- 1 Department of Pathobiological Sciences, University of Wisconsin–Madison, Madison, WI, USA
- 2 Department of Animal Sciences, University of Wisconsin–Madison, Madison, WI, USA
- 3 Laboratory of Bacterial Genomics, University of Wisconsin–Madison, Madison, WI, USA
More than 1.8 million people die annually from infection with Mycobacterium tuberculosis, the causative agent of tuberculosis. The ability of M. tuberculosis to obtain and distribute micronutrients, including biometals, is known to play a role in its intracellular survival and virulence within a host. Techniques to detect elemental distributions within M. tuberculosis cells have previously been limited to bulk detection methods or low-resolution analyses. Here, we present a method for determining the elemental distribution within M. tuberculosis on a single-cell level, at high (individual nanometer) resolution, using scanning transmission electron microscopy (STEM) in concert with energy-dispersive X-ray spectroscopy (EDS). Results revealed the presence of large polyphosphate granules in all strains of Mycobacteria tested. These persisted even through starvation conditions, and might play a role connected to elemental homeostasis in M. tuberculosis. Associated with the polyphosphate granules were micronutrients such as calcium and magnesium. In addition, we expanded the technique beyond Mycobacteria to show that STEM and EDS could be used as a simple screen to detect the presence or absence of concentrated elements on a single-cell level within all six other bacterial types tested, with minimal processing to the bacteria. Overall, we believe that this technique represents a first step in developing a better understanding of the role that components of the intracellular milieu, including polyphosphates and biometals, play in the pathogenesis of M. tuberculosis, with potential future applications for in vivo analysis.
Introduction
The accumulation as well as the detoxification of a number of elemental micronutrients is known to play an important role in the interactions between human hosts and invading pathogenic microbes (Agranoff and Krishna, 1998). For those pathogens that live within host cells, such as the intracellular human pathogen Mycobacterium tuberculosis, this can be particularly significant to pathogenesis (Botella et al., 2012). Early on, the availability of iron was shown to play a major role in the survival of pathogens in their microenvironments, including between macrophages and M. tuberculosis (De Voss et al., 2000; Schaible and Kaufmann, 2004; Rodriguez and Smith, 2006). More recently, the role of copper toxicity and extracellular export was shown to play a significant role in mycobacterial virulence (Ward et al., 2010; Wolschendorf et al., 2011). Available approaches to estimate intracellular levels of metals count mainly on indirect assays that use a large number of cells (e.g., fluorescent dyes or neutron activation; Vanhoe et al., 1989; Versieck, 1994; Barrios, 2006). Recently, advances in cellular techniques allowed the investigation of metal ion components of the mycobacterial phagosome on a single-cell level rather than the chemical analysis of cellular lysate. Using Synchrotron X-ray Fluorescence (SXRF) microscopy, the metal composition of the phagosomes formed by mycobacteria were analyzed (Wagner et al., 2005) with little differentiation between mycobacteria and its phagosome. Fortunately, a powerful technology based on scanning transmission electron microscopy (STEM) is currently available and could offer a higher resolution analysis of metal ions than SXRF in bacteria (Stewart et al., 1980; Gonzalez and Jensen, 1998). In this report, we developed an assay based on STEM to examine the elemental composition of mycobacterial species on a single-cell level.
Mycobacterial infections are caused by a group of highly successful pathogens that threaten both humans and animals. More than 1.8 million people die annually from infection with M. tuberculosis, the causative agent of tuberculosis (Corbett et al., 2003). Interestingly, the genome of the M. tuberculosis H37Rv strain indicated the presence of 28 metal transporters, among which 17 transporters were also encoded in the genome of M. leprae, suggesting their important role for intracellular survival (Agranoff and Krishna, 2004). Additionally, a non-specific transporter MntH belonging to the Nramp (natural resistance-associated macrophage protein) family of proteins was found to transport divalent cations such as Mn and Fe (Agranoff et al., 1999). Transcription of mntH was also found to be induced in M. tuberculosis during THP-1 macrophage infection and in response to low levels of Cu (0–70 μM range). However, the deletion of mntH did not affect the survival of the mutant M. tb in mouse lungs (Domenech et al., 2002), indicating that MntH was not an essential cation transporter and that there may be another transporter more essential to the survival of M. tb. Recently, a newly discovered gene in the M. tb genome, mymT, was found to encode a Cu metallothionein that protects M. tb against Cu toxicity by sequestering copper ions (Gold et al., 2008). However, mouse infections did not show a role for mymT in M. tb virulence, suggesting redundancy of the mymT gene (Gold et al., 2008). Generally, understanding the role of metal ions in intracellular enzymatic activities that control the phagosome microenvironments will help in understanding tuberculosis pathogenesis.
Despite the known importance of elemental interplay within bacterial pathogenesis, our knowledge of elemental concentrations and distributions within bacterial cells has previously been limited by the available technology. Recent advances in electron microscopy, including the development of scanning transmission electron microscopes capable of conducting elemental analysis via energy-dispersive X-ray spectroscopy (EDS), have allowed for the visualization of elemental concentrations at the single-cell level. To our knowledge, our group is the first to image M. tuberculosis using aberration corrected STEM in concert with high resolution EDS. Results show that many elements are not concentrated enough within mycobacterial cells to be quantified on a single-cell level using currently available technology. However, large polyphosphate granules were clearly visible in all mycobacteria tested, including M. tuberculosis, the bovine pathogen M. avium subsp. paratuberculosis, and the non-pathogenic strain M. smegmatis. Further, we show the utility of the STEM technique as a quick screen for profiling an elemental map for a variety of bacterial types and conditions. The data we present suggest that STEM microscopy could provide novel insights into the intracellular concentrations and localization of a variety of elements within mycobacteria.
