageal specification. Cell Stem Cell23, 501.e7–515.e7. 10.1016/j.stem.2018.08.008
342
Tsakmaki A. Fonseca Pedro P. Bewick G. A. (2017). 3D intestinal organoids in metabolic research: virtual reality in a dish. Curr. Opin. Pharmacol.37, 51–58. 10.1016/j.coph.2017.09.003
343
Tsamandouras N. Chen W. L. K. Edington C. D. Stokes C. L. Griffith L. G. Cirit M. (2017). Integrated gut and liver microphysiological systems for quantitative in vitro pharmacokinetic studies. AAPS J.19, 1499–1512. 10.1208/s12248-017-0122-4
344
Tziveleka L. A. Sapalidis A. Kikionis S. Aggelidou E. Demiri E. Kritis A. et al . (2020). Hybrid sponge-like scaffolds based on ulvan and gelatin: design, characterization and evaluation of their potential use in bone tissue engineering. Materials13, 1–20. 10.3390/ma13071763
345
Uchino T. Takezawa T. Ikarashi Y. (2009). Reconstruction of three-dimensional human skin model composed of dendritic cells, keratinocytes and fibroblasts utilizing a handy scaffold of collagen vitrigel membrane. Toxicol. Vitr.23, 333–337. 10.1016/j.tiv.2008.12.003
346
Ullah M. Liu D. D. Thakor A. S. (2019). Mesenchymal stromal cell homing: mechanisms and strategies for improvement. iScience15, 421–438. 10.1016/j.isci.2019.05.004
347
Ullah S. Chen X. (2020). Fabrication, applications and challenges of natural biomaterials in tissue engineering. Appl. Mater. Today20:100656. 10.1016/j.apmt.2020.100656
348
Ullah S. Zainol I. Chowdhury S. R. Fauzi M. B. (2018). Development of various composition multicomponent chitosan/fish collagen/glycerin 3D porous scaffolds: effect on morphology, mechanical strength, biostability and cytocompatibility. Int. J. Biol. Macromol.111, 158–168. 10.1016/j.ijbiomac.2017.12.136
349
Urbischek M. Rannikmae H. Foets T. Ravn K. Hyvönen M. de la Roche M. (2019). Organoid culture media formulated with growth factors of defined cellular activity. Sci. Rep.9:6193. 10.1038/s41598-019-42604-0
350
Vacanti C. A. (2006). The history of tissue engineering. J. Cell. Mol. Med.10, 569–576. 10.1111/j.1582-4934.2006.tb00421.x
351
van der Helm M. W. Henry O. Y. F. Bein A. Hamkins-Indik T. Cronce M. J. Leineweber W. D. et al . (2019). Non-invasive sensing of transepithelial barrier function and tissue differentiation in organs-on-chips using impedance spectroscopy. Lab Chip19, 452–463. 10.1039/C8LC00129D
352
Vatine G. D. Barrile R. Workman M. J. Sances S. Barriga B. K. Rahnama M. et al . (2019). Human iPSC-derived blood-brain barrier chips enable disease modeling and personalized medicine applications. Cell Stem Cell24, 995.e6–1005.e6. 10.1016/j.stem.2019.05.011
353
Vernetti L. Gough A. Baetz N. Blutt S. Broughman J. R. Brown J. A. et al . (2017). Functional coupling of human microphysiology systems: intestine, liver, kidney proximal tubule, blood-brain barrier and skeletal muscle. Sci. Rep.7:42296. 10.1038/srep42296
354
Vidal S. E. L. Tamamoto K. A. Nguyen H. Abbott R. D. Cairns D. M. Kaplan D. L. (2019). 3D biomaterial matrix to support long term, full thickness, immuno-competent human skin equivalents with nervous system components. Biomaterials198, 194–203. 10.1016/j.biomaterials.2018.04.044
355
Villasante A. Sakaguchi K. Kim J. Cheung N. K. Nakayama M. Parsa1 H. et al . (2017). Vascularized tissue-engineered model for studying drug resistance in neuroblastoma. Theranostics7:42296. 10.7150/thno.20730
356
Villenave R. Wales S. Q. Hamkins-Indik T. Papafragkou E. Weaver J. C. Ferrante T. C. et al . (2017). Human gut-on-a-chip supports polarized infection of coxsackie B1 virus in vitro. PLoS ONE12:e0169412. 10.1371/journal.pone.0169412
