- 1Key Laboratory of Cancer Proteomics of Chinese Ministry of Health, Xiangya Hospital, Central South University, Changsha, China
- 2Hunan Engineering Laboratory for Structural Biology and Drug Design, Xiangya Hospital, Central South University, Changsha, China
- 3State Local Joint Engineering Laboratory for Anticancer Drugs, Xiangya Hospital, Central South University, Changsha, China
- 4The State Key Laboratory of Medical Genetics, Central South University, Changsha, China
Pituitary adenoma (PA) is a common intracranial neoplasm that impacts on human health through interfering hypothalamus–pituitary–target organ axis systems. The development of proteomics gives great promises in the clarification of molecular mechanisms of a PA and discovery of effective biomarkers for prediction, prevention, early-stage diagnosis, and treatment for a PA. A great progress in the field of PA proteomics has been made in the past 10 years, including (i) the use of laser-capture microdissection, (ii) proteomics analyses of functional PAs (such as prolactinoma), invasive and non-invasive non-functional pituitary adenomas (NFPAs), protein post-translational modifications such as phosphorylation and tyrosine nitration, NFPA heterogeneity, and hormone isoforms, (iii) the use of protein antibody array, (iv) serum proteomics and peptidomics, (v) the integration of proteomics and other omics data, and (vi) the proposal of multi-parameter systematic strategy for a PA. This review will summarize these progresses of proteomics in PAs, point out the existing drawbacks, propose the future research directions, and address the clinical relevance of PA proteomics data, in order to achieve our long-term goal that is use of proteomics to clarify molecular mechanisms, construct molecular networks, and discover effective biomarkers.
Introduction
The pituitary gland, which is located in the bottom-middle of cranial cavity, functions in many biological processes. Pituitary is the most important endocrine gland of body and is composed of several kinds of cells, including corticotroph, thyrotroph, somatotroph, lactotroph, and gonadotroph, which is responsible for the secretion of corresponding hormone to regulate the endocrine system (1, 2). The endocrine functions are affected if any of the pituitary cell types goes wrong. Pituitary adenoma (PA) is the common neoplasm in pituitary gland and accounts for ca. 10% in intracranial tumors (3–5). Although most of the PAs are benign, its invasive features do exist and always comes with poor prognosis (6). The different types of PAs have different origin of cells and involve different endogenous and exogenous factors, which cause PA highly heterogeneous (3). PAs are clinically classified into functional and non-functional pituitary adenomas (FPAs and NFPAs). FPAs, commonly refer to monoclonal, originate from adrenocorticotropic hormone (ACTH) cell adenoma, thyrotropin-stimulating hormone (TSH) cell adenoma, growth hormone (GH) cell adenoma, and prolactin (PRL) cell adenoma, whereas the NFPAs commonly originate from gonadotroph cells (2, 7). NFPAs are subclassified into intact luteinizing hormone (LH+), follicle-stimulating hormone (FSH+), LH/FSH+, null cell, oncocytoma, silent corticotroph, and silent somatotroph subtypes (2). These differences in pathophysiological characteristics are the basis of proteomic heterogeneity in PAs. A series of proteomics strategies have been extensively used to study PAs, and the status and perspectives regarding PA proteomics were reviewed in detail by Zhan and Desiderio in the year 2004 (1). After that, significant progresses have been made in PA proteomics in the past 10 years, including sample preparation using laser-capture microdissection (LCM) (8, 9), proteomics analyses of FPA (such as prolactinoma) (10), invasive versus non-invasive PAs (6, 11), NFPA subtypes (12), serum (13, 14), protein variants/isoforms (3, 10, 15, 16), post-translational modifications (PTMs) (17–21), and protein molecular networks (22). This article reviews these progresses in PA proteomics in the past 10 years, the existing drawbacks, future perspectives, and the clinical relevance of PA proteomics data.
Progresses in PA Proteomics
Sample Preparation Using Laser-Capture Microdissection Coupled with Two-dimensional Liquid Chromatography
Two obvious drawbacks exist in two-dimensional gel electrophoresis (2DGE)-based quantitative comparison of proteomes between PAs and controls (1, 23). First, a PA tissue is commonly obtained from neurosurgery operation, and pituitary control tissue is commonly obtained from postmortem tissues died from other diseases or accidents. The origin of cells in PA is monoclonal, which is originated from corticotroph, somatotroph, thyrotroph, lactotroph, and gonadotroph in the pituitary gland (1). Thus, a PA tissue sample contains a relatively pure cell type, whereas postmortem pituitary control tissue sample contains multiple cell types, including corticotroph, somatotroph, thyrotroph, lactotroph, gonadotroph, and other types of cells. These differences in cell types affect the accurate comparison between PA and control tissue proteomes. Second, 2DGE with pH 3–10 IPG strip and 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) gel in separation of proteome is limited within a range of proteins with pH 4–8 and mass 15–150 kDa. These proteins with pI < 4 or pI > 8 and Mr > 150 kDa or Mr < 10 kDa are usually lost, which seriously impacts on the comparison between PA and control proteomes. Moreover, 2DGE has difficulty in the detection of low-abundance proteins in a proteome, which are usually associated with important physiological and pathological processes.
In order to overcome these drawbacks, LCM (24, 25) and multi-dimensional liquid chromatography (MDLC) (26) were used to analyze PA proteomes (8, 9, 27, 28). LCM can isolate, purify, and enrich tumor cells from PA tissues and the corresponding pituitary cells from control pituitaries, such as corticotroph, somatotroph, thyrotroph, lactotroph, and gonadotroph. This approach resolves the inconsistency in the origin of cells between PA and control pituitaries and achieves the accurate comparison between PA and control proteomes. Compared to 2DGE, MDLC has significant advantages in maximizing the coverage of a proteome because it is not to be limited in the detection and identification of low-abundance proteins, extremely acidic/basic proteins, extremely low/high-mass proteins, and hydrophobic proteins (26).
An FPA prolactinoma was analyzed using LCM and MDLC (8, 9, 27, 28). Prolactinoma tissue contains multiple cells, including lactotroph tumor cells, endothelial cells, fibroblasts, and other stromal cells (8). The lactotroph tumor cells were isolated and purified using immunohistochemistry-guided LCM (27, 28), and a total of 2243 unique proteins from LCM-enriched lactotroph tumor cells were identified using SDS-PAGE and 2D-LC-tandem mass spectrometry (MS/MS) (8), which form currently the biggest protein database for prolactinoma proteome. Moreover, immuno-LCM was used to enrich lactotroph cells from postmortem human pituitary tissues (9). A total of 1660 proteins from LCM-enriched lactotroph cells were identified using 2D-LC–MS/MS, which represent currently the biggest proteomic database of normal lactotroph cells. However, currently, no quantitative comparison study was carried out between LCM-enriched lactotroph tumor and normal cells to discover accurate differentially expressed proteins (DEPs) for the clarification of prolactinoma tumorigenesis.
Significant Extension of Proteomic Reference Mapping Data in PAs and Controls
Large-scale proteomic reference mapping data are essential to discover PA-related proteins for the understanding of molecular mechanisms and discovery of reliable biomarkers of a PA. (i) For a human pituitary proteome, a total of 1449 proteins were identified using multiple gel-based technology (MGT), including in-gel isoelectric focusing (IEF)–LC–MS/MS and SDS-PAGE–LC–MS/MS (29). Most of these proteins distribute within the range of pI 3.8–12.0 (75%) and Mr 20–100 kDa (70%), and lots of low-abundance proteins were identified, including 55 protein kinases (PKAs), 45 receptors, 29 transcription factors, and 43 Ras-family proteins such as 11 G proteins. Currently, these data form the biggest protein reference map of a human pituitary proteome. (ii) For an NFPA proteome, 2DGE-based proteomics technique was used to identify 111 unique proteins in an NFPA tissue (30) and 107 unique proteins in an FSH+-NFPA (31), which forms the protein reference map to serve for 2DGE-based comparative proteomics analysis of NFPAs. Furthermore, the proteomic reference map of FSH+-NFPA contains 107 unique proteins that were involved in multiple functional systems, including receptor signal pathways, transcription and translation regulating system, bioenzyme system, oxidative stress, glutathione metabolism, gap junction, and inflammation signaling pathway, which provide a reference to in-depth analysis of the heterogeneity of NFPA proteomes (31). (iii) For a prolactinoma proteome, immno-LCM in combination with 2D-LC–MS/MS was used to identify 2243 proteins from LCM-purified lactotroph tumor cells (8) and 1660 proteins from LCM-purified normal lactotroph cells (9); these data form the biggest proteomic database for purified lactotroph tumor cells and normal cells and benefit to analyze prolactinoma tumorigenesis. Moreover, 305 proteins were identified in LCM-purified normal lactotroph cells (9) and whole pituitary control tissues (29). These extended proteomic reference data will benefit analysis of different subtypes of FPA and NFPA proteomes to discover reliable biomarkers for the understanding of molecular mechanisms, early-stage diagnosis, and therapeutic targets.