Materials and Methods
Bacterial Cultures
A list of all bacterial strains and growth conditions used in this study is available in Table 1. All strains were grown at 37°C to late-log phase (OD600 = 1.2). Mycobacterial strains including an isogenic strain lacking the copper transporter CtpV (Ward et al., 2010) were grown in 7H9 liquid media (BD, Franklin Lakes, NJ, USA), supplemented with 10% albumin-catalase-dextrose (ADC) and, in the case of M. avium subsp. paratuberculosis, 2 μg/mL Mycobactin J (Allied Monitor, Fayette, MO, USA). In some experiments, CuCl2 was added to the growing media of M. smegmatis cultures at the final concentrations of 500 μM and 5 mM (Ward et al., 2008). In other variations, M. tuberculosis H37Rv cells were resuspended in ddH2O for 1 or 21 days, an established model for complete nutrient starvation (Parish, 2003). Escherichia coli, Staphylococcus aureus, and Bacillus subtilis were grown in liquid LB media (Life Technologies, Grand Island, NY, USA). Lactobacillus casei was grown in De Man–Rogosa–Sharpe (MRS) broth (Sigma-Aldrich, St. Louis, MO, USA). Neisseria lactamica cultures were grown in gonococcal base liquid medium with Kellogg’s supplements and 0.042% NaHCO3 (Dillard and Hackett, 2005). Helicobacter pylori cultures were grown in tryptic soy agar (MP Biomedicals, Solon, OH, USA) with defibrinated sheep blood in a microaerophilic chamber.
Sample Preparation
After culture growth to late-log phase, bacterial cells were fixed by pelleting and resuspension in a fixation buffer consisting of 4% paraformaldehyde and 2.5% glutaraldehyde in 0.1 M phosphate buffer according to our approved biosafety protocol and detailed before (Woo et al., 2007). Sodium cacodylate buffer (0.2 M), pH 7.2, was used for the phosphate-free buffer experiment. Cells were fixed for 1 h at room temperature prior to preparation for sample imaging.
Sample Imaging
Gold 200 mesh grids (Electron Microscopy Sciences, Hartfield, PA, USA) were coated with a thin layer (∼50 nm) of carbon as detailed before (Meyer et al., 2010; Albrecht et al., 2011). The carbon-filmed gold grids were then coated with poly-L-lysine. Fixed whole bacteria were placed on the grid for ten minutes, followed by two washes with double distilled water to remove unattached bacteria. The grids were dried via an ethanol series, ending with 100% sieve-dried ethanol, after which they were dried in a critical-point dryer (Tousimis Research Corp., Rockville, MD, USA). These whole-mounted bacterial samples were imaged at 200 kV using the FEI Titan 80-200 aberration corrected STEM (FEI, Hillsboro, OR, USA). Elemental spectra, utilizing EDS, were collected at a 15 s intervals also at 200 kV. Data analyses on collected spectra were performed using TIA software from FEI. Elemental spectra were collected from at least two points per cell, with a minimum of two bacteria per replicate imaged from at least two individual biological replicates. All imaging was performed via the UW Laboratory for Biological and Biomaterials Preparation, Imaging, and Characterization (BBPIC), and the Wisconsin Microscopy and Characterization Center.
Results
STEM imaging of M. tuberculosis
Early analysis of M. tuberculosis H37Rv indicated its ability to export copper through the induction of the Cu sensitive operon (cso; Ward et al., 2008). Further analysis indicated that a member of the cso, namely, ctpV, encodes an efflux pump for intracellular copper that is necessary for the survival of M. tuberculosis during infection (Ward et al., 2010). To better estimate intracellular levels of Cu on single M. tuberculosis bacilli, we employed a novel protocol for STEM. Analytical STEM has previously been shown to identify divalent cations in samples with sensitivity levels up to 1–10 Cu atoms in 0.2–0.3 nM sections (Cosgriffa et al., 2005; Wesenberg et al., 2007). Use of the aberration corrected STEM optics permitting sub-angstrom beam diameters and increased beam current should improve upon these previous levels. A preliminary protocol for micro-sectioning mycobacterial bacilli in presence or absence of toxic levels of Cu (up to 500 μM) did not yield reproducible elemental Cu maps of M. tuberculosis or M. smegmatis strains (unpublished data). Accordingly, we modified our protocol to use whole-mounted bacterial cells without sectioning. Interestingly, whole-mounted cells of M. tuberculosis and M. smegmatis bacteria yielded reproducible elemental maps. The most notable feature of the imaged M. tuberculosis was the presence of bright, electron dense areas within the cells, with sizes ranging from approximately 50 to 200 nm (Figure 1).