357
Vining K. H. Mooney D. J. (2017). Mechanical forces direct stem cell behaviour in development and regeneration. Nat. Rev. Mol. Cell Biol.18, 728–742. 10.1038/nrm.2017.108
358
Visone R. Talò G. Lopa S. Rasponi M. Moretti M. (2018). Enhancing all-in-one bioreactors by combining interstitial perfusion, electrical stimulation, on-line monitoring and testing within a single chamber for cardiac constructs. Sci. Rep.8:16944. 10.1038/s41598-018-35019-w
359
Vodovnik L. Miklavči,č D. Serša G. (1992). Modified cell proliferation due to electrical currents. Med. Biol. Eng. Comput.30, CE21–CE28. 10.1007/BF02446174
360
Vyas D. Baptista P. M. Brovold M. Moran E. Gaston B. Booth C. et al . (2018). Self-assembled liver organoids recapitulate hepatobiliary organogenesis in vitro. Hepatology67, 750–761. 10.1002/hep.29483
361
Wan A. M. D. Inal S. Williams T. Wang K. Leleux P. Estevez L. et al . (2015). 3D conducting polymer platforms for electrical control of protein conformation and cellular functions. J. Mater. Chem. B3, 5040–5048. 10.1039/c5tb00390c
362
Wan X. Li Z. Ye H. Cui Z. (2016). Three-dimensional perfused tumour spheroid model for anti-cancer drug screening. Biotechnol. Lett.38, 1389–1395. 10.1007/s10529-016-2035-1
363
Wang B. Yang W. McKittrick J. Meyers M. A. (2016). Keratin: structure, mechanical properties, occurrence in biological organisms, and efforts at bioinspiration. Prog. Mater. Sci.76, 229–318. 10.1016/j.pmatsci.2015.06.001
364
Wang C. Koo S. Park M. Vangelatos Z. Hoang P. Conklin B. R. et al . (2020). Maladaptive contractility of 3D human cardiac microtissues to mechanical nonuniformity. Adv. Healthc. Mater.9:1901373. 10.1002/adhm.201901373
365
Wang D. A. Varghese S. Sharma B. Strehin I. Fermanian S. Gorham J. et al . (2007). Multifunctional chondroitin sulphate for cartilage tissue-biomaterial integration. Nat. Mater.6, 385–392. 10.1038/nmat1890
366
Wang X. Guo C. Chen Y. Tozzi L. Szymkowiak S. Li C. et al . (2020). Developing a self-organized tubulogenesis model of human renal proximal tubular epithelial cells in vitro. J. Biomed. Mater. Res. Part A108, 795–804. 10.1002/jbm.a.36858
367
Wang X. Li W. Kumar V. (2006). A method for solvent-free fabrication of porous polymer using solid-state foaming and ultrasound for tissue engineering applications. Biomaterials27, 1924–1929. 10.1016/j.biomaterials.2005.09.029
368
Wang Y. I. Abaci H. E. Shuler M. L. (2017). Microfluidic blood-brain barrier model provides in vivo-like barrier properties for drug permeability screening. Biotechnol. Bioeng.114, 184–194. 10.1002/bit.26045
369
Wevers N. R. Kasi D. G. Gray T. Wilschut K. J. Smith B. van Vught R. et al . (2018). A perfused human blood–brain barrier on-a-chip for high-throughput assessment of barrier function and antibody transport. Fluids Barriers CNS15:23. 10.1186/s12987-018-0108-3
370
Wikswo J. P. Curtis E. L. Eagleton Z. E. Evans B. C. Kole A. Hofmeister L. H. et al . (2013). Scaling and systems biology for integrating multiple organs-on-a-chip. Lab Chip13, 3496–3511. 10.1039/c3lc50243k
371
Willerth S. M. Rader A. Sakiyama-Elbert S. E. (2008). The effect of controlled growth factor delivery on embryonic stem cell differentiation inside fibrin scaffolds. Stem Cell Res.1, 205–218. 10.1016/j.scr.2008.05.006
372
Wolter J. R. (1967). Proliferating pigment epithelium: producing a simple organoid structure in the subrentinal space of a human eye. Arch. Ophthalmol.77, 651–654. 10.1001/archopht.1967.00980020653016
373