Proteomics Analysis of Invasive Characteristics of NFPAs
Most of the PAs are benign. However, microscopic evidence shows that approximately 40% and even up to 80% of PAs have local invasion to surrounding structures (32–35), which causes difficulty in complete removal of a tumor through neurosurgery, increases the risks of complications and poor therapeutic outcome, and results in a need of adjuvant radiotherapy or medication after neurosurgery. Therefore, the invasive characteristics of PAs are a challenging clinical problem. Clarification of molecular mechanisms and discovery of protein biomarkers regarding invasiveness of PAs are necessary to better manage an invasive PA patient (36). Quantitative transcriptomics study revealed 346 differentially expressed genes (DEGs) (233 upregulated and 113 downregulated) in invasive relative to non-invasive NFPAs (36). A 2DGE-based comparative proteomics discovered 30 DEPs between invasive and non-invasive PAs (11). Furthermore, 2DGE proteomic comparison between 4 invasive and 4 non-invasive NFPAs (6) identified 57 DEPs (30 upregulated and 27 downregulated) from 103 differential 2D gel spots (64 upregulated and 39 downregulated), and these DEPs were involved in several invasiveness-related pathway networks, including oxidative stress, mitochondrial dysfunction, MAPK-signaling abnormality, ketogenesis and ketolysis, TR/RXR activation, proteolysis abnormality, CDK5 signaling abnormality, and amyloid processing. These findings provide important clues to insight into the molecular mechanisms of invasiveness of an NFPA and discovery of reliable protein biomarkers to guide adjuvant treatment for post-neurosurgery invasive NFPAs.
Proteomics Analysis of Heterogeneity of NFPAs
Non-functional pituitary adenoma is a highly heterogeneous neoplasm with different origin of cells and multiple hormone-expressed subtypes, including intact gonadotroph with LH/FSH+ (40–79%), silent somatotroph with GH expression (3%), silent corticotroph with ACTH expression (8%), null cell without hormone expression (17%), and oncocytoma without hormone expression (6%) (3, 37, 38). The intact gonadotroph may lead to several hormone-expressed subtypes of NFPAs, including LH+, FSH+, or LH/FSH+. The interesting thing is that silent hormone expression associates the invasiveness of an NFPA. For example, ACTH+-NFPA was more invasive and had higher recurrence rate than ACTH−-NFPA (39). The imbalance of ERα and ERβ expressions is involved in the pathological process and tumor biological behavior, such as invasiveness of an NFPA (40), and the ERα/ERβ ratio was positively related to invasive degree of an NFPA. Thereby, it is necessary to investigate proteomic heterogeneity among NFPA subtypes to discover the common and specific biomarkers for the accurate understanding of molecular mechanisms, early-stage diagnosis, and prognostic assessment and the therapeutic targets toward personalized and precise medicine. 2DGE-based comparative proteomics was used to identify 76 DEPs in NF−, LH+, FSH+, and LH/FSH+ (NF− means NFPA that had negative immunohistochemical stains for LH, FSH, ACTH, GH, PRL, and TSH) relative to pituitary controls (12). Of them, 44 DEPs (10 upregulated and 34 downregulated) were common to these 4 NFPA subtypes. The rest of DEPs were common or specific to some of the four NFPA subtypes. Four pathway–network systems, including mitochondrial dysfunction, oxidative stress, MAPK-signaling abnormality, and cell-cycle dysregulation, were common to four NFPA subtypes. However, many differences existed in each pathway–network system among four NFPA subtypes, including different expression levels of each protein node, different protein profiles, and different pathway–network profiles (12). These data provided a heterogeneous profile of human NFPA proteomes that would benefit in-depth clarification of NFPA heterogeneity, molecular mechanisms of NFPAs, and discovery of effective biomarkers for early-stage diagnosis, therapy, and prognosis for NFPA personalized and precise medicine practice.
Proteomics Analysis of Protein Variants/Isoforms and PTMs of PAs
The process of a gene-coded protein involves multiple post-transcriptional/post-translational regulations, including splicing, modifications, translocation, and spatial conformation (41–43). Protein variants/isoforms are mainly derived from splicing and PTMs. Protein variants/isoforms and PTMs are very important aspects in a proteome because they function in multiple physiological and pathological processes and would be biomarkers and therapeutic targets (44). Currently, exploration of protein variants/isoforms and PTMs is much insufficient in the width and depth in a PA proteome. By far, GH (or PRL) variants/isoforms in a human pituitary proteome have been documented (3, 10, 15, 16). PTMs regarding phosphorylation and nitration in PA and control proteomes have been reported (17–21).
Growth hormone is a very important hormone in the body. Abnormal secretions of GH are associated with many GH-related diseases, such as dwarfism, gigantism, acromegalia, and adenoma. The synthesis and release of human GH (hGH) are regulated under a series of factors (15). The hGH/IGF1 axis plays an important role in stimulating body growth (45), whereas some PTMs, such as phosphorylation, involve signal transduction pathway of hGH/IGF1 axis (46). Four hGH isoforms have been found in some GH-producing adenomas. Study (47) demonstrates that extensively molecular heterogeneity existed in hGH because of alternative splicing, different PTMs, and proteolytic processing and that circulating hGH composites of various proportions among three non-22-kDa GH isoforms 2–4 in various GH-related diseases. 2DGE-based proteomics found that 24 2D gel spots contained hGH, with significantly different expression proportions of these isoforms, namely, isoform 1 (87.5%) > isoform 2 (8.1%) > isoform 3 (3.3%) > isoform 4 (1.1%) (15). Moreover, three phosphorylation sites (Ser-77, Ser-176, and Ser-132) and a deamidation at Asn-178 were identified in the hGH amino acid sequence. Also, many studies demonstrate the ratio of non-22 kDa GH isoform might be a marker for normal and abnormal growth (48), and the evaluation of these GH isoforms could be a clinical monitoring for acromegalic patients (49). A 2DGE-based comparative proteomics (21) found at least 17 GH isoforms and 6 PRL isoforms in human pituitary proteomes, and not all these GH and PRL isoforms were associated with PAs.
Phosphorylation in a protein is an important PTM, which is involved in the regulation of multiple critical biological functions, such as protein–molecule (DNA, RNA, and protein) interactions, enzyme activity, protein trafficking, protein intracellular localization, and protein degradation, and is extensively associated with different diseases, such as cancer, inflammation, and neurodegenerative diseases (17). Phosphorylation occurs in Ser, Thr, and Tyr residues in a protein and is dynamically and reversibly regulated with PKAs and phosphatases that are the common drug targets, and much attention has been paid to protein phosphorylation event (50). Protein phosphorylation analysis was carried out in pituitary control tissues, which identified a total of 28 phosphoproteins from 2 different studies (17, 18), including 8 phosphorylation sites in 6 pituitary phosphoproteins that were characterized using immobilized metal ion affinity chromatography (IMAC)-based LC–MS/MS from unseparated digested pituitary protein mixture (18) and 50 phosphorylation sites in 26 pituitary phosphoproteins (73 phosphopeptides) that were characterized using a modified strategy-in-gel IEF–LC–MS/MS from digested pituitary protein mixture (17). These data expanded human pituitary phosphoprotein database. However, quantitative phosphoproteomics has not been performed between PAs and controls to discover PA-related phosphoproteins and phosphosites for in-depth understanding of biological significance of phosphorylation in PAs.