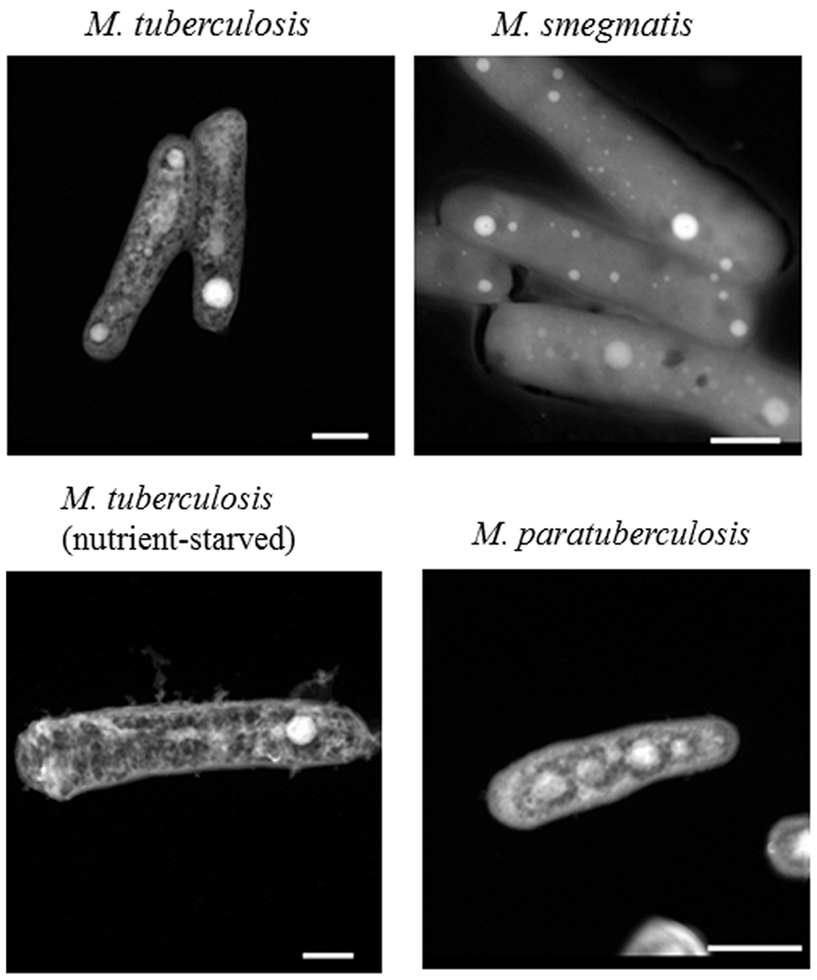
Figure 1. Polyphosphate granules in Mycobacteria. Mycobacterial isolates were fixed as whole-cell mounted, and imaged using an FEI Titan 80-200 STEM. Selected images show representative samples from at least two biological replicates, with multiple bacteria imaged within each replicate. Nutrient-starved M. tuberculosis was incubated in water for 21 days, and all other images are from bacteria grown to late-log phase in 7H9 liquid media. Scale bars (lower right) represent 500 nm.
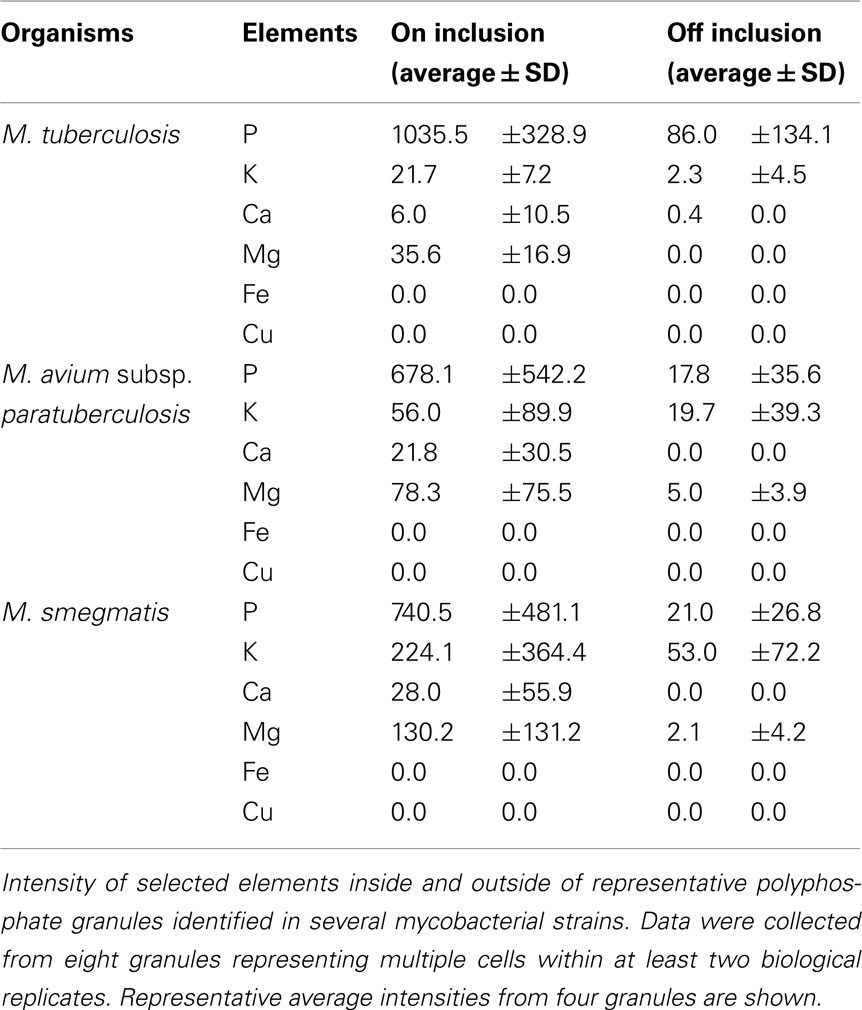
Further quantitative analysis of the collected images using an EDS protocol identified the presence of high levels of phosphorus, as well as lower levels of other elements such as magnesium, calcium, and potassium (Table 2). These granules were present in both virulent (M. tuberculosis) and avirulent (M. smegmatis) strains of mycobacteria. The identified elements (P, Mg, Ca, K) were concentrated mainly in granules and were generally either not detected or detected at much lower (background) levels outside of the granules (Table 2). Significant variations in element signals were observed among different granules contributing to the increased standard deviations. Surprisingly, despite supplying M. smegmatis cultures with high levels of CuCl2 (500 μM and 5 mM), STEM analysis did not detect significant levels of intracellular Cu above the background level. Overall, our protocol for STEM analysis was able to profile intracellular elements concentrated within granules but not dispersed inside cells.