Wu L. Magaz A. Huo S. Darbyshire A. Loizidou M. Emberton M. et al . (2020). Human airway-like multilayered tissue on 3D-TIPS printed thermoresponsive elastomer/collagen hybrid scaffolds. Acta Biomater.113, 177–195. 10.1016/j.actbio.2020.07.013
374
Wylie R. G. Ahsan S. Aizawa Y. Maxwell K. L. Morshead C. M. Shoichet M. S. (2011). Spatially controlled simultaneous patterning of multiple growth factors in three-dimensional hydrogels. Nat. Mater.10, 799–806. 10.1038/nmat3101
375
Xu G. Ding Z. Lu Q. Zhang X. Zhou X. Xiao L. et al . (2020). Electric field-driven building blocks for introducing multiple gradients to hydrogels. Protein Cell11, 267–285. 10.1007/s13238-020-00692-z
376
Xu R. Boudreau A. Bissell M. J. (2009). Tissue architecture and function: dynamic reciprocity via extra- and intra-cellular matrices. Cancer Metastasis Rev.28, 167–176. 10.1007/s10555-008-9178-z
377
Yan H. H. N. Siu H. C. Law S. Ho S. L. Yue S. S. K. Tsui W. Y. et al . (2018). A comprehensive human gastric cancer organoid biobank captures tumor subtype heterogeneity and enables therapeutic screening. Cell Stem Cell 23, 882.e11–897.e11. 10.1016/j.stem.2018.09.016
378
Yang Z. Xiong H.-R. (2012). Culture Conditions and Types of Growth Media for Mammalian Cells, in Biomedical Tissue Culture, eds Ceccherini-NelliL.MatteoliB. (InTech). 10.5772/52301
379
Yao T. Asayama Y. (2017). Animal-cell culture media: History, characteristics, and current issues. Reprod. Med. Biol.16, 99–117. 10.1002/rmb2.12024
380
Yeste J. García-Ramírez M. Illa X. Guimerà A. Hernández C. Simó R. et al . (2018). A compartmentalized microfluidic chip with crisscross microgrooves and electrophysiological electrodes for modeling the blood-retinal barrier. Lab Chip18, 95–105. 10.1039/c7lc00795g
381
Yin L. Du G. Zhang B. Zhang H. Yin R. Zhang W. et al . (2020). Efficient drug screening and nephrotoxicity assessment on co-culture microfluidic kidney chip. Sci. Rep.10:6568. 10.1038/s41598-020-63096-3
382
Yin X. Mead B. E. Safaee H. Langer R. Karp J. M. Levy O. (2016). Engineering stem cell organoids. Cell Stem Cell18, 25–38. 10.1016/j.stem.2015.12.005
383
Yip D. Cho C. H. (2013). A multicellular 3D heterospheroid model of liver tumor and stromal cells in collagen gel for anti-cancer drug testing. Biochem. Biophys. Res. Commun.433, 327–332. 10.1016/j.bbrc.2013.03.008
384
Yu J. Peng S. Luo D. March J. C. (2012). In vitro 3D human small intestinal villous model for drug permeability determination. Biotechnol. Bioeng.109, 2173–2178. 10.1002/bit.24518
385
Zbinden A. Marzi J. Schlünder K. Probst C. Urbanczyk M. Black S. et al . (2020). Non-invasive marker-independent high content analysis of a microphysiological human pancreas-on-a-chip model. Matrix Biol. 85–86, 205–220. 10.1016/j.matbio.2019.06.008
386
Zelikin A. N. Lynn D. M. Farhadi J. Martin I. Shastri V. Langer R. (2002). Erodible conducting polymers for potential biomedical applications. Angew. Chemie Int. Ed.41, 141–144. 10.1002/1521-3773(20020104)41:1<141::AID-ANIE141>3.0.CO;2-V
387
Zhang B. Montgomery M. Chamberlain M. D. Ogawa S. Korolj A. Pahnke A. et al . (2016a). Biodegradable scaffold with built-in vasculature for organ-on-a-chip engineering and direct surgical anastomosis. Nat. Mater.15, 669–678. 10.1038/nmat4570
388
Zhang B. Radisic M. (2017). Organ-on-a-chip devices advance to market. Lab Chip17, 2395–2420. 10.1039/c6lc01554a
389
Zhang C. Zhao Z. Abdul Rahim N. A. van Noort D. Yu H. (2009). Towards a human-on-chip: culturing multiple cell types on a chip with compartmentalized microenvironments. Lab Chip9:3185. 10.1039/b915147h