Tyrosine nitration in a protein is another important PTM that is mainly derived from in vivo peroxynitrite (ONOO−) pathway, myeloperoxidase pathway, and metalloperoxidase reaction pathway (19, 51, 52). Nitration decreases the electron density of a phenolic ring of tyrosine residue to affect the affinity ability between protein–protein interactions, such as enzyme–substrate, receptor–ligand, and antigen–antibody, and is extensively associated with different pathophysiological processes, such as cancer, neurodegenerative disease, and inflammation (19). Nitroproteomics was performed in pituitary controls (20, 53) and adenomas (21). Eight nitroproteins and eight nitrotyrosine sites were characterized from a pituitary control tissue using 2DGE-based nitrotyrosine Western blot coupled with MS/MS (20, 53), including cGMP-dependent PKA 2, stanniocalcin 1, synaptosomal-associated protein, actin, immunoglobulin α Fc receptor, mitochondrial co-chaperone protein HscB, progestin and adipoQ receptor family member III, and proteasome subunit α type 2. Nine nitroproteins and 10 nitrotyrosine sites were characterized from a PA tissue using nitrotyrosine affinity column (NATC)-based MALDI–MS/MS (21), including sphingosine-1-phosphate lyase 1, zinc finger protein 432, cAMP-dependent PKA type I-β regulatory subunit, Rho-GTPase-activating protein 5, leukocyte immunoglobulin-like receptor subfamily A member 4, centaurin-β 1, proteasome subunit α type 2 interleukin 1 family member 6, and rhophilin 2. Moreover, three nitroprotein–protein complexes were identified, including nitrated proteasome–ubiquitin complex, nitrated β-subunit of cAMP-dependent PKA complex, and nitrated interleukin 1 family member 6-interleukin 1 receptor–interleukin 1 receptor-associated kinase-like 2 (IL1F6–IL1R–IRAK2) (21). Moreover, protein domain/motif analysis found that most nitrotyrosine sites occurred within important protein domains and motifs (21), and pathway–network analysis found that these nitroproteins function in important signaling pathway-network systems (22), which demonstrated that tyrosine nitration might play important roles in the formation of a human PA. However, quantitative nitroproteomics has not been performed between PAs and controls to discover PA-related nitroproteins and nitrotyrosine sites for in-depth understanding of biological significance of tyrosine nitration in PAs.
Serum Proteomics and Peptidomics in NFPAs
No biomarker is used to early diagnose an NFPA, and NFPA is usually diagnosed exclusively using CT or MRI after appearance of compression symptoms or sometimes by chance (13). Serum is an important window to predict, early diagnose, and prognosis assess a PA. One study (13) analyzed 34 sera from NFPA patients and 34 sera from age-/sex-matched healthy controls using surface-enhanced laser desorption/ionization time-of-flight mass spectrometry (SELDI-TOF-MS) and identified 9 serum DEPs (7 upregulated and 2 downregulated) with both 82.4% sensitivity and specificity in discrimination of NFPAs from controls. Another study (14) quantitatively compared 65 sera from PAs and 90 sera from healthy controls using proteomic fingerprint technology that was magnetic beads-based MALDI-TOF-MS and identified 42 m/z peaks that were significantly related to PAs (14). A three-biomarker diagnostic model was established with 90.0% sensitivity and 88.3% specificity in discrimination of human PAs from controls, which provided a powerful and reliable method to diagnose a PA (14).
Protein Antibody Array Analysis of PAs
Protein antibody array is an effective method to detect alterations of known proteins in PAs and easily translated to biomarkers for prediction, diagnosis, and prognostic assessment of PAs. A protein antibody array with 1005 monoclonal antibodies (54) identified 316 proteins in GH-, ACTH-, and PRL-secreting FPAs, and in NFPAs. Also, a set of DEPs were identified and confirmed using immunohistochemistry and Western blotting.
Proteomic Data-Based Pathway Networks in NFPAs
Each protein functions in the corresponding pathway–network system to exert its biological roles. The IPA pathway analysis is a widely used pathway analysis system to identify statistically significant signaling pathway networks from a set of qualitative and quantitative proteomic data (22), including 111 proteins from a human NFPA tissue, 56 DEPs from human NFPA tissues and from prolactinoma tissues, 9 nitroproteins and 3 nitroprotein–protein complexes from a human NFPA tissue, and 8 nitroproteins from a pituitary control tissue. Comprehensive analysis of these pathway networks revealed four signaling pathway–network systems in NFPAs, including mitochondrial dysfunction, oxidative stress, cell-cycle dysregulation, and MAPK-signaling abnormality (22). These data provided potential biomarkers and contributed to the development of new efficacious treatment targets for PAs.
Clinical Relevance of PA Proteomics Data
A great methodological progress of PA proteomics and use of proteomics approach to study pathophysiological characteristics of PAs have been reviewed, as described above. Here, the clinical relevance of PA proteomics data (3, 6, 10–12, 55) is further addressed in the following aspects.
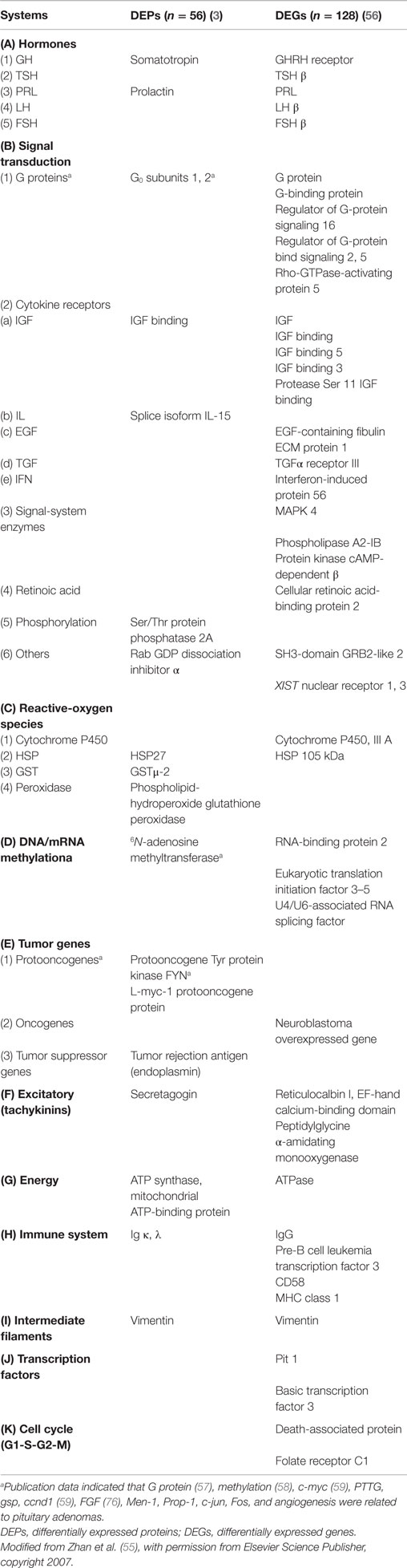
(i) Insights into molecular mechanisms of PAs. Proteomics offers a unique insight into molecular mechanisms of pituitary tumorigenesis at the level of proteome. Our multiple comparative proteomics studies have confirmed our initial hypothesis that the proteome differs between PAs and controls (3, 10), between invasive and non-invasive PAs (6, 11), and among different types of PAs (12); many DEPs and molecular networks were related to transcriptomics data (3, 10, 36, 56) and to the corresponding biological systems (6, 12, 22). These DEPs and molecular networks provided a basis to determine alterations in multiple protein systems that contribute to pituitary tumorigenesis. A set of DEP data (3) and DEG data (56) were correlated and obtained many protein systems that connect some critical aspects of PA formation (Table 1). First, the alterations in endocrine hormone levels are the significantly clinical characteristics of PAs. Many pituitary hormones were found to change, including GH, TSH, PRL, LH, and FSH. The synthesis and secretion of each hormone are the important function of pituitary organ and are closely regulated by various signaling pathways and factors. The cGMP-dependent PKA and nitric oxide are taken, for example, that significantly regulates water-soluble hormones. Proteomics study (20) discovered that the cGMP-dependent PKA 2 was nitrated in human pituitary tissues, which indicates that the nitration of cGMP-dependent PKA might be involved in the signal processing of water-soluble hormones. Second, the alterations in signal transduction systems significantly participate in the pathogenesis of PAs. Many signal pathways were found by omics data to change, including G proteins, cytokine receptors (IFG, IL, EGF, TGF, and IFN), and some signal system-regulated enzymes (MAPK4, phospholipase A2-IB, and cAMP-dependent PKA). G proteins have been confirmed to play an important role in PAs (57), and FGF family members are involved in pituitary pathogenesis (58). FGF receptor genes mediate FGF signal pathways, and these genes encode receptor tyrosine kinases. Our proteomic DEP data clearly have a Tyr PKA. In addition, phosphorylation is significantly involved in signal pathways, and a Ser/Thr protein phosphatase 2A was found in our proteomics data (3). Third, oxidative/nitrative stresses are significantly associated with multiple cancer pathogenesis. Proteomics DEP data (3) demonstrate that some reactive-oxygen species (ROS)-related proteins are changed in PAs, including cytochrome P450, heat shock protein, GST, and phospholipid-hydroperoxide glutathione peroxidase. Fourth, DNA methylation was found to participate in pituitary tumorigenesis (58); proteomics study (3) revealed a methylation-related DEP (Table 1), and three methylation-related DEGs were found by transcriptomics study (56). Fifth, oncogene activation in PAs (such as c-Myc) was found (59); protooncogene-related DEPs (3) and -related DEGs were found (56) (Table 1). Furthermore, multiple proteomics data-based molecular network studies revealed four important pathway–network systems that were altered in NFPAs, including mitochondrial dysfunction, oxidative stress, cell-cycle dysregulation, and MAPK-signaling system abnormality (22); and the last three pathway–network systems were involved in protein tyrosine nitration, for example, NRF2-mediated oxidative stress response, cell-cycle G2/M DNA damage checkpoint regulation, and p38 MAPK signaling were identified from PA nitroproteomic data (22). It shows that tyrosine nitration is involved in pituitary tumorigenesis. Finally, invasiveness of PAs is a challenging clinical problem. Fifty-seven DEPs (30 upregulated and 27 downregulated) were found by proteomic study to associate the invasive features of PAs (6), and invasiveness-related pathway networks were found including MAPK-signaling abnormality, mitochondrial dysfunction, oxidative stress, proteolysis abnormality, ketogenesis and ketolysis, TR/RXR activation, CDK5 signaling abnormality, and amyloid processing (6). These data demonstrate an overview of multiple protein systems that are involved in the pathogenesis of PAs.
(ii) Discovery of potential biomarkers and therapeutic targets for PAs. DEPs and DEGs (Table 1) (3, 56) and the consistently altered DEPs and DEGs in the levels of protein and mRNA (Table 2) (3, 10), which were identified with proteomics and transcriptomics studies, are a part of our novel source of candidate biomarkers and therapeutic targets. Secretagogin was taken, for example (60). Secretagogin is a type of Langerhans-specific Ca2+-binding protein (61) and is highly expressed in the secretory neurons of the anterior pituitary (62). Secretagogin can inhibit the transcription of excitatory amino acid substance P (SP) to lower the growth rate of RIN-5F tumor cells (61), which is consistent with Desiderio’s hypothesis that an imbalance between inhibitory (opioid) and excitatory (tachykinin) neuropeptidergic systems might contribute to the pathogenesis of a pituitary macroadenoma (63–67). Proteomics and transcriptomics analyses (60) revealed a significant downregulation of secretagogin in a set of human NFPAs, and its protein expression was significantly related to its mRNA expression. These data demonstrate that secretagogin might be a biomarker for PAs. Moreover, other candidate biomarkers, such as G proteins, tissue transglutaminase, isocitrate dehydrogenase (NADP) cytoplasmic (IDH1), calreticulin, cytokine receptors, and ROS-related proteins (Tables 1 and 2), need to be further studied for potential biomarkers and therapeutic targets.
(iii) Pituitary hormone variants/isoforms and PTMs in human PAs. Protein variants/isoforms result from splicing, PTMs, etc. The alterations in neuroendocrine hormone levels are the important characteristics of PAs. Proteomics studies demonstrate that human pituitary hormones had multiple variants/isoforms. Not all of variants/isoforms were associated with PAs (3). First, GH variants/isoforms and PTMs. Somatotropin was significantly downregulated at the mRNA and protein levels in NFPAs (3) and in prolactinomas (10), which was consistent with their monoclonal composition in origin (68, 69) – a NFPA generated from gonadotrophs and a prolactinoma from lactotrophs, whereas GH receptor gene was unchanged. These data showed that GH hyposecretion in NFPAs results from the hypoexpression of the GH gene. However, the interesting finding is that multiple GH variants/isoforms (17 spots contain somatotropin) were detected (3, 10); these data cannot be interpreted by transcriptomics method. The downregulated ratio of different GH isoforms was different in each cell type of PA relative to controls. These data suggested that the proportion of different GH isoforms changed in each cell type adenoma compared to controls. Other researchers showed that the proportion of the circulating GH isoforms significantly changed in PAs and other pituitary diseases (48, 49). The proportional change in the different GH isoforms might have an important value in the clinical evaluation of human PAs. Recently, we found that GH isoforms were derived from various splicing variants and PTMs, including phosphorylation. The phosphorylation of endogenous GH in the human pituitary (18) provided us with new insights into the mechanisms of GH that participates in the neuroendocrine signal pathways. Second, PRL variants/isoforms and PTMs. PRL is another important pituitary hormone. Proteomics detected six PRL variants/isoforms (3, 10). Similar to GH, each PRL variant/isoform was downregulated in each cell type of NFPA, with a different downregulated ratio relative to controls. There was no significant expression change in the PRL gene at the mRNA and protein levels in the prolactinomas relative to controls, which is consistent with a prolactinoma’s monoclonal composition in origin. However, the proportion of six PRL variants/isoforms was changed in prolactinomas compared to controls. Other researchers showed that glycosylation was an important PRL modification (70) that produces different variants/isoforms; glycosylation of human PRL might downregulate its hormone bioactivity and promote its metabolic clearance (71). Other studies showed that the main variant of PRL was non-glycosylated form of PRL in human prolactinomas (72), which is consistent with our result that some PRL isoforms (10) were downregulated in prolactinomas relative to controls. Thus, we speculated that PRL variants/isoforms in these downregulated spots were possibly glycosylated PRL. Therefore, our detailed study of different variants/isoforms and PTMs of GH and PRL could lead to new insights into the clinical importance of pituitary hormones in a PA. The ratio of the change of each variant/isoform in human pituitary tissue has value for clinical research. However, one must realize that studies regarding protein variants/isoforms and PTMs are much insufficient in the width and depth in PAs, much more efforts should be made to insight into this area in a PA proteome because protein variants/isoforms and PTMs play very essential roles in tumor pathogenesis and could be potential biomarkers and therapeutic targets (44).
(iv) Use of proteomic variations for personalized and precise medicine practice of PAs. We addressed in detail how to use proteomic variations for prediction, prevention, and personalized medicine in NFPAs (73) and use the integrated omics data for personalized and precise treatment for PAs with view point of multi-parameter systematic strategy (74, 75). Although there is a long distance to reach the extreme goal of personalized and precise medicine of PAs, proteomic variations in combination with integrated omic variations and the development of novel technologies offer the great promise to achieve the goal (73–75).