Polyphosphate Granules Inside Mycobacterial Bacilli
Previously, M. tuberculosis has been shown to contain polyphosphate granules (Winder and Denneny, 1957), and our elemental analysis with EDS confirmed that the electron dense areas consisted mainly of high concentrations of phosphate with the presence of other elements (Figure 2). To further examine the presence of such granules in other species of Mycobacteria, we examined a laboratory isolate of the bovine pathogen, M. avium subsp. paratuberculosis (M. paratuberculosis) that is distantly related to M. tuberculosis. As expected, polyphosphate granules were also detected within bacilli of M. paratuberculosis with a similar elemental distribution (inside/outside granule) to that present in M. tuberculosis (Figure 1). To ensure that granule visualization was not an artifact of the paraformaldehyde fixation process, M. paratuberculosis was imaged without the fixative step, which is required for deactivation of BSL-3 pathogenic bacteria. However, presence or absence of the fixative had no effect on polyphosphate granule formation (Figure 1). Finally, cultures of M. tuberculosis that were incubated in ddH2O for 1 or 21 days(a model of nutrient starvation) did not show any significant alteration of their granule structure or composition (Figure 1), a further indication of the stability of the polyphosphate granules.
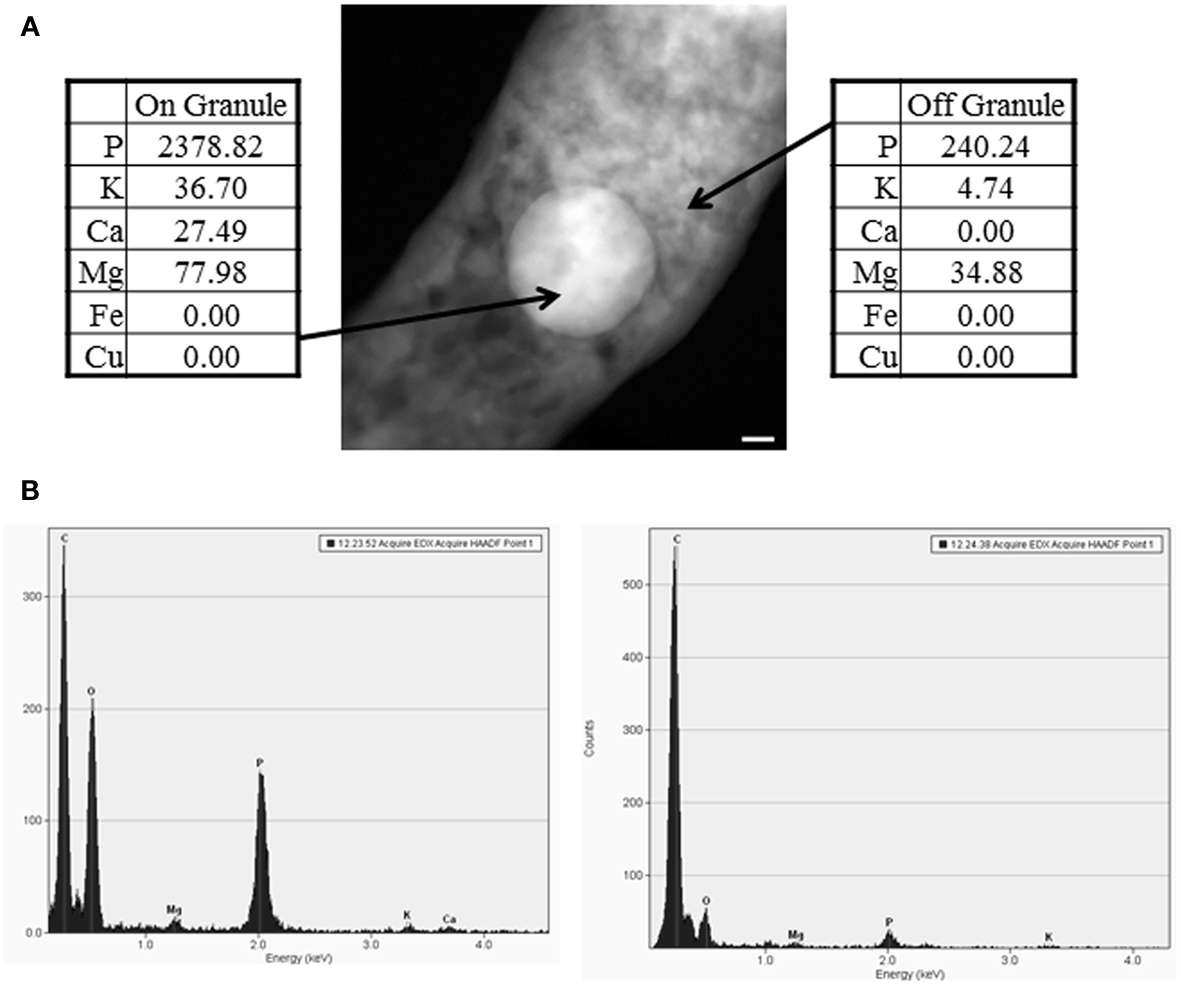
Figure 2. EDS analysis of polyphosphate granules. (A) A representative image of a polyphosphate granule with EDS analysis from wild-type M. tuberculosis. Scale bar (lower right of EM) represents 50 nm. (B) Spectra on the left and right represent EDS spectra on and off the granule, respectively. Data were collected from a minimum of two bacteria from at least two separate biological replicates, and data shown are representative. Elements presented in the spectra include C, for carbon, O, for oxygen, Ca for calcium, Na for sodium, Mg for magnesium, P for phosphorus, and K for potassium.
Because polyphosphate granules could play many roles for mycobacterial survival, we opted to examine whether these granules were present inside the cytoplasm or were attached to the membrane. For this set of experiments, we further examined M. smegmatis samples using multiple images from a 15° stage tilt to create a three-dimensional (3D) image. This analysis confirmed that the polyphosphate granules are located within the bacterial cytoplasm, and are not membrane-associated. A video clip illustrating the 3D image of M. smegmatis is provided as supplemental data. In general, granules composed mainly of polyphosphates as well as Mg, Ca, and K were detected in all examined mycobacterial strains.