390
Zhang J. Neoh K. G. Kang E. (2018). Electrical stimulation of adipose-derived mesenchymal stem cells and endothelial cells co-cultured in a conductive scaffold for potential orthopaedic applications. J. Tissue Eng. Regen. Med.12, 878–889. 10.1002/term.2441
391
Zhang J. Wehrle E. Adamek P. Paul G. R. Qin X. H. Rubert M. et al . (2020). Optimization of mechanical stiffness and cell density of 3D bioprinted cell-laden scaffolds improves extracellular matrix mineralization and cellular organization for bone tissue engineering. Acta Biomater.114, 307–322. 10.1016/j.actbio.2020.07.016
392
Zhang S. Wang H. (2019). Current progress in 3D bioprinting of tissue analogs. SLAS Technol.24, 70–78. 10.1177/2472630318799971
393
Zhang W. Wang Z. Xie C. Wang X. Luo F. Hong M. et al . (2019). Scaffold with micro/macro-architecture for myocardial alignment engineering into complex 3D cell patterns. Adv. Healthc. Mater.8:1901015. 10.1002/adhm.201901015
394
Zhang Y. Wang Q.-S. Yan K. Qi Y. Wang G.-F. Cui Y.-L. (2016c). Preparation, characterization, and evaluation of genipin crosslinked chitosan/gelatin three-dimensional scaffolds for liver tissue engineering applications. J. Biomed. Mater. Res. Part A104, 1863–1870. 10.1002/jbm.a.35717
395
Zhang Y. S. Aleman J. Shin S. R. Kilic T. Kim D. Shaegh S. A. M. et al . (2017). Multisensor-integrated organs-on-chips platform for automated and continual in situ monitoring of organoid behaviors. Proc. Natl. Acad. Sci. U.S.A.114, E2293–E2302. 10.1073/pnas.1612906114
396
Zhang Y. S. Arneri A. Bersini S. Shin S. R. Zhu K. Goli-Malekabadi Z. et al . (2016b). Bioprinting 3D microfibrous scaffolds for engineering endothelialized myocardium and heart-on-a-chip. Biomaterials110, 45–59. 10.1016/j.biomaterials.2016.09.003
397
Zhu W. Ye T. Lee S. J. Cui H. Miao S. Zhou X. et al . (2018). Enhanced neural stem cell functions in conductive annealed carbon nanofibrous scaffolds with electrical stimulation. Nanomedicine Nanotechnology, Biol. Med.14, 2485–2494. 10.1016/j.nano.2017.03.018
398
Zhu Y. Wang L. Yin F. Yu Y. Wang Y. Liu H. et al . (2017). A hollow fiber system for simple generation of human brain organoids. Integr. Biol.9, 774–781. 10.1039/c7ib00080d
399
Zidarič T. Milojević M. Gradišnik L. Kleinschek K. S. Maver U. Maver T. (2020). Polysaccharide-based bioink formulation for 3D bioprinting of an in vitro model of the human dermis. Nanomaterials10:733. 10.3390/nano10040733
400
Zohar B. Blinder Y. Mooney D. J. Levenberg S. (2018). Flow-induced vascular network formation and maturation in three-dimensional engineered tissue. ACS Biomater. Sci. Eng.4, 1265–1271. 10.1021/acsbiomaterials.7b00025
Summary
Keywords
tissue engineering, scaffold, hydrogel, 3D biology, organoid, organ-on-a-chip
Citation
Moysidou C-M, Barberio C and Owens RM (2021) Advances in Engineering Human Tissue Models. Front. Bioeng. Biotechnol. 8:620962. doi: 10.3389/fbioe.2020.620962
Received
24 October 2020
Accepted
22 December 2020
Published
28 January 2021
Volume
8 - 2020
Edited by
Davide Ricci, University of Genoa, Italy
Reviewed by
Moo-Yeal Lee, Cleveland State University, United States; Giorgia Imparato, Center for Advanced Biomaterials for Healthcare (IIT), Italy
Updates
Copyright
© 2021 Moysidou, Barberio and Owens.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Róisín Meabh Owens rmo37@cam.ac.uk
This article was submitted to Biomaterials, a section of the journal Frontiers in Bioengineering and Biotechnology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.