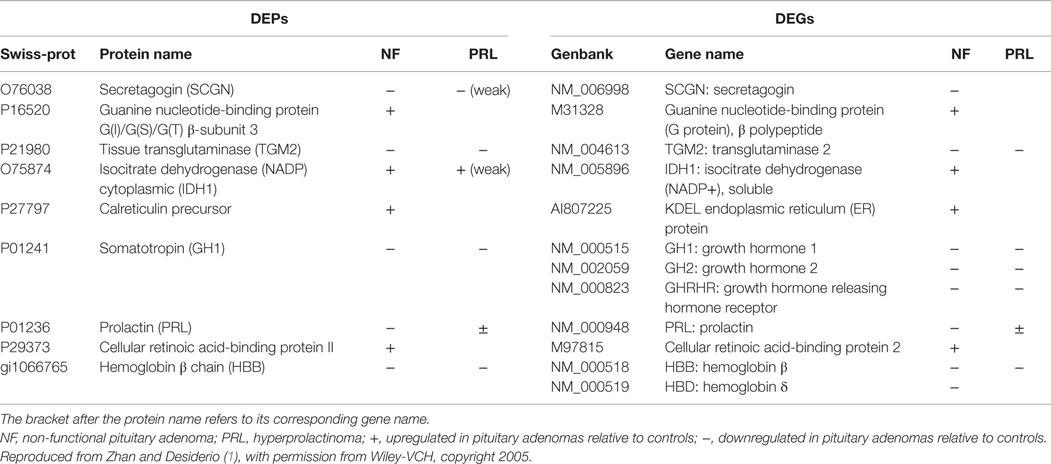
Table 2. Genes that altered consistently in the levels of protein and mRNA with comparison of proteomics data and transcriptomic data.
Future Perspectives in PA Proteomics
Although a great progress has been achieved in human PA proteomics, a series of unsolved issues are still existing in the following aspects. (i) High-throughput quantitative proteomics based on LCM-purified PA and control cells. LCM can effectively purify the PA cells and the corresponding pituitary cells for protein mapping analysis (8, 9) and minimize the inconsistency in cell types between PA tissues and postmortem pituitary control tissues. However, high-throughput quantitative proteomics strategies, such as isobaric tags for relative and absolute quantification (iTRAQ) (77, 78), label-free strategies (79, 80), or sequential window acquisition of all theoretical spectra (SWATH) (81, 82), have not been used to determine DEPs between LCM-purified PA cells and the corresponding control cells (somatotroph, corticotroph, lactrotroph, gonadotroph, and tyrotroph) for more accurate understanding of molecular mechanisms of different cell types of PAs and discovery of reliable biomarkers. (ii) PTM-quantitative proteomics in PAs. PTMs are very important aspects in a proteome (44). However, PTM proteomics studies in PAs are much insufficient in depth and width. Currently, only 28 phosphoproteins from pituitary control tissues (17, 18), 8 nitroproteins from a pituitary control tissue (20, 53), and 9 nitroproteins from an NFPA tissue (21) have been identified. PTM-quantitative proteomics (such as iTRAQ- and label-free strategies) has not been performed to discover PA-related PTMs, such as phosphorylation (83, 84), nitration (19, 52), acetylation (85, 86), glycosylation (87, 88), methylation (89), ubiquitylation (90), sumoylation (91), and succinylation (92); however, it is a very promising area in the field of PA proteomics. (iii) Structural proteomics of PAs. Three-dimensional (3D) structure of a protein determines its biological significance in a PA biological system (52). If 3D structure of a key PA protein can be reconstructed with X-ray crystallography data, then it would easily interpret its biological roles and is possible to design a small-molecule drug toward that 3D structure for the development of effective therapy strategy for PAs. (iv) Body fluid proteomics/peptidomics of PAs. PA is a whole-body disease with multiple factors, multiple processes, and multiple consequences (74). Body fluid, such as CSF and serum, is the window for prediction, diagnosis, and prognostic assessment of a PA (73). Body fluid proteomics/peptidomics is barely performed and is very promising research direction for PAs, especially NFPAs. (v) Omics data-based systems biology and molecular networks. PA is a complicated whole-body disease with alterations in multiple molecules in the levels of genome, transcriptome, proteome, and metabolome; and these molecules are mutually associated within a molecular network system (75). Multiple omics data-based systems biology and molecular network analyses might discover key molecular networks that are common or specific to different types of PAs and benefit discovery of effective and reliable biomarkers. (vi) Invasive mechanisms and biomarkers of FPAs and NFPAs. Proteomics studies regarding invasive characteristics of PAs analyzed only invasive and non-invasive NFPAs with 2DGE-quantiative proteomics (6, 11). Because of 2DGE disadvantages, further non-gel high-throughput quantitative proteomics is needed (26). Moreover, this type of study should expand to invasive and non-invase FPAs. The clarification of invasive mechanisms and determination of biomarkers might benefit management of invasive PAs. (vii) Heterogeneity of NFPAs. NFPA is highly heterogeneous. Current proteomics study focused only on NF−-, LH+-, FSH+-, and LH/FSH+-NFPA subtypes with 2DGE-quantitative proteomics (12). It should expand to other NFPA subtypes, such as silent corticotroph adenoma and silent somatotroph adenoma. Moreover, current proteomics studies of PAs mainly focused on NFPAs (1, 3, 6, 12), but barely analyzed FPAs (10). Therefore, in future, more efforts toward research on FPAs with proteomics strategies are required. The clarification of proteomic variations in different types of NFPAs and FPAs would benefit complete understanding of molecular mechanisms and discovery of biomarkers that are common and specific to different types of PAs and really realize personalized and precise medicine practice of PAs.
Conclusion
The use of proteomics to clarify molecular mechanisms of PAs and discover reliable biomarkers for prediction, prevention, and personalized treatment for PAs is our long-term goal. A great progress has been achieved in the field of PA proteomics in the past 10 years, including sample preparation with LCM and 2D-LC–MS/MS (8, 9), significant extension of proteomic reference mapping data in PAs and controls (8, 9, 29–31), proteomics analyses of invasive characteristics (6, 11), tumor heterogeneity (12), protein variants/isoforms (3, 10, 15, 16) and PTMs (17–21), and serum proteomics and peptidomics (13, 14) of PAs, protein antibody array analysis of PAs (54), and proteomic data-based pathway networks in NFPAs (22). These new progresses of pituitary proteomics are encouraging and provide new promise for insight into pathophysiological system of PAs. However, there are many aspects that have not been resolved and are worth studying in-depth including high-throughput quantitative proteomics based on LCM-enriched pituitary cells, PTM-quantitative proteomics, structural proteomics, body fluid proteomics and peptidomics, omics data-based integrative molecular networks, and proteomic characteristics of invasiveness and heterogeneity of PAs. Furthermore, one must realize that, although clinical relevance of PA proteomics data was discussed in this article, the real functional analysis regarding most of these interesting proteins from PA proteomics studies has not been carried out in the PA biological systems, which is very important aspect in the post-proteomics and must be strengthened in future. Resolving all these aspects will benefit systematical and complete understanding of PA pathogenesis and management of PAs toward personalized and precise medicine practice.
Author Contributions
XZ conceived of the concept and idea of this article, collected reference, wrote and revised the entire manuscript, was responsible for the corresponding work of the manuscript. XW collected references and drafted partial manuscript. TC participated in revision and edition of the manuscript. All authors read and approved the final manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors acknowledge the financial supports from China “863” Plan Project (Grant No. 2014AA020610-1 to XZ), the Xiangya Hospital Funds for Talent Introduction (to XZ), the National Natural Science Foundation of China (Grant No. 81272798 and 81572278 to XZ), and the Hunan Provincial Natural Science Foundation of China (Grant No. 14JJ7008 to XZ). The scientific contributions of Dominic M. Desiderio are acknowledged.