Polyphosphates in Other Bacteria
To determine whether the described STEM technique could be used for elemental analysis outside of Mycobacteria, we additionally imaged other bacterial strains representing Gram-negative (E. coli, N. lactamica, H. pylori) and Gram-positive (L. casei, B. subtilis, S. aureus). Although all strains were able to be imaged using the same whole-cell mounting technique as M. tuberculosis, most lacked discrete polyphosphate granules, with the exception of L. casei (Figure 3). The L. casei samples showed multiple polyphosphate granules per cell, similar to Mycobacteria. So far, STEM analysis indicated the presence of discrete granules composed mainly of polyphosphates in bacilli that belong to either Mycobacteria or Lactobacilli.
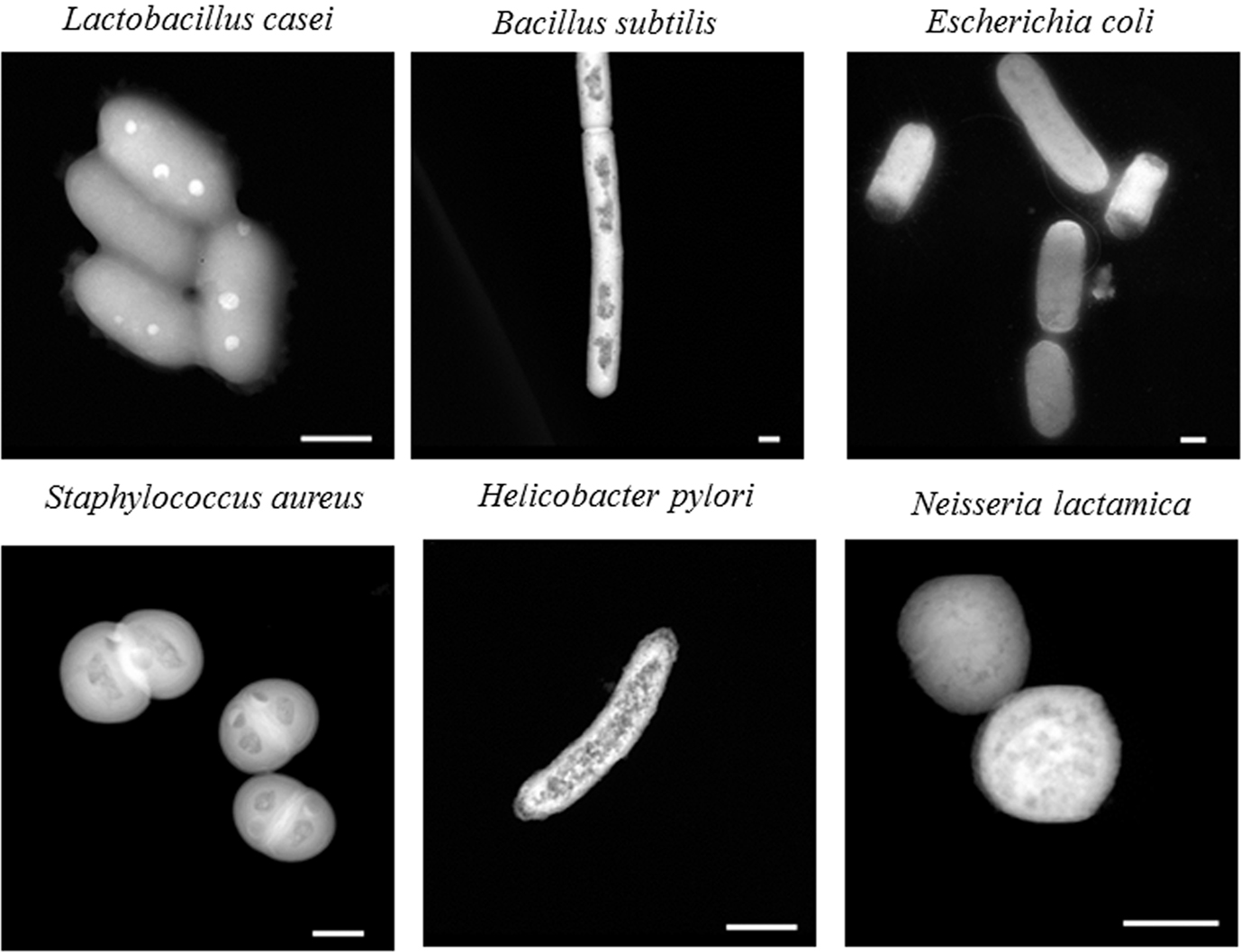
Figure 3. STEM imaging of a variety of bacterial strains. Bacteria were fixed; whole-cell mounted, and imaged using an FEI Titan 80-200 STEM. Selected images are representative samples from one biological replicate, with multiple bacteria imaged within each replicate. Scale bars (lower right) represent 500 nm.
Discussion
The contribution of metals to the pathogenesis of M. tuberculosis could help elucidate mechanisms behind persistent tuberculosis, a problem that impacts one-third of the world population (Dye et al., 2005). Our previous efforts geared toward the deciphering the impact of biometals on the survival of M. tuberculosis identified a novel class of metalloproteins represented by CsoR as a key regulator for metal ion transport, specifically, the divalent cation, Cu2+ (Liu et al., 2007; Ward et al., 2008). Moreover, one gene regulated by CsoR, the Cu transporter ctpV gene, was also shown to impact the survival of animals following aerosol infection with M. tuberculosis (Ward et al., 2008), another indication of the importance of biometals to the pathogenesis of tuberculosis. Most of the available techniques to examine intracellular levels of metals depend on either large number of cells (bulk samples) or the use of sophisticated, time consuming equipment (e.g., X-ray microscopy). Fortunately, we were able to adopt an approach based on STEM to profile the elemental map of Mycobacteria exposed to several stress conditions (e.g., nutrient starvation, high levels of Cu). Analytical STEM of M. tuberculosis (and other Mycobacteria) indicated the presence of discrete granules of several elements, mainly polyphosphates in addition to low levels of Mg and Ca. Other elements such as Cu and Fe were not detectable, most likely due to their dispersion throughout the cells. Previously, we were able to estimate intracellular levels of Cu, Fe, and other elements using bulk samples employing mass spectroscopy. It seems that the dispersion of these elements inside bacterial cells reduce the chances of detection using STEM approach. However, elements concentrated in polyphosphate granules could be easily analyzed using STEM. Fixation to stabilize bacterial structure and kill bacteria will result in ready diffusion of non-bound elemental species. Accordingly, internal and external concentrations of freely diffusing elements, would be expected to equilibrate. Washes and fixatives with low or no concentration of such elements would substantially deplete any concentration of non-bound elements within bacteria.
Polyphosphate has been associated with many diverse bacterial functions, including energy storage, motility, and biofilm formation (Kornberg, 1995; Rashid et al., 2000; Shi et al., 2011). Perhaps the most well-understood function of polyphosphate is its role in bacterial stringent response (Shiba et al., 1997), a phenomena that is particularly relevant to latent tuberculosis infections where M. tuberculosis can survive for decades in a persistent state within a host (Ahmad, 2011). It has been established in E. coli and confirmed in M. tuberculosis that accumulation of polyphosphate, which is triggered by exposure to specific stress conditions, including nutrient deprivation, osmotic stress, and oxidative stress, leads to activation of the stringent response (Ault-Rich et al., 1998; Sureka et al., 2007). In E. coli, more granules were observed in cells grown without inorganic phosphate or amino acids (Rao et al., 1998), an important indication for polyphosphate accumulation as part of stringent response to adverse growth conditions. In our hands, we observed large polyphosphate granules in M. tuberculosis exposed to conditions of nutrient starvation for up to three weeks. In general, STEM analysis applied here could further help elucidating the role of the relA regulon and other genes in polyphosphate formation.
In addition to its role in stringent response, polyphosphate accumulation has been previously associated with response to metal toxicity (Keasling, 1997; Wang et al., 2011). Interestingly, our observations show that the M. tuberculosis strain ΔctpV, which lacks a copper exporter required for copper detoxification (Ward et al., 2010), accumulates approximately twice as many polyphosphate granules (average of four/cell) as the wild-type strain (average of two/cell; unpublished observations). Subsequent studies will be directed toward confirming these preliminary results. Although the exact role that polyphosphate accumulation plays in pathogenesis is not yet understood, it has been shown that a knockout mutant of the enzyme responsible for polyphosphate synthesis in M. tuberculosis, Δppk1, had reduced survival in macrophages relative to wild-type bacteria (Sureka et al., 2007). Further, a strain lacking the enzyme required for polyphosphate degradation, MT0516:Tn, had reduced long-term survival in guinea pig lungs (Thayil et al., 2011). On the other hand, phosphate-limiting conditions have been shown to trigger a persistent state in M. tuberculosis, most probably through the 2-component regulatory system SenX3-RegX3 (Rifat et al., 2009). Despite what we know about the formation of polyphosphate granules and the regulatory genes involved in this process, we still do not have enough details on how Mycobacteria utilize polyphosphate during stress, or what role that plays in pathogenesis. Overall, the STEM analysis applied here could help elucidate the function of polyphosphate accumulation in M. tuberculosis pathogenesis and in vivo survival.
Although polyphosphate is present in all living cells, not all living cells form defined polyphosphate granules. For example, the model organism for polyphosphate utilization, E. coli, does not contain visible polyphosphate granules (Kornberg et al., 1999). In this report, polyphosphate granules were seen in both virulent and avirulent species of Mycobacteria as well as Lactobacilli, but defined granules were not detectable in the five other bacterial species tested. Notably, defined polyphosphate granules were not present in the H. pylori strain that was grown in a rich liquid media in our analysis. It has been shown previously that H. pylori accumulate polyphosphate granules differentially depending on growth conditions, including observed granule formation within in vivo-derived samples (Bode et al., 1993; Shirai et al., 2000), emphasizing the environment-dependent nature of polyphosphate granule formation.
We believe that our studies will eventually allow for the analysis of elemental profiles of M. tuberculosis in macrophages or in host tissue, possibly including the detection of dispersed elements with improvements to microscope sensitivity and/or data analysis. Recently, the presence of spore-like structures in Mycobacteria was suggested based on transmission electron microscopy (Ghosh et al., 2009; Lamont et al., 2012). However, this finding is still controversial and its evolutionary significance has been questioned (Traag et al., 2009). A STEM-based analysis of such structures could help in elucidating the observed phenotypes. Overall, we have shown that high resolution, field emission aberration corrected STEM analysis involving the use of extremely small beam diameter (<1 Å) has sufficient energy to penetrate whole-mounted specimens. This allows for high resolution EM images of bacterial cells. The small beam diameters also permit high spatial resolution EDS analysis of whole cells as well as elemental analysis of specific, down to the single nm, sized regions, or structures within cells. Such resolution will help studying the role of elemental distribution in the pathogenesis of M. tuberculosis as well as other intracellular pathogens.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Financial support was provided by NIH-R21AI081120. The authors thank Robin Kurtz and Katie Lyons Woodhams for providing bacterial cultures. The authors thank Sarah A. Marcus for reading the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at http://www.frontiersin.org/cellular_and_infection_microbiology/10.3389/fcimb.2012.00063/abstract
References
Agranoff, D., and Krishna, S. (2004). Metal ion transport and regulation in Mycobacterium tuberculosis. Front. Biosci. 9, 2996–3006.