Abbreviations
2D, two-dimension; 2DGE, two-dimensional gel electrophoresis; ACTH, adrenocorticotropic hormone; CSF, cerebrospinal fluid; DEG, differentially expressed gene; DEP, differentially expressed protein; FPA, functional pituitary adenoma; FSH, follicle-stimulating hormone; GH, growth hormone; IEF, isoelectric focusing; IL1, interleukin 1; IMAC, immobilized metal ion affinity chromatography; IPA, ingenuity pathway analysis; LC, liquid chromatography; LCM, laser-capture microdissection; LH, luteinizing hormone; MALDI, matrix-assisted laser desorption ionization; MDLC, multi-dimensional liquid chromatography; MGT, multiple gel-based technologies; MS/MS, tandem mass spectrometry; MS, mass spectrometry; NFPA, non-functional pituitary adenoma; NTAC, nitrotyrosine affinity column; PA, pituitary adenoma; PKA, cAMP-dependent protein kinase; PPPM, predictive, preventive and personalized medicine; PRL, prolactin; PTM, post-translational modification; SDS-PAGE, sodium dodecyl sulfate-polyacrylamide gel electrophoresis; SELDI, surface-enhanced laser desorption/ionization; TOF, time-of-flight; TSH, thyrotropin-stimulating hormone or thyrotropin.
References
1. Zhan X, Desiderio DM. Comparative proteomics analysis of human pituitary adenomas: current status and future perspectives. Mass Spectrom Rev (2005) 24(6):783–813. doi:10.1002/mas.20039
2. Melmed S. Mechanisms for pituitary tumorigenesis: the plastic pituitary. J Clin Invest (2003) 112(11):1603–18. doi:10.1172/JCI20401
3. Moreno CS, Evans CO, Zhan X, Okor M, Desiderio DM, Oyesiku NM. Novel molecular signaling and classification of human clinically nonfunctional pituitary adenomas identified by gene expression profiling and proteomic analyses. Cancer Res (2005) 65(22):10214–22. doi:10.1158/0008-5472.CAN-05-0884
4. Asa SL, Ezzat S. The pathogenesis of pituitary tumours. Nat Rev Cancer (2002) 2:836–49. doi:10.1038/nrc926
5. Asa SL, Ezzat S. Genetics and proteomics of pituitary tumors. Endocrine (2005) 28:43–7. doi:10.1385/ENDO:28:1:043
6. Zhan X, Desiderio DM, Wang X, Zhan X, Guo T, Li M, et al. Identification of the proteomic variations of invasive relative to non-invasive non-functional pituitary adenomas. Electrophoresis (2014) 35(15):2184–94. doi:10.1002/elps.201300590
7. Melmed S. Pathogenesis of pituitary tumors. Nat Rev Endocrinol (2011) 7(5):257–66. doi:10.1038/nrendo.2011.40
8. Liu Y, Wu J, Yan G, Hou R, Zhuang D, Chen L, et al. Proteomic analysis of prolactinoma cells by immuno-laser capture microdissection combined with online two-dimensional nano-scale liquid chromatography/mass spectrometry. Proteome Sci (2010) 8:2. doi:10.1186/1477-5956-8-2
9. Liu Y, Zhuang D, Hou R, Li J, Xu G, Song T, et al. Shotgun proteomic analysis of microdissected postmortem human pituitary using complementary two-dimensional liquid chromatography coupled with tandem mass spectrometer. Anal Chim Acta (2011) 688(2):183–90. doi:10.1016/j.aca.2010.12.032
10. Evans CO, Moreno CS, Zhan X, McCabe MT, Vertino PM, Desiderio DM, et al. Molecular pathogenesis of human prolactinomas identified by gene expression profiling, RT-qPCR, and proteomic analyses. Pituitary (2008) 11(3):231–45. doi:10.1007/s11102-007-0082-2
11. Liu Z, Liu Y, Fang W, Chen W, Li C, Xiao Z. Establishment of differential expression profiles from invasive and non-invasive pituitary adenomas. Zhong Nan Da Xue Xue Bao Yi Xue Ban (2009) 34:569–75. doi:1672-7437(2009)07-0569-07
12. Zhan X, Wang X, Long Y, Desiderio DM. Heterogeneity analysis of the proteomes in clinically nonfunctional pituitary adenomas. BMC Med Genomics (2014) 7:69. doi:10.1186/s12920-014-0069-6
13. Hu X, Zhang P, Shang A, Li Q, Xia Y, Jia G, et al. A primary proteomic analysis of serum from patients with nonfunctioning pituitary adenoma. J Int Med Res (2012) 40(1):95–104. doi:10.1177/147323001204000110
14. Zhou KY, Jin HH, Bai ZQ, Liu CB. Pituitary adenoma biomarkers identified using proteomic fingerprint technology. Asian Pac J Cancer Prev (2012) 13(8):4093–5. doi:10.7314/APJCP.2012.13.8.4093
15. Zhan X, Giorgianni F, Desiderio DM. Proteomics analysis of growth hormone isoforms in the human pituitary. Proteomics (2005) 5(5):1228–41. doi:10.1002/pmic.200400987
16. Kohler M, Thomas A, Püschel K, Schänzer W, Thevis M. Identification of human pituitary growth hormone variants by mass spectrometry. J Proteome Res (2009) 8(2):1071–6. doi:10.1021/pr800945b
17. Beranova-Giorgianni S, Zhao Y, Desiderio DM, Giorgianni F. Phosphoproteomic analysis of the human pituitary. Pituitary (2006) 9(2):109–20. doi:10.1007/s11102-006-8916-x
18. Giorgianni F, Beranova-Giorgianni S, Desiderio DM. Identification and characterization of phosphorylated proteins in the human pituitary. Proteomics (2004) 4(3):587–98. doi:10.1002/pmic.200300584
19. Zhan X, Wang X, Desiderio DM. Pituitary adenoma nitroproteomics: current status and perspectives. Oxid Med Cell Longev (2013) 2013:580710. doi:10.1155/2013/580710
20. Zhan X, Desiderio DM. The human pituitary nitroproteome: detection of nitrotyrosyl-proteins with two-dimensional Western blotting, and amino acid sequence determination with mass spectrometry. Biochem Biophys Res Commun (2004) 325(4):1180–6. doi:10.1016/j.bbrc.2004.10.169
21. Zhan X, Desiderio DM. Nitroproteins from a human pituitary adenoma tissue discovered with a nitrotyrosine affinity column and tandem mass spectrometry. Anal Biochem (2006) 354(2):279–89. doi:10.1016/j.ab.2006.05.024
22. Zhan X, Desiderio DM. Signaling pathway networks mined from human pituitary adenoma proteomics data. BMC Med Genomics (2010) 3:13. doi:10.1186/1755-8794-3-13
23. Zhan X, Desiderio DM. Heterogeneity analysis of the human pituitary proteome. Clin Chem (2003) 49:1740–51. doi:10.1373/49.10.1740
24. Emmert-Buck MR, Bonner RF, Smith PD, Chuaqui RF, Zhuang Z, Goldstein SR, et al. Laser capture microdissection. Science (1996) 274:998–1001. doi:10.1126/science.274.5289.998
25. Zhou G, Li H, DeCamp D, Chen S, Shu H, Gong Y, et al. 2D differential in-gel electrophoresis for the identification of esophageal scans cell cancer-specific protein markers. Mol Cell Proteomics (2002) 1:117–24. doi:10.1074/mcp.M100015-MCP200
26. Zhan X. Current status of two-dimensional gel electrophoresis and multi-dimensional liquid chromatography as proteomic separation techniques. Ann Chromatogr Sep Tech (2015) 1(2):1009.