Agranoff, D., Monahan, I. M., Mangan, J. A., Butcher, P. D., and Krishna, S. (1999). Mycobacterium tuberculosis expresses a novel pH-dependent divalent cation transporter belonging to the Nramp family. J. Exp. Med. 190, 717–724.
Agranoff, D. D., and Krishna, S. (1998). Metal ion homeostasis and intracellular parasitism. Mol. Microbiol. 28, 403–412.
Ahmad, S. (2011). Pathogenesis, immunology, and diagnosis of latent Mycobacterium tuberculosis infection. Clin. Dev. Immunol. 2011, 17.
Albrecht, R., Olorundare, O., liver, J., and Meyer, D. (2011). Nanoparticle labels for co-localization and correlative imaging at high spatial resolution. Microsc. Microanal. 17, 356–357.
Ault-Rich, D., Fraley, C. D., Tzeng, C. M., and Kornberg, A. (1998). Novel assay reveals multiple pathways regulating stress-induced accumulations of inorganic polyphosphate in Escherichia coli. J. Bacteriol. 180, 1841–1847.
Bode, G., Mauch, F., Ditschuneit, H., and Malfertheiner, P. (1993). Identification of structures containing polyphosphate in Helicobacter pylori. J. Gen. Microbiol. 139, 3029–3033.
Botella, H., Stadthagen, G., Lugo-Villarino, G., de Chastellier, C., and Neyrolles, O. (2012). Metallobiology of host-pathogen interactions: an intoxicating new insight. Trends Microbiol. 20, 106–112.
Corbett, E. L., Watt, C. J., Walker, N., Maher, D., Williams, B. G., Raviglione, M. C., and Dye, C. (2003). The growing burden of tuberculosis – global trends and interactions with the HIV epidemic. Arch. Intern. Med. 163, 1009–1021.
Cosgriffa, E. C., Oxleya, M. P., Allena, L. J., and Pennycookb, S. J. (2005). The spatial resolution of imaging using core-loss spectroscopy in the scanning transmission electron microscope. Ultramicroscopy 102, 317–326.
De Voss, J. J., Rutter, K., Schroeder, B. G., Su, H., Zhu, Y., and Barry, C. E. 3. (2000). The salicylate-derived mycobactin siderophores of Mycobacterium tuberculosis are essential for growth in macrophages. Proc. Natl. Acad. Sci. U.S.A. 97, 1252–1257.
Dillard, J. P., and Hackett, K. T. (2005). Mutations affecting peptidoglycan acetylation in Neisseria gonorrhoeae and Neisseria meningitidis. Infect. Immun. 73, 5697–5705.
Domenech, P., Pym, A. S., Cellier, M., Barry, C. E., and Cole, S. T. (2002). Inactivation of the Mycobacterium tuberculosis Nramp orthologue (mntH) does not affect virulence in a mouse model of tuberculosis. FEMS Microbiol. Lett. 207, 81–86.
Dye, C., Watt, C. J., Bleed, D. M., Hosseini, S. M., and Raviglione, M. C. (2005). Evolution of tuberculosis control and prospects for reducing tuberculosis incidence, prevalence, and deaths globally. JAMA 293, 2767–2775.
Ghosh, J., Larsson, P., Singh, B., Pettersson, B. M. F., Islam, N. M., Sarkar, S. N., Dasgupta, S., and Kirsebom, L. A. (2009). Sporulation in mycobacteria. Proc. Natl. Acad. Sci. U.S.A. 106, 10781–10786.
Gold, B., Deng, H., Bryk, R., Vargas, D., Eliezer, D., Roberts, J., Jiang, X., and Nathan, C. (2008). Identification of a copper-binding metallothionein in pathogenic mycobacteria. Nat. Chem. Biol. 4, 609–616.
Gonzalez, H., and Jensen, T. E. (1998). Nickel sequestering by polyphosphate bodies in Staphylococcus aureus. Microbios 93, 179–185.
Keasling, J. D. (1997). Regulation of intracellular toxic metals and other cations by hydrolysis of polyphosphate. Ann. N.Y. Acad. Sci. 829, 242–249.
Kornberg, A. (1995). Inorganic polyphosphate: toward making a forgotten polymer unforgettable. J. Bacteriol. 177, 491–496.
Kornberg, A., Rao, N. N., and Ault-Rich, D. (1999). Inorganic polyphosphates: a molecule of many functions. Annu. Rev. Biochem. 68, 89–125.
Lamont, E. A., Bannantine, J. P., Armin, A., Ariyakumar, D. S., and Sreevatsan, S. (2012). Identification and characterization of a spore-like morphotype in chronically starved Mycobacterium avium subsp. paratuberculosis cultures. PLoS ONE 7, e30648. doi:10.1371/journal.pone.0030648
Liu, T., Ramesh, A., Ma, Z., Ward, S. K., Zhang, L., George, G. N., Talaat, A. M., Sacchettini, J. C., and Giedroc, D. P. (2007). CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator. Nat. Chem. Biol. 3, 60–68.
Meyer, D., Heintz, J., and Albrecht, R. M. (2010). Detection of nanoparticles of differing composition for high resolution labeling. Microsc. Microanal. 16, 992–993.
Parish, T. (2003). Starvation survival response of Mycobacterium tuberculosis. J. Bacteriol. 185, 6702–6706.
Rao, N. N., Liu, S., and Kornberg, A. (1998). Inorganic polyphosphate in Escherichia coli: the phosphate regulon and the stringent response. J. Bacteriol. 180, 2186–2193.
Rashid, M. H., Rao, N. N., and Kornberg, A. (2000). Inorganic polyphosphate is required for motility of bacterial pathogens. J. Bacteriol. 182, 225–227.
Rifat, D., Bishai, W. R., and Karakousis, P. C. (2009). Phosphate depletion: a novel trigger for Mycobacterium tuberculosis persistence. J. Infect. Dis. 200, 1126–1135.
Rodriguez, G. M., and Smith, I. (2006). Identification of an ABC transporter required for iron acquisition and virulence in Mycobacterium tuberculosis. J. Bacteriol. 188, 424–430.
Schaible, M. E., and Kaufmann, S. H. E. (2004). Iron and microbial infection. Nat. Rev. Microbiol. 2, 946–953.
Shi, T., Fu, T., and Xie, J. (2011). Polyphosphate deficiency affects the sliding motility and biofilm formation of Mycobacterium smegmatis. Curr. Microbiol. 63, 470–476.
Shiba, T., Tsutsumi, K., Yano, H., Ihara, Y., Kameda, A., Tanaka, K., Takahashi, H., Munekata, M., Rao, N. N., and Kornberg, A. (1997). Inorganic polyphosphate and the induction of rpoS expression. Proc. Natl. Acad. Sci. U.S.A. 94, 11210–11215.
Shirai, M., Kakada, J., Shibata, K., Morshed, M. G., Matsushita, T., and Nakazawa, T. (2000). Accumulation of polyphosphate granules in Helicobacter pylori cells under anaerobic conditions. J. Med. Microbiol. 49, 513–519.
Stewart, M., Somlyo, A. P., Somlyo, A. V., Shuman, H., Lindsay, J. A., and Murrell, W. G. (1980). Distribution of calcium and other elements in cryosectioned Bacillus cereus T spores, determined by high-resolution scanning electron probe x-ray microanalysis. J. Bacteriol. 143, 481–491.
Sureka, K., Dey, S., Datta, P., Singh, A. K., Dasgupta, A., Rodrigue, S., Basu, J., and Kundu, M. (2007). Polyphosphate kinase is involved in stress-induced mprAB-sigE-rel signalling in mycobacteria. Mol. Microbiol. 65, 261–276.
Thayil, S. M., Morrison, N., Schechter, N., Rubin, H., and Karakousis, P. C. (2011). The role of the novel exopolyphosphatase MT0516 in Mycobacterium tuberculosis drug tolerance and persistence. PLoS ONE 6, e28076. doi:10.1371/journal.pone.0028076
Traag, B. A., Driks, A., Stragier, P., Bitter, W., Broussard, G., Hatfull, G., Chu, F., Adams, K. N., Ramakrishnan, L., and Losick, R. (2009). Do mycobacteria produce endospores? Proc. Natl. Acad. Sci. U.S.A. 107, 878–881.
Vanhoe, H., Vandecasteele, C., Versieck, J., and Dams, R. (1989). Determination of iron, cobalt, copper, zinc, rubidium, molybdenum, and cesium in human serum by inductively coupled plasma mass spectrometry. Anal. Chem. 61, 1851–1857.
Versieck, J. (1994). Neutron activation analysis for the determination of trace elements in biological materials. Biol. Trace Elem. Res. 43–45, 407–413.
Wagner, D., Maser, J., Lai, B., Cai, Z. H., Barry, C. E., Bentrup, K. H. Z., Russell, D. G., and Bermudez, L. E. (2005). Elemental analysis of Mycobacterium avium-, Mycobacterium tuberculosis-, and Mycobacterium smegmatis-containing phagosomes indicates pathogen-induced microenvironments within the host cell’s endosomal system. J. Immunol. 174, 1491–1500.
Wang, Y., Ren, Z., Jiang, F., Geng, J., He, W., and Yang, J. (2011). Effect of copper ion on the anaerobic and aerobic metabolism of phosphorus-accumulating organisms linked to intracellular storage compounds. J. Hazard. Mater. 186, 313–319.
Ward, S. K., Abomoelak, B., Hoye, E. A., Steinberg, H., and Talaat, A. M. (2010). CtpV: a putative copper exporter required for full virulence of Mycobacterium tuberculosis. Mol. Microbiol. 77, 1096–1110.
Ward, S. K., Hoye, E. A., and Talaat, A. M. (2008). The global responses of Mycobacterium tuberculosis to physiological levels of copper. J. Bacteriol. 190, 2939–2946.
Wesenberg, D., Bleuel, C., and Krauss, G.-J. (2007). “A glossary of microanalytical tools to assess the metallome,” in Molecular Microbiology of Heavy Metals, eds D. Nies and S. Silver (Berlin: Springer), 159–186.
Winder, F. G., and Denneny, J. M. (1957). The metabolism of inorganic polyphosphate in mycobacteria. J. Gen. Microbiol. 17, 573–585.
Wolschendorf, F., Ackart, D., Shrestha, T. B., Hascall-Dove, L., Nolan, S., Lamichhane, G., Wang, Y., Bossmann, S. H., Basaraba, R. J., and Niederweis, M. (2011). Copper resistance is essential for virulence of Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. U.S.A. 108, 1621–1626.
Keywords: tuberculosis, scanning transmission electron microscopy, metals, elements
Citation: Ward SK, Heintz JA, Albrecht RM and Talaat AM (2012) Single-cell elemental analysis of bacteria: quantitative analysis of polyphosphates in Mycobacterium tuberculosis. Front. Cell. Inf. Microbio. 2:63. doi: 10.3389/fcimb.2012.00063
Received: 24 February 2012; Paper pending published: 26 March 2012;
Accepted: 20 April 2012; Published online: 24 May 2012.
Edited by:
Rey Carabeo, Imperial College London, UKReviewed by:
Rey Carabeo, Imperial College London, UKPetros Karakousis, Johns Hopkins School of Medicine, USA
Copyright: © 2012 Ward, Heintz, Albrecht and Talaat. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Adel M. Talaat, Laboratory of Bacterial Genomics, Department of Pathobiological Sciences, University of Wisconsin–Madison, 1656 Linden Drive, Madison, WI 53706-1581, USA. e-mail:YXRhbGFhdEB3aXNjLmVkdQ==

 Joseph A. Heintz2
Joseph A. Heintz2