27. Jin L, Tsumanuma I, Ruebel KH, Bayliss JM, Lloyd RV. Analysis of homogeneous populations of anterior pituitary folliculostellate cells by laser capture microdissection and reverse transcription-polymerase chain reaction. Endocrinology (2001) 142(5):1703–9. doi:10.1210/en.142.5.1703
28. Liu Y, Wu J, Liu S, Zhuang D, Wang Y, Shou X, et al. Immuno-laser capture microdissection of frozen prolactioma sections to prepare proteomic samples. Colloids Surf B Biointerfaces (2009) 71(2):187–93. doi:10.1016/j.colsurfb.2009.02.005
29. Zhao Y, Giorgianni F, Desiderio DM, Fang B, Beranova-Giorgianni S. Toward a global analysis of the human pituitary proteome by multiple gel-based technology. Anal Chem (2005) 77(16):5324–31. doi:10.1021/ac050354e
30. Zhan X, Desiderio DM. A reference map of a pituitary adenoma proteome. Proteomics (2003) 3:699–713. doi:10.1002/pmic.200300408
31. Wang X, Guo T, Peng F, Long Y, Mu Y, Yang H, et al. Proteomic and functional profiles of a follicle-stimulating hormone-positive human nonfunctional pituitary adenoma. Electrophoresis (2015) 36(11–12):1289–304. doi:10.1002/elps.201500006
32. Hussaini IM, Trotter C, Zhao Y, Abdel-Fattah R, Amos S, Xiao A, et al. Matrix metalloproteinase-9 is differentially expressed in nonfunctioning invasive and noninvasive pituitary adenomas and increases invasion in human pituitary adenoma cell line. Am J Pathol (2007) 170(1):356–65. doi:10.2353/ajpath.2007.060736
33. Meij BP, Lopes MB, Ellegala DB, Alden TD, Laws ER Jr. The long-term significance of microscopic dural invasion in 354 patients with pituitary adenomas treated with transsphenoidal surgery. J Neurosurg (2002) 96(2):195–208. doi:10.3171/jns.2002.96.2.0195
34. Selman WR, Laws ER Jr, Scheithauer BW, Carpenter SM. The occurrence of dural invasion in pituitary adenomas. J Neurosurg (1986) 64(3):402–7. doi:10.3171/jns.1986.64.3.0402
35. Zhang X, Fei Z, Zhang W, Zhang JN, Liu WP, Fu LA, et al. Endoscopic endonasal transsphenoidal surgery for invasive pituitary adenoma. J Clin Neurosci (2008) 15(3):241–5. doi:10.1016/j.jocn.2007.03.008
36. Galland F, Lacroix L, Saulnier P, Dessen P, Meduri G, Bernier M, et al. Differential gene expression profiles of invasive and non-invasive non-functioning pituitary adenomas based on microarray analysis. Endocr Relat Cancer (2010) 17(2):361–71. doi:10.1677/ERC-10-0018
37. Zhan X. Proteomic heterogeneity of nonfunctional pituitary adenomas. Precis Med (2015) 2:e663. doi:10.14800/pm.663
38. Zhan X. Hormone-related proteomic and functional variations in human nonfunctional pituitary adenomas. Inflamm Cell Signal (2015) 2:e841. doi:10.14800/ics.841
39. Walsh MT, Couldwell WT. Symptomatic cystic degeneration of a clinically silent corticotroph tumor of the pituitary gland. Skull Base (2010) 20(5):367–70. doi:10.1055/s-0030-1253579
40. Zhou W, Song Y, Xu H, Zhou K, Zhang W, Chen J, et al. In nonfunctional pituitary adenomas, estrogen receptors and slug contribute to development of invasiveness. J Clin Endocrinol Metab (2011) 96(8):E1237–45. doi:10.1210/jc.2010-3040
41. Blencowe BJ. Alternative splicing: new insights from global analyses. Cell (2006) 126:37–47. doi:10.1016/j.cell.2006.06.023
42. Perrin BJ, Ervasti JM. The actin gene family: function follows isoform. Cytoskeleton (Hoboken) (2010) 67:630–4. doi:10.1002/cm.20475
43. Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu Rev Biochem (2003) 72:291–336. doi:10.1146/annurev.biochem.72.121801.161720
44. Zhan X. Insight into protein variants/isoforms and post-translational modifications in a proteome. Austin Proteomics (2015) 2(1):1009.
45. Robson H, Siebler T, Shalet SM, Williams GR. Interactions between GH, IGF-I, glucocorticoids, and thyroid hormones during skeletal growth. Pediatr Res (2002) 52(2):137–47. doi:10.1203/00006450-200208000-00003
46. Yoshizato H, Tanaka M, Nakai N, Nakao N, Nakashima K. Growth hormone (GH)-stimulated insulin-like growth factor I gene expression is mediated by a tyrosine phosphorylation pathway depending on C-terminal region of human GH receptor in human GH receptor-expressing Ba/F3 cells. Endocrinology (2004) 145(1):214–20. doi:10.1210/en.2003-0811
47. Boguszewski CL. Molecular heterogeneity of human GH: from basic research to clinical implications. J Endocrinol Invest (2003) 26(3):274–88. doi:10.1007/BF03345170
48. Boguszewski CL, Jansson C, Boguszewski MC, Rosberg S, Carlsson B, Albertsson-Wikland K, et al. Increased proportion of circulating non-22-kilodalton growth hormone isoforms in short children: a possible mechanism for growth failure. J Clin Endocrinol Metab (1997) 82(9):2944–9. doi:10.1210/jcem.82.9.4226
49. Boguszewski CL, Johannsson G, Bengtsson BA, Johansson A, Carlsson B, Carlsson LM. Circulating non-22-kilodalton growth hormone isoforms in acromegalic men before and after transsphenoidal surgery. J Clin Endocrinol Metab (1997) 82(5):1516–21. doi:10.1210/jcem.82.5.3911
50. Noble ME, Endicott JA, Johnson LN. Protein kinase inhibitors: insights into drug design from structure. Science (2004) 303(5665):1800–5. doi:10.1126/science.1095920
51. Khan J, Brennand DM, Bradley N, Gao B, Bruckdorfer R, Jacobs M. 3-Nitrotyrosine in the proteins of human plasma determined by an ELISA method. Biochem J (1998) 332(Pt 3):807–8. doi:10.1042/bj3320807v
52. Zhan X, Wang X, Desiderio DM. Mass spectrometry analysis of nitrotyrosine-containing proteins. Mass Spectrom Rev (2015) 34(4):423–48. doi:10.1002/mas.21413
53. Zhan X, Desiderio DM. Linear ion-trap mass spectrometric characterization of human pituitary nitrotyrosine-containing proteins. Int J Mass Spectrom (2007) 259:96–104. doi:10.1016/j.ijms.2006.06.009
54. Ribeiro-Oliveira AJ, Franchi G, Kola B, Dalino P, Pinheiro SV, Salahuddin N, et al. Protein western array analysis in human pituitary tumours: insights and limitations. Endocr Relat Cancer (2008) 15(4):1099–114. doi:10.1677/ERC-08-0003
55. Zhan X, Sacks H, Desiderio DM. The human pituitary proteome: clinical applications. In: Vekey K, Telekes A, Vertes A, editors. Medical Applications of Mass Spectrometry. Amsterdam: Elsevier Science Publisher (2007). p. 425–58.
56. Evans CO, Young AN, Brown MR, Brat DJ, Parks JS, Neish AS, et al. Novel patterns of gene expression in pituitary adenomas identified by complementary deoxyribonucleic acid microarrays and quantitative reverse transcription-polymerase chain reaction. J Clin Endocrinol Metab (2001) 86:3097–107. doi:10.1210/jcem.86.7.7616
57. Lania A, Montovani G, Spada A. Genetics of pituitary tumors: focus on G-protein mutations. Exp Biol Med (2003) 228:1004–7.
58. Alexander J. Tumor suppressor loss in pituitary tumors. Brain Pathol (2001) 11:342–55. doi:10.1111/j.1750-3639.2001.tb00404.x
59. Yu R, Melmed S. Oncogene activation in pituitary tumors. Brain Pathol (2001) 11:328–41. doi:10.1111/j.1750-3639.2001.tb00403.x
60. Zhan X, Evans CO, Oyesiku NM, Desiderio DM. Proteomics and transcriptomics analyses of secretagogin down-regulation in human non-functional pituitary adenoma. Pituitary (2003) 6:189–202. doi:10.1023/B:PITU.0000023426.99808.40
61. Wagner L, Oliyarnyk O, Gartner W, Nowotny P, Groeger M, Kaserer K, et al. Cloning and expression of secretagogin, a novel neuroendocrine- and pancreatic islet of Langerhans-specific Ca2+-binding protein. J Biol Chem (2000) 275:24740–51. doi:10.1074/jbc.M001974200
62. Gartner W, Lang W, Leutmetzer F, Domanovits H, Waldhausl W, Wagner L. Cerebral expression and serum detectability of secretagogin, a recently cloned EF-hand Ca2+-binding protein. Cereb Cortex (2001) 11:1161–9. doi:10.1093/cercor/11.12.1161
63. Desiderio DM. Mass spectrometric analysis of neuropeptidergic systems in the human pituitary and cerebrospinal fluid. J Chromatogr B Biomed Sci Appl (1999) 731:3–22. doi:10.1016/S0378-4347(99)00172-3
64. Zhu X, Robertson JT, Sacks HS, Dohan C Jr, Tseng JL, Desiderio DM. Opioid and tachykinin neuropeptides in prolactin-secreting human pituitary adenomas. Peptides (1995) 16:1097–107. doi:10.1016/0196-9781(95)00081-T
65. Zhu X, Desiderio DM. Effects of space flight stress on proopiomelanocortin, proenkephalin A, and tachykinin neuropeptidergic systems in the rat posterior pituitary. Life Sci (1994) 55:347–50. doi:10.1016/0024-3205(94)00644-X
66. Zhu X, Desiderio DM. Methionine enkephalin-like immunoreactivity, substance P-like immunoreactivity and beta-endorphin-like immunoreactivity postmortem stability in rat pituitary. J Chromatogr (1993) 616:175–87. doi:10.1016/0378-4347(93)80384-G
67. Desiderio DM, Kusmierz JJ, Zhu X, Dass C, Hilton D, Robertson JT, et al. Mass spectrometric analysis of opioid and tachykinin neuropeptides in nonsecreting and ACTH-secreting adenomas. Biol Mass Spectrom (1993) 22:89–97. doi:10.1002/bms.1200220112
68. Herman V, Fagin J, Gonsky R, Kovacs K, Melmed S. Clonal origin of pituitary adenomas. J Clin Endocrinol Metab (1990) 71:1427–33. doi:10.1210/jcem-71-6-1427
69. Alexander JM, Biller BM, Bikkal H, Zervas NT, Amold A, Klibanski A. Clinically nonfunctioning pituitary tumors are monoclonal in origin. J Clin Invest (1990) 86:336–40. doi:10.1172/JCI114705
70. Gambino GM, Beck-Peccoz P, Borgato S, Faglia G, Spada A, Persani L. Bioactivity and glycosylation of circulating prolactin in various physiological and pathological conditions. Pituitary (1999) 2:225–31. doi:10.1023/A:1009909513790
71. Hoffmann T, Penel C, Ronin C. Glycosylation of human prolactin regulates hormone bioactivity and metabolic clearance. J Endocrinol Invest (1993) 16:807–16. doi:10.1007/BF03348932
72. Hoffmann T, Gunz G, Brue T, Jaquet P, Ronin C. Prolactin isoforms secreted by human prolactinomas. Horm Res (1992) 38:164–70. doi:10.1159/000182534
73. Zhan X, Desiderio DM. The use of variations in proteomes to predict, prevent, personalize treatment for clinically non-functional pituitary adenomas. EPMA J (2010) 1:439–59. doi:10.1007/s13167-010-0028-z
74. Hu R, Wang X, Zhan X. Multi-parameter systematic strategy for predictive, preventive, and personalized medicine in cancer. EPMA J (2013) 4:2. doi:10.1186/1878-5085-4-2
75. Grech G, Zhan X, Yoo BC, Bubnov R, Hagan S, Danesi R, et al. EPMA position paper in cancer: current overview and future perspectives. EPMA J (2015) 6:9. doi:10.1186/s13167-015-0030-6
76. Ezzat S, Zheng L, Zhu XF, Wu GE, Asa SL. Targeted expression of a human pituitary tumor-derived isoform of FGF receptor-4 recapitulates pituitary tumorigenesis. J Clin Invest (2002) 109:69–78. doi:10.1172/JCI14036
77. Evans C, Noirel J, Ow SY, Salim M, Pereira-Medrano AG, Couto N, et al. An insight into iTRAQ: where do we stand now? Anal Bioanal Chem (2012) 404:1011–27. doi:10.1007/s00216-012-5918-6
78. Aggarwal K, Choe LH, Lee KH. Shotgun proteomics using the iTRAQ isobaric tags. Brief Funct Genomic Proteomic (2006) 5:112–20. doi:10.1093/bfgp/ell018
79. Guo H, Isserlin R, Lugowski A, Kuzmanov U, Emili A. Large-scale label-free phosphoproteomics: from technology to data interpretation. Bioanalysis (2014) 6:2403–20. doi:10.4155/bio.14.188
80. Matzke MM, Brown JN, Gritsenko MA, Metz TO, Pounds JG, Rodland KD, et al. A comparative analysis of computational approaches to relative protein quantification using peptide peak intensities in label-free LC-MS proteomics experiments. Proteomics (2013) 13:493–503. doi:10.1002/pmic.201200269
81. Gillet LC, Navarro P, Tate S, Röst H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics (2012) 11:O111. doi:10.1074/mcp.O111.016717
82. Lambert JP, Ivosev G, Couzens AL, Larsen B, Taipale M, Lin ZY, et al. Mapping differential interactomes by affinity purification coupled with data-independent mass spectrometry acquisition. Nat Methods (2013) 10:1239–45. doi:10.1038/nmeth.2702
83. von Stechow L, Francavilla C, Olsen JV. Recent findings and technological advances in phosphoproteomics for cells and tissues. Expert Rev Proteomics (2015) 12:469–87. doi:10.1586/14789450.2015.1078730
84. Ruprecht B, Lemeer S. Proteomic analysis of phosphorylation in cancer. Expert Rev Proteomics (2014) 11:259–67. doi:10.1586/14789450.2014.901156
85. Svinkina T, Gu H, Silva JC, Mertins P, Qiao J, Fereshetian S, et al. Deep, quantitative coverage of the lysine acetylome using novel anti-acetyl-lysine antibodies and an optimized proteomic workflow. Mol Cell Proteomics (2015) 14:2429–40. doi:10.1074/mcp.O114.047555
86. Rardin MJ, Newman JC, Held JM, Cusack MP, Sorensen DJ, Li B, et al. Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways. Proc Natl Acad Sci U S A (2013) 110:6601–6. doi:10.1073/pnas.1302961110
87. Christiansen MN, Chik J, Lee L, Anugraham M, Abrahams JL, Packer NH. Cell surface protein glycosylation in cancer. Proteomics (2014) 14:525–46. doi:10.1002/pmic.201300387
88. Pan S, Chen R, Aebersold R, Brentnall TA. Mass spectrometry based glycoproteomics – from a proteomics perspective. Mol Cell Proteomics (2011) 10:110. doi:10.1074/mcp.R110.003251
89. Carlson SM, Gozani O. Emerging technologies to map the protein methylome. J Mol Biol (2014) 426:3350–62. doi:10.1016/j.jmb.2014.04.024
90. Porras-Yakushi TR, Hess S. Recent advances in defining the ubiquitylome. Expert Rev Proteomics (2014) 11:477–90. doi:10.1586/14789450.2014.926223
91. Galisson F, Mahrouche L, Courcelles M, Bonneil E, Meloche S, Chelbi-Alix MK, et al. A novel proteomics approach to identify SUMOylated proteins and their modification sites in human cells. Mol Cell Proteomics (2011) 10:M110. doi:10.1074/mcp.M110.004796
Keywords: pituitary adenoma, proteome, heterogeneity, reference map, two-dimensional gel electrophoresis, mass spectrometry, bioinformatics
Citation: Zhan X, Wang X and Cheng T (2016) Human Pituitary Adenoma Proteomics: New Progresses and Perspectives. Front. Endocrinol. 7:54. doi: 10.3389/fendo.2016.00054
Received: 30 March 2016; Accepted: 17 May 2016;
Published: 31 May 2016
Edited by:
Carmen E. Georgescu, Iuliu Haţieganu University of Medicine and Pharmacy Cluj-Napoc, RomaniaReviewed by:
Laurence Katznelson, Stanford University, USAHidenori Fukuoka, Kobe University Hospital, Japan
Copyright: © 2016 Zhan, Wang and Cheng. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xianquan Zhan, eWp6aGFuMjAxMUBnbWFpbC5jb20=
 Xianquan Zhan
Xianquan Zhan Xiaowei Wang1,2,3
Xiaowei Wang1,2,3